Abstract
In shrimp aquaculture, white spot syndrome virus (WSSV) infections severely impact production. Previous research highlighted the crucial role of the Penaeus monodon Rab7 (PmRab7) protein in WSSV entry, specifically its interaction with the viral envelope protein VP28. PmRab7 exists in two conformations: GDP-bound (inactive) and GTP-bound (active). This study, using ELISA and isothermal titration calorimetry (ITC), reveals that the PmRab7-VP28 interaction occurs irrespective of the nucleotide binding state of PmRab7. Comparing the binding affinity between VP28 and different PmRab7 conformations, including wild-type (WT, 22.5 nM), a fast nucleotide exchange (L129F, 128 nM), a GDP-bound form (T22N, 334 nM), and a favorably GTP-bound form (Q67L, 1990 nM), PmRab7-WT exhibits the strongest binding affinity, especially at a lower temperature (25 °C). The binding of PmRab7-WT and VP28 in the presence of excess nucleotide (WT with excess GDP, 924 nM, and WT with excess GTP, 826 nM) shows a 2-fold higher binding affinity than in the absence (WT, 1920 nM) indicating that the addition of excess nucleotide for PmRab7-WT enhanced the affinity for VP28. Together, these findings support the potential of PmRab7-WT as a promising therapeutic candidate for WSSV control in shrimp. Furthermore, from an industrial point of view, the ITC platform developed to study the VP28-PmRab7 interactions provides a high-throughput method for screening additives for shrimp feed that can inhibit this interaction.
Similar content being viewed by others
Introduction
Globally, shrimp production has increased in recent years as shrimp has become one of the most commonly traded seafood commodities with Penaeus vannamei and P. monodon as the leading species1. The industrial output is significantly affected by pathogen outbreaks. Major diseases in shrimp farming include White Spot Disease (WSD) from White Spot Syndrome Virus (WSSV), Acute Hepatopancreatic Necrosis Disease (AHPND or EMS) from a specific strain of the bacterium Vibrio parahaemolyticus, Hepatopancreatic Microsporidiosis (HPM) from the microsporidian Enterocytozoon hepatopenaei (EHP) and White Feces Syndromes (WFS) of which causative agents may include, but not limited to, the bacteria in the genus Vibrio sp., and the bacterium Propionigenium in combination with EHP2,3,4.
WSSV causes a systemic infection that leads to a severe mortality rate of up to 80–100% within 5 to 10 days post infection5. It is a circular double-stranded DNA virus with a genome size of approximately 300 kb6. The virion consists of three layers: the innermost rod-shaped nucleocapsid, a central tegument, and the outermost envelope7. Four major envelope proteins, VP28, VP26, VP24, and VP19, have been identified with VP26 and VP28 as the most abundant8,9,10.
VP28 plays a major role in viral attachment and trafficking into shrimp cells11. In vivo and in vitro WSSV neutralization with a polyclonal antibody against VP28 confirmed that the protein is key to a systemic infection11,12. Most importantly, VP28 binds to a myriad of shrimp proteins including C-type lectins, heat-shock cognate protein 70 (Hsc70), and Rab7 to attach and enter host cells10,13,14,15,16. In addition to WSSV, Rab7 in shrimp has been implicated in the entry of other viruses including Yellow Head Virus (YHV), Taura Syndrome Virus (TSV), and Laem-Singh Virus (LSNV)17,18,19,20.
Rab7 in P. monodon (PmRab7) was identified as a WSSV-VP28 binding protein during viral entry15. It has been predicted that the switch II region on the surface of PmRab7 may play an important role in VP28 binding21. The pivotal role of Rab7 in WSSV has been extensively documented. Silencing of Rab7 gene expression in both P. monodon and P. vannamei using a specific double-stranded RNA (dsRNA) targeting Rab7 markedly decreased the mRNA expression of a WSSV marker to less than 5%18 and also reduced the mortality rate to around 50%22. An injection of a mixture of PmRab7 recombinant proteins or an antibody against PmRab7 with WSSV in P. monodon reduced the shrimp mortality rate by around 80% compared to the injection with WSSV alone15,23. Furthermore, feeding shrimp with feed supplemented with recombinant PmRab7 expressed in Pichia Pastoris as intracellular proteins or surface exposed proteins lowered the mortality rate by approximately 50%24,25. These experiments underscore that PmRab7 can be developed into an effective anti-WSSV agent. The use of the recombinant protein represents a promising strategy for reducing WSSV infection in shrimp on an industrial scale due to the low cost of production25. To manipulate the anti-WSSV activity of PmRab7, a further understanding of the PmRab7-VP28 interaction needs to be established.
Rab7 belongs to the Rab family of the Ras superfamily of small G proteins (GTPases). Small GTPases are recognized as essential regulators of protein trafficking among late endosomes and lysosomes26. Like other small GTPases, Rab7 alternates between a GDP-bound or inactive conformation and a GTP-bound or active conformation27. While mutations in many small GTPases lead to a variety of diseases28,29,30, they provide valuable tools for deciphering the mechanism of how these regulators associate with their partners. In a mutant classified as a dominant negative, the enzyme remains in an inactive GDP-bound state31,32,33. In Rab7, the T22N mutation loses the hydroxyl group of T22 which coordinates the divalent Mg2+ ion in the nucleotide binding site and accounts for the inability to exchange for a fresh molecule of GTP31,34,35. In contrast, the Q67L mutation in the switch II region of Rab7 results in a deficiency in GTP hydrolysis and a constitutively active phenotype which is characterized by an accumulation of the GTP bound conformation32,34,35,36,37. Another interesting mutation is an L129F missense mutation associated with Charcot-Marie-Tooth disease in humans35,38. While the Phe substitution still allows both GTP and GDP binding, it renders the nucleotide binding pocket shallower and is attributed to a quickened nucleotide exchange rate35,38. In terms of the interaction with binding partner, the L129F mutation retains its ability to interact with Rab-interacting lysosomal protein (RILP) which is characteristic of the GTP-bound state of Rab738. In the case of shrimp, the functional study of the L129F mutant has not been provided.
In humans, binding partners of these small GTPases that were investigated exhibit differing preferences for the nucleotide states of the small GTPases. Most partners exclusively bind to the GTP bound form such as the interaction between GAP and Ras p21, the interaction between the coiled-coil protein (CCDC186/CCCP-1) and Rab2a, the interaction between the metazoan-specific autophagy gene (EPG5) and Rab7, the interaction between RILP and Rab7, and the interaction between Rubicon and Rab739,40,41,42,43,44. On the other hand, a few number of partners are selectively associated with the GDP-bound conformation, namely the interaction between protrudin and Rab11 and the interaction between coronin 3 and Rab27a45,46. Last but not least, some effectors interact with small GTPases without discriminating between the GTP- and GDP-bound states; for instance, the interaction of α/β-tubulin heterodimer with the small GTPase domain of leucine-rich repeat kinase 2 (LRRK2), the interaction of π 4-kinase (PI4KB) with Rab11, the interaction of a phosphatidylinositol 30-kinase and its adaptor protein complex (hVPS34/p150) with Rab7, and the interaction of an N-terminal ankyrin repeat domain of oxysterol-binding protein-related protein 1 (ARDN of ORP1L) with Rab742,47,48,49,50. However, it is unclear which nucleotide binding state of PmRab7 that VP28 prefers.
In this study, the nucleotide binding states of PmRab7-WT and their mutants were characterized by fluorescence spectroscopy. Subsequently, the interactions between VP28 and the different PmRab7 forms were investigated using an enzyme-linked immunosorbent assay (ELISA) and isothermal titration calorimetry (ITC). Our results revealed that VP28 did not display any preference for any particular nucleotide binding state and the dissociation constant (KD) of PmRab7-WT to VP28 is stronger than that of other mutants. Moreover, the presence of either GTP or GDP further strengthens the PmRab7-WT and VP28 interaction. Understanding which particular nucleotide-bound form of PmRab7 is preferentially bound by VP28 will enable the design of recombinant PmRab7 proteins that can effectively inhibit WSSV infection by competing with the endogenous PmRab7 in shrimp. This information can guide the development of targeted therapies, such as engineered PmRab7 variants or small molecules that stabilize the preferred conformation, to disrupt the VP28-PmRab7 interaction in shrimp and prevent viral entry into host cells. Accordingly, this investigation recommends that PmRab7-WT should serve as the best candidate for further development of the WSSV control strategy.
Result
Q67L, T22N, and L129F PmRab7 mutants were candidates for studying the effect of the nucleotide binding state of PmRab7 on the PmRab7-VP28 interaction
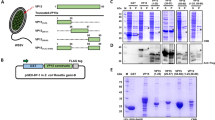
To observe the preference of VP28 on the nucleotide binding state PmRab7, three types of PmRab7 mutants were constructed by aligning the primary sequence of PmRab7 against that of Rab7 homologs and other members of the Ras protein superfamily (Fig. 1). The alignment showed that the percent identity of PmRab7 to human and rat Rab7 were 85.37% and 86.34%, respectively with 100% conservation at residues involved in GTP binding and catalysis. Thus, the mutations of PmRab7 were constructed based on previously published studies32,33,38,42. The constitutively active GTP-bound conformation was constructed as Q67L, while the dominant negative GDP-bound conformation was generated as T22N. The other candidate for the study was the fast-nucleotide-exchange L129F mutant. The WT and all mutants were purified and shown on SDS-PAGE with an expected size of approximately 23 kDa (Fig. 2). The purified PmRab7 proteins migrated as two major bands which were both identified as PmRab7 by a mass spectrometry analysis. The smaller band probably was a truncated PmRab7 that occurred from a TEV digestion (Supplementary Table S1).
PmRab7 mutants in this study were designed based on a multiple sequence alignment of PmRab7 against its homologs. Multiple sequence alignment of PmRab7 (DQ231062, Genbank), Rat_Rab7a (NP_076440.1, Genbank), human Rab7 (P51149, Uniprot), human Rab11a (P62491, Uniprot), human Rab11b (Q15907, Uniprot), human KRas; (P01116, Uniprot), human HRas (P01112, Uniprot), and human RhoA (P61586, Uniprot) was constructed using ClustalW program. The G1 to G5 domains of small GTPase proteins are boxes. A GTP/Mg2+ binding site is shown in grey. The switch II region is shown in blue. The star (*), single (.) and double (:) dots represent identical, conserved, and semi-conserved residues in the respective regions, respectively. Arrows indicate each mutated residue in T22N, Q67L, and L129F mutants. The positions of tryptophan residues are highlighted on a dark background.
Purified VP28 and PmRab7 on 12.5% SDS-PAGE electrophoresis. Lane M, protein marker; lane 1 purified VP28; lane 2 purified PmRab7-WT; lane 3 purified PmRab7-T22N; lane 4 purified PmRab7-Q67L; lane 5 purified PmRab7-L129F; lane 6 purified GST-PmRab7-WT; lane 7 previously verified fraction of PmRab7-WT (after the GST cleavage by the TEV protease).
The nucleotide binding states of PmRab7-WT were characterized by IFS
Prior to identifying the effect of nucleotide binding to PmRab7 on the PmRab7-VP28 interaction, an assay that can track the nucleotide binding state of PmRab7 must be established. An intrinsic tryptophan fluorescence spectroscopy (IFS) is a standard non-radioactive tool for studying conformational changes that occur upon nucleotide binding to small GTPases51,52,53,54. The conformational changes associated with the binding of a guanine nucleotide to PmRab7 shift the environments around three conserved Trp residues (W62, W102, W142) and alter the fluorescence emission spectra of the protein52,55 (Fig. 1).
To examine how the fluorescence emission spectra of PmRab7 are modulated by the binding of guanine nucleotides, the spectra of PmRab7-WT bound to either GDP or a non-hydrolyzable GTP analog, GppNHp, were compared to that of the nucleotide-free counterpart. The emission spectrum of the nucleotide-free enzyme was used to normalize the spectra of the protein in the presence of other guanine nucleotides or analogs (Fig. 3a). In the nucleotide-free condition, the maximum emission wavelength (λmax) of the Trp residues was 342 nm; the fluorescence intensity of this condition was set to 100%. The λmax in the presence of GDP occurred at 342 nm with 115% fluorescence. In contrast, the λmax of the GppNHp-bound conformation, which mimics the GTP-bound conformation, exhibited a blue shift to 338 nm with 91% fluorescence. These observations are consistent with previous reports that documented the migration of the λmax of the GTP bound conformation to a shorter wavelength compared to that of the GDP bound form, along with an increase in emission intensity at λmax in the GDP-bound conformation but a reduction in emission intensity at λmax in GTP-bound state52,53,54.
Fluorescence emission spectra of PmRab7 mutants were compared to the fluorescence profiles of PmRab7-WT to validate their dominant nucleotide binding state. Emission spectra of 1.5 µM PmRab7 WT (a), T22N (b), Q67L (c), and L129F (d) in the nucleotide-free (nf) state (dash) and after the addition of 25 µM GDP (grey) or GppNHp (black) at 37 ºC for 2 h.
The nucleotide-binding properties of PmRab7 mutants were validated using IFS
An investigation into the emission spectra of the PmRab7 mutants in the presence of various nucleotides was used to validate the nucleotide-binding nature of the mutants. To verify that the T22N mutant represented a GDP-bound conformation, we hypothesized that the mutant should display an increase in fluorescence intensity at λmax in the presence of GDP relative to that of the nucleotide-free form as in WT. The results indeed showed that, compared to PmRab7-WT (Table 1), the fluorescence emission at the λmax for the T22N mutant increased by 17% in the presence of GDP. (Fig. 3a and b). The increase in the fluorescence intensity of T22N in the presence of GDP implied that T22N adopted a similar conformation to the GDP-bound WT (Fig. 3b). Unexpectedly, a 20% rise in emission was observed in the GppNHp-bound form compared to the nucleotide-free counterpart. A previous study reported that T22N can bind to GTP, albeit with a lower affinity than GDP35,56. We hypothesized that the gain in the emission came from the binding of GTP to the GDP-bound conformation of T22N. Additionally, high concentrations of the GTP analog may overcome the weaker binding affinity of the T22N mutant, leading to detectable binding in our assays. Overall, these data underscored that PmRab7-T22N in this study adopted the GDP-bound conformation in the presence of guanine nucleotides.
As the Q67L mutant was designed based on the constitutively active GTP-bound conformation, it was hypothesized that the enzyme should exhibit a lower fluorescence intensity in the presence of GppNHp relative to the nucleotide-free as observed in WT, while in the presence of GDP, the fluorescence intensity should be slightly increased or comparable to the nucleotide-free condition (Fig. 3c). As previously discussed, a blue shift in λmax is characteristic of the GTP-bound conformation. Compared to other forms of PmRab7 in this study, the λmax of Q67L showed the largest blue shift of 6 nm. Compared to the nucleotide-free form of Q67L, the relative emission at λmax increased by 4% in the GDP-bound form, whereas a 12% decrease was observed when bound to GppNHp. Together, the result indicated that the PmRab7-Q67L protein produced in this study preferably binds to only GppNHp as previously reported.
The L129F mutant has been reported to bind to both GDP and GTP despite an increase in nucleotide exchange rate38. Therefore, we hypothesized that the fluorescence spectra of this mutant would be similar to WT which is capable of binding to both nucleotides. Compared to the fluorescence profile of other forms of PmRab7, the spectra of L129F exhibited the smallest blue shift of 1 nm when GDP is replaced by GppNHp (Table 1; Fig. 3d). As expected, when Rab7 is bound to GDP, the relative fluorescence intensity at λmax experienced a 14% increase relative to the nucleotide-free enzyme in the GDP-bound form as in the WT protein. Unlike in the spectrum of WT where a drop in emission was seen upon GppNHp association, there was a 2% increase in the GppNHp spectrum of L129F when compared to that of the nucleotide-free form. Since the steric hindrance, Leu-to-Phe switch at residue 129 made the bottom of the nucleotide binding pocket shallower38, it was speculated that while the mutant retains the capacity to bind both GDP and GTP, the binding to GTP might be compromised38. Accordingly, the high dissociation rate with GTP may account for the similar pattern between the GppNHp-bound and nucleotide-free spectra of L129F.
VP28 binds to both GDP- and GTP-bound forms of PmRab7
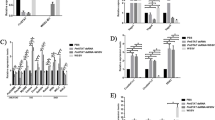
To determine whether VP28 prefers any particular nucleotide binding states of PmRab7, we performed ELISA by coating the 4 different recombinant PmRab7 onto a microplate and incubating them with purified VP28 (Fig. 4). The optical densities at 450 nm (OD450) of the PmRab7-VP28 interaction were subtracted with a non-specific binding between BSA-VP28. The purified VP28 protein exhibited equivalent binding efficiency for all PmRab7 conformations, suggesting that the VP28 protein binds both the GDP- and GTP-bound forms of PmRab7. Using ELISA with purified WSSV particles as a natural binding partner of PmRab7, similar results were observed (Fig. 5). Based on ELISA, VP28 could interact with either the GDP- or GTP-bound form of PmRab7.
The binding affinity of VP28 for PmRab7-WT is weakened by an increase in temperature
To gain a quantitative insight into the PmRab7-VP28 interaction, we hypothesized that an isothermal titration calorimetry (ITC) is a suitable technique for investigation. To test this hypothesis, we took advantage of the water temperature effect on WSSV infectivity in shrimp57,58,59,60. It has been reported that permissive temperatures between 16 ºC and 31 ºC allow WSSV replication in susceptible hosts59 with 25 ºC being the most favorable temperature57. In contrast, higher water temperatures (32–33 ºC) abate WSSV replication in shrimp and significantly curtail shrimp mortality from WSSV infection60. Accordingly, we hypothesized that lowering temperature strengthens the affinity between PmRab7 and VP28. Therefore, the thermodynamic parameters of PmRab7-WT and VP28 were investigated at the permissible temperature, 25 ºC, and the inhibitory temperature, 33 ºC.
At 25 ºC, the binding between PmRab7-WT and VP28 exhibited a dissociation constant (KD) of 22.5 nM (Table 2; Fig. 6a) which is derived from an enthalpy change (ΔH) of -22.1 kcal mol-1 and a free energy change (ΔG) of -10.4 kcal mol-1, consistent with the interaction being spontaneous. An entropy change (-TΔS) of 11.7 kcal mol-1 indicated that the association is accompanied by favorable hydrophobic interactions during conformational changes. As the temperature increased, the dissociation constant (KD) dropped by 16-fold from 22.5 nM at 25 ºC to 352 nM at 33 ºC, while the free energy of the interaction (ΔG) increased to -9.4 kcal mol-1 (Table 2; Figs. 7 and 6a).
The titration of VP28 to PmRab7-WT using isothermal titration calorimetry (ITC) showed the strongest interaction when compared to T22N, Q67L, and L129F mutants. ITC titration curve and its corresponding fitting curve of 200 µM of VP28 to 20 µM of PmRab7-WT (a), T22N (b), Q67L (c), and L129F (d) in 1X PBS with 5% glycerol at 25 ºC.
The thermodynamic parameters at 33 ºC compared to 25 ºC showed that the binding of PmRab7-WT and VP28 favorably occurs at low temperatures. (a) ITC titration curve (Above) and its corresponding fitting curve (Below) of 200 µM of VP28 to 20 µM of PmRab7-WT in 1X PBS with 5% glycerol at 33 ºC. (b) Comparison of the free energy change (ΔG), enthalpy change (ΔH), and entropy change (expressed as -TΔS) in the titration of VP28 to PmRab7-WT with two different temperatures at 25 ºC and 33 ºC.
Our result suggests that the weakened interaction between PmRab7, one of the WSSV receptors in shrimp, and VP28 constitutes one of the underlying reasons why WSSV becomes less virulent at higher temperatures. Furthermore, the result indicates that the ITC can be used to study the relative binding affinity between VP28 and different PmRab7 constructs.
VP28 binds with the highest affinity with PmRab7-WT
When compared to the dissociation constants of other PmRab7 conformations with VP28, the affinity of VP28 found PmRab7-WT was the strongest. As shown in Table 2; Fig. 6, the affinity of VP28 for PmRab7-L129F, PmRab7-T22N, and PmRab7-Q67L showed a 6-, 15- and 88-fold decrease in comparison to that of PmRab7-WT. Based on our findings, it is concluded that the best partner for VP28 binding is PmRab7-WT.
The nucleotide binding of PmRab7 enhances the PmRab7-VP28 affinity
As the WT enzyme exhibits the strongest binding affinity, we further hypothesized whether VP28 would prefer a particular nucleotide binding state of the wild-type enzyme. Accordingly, the ITC experiments between PmRab7-WT and VP28 in the presence of excess nucleotides were performed (Table 3; Fig. 8).
The titration of VP28 to PmRab7-WT in the presence of 0.5 mM nucleotide using ITC showed the addition of excess nucleotide (GDP or GTP) enhances the binding affinity. Isothermal titration calorimetry (ITC) titration curve and its corresponding fitting curve of 200 µM of VP28 to 20 µM of PmRab7-WT (a), PmRab7-WT + GDP (b), PmRab7-WT + GTP (c) in 50 mM Tris-HCl pH 8.0, 100 mM NaCl and 5% glycerol with 0.5 mM nucleotide at 25 ºC.
To measure the binding thermodynamics, excess nucleotide, either GDP or GTP, was added to the binding reaction to shift the population of the PmRab7-WT to the desired nucleotide binding state. In this experiment, a Tris-HCl-based buffer was used instead of a PBS-based buffer that was used in Table 3 because nucleotide binding reactions were commonly studied in this buffer48,61. The results showed that in the absence of excess nucleotide (Fig. 8a; the KD of 1920 nM), the binding affinity of VP28 and PmRab7-WT in the presence of co-purified nucleotides was slightly weaker by a 2-fold in comparison to that of in the presence of excess GDP and GTP (Fig. 8b and c; the KD of 924 nM and 826 nM, respectively). However, the nature of the excess nucleotide has no significant effect on the affinity. Either nucleotides presented in excess boosts the protein-protein interaction.
Discussion
The nucleotide binding states of PmRab7-WT and mutants could be characterized by the change in the fluorescence emission spectra
The fluorescence emission spectra of PmRab7-WT in the presence of guanine nucleotides are consistent with previous studies on human Rab5 and Rab751,52,53. Compared to the nucleotide-free state, the binding of GDP leads to an increase in fluorescence emission. On the contrary, the uptake of GTP or its analog, such as GppNHp, lowers the emission52,53.
The fluctuation in the fluorescence emission and the shift of the emission maximum is due to the change in the environment of Trp residues51. When Trp residues are in a more hydrophilic environment, such as solvent exposure, their fluorescence is more strongly quenched, hence lower intensity. Based on the crystallographic structures of Rab7-WT bound to GDP (PDB ID: 1VG1) and GppNHp (PDB ID: 1VG8), the solvent-accessible surface area (SASA) of Trp residues in the GDP-bound complex is greater than that of the GppNHp-bound structure, thereby accounting for the decrease in fluorescence intensity of GppNHp-bound state.
The thermodynamics of PmRab7-VP28 interaction is consistent with the high virulence of WSSV at low temperatures
When increasing the temperature to 33 ºC, the dissociation constant rose by 16-fold and the interaction became less entropically favorable as indicated by the more negative ΔS. (Table 2; Fig. 7). We proposed that the high temperature (33 ºC) mitigates WSSV virulence by diminishing the affinity between VP28 and PmRab7, and thereby partially hindering the viral entry. This correlates with previous reports that found a higher WSSV copy number at a lower temperature (16–31 ºC)57,59,60,62,63. However, the debilitating virulence is also attributable to the expression of other viral inhibitory factors such as aldehyde dehydrogenase (ALDH) and heat shock protein 70 (Hsp70) at higher temperature64. Thus, our finding indicated that the negative correlation between the VP28-PmRab7 affinity and water temperature may be one of the reasons why lower water temperature favors WSSV infection57,58,59,60.
PmRab7-WT is the best form of PmRab7 as an anti-WSSV agent
The ELISA and ITC results consistently showed that VP28 indiscriminately interacted with both GDP- and GTP-bound forms of PmRab7, albeit with differing affinity. Our analysis revealed that the PmRab7 conformations that adopted both GTP- and GDP-bound states (WT and L129F) interacted by VP28 with a higher affinity than those that were locked in either GDP-bound only (T22N) or GTP-bound only (Q67L) conformation. From an evolutionary standpoint, the lack of preference for any particular nucleotide binding PmRab7 states for VP28 binding may be beneficial for WSSV during viral entry as it increases the population of PmRab7 with which the virions can interact, thereby accelerating viral trafficking11,12.
When PmRab7 proteins were mutated to exclusively contain either GDP or GTP-bound conformation, their affinity drastically decreased by 15 and 88-found for the GDP-bound T22N mutant and the GTP-bound Q67L mutant, respectively. We hypothesize that the markedly weakened interaction suggested that the conformation flexibility that enables PmRab7 to switch between the two nucleotide binding states positively influences the VP28-PmRab7 interaction. The support for this hypothesis comes from a prior study, which found that the competitive inhibitor CID106770, targeting nucleotide binding pocket located in PmRab7, resulted in a reduction in the internalization of WSSV virions65. It was proposed in that investigation that the binding of competitive inhibitor results in the conformational rigidity that made PmRab7 incompatible with VP28 interaction.
Additionally, the 88-fold decrease in binding affinity of Q67L is supported by the molecular dynamics of the PmRab7-VP28 complex21. It was predicted that E68 and R69 on the surface of PmRab7 may play a crucial role in the PmRab7-VP28 interaction21. The vicinity of these two residues to the Q67L suggests that mutating Q67 may alter the interface for VP28-PmRab7 interaction, thus weakening the binding with VP28.
Nucleotide binding enhances the VP28-PmRab7 interaction
Knowing that the WT conformation is the best candidate, we asked whether any particular nucleotide binding states of the WT displayed a stronger affinity. Hence, excess nucleotides were added as shown in Table 3. The affinity of the interaction is strengthened by the presence of both excess nucleotides. This phenomenon is also observed in other pairs of Rab-binding partner interactions42,47,48,49,50. Consistent with our finding, a previous study showed that blocking the Rab7 nucleotide binding pocket by a small molecule inhibitor decreases the WSSV internalization65.
The observation that the presence of a small molecule, such as nucleotides, can modulate the affinity between the two proteins opens up an opportunity for shrimp-feed producing company to develop cost-effective small molecules that target the VP28-PmRab7 as a novel strategy to alleviate the economic loss by WSSV outbreak.
Applying the current ITC platform for high-throughput screening for inhibitors of the VP28-PmRab7 interaction
Previous research has demonstrated that recombinant PmRab7 and VP28 proteins can be orally administered to protect shrimp from WSSV infection25,67, under the hypothesis that these recombinant proteins compete with endogenous proteins for binding sites15. In our study, we found that the presence of excess guanine nucleotides significantly strengthens the interaction between recombinant PmRab7 and VP28 proteins. This suggests that adding guanine nucleotide analogs to shrimp feed, along with either recombinant PmRab7 or VP28, could enhance this competitive binding, thereby reducing WSSV infection.
However, the practical use of guanine nucleotides in shrimp farming is currently limited due to their high cost. Designing cost-effective nucleotide analogs could offer a viable solution. Agro-pharmaceutical companies, which often have extensive libraries of candidate compounds, have traditionally relied on labor-intensive and time-consuming bioassays in aquaculture to test the efficacy of these additives against WSSV25,67. Integrating ITC into the screening process would offer a more economical and high-throughput alternative, as ITC is widely used in the pharmaceutical industry for drug design and screening.
While KD values obtained through ITC may not directly reflect absolute binding affinities due to variations in pH and salt concentrations that affect the thermodynamic parameters of biochemical interactions66, the relative KD values derived from this method are sufficient for identifying potential small molecule inhibitors. This ITC-based approach could streamline the development of effective anti-WSSV agents for shrimp feed additives, benefiting not only shrimp farming but also paving the way for combating other viral diseases in aquaculture.
Conclusion
In conclusion, our biochemical investigation into the interaction between VP28 and PmRab7-WT, alongside its mutants, has provided valuable insights into the influence of PmRab7’s nucleotide binding state on its interaction with VP28. Our findings indicate that VP28 interacts with both GDP- and GTP-bound forms of PmRab7, with the strongest affinity observed for the wild-type. We also emphasize that the low temperature and addition of excess nucleotide enhance the binding of VP28. Moreover, we developed the ITC platform for screening potential anti-WSSV feed additives on an industrial scale.
Materials and methods
Materials
The expression vector pET28a was from Novagen Merck Millipore (Watford, UK). Restriction endonucleases (NcoI, NdeI, XbaI, and NotI) were from New England Biolabs (NEB) (Hitchin, UK). Chemically competent bacterial E. coli C41(DE3) cells were purchased from Merck Millipore (Darmstadt, Germany). LB media and LB agar were obtained from Merck Millipore (Darmstadt, Germany) and Kanamycin from Sigma-Aldrich (USA). Isopropyl β-D-1-thiogalactopyranoside (IPTG) was from Sigma-Aldrich (USA), PMSF, DNase I, and Lysozyme from hen egg from Sigma-Aldrich (USA).
Plasmid preparation
A recombinant pET28a-VP28-His6 plasmid was cloned by inserting the VP28 sequence15 between NcoI and XhoI of the pET28a plasmid. A recombinant pET28a-GST-PmRab7-WT was cloned by fusing the glutathione-S-transference sequence with PmRab715 and inserting it between NdeI and NotI of the pET28a plasmid. The PmRab7 mutants were constructed by a QuikChange mutagenesis using primers shown in Table 4. A TEV protease recognition site was inserted between GST and PmRab7 in the aforementioned plasmids for the QuikChange mutagenesis. All plasmid constructs were verified by DNA sequencing analysis.
Protein expression and purification of VP28
To express VP28-His6 proteins, the recombinant plasmid was transformed into the C41(DE3) Escherichia coli strain to overexpress. A glycerol stock containing the recombinant VP28 plasmid in the E. coli strain was inoculated in 10 ml of Luria-Bertani (LB) medium with 100 µg/ml kanamycin. The culture was grown overnight at 250 rpm and 37 ºC. The overnight culture was diluted to OD600 of 0.1 in LB media supplemented with 100 µg/ml kanamycin. The resulting culture was constantly shaken at 250 rpm and 37 ºC until the OD600 reached 0.6–0.8. Then, the protein expression was induced by adding 0.1 mM of IPTG at 30 ºC for 6 h. After the induction, the cells were harvested by centrifugation at 8000 ×g and 4 ºC for 5 min.
The pellet was resuspended with a lysis buffer (50 mM NaH2PO4 pH 8.0, 300 mM NaCl, 1% Triton X-100, 5% glycerol, 10 mM imidazole, 5 mM 2-βME, 1 mg/ml lysozyme, and 0.1 mM PMSF) and lysed by sonication with 15 min of 5 s pulse-on, 9 s pulse-off at a 40% amplitude at 4 ºC for 15 min (VCX 130 SONICS Vibra-Cell Ultrasonic Liquid Processor, Sonics). The lysate was centrifuged at 8000 ×g, 4 ºC for 15 min. The supernatant was subsequently loaded onto a polypropylene column (Qiagen, Germany) containing Ni sepharose (GE Healthcare, US). The soluble protein and beads were mixed by rocking at 4 ºC for 1 h. After that, the Ni-NTA column was washed with 10 column volumes (CV) of a wash buffer (50 mM NaH2PO4 pH 8.0, 500 mM NaCl, 1% Triton X-100, 5% glycerol, 10 mM imidazole, and 5 mM 2-βME). The VP28 protein was eluted with 10 CVs of an elution buffer (50 mM NaH2PO4 pH 8.0, 500 mM NaCl, 1% Triton X-100, 5% glycerol, 250 mM imidazole). The fractions containing the target protein were analyzed by SDS-PAGE followed by dialysis against 1X PBS pH 7.4 with 5% glycerol. The concentration of purified VP28 was measured by Bradford assay.
Protein expression and purification of PmRab7
The recombinant plasmids were transformed into the C41(DE3) Escherichia coli strain to overexpress the GST-PmRab7 proteins. A glycerol stock containing the recombinant PmRab7 plasmid in the E. coli strain was inoculated in 10 ml of Luria-Bertani (LB) medium with 100 µg/ml kanamycin. The culture was grown overnight at 250 rpm and 37 ºC. The overnight culture was diluted to OD600 of 0.1 in LB media supplemented with 100 µg/ml kanamycin. The resulting culture was constantly shaken at 250 rpm and 37 ºC until the OD600 reached 0.6–0.8. Then, the protein expression was induced by adding 0.1 mM of IPTG at 37 ºC for 4 h. After the induction, the cells were harvested by centrifugation at 8000 ×g, 4 ºC for 5 min.
The pellet was resuspended with a lysis buffer (1X PBS pH 7.4, 1 mM DTT, 5% glycerol, 1 mM EDTA, 1 mg/ml lysozyme, and 0.1 mM PMSF) and lysed by sonication as in the VP28 purification protocol. The lysate was centrifuged at 8000 ×g and 4 ºC for 15 min. The supernatant was subsequently loaded to a polypropylene column (Qiagen) containing Glutathione sepharose 4B resins (GE Healthcare). The soluble protein and beads were mixed by rocking at 4 ºC for 1 h. After that, the GST column was washed with 10 CVs of a wash buffer (1X PBS pH 7.4, 1 mM DTT, 5% glycerol, and 1 mM EDTA). The GST tag on the target protein was removed by an on-column cleavage using TEV proteases (0.2 mg of TEV per 2 g of wet cell weight) in 1 CV of the wash buffer at 4 ºC for 18 h. Fractions of flow-through (FT) and 2 CVs of wash buffer after digestion were collected. The fractions containing the target protein were analyzed by SDS-PAGE.
The protein was diluted 20-fold in a diluting buffer (50 mM Tris-HCl pH 8.0, 5% Glycerol, 2mM DTT, 100 mM NaCl) and further purified by anion exchange chromatography using a 5 ml HiTrap Q HP column (GE Healthcare). The protein purification was performed in a fast-protein liquid chromatography (FPLC) machine (ÄKTA start, GE Healthcare) at 4 ºC. The affinity column was washed with 5 CVs of a start buffer (50 mM Tris-HCl pH 8.0, 5% glycerol, and 2 mM DTT). Then, the protein was eluted with 6 CVs of an elution buffer (50 mM Tris-HCl pH 8.0, 5% glycerol, 2 mM DTT, and 200 mM NaCl). The fractions containing the target protein were pooled, concentrated, and dialyzed against 1X PBS pH 7.4 with 5% glycerol at 4 ºC for 8 h. The purified protein was analyzed by SDS-PAGE. The concentration of purified PmRab7 variants were measured by Bradford assay.
Intrinsic tryptophan fluorescence spectroscopy (IFS)
Intrinsic tryptophan fluorescence emission spectra were recorded using a spectrofluorophotometer (RF-5301PC, Shimadzu). In each measurement, 0.5 µM of PmRab7 was mixed with either 25 µM of GDP or GppNHp in a nucleotide binding buffer (40 mM HEPES pH 7.5, 10 mM MgCl2, and 2 mM DTT). The reactions were incubated for 2 h at 37 ºC before fluorescence emission measurements52,53. The excitation wavelength was set to 290 nm according to Pan et al., 1995 and Simon et al., 199652,53. The emission spectra were collected at 25 ºC between 305 and 450 nm. The collected data were normalized by the emission spectra of the nucleotide binding buffer in the absence and presence of guanine nucleotide. The nucleotide-free form was used to normalize the guanine nucleotide binding conditions.
Nucleotide-free Rab7 preparation
A nucleotide-free PmRab7 was used as a normalization in an intrinsic tryptophan fluorescence spectroscopy technique. To prepare nucleotide-free PmRab7, co-purified guanine nucleotides were extracted from the purified PmRab7 proteins according to the modified method from Res et al., 1990; Simon et al., 199652,68. First, tightly bound GDP was removed by chelating excess Mg2+ with 10 mM EDTA for 30 min at room temperature to accelerate the rate of nucleotide dissociation. Then, excess EDTA was removed using a PD-10 G-25 spin column to avoid inhibition of subsequent enzymatic reactions. After that, a 10-fold excess of GppCH2p was added to the reaction in the presence of 200 mM (NH4)2SO4 and incubated for 1 h at 22 ºC to promote GDP release from the protein. Antarctic Phosphatase (5U/mg) (NEB) was added to the reaction and incubated for 24 h at 22 ºC to hydrolyze GTP to GDP. Then, phosphodiesterase (5U/mg) was added and continued the incubation for 24 h to completely degrade the remaining GDP to guanosine. Finally, the reactions were shock-frozen and kept at -80 ºC. Before use, the free nucleotides were removed by centrifugation. The reactions were monitored using a fluorescence spectrophotometer (RF-5301PC, Shimadzu).
Enzyme-linked immunosorbent assay (ELISA)
For the ELISA experiment between recombinant VP28 and PmRab7, ninety-six well flat-bottom polystyrene ELISA plates were coated and incubated overnight at 4 ºC with 200 µl/well of 50 µg of purified PmRab7-WT or PmRab7 mutants or bovine serum albumin (BSA). All proteins were diluted with a coating buffer (0.05 M carbonate-bicarbonate buffer pH 9.6). Then, the coated plates were washed 4 times with 200 µl/well of a wash buffer (0.05% Tween-20 in 1X PBS pH 7.4). After that, the coated plates were blocked with 200 µl/well of a blocking buffer (10% fetal bovine serum (Gibco) in 1X PBS) for 2 h at room temperature to minimize non-specific binding. The plates were washed 3 times with the wash buffer followed by the addition of 200 µl/well of 100 µg VP28-His6 diluted with 1x PBS. After incubating the plates for 1 h at room temperature, the plates were washed 5 times with the wash buffer. The PmRab7-VP28 interaction was detected by adding 100 µl/well of rabbit anti-VP28 antibody diluted 1:1000 with 3% skim milk in 1X PBS and incubated for 1.5 h at room temperature. After 6 washes with the wash buffer, the plates were added with 100 µl/well of goat anti-rabbit (GAR)-HRP conjugate diluted 1:2000 with 3% skim milk in 1X PBS as a secondary antibody and incubated for 1 h at room temperature. The plates were washed 6 times as a final washing step, then each well was incubated with 50 µl of TMB solution (Merck) at room temperature on a gentle rocking. After 45 min, the color development was stopped by adding 50 µl/well of 2 M H2SO4. Absorbance was read at 450 nm using a microplate reader (Molecular Devices).
For the ELISA experiment between WSSV and PmRab7, the amounts of WSSV and PmRab7 were adapted from the previous study15. Ninety-six well flat-bottom polystyrene ELISA plates were coated and incubated overnight at 4 ºC with 200 µl/well of 0.5 × 104 copies of purified WSSV. Then, the coated plates were washed and blocked as in the ELISA experiment between recombinant VP28 and PmRab7. The plates were washed 3 times with the wash buffer followed by the addition of 200 µl/well of 10 µg PmRab7 conformations or BSA diluted with 1X PBS. After incubating the plates for 1 h at room temperature, the plates were washed 5 times with the wash buffer. The PmRab7-WSSV interaction was detected by adding 100 µl/well of rabbit anti-PmRab7 antibody diluted 1:1000 with 3% skim milk in 1X PBS and incubated for 1 h at room temperature. After 6 washes with the wash buffer, the plates were added with 100 µl/well of goat anti-rabbit (GAR)-HRP conjugate diluted 1:2000 with 3% skim milk in 1X PBS as a secondary antibody and incubated for 1 h at room temperature. The plates were washed 6 times as a final washing step, then each well was incubated with 50 µl of TMB substrate (Merck) at room temperature on a gentle rocking. After 10 min, the color development was stopped by adding 50 µl/well of 2 M H2SO4. Absorbance was read at 450 nm using a microplate reader (SpectraMax iD3 multi-mode microplate reader, molecular devices).
Isothermal titration calorimetry (ITC)
ITC experiments were performed in a buffer of 1X PBS pH 7.4 with 5% glycerol to verify which of the PmRab7 conformations binds VP28. PmRab7-WT and mutants were diluted at a concentration of 20 µM and then were titrated against a solution of 200 µM VP28-His6 using a MicroCal PEAQ-ITC calorimeter (Malvern Panalytical). The experiment parameters were set as follows: 19 times of injections, 0.2 µl of first injection volume, and 2 µl of sequential injection volume, with a spacing time of 120 s, at a reference temperature of 25 and 33 ºC. The heat from the initial injection was excluded from the data before analysis. Titration data were fitted with the one-site binding model using MicroCal PEAQ-ITC analysis software.
To validate the binding of PmRab7-WT and VP28 in the presence of excess nucleotides including GDP (Sigma-Aldrich) and GTP (Abcam), the buffers of purified PmRab7-WT and VP28 proteins were exchanged through a PD-10 column by eluting the column with 50 mM Tris-HCl pH 8.0, 100 mM NaCl and 5% glycerol prior to adding 0.5 mM the respective nucleotide before performing ITC experiments with the same parameters as above.
Data availability
All data generated or analyzed during this study are included in this published article and its Supplementary Information file.
References
FAO. The State of World Fisheries and Aquaculture 2020. FAO 32. FAO, (2020).
Thitamadee, S. et al. Review of current disease threats for cultivated penaeid shrimp in Asia. Aquaculture. 452, 69–87 (2016).
Subash, P. et al. White feces syndrome in Penaeus vannamei is potentially an Enterocytozoon hepatopenaei (EHP) associated pathobiome origin of Vibrio Spp. J. Invertebr Pathol. 198, 107932 (2023).
Munkongwongsiri, N. et al. Propionigenium and Vibrio species identified as possible component causes of shrimp white feces syndrome (WFS) associated with the microsporidian Enterocytozoon hepatopenaei. J. Invertebr Pathol. 192, 107784 (2022).
Qiu, W. et al. Toll receptor 2 (Toll2) positively regulates antibacterial immunity but promotes white spot syndrome virus (WSSV) infection in shrimp. Dev. Comp. Immunol. 115, 103878 (2021).
Leu, J. H. Springer Berlin Heidelberg, et al. Whispovirus. in Lesser Known Large dsDNA Viruses (ed. Van Etten, J. L.) 197–227 doi: (2009). https://doi.org/10.1007/978-3-540-68618-7_6
Tsai, J. M. et al. Identification of the nucleocapsid, tegument, and envelope proteins of the shrimp white spot syndrome virus virion. J. Virol. 80, 3021–3029 (2006).
Zhou, Q., Xu, L., Li, H., Qi, Y. P. & Yang, F. Four major envelope proteins of white spot syndrome virus bind to form a complex. J. Virol. 83, 4709–4712 (2009).
Sun, L., Su, Y., Zhao, Y., Fu, Z. Q. & Wu, Y. Crystal structure of major envelope protein VP24 from white spot syndrome virus. Sci. Rep. 6, 1–9 (2016).
Thomas, A., Sudheer, N. S., Kiron, V., Singh, B., Narayanan, R. B. & I. S. & Expression profile of key immune-related genes in Penaeus monodon juveniles after oral administration of recombinant envelope protein VP28 of white spot syndrome virus. Microb. Pathog. 96, 72–79 (2016).
Yi, G. et al. Vp28 of shrimp white spot syndrome virus is involved in the attachment and penetration into shrimp cells. J. Biochem. Mol. Biol. 37, 726–734 (2004).
Van Hulten, M. C. W., Witteveldt, J., Snippe, M. & Vlak, J. M. White spot syndrome virus envelope protein VP28 is involved in the systemic infection of shrimp. Virology. 285, 228–233 (2001).
Zhao, Z. Y. et al. A novel C-type lectin from the shrimp Litopenaeus vannamei possesses anti-white spot syndrome virus activity. J. Virol. 83, 347–356 (2009).
Wang, X. W., Xu, Y. H., Xu, J. D., Zhao, X. F. & Wang, J. X. Collaboration between a soluble C-type lectin and Calreticulin facilitates white spot syndrome virus infection in shrimp. J. Immunol. 193, 2106–2117 (2014).
Sritunyalucksana, K., Wannapapho, W., Lo, C. F. & Flegel, T. W. PmRab7 is a VP28-binding protein involved in white spot syndrome virus infection in shrimp. J. Virol. 80, 10734–10742 (2006).
Xu, H. et al. The interaction of white spot syndrome virus envelope protein VP28 with shrimp Hsc70 is specific and ATP-dependent. Fish. Shellfish Immunol. 26, 414–421 (2009).
Sanitt, P., Attasart, P. & Panyim, S. Protection of yellow head virus infection in shrimp by feeding of bacteria expressing dsRNAs. J. Biotechnol. 179, 26–31 (2014).
Ongvarrasopone, C., Chanasakulniyom, M., Sritunyalucksana, K. & Panyim, S. Suppression of PmRab7 by dsRNA inhibits WSSV or YHV infection in shrimp. Mar. Biotechnol. 10, 374–381 (2008).
Ongvarrasopone, C., Saejia, P., Chanasakulniyom, M. & Panyim, S. Inhibition of Taura syndrome virus replication in Litopenaeus vannamei through silencing the LvRab7 gene using double-stranded RNA. Arch. Virol. 156, 1117–1123 (2011).
Ongvarrasopone, C., Chomchay, E. & Panyim, S. Antiviral effect of PmRab7 knock-down on inhibition of Laem-Singh virus replication in black tiger shrimp. Antiviral Res. 88, 116–118 (2010).
Verma, A. K. et al. Interaction between shrimp and white spot syndrome virus through PmRab7-VP28 complex: an insight using simulation and docking studies. J. Mol. Model. 19, 1285–1294 (2013).
Attasart, P. et al. Inhibition of white spot syndrome virus replication in Penaeus monodon by combined silencing of viral rr2 and shrimp PmRab7. Virus Res. 145, 127–133 (2009).
Thagun, C., Srisala, J., Sritunyalucksana, K., Narangajavana, J. & Sojikul, P. Arabidopsis-derived shrimp viral-binding protein, PmRab7 can protect white spot syndrome virus infection in shrimp. J. Biotechnol. 161, 60–67 (2012).
Jupatanakul, N., Flegel, T. W., Withyachumnarnkul, B. & Sritunyalucksana, K. Preparation and application of recombinant PmRab7 in shrimp farm. in Proceedings of the 48th Kasetsart University Annual Conference, Kasetsart, 3–5 March, 2010. Subject: FisheriesKasetsart University, (2010).
Ananphongmanee, V. et al. Shrimp protected from a virus by feed containing yeast with a surface-displayed viral binding protein. J. Biotechnol. 342, 45–53 (2021).
Guerra, F. & Bucci, C. Multiple Roles of the Small GTPase Rab7. Cells 5, 34 (2016).
Goody, R. S., Müller, M. P. & Wu, Y. W. Mechanisms of action of Rab proteins, key regulators of intracellular vesicular transport. Biol. Chem. 398, 565–575 (2017).
Gibbs, J. B., Sigal, I. S., Poe, M. & Scolnick, E. M. Intrinsic GTPase activity distinguishes normal and oncogenic ras p21 molecules. Proc. Natl. Acad. Sci. U. S. A. 81, 5704–5708 (1984).
Aoki, Y. et al. Germline mutations in HRAS proto-oncogene cause Costello syndrome. Nat. Genet. 37, 1038–1040 (2005).
Niihori, T. et al. Germline KRAS and BRAF mutations in cardio-facio-cutaneous syndrome. Nat. Genet. 38, 294–296 (2006).
Nassar, N., Singh, K. & Garcia-Diaz, M. Structure of the dominant negative S17N mutant of Ras. Biochemistry. 49, 1970–1974 (2010).
Buczynski, G., Bush, J., Zhang, L., Rodriguez-Paris, J. & Cardelli, J. Evidence for a recycling role for Rab7 in regulating a late step in endocytosis and in retention of lysosomal enzymes in Dictyostelium Discoideum. Mol. Biol. Cell. 8, 1343–1360 (1997).
Gutierrez, M. G., Munafó, D. B., Berón, W. & Colombo, M. I. Rab7 is required for the normal progression of the autophagic pathway in mammalian cells. J. Cell. Sci. 117, 2687–2697 (2004).
Valencia, A., Chardin, P., Wittinghofer, A. & Sander, C. The ras protein family: evolutionary tree and role of conserved amino acids. Biochemistry. 30, 4637–4648 (1991).
Spinosa, M. R. et al. Functional characterization of Rab7 mutant proteins associated with Charcot-Marie-tooth type 2B disease. J. Neurosci. 28, 1640–1648 (2008).
Krengel, U. et al. Three-dimensional structures of H-ras p21 mutants: molecular basis for their inability to function as signal switch molecules. Cell. 62, 539–548 (1990).
Meresse, S., Gorvel, J. P. & Chavrier, P. The rab7 GTPase resides on a vesicular compartment connected to lysosomes. J. Cell. Sci. 108, 3349–3358 (1995).
McCray, B. A., Skordalakes, E. & Taylor, J. P. Disease mutations in Rab7 result in unregulated nucleotide exchange and inappropriate activation. Hum. Mol. Genet. 19, 1033–1047 (2010).
Frech, M. et al. Inhibition of GTPase activating protein stimulation of Ras-p21 GTPase by the Krev-1 gene product. Sci. (80-). 249, 169–171 (1990).
Herrmann, C., Martin, G. A. & Wittinghofer, A. Quantitative analysis of the complex between p21ras and the ras-binding domain of the human Raf-1 protein kinase. J. Biol. Chem. 270, 2901–2905 (1995).
Feig, L. A. & Cooper, G. M. Inhibition of NIH 3T3 cell proliferation by a mutant ras protein with preferential affinity for GDP. Mol. Cell. Biol. 8, 3235–3243 (1988).
Sun, Q., Westphal, W., Wong, K. N., Tan, I. & Zhong, Q. Rubicon controls endosome maturation as a Rab7 effector. Proc. Natl. Acad. Sci. U S A. 107, 19338–19343 (2010).
Cattin-Ortolá, J., Topalidou, I., Dosey, A., Merz, A. J. & Ailion, M. The dense-core vesicle maturation protein CCCP-1 binds RAB-2 and membranes through its C-terminal domain. Traffic. 18, 720–732 (2017).
Wang, Z. et al. The Vici syndrome protein EPG5 is a Rab7 effector that determines the fusion specificity of autophagosomes with late endosomes/lysosomes. Mol. Cell. 63, 781–795 (2016).
Shirane, M. & Nakayama, K. I. Protrudin induces neurite formation by directional membrane trafficking. Science. 314, 818–821 (2006).
Kimura, T. et al. The GDP-dependent Rab27a effector coronin 3 controls endocytosis of secretory membrane in insulin-secreting cell lines. J. Cell. Sci. 121, 3092–3098 (2008).
Gandhi, P. N., Wang, X., Zhu, X., Chen, S. G. & Wilson-Delfosse, A. L. The Roc domain of leucine-rich repeat kinase 2 is sufficient for interaction with microtubules. J. Neurosci. Res. 86, 1711–1720 (2008).
Ma, X. et al. A non-canonical GTPase interaction enables ORP1L-Rab7-RILP complex formation and late endosome positioning. J. Biol. Chem. 293, 14155–14164 (2018).
Burke, J. E. et al. Structures of PI4KIIIβ complexes show simultaneous recruitment of Rab11 and its effectors. Sci. (80-). 344, 1035–1038 (2014).
Stein, M. P., Feng, Y., Cooper, K. L., Welford, A. M. & Wandinger-Ness, A. Human VPS34 and p150 are Rab7 interacting partners. Traffic. 4, 754–771 (2003).
Brockhinke, A., Plessow, R., Kohse-Höinghaus, K. & Herrmann, C. Structural changes in the ras protein revealed by fluorescence spectroscopy. Phys. Chem. Chem. Phys. 5, 3498–3506 (2003).
Simon, I., Zerial, M. & Goody, R. S. Kinetics of interaction of Rab5 and Rab7 with nucleotides and magnesium ions. J. Biol. Chem. 271, 20470–20478 (1996).
Pan, J. Y., Sanford, J. C. & Wessling-Resnick, M. Effect of guanine nucleotide binding on the intrinsic tryptophan fluorescence properties of Rab5. J. Biol. Chem. 270, 24204–24208 (1995).
Sengottaiyan, P. et al. The intrinsic GTPase activity of the Gtr1 protein from Saccharomyces cerevisiae. BMC Biochem. 13, 1–8 (2012).
Ghisaidoobe, A. B. T. & Chung, S. J. Intrinsic tryptophan fluorescence in the detection and analysis of proteins: a focus on Förster resonance energy transfer techniques. Int. J. Mol. Sci. 15, 22518–22538 (2014).
John, J. et al. Kinetic and structural analysis of the Mg2+-binding site of the guanine nucleotide-binding protein p21(H-ras). J. Biol. Chem. 268, 923–929 (1993).
Thi Lua, D. & Hirono, I. Effect of low water temperature on the pathogenicity of white spot syndrome virus (WSSV) in kuruma shrimp (Marsupenaeus japonicus). J. Sci. Devel. 13, 1405–1414 (2015).
Escobedo-Bonilla, C. M. high water temperature affects WSSV management. Glob Aquac Advocate 74–75 (2006).
Wongmaneeprateep, S. et al. Effects of water temperature on the white spot syndrome virus infection in postlarvae Litopenaeus vannamei. Walailak J. Sci. Technol. 7, 127–134 (2011).
Rahman, M. M. et al. Effect of high water temperature (33°C) on the clinical and virological outcome of experimental infections with white spot syndrome virus (WSSV) in specific pathogen-free (SPF) Litopenaeus vannamei. Aquaculture 261, 842–849 (2006).
Tong, J., Tan, L., Chun, C. J. & Im, Y. J. Structural basis of human ORP1-Rab7 interaction for the late-endosome and lysosome targeting. PLoS One. 14, 1–13 (2019).
Granja, C. B., Aranguren, L. F., Vidal, O. M., Aragón, L. & Salazar, M. Does hyperthermia increase apoptosis in white spot syndrome virus (WSSV)-infected Litopenaeus vannamei? Dis. Aquat. Organ. 54, 73–78 (2003).
Granja, C. B., Vidal, O. M., Parra, G. & Salazar, M. Hyperthermia reduces viral load of white spot syndrome virus in Penaeus vannamei. Dis. Aquat. Organ. 68, 175–180 (2006).
Lin, Y. R. et al. The role of aldehyde dehydrogenase and Hsp70 in suppression of white spot syndrome virus replication at high temperature. J. Virol. 85, 3517–3525 (2011).
Kumar, A., Shekhar, S. & A, S. Role of PmRab7 regulation in WSSV infection and functional validation of small molecule as PmRab7 GTPase inhibitor. J. Aquac Res. Dev. 08, (2017).
Henderson, K. L., Boyles, D. K., Le, V. H., Lewis, E. A. & Emerson, J. P. ITC methods for assessing buffer/protein interactions from the perturbation of steady-state kinetics: a reactivity study of Homoprotocatechuate 2,3-dioxygenase. Methods Enzymol. 567, 257–278 (2016).
Taengchaiyaphum, S. et al. Vaccination with multimeric recombinant VP28 induces high protection against white spot syndrome virus in shrimp. Dev. Comp. Immunol. 76, (2017).
John, J. et al. Kinetics of interaction of nucleotides with nucleotide-free H-ras p21. Biochemistry. 29, 6058–6065 (1990).
Acknowledgements
PS is supported by the Science Achievement Scholarship of Thailand (SAST). DP acknowledges the funding support from the Thailand Research Fund (Grant No. RSA6280087). This project is funded by National Research Council of Thailand (NRCT): High-Potential Research Team Grant Program, Grant No. N42A650869 to KS. This project is additionally supported by Mahidol University (Fundamental Fund: fiscal year 2023 by National Science Research and Innovation Fund (NSRF)) (Grant No. FF-056/2566), and Mahidol University (Grant No. 191/2562).
Author information
Authors and Affiliations
Contributions
P.S. and O.I. designed the experiments, interpreted the data, and prepared the manuscript. P.S. performed the experiments. J.S. provided the WSSV stock and supervised the ELISA experiments. S.C. designed, interpreted, and supervised the ITC experiments. D.P., S.T., K.S., S.H., and O.I. supervised all experiments. P.S., D.P., S.T., .K.S, S.H., S.C., and O.I. edited and reviewed the manuscript.
Corresponding author
Ethics declarations
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Sudsat, P., Srisala, J., Pakotiprapha, D. et al. VP28 interacts with PmRab7 irrespective of its nucleotide state. Sci Rep 14, 27803 (2024). https://doi.org/10.1038/s41598-024-79310-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-79310-5
This article is cited by
-
VP39A is a novel host cellular binding protein of white spot syndrome virus and interacts with Na+, K+-ATPase alpha subunit
Journal of Oceanology and Limnology (2025)