Abstract
Antimicrobial resistance (AMR) databases enable the identification of AMR determinants from pathogen sequence data and the prediction of resistance profiles, enhancing AMR surveillance and informing a range of public health interventions. This review compares freely available and regularly updated AMR databases, explores their public health value and highlights key challenges to and opportunities for fully harnessing their potential.
Similar content being viewed by others
Introduction
Bacterial antimicrobial resistance (AMR) is one of the most significant challenges facing human health and requires urgent action1 Next generation sequencing technologies, including whole-genome sequencing (WGS), are increasingly used to monitor AMR and inform public health interventions, such as surveillance2,3, outbreak management4, infection prevention and control5, programmatic care6, and diagnostics development7. Genomic surveillance of AMR and knowledge of AMR determinants were identified as global strategic and research priorities8,9,10,including and rapid accurate methods to detect them11.
Resistance may be intrinsic (i.e. naturally found in a species) or acquired (e.g., though mutation or horizontal gene transfer). AMR determinants are genes and genetic modifications that confer resistance to one or more antibiotics and antibiotic classes. AMR genes encode proteins that modify the antibiotic (e.g., β-lactamases), modify and protect the antibiotic target site (e.g., ribosomal RNA methyltransferases), provide an additional target for which the antibiotic has reduced affinity (e.g., altered penicillin binding proteins), provide alternative metabolic pathways bypassing the antibiotic’s action (e.g., dihydrofolate reductases) or that extrude the antibiotic outside the cell (e.g., efflux pumps)12,13. Acquired AMR genes are often gained en bloc by horizontal gene transfer of mobile genetic elements that can move between DNA molecules, such as insertion sequences and transposons, or between bacterial cells, such as plasmids and integrative conjugative elements14.
Genetic modifications associated with resistance may change a single nucleotide (point mutations) or involve larger rearrangements (insertions or deletions) affecting either coding or regulatory regions. They may lead to alteration of the antibiotic target site (e.g., point mutations in rpoB leading to rifampicin resistance in M. tuberculosis), increased expression of an antibiotic target (e.g., point mutations in the promoter of folA encoding dihydrofolate reductase, the target of trimethoprim), or decreased antibiotic uptake (e.g., a small insertion in the ompK36 porin gene, or an in-frame deletion in the oprD porin gene leading to decreased antibiotic permeability)12,13. Replacement of natural gene promoters with stronger promoters from insertion sequences via horizontal gene transfer also leads to resistance. For example, insertion of ISAba1 upstream of naturally-occuring blaampC in Acinetobacter baumannii leads to cephalosporinase overproduction and cephalosporin resistance15.
Databases of AMR determinants are curated repositories that store reference nucleotide and protein sequences of genes known to confer resistance, as well as wild-type genes or sequences that can confer resistance when modified. They also include associated information needed to identify these sequences in pathogen genomes, such as rules, computational models and classification hierarchies. As such, they need to capture the diversity of genes and genetic modifications associated with resistance. However, currently available databases vary widely in size, scope, structure, content, annotation, curation approach and update frequency12,16,17,18,19. Some are species-specific (Table 1), while others span multiple or all WHO bacterial priority pathogens20. Species-agnostic databases often operate under the assumption that determinants have the same effect across pathogens, which may not be the case. Databases also differ in the range of antibiotics and antibiotic classes they cover. They may comprise genes, mutations, or both, but their inclusion and exclusion criteria may also differ.
Bioinformatic tools compare pathogen sequences to those in AMR databases, detect and identify AMR determinants, and report them with additional information such as the associated antibiotic, the position of the determinant in the genome and sequence similarity (the percentage match between the pathogen and reference sequence). This information can then be used to predict phenotypic resistance. Bioinformatic tools employ a variety of computational approaches to detect AMR determinants in pathogen sequences, differing in reporting format and content12,21,22,23,24,25,26,27,28. Both database and associated tools may be accessed through command-line interfaces for integration into user bioinformatic workflows, or through web-based tools. Web-based interfaces are generally more user-friendly and accessible to those without prior bioinformatic expertise. They streamline analyses by automating workflows, use off-site computing resources, and produce reports with quick turnaround times. Additionally, several high-quality external bioinformatic tools are available that leverage one or multiple of these databases12,17,18,24,25,29,30.
Methods
To identify AMR databases, we searched National Centre for Biotechnology Information (NCBI) PubMed using the term: (antimicrobial resistan* OR antibiotic resistan*) AND (mutation OR gene OR variant OR determinant) AND (catalogue OR database) for articles published between 1-January-2017 and 31-December-2023. We identified 1300 articles, of which 22 pertained directly to 12 AMR databases. Additional checks of reference lists and web searches pointed to an additional 12 databases. Of these 24 databases, 8 met these inclusion criteria: relevant for use with whole genome sequences (as opposed to metagenomes); free; regularly updated (defined as ≥1 update between 1-January-2024 and 1-January-2025); relevant to ≥1 WHO bacterial priority pathogen20; and spanning >1 AMR mechanism and/or >1 antibiotic and/or disinfection classes (Table 1).
Review of current databases
Overview
Other reviews have previously documented AMR databases12,17,18, either in the broader context of genomic methods and resources to study and monitor AMR or focussing on their gene content. Here, we compare and contrast AMR databases (Table 1) identified through literature and web searches and meeting pre-specified criteria, while providing a structured critical analysis framed from public health and policy perspectives.
Comprehensive antibiotic resistance database (CARD)
CARD includes a curated sequence database built on a structure known as the antibiotic resistance ontology (ARO) and a set of AMR detection models. The sequence database comprises AMR genes and mutations (not all clinically relevant), and biocide resistance genes.
The ARO refers to controlled vocabularies, categories and formal definitions indicating the relationships between database elements. Each AMR determinant sequence is classified by an ontological path, comprising categories by gene family, resistance mechanism, drug class and affected antibiotic26,31. In particular, the classification of β-lactam antibiotics also includes functional groups: monobactams, penicillins, cephalosporins (1st-5th generations), carbapenems and revised β-lactamase inhibitor combinations. The ARO is a key advantage of CARD; it improves data management, standardization, interoperability and quality of data, and contributes to consistent interpretation and genotype-phenotype associations.
CARD uses a variety of detection models to identify AMR determinants from sequences, based on manually curated similarity cut-offs and information about molecular interactions26,31. For an AMR determinant to be included in CARD, it must be described in a peer-reviewed scientific publication and have its DNA sequence available in GenBank, along with clear experimental evidence of elevated MIC over controls (except β-lactamases)26,31. The database is manually curated and updated, aided by machine learning methods (CARD-Shark)32 and curation quality control algorithms. Curation is done through cross-database harmonization (Fig. 1), and the contribution of external researchers, software developers and others via the AMR curation public repository33, also done by AMRFinderPlus34.
Arrows indicate the general flow of information, including both historical data inputs and ongoing exchanges between databases. ARDB Antibiotic Resistance Genes Database; ARG-ANNOT Antibiotic Resistance Gene-ANNOTation; BLDB Beta-Lactamase Database; CARD Comprehensive Antimicrobial Resistance Database; MTBc Mycobacterium tuberculosis complex. *database no longer actively maintained. Up to date as of 1st January 2025. Lahey is one of the first beta-lactamase databases and has been used to inform many subsequent ones such as BLDB and ARG-ANNOT but has migrated to NCBI.
The core CARD database is complemented by CARD-R, which stores AMR determinants not yet validated by clinical or experimental data –i.e. that were predicted in silico to confer resistance. CARD-R provides additional information on AMR determinants in public genomes, such as species and their location on chromosomes or plasmids26. CARD also features catalogues of AMR determinants specific to fungi (FungAMR) and M. tuberculosis (TB Mutations).
AMRFinderPlus
The AMRFinderPlus database has three components: the reference gene catalogue, the reference hidden Markov models catalogue (hidden Markov models are a set of probabilistic models used to identify patterns within sequences and classify them into AMR protein families using manually curated and validated cut-offs34) and the reference gene hierarchy23,34,35. The reference gene catalogue is a curated database comprising AMR genes, point mutations and their associated proteins (core catalogue), along with additional stress- and virulence-related genes and proteins (plus catalogue). The reference gene hierarchy is an ontology structure that organizes and relates hidden Markov models to AMR determinants and classifies them by antimicrobial class (e.g., β-lactam), subclass (e.g., carbapenem), similarity and function.
Each protein and protein family included in the database is assigned a name and a gene symbol, based on the internal hierarchy, cut-offs and rules34. Each gene symbol can then be associated with related protein sequences (e.g., blaKPC), and each allele with a single amino acid sequence variant (e.g., blaKPC-2)34. This standardized approach is key to enhance identification and naming of detected determinants and assigning new alleles (done by NCBI for β-lactamases36, Qnr and MCR alleles).
AMRFinderPlus gathers determinants via literature surveys, data exchange with external curated sources and other mechanisms34. Inclusion of a determinant requires experimental verification or a highly similar hit to an experimentally verified protein, but supporting evidence in the literature is not required for allele assignment34. AMRFinderPlus established different curation approaches for genes, alleles, point mutations and hierarchy levels, with most being manual. It also applies automated quality-control checks to the sequences that are deposited in the database and for resolving nomenclature conflicts34.
AMRFinderPlus is used in NCBI’s Pathogen Detection pipeline23,34,35, providing results to the National Database of Resistance Organisms (NDARO) and its associated isolate browser, antimicrobial susceptibility testing (AST) browser, and databases of microbial genetic and genomic elements (MicroBIGG-E) and geographical distribution/coverage by species (MicroBIGG-E Map)37.
ResFinder
ResFinder contains sequences of AMR genes22,38,39 and its associated database, PointFinder, contains information about chromosomal point mutations conferring resistance in selected bacterial species22,38,39,40. The primary focus has been to only include acquired AMR determinants. Additionally, DisinFinder –containing acquired disinfectant resistance genes– complements both AMR databases. ResFinder is supplemented by ResFinderFG which comprises AMR genes identified through functional cloning41. ResFinder initially drew on Antibiotic Resistance Genes Database (ARDB)42, and the Beta-Lactamase DataBase (BLDB)43 as data sources, but is currently manually curated by reviewing published literature38,39. Communication with researchers ensures that new AMR determinants meet requirements for inclusion22.
In addition to the focus on acquired resistance, another key difference with other databases is that the curation is focused on the in silico prediction of clinically relevant phenotypic antibiotic resistance profiles for selected bacterial species. For this purpose, ResFinder utilises genotype-to-phenotype tables for various antimicrobial classes and compounds, and species-specific panels for in silico antibiograms. The ResFinder software detects the presence of genetic determinants from all three databases and interprets specific combinations of multiple mutations or of genes and mutations that are needed to cause resistance in select species22,39.
Kleborate
Kleborate is a tool for the analysis of Klebsiella pneumoniae species complex genomes that performs quality control (QC), taxonomic assignment, multi-locus sequence typing (MLST), K and O locus typing, and detection of AMR determinants, virulence and serotype. The Kleborate AMR database is a species complex-specific database comprising intrinsic β-lactamase genes, horizontally acquired genes, point mutations, and gene truncations, as well as their associated antibiotic classes27,44. It was initially developed from a subset of AMR determinants found in CARD27 which were manually curated via exclusions, gene name changes, sequence replacements and additional inclusions of genetic resistance signatures from the Antibiotic Resistance Gene (ARG)-ANNOTation database45. The β-lactamases were classified by their enzyme activity (e.g., extended-spectrum β-lactamase, carbapenemase) and whether it is abolished by β-lactamase inhibitors43,45. The blaSHV alleles were further curated through the analysis of high-quality paired genotype and phenotype data and a literature systematic review of experimental evidence. This allowed the assignment of blaSHV alleles to functional classes associated with intrinsic or acquired resistance phenotypes (i.e., wild type resistance to ampicillin, extended-spectrum β-lactam resistance, or inhibitor-resistant β-lactam resistance), and the reclassification of alleles present in the NCBI reference gene catalogue and BLDB44.
AMR determinants included in the database must have associated resistance phenotypes with confirmed clinical relevance in published data. Unlike in other AMR databases, by providing a comprehensive classification and curation of AMR determinants Kleborate allows more detailed predictions of resistance in the K. pneumoniae complex. Understanding the resistance genotype is important as it determines the most appropriate antibiotic combinations to treat multidrug-resistant K. pneumoniae infections46.
Pathogenwatch
Pathogenwatch is a web platform for genomic surveillance. Through a web interface accessible to users with different bioinformatic expertise, Pathogenwatch provides QC, taxonomic assignment, MLST, serotype, AMR genes and mutations, AMR phenotype, plasmid replicon types, and trees showing genetic relatedness among genomes for select species.
Pathogenwatch AMR databases are curated for Campylobacter spp., Klebsiella spp., Neisseria gonorrhoeae, Staphylococcus aureus, Salmonella Typhi, Streptococcus pneumoniae, Vibrio cholerae and Enterococcus faecium28. Where available, Pathogenwatch uses existing, community-endorsed taxon-specific databases, such as for Klebsiella spp. (Kleborate)27,47 and E. faecium48. The remaining taxon-specific databases are developed and manually curated by compiling evidence from the literature, input from international consortia49,50, personal communications and other databases such CARD, ResFinder and AMRFinderPlus. To reduce false positive predictions, each gene or mutation is verified using available experimental resistance data and genome sequence data before incorporation into the final library. Like ResFinder, Pathogenwatch also predicts species-specific antibiotic susceptibility profiles, including resistance conferred by the presence of a combination of AMR determinants, plus inducible resistance, decreased susceptibility (intermediate resistance) and genetic elements that modify AMR determinants to restore susceptibility (e.g., deletions or disruptions by insertion elements). Like Kleborate, Pathogenwatch generates a QC report to ensure that AMR predictions are only considered for high-quality genomes.
WHO catalogue of mutations in Mycobacterium tuberculosis complex (MTBc) and their association with drug resistance
The WHO MTBc catalogue was developed through a systematic analysis of over 38,000 MTBc isolates from 45 countries. The data comprised high-quality phenotypic AST results for 13 anti-tuberculosis drugs matched with high-quality whole-genome sequences provided by several consortia and open data calls51,52. “Solo” mutations, i.e. single mutations within a set of genes of interest that best explain the observed drug resistance phenotype, were identified. The genotype-phenotype association was statistically evaluated, which enabled the calculation of odds ratios, positive predictive values, and confidence intervals to determine the certainty of the association with resistance. This enabled classification of mutations into five categories: associated with resistance, associated with resistance interim, uncertain significance, not associated with resistance, not associated with resistance interim. Interim categories reflect some uncertainty and they can be revised as new evidence becomes available.
The catalogue is released as a spreadsheet and variant call files (containing mutations, their genomic coordinates and associated antibiotics)53, with accompanying publications51,52,54. Independently developed bioinformatic tools, such as the resistance gene identifier (RGI)26, Mykrobe55 and TB Profiler56 use the catalogue to predict resistance from sequence data. The second edition of the catalogue increased the number of isolates and country representation, resulting in the addition of mutations associated with resistance to newly endorsed or repurposed drugs (e.g., bedaquiline, delamanid and linezolid)54.
Value and opportunities for public health
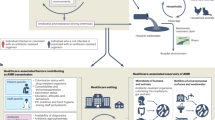
AMR databases can support public health activities directed to reducing the burden of antibiotic-resistant infections2,12,16 (Box 1). They allow detection of all known AMR determinants from sequence data, enabling more specific surveillance than phenotype alone2,16. AMR databases can capture and catalogue rare or emerging resistance variants—such as polymerase chain reaction (PCR)-positive blaKPC alleles with novel mutations that cause unusual or unexpected phenotypic resistance16,57—offering valuable insights into resistance mechanisms when conventional phenotypic and molecular tests are unavailable, unreliable or insensitive16,57. They can also facilitate the study of resistance determinant evolution and their geographical spread. Genomic surveillance in the Philippines found a nationally circulating clone carrying blaNDM-1 and blaNDM-7 on distinct plasmids, showing local and regional spread. Additionally, a hospital outbreak driven by a plasmid (IncFII) carrying the blaNDM-1 was spread across multiple genetic lineages in a neonatal intensive care unit3. AMR databases enable surveillance of the resistome through more recent approaches like untargeted or targeted metagenomic sequencing to understand how AMR determinants spread and evolve across clinical and environmental sources (e.g., wastewater surveillance)58,59. They have also supported shotgun proteomics for detecting AMR gene products60.
AMR databases play an important role in guiding the development of interventions, such as new diagnostic tests. These may include rapid tests such as PCR and loop-mediated isothermal amplification (LAMP) assays of particular utility in settings where conventional phenotypic tests or advanced testing such as panel-based PCR and genomics are unsuitable or still unattainable7,61,62,63. Notably, the WHO MTBc catalogue underpinned new WHO recommendations on the use of targeted next-generation sequencing (tNGS) tests for the diagnosis of drug-resistant tuberculosis as an alternative to conventional phenotypic methods64. By systematically documenting resistance-associated mutations, the catalogue harmonized the detection and interpretation of resistance across various drugs51,52,54 contributing to tNGS adoption, for example in Namibia6.
AMR genes can disseminate independently of host species through horizontal gene transfer and these events may remain undetected where surveillance relies on conventional speciation and phenotypic susceptibility testing alone16,65. Genomic surveillance revealed that a hospital outbreak was driven by a plasmid (IncFII) carrying blaNDM-1 spread across multiple genetic K. pneumoniae lineages in a neonatal intensive care unit3. AMR databases can facilitate prospective detection and timely response to outbreaks through identification of epidemiologically relevant determinants. Infection prevention and control measures can then be directed towards the specific resistance mechanism. Genomic surveillance of Pseudomonas aeruginosa ST621 revealed a decades-long hospital outbreak that persisted through sink-drain reservoirs and evolved multidrug resistance, but targeted infection-control interventions informed by these genomic findings ultimately suppressed transmission and ended the outbreak4. Databases of biocide resistance determinants (Table 1) can further support infection prevention and control measures5,23,26.
An understanding of specific AMR determinants in clinical specimens can guide patient care by informing treatment guidelines specific to the local context and epidemiology. For example, knowledge of certain rpoB mutations can guide the clinical management of rifampin-resistant TB by identifying rifabutin susceptibility and prompting testing for other second-line drugs54. Similarly, the treatment of carbapenemase-producing gram-negative organisms depends on the underlying carbapenemase, as certain β-lactam antibiotics have activity against specific carbapenemases46.
Challenges
Pathogen genomics is transforming public health efforts to combat AMR. Decreasing costs have enabled wider adoption, and ongoing advances in sequencing technologies and analytical tools are poised to further enhance its impact. However, realizing the full potential in the areas described above requires addressing several challenges, including those limiting the accurate prediction of resistance phenotypes from genomic data. AMR databases are a central component of these workflows and here we focus on the challenges related to their completeness, accuracy, standardization, timeliness and accessibility.
Completeness
Completeness refers to the extent to which all required AMR determinants and associated information are present, but this depends on the specific purpose or purposes the database serves –for example, clinical resistance prediction versus high-resolution molecular epidemiology. Despite collaboration and data sharing (Fig. 1), AMR databases differ in size, age, scope, structure and purpose (Table 1). Consequently, different databases contain overlapping but different sets of AMR determinants for different pathogens and antibiotics or antibiotic classes35 (see Fig. 2 for a comparison of mutation coverage). For example, CARD collects AMR determinants regardless of their clinical relevance, while ResFinder focuses primarily on clinically-relevant ones. The WHO MTBc catalogue, Kleborate and Pathogenwatch are curated to predict species-specific AMR, while PointFinder, CARD and AMRFinderPlus do this for point mutations, but not necessarily for all antibiotics. This variation in AMR databases and tools not only reflects the complexity of bacterial AMR but also demonstrates fragmentation between databases and the duplication of efforts. Understanding the suitability and limitations of different AMR databases for pathogen and antibiotic combinations is essential for using and interpreting their outputs for different purposes.
Rows represent pathogens within the WHO bacterial pathogen priority list, while columns represent antibiotic/antibiotic classes. Colour shading indicate the total number of AMR databases that provide information on point mutations for each pathogen-antibitioc/antibiotic class combination (yellow = 1, pink = 2, violet = 3, blue = 4, dark blue = 5 databases [AMRFinderPlus, Kleborate, Pathogenwatch, PointFinder, Comprehensive Antimicrobial Resistance Database (CARD)]). Up to date as of 1st January 2025.
Completeness is challenged by our insufficient understanding of the molecular mechanisms of resistance –AMR databases can only catalogue known determinants. AMR databases to date have been mostly developed using data from high-income settings, limiting global representativeness66. Local and regional variation in AMR determinants associated with specific resistance phenotypes, coupled with a lack of contextual information (e.g., geographical origin, treatment outcomes), can hinder the identification of uncharacterized genes, alleles, or mutations and affect downstream interpretation. For example, in a study from Valencia (Spain)67, and during the development of the second version of the WHO MTBc catalogue54, the analysis of M. tuberculosis genomes not included in the first version revealed relevant mutations that were previously unidentified. This highlights the value of large collections of well-characterized and genetically diverse high-quality genomes from diverse sources and regions (with matched AST phenotypes) to improve the completeness of AMR databases.
Accuracy
Accuracy refers to the correct association between AMR determinants and the inferred resistance phenotype. Databases providing detailed ontological and hierarchical relationships between AMR determinants, antibiotics and phenotypes, such as CARD and AMRFinderPlus, allow a structured way to analyse and interpret data23,26. However, it is broadly assumed that the presence of a resistance gene or mutation directly impacts the resistance phenotype to a particular drug or drug class, because of the strong genotype-phenotype concordance found for some species and antimicrobial classes48,49,51,68,69. AMR databases often include determinants whose presence alone does not necessarily confer clinically relevant resistance, such as some species-specific, intrinsic AMR genes (Table 1)27,51. For example, chromosomal blaOXA-51-like genes are ubiquitous in A. baumannii and encode weak carbapenemases which do not alter treatment options70. Differences between databases in cataloguing and reporting this additional information (clinical relevance, intrinsic resistance and species background) may affect interpretation and lead to inaccurate resistance phenotype predictions.
Moreover, databases do not completely cover the biological complexity of AMR in different species. The phenotypic expression of resistance can be affected by mobile genetic elements, mechanisms of gene regulation, changes in promoters/silencers, mutations in other intergenic regions, de novo ribosomal RNA mutations, increases in gene copy number and the additive effect of different AMR mechanisms and their interactions13. For example, the performance of CARD predictions for aminoglycoside and fluoroquinolone compounds in E. coli improved when compared to ResFinder, after exclusion of predictions of resistance based on determinants linked to efflux pump genes, whose expression is greatly influenced by transcription regulation71. So the mere presence of an AMR determinant does not guarantee expression sufficient to confer resistance detectable by standard phenotypic methods. Furthermore, differences in interpretive thresholds, such as epidemiological cut-offs versus clinical breakpoints (e.g., CLSI or EUCAST) affect the perceived concordance between genotypic and phenotypic resistance72,73.
Expert curation and the creation of expert rules72,73 are needed to improve the accuracy of genotype-phenotype associations in AMR databases, as demonstrated for Klebsiella spp.27, M. tuberculosis51, E. faecium48, Pseudomonas aeruginosa74 and Salmonella Typhi75. However, species-specific curation is time-intensive and particularly challenging for rarer and fastidious organisms where genotypic and phenotypic data are limited, such as Haemophilus influenzae76. In addition, identifying “solo” determinants, the basis of the genotype-phenotype associations in the MTBc catalogue, is considerably more challenging in organisms where resistance genes are frequently acquired en bloc through horizontal gene transfer.
Standardization
The absence of global unified standards for naming and identifying resistance genes and alleles hinders comparison and poses a significant challenge to local-to-global genomic surveillance of AMR77,78. Even with consistent content, databases may differ in gene and allele naming –either when different names are assigned to the same gene or protein, or when one database classifies a sequence as a gene and another one as an allele35. Older databases may not use the most up-to-date methods for curation and annotation.
Efforts to standardise nomenclature36,44, data structures, ontologies and hierarchies23,26 and harmonize gene name conventions between databases78,79 are important steps towards developing unified standards for more effective comparisons and data exchange. CARD and AMRFinderPlus participate in public discussion of nomenclature issues at the arpcard/amr_curation GitHub site80. Equally important are initiatives to standardize genomics workflows77, such as the ISO-certified AbritAMR30, and harmonization of AMR reports in a single consistent format81.
Timeliness
Regular database updates and revisions are crucial for timely inclusion of newly discovered AMR determinants and improved annotations, such as the reclassification of blaSHV alleles by Kleborate44. New versions are often released when new evidence requires substantial additions or revisions (Table 1). Manual curation of peer-reviewed experimental evidence ensures rigorous evaluation, but is labour-intensive, time-consuming and potentially subjective. As the number of publications grows16, manual curation and regular updates become increasingly time- and resource-consuming. Some database developers have addressed these challenges through machine learning methods to optimize literature review32 and the use of quality control algorithms34. Standardized data-driven approaches to curation would allow frequent analyses of large datasets; however, curation would still rely on human expertise82,83.
When databases are updated, the many bioinformatic pipelines implementing local copies also need updates. Despite database versioning –i.e. the release of discrete snapshots of a database over time with tracked changes51,54,84, outdated legacy databases and tools are still available online and their use may lead to inaccurate results.
Accessibility and usage
Choosing the most appropriate AMR database (Table 1), prediction tool, and parameters requires understanding of the available options and bioinformatics expertise, which may vary across different settings. Additionally, parameters used can significantly affect whether genes are matched from query sequences to databases, and ideal parameters for gene detection may change based on bacterial species and subtype85. To address this, AMR databases and their associated bioinformatic tools are available through a variety of web-based interfaces, which makes them well-suited for rapid response in resource-limited settings. However, they often provide less capacity to process large datasets and less customization compared to command-line tools.
Conclusion and future perspectives
This review highlighted the variability in AMR databases as well as their growing significance for public health. They play important roles in genomic surveillance, outbreak management, and infection prevention and control. To realize the full potential of AMR databases to enable these interventions, collaborative efforts should focus on improving data quality, standardization, global representativeness, and accessibility while avoiding duplication and fragmentation. The WHO MTBc catalogue is a powerful example of the impact of a community-driven, comprehensive, standardized catalogue, and similar initiatives are needed for all priority pathogens9,10.
WHO currently hosts the Global Antimicrobial Resistance Surveillance System (GLASS), which in the coming years will integrate genomic data for enhanced local-to global surveillance and situational awareness. GLASS will need centralized resources for AMR genomic data, including tools, guidance and a catalogue of AMR determinants. As a centralized, comprehensive resource, the catalogue may initially focus on selected priority pathogens and clinically significant determinants to support global surveillance. Compiled through collaboration and leveraging the many existing initiatives outlined in this review, the catalogue may evolve to include data from diverse settings, ensuring global representation. International collaboration and data sharing, according to data sharing principles86,87, will be vital for curating and harmonizing AMR databases with increased completeness, coverage and representation.
Future iterations may incorporate statistical models with certainty estimates to support clinical care. The integration of genomic and phenotypic data will be essential for developing models to predict phenotypic resistance from genome sequences44,72,82,83,88,89. Machine learning methods and artificial intelligence are poised to improve phenotype prediction by identifying low-certainty calls26,37,44,51,52, novel AMR mechanisms, and interactions between mechanisms90. Large, high-quality datasets spanning different sources and regions, and encompassing a wide range of pathogen lineages, will be needed to inform robust models91,92. Implementation of data standards and quality control will ensure validity of the catalogue51,54,93,94,95,96,97,98.
Centralized resources accessible to users with varied expertise will ensure that public-health benefits are equitably distributed, including the identification of geographically relevant targets for drug, diagnostics and vaccine development. To accelerate progress, we call on governments, public health agencies, researchers and funders to commit sustained support for building and maintaining standardized AMR genomic resources, a global priority8,9,10. This ambitious vision promises to significantly advance our understanding and management of AMR on a global scale.
Data availability
The datasets and code used to construct Table 1 and Fig. 2 can be accessed here: https://worldhealthorg-my.sharepoint.com/:f:/g/personal/centnerc_who_int/IgC8e2i67D5yQrzOXxD2ChKPARK2uRA8K8CqXHIxd26fdSI?e=DHahWP. They were generated using datasets obtained from CARD [https://card.mcmaster.ca/download]; AMRFinderplus [https://github.com/ncbi/amr/wiki/AMRFinderPlus-database]; ResFinder [https://bitbucket.org/genomicepidemiology/resfinder_db/src/master/]; PointFinder [https://bitbucket.org/genomicepidemiology/pointfinder_db/src/master/]; Kleborate [https://github.com/klebgenomics/Kleborate], Pathogenwatch [https://github.com/pathogenwatch-oss/amr-libraries]; and the WHO MTBc catalogue [https://github.com/GTB-tbsequencing/mutation-catalogue-2023/tree/main/Final%20Result%20Files] repositories.
References
UN General Assembly (79th sess : 2024-2025). Political Declaration of the High-Level Meeting on Antimicrobial Resistance: resolution A/RES/79/2: adopted by the General Assembly. (2024).
Baker, K. S. et al. Evidence review and recommendations for the implementation of genomics for antimicrobial resistance surveillance: reports from an international expert group. Lancet Microbe 4, e1035–e1039 (2023).
Argimón, S. et al. Integrating whole-genome sequencing within the National Antimicrobial Resistance Surveillance Program in the Philippines. Nat. Commun. 11, 2719 (2020).
Stribling, W. et al. Detecting, mapping, and suppressing the spread of a decade-long Pseudomonas aeruginosa nosocomial outbreak with genomics. eLife 13, https://doi.org/10.7554/eLife.93181 (2025).
Kelemen, J. et al. Complex infection-control measures with disinfectant switch help the successful early control of carbapenem-resistant Acinetobacter baumannii outbreak in intensive care unit. Antibiotics 13, 869 (2024).
de Araujo, L. et al. Implementation of targeted next-generation sequencing for the diagnosis of drug-resistant tuberculosis in low-resource settings: a programmatic model, challenges, and initial outcomes. Front Public Health 11, 1204064 (2023).
Okeke, I. N. et al. Leapfrogging laboratories: the promise and pitfalls of high-tech solutions for antimicrobial resistance surveillance in low-income settings. BMJ Glob. Health 5, e003622 (2020).
Seventy-Seventh World Health Assembly. Antimicrobial resistance: accelerating national and global responses: WHO strategic and operational priorities to address drug-resistant bacterial infections in the human health sector, 2025–2035 (A77/5), https://apps.who.int/gb/ebwha/pdf_files/WHA77/A77_5-en.pdf (2024).
World Health Organization. Global research agenda for antimicrobial resistance in human health. (World Health Organization, 2024).
Bertagnolio, S. et al. WHO global research priorities for antimicrobial resistance in human health. Lancet Microbe 5, 100902 (2024).
Ellington, M. J. et al. The role of whole genome sequencing in antimicrobial susceptibility testing of bacteria: report from the EUCAST Subcommittee. Clin. Microbiol. Infect. 23, 2–22 (2017).
Boolchandani, M., D’Souza, A. W. & Dantas, G. Sequencing-based methods and resources to study antimicrobial resistance. Nat. Rev. Genet. 20, 356–370 (2019).
Darby, E. M. et al. Molecular mechanisms of antibiotic resistance revisited. Nat. Rev. Microbiol. 21, 280–295 (2023).
Partridge, S. R., Kwong, S. M., Firth, N. & Jensen, S. O. Mobile genetic elements associated with antimicrobial resistance. Clin. Microbiol. Rev. 31, https://doi.org/10.1128/cmr.00088-17 (2018).
Heritier, C., Poirel, L. & Nordmann, P. Cephalosporinase over-expression resulting from insertion of ISAba1 in Acinetobacter baumannii. Clin. Microbiol. Infect. 12, 123–130 (2006).
World Health Organization. GLASS whole-genome sequencing for surveillance of antimicrobial resistance. (World Health Organization, 2020).
Papp, M. & Solymosi, N. Review and comparison of antimicrobial resistance gene databases. Antibiotics 11, 339 (2022).
Hendriksen, R. S. et al. Using genomics to track global antimicrobial resistance. Front Public Health 7, 242 (2019).
Ferreira, I. et al. Species identification and antibiotic resistance prediction by analysis of whole-genome sequence data by use of ARESdb: an analysis of isolates from the unyvero lower respiratory tract infection trial. J. Clin. Microbiol. 58, https://doi.org/10.1128/JCM.00273-20 (2020).
World Health Organization. WHO Bacterial Priority Pathogens List, 2024: bacterial pathogens of public health importance to guide research, development and strategies to prevent and control antimicrobial resistance. (World Health Organization, 2024).
Davies, T. J. et al. Discordance between different bioinformatic methods for identifying resistance genes from short-read genomic data, with a focus on Escherichia coli. Microb Genom 9, https://doi.org/10.1099/mgen.0.001151 (2023).
Florensa, A. F., Kaas, R. S., Clausen, P., Aytan-Aktug, D. & Aarestrup, F. M. ResFinder - an open online resource for identification of antimicrobial resistance genes in next-generation sequencing data and prediction of phenotypes from genotypes. Microb. Genom. 8, https://doi.org/10.1099/mgen.0.000748 (2022).
Feldgarden, M. et al. AMRFinderPlus and the Reference Gene Catalog facilitate examination of the genomic links among antimicrobial resistance, stress response, and virulence. Sci. Rep. 11, 12728 (2021).
Hunt, M. et al. ARIBA: rapid antimicrobial resistance genotyping directly from sequencing reads. Micro. Genom. 3, e000131 (2017).
Seemann, T. Abricate, https://github.com/tseemann/abricate (2017).
Alcock, B. P. et al. CARD 2023: expanded curation, support for machine learning, and resistome prediction at the Comprehensive Antibiotic Resistance Database. Nucleic Acids Res. 51, D690–D699 (2023).
Lam, M. M. C. et al. A genomic surveillance framework and genotyping tool for Klebsiella pneumoniae and its related species complex. Nat. Commun. 12, 4188 (2021).
Centre for Genomic Pathogen Surveillance. Pathogenwatch, https://pathogen.watch/.
Kaprou, G. D., Bergspica, I., Alexa, E. A., Alvarez-Ordonez, A. & Prieto, M. Rapid Methods for Antimicrobial Resistance Diagnostics. Antibiotics 10, https://doi.org/10.3390/antibiotics10020209 (2021).
Sherry, N. L. et al. An ISO-certified genomics workflow for identification and surveillance of antimicrobial resistance. Nat. Commun. 14, https://doi.org/10.1038/s41467-022-35713-4 (2023).
Alcock, B. P. et al. CARD 2020: antibiotic resistome surveillance with the comprehensive antibiotic resistance database. Nucleic Acids Res. 48, D517–d525 (2020).
Edalatmand, A. & McArthur, A. G. CARD*Shark: automated prioritization of literature curation for the Comprehensive Antibiotic Resistance Database. Database 2023, https://doi.org/10.1093/database/baad023 (2023).
CARD Curation Team. AMR curation, https://github.com/arpcard/amr_curation (2019).
Feldgarden, M. et al. Curation of the AMRFinderPlus databases: applications, functionality and impact. Microb. Genom. 8, https://doi.org/10.1099/mgen.0.000832 (2022).
Feldgarden, M. et al. Validating the AMRFinder tool and resistance gene database by using antimicrobial resistance genotype-phenotype correlations in a collection of isolates. Antimicrob. Agents Chemother. 63, e00483–19 (2019).
Bradford, P. A. et al. Consensus on beta-lactamase nomenclature. Antimicrob. Agents Chemother. 66, e0033322 (2022).
Sayers, E. W. et al. Database resources of the National Center for Biotechnology Information in 2025. Nucleic Acids Res. 53, D20–D29 (2024).
Zankari, E. et al. Identification of acquired antimicrobial resistance genes. J. Antimicrob. Chemother. 67, 2640–2644 (2012).
Bortolaia, V. et al. ResFinder 4.0 for predictions of phenotypes from genotypes. J. Antimicrob. Chemother. 75, 3491–3500 (2020).
Zankari, E. et al. PointFinder: a novel web tool for WGS-based detection of antimicrobial resistance associated with chromosomal point mutations in bacterial pathogens. J. Antimicrob. Chemother. 72, 2764–2768 (2017).
Gschwind, R. et al. ResFinderFG v2.0: a database of antibiotic resistance genes obtained by functional metagenomics. Nucleic Acids Res. 51, W493–W500 (2023).
Liu, B. & Pop, M. ARDB-antibiotic resistance genes database. Nucleic Acids Res. 37, D443–D447 (2009).
Naas, T. et al. Beta-lactamase database (BLDB)—structure and function. J. Enzym. Inhib. Med. Chem. 32, 917–919 (2017).
Tsang, K. K. et al. Diversity, functional classification and genotyping of SHV beta-lactamases in Klebsiella pneumoniae. Microb. Genom. 10, https://doi.org/10.1099/mgen.0.001294 (2024).
Gupta, S. K. et al. ARG-ANNOT, a new bioinformatic tool to discover antibiotic resistance genes in bacterial genomes. Antimicrob. Agents Chemother. 58, 212–220 (2014).
Tamma, P. D. et al. Infectious diseases society of america 2024 guidance on the treatment of antimicrobial-resistant gram-negative infections. Clin. Infect Dis., https://doi.org/10.1093/cid/ciae403 (2024).
Argimon, S. et al. Rapid Genomic Characterization and Global Surveillance of Klebsiella Using Pathogenwatch. Clin. Infect. Dis. 73, S325–S335 (2021).
Coll, F. et al. Antibiotic resistance determination using Enterococcus faecium whole-genome sequences: a diagnostic accuracy study using genotypic and phenotypic data. Lancet Microbe 5, e151–e163 (2024).
Argimón, S. et al. A global resource for genomic predictions of antimicrobial resistance and surveillance of Salmonella Typhi at pathogenwatch. Nat. Commun. 12, https://doi.org/10.1038/s41467-021-23091-2 (2021).
Sánchez-Busó, L. et al. A community-driven resource for genomic epidemiology and antimicrobial resistance prediction of Neisseria gonorrhoeae at Pathogenwatch. Genome Med. 13, https://doi.org/10.1186/s13073-021-00858-2 (2021).
Walker, T. M. et al. The 2021 WHO catalogue of Mycobacterium tuberculosis complex mutations associated with drug resistance: a genotypic analysis. Lancet Microbe 3, e265–e273 (2022).
World Health Organization. Catalogue of mutations in Mycobacterium tuberculosis complex and their association with drug resistance. (World Health Organiation, 2021).
World Health Organization. TB Sequencing, Mutations (2023).
World Health Organization. Catalogue of mutations in Mycobacterium tuberculosis complex and their association with drug resistance. 2nd edn, (World Health Organization, 2023).
Hunt, M. et al. Antibiotic resistance prediction for Mycobacterium tuberculosis from genome sequence data with Mykrobe. Wellcome Open Res. 4, 191 (2019).
Phelan, J. E. et al. Integrating informatics tools and portable sequencing technology for rapid detection of resistance to anti-tuberculous drugs. Genome Med. 11, 41 (2019).
World Health Organization. Global Antimicrobial Resistance Surveillance System (GLASS): molecular methods for antimicrobial resistance (AMR) diagnostics to enhance the Global Antimicrobial Resistance Surveillance System. (World Health Organization, 2019).
Guitor, A. K. et al. Capturing the Resistome: a Targeted Capture Method To Reveal Antibiotic Resistance Determinants in Metagenomes. Antimicrob Agents Chemother 64, https://doi.org/10.1128/AAC.01324-19 (2019).
Knight, M. E. et al. National-scale antimicrobial resistance surveillance in wastewater: a comparative analysis of HT qPCR and metagenomic approaches. Water Res 262, 121989 (2024).
Chen, C. Y. et al. Detection of antimicrobial resistance using proteomics and the comprehensive antibiotic resistance database: a case study. Proteom. Clin. Appl 14, e1800182 (2020).
Cerezales, M. et al. Novel multiplex PCRs for detection of the most prevalent carbapenemase genes in Gram-negative bacteria within Germany. J. Med. Microbiol. 70, https://doi.org/10.1099/jmm.0.001310 (2021).
Garciglia Mercado, C. et al. Development of a LAMP method for detection of carbapenem-resistant Acinetobacter baumannii during a hospital outbreak. J. Infect. Dev. Ctries 14, 494–501 (2020).
Boutal, H. et al. The Revolution of Lateral Flow Assay in the Field of AMR Detection. Diagnostics 12, https://doi.org/10.3390/diagnostics12071744 (2022).
World Health Organization. WHO consolidated guidelines on tuberculosis: module 3: diagnosis: rapid diagnostics for tuberculosis detection, 3rd ed. (World Health Organization, 2024).
Parcell, B. J., Gillespie, S. H., Pettigrew, K. A. & Holden, M. T. G. Clinical perspectives in integrating whole-genome sequencing into the investigation of healthcare and public health outbreaks—hype or help? J. Hospital Infect. 109, 1–9 (2021).
Nurjadi, D., Boutin, S. & Velavan, T. P. Are low-income and middle-income countries being neglected in genomic surveillance of the emergence and spread of multidrug-resistant Enterobacterales? Lancet Microbe, https://doi.org/10.1016/S2666-5247(24)00069-7 (2024).
Garcia-Marin, A. M. et al. Role of the first WHO mutation catalogue in the diagnosis of antibiotic resistance in Mycobacterium tuberculosis in the Valencia Region, Spain: a retrospective genomic analysis. Lancet Microbe 5, e43–e51 (2024).
Aanensen, D. M. et al. Whole-Genome Sequencing for Routine Pathogen Surveillance in Public Health: a Population Snapshot of Invasive Staphylococcus aureus in Europe. mBio 7, e00444–00416 (2016).
Rebelo, A. R. et al. One day in Denmark: comparison of phenotypic and genotypic antimicrobial susceptibility testing in bacterial isolates from clinical settings. Front Microbiol 13, 804627 (2022).
Turton, J. F. et al. The role of ISAba1 in expression of OXA carbapenemase genes in Acinetobacter baumannii. FEMS Microbiol Lett. 258, 72–77 (2006).
Mahfouz, N., Ferreira, I., Beisken, S., von Haeseler, A. & Posch, A. E. Large-scale assessment of antimicrobial resistance marker databases for genetic phenotype prediction: a systematic review. J. Antimicrob. Chemother. 75, 3099–3108 (2020).
Holt, K. & Hawkey, J. AMRRules, <https://github.com/AMRverse/AMRrules> (2024).
EUCAST. The EUCAST Subcommittee on WGS and Phenotypic AST, <https://www.eucast.org/organization/subcommittees/wgs-and-phenotypic-testing> (2025).
Madden, D. E. et al. Keeping up with the pathogens: improved antimicrobial resistance detection and prediction from Pseudomonas aeruginosa genomes. Genome Med. 16, 78 (2024).
Ingle, D. J. et al. Typhi Mykrobe: fast and accurate lineage identification and antimicrobial resistance genotyping directly from sequence reads for the typhoid fever agent Salmonella Typhi. bioRxiv, https://doi.org/10.1101/2024.09.30.613582 (2024).
Diricks, M. et al. Revisiting mutational resistance to ampicillin and cefotaxime in Haemophilus influenzae. Genome Med 16, 140 (2024).
Nunez-Garcia, J. et al. Harmonisation of in-silico next-generation sequencing based methods for diagnostics and surveillance. Sci. Rep. 12, 14372 (2022).
Ugarcina Perovic, S. et al. argNorm: normalization of antibiotic resistance gene annotations to the Antibiotic Resistance Ontology (ARO). Bioinformatics, https://doi.org/10.1093/bioinformatics/btaf173 (2025).
Anthony Underwood, A. W., Alex Manuele, Inês Mendes, Thanh Le Viet, Trestan Pillonel, Varun Shamanna. CharmeDb, <https://gitlab.com/antunderwood/chamredb> (2021).
arpcard/amr_curation, <https://github.com/arpcard/amr_curation/issues> (2025).
Mendes, I. et al. hAMRonization: Enhancing antimicrobial resistance prediction using the PHA4GE AMR detection specification and tooling. bioRxiv, https://doi.org/10.1101/2024.03.07.583950 (2024).
Kim, J. I. et al. Machine learning for antimicrobial resistance prediction: current practice, limitations, and clinical perspective. Clin. Microbiol. Rev. 35, e0017921 (2022).
Wheeler, N. E. et al. Innovations in genomic antimicrobial resistance surveillance. Lancet Microbe 4, e1063–e1070 (2023).
Libuit, K. et al. Ten Best Practices for Public Health Bioinformatics Pipelines, <https://github.com/pha4ge/public-health-pipeline-best-practices/blob/main/docs/pipeline-best-practices.md> (2024).
Hetman, B. M., Pearl, D., Reid-Smith, R., Parmley, E. & Taboada, E. An epidemiological framework for improving the accuracy of whole-genome sequence-based antimicrobial resistance surveillance in Salmonella. Can. J. Microbiol. 71, 1–17 (2025).
World Health Organization. WHO guiding principles for pathogen genome data sharing. (World Health Organization, 2022).
Griffiths, E. J. et al. The PHA4GE Microbial Data-Sharing Accord: establishing baseline consensus microbial data-sharing norms to facilitate cross-sectoral collaboration. BMJ Glob Health 9, https://doi.org/10.1136/bmjgh-2024-016474 (2024).
Hu, K. et al. Assessing computational predictions of antimicrobial resistance phenotypes from microbial genomes. Brief Bioinform. 25, https://doi.org/10.1093/bib/bbae206 (2024).
Ren, Y. et al. Prediction of antimicrobial resistance based on whole-genome sequencing and machine learning. Bioinformatics 38, 325–334 (2022).
Batisti Biffignandi, G. et al. Optimising machine learning prediction of minimum inhibitory concentrations in Klebsiella pneumoniae. Microb. Genom. 10, https://doi.org/10.1099/mgen.0.001222 (2024).
Nsubuga, M., Galiwango, R., Jjingo, D. & Mboowa, G. Generalizability of machine learning in predicting antimicrobial resistance in E. coli: a multi-country case study in Africa. BMC Genomics 25, 287 (2024).
Hall, M. B. & Coin, L. J. M. Assessment of the 2021 WHO Mycobacterium tuberculosis drug resistance mutation catalogue on an independent dataset. Lancet Microbe 3, e645 (2022).
Kristensen, T. et al. Results of the 2020 Genomic Proficiency Test for the network of European Union Reference Laboratory for Antimicrobial Resistance assessing whole-genome-sequencing capacities. Microb Genom. 9, https://doi.org/10.1099/mgen.0.001076 (2023).
Sorensen, L. H. et al. Whole-genome sequencing for antimicrobial surveillance: species-specific quality thresholds and data evaluation from the network of the European Union Reference Laboratory for Antimicrobial Resistance genomic proficiency tests of 2021 and 2022. mSystems, e0016024, (2024).
Lau, K. A. et al. Proficiency testing for bacterial whole genome sequencing in assuring the quality of microbiology diagnostics in clinical and public health laboratories. Pathology 53, 902–911 (2021).
Petrillo, M. et al. A roadmap for the generation of benchmarking resources for antimicrobial resistance detection using next generation sequencing. F1000Res 10, 80 (2021).
Raphenya, A. R. et al. Datasets for benchmarking antimicrobial resistance genes in bacterial metagenomic and whole genome sequencing. Sci. Data 9, https://doi.org/10.1038/s41597-022-01463-7 (2022).
Strepis, N. et al. BenchAMRking: a Galaxy-based platform for illustrating the major issues associated with current antimicrobial resistance (AMR) gene prediction workflows. BMC Genomics 26, 27 (2025).
Acknowledgements
The authors acknowledge the input of David Aanensen, Frank Møller Aarestrup, Kathryn Holt, Yvan Hutin, Nazir Ismail, Andrew McArthur, Arjun Prasad and René S. Hendricksen. The salary of SDG was supported by a grant from the Fleming Fund and the salary for AB was supported by a grant from the Centers for Disease Control and Prevention. The funder played no role in study design, data collection, analysis and interpretation of data, or the writing of this manuscript. The findings and conclusions in this article are those of the authors and do not necessarily reflect the official positions of WHO or any other institutions mentioned.
Author information
Authors and Affiliations
Contributions
C.C.: writing—original draft, writing—review and editing, conceptualization, methodology, project administration, supervision, visualization; S.D.G.: writing—original draft, writing—review and editing, conceptualization, methodology, data curation, formal analysis, visualization; A.D.: Writing—review and editing, conceptualization; A.B.: writing—original draft, writing—review and editing, conceptualization, visualization, methodology, data curation; S.A.: writing—review and editing, conceptualization; D.M.: writing—review and editing, conceptualization; S.B.: writing—review and editing, conceptualization, methodology, project administration, supervision.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Centner, C.M., Di Gregorio, S., Argimón, S. et al. Antimicrobial resistance databases: opportunities and challenges for public health. npj Antimicrob Resist 4, 1 (2026). https://doi.org/10.1038/s44259-025-00169-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s44259-025-00169-1