Abstract
Single-cell analysis is crucial for deciphering cellular heterogeneity and understanding complex biological systems. However, most existing single-cell sample manipulation (SCSM) systems suffer from various drawbacks such as high cost, low throughput, and heavy reliance on human interventions. Currently, large language models (LLMs) have been used in robotic platforms, but a limited number of studies have reported the application of LLMs in the field of lab-on-a-chip automation. Consequently, we have developed an active-matrix digital microfluidic (AM-DMF) platform that realizes fully automated biological procedures for intelligent SCSM. By combining this with a fully programmable lab-on-a-chip system, we present a breakthrough for SCSM by combining LLMs and object detection technologies. With the proposed platform, the single-cell sample generation rate and identification precision reach up to 25% and 98%, respectively, which are much higher than the existing platforms in terms of SCSM efficiency and performance. Furthermore, a three-class detection method considering droplet edges is implemented to realize the automatic identification of cells and oil bubbles. This method achieves a 1.0% improvement in cell recognition accuracy according to the \({\rm{AP}}_{75}^{\rm{test}}\) metric, while efficiently distinguishing obscured cells at droplet edges, where approximately 20% of all droplets contain cells at their edges. More importantly, as the first attempt, a ubiquitous tool for automatic SCSM workflow generation is developed based on the LLMs, thus advancing the development and progression of the field of single-cell analysis in the life sciences.

Similar content being viewed by others
Introduction
Cells and their interactions within the human body are both complex and diverse, which necessitates the focus on single-cell analysis in fields such as genomics1,2, transcriptomics3,4, proteomics5,6,7, and metabolomics8,9. These single-cell researches enable the discovery of gene regulation mechanisms and protein expression dynamics, which have completely transformed how we understand the role of cell heterogeneity in health and disease10,11,12. Many technologies and platforms have been developed for single-cell analysis, such as flow cytometry13, microwell microfluidics14, microdroplet microfluidics15, optoelectronic tweezers16, and digital microfluidics (DMF)17 (Table S1). A DMF system offers several benefits over other platforms, as it can simultaneously perform sample separation, real-time manipulation, and in situ analysis in parallel on a two-dimensional surface18,19,20. Furthermore, DMF systems are compatible with various detection modalities (e.g., optical or electrochemical detection), making it particularly useful in single-cell research applications21,22,23. DMF systems are comprised of a DMF driver and a DMF chip, which manipulates the droplets. Most DMF chips feature a passive matrix (PM) structure, which utilizes a grid of large electrodes, individually connected to control lines to produce the droplet movement. However, PM-DMF systems are constrained by their wiring designs and therefore, their electrode density, resulting in a limited number of individually controlled droplets24. To address this issue, researchers have developed active-matrix digital microfluidic (AM-DMF) systems with integrated thin-film transistors at each pixel, allowing each pixel to be individually addressed by scanning the row and column control signals and enabling the parallel manipulation of thousands of cell-containing droplets25,26,27.
On the AM-DMF systems, great challenges exist in achieving intelligent single-cell sample manipulation (SCSM), which often include generating nanolitre-scale cell-captured droplets, sorting single-cell droplets, and assigning the sorted droplets into desired locations for further experiments. Nowadays, the SCSM workflow continues to rely heavily on manual input, particularly in the manual editing of droplet paths and the manual sorting of single-cell droplets. As the number of droplets increases, these processes become increasingly time-consuming and inefficient. Therefore, it is essential to enhance the development of automated workflows for efficient and reliable SCSM. The existing studies have demonstrated the successful application of artificial intelligence (AI) on automation design for biology28,29,30 and the microfluidics fields31,32,33,34,35,36,37. However, fewer studies have applied AI in AM-DMF systems. In our previous work, we integrated AM-DMF technology with AI to achieve automated biosample determination38, which supports single-cell recognition. However, cell recognition can be affected by oil bubbles within droplets. Moreover, the droplet path design was inefficient due to manual editing processes.
Previous studies on droplet path planning primarily focused on clearly defined bounded problems within specific scenarios. Compiling-based approaches reduce the droplet routing problem to well-known problems, such as integer linear programming39 and boolean satisfiability40. These algorithms are complete, optimal, and efficient for small-scale problems. However, their computational complexity grows exponentially with problem scale. Moreover, they primarily aim to minimize completion time and are not suitable for other optimization criteria, such as minimizing path crossover to reduce the potential for cross-contamination. Numerous priority-based methods that sequentially generate routes for individual droplets are proposed with varying priority rules and strategies to avoid detours or deadlocks41,42. They are fast and flexible, and easily integrate with other goals such as minimizing active electrodes or cross-contamination. However, their effectiveness has been demonstrated only in small-scale scenarios. Reinforcement learning-based methods are proposed to deal with online dynamic situations such as movement failure caused by electrode degradation43,44. These techniques are therefore neither complete nor optimal, facing challenges with sparse rewards and unstable training. In recent years, large language models (LLMs) have become a powerful tool that has been widely adopted in both academia and industry, due to their unprecedented performance and flexibility in a range of applications45,46. They have shown great potential in robotic-based platforms by simplifying and minimizing the labor effort45,47,48. The emergent capabilities of LLMs provide new perspectives for droplet path planning. They enable users to obtain solutions that meet specific requirements through human–computer interaction without redefining the problem. Given the experimental conditions and objectives, LLMs can generate corresponding path planning strategies. However, their application in droplet path planning still faces challenges, including high data requirements and substantial computational resources for training and inference. In this work, we take the first step forward and develop an AM-DMF platform that realizes fully automated biological procedures for intelligent SCSM. The platform employs an AI-based model for the sorting of single-cell droplets and an LLMs-based droplet path generation (DPG) model for generating nanolitre-scale droplets and path planning of single-cell droplets to designated locations. Based on a fully programmable AM-DMF system, we represent a breakthrough for SCSM by combining LLMs and object detection technologies, significantly enhancing experimental efficiency and broadening the horizons of AI applications in the life sciences. The automated workflow combines the DPG model and the cell recognition model for intelligent SCSM, as illustrated in Fig. 1. By integrating these methods, our AM-DMF platform enhances the accuracy and reliability of SCSM while also increasing the flexibility and automation of its workflow. This work can be summarized according to the following research highlights.
The automated workflow combines the DPG model and the cell recognition model for intelligent SCSM
Advancement of the platform
As a pioneering work, we introduce a fully automated AM-DMF platform for SCSM based on LLMs that processes 1600–1700 droplets/h, achieves a single-cell sample generation rate of over 25%, and attains a model identification precision exceeding 98%.
Novelty of the proposed method
We have addressed several challenges in the DMF field. The three-class detection method enhances cell recognition accuracy by identifying oil bubbles, recognizing cells concealed at the edges of droplets using the proposed droplet movement method, and automatically generating workflows using the DPG model, eliminating the need for manual editing.
Relevance and potential to advance new biological applications
The proposed method serves as a powerful platform for intelligent SCSM, supporting a wider range of applications in single-cell research and extending its utility to various research areas, such as biological sciences and chemistry.
Results and discussion
Distinguishing cells and oil bubbles
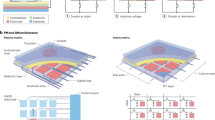
Droplets are enveloped in the medium oil on the AM-DMF chip. During droplet transportation, a phenomenon is observed where the oil film sandwiched between the droplet and the chip may break up, forming oil bubbles49. Oil bubbles under droplets form due to the instability of an entrapped thin oil film caused by electrostatic pressure and surface tension, with their size influenced by the applied voltage, as confirmed through both theoretical analyses and numerical simulations50. During the experiments evaluating the previously proposed model for cell recognition38, we observed that some oil bubbles and cells exhibited similar appearances, causing the cell recognition algorithm to misidentify oil bubbles as cells. This misidentification problem can be challenging to correct, even for a human observer. To increase the accuracy of the model in terms of recognizing cells, we developed a three-class method that builds on the previous two-class recognition algorithm (droplets, cells) by adding an oil bubble identification mechanism. To ensure the accuracy of data labeling, we implemented a rigorous annotation process. Each image was independently annotated by multiple experienced experts. Only images with unanimous annotations were included in the training dataset, while those with discrepancies were discarded. The three-class detection model was trained for 100 epochs. Figure 2a–f shows the training and test set loss functions for the droplet, cell, and oil bubble detection tasks. The results indicate that the three-class detection model has converged and achieved optimal performance. Figure 2g shows the confusion matrix of the three-class model. To further verify the effectiveness of our proposed three-class method, we carried out a model comparison experiment on the same dataset to test the two-class and three-class methods separately. The two-class model was trained on a dataset containing only droplets and cells, whereas the dataset for the three-class model included an additional category for manually labeled bubbles. The results shown in Fig. 2h reveal that the two-class method achieved an average precision on the test set \({\rm{AP}}_{50}^{\rm{test}}\) of 96.4% for cells, whereas the three-class method slightly increased this value to 96.8%, representing an improvement of 0.4%. In terms of \({\rm{AP}}_{75}^{\rm{test}}\), the two-class method achieved 90.9% for cells, with the three-class method enhancing this value to 91.9%, representing an improvement of 1.0%. Finally, regarding the overall \({\rm{AP}}_{50:95}^{\rm{test}}\), the two-class method reached 74.8% for cells, and the three-class method improved this score to 75.5%, representing an improvement of 0.7%. Table S2 compares the cell category recognition performance of two-class and three-class methods across different models. As illustrated in Fig. 2i, the two-class detection model identified 5 cells, whereas the three-class detection model identified 4 cells and 13 oil bubbles, revealing that one of the oil bubbles was misclassified as a cell in the two-class detection results. Figure 2j shows that in the two-class detection task, 5 cells were detected, whereas in the three-class detection task, 3 cells and 19 oil bubbles were identified, correcting the misclassification of two oil bubbles as cells in the two-class detection task. Finally, Fig. 2k indicates that two-class detection resulted in 2 cells, whereas three-class detection resulted in 1 cell and 4 oil bubbles, avoiding the mislabelling of an oil bubble as a cell in the two-class detection task.
The loss functions of the cell detection model on the training dataset (a–c) and test dataset (d–f). g The confusion matrix of the three-class model. h Cell category recognition performance comparison between two-class and three-class models. i–k The red boxes indicate the droplets predicted by the model, the pink boxes represent the cells predicted by the model, and the orange boxes denote the oil bubbles predicted by the model
Multi-scenario cell recognition testing and potential application analysis
In practice, cell recognition models may encounter various challenging situations, such as unclear images or conducting experiments on different chip substrates. When screening single-cell samples, the autofocus algorithm of the AM-DMF system is capable of controlling the imaging plane of the microscope within ±25 μm of the focal plane. We conducted a series of tests examining images with varying focus levels to assess the performance of the model. Figure 3a shows a schematic diagram of the model’s predictions produced for droplets (red boxes), cells (pink boxes), and oil bubbles (orange boxes) at various imaging clarity levels. Specifically, we conducted 20 groups of tests. In each group, we fix the x–y position of the system to target droplets on the chip, while moving the microscope upwards and downwards, in the z-axis, to vary the focus of the captured image. The clearest image was identified as a focal plane, followed by moving the microscope upwards and downwards with a step size of 5 μm at the same position, moving up to 100 μm in both directions. The results of the droplet recognition test are shown in Fig. 3b. The model could correctly detect droplets across all 20 positions, regardless of the image clarity level. The model maintained this accuracy for an average distance of 100 μm above and below the focal plane. Figure 3c depicts the results of the cell recognition tests. The model exhibited a slight accuracy variance across the 20 positions for images with different clarity levels. The model could correctly detect cells at average distances of 78 μm above the focal plane and 72.75 μm below it. Notably, within the controllable focusing range of ±25 μm of the AM-DMF system, the model demonstrated high accuracy in terms of recognizing droplets and cells, laying the foundation for a fully automated algorithm that can manipulate high-throughput single-cell samples in the next section.
a Model’s predictions were produced for droplets (red boxes), cells (pink boxes), and oil bubbles (orange boxes) at different imaging planes. b Model precision rates achieved for droplet recognition at various distances from the focal plane. c Model precision rates achieved for cell recognition at various distances from the focal plane. Marginal histograms of droplet (d) and cell (e) heights and widths obtained by the cell recognition model and manual measurements
Additionally, we carried out real-time analysis of droplet morphological features, cell morphological features, and cell density on the AM-DMF system using the cell recognition model. Specifically, we analyzed 212 locations under a high-magnification field, covering 212 droplets and 310 cells. Figure 3d presents the marginal histograms of droplet heights and widths obtained from the model’s predictions and manual measurements, demonstrating the effectiveness of our proposed model. More than 80% of the droplets have diameters ranging from 110 to 130 μm. A linear regression analysis of the droplet height and width data resulted in an aspect ratio of 0.9702, indicating excellent circularity of the droplets. This demonstrates that our AM-DMF chip offers consistent electrode control. Similarly, Fig. 3e depicts the marginal histograms of cell heights and widths obtained from the model’s predictions and manual measurements, demonstrating the effectiveness of our proposed model. More than 80% of the cells have diameters ranging from 7 to 11 μm. A linear regression analysis of the cell height and width data from the model revealed an aspect ratio of 0.9754, suggesting a high level of cell circularity. This finding demonstrates that the tested cells are uniform in shape and exhibit high viability, indicating that DMF actuation is gentle on cells. Additionally, as shown in Fig. S1, we conducted 28 sets of cell density tests. The results demonstrate the significant potential of our platform in measuring these parameters, contributing to the analysis of electrode control uniformity, droplet morphological features, cell morphological features, and cell density.
Moreover, we conducted additional tests to assess the robustness of the three-class cell detection model. Figure 4a displays the model predictions produced on various AM-DMF chip backgrounds, demonstrating its good performance. Figure 4b–f illustrates the precision, recall, mAP50, mAP75, and mAP50:95 functions achieved by the three-class recognition model on the test set. Figure 4g presents the performance metrics achieved by the model on the test set. We conducted testing on 1482 locations under a high-magnification field, encompassing 10,574 boxes. The overall performance metrics are as follows: the precision metric achieved 92.7%, the recall metric achieved 93.4%, the mAP50 metric achieved 96.1%, the mAP75 metric achieved 83.4%, and the mAP50:95 metric achieved 76.4%. For the droplet category, which has 1485 instances, the precision metric was 99.7%, the recall metric was 99.6%, the AP50 metric was 99.4%, the AP75 metric was 99.4%, and the AP50:95 metric was 98.3%. For the cell category, which has 2157 instances, the precision metric reached 93.1%, the recall metric reached 95.2%, the AP50 metric reached 96.8%, the AP75 metric reached 91.9%, and the AP50:95 metric reached 75.5%. For the bubble category, which has 6,932 instances, the precision metric achieved 85.3%, the recall metric achieved 85.5%, the AP50 metric achieved 92.1%, the AP75 metric achieved 59.1%, and the AP50:95 metric achieved 55.3%. The results demonstrate that the model exhibits robust performance in recognizing droplets, cells, and oil bubbles. Although the performance metrics for the oil bubble category are less favorable than those for the droplet and cell categories, this difference is expected, as the dataset does not label minute oil bubbles that do not impact cell recognition. In practical applications, we can increase the confidence threshold (e.g., above 0.45) to filter out minute bubbles, which typically have low confidence scores, thereby minimizing their impact on downstream analysis. We conducted further tests to evaluate the platform’s performance on various cell types, as shown in Fig. S2. The results show the platform’s applicability across a broader range of cell types.
a The model’s predictions were evaluated across different AM-DMF chip backgrounds. The precision metric function (b), the recall metric function (c), the mAP function at 0.5 IoU (d), the mAP function at 0.75 IoU (e), and the mAP function at a range of 0.5–0.95 IoU (f) of the model on the test dataset. g Model performance metrics values
Recognizing cells concealed at the edges of droplets
Theoretically, the encapsulation of cells within droplets is expected to follow the Poisson distribution51. The positions of the cells are distributed randomly within the droplets. Therefore, we noticed that cells positioned near the edges of droplets were often obscured, making it very difficult for the model and manual inspection to identify the cells. To determine whether cells are at the edge, we performed multiple movements of 500 droplets in four directions: up, left, down, and right (Fig. S3). If additional cells were detected during any of these movements, we concluded that the cells were located at the edge of the droplet. As illustrated in Fig. 5a, a statistical analysis was performed, which revealed that approximately 76% of all droplets exhibited no cells at their edges, while around 24% of the droplets contained cells at their edges. Within the subset of droplets containing cells at their edges, it was found that 79.2% had one cell present, 16.7% had two cells present, and 4.2% had more than two cells present. Due to their concealment at the edges of droplets, the chosen single-cell droplets may actually contain two or more cells, and the presence of this phenomenon affects the reliability of droplet-based single-cell research. Therefore, we propose a droplet movement method to improve the cell recognition performance. This approach reduces model detection failures for cells located at the edges of droplets. By applying a voltage to the electrodes adjacent to a droplet, a movement was induced for the target droplet. This movement displaces the cells from the edge of the droplet, due to the difference in their momenta, to its interior, enhancing the detection ability of the algorithm. Figure 5b shows that the model detected 1 cell and 7 oil bubbles before the droplet movement occurred. In comparison, the model identified 2 cells and 7 oil bubbles after the droplet moved to the left, effectively resolving the issue of cells hidden at the droplet edge. Similarly, Fig. 5c shows that prior to droplet movement, the model detected 0 cells and 12 oil bubbles. However, after shifting the droplet upward, the model detected 1 cell and 7 oil bubbles, thus avoiding the problem of cells being obscured by the droplet contour and being undetected by the model.
a A statistical analysis of the percentage of cell positions distributed within the droplets. b Before moving the droplet to the left, the model detected one cell; after the droplet moved, two cells were detected. c Before moving the droplet upward, the model detected no cells; after the droplet moved, one cell was detected
Intelligent DPG
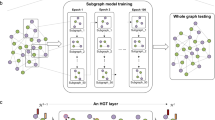
An automated path design method for DMF systems can significantly reduce the time required for manual path design, simplifying the task of planning and executing experimental procedures, and enhancing operational efficiency. Therefore, we propose a DPG model to automatically design experimental paths. DPG is a Llama 3-based droplet path generation model that can automatically generate droplet movement and splitting paths. We utilized Llama 3-8B as the pre-trained model and employed the low-rank adaptation (LoRA) fine-tuning technique to freeze the weights of the pre-trained Llama 3-8B model. Figure 6 shows the droplet paths generated by the DPG model through a series of simulations and experiments. The DPG model was trained for 31,432 steps. Figure 6a, b illustrates the training and evaluation loss functions during the training process. We compared the performance of fine-tuned Llama 3.2-3B and Llama 3-8B. The results indicate that fine-tuned Llama 3-8B demonstrates superior performance in the droplet path generation task. Figure 6c illustrates the training and evaluation results of the DPG model. Throughout the training process, the total floating point operations (flops) reached 3.76 × 1018, with a training loss of 4.65 × 10−3. The training runtime lasted for 41,903 s, achieving an average processing rate of 3.001 samples/s and 0.75 steps/s. In the evaluation phase, the evaluation loss was recorded as 1.69 × 10−6, with a runtime of 979 s and an average processing rate of 14.267 samples/s and 7.134 steps/s. These results indicate that the model has achieved optimal performance in both the training and evaluation processes. Figure 6d shows the droplet movement paths generated by DPG in the y-axis direction, the x-axis direction, and a combination of both the x- and y-axis directions. Figure 6e displays subdroplet generation paths generated by DPG, illustrating the process of generating subdroplets with widths of 2, heights of 2, row spacings of 2, and column spacings of 4. Figure 6f depicts the droplet movement and splitting paths generated by DPG, illustrating the process from the starting droplet (row 60, column 36) to the end droplet (row 54, column 42). At the end droplet position, subdroplets were generated with widths of 1, heights of 2, row spacings of 2, and column spacings of 2. Figure 6g shows the paths generated by DPG for droplets assigned to designated locations. Five designated droplets were assigned to specified locations with a column movement of −4, a row movement of −12, and a subdroplet column spacing of 3 in a 4 × 4 droplet array. In the example depicted in Fig. 6g, with regard to the output time issue of the DPG model, we propose a new droplet path output format. As illustrated in Tables S3 and S4, the original format required 2882 characters, whereas our new format only demands 788 characters, representing a reduction by a factor of 3.66. This substantial decrease in the number of output characters greatly enhances the temporal efficiency of the path-planning process executed by the model. Additional details regarding the user prompts, DPG model responses, the concepts used in the prompts and responses, and a series of simulations can be found in Tables S4–S9 and Movie S1. We conducted a series of tests to evaluate the performance of the DPG model in generating droplet paths. As shown in Fig. 6h, the user prompt and the corresponding responses from the DPG, GPT-4o, Gemini 2.0, and DeepSeek-V3 models are presented. Of these, only the DPG model successfully generated the correct path, while the other models failed to generate the correct path.
a The training loss function of the DPG model. b The evaluation loss function of the DPG model. c Statistics on the training and evaluation results of the DPG model. d Droplet movement paths are generated by DPG in the y-axis direction, the x-axis direction, and both the x- and y-axis directions. e Subdroplet generation paths generated by DPG. f Both the droplet movement and splitting paths were generated by DPG. g Paths generated by DPG for droplets assigned to designated locations. h The user prompt and corresponding responses from the DPG, GPT-4o, Gemini 2.0, and DeepSeek-V3 models
The solution for intelligent SCSM
We developed a solution for high-precision and high-throughput SCSM, addressing the challenges traditionally associated with manual operation, which often leads to inconsistencies and inefficiencies. The proposed solution can achieve a precision rate of over 98% in terms of cell identification and a single-cell sample generation rate exceeding 25%. The proposed solution is detailed in Movie S2 and illustrated in the flowchart shown in Fig. 7a, including generating a droplet array, sorting out single-cell droplets, and moving the single-cell droplets into the desired location. Figure 7b illustrates the user prompt, the response of the DPG model, and the droplet array generation test conducted on the AM-DMF chip. We generated an 8 × 8 array of subdroplets. Then, the single-cell sample sorting stage was performed by combining the three-class method with the droplet movement method (Fig. 7a, c). A motion control card (MCC) guides the microscope to the specified droplet via the x- and y-axes. The z-axis of the MCC, combined with the microscope module, activates the autofocus function to ensure image clarity. These images were fed into the cell recognition model for forward inference, with cell counts recorded via postprocessing algorithms such as nonmaximum suppression (NMS). For single-cell detection, the process involved manipulating the position of the droplet: if more than one cell was detected at a position, the process is stopped, and the camera is moved to the next location. Otherwise, the droplet was moved to the right, and the camera captured a new image for cell counting. If more than one cell still remained, the process was stopped; otherwise, the droplet was moved to the left. After three attempts, if no cells were detected, the process moved on. If a single cell was identified, its location was recorded for further analysis. Finally, the DPG model generated paths according to the user prompt for guiding single-cell samples to their designated locations after the sorting workflow was completed (Fig. 7d). The DPG model is designed to automatically generate experimental paths. Further details on addressing real-world variability factors, such as electrode failures or unsuccessful droplet splitting events, can be found in our previous work38.
a A fully automated algorithmic flowchart for SCSM. b The droplet path generated with 8 × 8 subdroplets by the DPG model on the AM-DMF chip (128 rows × 128 columns). c The process of sorting the droplets containing one cell by combining the three-class method with the droplet movement method. d The path for the assigned single-cell sample to their designated locations is generated by the DPG model
Conclusion
In this work, we take the first step forward and present a platform for intelligent SCSM via an AM-DMF system, employing a DPG model and a cell recognition model in combination with specialized pre- and postprocessing techniques. This platform offers a fully automated solution for efficiently manipulating single-cell samples, thus addressing the current demands of single-cell research and significantly outperforming existing platforms in terms of efficiency and performance. We conducted a comprehensive intelligent SCSM experiment, including generating paths for droplets containing cells via the DPG model, performing cell recognition via the three-class classification method and droplet movement method, and generating paths for assigning single-cell droplets to designated locations based on the DPG model. The proposed method was able to achieve a single-cell sample generation rate of over 25% and a model identification precision rate exceeding 98%. The novelty of the proposed method include (1) an AI-based algorithm capable of detecting and distinguishing between droplets, cells, and oil bubbles, which improved the cell recognition accuracy of the model by 1.0% in terms of the \({\rm{AP}}_{75}^{\rm{test}}\) metric; (2) a technique that leverages droplet movement to recognize cells obscured by droplet edges, with approximately 20% of all droplets contained cells at their edges; (3) a DPG model for automatically generating experimental paths, replacing the manual design process; (4) a fully automated solution for SCSM; and (5) two comprehensive annotated datasets for an AM-DMF chip. We anticipate that the integration and proliferation of AM-DMF technology, LLMs, and object detection technologies, combined with the ongoing improvement and expansion of related functional modules, will establish the AM-DMF system as a powerful platform for SCSM. This platform is expected to support a wide range of applications in single-cell research and extend its utility to various research areas, such as biological sciences and chemistry.
Methods and materials
Platform and reagents
The AM-DMF system (DM sys) was developed by Guangdong ACXEL Micro & Nano Tech (Foshan, China) and ACX Instruments Ltd. (Cambridge, UK) (as shown in Fig. S4). In this study, the AM-DMF chip contains 16,384 electrodes (128 × 128), with an electrode pitch of 250 μm and a two-plate gap size of 50 μm (as shown in Fig. S5). A control voltage of 45 V was applied to the droplets throughout the experiments. The medium oil used in this study was silicone oil (2 cSt) obtained from Dow Corporate. Human cervical cancer (HeLa) cells were obtained from Cellverse Bioscience Technology Co., Ltd. The HeLa cells were cultured in a cell culture incubator (5% CO2, 75% atmosphere, 37 °C). The concentration of the HeLa cells was 4.5 × 105 cells/ml.
Data collection and composition
In this work, the original dataset used for training and testing the cell recognition model was obtained from images and videos. After the droplets were split via the AM-DMF chip, a microscope with a 7.5-fold objective lens was used to observe the cell-containing droplets. High-magnification microscope with pixel resolutions of 2448 × 2048 captured the cell images. We focused on fields of view with resolutions of 2048 × 2048 or 1280 × 1280 for analysis purposes. The images were displayed and saved using custom software. The exposure time was set to 20 ms, and the gain was set to 500%. We randomly divided the samples, allocating 70% to the training set and 30% to the test set. A comprehensive overview of the dataset is provided in Table 1, which includes the total number of labels and images for the cell detection models. The manual dataset annotations, which categorized droplets, cells, and oil bubbles, consisted of 4940 images, 4950 droplet labels, 7102 cell labels, and 22,718 bubble labels. The training set consisted of 3458 images, with 3465 droplet category labels, 4945 cell category labels, and 15,786 bubble category labels. Moreover, the testing set included 1482 images, with 1485 droplet category labels, 2157 cell category labels, and 6932 bubble category labels.
For the DPG model dataset, we initially prepared 140,000 data instances, comprising 50,000 droplet movement paths, 50,000 droplet split paths, and 40,000 combined droplet movement and split paths. After filtering out the noncompliant data, we retained a total of 139,701 instances for training and testing the DPG model. All droplet paths and prompts were saved in text files. Finally, these 139,701 sets of droplet paths and prompts were organized into a single JSON file format to create a conversational-style dataset through postprocessing.
Model architecture
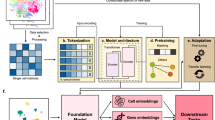
Figure S6 shows the detailed architecture of the three-class detector. The model’s output stage provides class probabilities for droplets, cells, and oil bubbles. The inference time of the cell recognition model is approximately 350 ms on an Intel Core i7 processor. Further details regarding the cell recognition model can be found in our previous work38. GPT-4o (Omni) was launched in May 2024 and developed by OpenAI52. Gemini was launched in December 2023 and developed by the Gemini Team, Google53,54. DeepSeek-V3 was launched in December 2024 and developed by Hangzhou Deepseek Artificial Intelligence Basic Technology Research Co., Ltd.55. As shown in Fig. S7, the DPG model was developed on the basis of the Llama 3 architecture and the LoRA technique. Llama 3 is an open-source LLMs based on a decoder transformer architecture that was launched in April 2024 by Meta56. The LoRA is an LLMs fine-tuning technique that freezes the weights of a pretrained LLMs and injects a trainable low-rank matrix factorization into each layer of the transformer architecture57. This approach reduces the number of trainable parameters for the droplet path generation task. On the Nvidia GeForce RTX 4090, the time from user input to the first token output is approximately 2 s. After that, the DPG model outputs paths line by line, where each line corresponds to one step of movement for all droplets. The DPG model generates each line of the path faster than the time required for each step of the droplet movement.
Model training parameter setup
The cell recognition model was implemented via Python 3.9.0 and the PyTorch 2.0.1 framework on a machine equipped with a 24-GB Nvidia GeForce RTX 4090 graphics card. The total number of parameters in the cell recognition model is 8,625,065. The DPG model was implemented via Python 3.12.0 and the PyTorch 2.3.1 framework on a machine equipped with two 24 GB Nvidia GeForce RTX 4090 GPUs. All the models were implemented on an Intel® X®(R) Gold 6133 CPU @ 2.50 GHz with 125 GB of RAM and run in the Ubuntu 20.04.5 LTS environment. The parameters of the Llama 3-8B model are 8,051,232,768. In the DPG model, the rank of the LoRA was set to 8. Consequently, the DPG model comprises 20,971,520 trainable parameters, representing 0.26% of the total number of parameters. During the training process, the other experimental parameter settings for the cell recognition model and DPG model (hyperparameters of LORA) were as shown in Table S10.
Model performance evaluation metrics
The research community often relies on average precision (AP) as the primary performance comparison metric for object detection models. The AP metric is derived from precision and recall. Precision is determined by the number of true positives divided by the sum of the true positives and false positives, whereas recall is calculated as the number of true positives divided by the sum of the true positives and false negatives. AP50 is the AP at an intersection over union (IoU) threshold of 0.5; AP75 is the AP at an IoU threshold of 0.75; AP50:95 is the AP at an IoU ranging from 0.5-0.95; the mAP for object detection is the mean of the AP over all object classes. The AP is defined as:
The precision and recall metrics are defined as follows
where TP represents true positives, FP represents false positives, and FN represents false negatives.
Data availability
The authors declare that all relevant data are available in the paper and its Supplementary Information Files, or from Z.J. (jiazq@sibet.ac.cn) on request.
Code availability
The program code is available for download from GitHub.
References
Kanton, S. et al. Organoid single-cell genomic atlas uncovers human-specific features of brain development. Nature 574, 418–422 (2019).
Luecken, M. D. et al. Benchmarking atlas-level data integration in single-cell genomics. Nat. Methods 19, 41–50 (2022).
Aldridge, S. & Teichmann, S. A. Single cell transcriptomics comes of age. Nat. Commun. 11, 4307 (2020).
Piwecka, M., Rajewsky, N. & Rybak-Wolf, A. Single-cell and spatial transcriptomics: deciphering brain complexity in health and disease. Nat. Rev. Neurol. 19, 346–362 (2023).
Marx, V. A dream of single-cell proteomics. Nat. Methods 16, 809–812 (2019).
Gatto, L. et al. Initial recommendations for performing, benchmarking and reporting single-cell proteomics experiments. Nat. Methods 20, 375–386 (2023).
Li, H. et al. Droplet-based microfluidics with mass spectrometry for microproteomics. Engineering 43, 37–53 (2024).
Sun, M., Chen, X. & Yang, Z. Single cell mass spectrometry studies reveal metabolomic features and potential mechanisms of drug-resistant cancer cell lines. Anal. Chim. Acta 1206, 339761 (2022).
Nguyen, T. D. et al. Single-cell mass spectrometry enables insight into heterogeneity in infectious disease. Anal. Chem. 94, 10567–10572 (2022).
Baysoy, A., Bai, Z., Satija, R. & Fan, R. The technological landscape and applications of single-cell multi-omics. Nat. Rev. Mol. Cell Biol. 24, 695–713 (2023).
Guo, Z. et al. Deep learning‐assisted label‐free parallel cell sorting with digital microfluidics. Adv. Sci. 12, 2408353 (2025).
Yang, Z. et al. AM-DMF-SCP: integrated single-cell proteomics analysis on an active matrix digital microfluidic chip. JACS Au 4, 1811–1823 (2024).
Hua, X. et al. Light-field flow cytometry for high-resolution, volumetric and multiparametric 3D single-cell analysis. Nat. Commun. 15, 1975 (2024).
Park, J., Park, C., Sugitani, Y., Fujii, T. & Kim, S. H. An electroactive microwell array device to realize simultaneous trapping of single cancer cells and clusters. Lab Chip 22, 3000–3007 (2022).
Joensson, H. N. & Andersson Svahn, H. Droplet microfluidics—a tool for single‐cell analysis. Angew. Chem. Int. Ed. 51, 12176–12192 (2012).
Huang, K.-W., Wu, Y.-C., Lee, J.-A. & Chiou, P.-Y. Microfluidic integrated optoelectronic tweezers for single-cell preparation and analysis. Lab Chip 13, 3721–3727 (2013).
Lamanna, J. et al. Digital microfluidic isolation of single cells for-Omics. Nat. Commun. 11, 5632 (2020).
Peng, J. et al. All-in-One digital microfluidics pipeline for proteomic sample preparation and analysis. Chem. Sci. 14, 2887–2900 (2023).
Choi, K., Ng, A. H., Fobel, R. & Wheeler, A. R. Digital microfluidics. Annu. Rev. Anal. Chem. 5, 413–440 (2012).
Dong, W. et al. Digital microfluidics with integrated Raman sensor for high-sensitivity in-situ bioanalysis. Biosens. Bioelectron. 271, 117036 (2025).
Xu, X. et al. Digital microfluidics for biological analysis and applications. Lab Chip 23, 1169–1191 (2023).
Hu, S. et al. Large-area electronics-enabled high-resolution digital microfluidics for parallel single-cell manipulation. Anal. Chem. 95, 6905–6914 (2023).
Zhou, T. et al. Self-interference digital optofluidic genotyping for integrated and automated label-free pathogen detection. ACS Sens. 9, 6411–6420 (2024).
Ma, H. et al. Large-area manufacturable active matrix digital microfluidics platform for high-throughput biosample handling. In 2020 IEEE International Electron Devices Meeting (IEDM) 35.5.1–35.5.4 (IEEE, 2020)
Hadwen, B. et al. Programmable large area digital microfluidic array with integrated droplet sensing for bioassays. Lab Chip 12, 3305–3313 (2012).
Noh, J. H., Noh, J., Kreit, E., Heikenfeld, J. & Rack, P. D. Toward active-matrix lab-on-a-chip: programmable electrofluidic control enabled by arrayed oxide thin film transistors. Lab Chip 12, 353–360 (2012).
Yu, J. et al. Field programmable digital microfluidics chip for high-throughput droplet array manipulation. In 2023 International Electron Devices Meeting (IEDM) 1–4 (IEEE, 2023).
Yip, M. et al. Artificial intelligence meets medical robotics. Science 381, 141–146 (2023).
Theodoris, C. V. et al. Transfer learning enables predictions in network biology. Nature 618, 616–624 (2023).
Chen, A. I., Balter, M. L., Maguire, T. J. & Yarmush, M. L. Deep learning robotic guidance for autonomous vascular access. Nat. Mach. Intell. 2, 104–115 (2020).
Gardner, K. et al. Deep learning detector for high precision monitoring of cell encapsulation statistics in microfluidic droplets. Lab Chip 22, 4067–4080 (2022).
Siemenn, A. E. et al. A machine learning and computer vision approach to rapidly optimize multiscale droplet generation. ACS Appl. Mater. Interfaces 14, 4668–4679 (2022).
Chu, A. et al. Automated detection and sorting of microencapsulation via machine learning. Lab Chip 19, 1808–1817 (2019).
Arjun, A., Ajith, R. & Kumar Ranjith, S. Mixing characterization of binary-coalesced droplets in microchannels using deep neural network. Biomicrofluidics 14, 034111 (2020).
Anagnostidis, V. et al. Deep learning guided image-based droplet sorting for on-demand selection and analysis of single cells and 3D cell cultures. Lab Chip 20, 889–900 (2020).
Sesen, M. & Whyte, G. Image-based single cell sorting automation in droplet microfluidics. Sci. Rep. 10, 8736 (2020).
Howell, L., Anagnostidis, V. & Gielen, F. Multi‐Object detector yolov4‐tiny enables high‐throughput combinatorial and spatially‐resolved sorting of cells in microdroplets. Adv. Mater. Technol. 7, 2101053 (2022).
Jia, Z. et al. Artificial intelligence-enabled multipurpose smart detection in active-matrix electrowetting-on-dielectric digital microfluidics. Microsyst. Nanoeng. 10, 139 (2024).
Zhao, Y. & Chakrabarty, K. Simultaneous optimization of droplet routing and control-pin mapping to electrodes in digital microfluidic biochips. IEEE Trans. Comput. Aided Des. Integr. Circuits Syst. 31, 242–254 (2012).
Keszocze, O., Wille, R., Chakrabarty, K. & Drechsler, R. A general and exact routing methodology for digital microfluidic biochips. In 2015 IEEE/ACM International Conference on Computer-Aided Design (ICCAD) 874–881 (IEEE, 2015).
Yuh, P.-H., Yang, C.-L. & Chang, Y.-W. BioRoute: a network-flow-based routing algorithm for the synthesis of digital microfluidic biochips. IEEE Trans. Comput. Aided Des. Integr. Circuits Syst. 27, 1928–1941 (2008).
Huang, T.-W. & Ho, T.-Y. A fast routability-and performance-driven droplet routing algorithm for digital microfluidic biochips. In 2009 IEEE International Conference on Computer Design 445-450 (IEEE, 2009).
Liang, T.-C. & Zhong, Z. Adaptive droplet routing in digital microfluidic biochips using deep reinforcement learning. In Proceedings of the 37th International Conference on Machine Learning 6050–6060 (PMLR, 2020).
Jiang, C. et al. A cooperative multiagent reinforcement learning framework for droplet routing in digital microfluidic biochips. IEEE Trans. Comput. Aided Des. Integr. Circuits Syst. 42, 3007–3020 (2022).
Boiko, D. A., MacKnight, R., Kline, B. & Gomes, G. Autonomous chemical research with large language models. Nature 624, 570–578 (2023).
Lin, Z. et al. Evolutionary-scale prediction of atomic-level protein structure with a language model. Science 379, 1123–1130 (2023).
Szymanski, N. J. et al. An autonomous laboratory for the accelerated synthesis of novel materials. Nature 624, 86–91 (2023).
Slattery, A. et al. Automated self-optimization, intensification, and scale-up of photocatalysis in flow. Science 383, eadj1817 (2024).
Ruiz-Gutiérrez, É. & Ledesma-Aguilar, R. Lattice-Boltzmann simulations of electrowetting phenomena. Langmuir 35, 4849–4859 (2019).
Staicu, A. & Mugele, F. Electrowetting-induced oil film entrapment and instability. Phys. Rev. Lett. 97, 167801 (2006).
Collins, D. J., Neild, A., DeMello, A., Liu, A.-Q. & Ai, Y. The Poisson distribution and beyond: methods for microfluidic droplet production and single cell encapsulation. Lab Chip 15, 3439–3459 (2015).
Hurst, A. et al. Gpt-4o system card. Preprint at arXiv https://arxiv.org/abs/2410.21276 (2024).
Team, G. et al. Gemini: a family of highly capable multimodal models. Preprint at arXiv https://arxiv.org/abs/2312.11805 (2023).
Team, G. et al. Gemini 1.5: Unlocking multimodal understanding across millions of tokens of context. Preprint at arXiv https://arxiv.org/abs/2403.05530 (2024).
Liu, A. et al. Deepseek-v3 technical report. Preprint at arXiv https://arxiv.org/abs/2412.19437 (2024).
Grattafiori, A. et al. The llama 3 herd of models. Preprint at arXiv https://arxiv.org/abs/2407.21783 (2024).
Hu, E. J. et al. Lora: Low-rank adaptation of large language models. Preprint at arXiv https://arxiv.org/abs/2106.09685 (2021).
Acknowledgements
This research was funded by the National Key R&D Program of China (2023YFF0721500). The National Natural Science Foundation of China (Nos. 62374102, 82172077, 22201298, and 62027825). The Innovation and Entrepreneurship Team of Jiangsu Province (JSSCTD202145). The Science and Technology Innovation Project of Foshan, Guangdong Province, China (No. 1920001000047). The Science and Technology Development Project of Jilin Province (No. 20210204110YY and 20250204092YY). The Suzhou Basic Research Project (SSD2023013).
Author information
Authors and Affiliations
Contributions
Z.J., C.J., W.D., Z.Y. and H.M. conceived the concept and experiments. Z.J., J.L., L.L., T.H., B.Y., S.H., X.H., T.H. and M.G. performed the research. Z.J., C.J., J.L., B.Y., L.L., M.G. and S.H. analyzed the data. Z.J., C.J., J.L., X.H., T.H., Z.Y., W.D. and H.M. wrote the paper.
Corresponding authors
Ethics declarations
Conflict of interest
Z.J. is a PhD student at Changchun University of Science and Technology and is currently a visiting student at the Chinese Academy of Sciences and the Chinese University of Hong Kong. Z.J. contributed to this work while interning part-time at Guangdong ACXEL Micro & Nano Tech Co., Ltd. W.D. is a PhD supervisor at Changchun University of Science and Technology.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Jia, Z., Jiang, C., Li, J. et al. Intelligent single-cell manipulation: LLMs- and object detection-enhanced active-matrix digital microfluidics. Microsyst Nanoeng 11, 133 (2025). https://doi.org/10.1038/s41378-025-00962-y
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41378-025-00962-y
This article is cited by
-
Active-matrix digital microfluidics for high-throughput, precise droplet handling
Nature Reviews Electrical Engineering (2025)