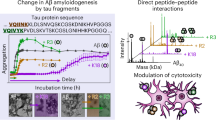
Abstract
Accumulation of Amyloid β (Aβ), a peptide derived from endocytic processing of the amyloid precursor protein (APP), is a critical initial step in the development of Alzheimer’s disease (AD). While the APP695E590D mutation was previously discovered in 2 pathologically confirmed AD patients, the pathogenicity of this mutation has remained uncertain due to its exceptional rarity. Here, we characterize the APP695E590D mutation by evaluating multiple APP metabolites and determining its effects on tauopathy in cellular and animal models. We show that APP695E590D not only increases Aβ through endocytic β-secretase processing but also increases Aη, an alternative APP-derived synaptotoxic peptide. We further demonstrate that APP695E590D promotes tauopathy by increasing tau seeding and aggregation in cellular models and exacerbating phospho-tau pathology and neuroinflammation in tauP301S mice. These results reveal a unique modality by which APP695E590D impinges on AD pathology by enhancing both Aβ and Aη generation and accelerating tauopathy.
Similar content being viewed by others
Introduction
The accumulation of misfolded amyloid β (Aβ) and tau are widely recognized as disease-driving pathological hallmarks of Alzheimer’s disease (AD)1,2,3,4. Aβ generation occurs through sequential β-secretase (BACE1) and γ-secretase (presenilin complex) processing of the amyloid precursor protein (APP), steps that require the endocytosis of APP from the plasma membrane5,6,7. Indeed, disease-causing APP mutations, which segregate with early-onset familial AD (FAD)8,9,10,11, drive increased Aβ production and/or aggregation of Aβ10,12,13,14,15,16. Although such mutations are rare (<1% of AD), genetic and biochemical studies support Aβ as a crucial pathogenic factor in AD. Beyond Aβ, however, APP also plays important roles in physiological processes, such as neurogenesis, neurite outgrowth, and synaptic function, as evidenced by loss-of-function and gain-of-toxic function studies using multiple models17,18. Notably, β-secretase inhibition, which has shown lack of efficacy in AD clinical trials19, increases the alternative η-cleavage of APP occurring ~93 residues N-terminal to the β-cleavage. This results in the generation of Aη, a neurotoxic ~15 kDa peptide that impairs synaptic plasticity, including hippocampal long-term potentiation (LTP)20.
Beyond the ~50 pathogenic APP mutations identified to date, another 50 + APP mutations identified are of unknown or uncertain pathogenic significance21,22,23,24,25,26. In 1994, APP E665D mutation (based on APP770 isoform) was first identified in a late-onset AD patient, whom upon postmortem examination, displayed marked neuronal loss, astrogliosis, neuritic plaques, and neurofibrillary tangles27. In 2020, Hsu and colleagues evaluated 22 APP mutations and 42 PSEN1/2 mutations of unknown significance for Aβ42 and Aβ40 levels in transfected cells by ELISA, which included the APP E665D mutation28. In this study, APP-E665D increased Aβ42 secretion by ~28% compared to APP-WT, which fell slightly short of significance28. Although it was not clear whether secreted Aβ was normalized to transfected intracellular APP levels, this study suggested that APP E665D mutation may be benign28. In 2021, however, APP E665D mutation was identified again in an early-onset AD patient (45 years old) with progressive memory decline and behavioral dysfunction29. PET amyloid imaging showed elevated Aβ deposition, and MRI revealed white matter hyperintensities and scattered cortical lesions consistent with cerebral amyloid angiopathy (CAA)29. In addition, Aβ and phospho-tau levels in the cerebrospinal fluid (CSF) of this patient were consistent with AD29. Further, the patient developed cognitive and neuropsychological dysfunction starting from childhood, suggesting that APP E665D mutation may be pathogenic, perhaps driving disease phenotypes even beyond Aβ accumulation. Based on these observations, we performed an extensive assessment of multiple APP metabolites derived from the APP E665D mutation (E590D in APP695 isoform) by Western blotting. Our new findings show that APP695E590D not only increases Aβ through faster APP internalization and β-cleavage but also increases APP η-cleavage and elevates Aη, an alternative neurotoxic peptide derived from APP20. We further demonstrate that APP695E590D expression promotes tau seeding and aggregation in cellular and animal models, collectively supporting a pathogenic role of this mutation and establishing a unique modality by which APP links to AD pathology.
Results
APPE590D increases Aβ and Aη generation
Fig. 1A illustrates the corresponding position of the E665D and E590D mutation in APP770 and APP695 isoforms, respectively, surrounded by α, β, and η cleavage sites and recognition epitopes of the antibodies used to determine APP metabolites (Fig. 1A). We used the neuronal isoform APP695 to generate the E590D mutation by site-directed mutagenesis to test whether this mutation alters APP processing. We transiently transfected HEK293T cells with equal amounts of wild type (WT) APP695WT or APP695E590D and collected the media and lysates to detect full-length APP and its proteolytic products. Compared to APP695WT, APP695E590D exhibited a ~ 2.4-fold increase in Aβ secretion normalized to full-length APP (Fig. 1B, C), concomitant with ~1.8-fold and ~2.3-fold increases in secreted APPβ (sAPPβ) and intracellular ~12 kDa C-terminal fragment-β (CTFβ) (Fig. 1B–E), indicating that the E590D mutation promotes β-secretase processing of APP via BACE1. Alternative η- and α/β-processing of APP results in secretion of ~15 kDa Aη peptides, of which Aη-α increases with BACE1 inhibition20. We observed ~15 kDa doublet Aη peptides in the medium as detected by the D54D2 antibody, which significantly increased by ~4-fold in cells transfected with APP695E590D compared to APP695WT, normalized to full-length APP (Fig. 1A, B, F). As a substantial proportion of η-cleaved APP can escape α- and β-cleavages, APP695E590D cells exhibited a marked increase in the ~25 kDa APP-CTFη (Fig. 1B, G), a CTF that can subsequently serve as a substrate for more Aη and/or Aβ generation20.
A Schematic of APP770, APP695, and sAPPβ with corresponding domains, cleavage sites (η, β, & α), mutation site (E665D or E590D), and antibody epitopes. B Representative blots of HEK293T cells transfected with vector control (VC), APP695WT, or APP695E590D and subjected to Western blotting from the medium and lysates with D54D2 antibody (medium : Aη & Aβ), sAPPβ antibody (medium), 6E10 antibody (lysate: APP, CTFη, & CTFβ), and actin antibody (lysate). Quantification of (C) Aβ, (D) CTFβ, (E) sAPPβ, (F) Aη and (G) CTFη levels normalized to total APP expression (t test, **P < 0.01, ****P < 0.0001, n = 6–9 / group). H Blots of mouse primary neurons transduced with rAAV9 expressing vector control (VC), APP695WT, or APP695E590D at DIV14 for 7 days and subjected to Western blotting from the medium and lysates with D54D2 antibody (medium : Aη & Aβ), sAPPβ antibody (medium), 6E10 antibody (lysate: APP), and actin antibody (lysate). Quantification of (I) Aβ, (J) sAPPβ, and (K) Aη levels normalized to total APP expression (t-test, *P < 0.05, **P < 0.01, n = 3 / group).
To determine whether APPE590D similarly alters APP processing in neurons, we transduced mouse primary neurons with rAAV9 expressing vector control, APP695WT, or APP695E590D. Indeed, we observed significant increases in Aβ and sAPPβ by APP695E590D compared to APP695WT normalized to full-length APP (Fig. 1H–J). Despite the lower levels of secreted Aη in mouse primary neurons, we also detected close to threefold increase in Aη by APP695E590D compared to APP695WT normalized to full-length APP (Fig. 1H, K). These results indicate that the E590D mutation promotes both BACE1-dependent Aβ and BACE1-independent Aη production in non-neuronal and neuronal cells.
APPE590D reduces cell surface APP and enhances APP internalization
While α- and η-secretase processing of APP are thought to occur on the cell surface via ADAM10/17 and metalloproteases (i.e., MT5-MPP), respectively, to generate Aη-α20, β- and γ-secretase processing to generate Aβ requires APP internalization from the cell surface30,31,32,33,34,35,36,37,38,39,40,41. Thus, we next assessed whether the E590D mutation alters surface APP levels. First, we measured cell surface APP levels of transfected cells by immunocytochemistry (ICC) using the 6E10 antibody at 4 oC without membrane permeabilization, which showed significantly reduced surface APP695E590D compared to APP695WT (Fig. 2A, B). Second, we used a biochemical approach to biotinylate the cell surface on ice, followed by immunoprecipitation (IP) for biotin and detection of APP. Normalized to total full-length APP, we detected a highly significant ~60% reduction in surface APP695E590D compared to surface APP695WT (Fig. 2C, D).
A Representative images of HEK293T cells transfected with equal amounts of APP695WT or APP695E590D and subjected to staining for surface APP with 6E10 antibody without membrane permeabilization. B Quantification of surface APP levels (t test, ****P < 0.0001, n = 30–31 images / group from 4 repeats). C Representative blots of HEK293T cells transfected with vector control (VC), APP695WT, or APP695E590D and subjected to cell surface biotinylation and immunoprecipitation (IP) for biotin and immunoblotting (IB) for APP (6E10 antibody). Lower 2 panels showing total APP and actin levels in lysates (input). D Quantification of surface biotinylated APP normalized to total APP expression (t test, ****P < 0.0001, n = 6 / group). E Representative images of HEK293T cells transfected with vector control (VC), APP695WT, or APP695E590D for 48 h and subjected to incubation with 6E10 antibody on ice for 30 min to label surface APP, resumption of endocytosis for 10 min at 37 oC, fixation, and ICC for EEA1 (early endosome marker) and visualization of 6E10-labeled APP. F Quantification of % APP colocalization with EEA1 (t test, ****P < 0.0001, n = 148-153 images / group from 4 repeats). G Representative blots of HEK293T cells transfected with vector control (VC), APP695WT, or APP695E590D for 48 h and subjected to cell surface biotinylation at 4 oC, followed by endocytosis at 37 oC for 10 min and removal of surface biotin at 4 oC. Internalized biotinylated proteins immunoprecipitated (IP) with biotin antibody and internalized APP detected with 6E10 antibody. Lower 2 panels showing total APP (6E10) and actin levels in lysates (input). H Quantification of internalized APP normalized to total APP expression (t test, **P < 0.01, n = 6 / group).
Upon APP internalization from the cell surface, APP is localized in early endosomes, where β- and γ-secretase activities mediate Aβ generation42. To detect APP in early endosomes, we incubated transfected cells on ice for 30 min with 6E10 antibody to label surface APP. Cells were then incubated at 37 oC for 10 min to allow endocytosis, after which, cells were stained with the early endosome marker EEA1 together with visualization of 6E10-bound APP. Consistent with reduced levels of surface APP695E590D, APP695E590D exhibited >50% increase in colocalization with EEA1 puncta compared to APP695WT (Fig. 2E, F), indicating faster internalization of APP695E590D to early endosomes. We also performed biotin internalization experiments, where we biotinylated surface proteins on ice, followed by internalization at 37 °C for 10 min. After removal of surface biotinylated proteins on ice, we then performed immunoprecipitation of biotinylated proteins to detect internalized APP. Similar to that observed with immunofluorescence-based internalization experiments, we observed significantly greater internalization of biotinylated APP695E590D compared to APP695E590D, normalized to total APP levels (Fig. 2G, H).
APPE590D increases tau seeding and aggregation
Previous studies have shown that mutant APP or Aβ enhances tau seeding and aggregation2,43,44,45,46,47. We next determined whether APP695E590D can influence tau seeding using HEK293T Tau-RD-P301S-CFP/YFP biosensor cells, in which the uptake of proteopathic tau seeds stimulates the aggregation of RD-CFP and RD-YFP, producing a strong fluorescent punctate signal48,49,50. Hence, after transfecting Tau-RD-P301S-CFP/YFP biosensor cells with vector control, APP695WT, or APP695E590D for 24 h, we treated cells with sonicated tau seeds from 8-month-old tauP301S mouse brains for 24 h. We then stained fixed cells for APP (6E10) and visualized tau-RD fluorescent puncta. Indeed, we observed a significant increase in tau-RD puncta (green) in APP695E590D cells (red) compared to both APP695WT (red) and control cells (Fig. 3A, B), indicating increased tau seeding by APP695E590D. To further assess the effects of APP695E590D on tau aggregation, we transfected HeLa-V5-tau cells stably expressing wild-type 4R tau with vector control, APP695WT, or APP695E590D. In RIPA-soluble lysates, tau levels appeared to be reduced in APP695E590D cells compared to APP695WT or vector control cells (Fig. 3C, D). To determine if this is due to changes in tau insolubility and aggregation, we performed filter trap assay to capture tau aggregates in the RIPA-insoluble fraction. Indeed, we observed a significant increase in captured tau aggregates in APP695E590D cells compared to APP695WT or vector control cells (Fig.3E, F). These results therefore indicate that APP695E590D increases tau seeding and aggregation beyond its WT counterpart.
A Representative images of Tau-RD biosensor cells transfected with vector control (VC), APP695WT, or APP695E590D for 24 h and subjected to treatment with sonicated tau seeds from 8-month-old tauP301S mouse brain for 24 h. Representative images showing tau-RD puncta (green) and APP staining (red, 6E10). Yellow arrows showing tau-RD puncta (green) in APP695E590D cells (red). B Quantification of tau-RD puncta intensity normalized by APP expression (red) (1-way ANOVA, post hoc Tukey, *P < 0.05, **P < 0.001, ns=not significant, n = 3, 20 images/replicate). C–F Hela-V5-tau cells stably expressing 4R tau transfected with vector control (VC), APP695WT, or APP695E590D for 48 h and subjected to (C) Western blotting for APP (6E10), tau (A-10), and actin from the RIPA-soluble fraction (1-way ANOVA, post hoc Tukey, *P < 0.05, n = 3 / group); (D) quantification of soluble tau from the RIPA-soluble fraction and (E) filter-trap assay from the RIPA-insoluble fraction to capture tau aggregates. F Quantification of tau aggregates captured by filter-trap assay from the RIPA-insoluble fraction (1-way ANOVA, post hoc Tukey, **P < 0.01, ***P < 0.001, n = 3 / group).
APP695E590D exacerbates tauopathy and neuroinflammation in tauP301S mice
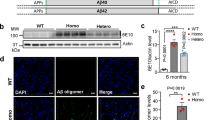
Having determined that the overexpression of APP695E590D but not APP695WT increases tau seeding and aggregation, we subsequently investigated whether overexpression of APP695E590D also exacerbates tauopathy in the brains of tauP301S (PS19) mice. Thus, we performed stereotaxic injections of control AAV9, AAV9-APP695WT, or AAV9-APP695E590D into the hippocampus of 5-month-old tauP301S mice. Three months post-injections, our findings demonstrated that overexpression of APP695E590D elevated both RIPA-soluble and RIPA-insoluble phospho-tau (pS199/202) as identified through immunoblotting (Fig. 4A–C). Likewise, immunohistochemistry revealed elevated pS199/202-tau in both the cortex and hippocampus of tauP301S mouse brains by the overexpression of APP695E590D (Fig. 4E–J). Notably, unlike in vitro conditions, Aβ monomer (below 10kda) was not detectable in brains from either APP695WT or APP695E590D AAV-injected PS19 mice. A distinct ~17-kDa APP-derived fragment was observed only in APP695E590D-injected mice (Fig. 4A, D), suggesting altered APP processing by APP695E590D in vivo. As anticipated, these observations were associated with elevated astrogliosis (GFAP) and microgliosis (IBA1) by APP695E590D overexpression compared to control and APP695WT (Fig. 4H–J). Therefore, overexpression of APP695 carrying E590D mutation markedly exacerbates tauopathy and ensuing neuroinflammation in the brains of tauP301S mice.
A–C TauP301S mice (5 mo) transduced with purified high-titer APP695E590D rAAV9, APP695WT, or control by stereotaxic injection into the hippocampus for 3 months. Equal amounts of RIPA soluble and insoluble protein lysates from the hippocampus subjected to immunoblotting for APP (6E10), phospho-tau (pS199/202), and actin. B–D Quantification of soluble and insoluble phospho-tau (pS199/202), and Aη (1-way ANOVA, post hoc Tukey, *P < 0.05, ** P < 0.01, n = 3 mice for control and APP695E590D groups (one mouse died from each group after injections), n = 4 mice for APP695WT). E Representative images of brain slices from the hippocampus and cortex processed for confocal microscopy after IHC for phospho-tau (pS199/202, green). F, G Quantification of phospho-tau (pS199/202) from the cortex and hippocampus (1-way ANOVA, post hoc Tukey, *P < 0.05, ** P < 0.01, n = 3 mice/group, 15–19 images/mouse). H Representative hippocampal images of brain slices processed for confocal microscopy after IHC for IBA1 (green), and GFAP (red). I, J Quantification of hippocampal IBA1 and GFAP intensity (1-way ANOVA, post hoc Tukey, *P < 0.05, ** P < 0.01, n = 3 mice/group, 11–12 images/group).
Discussion
Many studies have shown that rare mutations in APP drive AD by increasing Aβ generation and/or promoting Aβ aggregation21,51,52, forming a foundational pillar for the amyloid hypothesis of AD53,54. Among APP mutations of unknown significance, the APP695E590D (or APP770E665D) mutation has thus far been reported in 2 AD cases to date. In the first AD case reported in 1994, postmortem examination confirmed the diagnosis of AD27. However, the availability of historical medical records was highly limited, and neither antemortem amyloid PET nor MRI examinations were performed27. In the second AD case reported in 2021, multiple measures, including CSF Aβ and tau, amyloid PET, MRI, and historical medical records were consistent with AD and supported the possibility that the APP695E590D mutation is pathogenic29. In this study, we conducted a thorough assessment of the APP695E590D mutation, including Aβ, Aη, sAPPβ, CTFβ, CTFη, APP endocytosis, and tau seeding and aggregation. Our results showed that the APP695E590D mutation not only increases Aβ, sAPPβ, and CTFβ, but also increases η-processing of APP to promote Aη generation, a previously characterized synaptotoxic peptide20. These results, together with the observed tauopathy-promoting effects of the APP695E590D mutation, strongly support the notion that this mutation is pathogenic and promotes AD pathological phenotype.
An earlier study examined 22 APP and 42 PSEN1/2 mutations for Aβ42 and Aβ40 levels by ELISA in transiently transfected N2a cells, in which the APP695E590D mutation was found to increase Aβ42 by ~28%, despite falling slightly shy of significance28. However, they did not state the numbers of experimental samples examined per mutation, and ELISA-based Aβ levels were not normalized to total APP expression28. In the current study, we measured total Aβ and did not distinguish Aβ species, which ranges from 36-43 amino acids55, and normalized all APP metabolites to total APP expression. Hence, ELISAs used to detect Aβ42 and Aβ40 in the previous study28 missed multiple different Aβ peptides and did not control for potential variations in APP expression. Second, we used HEK293T cells and primary neurons, while the previous study used mouse N2a cells28. Third, we assessed APP metabolites other than Aβ, such as Aη, sAPPβ, and APP-CTFs, whereas the previous study did not28. Therefore, directly comparing the results of the previous study to the current study is not possible. However, we note that the directional trend for increasing Aβ by APP695E590D mutation was seen in both studies. A significant novelty of the current study, however, is that APP695E590D mutation not only increased Aβ but also Aη and CTFη (Fig. 5, schematic), that latter that can serve as a substrate to generate more Aβ and/or Aη20.
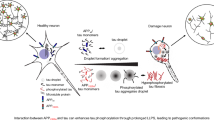
APP695E590D increases η-processing of APP on the cell surface (i.e., presynaptic), resulting in elevated Aη generation. APP695E590D also increases APP internalization, promoting Aβ generation through β-secretase processing (BACE1). The combination of Aη, Aβ, and/or APP695E590D ectodomain drives tau seeding and aggregation, contributing to synaptic dysfunction (i.e,. postsynaptic).
To our knowledge, APP695E590D mutation is the only mutation known to date that increases both Aβ and Aη levels, raising the intriguing possibility that elevated Aη peptide may contribute to unique brain dysfunctions beyond Aβ deposition. Notably, Aη is not deposited in plaque cores but decorates dystrophic neurites and impairs synaptic plasticity, including LTP20. Such activity of Aη in deregulating synaptic activity is consistent with the early-onset AD APP695E590D patient, who developed cognitive and neuropsychological dysfunction from early childhood29. We speculate that the combined effects of Aβ, Aη, and possibly other ectodomain products of APP695E590D promote tau seeding and aggregation, as seen in the current study (Fig. 5, schematic). Whether this occurs through intracellular and/or secretory mechanisms requires further investigation.
Unlike classical AD-linked APP missense mutations, which are closely juxtaposed to β- or γ-cleavage sites, the E590D mutation lies 6 residues N-terminal to the β-cleavage site within the Aη sequence. Therefore, it is conceivable that Aη peptides containing the E590D mutation may exert differential activity on synaptic plasticity and tauopathy, which will be highly informative to test in future studies. While this mutation conservatively alters the Aη peptide, resulting in one less methylene group in the aspartate side chain29, Aη sequence is known to be highly O-glycosylated on Thr558, Thr576, Thr577, Thr584, Thr588, and Ser59256,57. Hence, it is conceivable that the E590D mutation may alter O-glycosylation of nearby residues, such as Thr588 and Ser592. This, in turn, may shift APP695E590D accessibility to β-secretase (BACE1) and/or η-secretase. Indeed, a recent study showed that mucin-type O-glycosylation of APP on Thr588 and Ser592 changes the conformation of this region and increases BACE1-mediated cleavage in vitro58. While further studies are required to define the precise mechanism, the results of this study overall indicate that the overexpression of APP695 carrying E590D mutation is pathogenic by increasing both Aη and Aβ levels as well as promoting tau aggregation.
This study has several limitations. As we did not quantify α-secretase products (sAPPα, CTFα) in the nonamyloidogenic α-secretase pathway and the Aβ42:Aβ40 ratio from amyloidogenic γ-secretase pathway, our APP processing data do not fully define the balance of APP processing across all pathways. The inclusion of APP695WT AAV expression in vivo enabled differences between mutation-specific effects and APP overexpression alone, revealing no significant impact of APP695WT on tau pathology or glial activation in PS19 mice. Although APP processing products were assessed in brain tissue, canonical Aβ species were not detected, and a distinct ~17-kDa APP-derived fragment was observed selectively in APP695E590D-injected mice. Thus, while these findings support altered APP processing associated with the E590D variant in vivo, the precise molecular mechanisms linking APP695E590D to tau pathology remain to be fully elucidated and are supported in part by mechanistic insights from cell-based models. We also did not distinguish whether the effects of APPE590D on tau seeding and aggregation occur through intracellular or extracellular APP metabolites. Given that the E590D mutation resides in the ectodomain of APP, it is conceivable that this effect occurs, at least in part, through secreted paracrine and/or propagation pathways. While we focused on the pS199/pS202 phospho-tau epitope in this study, evaluation of additional phospho-tau epitopes could be informative future investigations. Nonetheless, these limitations do not impact our primary conclusion that APP695E590D promotes enhanced APP processing (Aβ & Aη) and tau seeding/aggregation. Rather, the current results lay the groundwork for future use of APPE590D knock-in or transgenic models to better understand the contribution of Aη together with Aβ in neurophysiology, neurodegeneration, and AD pathology.
Methods
Ethics approval
The Institutional Animal Care and Use Committee (IACUC) approved all methods and protocols used in this mouse study. All methods were carried out in accordance with the relevant guidelines and regulations, which were also approved by the IACUC and Institutional Biosafety Committees (IBC).
Antibodies and reagents
Monoclonal antibodies to β-actin (Cat# A3854-200UL) was purchased from MilliporeSigma (Burlington, MA, USA), and APP (6E10, Cat# 803003) was purchased from BioLegend (CA, USA). Anti-Aβ (D54D2, Cat#8243) and anti-EEA1(Cat#2411) antibodies were purchased from Cell Signaling Technology (Danvers, MA, USA). The anti-sAPP-β (6A1, Cat#10321) antibody was purchased from IBL America (Minneapolis, MN, USA). Anti-tau (A-10, sc390476) antibody was purchased from Santa Cruz (Dallas, Texas, USA). The anti-GFAP (Cat # 14-9892-82) and anti-phospho-Tau Ser199/ser202 (Cat # 44-768 G) antibodies were purchased from Thermo Scientific (MA, USA). Anti-Iba1(Cat# 019-19741) antibody was purchased from Wako Chemicals USA (Richmond, VA, USA).
Cell lines and primary neurons
HEK293T, Tau-RD-FRET, and Hela-V5-tau cells were cultured in Dulbecco’s modified Eagle’s medium (DMEM, Thermo Scientific, MA, USA), which was supplemented with 1% penicillin/streptomycin (P/S) and 10% fetal bovine serum (FBS). The procedure for preparing cultures of primary cortical neurons from P0 pups has been described previously59,60,61. Briefly, the cortex was isolated and dissected in ice-cold HBSS prior to trypsin digestion. Plates or coverslips containing poly-D-lysine (Sigma-Aldrich, St. Louis, MO, USA) in neurobasal medium (Invitrogen, Carlsbad, CA, USA), with 2% glutamax, and 2% B27 supplement were utilized to culture mouse neurons (Invitrogen, Carlsbad, CA, USA). At 37 °C, all cells were cultured in a humidified atmosphere with 5% CO2.
DNA constructs, transfections, and rAAV9 generation
The pLHCX-APP695-WT plasmid was used as template to perform mutagenesis to generate pLHCX-APP-E590D construct using Q5-site-Directed Mutagenesis Kit (New England Biolabs, Ipswich, MA, USA). To introduce variants of APP into the rAAV9 plasmid, APP695-WT and APP-E590D were subcloned into the pTR12.1-MCSW vector using restriction sites HindIII and NotI. Sequencing was performed to validate all cloned or subcloned plasmids before their application in experiments. For DNA plasmid transfection, Fugene HD (Promega, Madison, WI, USA) was utilized to transfect cells with DNA plasmids into Opti-MEM I (Invitrogen, Carlsbad, CA, USA). The cells were harvested 48 h after transfection in accordance with the manufacturer’s guidelines.
For AAV9 production, recombinant AAV9 viruses were produced in HEK293T cells via co-transfection of a serotype vector containing the target gene along with pAAV9 and pXX6. The produced viruses were then purified according to the referenced procedures62.
Cell lysis, filter-trap assay, and SDS-PAGE
Cells were lysed with RIPA lysis buffer (50 mM Tris pH 7.4, 150 mM NaCl, 2 mM EDTA, 1% NP-40, 0.1% sodium dodecyl sulfate)61,63. The concentrations of total protein were quantified using a colorimetric detection assay (BCA Protein Assay, Pierce, USA). Protein lysates in equal amounts were separated by sodium dodecyl sulfate (SDS)-polyacrylamide gel electrophoresis (PAGE) and transferred to a nitrocellulose membrane (Millipore Corporation, Bedford, MA, USA). For filter-trap assays (FTA), equal amounts of sonicated RIPA-insoluble extracts were filtered through 0.2 μm cellulose acetate membranes61,63,64 (ThermoFisher Scientific, MA, USA) using a 96-well vacuum dot blot apparatus (Bio-Rad, Hercules, CA, USA), followed by PBS washing and 20% methanol fixation. Interested proteins were probed with primary antibodies, followed by peroxidase-conjugated secondary antibodies and ECL detection (Merck Millipore Corporation, Darmstadt, Germany). All immunoblot images were acquired by LAS-4000 (GE Healthcare Biosciences, Pittsburgh, PA) or ImageQuant 800 (Amersham, Chicago, IL) and quantified using ImageJ (NIH, Bethesda, MD).
Surface biotinylation and Internalization assay
Surface biotinylation and internalization assays were performed as previously described65,66. Briefly, to initiate the process of surface biotinylation, cells were seeded into 6-well plates and subjected to treatment with 2 mg/ml sulfo-NHS-LC-biotin in phosphate-buffered saline (pH 8.0) while being gently agitated for an hour on ice after three washes (PBS). Following three washes with PBS, the cells were lysed in the lysis buffer containing 1% Nonidet P-40. Anti-biotin antibody pulldown was utilized in conjunction with anti-mouse agarose beads to isolate biotinylated proteins. For biotin internalization assay, cells in 6-well plate and subjected to three 20-min incubations in the solution containing 2 mg/ml sulfosuccinimidyl 2-(biotinamido)ethyl-1,3-dithiopropionate (Pierce, Rockford, IL, USA) at 4 °C. Following rinsing with cold PBS containing 0.1 M glycine, the cells were washed with cold PBS three times. To facilitate the internalization of the labeled proteins, cells were incubated at 37 °C for 10 min in culture medium. Internalization was halted as a result of the rapid cooling caused by ice. To cleave biotin that was exposed at the cell surface, the cells underwent three 20-min incubations at 4 °C with 50 mM 2-mercaptoethanesulfonic acid (Sigma) in 50 mM Tris-HCl (pH 8.7), 100 mM NaCl, and 2.5 mM CaCl2. After subjecting the cells to a thorough rinsing process using PBS supplemented with 20 mM HEPES, they were lysed in a 1% Nonidet P-40 buffer. Then, the biotinylated internalized proteins were isolated by immunoprecipitation with anti-biotin antibody and analyzed for APP protein level by immunoblotting.
Tau seeding activity assay
Tau seeding experiments were conducted using HEK293T Tau RD P301S FRET Biosensor (TauRD) cells, stably expressing the P301S tau repeat domain67. The cells were treated with tau seeds obtained from tauP301S mouse brain for 24 h, and tau seeding activity was evaluated by direct fluorescence intensity. The preparation of tau seeds has been previously documented68. Briefly, an 8-month-old tauP301S mouse brain was homogenized (10% w/v) in DPBS and centrifuged at 500 g for 5 min. The supernatant was centrifuged at 1000 × g for 5 min before protein concentration measurement. Tau seeds were sonicated before the treatment of TauRD cells.
Immunocytochemistry, immunohistochemistry, and fluorescent microscopy
For immunocytochemistry (ICC), cells were washed with PBS and fixed for 15 min at room temperature with 4% paraformaldehyde (PFA)61. Fixed cells were washed with PBS, incubated with blocking solution (0.2% Triton X-100, 3% normal goat serum) for 1 h, incubated with primary antibody at 4 °C overnight, washed three times with PBST, and incubated with Alexa-488 or Alexa-594-conjugated secondary IgG antibodies for 1 h at room temperature (Vector Laboratories, Burlingame, CA). The slides were then washed three times with PBST before being mounted with a fluorochrome mounting solution (Vector Laboratories). For immunohistochemistry (IHC), mice were transcardially perfused with phosphate-buffered saline (PBS). One hemisphere of each brain was snap-frozen and stored at −80 °C for subsequent biochemical analyses, while the other hemisphere was fixed in 4% paraformaldehyde at 4 °C for 24 h and then cryoprotected in 30% sucrose. Coronal brain sections (25 μm thick) were prepared and blocked with 3% normal goat serum with 0.2% Triton X-100 in PBS for 1 h at room temperature. Sections were incubated overnight at 4 °C with primary antibodies against the interested proteins. After washing, sections were incubated with Alexa Fluor-conjugated secondary antibodies (Alexa-488 or Alexa-594) for 1 h at room temperature, followed by mounting. All images were captured with Nikon AX Ti2 confocal (Tokyo, Japan), and ImageJ software (NIH, Bethesda, MD) was used to quantify the immunoreactivities. In all ICC and IHC experiments, all comparison images had the same laser intensity, exposure time, and filter. During image acquisition and quantification, investigators were blinded to the experimental conditions, and regions of interest were selected randomly. Brightness/contrast adjustments were applied uniformly to all comparison images.
Generation of rAAV9 and stereotaxic injections in mice
Recombinant AAV9 virus was produced by co-transfecting HEK293 cells with the APPE695E590D or APPE695WT expressed plasmid pTR12.1-MCS, pAAV9, and pXX6 helper plasmid, followed by purification according to established protocols62. TauP301S (PS19) transgenic mice on a C57BL/6 background were maintained under standard vivarium conditions with ad libitum access to food and water and a 12-h light/dark cycle. For stereotaxic injections, five-month-old mice (sex-balanced across conditions) were anesthetized with isoflurane and secured in a stereotaxic apparatus. Small burr holes were drilled in the skull using a dental drill (SSW HP-3; SSWhite Burs, Lakewood, NJ, USA). Bilateral injections targeting the hippocampus were performed using a 26-gauge needle connected to a 10-μL Hamilton syringe (Hamilton, Reno, NV, USA). The injection coordinates relative to bregma were: anteroposterior −2.7 mm, lateral −2.7 mm, and dorsoventral 3.0 mm. A total of 2.5 μL of purified rAAV9 (1 × 10¹¹ vg/mL) was delivered per site over a 2-min period using convection-enhanced delivery. Following the injection, incisions were sutured and cleaned, and animals were monitored until recovery, typically within 20 min. Mice were housed individually and sacrificed 3 months post-injection for downstream analyses.
Statistical analysis
Statistical analysis of all graphs was conducted using GraphPad Prism 8.0 (GraphPad Software, San Diego, CA, USA), which included the following methods: 1-way analysis of variance (ANOVA) with Dunnett or Sidak post hoc tests or student’s t test. Error bars in graphs represent SEM with significant set at P < 0.05.
Data availability
All data needed to evaluate the conclusions of the paper are present in the paper.
References
Hampel, H. et al. The amyloid-beta pathway in Alzheimer’s disease. Mol. Psychiatry 26, 5481–5503 (2021).
Bloom, G. S. Amyloid-beta and tau: the trigger and bullet in Alzheimer disease pathogenesis. JAMA Neurol. 71, 505–508 (2014).
Iqbal, K., Liu, F., Gong, C. X. & Grundke-Iqbal, I. Tau in Alzheimer disease and related tauopathies. Curr. Alzheimer Res. 7, 656–664 (2010).
Iqbal, K. et al. Tau pathology in Alzheimer disease and other tauopathies. Biochim Biophys. Acta. 1739, 198–210 (2005).
Octave, J. N. et al. The role of presenilin-1 in the gamma-secretase cleavage of the amyloid precursor protein of Alzheimer’s disease. J. Biol. Chem. 275, 1525–1528 (2000).
Cole, S. L. & Vassar, R. The Alzheimer’s disease beta-secretase enzyme, BACE1. Mol. Neurodegener. 2, 22 (2007).
Thinakaran, G. & Koo, E. H. Amyloid precursor protein trafficking, processing, and function. J. Biol. Chem. 283, 29615–29619 (2008).
Cacace, R., Sleegers, K. & Van Broeckhoven, C. Molecular genetics of early-onset Alzheimer’s disease revisited. Alzheimers Dement 12, 733–748 (2016).
Lanoiselee, H. M. et al. APP, PSEN1, and PSEN2 mutations in early-onset Alzheimer disease: A genetic screening study of familial and sporadic cases. PLoS Med. 14, e1002270 (2017).
Goate, A. et al. Segregation of a missense mutation in the amyloid precursor protein gene with familial Alzheimer’s disease. Nature 349, 704–706 (1991).
Rosenberg, R. N., Lambracht-Washington, D., Yu, G. & Xia, W. Genomics of Alzheimer disease: A review. JAMA Neurol. 73, 867–874 (2016).
Nguyen, M., Wong, Y. C., Ysselstein, D., Severino, A. & Krainc, D. Synaptic, mitochondrial, and lysosomal dysfunction in Parkinson’s disease. Trends Neurosci. 42, 140–149 (2019).
Dorszewska, J., Prendecki, M., Oczkowska, A., Dezor, M. & Kozubski, W. Molecular basis of familial and sporadic Alzheimer’s disease. Curr. Alzheimer Res. 13, 952–963 (2016).
Devkota, S., Williams, T. D. & Wolfe, M. S. Familial Alzheimer’s disease mutations in amyloid protein precursor alter proteolysis by gamma-secretase to increase amyloid beta-peptides of >/=45 residues. J. Biol. Chem. 296, 100281 (2021).
Shepherd, C., McCann, H. & Halliday, G. M. Variations in the neuropathology of familial Alzheimer’s disease. Acta Neuropathol. 118, 37–52 (2009).
Scheuner, D. et al. Secreted amyloid beta-protein similar to that in the senile plaques of Alzheimer’s disease is increased in vivo by the presenilin 1 and 2 and APP mutations linked to familial Alzheimer’s disease. Nat. Med. 2, 864–870 (1996).
Guo, Q., Wang, Z., Li, H., Wiese, M. & Zheng, H. APP physiological and pathophysiological functions: insights from animal models. Cell Res. 22, 78–89 (2012).
Dai, M. H., Zheng, H., Zeng, L. D. & Zhang, Y. The genes associated with early-onset Alzheimer’s disease. Oncotarget 9, 15132–15143 (2018).
Volloch, V. & Rits, S. Results of beta secretase-inhibitor clinical trials support amyloid precursor protein-independent generation of beta amyloid in sporadic Alzheimer’s Disease. Med. Sci. (Basel) 6, https://doi.org/10.3390/medsci6020045 (2018).
Willem, M. et al. eta-Secretase processing of APP inhibits neuronal activity in the hippocampus. Nature 526, 443–447 (2015).
Tcw, J. & Goate, A. M. Genetics of beta-amyloid precursor protein in Alzheimer’s Disease. Cold Spring Harb. Perspect Med. 7, https://doi.org/10.1101/cshperspect.a024539 (2017).
Cahill, M. K. & Huang, E. J. Testing the amyloid hypothesis with a humanized AD mouse model. Neuron. 93, 987–989 (2017).
Bagyinszky, E., Youn, Y. C., An, S. S. & Kim, S. Mutations, associated with early-onset Alzheimer’s disease, discovered in Asian countries. Clin. Inter. Aging 11, 1467–1488 (2016).
Wu, L. et al. Early-onset familial Alzheimer’s disease (EOFAD). Can. J. Neurol. Sci. 39, 436–445 (2012).
Cruchaga, C. et al. Rare variants in APP, PSEN1 and PSEN2 increase risk for AD in late-onset Alzheimer’s disease families. PLoS One 7, e31039 (2012).
Mullan, M. et al. A pathogenic mutation for probable Alzheimer’s disease in the APP gene at the N-terminus of beta-amyloid. Nat. Genet 1, 345–347 (1992).
Peacock, M. L. et al. Novel amyloid precursor protein gene mutation (codon 665Asp) in a patient with late-onset Alzheimer’s disease. Ann. Neurol. 35, 432–438 (1994).
Hsu, S. et al. Systematic validation of variants of unknown significance in APP, PSEN1 and PSEN2. Neurobiol. Dis. 139, 104817 (2020).
Abbatemarco, J. R. et al. Amyloid precursor protein variant, E665D, associated with unique clinical and biomarker phenotype. Am. J. Alzheimers Dis. Other Demen. 36, 1533317520981225, https://doi.org/10.1177/1533317520981225 (2021).
Kimura, N. & Yanagisawa, K. Traffic jam hypothesis: Relationship between endocytic dysfunction and Alzheimer’s disease. Neurochem Int 119, 35–41 (2018).
Sosa, L. J. et al. The physiological role of the amyloid precursor protein as an adhesion molecule in the developing nervous system. J. Neurochem. 143, 11–29 (2017).
Hoe, H. S. et al. The effects of amyloid precursor protein on postsynaptic composition and activity. J. Biol. Chem. 284, 8495–8506 (2009).
Jaeger, S. & Pietrzik, C. U. Functional role of lipoprotein receptors in Alzheimer’s disease. Curr. Alzheimer Res. 5, 15–25 (2008).
Muller, U. C. & Deller, T. Editorial: The physiological functions of the APP gene family. Front Mol. Neurosci. 10, 334 (2017).
Copenhaver, P. F. & Kogel, D. Role of APP interactions with heterotrimeric G proteins: Physiological functions and pathological consequences. Front Mol. Neurosci. 10, 3 (2017).
Muller, U. C. & Zheng, H. Physiological functions of APP family proteins. Cold Spring Harb. Perspect. Med. 2, a006288 (2012).
Muller, U. C., Pietrzik, C. U. & Deller, T. The physiological functions of the beta-amyloid precursor protein APP. Exp. Brain Res 217, 325–329 (2012).
Swaminathan, G., Zhu, W. & Plowey, E. D. BECN1/Beclin 1 sorts cell-surface APP/amyloid beta precursor protein for lysosomal degradation. Autophagy 12, 2404–2419 (2016).
Feng, T. et al. SNX15 regulates cell surface recycling of APP and Abeta generation. Mol. Neurobiol. 53, 3690–3701 (2016).
Choy, R. W., Cheng, Z. & Schekman, R. Amyloid precursor protein (APP) traffics from the cell surface via endosomes for amyloid beta (Abeta) production in the trans-Golgi network. Proc. Natl. Acad. Sci. USA 109, E2077–E2082 (2012).
Talamagas, A. A., Efthimiopoulos, S., Tsilibary, E. C., Figueiredo-Pereira, M. E. & Tzinia, A. K. Abeta(1-40)-induced secretion of matrix metalloproteinase-9 results in sAPPalpha release by association with cell surface APP. Neurobiol. Dis. 28, 304–315 (2007).
Zhang, X. & Song, W. The role of APP and BACE1 trafficking in APP processing and amyloid-beta generation. Alzheimers Res. Ther. 5, 46 (2013).
LaFerla, F. M. Pathways linking Abeta and tau pathologies. Biochem Soc. Trans. 38, 993–995 (2010).
Selenica, M. L. et al. Amyloid oligomers exacerbate tau pathology in a mouse model of tauopathy. Neurodegener. Dis. 11, 165–181 (2013).
Pickett, E. K. et al. Amyloid beta and tau cooperate to cause reversible behavioral and transcriptional deficits in a model of Alzheimer’s disease. Cell Rep. 29, 3592–3604 e3595 (2019).
Imbimbo, B. P., Ippati, S., Watling, M. & Balducci, C. A critical appraisal of tau-targeting therapies for primary and secondary tauopathies. Alzheimers Dement 18, 1008–1037 (2022).
Pang, K. et al. An App knock-in rat model for Alzheimer’s disease exhibiting Abeta and tau pathologies, neuronal death and cognitive impairments. Cell Res 32, 157–175 (2022).
Carlomagno, Y. et al. The AD tau core spontaneously self-assembles and recruits full-length tau to filaments. Cell Rep. 34, 108843 (2021).
Kfoury, N., Holmes, B. B., Jiang, H., Holtzman, D. M. & Diamond, M. I. Trans-cellular propagation of Tau aggregation by fibrillar species. J. Biol. Chem. 287, 19440–19451 (2012).
Yuste-Checa, P. et al. The extracellular chaperone Clusterin enhances Tau aggregate seeding in a cellular model. Nat. Commun. 12, 4863 (2021).
Van Cauwenberghe, C., Van Broeckhoven, C. & Sleegers, K. The genetic landscape of Alzheimer disease: clinical implications and perspectives. Genet Med. 18, 421–430 (2016).
Selkoe, D. J. & Hardy, J. The amyloid hypothesis of Alzheimer’s disease at 25 years. EMBO Mol. Med. 8, 595–608 (2016).
Tanzi, R. E. & Bertram, L. Twenty years of the Alzheimer’s disease amyloid hypothesis: a genetic perspective. Cell 120, 545–555 (2005).
Kepp, K. P., Robakis, N. K., Hoilund-Carlsen, P. F., Sensi, S. L. & Vissel, B. The amyloid cascade hypothesis: an updated critical review. Brain 146, 3969–3990 (2023).
Chen, G. F. et al. Amyloid beta: Structure, biology and structure-based therapeutic development. Acta Pharm. Sin. 38, 1205–1235 (2017).
Halim, A. et al. Site-specific characterization of threonine, serine, and tyrosine glycosylations of amyloid precursor protein/amyloid beta-peptides in human cerebrospinal fluid. Proc. Natl. Acad. Sci. USA 108, 11848–11853 (2011).
Shi, J. et al. Comprehensive analysis of O-glycosylation of amyloid precursor protein (APP) using targeted and multi-fragmentation MS strategy. Biochim Biophys. Acta Gen. Subj. 1865, 129954 (2021).
Singh, Y. et al. Mucin-type O-glycosylation proximal to beta-secretase cleavage site affects APP processing and aggregation fate. Front Chem. 10, 859822 (2022).
Woo, J. A. et al. Activated cofilin exacerbates tau pathology by impairing tau-mediated microtubule dynamics. Commun. Biol. 2, 112 (2019).
Fang, C. et al. SSH1 impedes SQSTM1/p62 flux and MAPT/Tau clearance independent of CFL (cofilin) activation. Autophagy 17, 2144–2165 (2021).
Liu, T. et al. Modulation of synaptic plasticity, motor unit physiology, and TDP-43 pathology by CHCHD10. Acta Neuropathol. Commun. 10, 95 (2022).
Woo, J. A. et al. Loss of function CHCHD10 mutations in cytoplasmic TDP-43 accumulation and synaptic integrity. Nat. Commun. 8, 15558 (2017).
Liu, T. et al. CHCHD10-regulated OPA1-mitofilin complex mediates TDP-43-induced mitochondrial phenotypes associated with frontotemporal dementia. FASEB J. 34, 8493–8509 (2020).
Yan, Y. et al. X-linked ubiquitin-specific peptidase 11 increases tauopathy vulnerability in women. Cell 185, 3913–3930 e3919 (2022).
Lakshmana, M. K. et al. Novel role of RanBP9 in BACE1 processing of amyloid precursor protein and amyloid beta peptide generation. J. Biol. Chem. 284, 11863–11872 (2009).
Woo, J. A. et al. Slingshot-Cofilin activation mediates mitochondrial and synaptic dysfunction via Abeta ligation to beta1-integrin conformers. Cell Death Differ. 22, 1069–1070 (2015).
Holmes, B. B. et al. Proteopathic tau seeding predicts tauopathy in vivo. Proc. Natl. Acad. Sci. USA 111, E4376–E4385 (2014).
McCarthy, J. M. et al. Development of P301S tau seeded organotypic hippocampal slice cultures to study potential therapeutics. Sci. Rep. 11, 10309 (2021).
Acknowledgements
This work was supported by grants from the National Institutes of Health (NIH) to T.L. (R01AG086365, R03AG084948, R21AG095389 & R21AG070299), to D.E.K. (R01NS122350 & R01AG080924), to J.A.W. (RF1NS134638), and to D.E.K. and J.A.W. (R01AG067741 & RF1NS122218); and Veterans Affairs to D.E.K. (BX006539). D.E.K. is also supported by the Howard T. Karsner Professorship in Pathology, CWRU. T.L. was supported by the Research Education component of the Cleveland Alzheimer’s Disease Research Center (NIA P30 AG072959).
Author information
Authors and Affiliations
Contributions
Conceptualization: T.L., D.E.K. Investigation: T.L., L.W., D.R., Z.K., Z.Z., P.K., V.G., H.J. Visualization: T.L., J.A.W., D.E.K. Formal analysis: T.L., D.E.K. Project administration: T.L., J.A.W., D.E.K. Supervision: T.L., J.A.W., D.E.K. Funding acquisition: T.L., J.A.W., D.E.K. Writing-original draft: T.L., D.E.K. Writing-review and editing: T.L., J.A.W., D.E.K.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Liu, T., Wetzel, L., Roy, D. et al. APP E590D mutation increases generation of Aβ and Aη peptides and exacerbates tauopathy. npj Dement. 2, 21 (2026). https://doi.org/10.1038/s44400-026-00069-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s44400-026-00069-9