Abstract
DDX3X, a DEAD-box RNA helicase, has been identified as a risk gene for autism spectrum disorder (ASD). To elucidate the role of DDX3X mutations in ASD pathogenesis, HT22 cell models and mouse models with Ddx3x knockdown specifically in the medial prefrontal cortex were established. Ddx3x knockdown in HT22 cells resulted in slower growth, while in mice, it induced autism-like behaviors. Proteomic analysis in cortex revealed that many down-regulated proteins were involved in synaptic plasticity. The differentially expressed proteins (DEPs) were associated with long-term potentiation, glutamatergic and GABAergic synapses, postsynaptic density, branched-chain amino acid degradation, and the oxytocin pathway. Integrating cellular and cortical omics studies revealed overlaps in the ubiquitin–proteasome system and the ‘de novo’ protein folding pathway. These pathways were inhibited in the cortex of the model mice. The expression of several important proteins was validated. Confocal microscopy and transmission electron microscopy demonstrated a significant reduction in dendritic spine density and postsynaptic density of mouse models. Electrophysiological experiments showed that the frequency of miniature excitatory postsynaptic currents was significantly reduced. These results suggest that DDX3X deficiency impairs synaptic function, thereby affecting neurodevelopment and social abilities. Abnormal synaptic plasticity may contribute to the pathogenesis of ASD with DDX3X gene mutations.
Similar content being viewed by others
Introduction
Autism spectrum disorder (ASD) is a heterogeneous group of neurodevelopmental syndromes in which individuals have deficits in social communication and the manifestation of limited and repetitive behaviors [1, 2]. A significant proportion of ASD individuals may have complications such as intellectual disability (ID), global developmental delay, and epilepsy, accompanied by ultra-rare genetic variants [3]. In the past two decades, the prevalence of ASD has continued to rise, and it has become a serious global public health problem.
ASD is highly heritable, as evidenced by its increased prevalence among relatives of individuals with ASD compared to the general population [4, 5]. With the advancement of genomics research, over 1200 genes have been identified as susceptibility genes for ASD in the SFARI Gene database (https://www.sfari.org/resource/sfari-gene/). More than 100 risk genes have been confirmed, and the discovery of new genes is ongoing [6]. Despite these significant achievements, several challenging questions remain. Firstly, the precise relationship between these identified genes and the pathogenesis of ASD requires further elucidation [7, 8]. Secondly, given the genetic heterogeneity of ASD, it is crucial to identify common characteristics or underlying mechanisms that may link these diverse genetic factors [9,10,11]. Moreover, the occurrence of ASD is also associated with environmental factors, the interaction between genetic and environmental factors, and epigenetic regulation [9, 12, 13].
In the previous studies, we identified an ASD child carrying a DDX3X risk variant via targeted sequencing [14, 15]. DDX3X, also known as DBX or DDX3, is a DEAD-box helicase involved in translation initiation and multiple stages of RNA metabolism, with critical roles in neurite growth, synaptogenesis, and cortical neurogenesis [16,17,18]. Mutations in DDX3X occur in ~1–3% of individuals with unexplained ID [19,20,21], and large-scale exome sequencing studies have implicated DDX3X in severe developmental disorders [19, 22, 23]. DDX3X syndrome is now recognized as a neurodevelopmental disorder associated with ASD, ID, and various comorbidities [24], with loss-of-function mutations classified as ASD risk variants [25].
Existing studies have confirmed that DDX3X is a core risk gene for ASD. Deficiency in DDX3X disrupts neurodevelopment through multiple mechanisms. These include impaired RNA helicase activity [18], altered translation of synaptic mRNAs [18, 26], and reduced crebbp mRNA stability [16]. Additionally, deletion of Ddx3x leads to embryonic lethality in mice, which is associated with its critical role in placentation and embryogenesis [17]. At the cellular level, Ddx3x depletion prolongs neural progenitor cell cycles, reduces neurogenic divisions, and disrupts the excitatory/inhibitory balance [16, 27], while knockout in neural stem cells impairs hindbrain development [28]. Phenotypically, DDX3X-deficient mice and zebrafish exhibit ASD behaviors and brain abnormalities (e.g., microcephaly) [16, 29].
Notably, although DDX3X syndrome is more common in females, males with hypomorphic missense mutations also show neurodevelopmental deficits (e.g., ID, autistic features) [17, 30]. Critically, Ddx3y (Y-homolog) is lowly expressed in the mouse brain and unable to fully compensate for Ddx3x loss [28, 30], while DDX3X’s partial escape from X-chromosome inactivation (XCI) leads to mosaic expression in female tissues [17, 29]. Combined with female estrous cycle-induced behavioral variability [31, 32], we thus used male mice and targeted the mPFC—a key region for ASD-related core functions including social behavior, cognition, and memory [33, 34]—to establish a Ddx3x knockdown mouse model. We also constructed an in vitro Ddx3x knockdown cellular model. Subsequently, we combined behavioral, proteomic, biochemical, and electrophysiological approaches to investigate the link between DDX3X deficiency and ASD as well as the underlying mechanisms.
Material and methods
Cell culture and lentivirus transfection
The experimental flow of this study is shown in Fig. 1. The mouse hippocampal neuron cells (HT22) purchased from Wuhan university cell bank (Wuhan, China). HT22 cells were cultured in high-glucose DMEM medium with 10% fetal bovine serum and placed in in a stable-temperature incubator at 37 °C, 5% CO2, and saturated humidity. The Ddx3x-knockdown and empty lentivirus expressed green fluorescent protein (GFP) were purchased from Jikai Gene (Shanghai, China). The sequence of RNA interference screen for the knockdown of Ddx3x and GCAAGCAAAGGGCGTTATATC was selected as targeted sequence (Supplementary Fig. 1). The Ddx3x-knockdown lentivirus was seeded in HT22 cells with an MOI (multiplicity of infection) of 8 as sh-DDX3X group, and the empty lentivirus was used as control group. The cell fluorescence was observed with a fluorescence microscope after 24–48 h. The cells were counted manually. The cells were harvested after 96 h.
FASP: filter-aided sample preparation, mPFC: medial prefrontal cortex, iTRAQ: isobaric tags for relative and absolute quantitation, SWATH: sequence window to obtain all theoretical fragment ions, TEM: transmission electron microscope, mEPSC: miniature excitatory postsynaptic currents. This figure panel was created using BioRender.
Animals and stereotaxic surgeries
Since previous neuroimaging studies on functional connectivity suggested that social task-related is lack of activation of medial prefrontal cortex (mPFC) [35, 36]. Here, the mPFC brain region of C57/B6J mice was treated with adeno-associated virus (AAV) to reduce DDX3X levels and generate an ASD-like phenotype mouse. Wild type (WT, C57BL/6J strain) were obtained from Bestest Biotechnology company (Zhuhai, China). All the animals were housed in an air-conditioned room (22 ± 2 °C, 12-h light, and 12-h dark cycle) with food and water ad libitum. The recombinant AAV including AAV-hSyn-EGFP-Ddx3x-RNAi (103688-1) and AAV-hSyn-EGFP-MCS were manufactured by Jikai Gene (Shanghai, China). The target sequence for siRNAs is GCAAGCAAAGGGCGTTATATC. The hSyn promoter, derived from the human SYN1 gene, is known for its specificity in driving gene expression in neurons [37, 38]. Male 20-days-old WT mice were randomly divided into two groups. The mice (n = 18, male) in control group were injected AAV-hSyn-EGFP-MCS and the mice (n = 20, male) in sh-DDX3X group were injected AAV-hSyn-EGFP-Ddx3x-RNAi (103688-1). The mice were anesthetized with isoflurane (3% initial dose, 1.5% maintenance dose) and the mouse brains were injected with 2 µL of the AAV at a rate of 0.25 µL/min in mPFC (from Bregma: AP, + 1.75 mm; ML, ± 0.30 mm; DV, −1.75 mm). Before behavioral tests, mice were allowed to grow to 9-weeks-old of age. Injection sites were confirmed at the end of the behavior experiments and incorrect injection placement were excluded.
Mouse behavioral tests
The open field test
For the open field test, mice were placed in an open field apparatus. Mice were recorded with a video camera for 10 min, and the total distance traveled was calculated using a camera and further analyzed. This was to exclude confounding factors like motor dysfunction and excessive anxiety, ensuring subsequent observed social abnormalities were specific to ASD-related deficits.
Three-chamber social test
The three-chamber social test was performed as previously described with minor modifications [39, 40], which is a classic assay for evaluating core social deficits in ASD rodent models. Time spent in close interaction with a stranger mouse were recorded for analysis. Target subjects (stranger 1 and stranger 2) were 9-week-old male mice habituated to being placed inside the cages for a week before the test. (A) Habituation: the test mouse was placed in the middle chamber and allowed to explore for 10 min. (B) Sociability: an age- and sex-matched stranger 1 was introduced to the wire cage located on one side. An inanimate object was introduced on opposite side. The test mouse was placed in the center chamber and allowed to explore the entire apparatus for 10 min. (C) Social novelty: a new stranger 2 was replaced to object. The test mouse spent another 10 min in exploring the entire apparatus.
Y-maze test
This assay was used to assess spatial recognition memory, a common cognitive comorbidity in ASD. Mice were food-deprived to 85% of their free-feeding body weight before undergoing the Y-maze test [41]. The Y-maze apparatus included a start arm and two new arms enclosed by a 10 cm high wall. (A) Mice were habituated for 2 days to obtain food from the ends of the two arms. (B) Training trials were conducted three times daily for 4 days. During forced runs, mice were placed in the start arm with one arm blocked and the other open for food access. In test runs, mice chose between the two arms after the block was removed, with correct choices rewarded by food consumption and incorrect choices unrewarded. (C) In tested trials, the interval between forced and test runs was extended to 1 min and 3 min.
Social interaction with a juvenile mouse
This assay was used to quantify micro-level social behaviors and further confirm ASD-related social recognition impairments. An individual tested mouse was placed in a clean and empty arena; stimulus animals were 20 days old male wild-type C57BL/6 J mice. Subjects and stimulus partners were individually housed in a clean cage for 1 h before the test. They were allowed to freely interact for 10 min and recorded behaviors. Parameters were routinely evaluated include nose-to-nose sniffing, nose-to-anogenital sniffing, and following.
Marble burying test (MBT)
To assess ASD’s core restricted, repetitive behaviors, the marble burying test was conducted. Experiments were conducted in a 29 × 18 × 15 cm cage with a uniform layer of cereal bedding to a depth of 5 cm. Each mouse was first allowed to explore the cage for 10 min without marbles, and then 15 glass marbles were spaced evenly in a 3 × 5 grid. Each mouse was recorded for 30 min and the glass marbles buried in the bedding (to 2/3 of their depth) were counted [42].
Proteomics analysis
In proteomics research, we used iTRAQ (isobaric tags for relative and absolute quantitation) and SWATH (sequence window to obtain all theoretical fragment ions) proteomics methods for the study of Ddx3x knockdown HT22 cells and mouse mPFC tissues, respectively.
Proteomic study of DDX3X gene interference cell model based on iTRAQ labeling
iTRAQ analysis was performed as previously described [43]. HT22 cell lysates were prepared using RIPA buffer (Beyotime, China). Proteins from control (n = 3) and sh-DDX3X (n = 3) groups were digested with trypsin, labeled with iTRAQ tags (controls: 115, 116, 117; sh-DDX3X: 118, 119, 121), and mixed. The samples were analyzed using a Triple TOF 6500 system (AB Sciex, Foster City, CA) in data-dependent acquisition (DDA) mode. Protein identification and quantification were conducted using ProteinPilot v4.5 (AB Sciex) against the UniProt Mus musculus database, with a false discovery rate (FDR) < 1 and a 95% confidence level. Proteins were identified with at least one unique peptide and quantified with at least two peptides.
Proteomic analysis of mPFC tissue by SWATH-MS: protein extraction and filter-aided sample preparation (FASP) digestion
The mouse from sh-DDX3X (n = 8) and control group (n = 8) were anesthetized by isoflurane at 14-weeks-old after behavior test. Sequentially, they were euthanized by cervical dislocation and the brains were collected. The mPFC tissue was dissected from coronal sections using anatomical landmarks and snaped frozen in liquid nitrogen. The mPFC tissue were lysed with RIPA lysis buffer (Beyotime, No. P0013B) and were homogenized with a freeze grinder (Jingxin, Shanghai, China) for 3 min, 90 hz. After incubation in 4 °C for 1 h, the samples were centrifuged at 12,000 × g and collect the supernatant to determine protein concentration with a BCA kit (KeyGen Biotech, China, No. KGP903). Trypsin digestion was performed using a previously described FASP method with slight modifications [44].
Information dependent acquisition (IDA) mode to create reference spectral library for SWATH
A library of reference spectra was required for SWATH-mass spectrum (MS) analysis. We collected 20 μg of peptides from each sample to obtain a pooled sample of all samples for quality control (QC). A liquid phase system was used to carry out liquid chromatography (LC) to collect peptides at 1-min intervals in 20 fractions. The data were acquired by IDA mode in Triple TOF 6600 (AB Sciex) mass spectrometer. Following each survey scan the top 40 most intense ions with multiple charged ions (+2– + 5) sequentially.
SWATH-MS analysis
A SWATH-MS analysis was performed in IDA mode, and peptides were separated after enzymatic hydrolysis for each sample [45, 46]. During the IDA detection process, samples from two groups were tested alternately. In subsequent quality control analyses, QC samples were detected every five samples. Following the acquisition process, the ParagonTM algorithm process IDA data files by ProteinPilot version 5.0 (AB Sciex), while SWATH data files undergo processing via PeakView software (SWATH acquisition Micro APP 2.0).
Analysis of proteomic characteristics and identification of differentially expressed proteins (DEPs) between groups
Data were analyzed using ProteinPilot v4.5 (AB Sciex). Principal coordinates analysis (PCoA) was conducted by R package Vegan (2.6–4) and partial least squares discriminant analysis (PLS-DA), orthogonal partial least squares discriminant analysis (OPLS-DA), and permutation test were performed using the SIMCA-P software (version14.1, Sartorius Stedim Data Analytics AB, Umea, Sweden). The DEPs were calculated using the R package Limma (v3.48.3). The cutoff value for up-regulation was 1.2-fold change and for down-regulation was 0.83- fold change, and P-value < 0.05 was established for DEPs between sh-DDX3X and control group.
Bioinformatic analysis
Gene ontology (GO), Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis, and protein- protein interaction (PPI) network were analyzed by R package clusterProfiler (v4.2.1), STRING (https://string-db.org/) and Cytoscape software (V 3.8.2). The Volcano plot and heatmaps were drawn using the ggplot2 (v3.3.5) and R package pheatmap (v1.0.12). The sunburst plots were generated based on synaptic gene ontologies (https://syngoportal.org). The hub proteins were calculated by CytoHubba plugin and R package RobustRankAggreg (v1.1).
Western blot analysis
Protein samples were boiled for 10 min in loading buffer and separated by electrophoresis. Proteins were then transferred to a polyvinylidene fluoride (PVDF) membrane (Millipore, No. IPVH00010, USA) and blocked with 5% non-fat milk in tris-buffered saline (TBS: 100 mM Tris, and 1.5 M NaCl, pH 7.6) containing 0.4% (v/v) tween 20 (TBST) for 1 h. The membrane was incubated overnight at 4 °C with primary antibodies against DDX3X (Abcam, No. ab271002, 1:2000, UK), MAPK1 (Abclonal, No. A0229, 1:1000, China), CAMKII-delta (Proteintech, No. 20667-1-AP, 1:1000, China), NMDAR (Huabio, No. JM11-26, 1:600, China), PSD95 (Proteintech, No. 20665-1-AP, 1:1000), PRRT1 (Proteintech, No. 66031-1-lg, 1:600), GAPDH (Abmart, No. M20050, 1:5000, China), TUBA (Proteintech, No. 66031-1-lg, 1:10000), and VCL (Proteintech, No. 66305-1-lg, 1:15000). Secondary antibodies included anti-rabbit (Beyotime, No. A0208, 1:1000, China) and anti-mouse (Abmart, No. M21001, 1:5000, China), incubated for 1 h at room temperature. Protein expression levels were normalized to α-tubulin or vinculin. Specific proteins were detected using chemiluminescence with horseradish peroxidase (HRP) substrate (Millipore, No. WBKLS0100), and band intensity was quantified by densitometry using ImageJ software (Bio-Rad, Hercules, CA).
Immunofluorescence detection of representative proteins, morphological analysis of dendritic spines, and the detection of synaptic ultrastructure
Mice were perfused with PBS (phosphate buffered saline) and fixed with 4% PFA (paraformaldehyde) under anesthesia. Brains were postfixed in 4% PFA overnight at 4 °C and dehydrated in 30% sucrose. Coronal sections (20 μm) were prepared from OCT (optimal cutting temperature compound (Sakura, Japan))-embedded brains using a cryostat (Leica CM 3050S). Sections were washed with PBS, permeabilized with 0.3% Triton-X 100 for 30 min, and blocked with PBS containing 1% BSA, 0.1% Triton-X 100, and 10% normal goat serum. Sections were incubated with primary antibodies against DDX3X, MAPK1, and PSD95 overnight at 4 °C, followed by incubation with a fluorescence-conjugated secondary antibody (Abcam, No. ab150088, 1:200) for 2 h at room temperature. Nuclei were counterstained with DAPI (4’,6-Diamidino-2-phenylindole; Beyotime, No. C1002, 1:2000) for 3 min. Confocal images were captured using a Zeiss LSM880 microscope.
For the spine analysis, dendritic were randomly chosen from AAV infected neurons expressing GFP and spines on dendrites were identified and counted in the orthogonal projection images using software Zen blue (ZEISS). Counting of dendritic spines along traced dendrites was performed manually. Stubby spines were defined as short protrusions without a neck, thin as long protrusions without a wide head, and mushroom spines as protrusions with a short neck and a distinguishable head.
To observe ultramicroscopic structure, samples were fixed in 2.5% glutaraldehyde (pH 7.4) for 2 h, washed with 0.1 M phosphate buffer (pH 7.2), and postfixed in 1% osmium tetroxide at 4 °C for 2 h. After gradient ethanol dehydration, samples were embedded in Epon-Araldite resin, sectioned, and stained with 3% uranyl acetate and 2.7% lead citrate. Ultrathin sections were analyzed using a JEM1400 transmission electron microscope (Hitachi, Japan).
Electrophysiology for miniature excitatory postsynaptic currents (mEPSCs)
In brief, mice were deeply anesthetized and transcardially perfused with 95% O2 and 5% CO2 oxygenated ice-cold cutting solution. The brain was quickly removed and immersed in ice-cold cutting solution. Coronal brain slices containing mPFC (250 μm) were cut with a Leica VT1200s vibratome and then maintained at 25 °C in oxygenated artificial cerebrospinal fluid (ACSF) for 1 h until electrophysiological recordings.
Slices were transferred to the recording chamber with oxygenated ACSF (~ 3 mL/min). Neurons in the mPFC were identified by their location and fluorescence. For whole-cell recording of mEPSCs, pipettes were filled with a solution containing: 135 mM K-gluconate, 5 mM KCl, 0.5 mM CaCl2, 10 mM HEPES, 2 mM Mg-ATP, 0.1 mM GTP, and 5 mM EGTA, 300 mOsm (pH adjusted to 7.3 with KOH). Bicuculline (10 μM) and TTX (1 μM) were used to block the GABAA receptors and Na+ channel, respectively. Peak events were detected automatically using an amplitude threshold of 10 pA. The holding potential was −70 mV during mEPSC recording.
All signals were acquired with a MultiClamp 700B amplifier (Molecular Devices, USA), filtered at 1 kHz, and sampled at 5 kHz with a Digidata 1440 A interface using Clampex 10.2 (Molecular Devices). Data were accepted when series resistance fluctuated within 15% of initial values (25–35 MΩ). mEPSCs were analyzed using MiniAnalysis 6.07 software (Synaptosoft, Decatur, GA). A 5-min recording duration was used for frequency and amplitude analyses.
Statistical analysis
Statistical analysis was calculated with GraphPad Prism Version 9.0 (GraphPad Software, San Diego, USA). Data were presented as the mean ± standard deviation (SD). The non-paired and 2-tailed of t test was applied to determine whether a significant (p < 0.05), and have ensured that the homogeneity of variance and normal distribution status of the data were in accordance with the testing requirements.
Results
The knockdown of Ddx3x decreased the growth rate of HT22 cells
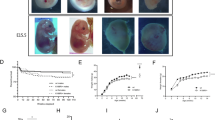
The sh-Ddx3x and control lentivirus infected HT22 cells successfully and expressed GFP (Fig. 2A). The protein expression level of DDX3X was significantly decreased in sh-DDX3X group (Fig. 2B). As shown in Fig. 2C, the processes of the cells in the shDDX3X group developed more slowly than those in the control group, and at the same incubation time, the cells in the control group already appeared to be connected, whereas the cells in the sh-DDX3X group still existed singly. The DDX3X deficiency induced cell growth rate significantly decreased compared to the control group.
A Images of HT22 cells infected by lentivirus and expressed GFP. B Representative images of HT22 cells growth for different time periods and quantification analysis. scale bar: 100 μm. C Western blot bands showing DDX3X protein levels in HT22 cells compared to control group. D PcoA plot between control and sh-DDX3X group. E PLS-DA score plot between control and sh-DDX3X. F PLS-DA validation plot (permutation = 200). G The volcano plot of DEPs from iTRAQ analysis. H The number of up-regulated and down-regulated DEPs from iTRAQ analysis. I The cluster heatmap of DEPs from iTRAQ analysis. WL: white light. sh: sh-DDX3X, CTR: control.
iTRAQ-based proteomic analysis of sh-Ddx3x HT22 cells and controls
iTRAQ analysis identified a total of 7861 proteins, of which 5451 were quantified (Supplementary Table 1). Unsupervised PCoA analysis revealed a general trend towards separation between the control and sh-DDX3X groups (Fig. 2D). The PLS-DA analysis showed that the two groups were clearly distinguishable, and the model exhibited robustness without overfitting (Figs. 2E and 2F). A total of 242 DEPs were identified between sh-Ddx3x HT22 cells and controls, including 114 up-regulated and 128 down-regulated proteins (Figs. 2G, 2H and Supplementary Table 2). Clustering analysis of these DEPs completely distinguished the sh-DDX3X group from the control group (Fig. 2I).
GO analysis indicated that the DEPs were primarily involved in biological processes (BPs) such as protein folding, tRNA metabolic process, Golgi organization, and chaperone cofactor-dependent protein refolding. In terms of molecular function (MF), significant changes were observed in catalytic activity, single-stranded telomeric DNA binding, chaperone binding, and aminoacyl-tRNA ligase activity (Supplementary Fig. 2). KEGG analysis revealed enrichment in pathways including metabolic pathways, mTOR signaling pathway, ErbB signaling pathway, aminoacyl-tRNA biosynthesis, and adherens junction (Supplementary Fig. 3).
Validation of shRNA AAV against Ddx3x gene in mPFC of C57/B6J mice
The Ddx3x shRNA AAV was delivered to mPFC bilaterally of C57/B6J mice (Fig. 3A). In vivo knockdown was confirmed by the significantly lower level of DDX3X protein in mPFC of sh-DDX3X group by Western blot analysis (Figs. 3B and C). The lower overlap DDX3X and GFP was found in neuron indicating the decrease of DDX3X (Figs. 3D and E).
A An image showing the viral-infected mPFC region (expressed GFP) from a mouse with the stereotaxic injection of Ddx3x shRNA AAV. Scale bar: 100 μm. B and C Western blot data showing DDX3X protein levels in mPFC of mice in sh-DDX3X group compared to control group. D The confocal images of immunostaining of DDX3X (red) in mPFC neurons infected with Ddx3x shRNA AAV. E The representative image of DDX3X in cell between sh-DDX3X and control group. Scale bar: 10 μm. F Representative motion map of control and sh-DDX3X group mice in open filed test. G Results of total distance, enter zone number, edge time, and edge distance in open filed. *p < 0.05, ** p < 0.01, *** p < 0.001, two-tailed t test.
Behavior test to study the sociability
To investigate the relationship between decreased DDX3X and social deficits, we conducted behavioral tests on the two groups. In the open field test, sh-DDX3X mice showed no significant differences in total distance traveled, number of zone entries, edge time, or edge distance compared to controls (Figs. 3F and G), indicating that reduced DDX3X levels did not affect mouse mobility or anxiety.
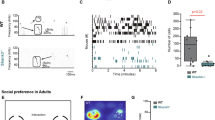
In the three-chamber social test, control mice exhibited a preference for a stranger mouse (S1) over a novel object, whereas sh-DDX3X mice showed equivalent interaction times with both (Fig. 4A–D). When a new stranger mouse (S2) replaced the object to test social novelty preference, control mice interacted more with S2 than with the familiar S1, while sh-DDX3X mice did not show a significant preference for S2 (Fig. 4E–G). These results suggest that both social preference and social novelty preference were impaired in the sh-DDX3X group.
A The flow diagram of three-chamber social test. B The interaction time in testing sociality. C Representative motion map of control and sh-DDX3X groups mice in testing sociality. D The preference index in testing sociality. E The interaction time in testing social novelty. F Representative motion map of control and sh-DDX3X groups mice in testing social novelty. G The preference index in testing social novelty. H The flow diagram of appetitively-motivated Y-maze test. I The correct rate of Y-maze test in differently interval time. J-O Results of number of nose-anogenital, time spent of nose-anogenital, number of social sniffing, time spent of social sniffing, number of followings, and time spent of following in juvenile mouse interaction test. P Results of marble burying test. In the three-chamber social test (B-G), the Y-maze (I), the social interaction with a juvenile mouse (J-O), the number of mice was Control:sh-DDX3X = 10:10, whereas in the marble burying test (P) this number was 8:9. *p < 0.05, ** p < 0.01, ***, p < 0.001, two-tailed t test.
Additionally, a food-restricted Y-maze test was used to assess spatial recognition memory. The correct rate between control and sh-DDX3X groups was similar within a 1-min interval, but sh-DDX3X mice exhibited decreased accuracy within a 3-min interval (Fig. 4H and I), indicating impaired spatial recognition memory in sh-DDX3X mice over longer intervals.
In social interaction tests with juvenile mice, sh-DDX3X mice displayed a significant reduction in nose-to-anogenital contacts compared to controls (Fig. 4J) and tended to initiate fewer such interactions (Fig. 4K). However, no significant differences were observed in social sniffing, time spent sniffing, following behavior, or time spent following between the two groups (Fig. 4L–O). Finally, in the marble burying test, sh-DDX3X mice buried more marbles than controls (Fig. 4P), suggesting increased repetitive behaviors in this group.
The results of proteomic analysis the medial prefrontal cortex in Ddx3x gene knockdown mice
A total of 3087 quantitative proteins were identified by SWATH-MS analysis, of which 2480 quantitative proteins met the quality control requirements after QC correction (Supplementary Table 3). In unsupervised PCoA analysis, there was some overlap between sh-DDX3X and control groups despite the general trend towards separation of protein profiles, and QC samples cluster tightly together indicating the reliability and stability of the data (Fig. 5A). The contribution of variables was characterized using the supervised OPLS-DA model, which showed that there were significant differences between proteins of sh-DDX3X and control group and the model was validated by 200 times permutation tests, which showed low Q2 values and a negative intercept of the regression line on the y-axis, indicating that the model was robust and not over-fitted (Figs. 5B and 5C).
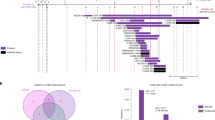
A PcoA plot among QC, control and sh-DDX3X group. B OPLS-DA score plot between control and sh-DDX3X. C OPLS-DA validation plot (permutation = 200). D The number of up-regulated and down-regulated DEPs. E The volcano plot of DEPs. F The cluster heatmap of DEPs. G The network plot of enriched pathways based on KEGG analysis. H The z-score of ASD-related pathways based on KEGG analysis. The positive and negative z-score means the pathway is activated and inhibited, respectively. I The PPI network among LTP, glutamatergic synapse, and GABAergic synapse. J The sunburst plot of DEPs in presynapse. K The sunburst plot of DEPs in postsynapse. L The PPI network from four term based on SynGO. CTR: control, sh:sh-DDX3X.
Volcano plot analysis revealed that a total of 285 DEPs were identified including 265 down-regulated proteins, and up-regulated 20 proteins (Figs. 5D, E, and Supplementary Table 4). These DEPs were used for clustering analysis, the results showed that the majority of sh-DDX3X group could distinguished from the control group (Fig. 5F).
GO analysis showed that DEPs, in BP network, have significant alteration in small molecule catabolic process, carboxylic acid catabolic process, response to oxidative stress, peptidyl-serine phosphorylation, and generation of precursor metabolites and energy, etc. Several network clusters in cellular components (CC) showed presynapse, growth cone, postsynaptic density, proteasome accessory complex, actomyosin, and mitochondrial matrix, etc. In MF network, the pathways including lyase activity, scaffold protein binding, ubiquitin protein ligase binding, and endopeptidase regulator activity, were significantly enriched in the sh-DDX3X group (Supplementary Fig. 4).
KEGG analysis showed that the DEPs were mostly enriched in valine, leucine and isoleucine degradation, carbon metabolism, long-term potentiation (LTP), Parkinson disease, and pathways of neurodegeneration - multiple diseases, cholinergic synapse, oxytocin signaling pathway, and GABAergic synapse, etc. The network clusters showed that LTP has strong correlation with other significant pathways (Fig. 5G).
We calculated the z-scores of KEGG pathways related to ASD, including valine, leucine, and isoleucine degradation, LTP, oxytocin signaling pathway, GABAergic synapse, long-term depression, ErbB signaling pathway, glutamatergic synapse, mTOR signaling pathway, synaptic vesicle cycle, and autophagy-animal pathway. These pathways were found to be inhibited in the sh-DDX3X group (Fig. 5H). The relationship of proteins involved in LTP, glutamatergic synapse, and GABAergic synapse was showed in PPI network, and PKCA, PRKACA, MAPK1, MAPK3, CACNA1A, and GNG4 were repeated among these pathways (Fig. 5I). The results suggest that a series of pathways were not sufficient activation with deficiency of DDX3X.
We further annotated the DEPs obtained from the mouse cortex using the synaptic protein annotation database. The results showed that 49 DEPs were mapped to 49 unique SynGO annotated genes. The sunburst plot showed that 23 DEPs were categorized into presynapse, and 9 DEPs were enriched in synaptic vesicle compared to control group (Fig. 5J). Twenty-eight (28) DEPs were categorized into postsynapse, and 11 DEPs were enriched in postsynaptic density (PSD) by DDX3X deficiency (Fig. 5K). The PPI network revealed the interplay between DEPs originating from the PSD, integral of postsynaptic membrane, postsynaptic ribosome, and postsynaptic actin cytoskeleton (Fig. 5L). These results indicated that the deficiency of Ddx3x expression in the mPFC impacted synaptic signaling and structures, which may underlie the abnormal behavioral phenotypes observed.
The shared dysregulated pathways observed in mouse brain and HT22 cells with decreased DDX3X expression
An integrated analysis of the proteomic data from HT22 cells and mouse cortex revealed that 9 proteins were commonly dysregulated, including Pebp1, Osbpl8, Echdc1, Pdap1, Ppia, Sod1, Prdx5, Atp6v1g1, and Myg1. Among them, the expression of Echdc1, Osbpl8, and Myg1 exhibited consistent down-regulation (Figs. 6A and B). These overlapping DEPs were enriched in acetylation and antioxidant pathways (Fig. 6C).
A The Venn diagram of overlapping DEPs between mouse and HT22 cells. B The log2FC value of overlapping DEPs. C The PPI network of overlapping DEPs. D The scatter plot based on the z-score of overlapping pathways from GO analysis. E The scatter plot based on the z-score of overlapping pathways from KEGG analysis. F. The PPI network of DEPs from four overlapping pathways from KEGG analysis. The positive and negative z-score means the pathway is activated and inhibited, respectively.
Consequently, we calculated z-scores to assess the activation or inhibition of shared and significant pathways in both proteomics results. Based on GO analysis, 16 overlapping pathways were identified, of which 7 pathways including chaperone-mediated protein folding requiring cofactor, chaperone-mediated protein folding, ‘de novo’ posttranslational protein folding, ‘de novo’ protein folding, unfolded protein binding, hydro-lyase activity, and carbon-oxygen lyase activity were inhibited in the context of deficiency of DDX3X (Fig. 6D). Moreover, four overlapping pathways were found in KEGG analysis, i.e., ubiquitin-mediated proteolysis, EGFR tyrosine kinase inhibitor resistance, mTOR signaling, and ErbB signaling pathway. Among these, the ErbB signaling pathway and the ubiquitin-mediated proteolysis pathway were both inhibited compared to the control group (Fig. 6E). The PPI network analysis showed that DEPs involved mTOR signaling pathway, EGFR tyrosine kinase inhibitor resistance, and ErbB signaling pathway exhibited stronger interaction. Additionally, DEPs from the ubiquitin-mediated proteolysis pathway consistently showed significant inhibition in both mouse and HT22 cells (Fig. 6L).
The decreased expression of proteins related to synaptic plasticity
The top 20 hub proteins identified by CytoHubba and robust rank aggregation included MAPK3, MAPK1, MAP2K1, PSD95, PRKCA, and PRKACA, which were belonged to LTP and glutamatergic synapse pathways (Fig. 7A). Seven key proteins, namely MAPK1, PKCA, PSD95, PRRT1, GRIA2, GRIN1, and CAMKIID, were selected for Western blot analysis validation. Among these, MAPK1, PKCA, GRIA2, GRIN1, and CAMKIID were involved in the LTP pathway, while PSD95 and PRRT1 were associated with the postsynapse according to SynGO analysis. Their expression levels were down-regulated compared to the control group, consistent with the proteomics data (Figs. 7B and C). Furthermore, immunofluorescence staining was employed to examine the expression of MAPK1 and PSD95. Their fluorescence intensity in the sh-DDX3X group was significantly lower than that in the control group, which corroborated the proteomics findings (Figs. 7D and E).
A The rank of hub proteins identified by robust rank aggregation. B Representative images of Western blot. C Quantification of Western blot bands. D The representative confocal images of MAPK1 stanning and quantification of fluorescence intensity. Scale bar: 10 μm. E The representative confocal images of PSD95 stanning and quantification of fluorescence intensity. Scale bar: 10 μm. F The representative images of dendrite spine. Scale bar: 10 μm. G The quantitative analysis of the total spine density. H-J The density of the stubby-, mushroom-, and thin-shaped spines. K Representative transmission electron microscopy image of the synapse ultrastructure. Scale bar: 200 nm. L-O The quantitative analysis of PSD thickness, PSD length, PSD area, and cleft width. P Representative samples of mEPSC from control and sh-DDX3X group. Q-R Quantitative analyses of the mEPSC frequency and amplitude (n = 10 from 3 mice per group) from control and sh-DDX3X group. *p < 0.05, ** p < 0.01, *** p < 0.001, two-tailed t test.
The alteration of synapse structure and decreased excitatory induced by the deficiency of DDX3X
Confocal microscopy revealed a significant decrease in total spine density in the sh-DDX3X group compared to controls (Fig. 7F-G), with reductions specifically observed in stubby and mushroom-type spines (Fig. 7H-I), but not in thin spines (Fig. 7J). This suggests that DDX3X deficiency selectively impacts mature spines. Transmission electron microscopy showed that DDX3X deficiency in excitatory synapses reduced PSD thickness and area (Fig. 7K-L), while PSD length and synaptic cleft width were unaffected (Fig. 7M-O), indicating a modest impact on synaptic structure. Moreover, mEPSC recordings in the mPFC demonstrated a significant reduction in frequency but not amplitude in the sh-DDX3X group compared to controls (Fig. 7P-R). These findings collectively suggest that DDX3X deficiency impairs synaptic structure and function, leading to synaptic dysplasia.
Discussion
The DDX3X gene encodes a conservatively expressed ATP-dependent DEAD-box RNA helicase that play an important role in mammalian development [17, 47]. The mutation of DDX3X contributes to a significant portion of ASD patients associated with ID, global developmental delay, and epilepsy [3]. At present, there are rare reports about DDX3X deficiency in animals and cultured neuronal cells in vitro. In this study, we selected an effective targeted sequence to knockdown Ddx3x expression. In HT22 cells, DDX3X deficiency significantly inhibited cell growth. In mice, Ddx3x knockdown did not affect open field behavior but decreased social preference and social novelty preference in the three-chamber test. Additionally, the sh-DDX3X group exhibited impaired spatial recognition memory in the Y-maze test and increased anxiety and repetitive behaviors in the marble burying test. These results indicate that knockdown of the Ddx3x gene in the mPFC of male mice induces social deficits and repetitive behaviors. These phenotypes are similar to the reduced social interaction observed in Ddx3x insufficient mice [29] and the decreased social cohesion reported in ddx3x-deficient zebrafish [16] in previous studies, suggesting that DDX3X deficiency may contribute to ASD-like behaviors and moderate social deficits in mice.
The proteomic analysis of sh-Ddx3x HT22 cells revealed that DEPs between the sh-DDX3X group and the control group were predominantly linked to pathways associated with protein folding, viral genome replication, tRNA metabolism, and DNA binding. These findings are consistent with previous studies on DDX3X-related RNA transcriptional dysregulation [21, 48, 49]. Proteomic analysis of the mPFC in Ddx3x knockdown mice revealed that DEPs were primarily enriched in pathways associated with neurological diseases, including valine, leucine, and isoleucine degradation, long-term potentiation, Parkinson’s disease, GABAergic synapses, and glutamatergic synapses.
Valine, leucine, and isoleucine are branched-chain amino acids (BCAAs) that are essential for protein synthesis and various physiological processes. They serve as key metabolic intermediates in mitochondrial metabolism, influencing glucose and energy metabolism, fatty acid metabolism, and acting as amino acid donors for glutamate synthesis in the brain [50]. BCAA degradation involves a series of enzymatic reactions that ultimately produce ATP. The initial step of BCAA catabolism is catalyzed by two isoenzymes, BCAT1 and BCAT2, with BCAT2 deficiency linked to developmental delays and ASD [51]. In this study, BCAT1 was down-regulated in the Ddx3x knockdown group. Elevated levels of valine, leucine, and isoleucine have been observed in the brains of individuals with ASD [52] and in the hippocampus of valproate (VPA) and thalidomide (THAL)-induced ASD mouse models [53], potentially due to blocked metabolic pathways leading to BCAA accumulation. Our results indicate that reduced DDX3X levels in the mPFC can alter the expression of proteins involved in valine, leucine, and isoleucine degradation pathways, causing BCAA metabolic abnormalities and contributing to social deficits and memory decline in mice.
In the LTP pathway, intracellular kinases, including MAPK1, MAP2K1, MAPK3, CAMKIID, CAMKIIB, PKCA, and PRKACA, were significantly reduced, while gated channel proteins and ion receptors, such as CACNA1A, GRIA2, and GRIN1, were down-regulated. Numerous studies have suggested that MAPK signaling is broadly altered in ASD [54,55,56,57,58]. Dysfunction of the MAPK pathway may contribute to synaptic pathology in ASD-like phenotypes [59, 60]. In addition, several physiologically relevant MAPK substrates have been identified at synaptic sites, including key synaptic scaffold proteins (PSD-95 and PSD-93), Kv4.2 K⁺ channels, and metabotropic glutamate receptors [61]. Activation of the Ras-ERK pathway by NMDA induces a series of transmissions that influence postsynaptic scaffolding proteins, such as PSD-95 [62]. In this study, the down-regulation of MAPK1 and PSD95 in the sh-DDX3X group suggests an underlying relationship between kinase activity and scaffold protein function.
The CaMKII family is a central molecule involved in organizing crucial events for synaptic plasticity, including receptor trafficking, localization and activity, and actin cytoskeletal dynamics [63]. While CAMKIIB has been implicated in neurodevelopmental disorders, CAMKIID has rarely been reported in this context [64, 65]. Maintaining the activity of these protein kinases is essential for synaptic excitation and dendritic spine remodeling, including increased spine volume, stability, and clustering [55, 63, 66,67,68,69].
Previous studies have shown that DDX3X deficiency alters the balance of glutamate neurons in the developing mouse cortex [34]. The interplay among glutamate receptors, CaMKII, postsynaptic density, and MAPK is critical for regulating learning and memory, as well as synaptic plasticity [59]. Additionally, polymorphisms in CACNA1A have been associated with ASD in Chinese populations [60]. Here, the reduced expression of MAPK1, CAMKIID, CAMKIIB, and PKCA in the sh-DDX3X group, which are involved in LTP, was validated by immunoblotting or immunofluorescence. Collectively, these results demonstrate that DDX3X deficiency is associated with a widespread down-regulation of proteins involved in synaptic plasticity. This finding further highlights the key role of the CaMKII family, or kinase-like proteins, in neuroregulation.
Eight DEPs in the GABAergic signaling pathway were significantly down-regulated, namely GABARAPL2, SLC6A11, PRKCA, GNG4, PRKACA, CACNA1A, GAD2, and ABAT. Among them, GAD2 and ABAT are associated with GABA production and metabolism. GAD2 is a regulatory target for cellular activity in various brain functions and GABA-related neurological diseases, catalyzing the conversion of glutamate to GABA. ABAT is a key enzyme responsible for GABA catabolism [70]. This suggests that Ddx3x knockdown may lead to GABA concentration dysregulation and metabolic disturbances in cortical neurons. Changes in GABAergic neurons in the cerebral cortex and striatum are common in autism and schizophrenia [71]. However, in the present study, GABA-related receptors, such as GABRA1 and GABRA3, showed no significant changes. This is consistent with a previous study indicating that GABAA receptors are normal in number and function in individuals with ASD and three types of autism model mice. The autism phenotype may result from other disruptions in the GABAergic pathway, such as reduced GABA release [72]. Therefore, the reduced social behavior due to down-regulated Ddx3x gene expression is likely more related to impaired GABA release than to GABAergic receptors [73].
SynGO analysis revealed that DEPs were enriched in PSD, integral of postsynaptic membrane, postsynaptic ribosome, and postsynaptic actin cytoskeleton. In the context of DDX3X deficiency, the protein expression of PSD95, PRRT1, and SIPA1L1 was significantly decreased. PSD95 is crucial for the formation and function of dendritic spines. It can anchor receptors on the postsynaptic membrane, thereby regulating the quantity, transport, and localization of NMDAR and AMPAR in the postsynaptic membrane [74,75,76,77,78]. PSD95 mutations have been implicated in ASD [79, 80]. PRRT1 regulates AMPAR levels and membrane trafficking [81], while SIPA1L1 is essential for spine growth, synaptic contraction, and the PSD-95/NMDAR complex [82]. The reduced levels of NMDAR and AMPAR in the postsynaptic membrane due to Ddx3x knockdown may decrease neuronal excitability and impair synaptic plasticity [83, 84]. This aberrant synaptic function could disrupt synaptic homeostasis and increase ASD risk.
Indeed, many ASD-related genes encode synaptic proteins or are involved in chromatin remodeling, transcription, protein turnover, and cytoskeleton dynamics. Changes in these proteins can alter synaptic strength and structure, reduce plasticity, and impact neuronal connectivity in the brain [85], and inadequate neuronal connectivity and synaptic development in specific brain regions are considered significant pathogenic factors in ASD [86]. Our results suggest that DDX3X deficiency impairs neurodevelopment and synaptic plasticity by altering the protein profile. Consistently, a previous study showed that silencing Ddx3x expression reduced dendritic spine density, decreased mature mushroom-like spines, and increased immature filamentous and irregular spines [17]. These changes may trigger ASD by disrupting synapse formation and neuronal connectivity. Previous studies have shown that the pathological characteristics of ASD are not caused by a single change in synaptic excitation or a single change in synaptic inhibition, but rather by the combined effect of an imbalance between excitation and inhibition (E/I) resulting from alterations in glutamatergic or GABAergic synapses [9, 87,88,89]. Disruption of the Ddx3x gene, which results in a decrease in excitatory synapses and potential impairment of GABA release, may lead to an E/I imbalance and thereby contribute to the development of ASD.
Over all, proteomic analysis of the mPFC revealed that Ddx3x knockdown significantly impacts postsynaptic signaling pathways. To verify functional synaptic abnormalities, we conducted electrophysiological recordings and confocal microscopy, observing reduced mEPSC frequency, decreased mature dendritic spine density, and smaller PSD size. These findings align with previous studies showing that Ddx3x mutations disrupt LTP-related kinase activity [13], Ddx3x regulates Rac1-mediated axonal growth and synaptogenesis [14], and Ddx3x haploinsufficient mice exhibit reduced cortical PSD95 and impaired dendritic spine maturation [15]. Collectively, these results confirm that Ddx3x deficiency inhibits excitatory synaptic transmission by disrupting postsynaptic structure and function. Importantly, the reduced mEPSC frequency correlates with decreased spine density and may reflect impaired neuronal activity, which is crucial for learning and memory [86].
Integrated analysis of cellular and mouse brain proteomics suggests that ubiquitin-related pathways play a crucial role in ASD pathogenesis. The ubiquitin–proteasome system (UPS) is essential for regulating protein degradation, synapse composition, assembly, and elimination [76], as well as neural development processes such as axonal initiation, dendritic maturation, and synaptic pruning, which are often impaired in ASD [77]. As a regulatory hub intersecting with pathways like mTOR [78], UPS dysfunction can broadly disrupt neural development. Supporting this, brain-specific Uba6 knockout mice exhibit ASD-like behaviors, including increased anxiety, reduced social interaction, and impaired communication [79], while our model mice showed down-regulated cortical UBA3, UBA5, and UBA6 (Supplementary Table 4).
De novo protein synthesis is critical for synaptic plasticity and memory [80], and the UPS mediates most eukaryotic protein turnover [80, 81]. Protein chaperones prevent misfolding by directing proteins to the UPS or autophagy [82]. Abnormalities in postsynaptic ribosomal proteins, such as RPS27A (which mediates eEF1A ubiquitination for translation termination [83]), may disrupt this balance. Consistent with this, sh-DDX3X mice had reduced cortical RPS27A (Supplementary Table 4), and existing studies confirm that DDX3X regulates cortical mRNA translation [13, 34, 84, 85].
These results indicate that Ddx3x deficiency coordinately suppresses the ubiquitin-mediated proteolytic pathway (via UBA3/5/6 downregulation) and de novo protein folding pathway (via RPS27A downregulation). This aligns with previous findings of UPS protein abnormalities in Ddx3x-knockdown mice [35] and the disruption of embryonic protein folding pathways by germline Ddx3x inactivation [14]. Collectively, these findings suggest that inhibition of the UPS and protein folding pathways may represent a conserved mechanism of Ddx3x deficiency, contributing to ASD pathogenesis by disrupting synaptic protein homeostasis.
In summary, this study established a male mouse model with mPFC neuron-specific Ddx3x knockdown, suggesting that Ddx3x deficiency in the mPFC is associated with abnormal synaptic plasticity and core ASD phenotypes. The findings revealed that Ddx3x knockdown can regulate the structure and function of the PSD by affecting the expression of key proteins in postsynaptic signaling pathways; meanwhile, it may exacerbate abnormalities in synaptic protein quality control by inhibiting the UPS and protein folding pathways, a conserved mechanism. These results provide new insights into understanding the potential role of Ddx3x in ASD and the pathological significance of synaptic protein homeostasis imbalance.
However, this study has limitations: the absence of rescue experiments makes it difficult to fully establish the causal relationship between DDX3X deficiency and ASD-like phenotypes; Ddx3x’s role in non-neuronal cells (e.g., astrocytes and microglia) is unexplored; and other ASD-related regions (e.g., the hippocampus and amygdala) lack systematic study. Future work will verify causality via conditional knockout or re-expression experiments, analyze Ddx3x across cell types, and extend to female mice and key regions to deepen understanding of DDX3X’s mechanisms.
Conclusion
In this study, an HT22 cell model with Ddx3x gene knockdown and a mouse model with Ddx3x gene knockdown in the mPFC were successfully established. The results demonstrated that Ddx3x gene knockdown led to retarded growth of HT22 cells and induced autism-like behaviors in mice. Proteomic, morphological, and electrophysiological analyses revealed: DEPs enriched in synaptic plasticity, PSD, and UPS pathways; reduced dendritic spine density/PSD size; and decreased mEPSC frequency. These confirm Ddx3x deficiency impairs postsynaptic structure/function, inhibits excitatory synaptic transmission, and disrupts synaptic protein homeostasis (via suppressed UPS and protein folding)—key mechanisms linking Ddx3x mutation to ASD. Future studies will verify causality via knockout/re-expression, explore Ddx3x in non-neuronal cells and female mice, and clarify how DDX3X specifically regulates the expression and function of synaptic plasticity-related proteins as well as their impacts on neuronal communication and network function—deepening insights into DDX3X-related ASD pathogenesis.
Data availability
All raw data and corresponding parameter have been deposited as online resource to the Figshare database with the name: “The Knockdown of Ddx3x in mPFC Causes Autistic-like Phenotype in Mice with Altered Synaptic Plasticity”. (10.6084/m9.figshare.23684220).
References
Lawrence KE, Hernandez LM, Fuster E, Padgaonkar NT, Patterson G, Jung J, et al. Impact of autism genetic risk on brain connectivity: a mechanism for the female protective effect. Brain. 2022;145:378–87.
Ellingford RA, Panasiuk MJ, de Meritens ER, Shaunak R, Naybour L, Browne L, et al. Cell-type-specific synaptic imbalance and disrupted homeostatic plasticity in cortical circuits of ASD-associated Chd8 haploinsufficient mice. Mol Psychiatry. 2021;26:3614–24.
Tang L, Levy T, Guillory S, Halpern D, Zweifach J, Giserman-Kiss I, et al. Prospective and detailed behavioral phenotyping in DDX3X syndrome. Mol Autism. 2021;12:36.
Kara T, Akaltun İ, Cakmakoglu B, Kaya İ, Zoroğlu S. An Investigation of SDF1/CXCR4 Gene Polymorphisms in Autism spectrum disorder: a family-based study. Psychiatry Investig. 2018;15:300–5.
Sasson NJ, Lam KS, Parlier M, Daniels JL, Piven J. Autism and the broad autism phenotype: familial patterns and intergenerational transmission. J Neurodev Disord. 2013;5:11.
Havdahl A, Niarchou M, Starnawska A, Uddin M, van der Merwe C, Warrier V. Genetic contributions to autism spectrum disorder. Psychological Med. 2021;51:2260–73.
Rosti RO, Sadek AA, Vaux KK, Gleeson JG. The genetic landscape of autism spectrum disorders. Dev Med Child Neurol. 2014;56:12–8.
Gazzellone MJ, Zhou X, Lionel AC, Uddin M, Thiruvahindrapuram B, Liang S, et al. Copy number variation in Han Chinese individuals with autism spectrum disorder. J Neurodev Disord. 2014;6:34.
Zhuang H, Liang Z, Ma G, Qureshi A, Ran X, Feng C, et al. Autism spectrum disorder: pathogenesis, biomarker, and intervention therapy. MedComm. 2024;5:e497.
Busch RM, Srivastava S, Hogue O, Frazier TW, Klaas P, Hardan A, et al. Neurobehavioral phenotype of autism spectrum disorder associated with germline heterozygous mutations in PTEN. Transl Psychiatry. 2019;9:253.
Guo H, Wang T, Wu H, Long M, Coe BP, Li H, et al. Inherited and multiple de novo mutations in autism/developmental delay risk genes suggest a multifactorial model. Mol Autism. 2018;9:64.
Yao TT, Chen L, Du Y, Jiang ZY, Cheng Y. MicroRNAs as regulators, biomarkers, and therapeutic targets in autism spectrum disorder. Mol Neurobiol. 2025;62:5039–56.
Chen L, Fu Q, Du Y, Jiang ZY, Cheng Y. Transcriptome analysis and epigenetics regulation in the hippocampus and the prefrontal cortex of VPA-Induced rat model. Mol Neurobiol. 2024;61:167–74.
Tang X, Feng C, Zhao Y, Zhang H, Gao Y, Cao X, et al. A study of genetic heterogeneity in autism spectrum disorders based on plasma proteomic and metabolomic analysis: multiomics study of autism heterogeneity. MedComm. 2023;4:e380.
Shen L, Zhang H, Lin J, Gao Y, Chen M, Khan NU, et al. A combined proteomics and metabolomics profiling to investigate the genetic heterogeneity of autistic children. Mol Neurobiol. 2022;59:3529–45.
Duan W, Huang G, Sui Y, Wang K, Yu Y, Chu X, et al. Deficiency of DDX3X results in neurogenesis defects and abnormal behaviors via dysfunction of the Notch signaling. Proc Natl Acad Sci USA. 2024;121:e2404173121.
Chen CY, Chan CH, Chen CM, Tsai YS, Tsai TY, Wu Lee YH, et al. Targeted inactivation of murine Ddx3x: essential roles of Ddx3x in placentation and embryogenesis. Hum Mol Genet. 2016;25:2905–22.
Lennox AL, Hoye ML, Jiang R, Johnson-Kerner BL, Suit LA, Venkataramanan S, et al. Pathogenic DDX3X mutations impair RNA metabolism and neurogenesis during fetal cortical development. Neuron. 2020;106:404–20.e8.
Deciphering Developmental Disorders Study. Large-scale discovery of novel genetic causes of developmental disorders. Nature. 2015;519:223–8.
Snijders Blok L, Madsen E, Juusola J, Gilissen C, Baralle D, Reijnders MR, et al. Mutations in DDX3X Are a Common Cause of Unexplained Intellectual Disability with Gender-Specific Effects on Wnt Signaling. Am J Hum Genet. 2015;97:343–52.
Kellaris G, Khan K, Baig SM, Tsai IC, Zamora FM, Ruggieri P, et al. A hypomorphic inherited pathogenic variant in DDX3X causes male intellectual disability with additional neurodevelopmental and neurodegenerative features. Hum Genomics. 2018;12:11.
Martin HC, Gardner EJ, Samocha KE, Kaplanis J, Akawi N, Sifrim A, et al. The contribution of X-linked coding variation to severe developmental disorders. Nat Commun. 2021;12:627.
Ruzzo EK, Pérez-Cano L, Jung JY, Wang LK, Kashef-Haghighi D, Hartl C, et al. Inherited and de novo genetic risk for autism impacts shared networks. Cell. 2019;178:850–66.e26.
Levy T, Siper PM, Lerman B, Halpern D, Zweifach J, Belani P, et al. DDX3X Syndrome: summary of findings and recommendations for evaluation and care. Pediatr Neurol. 2023;138:87–94.
Iossifov I, O’Roak BJ, Sanders SJ, Ronemus M, Krumm N, Levy D, et al. The contribution of de novo coding mutations to autism spectrum disorder. Nature. 2014;515:216–21.
Chen HH, Yu HI, Tarn WY. DDX3 modulates neurite development via translationally activating an RNA regulon involved in rac1 activation. J Neurosci. 2016;36:9792–804.
Hoye ML, Calviello L, Poff AJ, Ejimogu NE, Newman CR, Montgomery MD, et al. Aberrant cortical development is driven by impaired cell cycle and translational control in a DDX3X syndrome model. Elife. 2022;11:e78203.
Mossa A, Dierdorff L, Lukin J, Garcia-Forn M, Wang W, Mamashli F, et al. Sex-specific perturbations of neuronal development caused by mutations in the autism risk gene DDX3X. Nat Commun. 2025;16:4512.
Boitnott A, Garcia-Forn M, Ung DC, Niblo K, Mendonca D, Park Y, et al. Developmental and behavioral phenotypes in a mouse model of DDX3X syndrome. Biol Psychiatry. 2021;90:742–55.
Kennis MGP, Rots D, Bouman A, Ockeloen CW, Boelen C, Marcelis CLM, et al. DDX3X-related neurodevelopmental disorder in males - presenting a new cohort of 19 males and a literature review. Eur J Hum Genet. 2025;33:980–8.
Lovick TA. GABA in the female brain - oestrous cycle-related changes in GABAergic function in the periaqueductal grey matter. Pharmacol Biochem Behav. 2008;90:43–50.
Murakawa T, Kogure L, Hata K, Hasunuma K, Takenawa S, Sano K, et al. Estrous cycle-dependent modulation of sexual receptivity in female mice by estrogen receptor beta-expressing cells in the dorsal raphe nucleus. J Neurosci. 2024;44:e11374242024.
Guo Z, Tang X, Xiao S, Yan H, Sun S, Yang Z, et al. Systematic review and meta-analysis: multimodal functional and anatomical neural alterations in autism spectrum disorder. Mol Autism. 2024;15:16.
Chen L, Xiong XY, Yao TT, Gui LN, Luo F, Du Y, et al. Blood exosome sensing via neuronal insulin-like growth factor-1 regulates autism-related phenotypes. Pharmacol Res. 2023;197:106965.
Padilla-Coreano N, Batra K, Patarino M, Chen Z, Rock RR, Zhang R, et al. Cortical ensembles orchestrate social competition through hypothalamic outputs. Nature. 2022;603:667–71.
Kelly E, Meng F, Fujita H, Morgado F, Kazemi Y, Rice LC, et al. Regulation of autism-relevant behaviors by cerebellar-prefrontal cortical circuits. Nat Neurosci. 2020;23:1102–10.
Kügler S, Kilic E, Bähr M. Human synapsin 1 gene promoter confers highly neuron-specific long-term transgene expression from an adenoviral vector in the adult rat brain depending on the transduced area. Gene Ther. 2003;10:337–47.
Minderer M, Liu W, Sumanovski LT, Kügler S, Helmchen F, Margolis DJ. Chronic imaging of cortical sensory map dynamics using a genetically encoded calcium indicator. J Physiol. 2012;590:99–107.
Bian WJ, Brewer CL, Kauer JA, de Lecea L. Adolescent sleep shapes social novelty preference in mice. Nat Neurosci. 2022;25:912–23.
Mehta A, Shirai Y, Kouyama-Suzuki E, Zhou M, Yoshizawa T, Yanagawa T, et al. IQSEC2 deficiency results in abnormal social behaviors relevant to autism by affecting functions of neural circuits in the medial prefrontal cortex. Cells. 2021;10:2724.
Jung EM, Moffat JJ, Liu J, Dravid SM, Gurumurthy CB, Kim WY. Arid1b haploinsufficiency disrupts cortical interneuron development and mouse behavior. Nat Neurosci. 2017;20:1694–707.
Liu H, Huang X, Li Y, Xi K, Han Y, Mao H, et al. TNF signaling pathway-mediated microglial activation in the PFC underlies acute paradoxical sleep deprivation-induced anxiety-like behaviors in mice. Brain Behav Immun. 2022;100:254–66.
Lin J, Zhang K, Cao X, Zhao Y, Ullah Khan N, Liu X, et al. iTRAQ-Based proteomics analysis of rat cerebral cortex exposed to valproic acid before delivery. ACS Chem Neurosci. 2022;13:648–63.
Wiśniewski JR, Zougman A, Nagaraj N, Mann M. Universal sample preparation method for proteome analysis. Nat Methods. 2009;6:359–62.
Zhang X, Yang H, Wang Y, Zhang J, Zhang H, Cao X, et al. Proteomic study on the mechanism of arsenic neurotoxicity in the rat cerebral cortex and the protective mechanism of dictyophora polysaccharides against arsenic neurotoxicity. ACS Chem Neurosci. 2023;14:2302–19.
Gillet LC, Navarro P, Tate S, Röst H, Selevsek N, Reiter L, et al. Targeted data extraction of the MS/MS spectra generated by data-independent acquisition: a new concept for consistent and accurate proteome analysis. Mol Cell Proteom. 2012;11:O111.016717.
Sharma D, Jankowsky E. The Ded1/DDX3 subfamily of DEAD-box RNA helicases. Crit Rev Biochem Mol Biol. 2014;49:343–60.
Khadivjam B, Bonneil É, Thibault P, Lippé R. RNA helicase DDX3X modulates herpes simplex virus 1 nuclear egress. Commun Biol. 2023;6:134.
Valentin-Vega YA, Wang YD, Parker M, Patmore DM, Kanagaraj A, Moore J, et al. Cancer-associated DDX3X mutations drive stress granule assembly and impair global translation. Sci Rep. 2016;6:25996.
Shimomura Y, Kitaura Y. Physiological and pathological roles of branched-chain amino acids in the regulation of protein and energy metabolism and neurological functions. Pharmacol Res. 2018;133:215–7.
Knerr I, Colombo R, Urquhart J, Morais A, Merinero B, Oyarzabal A, et al. Expanding the genetic and phenotypic spectrum of branched-chain amino acid transferase 2 deficiency. J Inherit Metab Dis. 2019;42:809–17.
Graham SF, Turkoglu O, Yilmaz A, Ustun I, Ugur Z, Bjorndhal T, et al. Targeted metabolomics highlights perturbed metabolism in the brain of autism spectrum disorder sufferers. Metabolomics. 2020;16:59.
Toczylowska B, Zieminska E, Senator P, Lazarewicz JW. Hippocampal metabolite profiles in two rat models of autism: NMR-Based metabolomics studies. Mol Neurobiol. 2020;57:3089–105.
Wen Y, Alshikho MJ, Herbert MR. Pathway network analyses for autism reveal multisystem involvement, major overlaps with other diseases and convergence upon MAPK and calcium signaling. PLoS One. 2016;11:e0153329.
Thomas GM, Huganir RL. MAPK cascade signalling and synaptic plasticity. Nat Rev Neurosci. 2004;5:173–83.
Xing L, Larsen RS, Bjorklund GR, Li X, Wu Y, Philpot BD, et al. Layer specific and general requirements for ERK/MAPK signaling in the developing neocortex. Elife. 2016;5:e11123.
Faridar A, Jones-Davis D, Rider E, Li J, Gobius I, Morcom L, et al. Mapk/Erk activation in an animal model of social deficits shows a possible link to autism. Mol Autism. 2014;5:57.
Satoh Y, Endo S, Nakata T, Kobayashi Y, Yamada K, Ikeda T, et al. ERK2 contributes to the control of social behaviors in mice. J Neurosci. 2011;31:11953–67.
Tiwari A, Rahi S, Mehan S. Elucidation of abnormal extracellular regulated kinase (ERK) signaling and associations with syndromic and non-syndromic autism. Curr Drug Targets. 2021;22:1071–86.
Engel SR, Creson TK, Hao Y, Shen Y, Maeng S, Nekrasova T, et al. The extracellular signal-regulated kinase pathway contributes to the control of behavioral excitement. Mol Psychiatry. 2009;14:448–61.
Mao LM, Wang JQ. Synaptically localized mitogen-activated protein kinases: local substrates and regulation. Mol Neurobiol. 2016;53:6309–15.
Wu M, Funahashi Y, Takano T, Hossen E, Ahammad RU, Tsuboi D, et al. Rho-Rho-Kinase regulates Ras-ERK signaling through SynGAP1 for dendritic spine morphology. Neurochem Res. 2022;47:2757–72.
Yasuda R, Hayashi Y, Hell JW. CaMKII: a central molecular organizer of synaptic plasticity, learning and memory. Nat Rev Neurosci. 2022;23:666–82.
Küry S, van Woerden GM, Besnard T, Proietti Onori M, Latypova X, Towne MC, et al. De novo mutations in protein kinase genes CAMK2A and CAMK2B cause intellectual disability. Am J Hum Genet. 2017;101:768–88.
Nicole O, Bell DM, Leste-Lasserre T, Doat H, Guillemot F, Pacary E. A novel role for CAMKIIβ in the regulation of cortical neuron migration: implications for neurodevelopmental disorders. Mol Psychiatry. 2018;23:2209–26.
Colgan LA, Hu M, Misler JA, Parra-Bueno P, Moran CM, Leitges M, et al. PKCα integrates spatiotemporally distinct Ca(2+) and autocrine BDNF signaling to facilitate synaptic plasticity. Nat Neurosci. 2018;21:1027–37.
Stampanoni Bassi M, Iezzi E, Gilio L, Centonze D, Buttari F. Synaptic plasticity shapes brain connectivity: implications for network topology. Int J Mol Sci. 2019;20:6193.
Engert F, Bonhoeffer T. Dendritic spine changes associated with hippocampal long-term synaptic plasticity. Nature. 1999;399:66–70.
Gatta V, Drago D, Fincati K, Valenti MT, Dalle Carbonare L, Sensi SL, et al. Microarray analysis on human neuroblastoma cells exposed to aluminum, β(1-42)-amyloid or the β(1-42)-amyloid aluminum complex. PLoS One. 2011;6:e15965.
Besse A, Wu P, Bruni F, Donti T, Graham BH, Craigen WJ, et al. The GABA transaminase, ABAT, is essential for mitochondrial nucleoside metabolism. Cell Metab. 2015;21:417–27.
Canitano R, Pallagrosi M. Autism spectrum disorders and schizophrenia spectrum disorders: excitation/inhibition imbalance and developmental trajectories. Front Psychiatry. 2017;8:69.
Zhao H, Mao X, Zhu C, Zou X, Peng F, Yang W, et al. GABAergic system dysfunction in autism spectrum disorders. Front Cell Dev Biol. 2021;9:781327.
Horder J, Andersson M, Mendez MA, Singh N, Tangen Ä, Lundberg J, et al. GABA(A) receptor availability is not altered in adults with autism spectrum disorder or in mouse models. Sci Transl Med. 2018;10:eaam8434.
Rodríguez-Palmero A, Boerrigter MM, Gómez-Andrés D, Aldinger KA, Marcos-Alcalde Í, Popp B, et al. DLG4-related synaptopathy: a new rare brain disorder. Genet Med. 2021;23:888–99.
Brown JC, Higgins ES, George MS. Synaptic plasticity 101: the story of the AMPA receptor for the brain stimulation practitioner. Neuromodulation. 2022;25:1289–98.
Yao M, Meng M, Yang X, Wang S, Zhang H, Zhang F, et al. POSH regulates assembly of the NMDAR/PSD-95/Shank complex and synaptic function. Cell Rep. 2022;39:110642.
Zhao JP, Murata Y, Constantine-Paton M. Eye opening and PSD95 are required for long-term potentiation in developing superior colliculus. Proc Natl Acad Sci USA. 2013;110:707–12.
Compans B, Camus C, Kallergi E, Sposini S, Martineau M, Butler C, et al. NMDAR-dependent long-term depression is associated with increased short term plasticity through autophagy mediated loss of PSD-95. Nat Commun. 2021;12:2849.
Li J, Shi M, Ma Z, Zhao S, Euskirchen G, Ziskin J, et al. Integrated systems analysis reveals a molecular network underlying autism spectrum disorders. Mol Syst Biol. 2014;10:774.
Xing J, Kimura H, Wang C, Ishizuka K, Kushima I, Arioka Y, et al. Resequencing and association analysis of six PSD-95-Related genes as possible susceptibility genes for schizophrenia and autism spectrum disorders. Sci Rep. 2016;6:27491.
Troyano-Rodriguez E, Mann S, Ullah R, Ahmad M. PRRT1 regulates basal and plasticity-induced AMPA receptor trafficking. Mol Cell Neurosci. 2019;98:155–63.
Pak DT, Yang S, Rudolph-Correia S, Kim E, Sheng M. Regulation of dendritic spine morphology by SPAR, a PSD-95-associated RapGAP. Neuron. 2001;31:289–303.
Iasevoli F, Tomasetti C, de Bartolomeis A. Scaffolding proteins of the post-synaptic density contribute to synaptic plasticity by regulating receptor localization and distribution: relevance for neuropsychiatric diseases. Neurochem Res. 2013;38:1–22.
Chen Y, Wang Y, Ertürk A, Kallop D, Jiang Z, Weimer RM, et al. Activity-induced Nr4a1 regulates spine density and distribution pattern of excitatory synapses in pyramidal neurons. Neuron. 2014;83:431–43.
Bourgeron T. From the genetic architecture to synaptic plasticity in autism spectrum disorder. Nat Rev Neurosci. 2015;16:551–63.
Alaerts K, Swinnen SP, Wenderoth N. Neural processing of biological motion in autism: An investigation of brain activity and effective connectivity. Sci Rep. 2017;7:5612.
Rubinstein M, Westenbroek RE, Yu FH, Jones CJ, Scheuer T, Catterall WA. Genetic background modulates impaired excitability of inhibitory neurons in a mouse model of Dravet syndrome. Neurobiol Dis. 2015;73:106–17.
Rubenstein JL, Merzenich MM. Model of autism: increased ratio of excitation/inhibition in key neural systems. Genes Brain Behav. 2003;2:255–67.
Nelson SB, Valakh V. Excitatory/Inhibitory balance and circuit homeostasis in autism spectrum disorders. Neuron. 2015;87:684–98.
Acknowledgements
We thank the Center for Instrumental Analysis, Shenzhen University. We also grateful to the BioRender website and PowerPoint for providing the image materials.
Funding
This study was supported by the National Natural Science Foundation of China (Grant No. 31870825), the Shenzhen Bureau of Science, Technology and Information (No. JCYJ20170412110026229), the Shenzhen-Hong Kong Institute of Brain Science-Shenzhen Fundamental Research Institutions (2023SHIBS0003), and the Graduate Student Innovation Cultivation Foundation of Shenzhen University.
Author information
Authors and Affiliations
Contributions
HZ and XC wrote the original draft and contributed to visualization, methodology, and investigation. XT contributed to writing—review & editing, methodology, and visualization. ZL contributed to writing—review & editing and visualization. HW contributed to writing—review & editing and methodology. QL contributed to writing—review & editing and investigation. XY, MW, and YH contributed to writing—review & editing. XL and XS contributed to writing—review & editing and methodology. DL and YW contributed to methodology. LS contributed to writing—original draft, writing—review & editing, methodology, investigation, supervision, and funding acquisition. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
All authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Ethics approval and consent to participate
All methods were performed in accordance with the relevant guidelines and regulations. The study was approved by the Animal Ethical and Welfare Committee of Shenzhen University (Permit Number: AEWC-20140615-002). This study did not involve human participants or identifiable human images, so informed consent and consent for publication are not applicable.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Consent for publication
All authors have agreed with the content of the research article and agreed to publish the outcomes.
Supplementary information
41398_2026_3945_MOESM6_ESM.xlsx (download XLSX )
Supplementary Table4. The results of differential analysis between sh-DDX3X and control group in mouse prefrontal cortex.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhuang, H., Cao, X., Tang, X. et al. Knockdown of Ddx3x in mPFC induces autistic-like phenotype in mice via altered synaptic plasticity. Transl Psychiatry 16, 216 (2026). https://doi.org/10.1038/s41398-026-03945-3
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41398-026-03945-3