Abstract
Multiple myeloma (MM) is a plasma cell (PC) malignancy characterized by cytogenetic abnormalities, such as t(11;14)(q13;q32), resulting in CCND1 overexpression. The rs9344 G allele within CCND1 is the most significant susceptibility allele for t(11;14). Sequencing data from 2 independent cohorts, CoMMpass (n = 698) and Mayo Clinic (n = 661), confirm the positive association between the G allele and t(11;14). Among 80% of individuals heterozygous for rs9344 with t(11;14), the t(11;14) event occurs on the G allele, demonstrating a biological preference for the G allele in t(11;14). Within t(11;14), the G allele is associated with higher CCND1 expression and elevated H3K27ac and H3K4me3. CRISPR/Cas9 mediated A to G conversion resulted in increased H3K27ac over CCND1 and elevated CCND1 expression. ENCODE ChIP-seq data supported a PAX5 binding site within the enhancer region covering rs9344, showing preferential binding to the G allele. Overexpression of PAX5 resulted in increased CCND1 expression. These results support the importance of rs9344 G enhancer in increasing CCND1 expression in MM.

Similar content being viewed by others
Introduction
Multiple myeloma (MM) is an incurable plasma cell (PC) malignancy characterized by cytogenetic abnormalities that drive clinical manifestations, response to treatment, and overall prognosis [1]. Translocation t(11;14)(q13;q32), leading to juxtaposition of the immunoglobulin heavy chain (IGH) near CCND1, is the most common IGH translocation in MM [1]. Interestingly, MM with t(11;14) is the only subtype with a known germline susceptibility locus at 11q13.3, as identified by genome-wide association studies (GWAS) [2]. The G allele of rs9344 (formerly rs603965), located within the last nucleotide of exon 4 of CCND1 (c.723 G > A, p.Pro241 = , NM_053056), has been reported to be the most significant and strongest germline risk factor identified in association with an increased risk of t(11;14) MM. The G allele of rs9344 has an observed odds ratio (OR) of 1.82 (p = 7.96 × 10−11) compared to healthy controls [2] and 1.95 (p = 2.07 × 10−11) compared to non-t(11;14) MM cases [2]. We previously confirmed this association in the Relating Clinical Outcomes in MM to Personal Assessment of Genetic Profile (CoMMpass) and Mayo Clinic cohorts [3]. Interestingly, rs9344 is not associated with an increased risk of other B-cell malignancies [4], like mantle cell lymphoma (MCL) [2], which is also associated with t(11;14), suggesting a unique role of rs9344 in the pathology of t(11;14)-driven MM. However, the molecular mechanism by which rs9344 influences the development of t(11;14) MM has remained unknown. Here, we utilized genetic, epigenetic, and gene-editing approaches to identify rs9344 as the likely causal SNP at 11q13.3. This study demonstrated the G allele of rs9344 along with the transcription factor PAX5 promotes CCND1 expression following t(11;14) in MM.
Materials and methods
Study cohorts
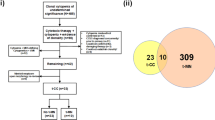
We performed a retrospective study of newly diagnosed MM patients from the publicly available international MM cohort, the Relating Clinical Outcomes in the CoMMpass study (NCT01454297) and a Mayo Clinic cohort. The CoMMPass study was approved by ethics committees or institutional review boards (IRB) at individual study sites and by the IRB at the Mayo Clinic. Samples were collected with written informed consent from the patients in accordance with ethical guidelines. The research involving human material and human data is in accordance with the Declaration of Helsinki. Patients with available clinical and demographic data, and with whole genome sequencing (WGS), whole exome sequencing (WES), and RNA sequencing (RNA-seq) data from samples collected at diagnosis in either cohort were selected. This study included 698 patients in the CoMMpass cohort and 661 in the Mayo cohort. MM-specific cytogenetic abnormalities were identified using FISH as previously described by Smadbeck et al. [5] in the Mayo cohort and by WGS in the CoMMpass cohort [6].
Sample collection, library construction, and sequencing
For the CoMMpass cohort, sample collection, DNA extraction, WES, RNA extraction, and RNA-seq were performed as previously described [7]. For the Mayo cohort, PCs were enriched using anti-CD138+ beads and stored as frozen pellets or in TRIzol. WES and RNA-seq were performed as described in supplementary methods.
WES and RNA-seq data analysis
Sequencing data from both cohorts, were analyzed as described in supplementary methods.
Chromatin immunoprecipitation and sequencing (ChIP-seq) and Assay for transposase-accessible chromatin using sequencing (ATAC-seq)
ChIP-seq and ATAC-seq using CD138+ MM tumor samples were performed following previously published methods [8, 9] with slight modifications, as described in supplementary methods.
Human myeloma cell lines and overexpression of PAX5
Overexpression of PAX5 was performed in U266B1 and KMS12PE cell lines. U266B1, a human MM cell line with t(11;14) carrying the rs9344 AG genotype, was obtained from the American Type Culture Collection (ATCC, Manassas, VA, USA) and was mycoplasma negative. The U266B1 cell line was authenticated using STR testing (Labcorp, Burlington, NC, USA). The KMS12PE human MM cell line with t(11;14) was obtained from Creative Bioarray (Shirley, NY, USA) was mycoplasma negative. The WT line carries the rs9344 AA genotype. Authentication of the KMS12PE cell line was performed using STR testing (Labcorp) and a customized PCR strategy to differentiate KMS12PE from KMS12BM. The substitution of the A allele by the G allele at rs9344 was performed by Synthego (Redwood City, CA, USA) using the CRISPR/Cas9 system [10] (see supplementary methods) and confirmed to be mycoplasma negative after CRISPR/Cas9 editing. The cell lines were cultured in RPMI-1640 (30-2001, ATCC) supplemented with 15% fetal bovine serum (FBS; 35-010-CV, Corning, Glendale, AZ) and 1% penicillin-streptomycin (15070063, Thermo Fischer). For PAX5 overexpression studies, the pLV-EGFP-hPAX5 overexpression plasmid (vector ID: VB900139-7742zer) and control pLV-EGFP plasmid (VB010000-9292bku) were purchased from Vector Builder (Chicago, IL, USA). Transfection, quantitative PCR and Western blotting was performed as described in supplementary methods.
Statistical analysis
Test of associations between rs9344 genotypes and expressed alleles, CCND1 and PAX5 expression including quantification of interaction effects was analyzed using generalized linear regression model (GLRM) as described in supplementary methods. For rs9344 genotypes and expressed alleles, overall survival analysis was performed using the Cox Proportional Hazards Regression model built into the Survival R package, with adjustment for age (continuous), sex (1 male, 0 female), and race (1 black, 0 non-black). A Logistic Regression model was used to test the associations with a binary outcome. The difference between two groups of continuous variables following normal distribution was tested using Student’s t test conditional on equal variance or the Welch test conditional on unequal variance. The Mann–Whitney U test was applied to data deviating from the normal distribution. The chi-square test was used to test the rate or ratio differences. All tests were two-sided, and a p < 0.05 in both cohorts was considered statistically significant. The analysis was performed using R software (v4.2.1).
Results
The G allele at rs9344 is preferentially associated with t(11;14) in MM
We and others have identified a positive association between the G allele of rs9344 and the t(11;14)(q13;q32) subtype of MM [2, 3]. We evaluated this association using 1,359 patients with MM from two large and independent cohorts, CoMMpass and Mayo Clinic. The distributions were similar for age, sex, rs9344 genotype, and frequency of t(11;14) between the two cohorts (p > 0.05) (Table 1).
Consistent with previous reports [2, 3], the G allele of rs9344 showed significant positive association with t(11;14) in both the CoMMpass (OR = 1.88, 95%CI:1.40–2.53, p = 3.2 × 10−5) and Mayo (OR = 1.54, 95%CI:1.17–2.04, p = 0.002) cohorts. In the t(11;14) group, 117 individuals were heterozygous for the rs9344 SNP. Among 80.34% of individuals (95%CI:64.92–98.32%, p = 4.2 × 10−15), t(11;14) involved the G allele based on RNA-seq data, demonstrating a ~ 4-fold enrichment relative to the A allele.
The G allele at rs9344 is associated with increased CCND1 gene expression
As previously reported [11], CCND1 gene expression was increased by ~40–50-fold in t(11;14) MM relative to non-t(11;14) MM (p = 1.3 × 10_13, Table 1). No other cytogenetic abnormalities significantly affected CCND1 expression (Supplementary Table 1.1). Given the positive association of the G allele at rs9344 with t(11;14)-derived MM and the association of t(11;14) with increased CCND1 expression, we investigated whether the G allele was associated with increased CCND1 gene expression in MM. Using a univariate analysis of the whole cohort, we identified a positive association of the G allele with CCND1 gene expression in CoMMpass (p = 3.0 × 10-5) and Mayo (p = 0.003) cohorts (Supplementary Table 1.2).
To further evaluate the unique association between the G allele at rs9344 and increased CCND1 gene expression in t(11;14)-derived MM, we investigated whether the G allele at rs9344 “interacts” with t(11;14) to further increase CCND1 expression. Following the concept of “gene-environment interactions” [12], we asked whether there exists an interplay between the rs9344 G risk allele, which stands for the “gene”, and the presence of t(11;14), which represents the “environment”, on CCND1 expression. For example, we asked whether the rs9344 G genotype increased CCND1 expression only in the presence of t(11;14). To test this possibility, we performed a GLRM including rs9344 genotype, cytogenetic abnormalities, and their interactions with adjustment for age, race and sex (Supplementary Table 1.3). Results from this model demonstrated the G allele (AG/GG genotypes) at rs9344 significantly increased CCND1 expression in combination with t(11;14) in both CoMMpass (p = 0.0053) and Mayo (p = 0.034) cohorts, but not in combination with other cytogenetic abnormalities (Supplementary Table 1.3–1.4). Consistent with these observations, the multivariate analysis without considering interactions failed to identify a significant association between the rs9344 genotype and CCND1 expression (Supplementary Table 1.1), further suggesting a lack of a main effect for rs9344 genotype on CCND1 expression.
We next evaluated the interaction separately in t(11;14) and non-t(11;14) cases. The AG and GG genotype groups had higher CCND1 expression than the AA group in both cohorts (Table 2). Importantly, in the non-t(11;14) group, the rs9344 genotype was not associated with increased CCND1 levels, as described above (Table 2). For GG genotype within the t(11;14) group, the interaction effect accounted for ~20% of CCND1 expression, which was higher than in the AG genotype group, further suggesting the existence of an interaction effect of the genotype with t(11;14) on CCND1 expression (Supplementary Table 1.5).
Gain of a super-enhancer at the risk locus in MM with t(11;14)
Given the increased expression of CCND1 in association with the G allele of rs9344 within t(11;14) MM, we evaluated whether rs9344 influenced the regulatory elements at the 11q13.3 locus. We first extracted variants in LD (r2 ≥ 0.5) with rs9344 based on 2504 individuals of European genetic ancestry (EUR) from the 1000 Genomes Project. The entire LD block included 11 variants with r2 ≥ 0.8, with minor allele frequency (MAF) > 0.41, covering a ~ 12.3 kb region [13] (Fig. 1A, Supplementary Figure 1in AFR), which is located within the same topologically associating domain based on Hi-C data [14, 15]. All variants in LD with rs9344 were in the intergenic or intronic regions of CCND1.
A LD zoom plot for rs9344 in a population with European genetic ancestry (EUR) from the 1000 Genomes Project. B Signal tracks for ATAC-seq and ChIP-seq of three histone modifications and PAX5. For the 28 MM cases with t(11;14) that had ChIP-seq or ATAC-seq data, only five samples being heterozygous for rs9344 were selected to display. All cases are displayed in Supplementary Fig. 4. In these heterozygous samples, at least one of the marks showed allelic imbalance at rs9344. At this locus, a total of 22 variants are linked to the index SNP rs9344 (r2 ≥ 0.5) in the EUR population. Twenty-two of the 23 variants reside in a 17-kb region, with the remaining SNP rs1231288366, which is not shown here, being located 30,607 bp away, downstream of CCND1. This SNP overlapped none of the epigenetic marks included in the study. Within the 17-kb region, 19 of the variants fell within regulatory regions (RE2 to RE6) in MM with t(11;14). Among them, 11 overlapped a broad H3K27ac peak region that was also enriched with H3K4me3 (RE3 to RE6), including ten in three open chromatin regions identified by ATAC-seq (E3, E5, and E6); the other eight variants were located in a broad H3K4me3 peak. In addition, rs9344 was in a PAX5 binding site in GM12878. Y-axis represents normalized signal in reads-per-million. ATAC-seq peak in the CCND1 promoter is boxed in gray. Regulatory regions carrying rs9344-linked variants are highlighted. RE, regulatory element. C Bar plot showing allelic imbalance at rs9344. The risk G allele was associated with significant increase of H3K27ac, H3K4me3, H3K4me1 and chromatin accessibility in one to five MM cases (Benjamini-Hochberg adjusted p value: 9.23E-03 – 7.33E-16, binomial test), except H3K4me3 in N23C (p = 7.81E-02). In GM12878, the G allele showed the tendency of association with increased PAX5 signal relative to the A allele (p = 1.09E-01). D Average chromatin signal profiles across the CCND1 locus. ChIP-seq data from 8 primary MM samples and ATAC-seq data from 28 primary MM samples, all with t(11;14), were represented in the top four panels and H3K27ac data from KMS12PE KI and WT were shown in (E). RPM, reads-per-million; A and G represent the expressed alleles at rs9344 based on RNA-seq data. Arrow indicates the position of rs9344. F Quantification of the H3K27ac signals from (E). G Relative mRNA expression of CCND1 in KMS12PE KI rs9344 GG clones compared to KMS12PE WT with rs9344 AA. The CT values of CCND1 and GAPDH mRNAs were determined in triplicate using real-time one step RT-qPCR quantification. The mRNA level of CCND1 was normalized to GAPDH. The normalized CT values in the KMS12PE KI clones was compared with KMS12PE WT using paired Student’s t test with α = 0.05. *P < 0.05, **P ≤ 0.01. H Transcription factor PAX5 binding motif. The G allele is adjacent to the PAX5 motif. Arrow indicates the index SNP rs9344.
Putative regulatory variants were identified by utilizing peaks from public ChIP-seq and ATAC-seq repositories of MM and healthy immune samples from the Roadmap Epigenomics Consortium (Supplementary Table 2) [16,17,18,19]. We identified 6 regulatory regions (RE1-RE6) within 17 kb that together covered 22 (from rs1683847 to rs7177) of the 23 variants with r2 ≥ 0.5 (Fig. 1B, Supplementary Table 3).
We compared the ChIP-seq and ATAC-seq profiles within 17 kb covering the 22 variants described above from 28 internal t(11;14) MM samples with healthy immune samples. Of the 6 regulatory regions (RE1-RE6, Fig. 1B), the healthy samples showed enrichment of the repressive mark H3K27me3 in RE1 and enrichment of H3K4me1 and H3K27me3 in RE2 and RE3, suggesting a poised chromatin state (Supplementary Fig. 2). In contrast, in 8 of the internal t(11;14) MM samples, this region was enriched with H3K4me3, H3K4me1, H3K27ac and ATAC-seq signals (Fig. 1B). Of the remaining 20 samples with only ATAC-seq data, all but 3 (N29, N49 and N15b) showed enrichment. To understand whether increased chromatin accessibility was t(11;14) specific, we analyzed additional publicly available ATAC-seq data from 21 MM patients and 5 MM cell lines [16]. The observed chromatin accessibility pattern appeared to be specific to t(11;14), as it was not identified in MM with other chromosomal abnormalities (Supplementary Fig. 3A).
Of the t(11;14) MM samples with increased chromatin accessibility, high levels of H3K27ac were observed (Fig. 1B and Supplementary Fig. 3A), suggesting that the CCND1 locus within t(11;14) formed a super-enhancer (SE) associated with high H3K27ac levels and high gene expression [20]. To test this possibility, we used the “Rank Ordering of Super-Enhancers” (ROSE) pipeline and identified SEs of 13.7 and 18.9 kb in size, respectively, in the t(11;14) samples N23C and N39C, covering the CCND1 gene (Supplementary Fig. 3B). To assess cell type specificity, we downloaded a collection of ROSE-identified SEs in 86 cell and tissue types [20]. This SE in N23C and N39C was not identified in any of the 24 healthy samples [20], indicating it was a de novo SE gained in MM with t(11;14). Based on the 111 reference epigenomes from the Roadmap Epigenomics Consortium [19], none of the immune cells had H3K27ac. Instead, most healthy samples showed both H3K4me1 and H3K27me3, suggesting that this region is a poised enhancer (Supplementary Fig. 2). To understand whether this SE was only present in MM with t(11;14), we called SEs as described above from 7 MM cases and 7 MM cell lines [18]. The 2 cell lines (U266B1 and KMS12BM) with t(11;14) had SEs (11.6-19.4 kb) in this region (Supplementary Figure 3C). The 2 MM cases with t(11;14) (MM3 and MM9) also showed broad H3K27ac, although below the SE calling cutoff. None of the 10 MM samples with other cytogenetic abnormalities had a SE in this region suggesting that this SE is highly cell type-specific and only detectable in MM with t(11;14), consistent with other studies [21,22,23]. Using both ChIP-seq and ATAC-seq data from 8 MM cases with t(11;14) (Supplementary Fig. 4), we identified the IGH SEs were inserted or translocated into the 144-577 kb upstream of CCND1 (Supplementary results, Supplementary Fig. 5).
The G allele of rs9344 altered the local chromatin state in MM with t(11;14)
Previous studies have demonstrated that coding exons can also function as active enhancers [24,25,26]. Hypothesizing that rs9344 plays a regulatory role through an epigenetic mechanism, we focused on RE5, which spans the rs9344 SNP. Of the 24 healthy samples, RE5 showed only weak H3K4me1 in CD19+ B and CD34+ hematopoietic stem/progenitor cells (Supplementary Fig. 2), suggesting that RE5 is a primed enhancer. In contrast, an ATAC-seq and H3K27ac peak in RE5 was observed in most of the t(11;14) MM cases (Fig. 1B, Supplementary Fig. 3A and Supplementary Fig. 4). None of the public data from non-t(11;14) MM had a peak in RE5 (Supplementary Fig. 3). Of the ATAC-seq data from the 28 internal MM cases with t(11;14), 2 had AA, 10 had GG, and 16 had the AG genotype at rs9344 (Supplementary Fig. 4). Four of the 15 samples with an expressed G allele in the AG genotype had a peak overlapping rs9344, and all four showed allelic imbalance, with the G allele associated with increased chromatin accessibility (p = 4.45 × 10−15–7.33 × 10−16, Fig. 1C and Supplementary Fig. 4). Of the 8 internal MM cases with t(11;14) with ChIP-seq data, those with GG or AG were more likely to have peaks covering rs9344 in comparison to AA samples, where no peaks for all three histone marks were found. Importantly, the peaks in the samples with the AG genotype all showed allelic imbalance, with the G allele associated with increased histone modifications (p = 7.8 × 10−2 for N23C H3K4me3 and p = 9.23 × 10−3–1.91 × 10−6 for the others, Fig. 1C and Supplementary Fig. 4). The overall increase in H3K27ac and H3K4me3 levels in association with the expressed G allele suggests a regulatory role in t(11;14) (Fig. 1D).
To validate the allele-specific enhancer effect of rs9344 and its impact on CCND1 gene expression, we analyzed two independent CRISPR/Cas9 base edited KMS12PE knock-in (KI) clones that were generated with a substitution of the wild-type A allele with the G allele at rs9344. Indeed, the A > G conversion within KI clones increased H3K27ac over the CCND1 locus compared to WT (Fig. 1E, F). Within RE5, there were twice as many reads carrying the G-allele in KI than reads carrying the A-allele in WT. Further, the expression of CCND1 increased by ~2-fold in KMS12PE KI GG clones compared to KMS12PE WT AA cells (Fig. 1G) supporting the role for the G allele in modulating enhancer activity of t(11;14) MM. To understand whether the allelic imbalance at rs9344 was detectable in other cell types without t(11;14), we evaluated publicly available data (Supplementary Table 2), including ATAC-seq (n = 22) and H3K27ac ChIP-seq data (n = 10) from non-t(11;14) MM samples, DNase-seq (n = 6) and H3K27ac ChIP-seq data (n = 8) from normal B cells, as well as H3K27ac and ChIP-seq data from lymphoblastoid cell lines (n = 5, all with G/A genotype). None of the non-t(11;14) MM (Supplementary Fig. 3) and normal samples had a peak that spanned rs9344 in RE5, excluding the possibility of genotype inference and an allelic imbalance test in these samples. This suggests that rs9344 alone without t(11;14) does not appear to promote H3K27ac or chromatin accessibility. Rather the imbalance of histone modifications and chromatin accessibility associated with rs9344 alleles are observed only after t(11;14) event.
PAX5 associates with the CCND1 locus
We next evaluated whether differences in the recruitment of transcription factors to the CCND1 locus may be associated with increased CCND1 gene expression. We utilized ENCODE ChIP-seq data from the GM12878 cell line. Of 76 transcription factors and activators, PAX5 showed a clear binding peak over rs9344 with high confidence, followed by ELF1 and PU.1 (Fig. 1B and Supplementary Fig. 6). Rs9344 was located within a PAX5 binding site in both replicates, showing high reproducibility (r = 0.94) (Fig. 1B). To support PAX5 binding in this region, we identified a PAX5 binding motif immediately upstream of rs9344 (Fig. 1H). We observed a trend toward an association between the rs9344 G allele and an increase in PAX5 binding (p = 1.09 × 10−1, Fig. 1C) in GM12878 that is heterozygous for rs9344.
We evaluated whether PAX5 was associated with increased CCND1 expression within t(11;14). Similar to CCND1, the expression of PAX5 significantly deviated from a normal distribution (p ≤ 8.8 × 10−36), where t(11;14) MM had a ~ 3–4-fold higher PAX5 expression compared to non-t(11;14) (p ≤ 1.3 × 10−13) in both cohorts (Table 1 and Fig. 2A). To understand which variables may be associated with increased PAX5 expression in MM cells, we performed a multivariate analysis considering the rs9344 genotype, age, sex, race and cytogenetic abnormalities. The analysis revealed that only t(11;14) was associated with a significant increase in PAX5 expression (p = 1.2 × 10−12 in CoMMpass and p = 4.1 × 10−16 in Mayo cohorts) (Supplementary Table 4.1). Considering the strong correlation between t(11;14) and PAX5 expression and the preference for the rs9344 G allele in t(11;14), we tested whether the rs9344 genotype was associated with increased PAX5 expression. No significant association between the rs9344 genotypes and PAX5 expression was found in t(11;14) (Supplementary Table 4.2). In addition, there was no interaction between the rs9344 genotypes and t(11;14) on PAX5 expression (Supplementary Table 4.2). Consistent with this finding, no increase in PAX5 expression was observed in KMS12PE WT and KI clones in association with the rs9344 genotype (Supplementary Fig. 7A).
A Expression of PAX5 in t(11;14) compared to non-t(11;14) samples in the CoMMpass cohort (top) and the Mayo cohort (bottom). B Interactions of rs9344 genotype by t(11;14) (top) and of expressed allele by t(11;14) (bottom) in the CoMMpass cohort. X-axis: PAX5 expression in transcripts-per-million (TPM); y-axis: CCND1 expression in TPM. Expression of CCND1 (TPM) in relation to rs9344 genotype in (C) or in relation to the expressed allele in (D) in the CoMMpass (top) and Mayo (bottom) cohorts. E Relative mRNA expression (left) or protein expression (right) of PAX5 and CCND1 following induced overexpression of PAX5 in U266B1 cells. Overexpression of PAX5 was induced by plasmid nucleofection. WT U266B1 transfected with empty (GFP) plasmid were used as negative control. The level of PAX5 and CCND1 mRNA in each sample was normalized to that of housekeeping gene GAPDH mRNAs. Normalized CT values for experimental sample and control sample were compared with paired Student’s t test with 0.05 as level of statistical significance. *P < 0.05, **P ≤ 0.01, ***P ≤ 0.001.
PAX5 regulates CCND1 expression in t(11;14)
Given the interaction of AG and GG genotypes of rs9344 with t(11;14) on CCND1 expression, we plotted the expression of PAX5 versus that of CCND1 in t(11;14) (separated by rs9344 genotypes or alleles) and non-t(11;14) cases (Fig. 2B). The plots suggested a stronger effect of PAX5 on CCND1 expression in the AG and GG genotypes groups within t(11;14). These observations were supported by the analysis with the full model demonstrating a positive association of PAX5 and CCND1 expression in relation to rs9344 genotype, with a higher expression of CCND1 respectively for AG and GG genotypes in the t(11;14) group than in the non-t(11;14) group (Table 3), which were consistent with Fig. 2B, C and Supplementary Table 4.3. No interaction effect was observed for t(11;14) cases having AA genotype.
Given that the rs9344 G allele had a much higher (4-fold) overall chance of being involved in the translocation than the A allele in heterogeneous (AG) cases, we determined the genotype of the expressed alleles from those cases. In homozygous AA and GG cases, the A or G allele was treated as the expressed allele and the expressed G allele was associated with increased CCND1 expression (Fig. 2D). The full model showed that the interaction of PAX5 with the expressed G allele conferred a much larger effect size compared to the A allele in CoMMpass (p = 1.3 × 10−26) and Mayo (p = 5.2 × 10−8) cohorts (Table 3 and Supplementary Table 4.4), demonstrating higher CCND1 expression in t(11;14) cases with the G allele than in non-t(11;14) cases but no effect on CCND1 expression in t(11;14) cases with the A allele (Table 3 and Supplementary Table 4.4). In the same full model, within the t(11;14) group the t(11;14) event alone accounted for 75% and 85% of CCND1 expression and the combined effect of t(11;14) and its interaction with the rs9344 G allele accounted for 98% of CCND1 expression in both cohorts. We next perturbed the expression of PAX5 in the t(11;14)-positive U266B1 cell line, which has the rs9344 AG germline genotype (Supplementary Fig. 8A). Overexpression of PAX5 ( + PAX5) using a PAX5 expression plasmid resulted in an ~2-fold increase in CCND1 expression. Sequencing of the overexpressed CCND1 allele revealed the presence of only genotype G (Fig. 2E, Supplementary Fig. 8A). Overexpression of PAX5 ( + PAX5) did not increase CCND1 expression in KMS12WT (AA) control and the KMS12PE KI cell lines (Supplementary Fig. 7B).
Discussion
We identified the interaction between the rs9344 G risk allele and t(11;14) contributed to increased active histone modifications and overexpression of CCND1 in MM, supporting rs9344 as a putative causal driver mediated by the IGH enhancer and PAX5. Although this study focused on t(11;14) MM irrespective of patient race, the G risk allele was found enriched in both healthy individuals of African genetic ancestry (Supplementary Fig. 9) and African American individuals with MM [3]. Resolving the mechanism of how rs9344 is associated with t(11;14) may enable a deep understanding of t(11;14) MM pathogenesis, particularly in populations with a high risk allele frequency.
Previous studies in non-t(11;14) samples have focused on the role of rs9344 in alternative splicing of CCND1 [27, 28]. We identified a ~1.5-2-fold increase in retention of intron 4 in association with genotype A (forming cyclin D1b) compared to G in primary t(11;14) MM samples and in the KMS12PE KI cell lines (Supplementary Fig. 10). However, mass spectrometry-based proteomics analysis did not detect any peptide specific to cyclin D1b (VSEGDVPGSLAGAYRGRHLVPRKCRGWCQGPQG) in U266B1, KMS12PE WT and KI cell lines and in 7 primary t(11;14) MM samples (data not shown) suggesting that MM cells may only express cyclin D1a similar to MCL [29] or that expression levels are extremely low and below the limit of detection. Thus, we focused our attention on the novel role of rs9344 within an active regulatory region of CCND1.
As the most prevalent translocation in MM [1], t(11;14) brings enhancer(s) from the IGH locus to the vicinity of CCND1 [16, 22, 30, 31], an event associated with its overexpression [22, 30]. Interestingly, 293T cells carrying CRISPR/Cas9 induced t(11;14), generated by placing the IGH Eµ sequence together with a 15-kb upstream region of CCND1, had low CCND1 expression [32] and overexpression of PAX5 in 293T cells did not result in increased CCND1 expression (data not shown), indicating an essential role of the active IGH enhancer in CCND1 overexpression. In contrast to normal immune cells, we identified three active enhancers in MM with t(11;14) [17, 18] and found evidence for the hijacking of IGH enhancer(s) upstream of CCND1 as previously reported [18, 22, 23]. De novo formation of the broad active domain reflects local chromatin reprogramming, presumably caused by the t(11;14) event [18].
We found that PAX5, whose expression is repressed in normal PCs [33], is retained in t(11;14) MM [34, 35] and plays a novel role in CCND1 overexpression. This finding is consistent with a recent study revealing increased chromatin accessibility at the PAX5 motif in MM samples with t(11;14) relative to non-t(11;14) [36]. Interestingly, PAX5 expression is often associated with MS4A1 (CD20) expression, small mature PC morphology [37, 38] and the recently described “B cell” like epigenetic signature [36]. We observed that t(11;14) MM was also correlated with CD20 expression (Supplementary Table 4.5) supporting the observation that t(11;14) MM maintains B-cell-like features [38]. This feature appears to be independent of rs9344 status, as we found no significant association between the rs9344 genotype and PAX5 expression (Supplementary Table 4.2, Supplementary Fig. 7A). Given that PAX5 and CD20 expression are important in defining the CD1 and CD2 expression subtypes of t(11;14) [39], we observed no differences in the distribution of these subtypes in relation to rs9344 allele status (Table 1, Supplementary Fig. 11).
Notably, rs9344 is one base pair away from a PAX5 binding motif. While TF binding is frequently altered by variants within the binding sites that create or disrupt the binding motif, adjacent base pairs can cause allele-specific TF binding [40] and further impact enhancer function [41, 42] by presumably impacting DNA shape, chromatin accessibility, or the binding of interacting TFs. Pioneer TFs can bind to nucleosomal DNA to open local chromatin, facilitating the binding of other TFs [43] and can directly recruit activating and repressive chromatin regulators to binding sites to regulate the gene expression. Thus, genetic variants that alter the binding sites of pioneer TFs cause allele-specific chromatin accessibility [44, 45]. While PAX5 binding induces or eliminates chromatin accessibility in the regulatory regions of its target genes [46, 47], the extent to which its binding is rewired in t(11;14) MM and how it may contribute to CCND1 expression remains unclear. Furthermore, the inability to increase CCND1 expression after PAX5 overexpression in the KMS12PE KI lines (GG genotype) was surprising. Interestingly, this cell line expresses very low endogenous PAX5 but expresses a PAX5 paralog, PAX8 (data not shown), suggesting that PAX8 could substitute for PAX5 in promoting CCND1 expression [48].
It remains unknown how precisely the t(11;14) translocation event occurs and whether the rs9344 G risk allele enhances translocation susceptibility or exerts its effect after the t(11;14) event. The higher CCND1 expression mediated by the G allele after the t(11;14) event may create a selective advantage for PCs to undergo malignant transformation. Surprisingly, we did not observe significant differences between G and A alleles in the acquisition of somatic mutations or differences in overall survival of t(11;14) MM patients (Supplementary Figure 12, Supplementary Tables 5.1–5.5 and 6).
Our study is limited by using two RNA-seq datasets with different designs and library preparations resulting in differences in sequencing depth, with or without coverage of intron and/or UTR regions. Other limitations include a lack of genotype information for variants in LD with rs9344, as these variants were located within intronic or intergenic regions that were not sequenced. However, we identified null interactions of the linked variants rs7177 and rs678653 with t(11;14), further supporting rs9344 as a candidate causal SNP. Further, due to the biological rarity of t(11;14) MM samples expressing A allele, the small sample size for this subtype resulted in a lower statistical power.
Taken together, we identified the GWAS lead SNP rs9344 as a potential causal variant in two independent MM patient cohorts. Rs9344 is located in an active enhancer that is bound by PAX5 in GM12878, which interacts with translocated IGH super enhancer(s) to upregulate CCND1 expression within t(11;14) MM. The study advances the understanding of t(11;14) MM and identifies a potential molecular mechanism linking rs9344 with increased t(11;14) MM risk.
Data access statement
All CoMMpass data used in this study are publicly available [database of Genotypes and Phenotypes (dbGap): phs000748.v1.p1 and EGAS00001001178] and also http://www.ncbi.nlm.nih.gov/bioproject/248538. Accession numbers for all other publicly available data sets using in this study are included in Supplementary Table 2. The ChIP-seq and ATAC-seq data have been submitted to dbGap. CCND1 and PAX5 TPM data from RNA-seq are available in Supplementary Tables 8–11.
References
Kumar SK, Rajkumar SV. The multiple myelomas - current concepts in cytogenetic classification and therapy. Nat Rev Clin Oncol. 2018;15:409–21.
Weinhold N, Johnson DC, Chubb D, Chen B, Forsti A, Hosking FJ, et al. The CCND1 c.870G>A polymorphism is a risk factor for t(11;14)(q13;q32) multiple myeloma. Nat Genet. 2013;45:522–5.
Baughn LB, Li Z, Pearce K, Vachon CM, Polley MY, Keats J, et al. The CCND1 c.870G risk allele is enriched in individuals of African ancestry with plasma cell dyscrasias. Blood Cancer J. 2020;10:39.
Seto M, Yamamoto K, Iida S, Akao Y, Utsumi KR, Kubonishi I, et al. Gene rearrangement and overexpression of PRAD1 in lymphoid malignancy with t(11;14)(q13;q32) translocation. Oncogene 1992;7:1401–6.
Smadbeck J, Peterson JF, Pearce KE, Pitel BA, Figueroa AL, Timm M, et al. Mate pair sequencing outperforms fluorescence in situ hybridization in the genomic characterization of multiple myeloma. Blood Cancer J. 2019;9:103.
Sharma N, Smadbeck JB, Abdallah N, Zepeda-Mendoza C, Binder M, Pearce KE, et al. The prognostic role of MYC structural variants identified by NGS and FISH in multiple myeloma. Clin Cancer Res. 2021;27:5430–9.
Skerget S, Penaherrera D, Chari A, Jagannath S, Siegel DS, Vij R, et al. Comprehensive molecular profiling of multiple myeloma identifies refined copy number and expression subtypes. Nat Genet. 2024;56:1878–89.
Zhong J, Ye Z, Lenz SW, Clark CR, Bharucha A, Farrugia G, et al. Purification of nanogram-range immunoprecipitated DNA in ChIP-seq application. BMC Genom. 2017;18:985.
Corces MR, Trevino AE, Hamilton EG, Greenside PG, Sinnott-Armstrong NA, Vesuna S, et al. An improved ATAC-seq protocol reduces background and enables interrogation of frozen tissues. Nat Methods. 2017;14:959–62.
Ran FA, Hsu PD, Wright J, Agarwala V, Scott DA, Zhang F. Genome engineering using the CRISPR-Cas9 system. Nat Protoc. 2013;8:2281–308.
Bergsagel PL, Kuehl WM, Zhan F, Sawyer J, Barlogie B, Shaughnessy J Jr. Cyclin D dysregulation: an early and unifying pathogenic event in multiple myeloma. Blood. 2005;106:296–303.
Herrera-Luis E, Benke K, Volk H, Ladd-Acosta C, Wojcik GL. Gene–environment interactions in human health. Nat Rev Genet. 2024:1–17. https://doi.org/10.1038/s41576-024-00731-z.
Machiela MJ, Chanock SJ. LDlink: a web-based application for exploring population-specific haplotype structure and linking correlated alleles of possible functional variants. Bioinformatics. 2015;31:3555–7.
Wang Y, Song F, Zhang B, Zhang L, Xu J, Kuang D, et al. The 3D genome browser: a web-based browser for visualizing 3D genome organization and long-range chromatin interactions. Genome Biol. 2018;19:151.
Rao SS, Huntley MH, Durand NC, Stamenova EK, Bochkov ID, Robinson JT, et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell. 2014;159:1665–80.
Alvarez-Benayas J, Trasanidis N, Katsarou A, Ponnusamy K, Chaidos A, May PC, et al. Chromatin-based, in cis and in trans regulatory rewiring underpins distinct oncogenic transcriptomes in multiple myeloma. Nat Commun. 2021;12:5450.
Fulciniti M, Lin CY, Samur MK, Lopez MA, Singh I, Lawlor MA, et al. Non-overlapping Control of transcriptome by promoter- and super-enhancer-associated dependencies in multiple myeloma. Cell Rep. 2018;25:3693–705 e3696.
Jia Y, Zhou J, Tan TK, Chung TH, Wong RWJ, Chooi JY, et al. Myeloma-specific superenhancers affect genes of biological and clinical relevance in myeloma. Blood Cancer J. 2021;11:32.
Roadmap Epigenomics C, Kundaje A, Meuleman W, Ernst J, Bilenky M, Yen A, et al. Integrative analysis of 111 reference human epigenomes. Nature. 2015;518:317–30.
Hnisz D, Abraham BJ, Lee TI, Lau A, Saint-Andre V, Sigova AA, et al. Super-enhancers in the control of cell identity and disease. Cell. 2013;155:934–47.
Alaterre E, Ovejero S, Herviou L, de Boussac H, Papadopoulos G, Kulis M, et al. Comprehensive characterization of the epigenetic landscape in Multiple Myeloma. Theranostics. 2022;12:1715–29.
Choudhury SR, Ashby C, Tytarenko R, Bauer M, Wang Y, Deshpande S, et al. The functional epigenetic landscape of aberrant gene expression in molecular subgroups of newly diagnosed multiple myeloma. J Hematol Oncol. 2020;13:108.
Mikulasova A, Kent D, Trevisan-Herraz M, Karataraki N, Fung KTM, Ashby C, et al. Epigenomic translocation of H3K4me3 broad domains over oncogenes following hijacking of super-enhancers. Genome Res. 2022;32:1343–54.
Stergachis AB, Haugen E, Shafer A, Fu W, Vernot B, Reynolds A, et al. Exonic transcription factor binding directs codon choice and affects protein evolution. Science. 2013;342:1367–72.
Birnbaum RY, Clowney EJ, Agamy O, Kim MJ, Zhao J, Yamanaka T, et al. Coding exons function as tissue-specific enhancers of nearby genes. Genome Res. 2012;22:1059–68.
Birnbaum RY, Patwardhan RP, Kim MJ, Findlay GM, Martin B, Zhao J, et al. Systematic dissection of coding exons at single nucleotide resolution supports an additional role in cell-specific transcriptional regulation. PLoS Genet. 2014;10:e1004592.
Knudsen KE, Diehl JA, Haiman CA, Knudsen ES. Cyclin D1: polymorphism, aberrant splicing and cancer risk. Oncogene 2006;25:1620–8.
Pestell RG. New roles of cyclin D1. Am J Pathol. 2013;183:3–9.
Marzec M, Kasprzycka M, Lai R, Gladden AB, Wlodarski P, Tomczak E, et al. Mantle cell lymphoma cells express predominantly cyclin D1a isoform and are highly sensitive to selective inhibition of CDK4 kinase activity. Blood. 2006;108:1744–50.
Mikulasova A, Ashby C, Tytarenko RG, Qu P, Rosenthal A, Dent JA, et al. Microhomology-mediated end joining drives complex rearrangements and overexpression of MYC and PVT1 in multiple myeloma. Haematologica. 2020;105:1055–66.
Rico D, Kent D, Karataraki N, Mikulasova A, Berlinguer-Palmini R, Walker BA, et al. High-resolution simulations of chromatin folding at genomic rearrangements in malignant B cells provide mechanistic insights into proto-oncogene deregulation. Genome Res. 2022;32:1355–66.
Tsuyama N, Abe Y, Yanagi A, Yanai Y, Sugai M, Katafuchi A, et al. Induction of t(11;14) IgH enhancer/promoter-cyclin D1 gene translocation using CRISPR/Cas9. Oncol Lett. 2019;18:275–82.
Fuxa M, Busslinger M. Reporter gene insertions reveal a strictly B lymphoid-specific expression pattern of Pax5 in support of its B cell identity function. J Immunol. 2007;178:3031–7.
Borson ND, Lacy MQ, Wettstein PJ. Altered mRNA expression of Pax5 and Blimp-1 in B cells in multiple myeloma. Blood. 2002;100:4629–39.
Bal S, Kumar SK, Fonseca R, Gay F, Hungria VT, Dogan A, et al. Multiple myeloma with t(11;14): unique biology and evolving landscape. Am J Cancer Res. 2022;12:2950–65.
Leblay N, Ahn S, Tilmont R, Poorebrahim M, Maity R, Lee H, et al. Integrated epigenetic and transcriptional single-cell analysis of t(11;14) multiple myeloma and Its BCL2 dependency. Blood. 2023;143:42–56.
Robillard N, Avet-Loiseau H, Garand R, Moreau P, Pineau D, Rapp MJ, et al. CD20 is associated with a small mature plasma cell morphology and t(11;14) in multiple myeloma. Blood. 2003;102:1070–1.
Lin P, Mahdavy M, Zhan F, Zhang HZ, Katz RL, Shaughnessy JD. Expression of PAX5 in CD20-positive multiple myeloma assessed by immunohistochemistry and oligonucleotide microarray. Mod Pathol. 2004;17:1217–22.
Zhan F, Huang Y, Colla S, Stewart JP, Hanamura I, Gupta S, et al. The molecular classification of multiple myeloma. Blood. 2006;108:2020–8.
Lu X, Chen X, Forney C, Donmez O, Miller D, Parameswaran S, et al. Global discovery of lupus genetic risk variant allelic enhancer activity. Nat Commun. 2021;12:1611.
Farley EK, Olson KM, Zhang W, Brandt AJ, Rokhsar DS, Levine MS. Suboptimization of developmental enhancers. Science. 2015;350:325–8.
Yang MG, Ling E, Cowley CJ, Greenberg ME, Vierbuchen T. Characterization of sequence determinants of enhancer function using natural genetic variation. Elife. 2022;11:e76500.
Cirillo LA, Lin FR, Cuesta I, Friedman D, Jarnik M, Zaret KS. Opening of compacted chromatin by early developmental transcription factors HNF3 (FoxA) and GATA-4. Mol Cell. 2002;9:279–89.
Jacobs J, Atkins M, Davie K, Imrichova H, Romanelli L, Christiaens V, et al. The transcription factor Grainy head primes epithelial enhancers for spatiotemporal activation by displacing nucleosomes. Nat Genet. 2018;50:1011–20.
Atak ZK, Taskiran II, Demeulemeester J, Flerin C, Mauduit D, Minnoye L, et al. Interpretation of allele-specific chromatin accessibility using cell state-aware deep learning. Genome Res. 2021;31:1082–96.
Revilla IDR, Bilic I, Vilagos B, Tagoh H, Ebert A, Tamir IM, et al. The B-cell identity factor Pax5 regulates distinct transcriptional programmes in early and late B lymphopoiesis. EMBO J. 2012;31:3130–46.
Gruenbacher S, Jaritz M, Hill L, Schafer M, Busslinger M. Essential role of the Pax5 C-terminal domain in controlling B cell commitment and development. J Exp Med. 2023;220:e20230260.
Hart MR, Anderson DJ, Porter CC, Neff T, Levin M, Horwitz MS. Activating PAX gene family paralogs to complement PAX5 leukemia driver mutations. PLoS Genet. 2018;14:e1007642.
Acknowledgements
We thank Dr. Jeong-Heon Lee and other members of the Epigenomics Development Laboratory at the Mayo Clinic. The research reported in this publication was partially supported by the Department of Laboratory Medicine and Pathology, Mayo Clinic Cancer Center, the National Cancer Institute of the National Institutes of Health under Award Number R37CA272883 (LBB) and also supported in part by grants to A.P. (U01CA271410 and P30CA15083). The content is the sole responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
Conceived and designed the analysis: LBB, HT, HY, SS, NS, JS, EE, and AGM; Collected the data/performed the analysis: LBB, HT, HY, SS, SL, NS, JS, YA, CV, JSH, RK, AP, MK, FM, AM, and EE; Contributed data/samples: JK, PLB, RF, AKS, SVR, SKK, and EB; Completed first draft of manuscript: LBB, HT, HY, SS, and EE.
Corresponding author
Ethics declarations
Competing interests
SKK served as a consultant for Celgene, Takeda, Amgen, Janssen, and Bristol-Myers Squibb, and received research funding from Celgene, Takeda, Novartis, Amgen, AbbVie, Janssen, and Bristol-Myers Squibb. LBB served as a consultant for Genentech. Eran Elhaik is a consultant at the DNA Diagnostics Centre. AKS Amgen, Glaxo Smith Klein, Genentech, Janssen, Ono, Skyline Diagnostics, Tempus Labs Inc. RF Consulting: AbbVie, Adaptive Biotechnologies, AMGEN, AZeneca, Bayer, Binding Site, BMS (Celgene), Millenium Takeda, Jansen, Juno, Kite, Merck, Pfizer, Pharmacyclics, RA Capital, Regeneron, Sanofi. Scientific Advisory Boards: Adaptive Biotechnologies, Caris Life Sciences, Oncotracker. Board of Directors: Antegene (for profit), AZBio (not for Profit), patent for FISH in MM. The remaining authors declare no competing financial interests relevant to the project.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Tang, H., Yan, H., Shivaram, S. et al. Functional variant rs9344 at 11q13.3 regulates CCND1 expression in multiple myeloma with t(11;14). Leukemia 39, 42–50 (2025). https://doi.org/10.1038/s41375-024-02363-y
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41375-024-02363-y