Abstract
Pathological cardiac hypertrophy is associated with intricate changes that can eventually lead to heart failure. Galangin is a natural flavonoid derived from the rhizome of the galangal plant that has shown protective effects against a variety of cardiomyopathies. Galangin undergoes methylation at the third hydroxyl position to form Galangin 3-methyl ether (G3-ME), thus increasing its lipophilicity, thereby facilitating its cellular entry and regulatory effects. In this study, we investigated the effects of G3-ME on hypertrophic cardiomyopathy (HCM) and the underlying mechanisms. Two established transgenic mouse models of HCM, i.e., Myh6R404Q and Tnnt2R109Q mice, were used for in vivo studies. For in vitro experiments, H9c2, a rat cardiomyocyte cell line, along with human embryonic stem cell-derived cardiomyocytes (hESC-CMs), were exposed to angiotensin II (Ang II, 1 μM). We showed that G3-ME (2.5–20 μM) dose-dependently inhibited the expression of pathological cardiac hypertrophy marker genes Nppa and Nppb in Ang II-treated H9c2 cells. G3-ME (5 μM) effectively mitigated the Ang II-induced expansion of H9c2 cells and hESC-CMs. In the two HCM mouse models, administration of G3-ME (20 or 50 mg·kg−1·d−1, i.g.) for 4 weeks significantly improved the cardiac hypertrophy phenotype. By conducting RNA sequencing combined with network pharmacology, we identified histone deacetylase 2 (HDAC2) as the direct target of G3-ME in H9c2 cells. We demonstrated that G3-ME inhibited both the phosphorylation of HDAC2 and the PI3K-AKT signaling pathway during the cardiac hypertrophy process in vivo and in vitro. Knockdown of HDAC2 resulted in inhibition of cardiac hypertrophy, whereas its overexpression facilitated cardiac hypertrophy through the PI3K-AKT signaling pathway. In conclusion, G3-ME attenuates pathological cardiac hypertrophy in vivo and in vitro by targeting HDAC2, leading to the inactivation of the PI3K-AKT signaling pathway. G3-ME may serve as a promising option for addressing hypertrophic cardiomyopathy.
Similar content being viewed by others
Introduction
The heart typically possesses a robust capacity for compensation [1]. Cardiac hypertrophy that is physiologically compensated represents an adaptive mechanism initiated by specific physiological triggers and is generally not associated with cardiac dysfunction [2, 3]. In contrast, pathological cardiac hypertrophy involves structural alterations in the heart resulting from certain diseases or pathological conditions, and is generally linked to arrhythmia and even heart failure [2, 4, 5]. Mechanistically, pathological cardiac hypertrophy can be secondary to various hypertrophic stimulus factors, and is also a primary clinical manifestation in patients with hereditary hypertrophic cardiomyopathy (HCM) [6, 7]. Currently, there are no clinically effective methods available to cure cardiac hypertrophy, particularly in HCM. As such, it is crucial to explore viable targets or agents that can inhibit cardiac hypertrophy and HCM for therapeutic purposes.
Pathological cardiac hypertrophy development is associated with the dysregulation of various signaling pathways, among which the phosphoinositide 3-kinase (PI3K) and protein kinase B (AKT) pathways are particularly important [8, 9]. PI3K, a heterodimeric lipid kinase found at the plasma membrane, catalyzes the production of phosphatidylinositol-3,4,5-trisphosphate [10]. The activity of AKT, which is a serine/threonine protein kinase, is predominantly regulated by membrane recruitment facilitated by phosphatidylinositol-3,4,5-trisphosphate [11]. Various factors can induce cardiomyocyte hypertrophy, which is typically associated with differing levels of persistent abnormal upregulation of PI3K-AKT signaling [8, 12].
A notable aspect of cardiac hypertrophy is the reactivation of a “fetal gene program” [13]. The specific molecular mechanisms that affect this program are largely unclear; however, the regulation of gene transcription at a relatively high level is at least somewhat directed by a sophisticated array of modifications to core histones, which are proteins that interact with DNA to create chromatin [14]. Changes in histone acetylation and deacetylation, which are performed by histone acetyltransferases and histone deacetylases (HDACs), respectively, result in modifications to the chromatin structure and subsequent alterations in gene expression [15, 16]. The HDAC family consists of a well-conserved group that includes three distinct subfamilies [15, 17]. The importance of the HDAC family in maintaining cardiomyocyte homeostasis is both crucial and intricate [18]. Multiple studies have shown that class I and class II HDACs play contrasting roles in the control of hypertrophic pathways: class I HDACs (e.g., HDAC2) promote certain hypertrophic responses [15, 19,20,21]. Current evidence implies that HDAC2 activates a signaling cascade within the cell that is downstream of PI3K-AKT, ultimately leading to cardiac hypertrophy [15, 22,23,24]. Inhibition of the catalytic activity of class I HDACs effectively hinders hypertrophic responses in both isolated myocytes and mice, suggesting that focusing on class I HDACs, such as HDAC2, may offer a promising approach to mitigate pathological cardiac hypertrophy and even heart failure [25]. However, developing effective methods to inhibit HDAC2 activation and the associated hypertrophic response remains challenging.
Galangin 3-methyl ether (G3-ME), a structural analog of galangin, is extracted from the rhizome of galangal. Galangin, also known as 3,5,7-trihydroxyflavone, is methylated at the third hydroxyl position to form G3-ME. This methylation process reduces the polarity of G3-ME while increasing its lipophilicity, thereby facilitating its cellular entry and regulatory effects. Furthermore, the methylation of the hydroxyl group enhances the stability of G3-ME against oxidative stress and metabolic degradation, contributing to its sustained efficacy, which includes significant anti-inflammatory and antitumor effects. Galangin has been shown to protect against myocardial ischemia-reperfusion injury [26, 27], cardiac inflammation [28, 29], isoproterenol-induced cardiac remodeling [30], and doxorubicin-induced cardiotoxicity [31, 32]. However, the mechanism of action of G3-ME in cardiac diseases, particularly in pathological cardiac hypertrophy, has never been defined. The present study aimed to investigate the roles of G3-ME in HDAC2-mediated PI3K-AKT signaling and to elucidate the underlying mechanisms involved in cardiac hypertrophy.
Materials and methods
HCM mouse models
This research employed two established transgenic mouse models of HCM, resulting from a missense mutation in myosin heavy chain 6 (Myh6) at the 404th amino acid residue (Myh6R404Q) and a missense mutation in cardiac troponin T2 (Tnnt2) at the 109th amino acid residue (Tnnt2R109Q). Myh6R404Q mice are characterized by a heterozygous arginine (R)-to-glutamine (Q) mutation in Myh6 at position 404 [33]. The murine Myh6 gene is orthologous to the human MYH7 gene, which is the gene most commonly associated with HCM. The sequence similarity between the murine MYH6 protein and the human MYH7 protein exceeds 92%. Notably, the 404th amino acid of the mouse MYH6 protein corresponds to the 403rd amino acid of the human MYH7 protein. The R-to-Q mutation at position 403 in humans is recognized as one of the primary causes of HCM [34,35,36]. Sequencing analysis revealed that Myh6R404Q mice exhibited alterations in codons 1210–1212, which changed from CGG to CAG (Fig. 3a).
Tnnt2R109Q mice were characterized by a heterozygous R-to-Q mutation in the mouse Tnnt2 at residue 109. The sequence similarity between murine and human TNNT2 is approximately 90%. Notably, HCM caused by mutations in the TNNT2 gene accounts for approximately 5% of cases [37], with the R-to-Q mutation at position 92 in human TNNT2 recognized as one of the primary causes of HCM [38,39,40]. Notably, human amino acid 92 of TNNT2 corresponds to mouse amino acid 109; thus, we constructed the mouse Tnnt2 gene 325–327 from AGG to CAG (Fig. 5a).
Animal experimental procedure
Eight-week-old male wild-type C57BL/6J mice (WT) and HCM mice were procured from GemPharmatech Co., Ltd (Nanjing, China). These mice were kept in a temperature-regulated environment and maintained at 22–26 °C with humidity levels of 40%–50%, under a 12-h light/dark cycle. They had unrestricted access to standard chow and tap water. All research adhered to the principles outlined in the Guide for the Care and Use of Laboratory Animals (NIH, 8th edition, 2011) and received approval from the Laboratory Animal Welfare & Ethics Committee at Jiangnan University (approval number: JN. No. 20250228m0800901[045]). To assess the therapeutic effects of G3-ME on cardiac hypertrophy, normal WT and Myh6R404Q mice were randomly assigned to receive vehicle (Veh), a low dose of G3-ME (20 mg·kg−1·d−1), or a high dose of G3-ME (50 mg·kg−1·d−1) via intragastric administration for four consecutive weeks (Fig. 3a). Additionally, Tnnt2R109Q mice were selected to further confirm the effects of G3-ME on pathological cardiac hypertrophy. WT or Tnnt2R109Q mice were administered 20 mg·kg−1·d−1 G3-ME intragastrically for four consecutive weeks (Fig. 5a).
Reagents and chemicals
G3-ME (HY-N4167), CHIR-99021 (HY-10182), IWR-1 (HY-12238), F-127 (HY-D1005), and G418 (HY-17561) were purchased from MedChem Express (Monmouth Junction, NJ, USA). Angiotensin II (05-23-0101) and protease XIV (P5147) were purchased from Sigma Chemical Co. (St Louis, USA). Wheat germ agglutinin (WGA, 7024), Hoechst-33342 (C10246), DAPI (D21490), the Pierce™ BCA Protein Assay Kit (23225), and radioimmunoprecipitation assay lysis buffer (89900) were obtained from Thermo Fisher Scientific (Carlsbad, CA, USA). A Cal-520®-AM probe (21130) was obtained from AAT Bioquest (Pleasanton, CA, USA). Antibodies against ANP (WL02666) and BNP (WL02126) were gained from WanleiBio (Shenyang, China). Antibodies against AKT (10176-2-AP), phospho-AKT (Ser473, 66444-1-lg), cardiac troponin T (cTNT, 15513-1-AP), GAPDH (60004-1-lg), and β-tubulin (10094-1-AP) were purchased from Proteintech (Chicago, USA). Antibodies against PI3K p85α (AF6241), phospho-PI3K p85α (Tyr607, AF3241), HDAC2 (AF6470), and phospho-HDAC2 (Ser394, AF3470) were purchased from Affinity Biosciences (Cincinnati, USA). Hematoxylin & eosin (H&E; C0105) was procured from Beyotime Biotechnology (Shanghai, China). A Cell Counting Kit-8 (CCK-8, K1018) was obtained from APExBio Technology LLC (Houston, TX, USA). Masson’s trichrome (G1006) was obtained from ServiceBio (Wuhan, China). The hESC line H7 was obtained from the WiCell Research Institute (Madison, Wisconsin, USA). The primers used were provided by Sangon Biotech (Shanghai, China), and the sequences are provided in Supplementary Table 1.
hESC culture and differentiation
As previously reported, we employed TALEN-mediated genomic engineering to integrate the neomycin selection cassette immediately following the expression of the myosin light chain 2 (MYL2) gene, a gene with ventricular-specific expression, in hESCs [41]. In this way, neomycin can be coexpressed with MYL2 in hESC-differentiated cardiomyocytes. We then used G418 selection to obtain high-purity hESC-CMs (>98%). The differentiation of hESCs into cardiomyocytes was accomplished using a previously outlined method [42]. In summary, once the hESCs reached 80%–90% confluence, we sequentially treated the cells with CHIR-99021 and IWR-1. After 7–9 days of differentiation, beating clusters could be observed. On the 21st day post-differentiation, these beating clusters were digested with collagenase-I and reseeded into a 6-well or 12-well plate. G418 (100 μg/mL) was then used for the selection of hESC-CMs for 8 days. After around 30 days of differentiation and purification, the hESC-CMs were treated with different stimuli, and the corresponding assays were performed (Fig. 2A).
Evaluation of cell viability
The viability of H9c2 cells or hESC-CMs was assessed using a CCK-8 assay as previously described [43]. In summary, the cells were placed in 96-well plates for cultivation. Once the confluence of the cells reached approximately 80%, G3-ME was added to the culture medium for 24 h. Afterward, 10 μL of CCK-8 solution was added to each well and incubated for 2 h at 37 °C within a humidified atmosphere containing 5% CO2. The absorbance was measured at 450 nm using a Multimode Plate Reader (EnSight, PerkinElmer, Waltham, MA, USA).
Echocardiography
Echocardiographic assessments in mice were conducted utilizing a high-resolution in vivo imaging system (Vevo3100LT, Toronto, Canada). Anesthesia was achieved with a flow of 1.5%–2% isoflurane, ensuring that the heart rates were maintained within 450–480 BPM for precise measurements. From the echocardiograms, the ejection fraction (EF), fractional shortening (FS) of the left ventricle (LV), LV mass, and thickness of the anterior and posterior walls of the LV during both systole and diastole (LVAW;s, LVAW;d, LVPW;s, and LVPW;d) were calculated. Additionally, the LV diameter in systole and diastole (LVD;s, LVD;d) and the LV volume in systole and diastole (LVV;s, LVV;d) were also measured. Comprehensive data can be found in Supplementary Table 3 and Table 4.
Immunofluorescence staining
Immunofluorescence staining of cTNT was performed to assess cardiomyocyte size as previously described [44]. Briefly, H9c2 cells or hESC-CMs were seeded onto glass coverslips in 48-well plates and subjected to various treatments. Following a specific incubation time, the cells were subjected to a series of treatments: paraformaldehyde (4%) for 15–20 min, Triton X-100 (0.2%) for 5 min, and finally BSA (1%) for 2 h. Next, the cells were allowed to incubate overnight at 4 °C with primary antibodies targeting cTNT (1:200). The samples were subsequently treated with fluorescence-conjugated secondary antibodies (1:1000) for 1 h at room temperature. DAPI (1:1000) was used for nuclear staining. Immunofluorescence images were obtained with an RVL2-K inverted fluorescence microscope (USA), and the cell area was analyzed using ImageJ software (v1.80; NIH; Maryland, USA).
Histological analysis
The mouse heart was quickly isolated following euthanasia and then successively fixed with 4% paraformaldehyde, dehydrated with ethanol, and embedded in paraffin. Sections (5 μm) were subjected to H&E staining, Masson’s staining, and WGA staining according to the manufacturer’s instructions as previously described [45]. Images of H&E and Masson’s staining were captured with a light microscope (BX-51; Olympus, Tokyo, Japan). The cross-sectional area and relative fibrosis area of the heart were quantified using ImageJ software. Images of WGA staining were obtained with a fluorescence microscope, and the areas of cardiomyocytes were analyzed using ImageJ software.
Calcium (Ca2+) transient
On day 30 of differentiation, the hESC-CMs were dissociated into individual cells and placed onto 20 mm cover glass-bottom dishes. The beating cardiomyocytes were treated with 5 μM Cal-520®-AM and 0.02% F-127 in a solution of 10% FBS Hank’s at 37 °C for a duration of 20 min. Calcium signaling was captured at 37 °C using the line-scan mode with a laser scanning confocal microscope (Zeiss, LSM-880) and was analyzed through a semiautomated software specifically developed for examining local Ca2+ spark release [46].
Quantitative real-time polymerase chain reaction (qRT-PCR)
Total RNA was isolated from H9c2 cells, hESC-derived cardiomyocytes, and LV tissues using TRIzol reagent (Vazyme Biotech Co., Ltd; Nanjing, China). The RNA concentration was subsequently quantified with a NanoDrop spectrophotometer (Thermo Fisher Scientific; MA, USA). For reverse transcription-PCR, 1 μg of RNA was used along with a PrimeScript RT Master Mix kit (Takara; Kyoto, Japan). In the case of qRT-PCR, SYBR Green Master Mix (Takara) was employed on a LightCycler® R96 Instrument (Basel, Switzerland). The sequences of the primers used in this research can be found in Supplementary Table 1.
Western blot analysis
First, total protein was extracted with radioimmunoprecipitation assay lysis buffer, and the protein concentration was determined using a Pierce™ BCA protein assay kit. Total protein (20 μg) was subsequently separated by SDS-PAGE and transferred onto PVDF membranes. These membranes were then incubated with specific primary antibodies, followed by the application of HRP-conjugated secondary antibodies. The blot analysis was conducted with a fully automatic digital gel/chemiluminescence imaging system (Tanon; Shanghai, China). For Western blot analysis, anti-GAPDH and anti-β-tubulin were diluted at a ratio of 1:5000, whereas all other antibodies were diluted to 1:1000. The intensity of the staining of the blot was quantified using ImageJ software.
Enzyme-linked immunosorbent assay (ELISA)
Serum N-terminal pro-brain natriuretic peptide (NT-proBNP) serves as an important biomarker for assessing cardiac load and functional status in patients with pathological myocardial hypertrophy [47]. NT-proBNP levels in the serum of mice were measured using a commercial mouse NT-proBNP ELISA kit (E-EL-M0834) from Elabscience Biotechnology Co., Ltd (Wuhan, China). Whole blood samples from the mice were incubated at 4 °C for 30 min and then centrifuged at 5000 rpm for 20 min to obtain the supernatant for subsequent experiments. The detection of plasma NT-proBNP levels was conducted according to the instructions provided.
Plasmids and transfection
The full-length cDNA of Hdac2 tagged with FLAG was amplified by standard PCR and inserted into the pcDNA 3.1 vector, and the pcDNA 3.1 plasmid was used as a control. The plasmids were transfected into H9c2 cells using Lipofectamine 3000 (Invitrogen, Carlsbad, CA, USA) following the manufacturer’s instructions. The sequences used in the overexpression plasmid are provided in Supplementary Table 2.
HDAC2 and inositol polyphosphate-5-phosphatase f (INPP5F) stable knockdown cell line construction
The pLKO.1-Puro-shHdac2 and pLKO.1-Puro-shInpp5f plasmids were constructed, with the control plasmid being pLKO.1-Puro. HEK-293T cells were cotransfected with the packaging plasmids psPAX2 and pMD2.G, and the pLKO.1-Puro-shRNA plasmids to produce lentiviral particles. The resulting virus was collected and used to infect cells for 24 h. Infected cells were screened using 10 μg/mL puromycin, and the surviving cells were identified as HDAC2 or INPP5F stable knockdown cells. The shRNA sequences are detailed in Supplementary Table 2.
Drug affinity reaction target stability (DARTs)
DARTs experiment was performed in vitro or in vivo. For the in vitro model, H9c2 cells or hESC-CMs were treated with 1% NP-40 lysis buffer, followed by centrifugation at 4 °C (12,000 rpm), after which the supernatants were collected. The cell lysates were then divided into two equal portions, which were incubated under normal control conditions (Veh) or with G3-ME (50 μM) for 2 h at room temperature. The Veh- or G3-ME-treated lysates were subsequently divided into two equal portions, each of which was incubated with PBS or protease XIV (100 μg/mL) for 5 min. An equal volume of loading buffer (5×) was added to all the samples, which were subsequently subjected to a Western blot at 100 °C for 3 min.
To establish an in vivo model, H9c2 cells or hESC-CMs were pretreated with Veh or G3-ME (50 μM) for 24 h of culture at 5% CO2 at 37 °C. Total protein was subsequently extracted from cells using 1% NP-40 lysis buffer, followed by centrifugation at 12,000 rpm at 4 °C. The Veh- or G3-ME-treated lysates were subsequently divided into two equal portions, each of which was incubated with PBS or protease XIV (100 μg/mL) for 5 min. An equal volume of loading buffer (5×) was added to all the samples, and after heating at 100 °C for 3 min, they were subjected to perform Western blotting.
Cellular thermal shift assay (CETSA)
H9c2 cells and hESC-CMs were treated with 1% NP-40 lysis buffer, followed by centrifugation at 12,000 rpm at 4 °C, after which the supernatants were collected. The cell lysates were then divided into two equal portions, which were incubated with Veh or G3-ME (50 μM) for 2 h at room temperature. The Veh- or G3-ME-treated lysates were subsequently divided into seven equal portions, each subjected to individual treatments at the specified temperatures (40–70 °C) for 7 min. An equal volume of loading buffer (5×) was added to all the samples, and after they were heated at 100 °C for 3 min, they were subjected to Western blotting.
Surface plasmon resonance (SPR) assay
SPR technology was used to experimentally validate the binding between G3-ME and HDAC2. The protein sample was immobilized onto a CM5 chip, and the protein fixation parameters were set as follows: 30 μg/mL, pH 4.5, flow rate 10 μL/min, and 25 °C. Subsequently, ethanolamine was used to block the chip at a speed of 10 μL/min for 420 s. G3-ME was serially diluted (0.04–10 μM, 0.1% DMSO) and then injected at a flow rate of 30 μL/min for 90 s (contact phase) followed by 180 s (dissociation phase) at 25 °C. Finally, the data were collected using Biacore 1 K evaluation software.
RNA sequencing (RNA-Seq)
RNA-Seq was conducted on hESC-CMs to investigate the underlying mechanism of G3-ME in pathological cardiac hypertrophy. Briefly, 1 μg of total RNA was used to construct a cDNA library using the TrueLib mRNA Library Prep Kit for Illumina (Vazyme). The cDNA libraries were then sequenced on an Illumina HiSeq (Illumina; Shanghai, China). The mRNA abundance was measured as fragments per kilobase of exon per million reads mapped (FPKM), and the primary FPKM data were normalized using R (version 4.1.0). Quality control was performed by principal component analysis (PCA). Additionally, Student’s t-test and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis were conducted using relevant R packages.
Network pharmacology analysis
G3-ME targets were predicted using the SwissTargetPrediction, PharmMapper, and TCMSP databases. The potential targets of HCM were searched in the OMIM, DisGeNET, and GeneCards databases. The common targets of G3-ME and HCM were further identified, and analysis of protein-protein interaction networks and KEGG pathway enrichment was performed utilizing appropriate R packages.
Molecular docking
Prior to analysis, the crystal structure of the HDAC2 protein (PDB ID: 7ZZT) was obtained from the RCSB PDB database, and the G3-ME structure (CID: 5281946) was sourced from the PubChem database. Afterward, the HDAC2 protein crystal structure was preprocessed using the PyMOL program to eliminate heteroatoms. The processed HDAC2 protein was subsequently globally docked with G3-ME using the AutoDock Vina 1.2.5 program. The grid calculation spacing parameter in AutoDock Vina was set to 0.375, whereas the exhaustiveness parameter was set to 8. All the docking conformation scores are provided in Supplementary Table 5.
Statistical analysis
The research was carried out using a randomized, double-blind approach. The results are shown as the mean ± SEM. Student’s t-test was used to compare differences between two groups. One-way ANOVA followed by the Bonferroni post hoc correction was used to test the effect of one independent variable or factor. Two-way ANOVA followed by the Bonferroni post hoc correction was used to examine the effect of two independent variables or factors on a dependent variable. Before ANOVA was performed, the distribution of all the data was evaluated. All the data in this study were normally distributed. A P-value less than 0.05 was considered statistically significant.
Results
G3-ME treatment alleviates Ang II-induced cardiomyocyte hypertrophy in H9c2 cells
We utilized various concentrations of G3-ME to treat H9c2 cells in vitro and observed that G3-ME exhibited low toxicity to cardiomyocytes, with an IC50 of 175.3 μM for H9c2 cells (Fig. 1b). Ang II is the most critical endocrine ligand in the renin-angiotensin system, and abnormal elevations in Ang II contribute to a wide spectrum of cardiovascular diseases, including hypertrophy [48]. In this study, cardiomyocytes were stimulated with Ang II to construct a pathological cardiac hypertrophy model in vitro. We subsequently combined different concentrations of G3-ME with Ang II to treat H9c2 cells and found that a concentration of 2.5 µM G3-ME attenuated the expression of the pathological hypertrophy markers, Nppa and Nppb, whereas a concentration of 5 µM resulted in a significant inhibitory effect (Fig. 1c). We further evaluated the effects of 5 µM G3-ME on the expression of Nppa and Nppb in Ang II-induced H9c2 cells at different time points and reported that G3-ME exerted optimal therapeutic effects after 24 h of treatment (Fig. S1). Consequently, we selected 5 µM G3-ME, which is significantly lower than the IC50 concentration, to treat cardiomyocytes for 24 h in subsequent in vitro experiments. G3-ME treatment effectively inhibited the Ang II-induced increase in ANP and BNP protein levels (Fig. 1d). More intuitively, G3-ME significantly reduced the area of cardiomyocytes induced by Ang II (Fig. 1e). These results indicate that G3-ME is beneficial for inhibiting pathological cardiomyocyte hypertrophy.
a Chemical structural formula of G3-ME. b The toxic effects of G3-ME in H9c2 were determined using the CCK-8 assay. n = 6 per group. c The mRNA expression levels of pathological cardiac hypertrophy marker genes Nppa and Nppb. n = 3 per group. d The protein expression levels of ANP and BNP. n = 4 per group. e Representative images of H9c2 cell morphology were obtained through immunofluorescent staining of cTNT (green), and cardiomyocyte size was analyzed from 10 confocal dishes, with all cardiomyocytes in the different groups being 135, 160, 97, and 153, respectively. The values are mean ± SEM. *P < 0.05 vs. control and #P < 0.05 vs. Ang II in (c). *P < 0.05 in (d, e).
G3-ME ameliorates Ang II-induced human cardiomyocyte hypertrophy
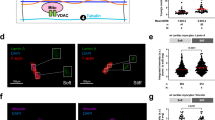
Considering the substantial differences in structure and function between rodent and human cardiomyocytes, we derived human cardiomyocytes from hESCs using a 2D monolayer-based differentiation method (Fig. 2a). Similar results were obtained with H9c2, in which G3-ME showed low toxicity to hESC-CMs, with an IC50 of approximately 245.4 μM (Fig. 2b). Ang II promoted the mRNA and protein expression of NPPA and NPPB in hESC-CMs, and these effects were reversed by G3-ME treatment (Fig. 2c, d). Additionally, G3-ME effectively inhibited the increase in hESC-CM area induced by Ang II (Fig. 2e). Ang II and other hypertrophic stimuli stimulate an increase in intracellular Ca2+ activity, which originates from the perinuclear sarco/endoplasmic reticulum (SR/ER) Ca2+ store. Prolonged Ca2+ activation results in maladaptive transcriptional effects that contribute to the progression of pathological cardiac hypertrophy and heart failure. Notably, G3-ME significantly alleviated Ca2+ activity overload (Fig. 2f). These results suggest that G3-ME has beneficial effects against cardiomyocyte hypertrophy in human and rodent cardiomyocytes.
a Schematic diagram of the induced differentiation of human embryonic stem cells (hESC) into cardiomyocytes (hESC-CMs). b The toxic effects of G3-ME in hESC-CMs were evaluated using the CCK-8 assay. n = 6 per group. c The protein expression levels of ANP and BNP. n = 5 per group. d The mRNA expression levels of NPPA and NPPB. n = 4 per group. e Representative images of hESC-CM morphology were obtained through immunofluorescent staining of cTNT (green), and cardiomyocyte size was analyzed from 10 confocal dishes, with all cardiomyocytes in the different groups being 34, 39, 40, and 44, respectively. f Representative images of calcium (Ca2+) transients were recorded in single cells, and Ca2+ transient amplitude, time to peak, peak to peak, and transient duration 50% in hESC-CMs were analyzed from 6 confocal dishes, with all transient cycles in the different groups being 24, 24, 32, and 25, respectively. The values are mean ± SEM. *P < 0.05.
G3-ME supplementation restores both cardiac systolic and diastolic function in Myh6 R404Q mice with pathological cardiac hypertrophy
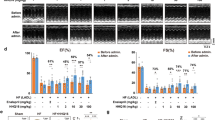
To investigate the role of G3-ME in pathological cardiac hypertrophy in vivo, WT and Myh6R404Q mice were administered different doses of G3-ME (Fig. 3a). The lack of impact of G3-ME treatment on mouse body weight suggested that the in vivo toxicity of G3-ME is minimal (Fig. S2a). Myh6R404Q mice exhibited significant increases in heart volume and weight, which were reversed by both low and high doses of G3-ME (Fig. 3b, c). Echocardiographic assessments revealed that Myh6R404Q mice displayed marked increases in LV mass and wall thickness, and notable decreases in LV volume and diameter. Both low and high doses of G3-ME effectively ameliorated these characteristics in Myh6R404Q mice (Fig. 3d–h). The EF and FS of Myh6R404Q mice are abnormally elevated as a compensatory response to the impaired ejection function resulting from the reduction in LV volume and diameter. Treatment with G3-ME effectively restored both EF and FS while concurrently reducing ventricular wall thickness and increasing ventricular volume (Fig. 3i and Fig. S2b). Additionally, the E peak reflects early diastolic blood flow in the LV and is associated with early diastolic active relaxation, whereas the A peak represents late left ventricular diastolic blood flow, with its level correlated with ventricular compliance. The ratio of the LV E peak to the A peak (E/A) typically ranges from 1.5 to 2.0 and serves as an indicator of cardiac diastolic function. Compared with that in normal WT mice, the E/A ratio in Myh6R404Q mice was significantly lower. However, both low and high doses of G3-ME effectively restored the E/A ratio (Fig. 3d, j). These data indicate that both low-dose and high-dose G3-ME supplementation effectively restored cardiac systolic and diastolic function in Myh6R404Q mice.
a Schematic diagram of the mice experimental procedure. b Representative images of gross heart morphology. c The ratio of heart weight (HW) to tibial length (TL). d Representative images of echocardiograms. e The left ventricular (LV) mass. f Echocardiographic analysis of LV anterior wall (LVAW) or posterior wall (LVPW) thickness in systolic and diastolic periods. g Echocardiographic analysis of LV diameter (LVD). h Echocardiographic analysis of LV volume (LVV). i Echocardiographic analysis of LV ejection fraction. j The ratio of E peak to A peak (E/A) of mice. n = 6 per group. The values are mean ± SEM. *P < 0.05.
G3-ME improves cardiac injury and remodeling in Myh6 R404Q mice
Elevated expression of NPPA and NPPB is commonly used as an indicator of pathological cardiac function impairment [49]. Western blot and qPCR analyses revealed that their expression at both the protein and mRNA level was significantly increased in the LV of Myh6R404Q mice, but was effectively reduced by treatment with varying doses of G3-ME (Fig. 4a, d). Serum NT-proBNP serves as an important biomarker for assessing cardiac load and functional status in patients with pathological myocardial hypertrophy [47]. The results of the serum NT-proBNP test further indicated that G3-ME effectively decreased the cardiac load in Myh6R404Q mice (Fig. 4c). We also examined the effects of G3-ME on pathological cardiac remodeling. H&E staining revealed significant enlargement of the cardiac cross-section and cardiomyocyte area in Myh6R404Q mice, accompanied by a disorganized myofiber arrangement and partial cardiomyocyte damage, whereas both low-dose and high-dose G3-ME intervention effectively ameliorated these phenotypes (Fig. 4f, g). Myocardial fibrosis is frequently considered one of the initial manifestations of cardiac hypertrophy. Masson’s staining revealed significant cardiac fibrosis in Myh6R404Q mice, whereas G3-ME treatment effectively inhibited the development of pathological fibrosis (Fig. 4h, i). Importantly, the expression levels of the myocardial fibrosis markers Acta2, Col1a1, and Col3a1 were significantly elevated in the hearts of Myh6R404Q mice. In contrast, both low-dose and high-dose G3-ME effectively suppressed the expression of these markers (Fig. 4e). Additionally, the results of cardiac WGA staining revealed that treatment with different doses of G3-ME reduced the myocardial cell area in Myh6R404Q mice (Fig. 4j, k). These results collectively suggest that G3-ME treatment plays a crucial role in curbing pathological myocardial remodeling. Consequently, we opted to further evaluate the therapeutic effects of low-dose G3-ME across different models of pathological cardiac hypertrophy.
a, b The protein expression levels of ANP and BNP in cardiac tissues. n = 4 per group. c The serum NT-proBNP levels. n = 5 per group. d, e The mRNA expression levels of Nppa, Nppb, Acta2, Col1a1, and Col3a1 in cardiac tissues. n = 4 per group. f Representative images of cardiac H&E staining. g The cardiac cross-sectional area determined by H&E staining. n = 6 per group. h Representative images of the cardiac fibrosis area using Masson’s staining. i Quantification of fibrosis area by Masson’s staining. n = 6 per group. j Representative images of cardiomyocytes by WGA staining. k Analysis of cardiomyocyte size using WGA staining from 6 mice, with all cardiomyocytes in the same area analyzed in each section. n = 375, 341, 328, 283, 353, and 340, respectively. The values are mean ± SEM. *P < 0.05.
G3-ME improves both cardiac function and structure in Tnnt2 R109Q mice with pathological cardiac hypertrophy
To further elucidate the role of G3-ME in pathological cardiac hypertrophy and HCM, we administered G3-ME to Tnnt2R109Q mice, another well-established mouse model of HCM, and assessed its effects on both cardiac function and structure in pathological cardiac hypertrophy (Fig. 5a). Similarly, G3-ME did not significantly affect the body weight of the mice (Fig. S2c). Tnnt2R109Q mice also exhibited significant increases in heart volume and weight, LV mass, EF, FS, and wall thickness, all of which were reversed by G3-ME treatment (Fig. 5b–d and Fig. S2d, e). In addition, G3-ME supplementation effectively restored the E/A ratio (Fig. 5c, d) and significantly reduced serum NT-proBNP levels in Tnnt2R109Q mice (Fig. 5e). H&E staining, Masson’s staining, and WGA staining revealed that the cardiac cross-sectional area, cardiomyocyte size, and degree of cardiac fibrosis in Tnnt2R109Q mice were significantly altered, accompanied by disorganized myofiber arrangement and partial cardiomyocyte damage, whereas G3-ME intervention effectively ameliorated these phenotypes (Fig. 5f–k). Notably, G3-ME also reduced the expression of the cardiac injury markers Nppa and Nppb (Fig. S2f) and the myocardial fibrosis markers Acta2, Col1a1, and Col3a1 in Tnnt2R109Q mice (Fig. S2g). These results further reinforce the protective effects of G3-ME on HCM and pathological cardiac hypertrophy, suggesting its potential as an effective therapeutic agent for this condition.
a Schematic diagram of the mice experimental procedure. b The ratio of heart weight (HW) to tibial length (TL). c Representative images of echocardiograms. d Echocardiographic analysis of left ventricular (LV) anterior wall (LVAW) or posterior wall (LVPW) thickness in systolic and diastolic periods, and the ratio of E peak to A peak (E/A). n = 6 per group. e The serum NT-proBNP levels. n = 5 per group. f Representative images of cardiac H&E staining. g The cardiac cross-sectional area determined by H&E staining. n = 5 per group. h Representative images of the cardiac fibrosis area using Masson’s staining. i Quantification of fibrosis area by Masson’s staining. n = 5 per group. j Representative images of cardiomyocytes by WGA staining. k Analysis of cardiomyocyte size using WGA staining from 6 mice, with all cardiomyocytes in the same area analyzed in each section. n = 332, 348, 292, and 319, respectively. The values are mean ± SEM. *P < 0.05.
G3-ME inhibits pathological cardiac hypertrophy by regulating the PI3K-AKT signaling pathway
To investigate the mechanisms underlying the potential beneficial effects of G3-ME in relieving cardiac hypertrophy, we treated hESC-CMs with Ang II or a combination of G3-ME and Ang II for 12 h, followed by RNA-Seq analysis (Fig. 6a). PCA and the heatmap demonstrated clear separation of gene expression between the different groups (Fig. 6b, c). We subsequently conducted KEGG pathway enrichment analysis of the differentially expressed genes (P < 0.05 and fold change ≥1.5) in hESC-CMs. The results of the KEGG analysis revealed significant alterations in the PI3K-AKT signaling pathway upon G3-ME treatment, which were closely associated with myocardial hypertrophy and HCM (Fig. 6d).
a Schematic diagram for exploring the key targets and signaling pathways of G3-ME. b Principal component analysis (PCA) of the angiotensin II (Ang II)-treated and the G3-ME + Ang II-treated RNA sequence. c Heatmap of differential genes. d KEGG enrichment analysis according to differential genes. e–i The protein expression levels of p-P85, P85, p-AKT, and AKT in H9c2, hESC-CMs, and cardiac tissues of wild-type (WT) or Myh6R404Q mice. n = 4 per group. The values are mean ± SEM. *P < 0.05.
We further investigated the effects of G3-ME on the PI3K-AKT signaling pathway in Ang II-induced hypertrophic cardiomyocytes and in the hearts of low-dose G3-ME-treated WT or Myh6R404Q mice. The results indicated that Ang II activated the PI3K-AKT signaling pathway in both H9c2 cells and hESC-CMs, as evidenced by increased phosphorylation levels of PI3K P85 and AKT. However, G3-ME effectively inhibited the phosphorylation of both P85 and AKT (Fig. 6e–i). Furthermore, a low dose of G3-ME significantly prevented the abnormal increase in the phosphorylation of P85 and AKT in the hearts of Myh6R404Q mice (Fig. 6g–i). These results collectively reveal that the ability of G3-ME to reduce hypertrophy is related to the deactivation of the PI3K-AKT pathway.
G3-ME prevents pathological cardiac hypertrophy by directly targeting HDAC2
Surprisingly, as an analog of galangin, the potential biological targets of G3-ME have never been reported. To investigate the direct target of G3-ME in the regulation of pathological cardiac hypertrophy, we screened 365 potential targets of G3-ME from various drug target databases on the basis of G3-ME structural characteristics. Concurrently, we identified 1149 targets associated with the onset and progression of HCM from three disease-related target databases. Through a co-analysis of the potential targets of G3-ME and HCM-related genes, we identified 42 possible targets of G3-ME that may play crucial roles in regulating pathological cardiac hypertrophy (Fig. 7a). KEGG pathway enrichment analysis revealed that these 42 genes were also significantly enriched in the PI3K-AKT signaling pathway (Fig. 7b). These findings were consistent with the RNA-Seq results, suggesting that G3-ME indeed inhibited pathological cardiac hypertrophy by negatively regulating the PI3K-AKT signaling pathway. Furthermore, we singly performed molecular docking on the 42 potential targets of G3-ME. The results indicated that HDAC2 exhibited the highest binding energy with G3-ME (Fig. 7c and Supplementary Table 5), with HDAC2 capable of forming hydrogen bonds with the HIS62, ARG60, and TYR19 residues, thus creating a binding pocket (Fig. 7d).
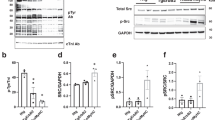
a The flowchart of network pharmacology and the protein-protein interaction diagram. b KEGG enrichment analysis according to 42 potential targets of G3-ME. c The binding energy between G3-ME and potential targets enriched in the PI3K-AKT signaling pathway. d Representative schematic diagram of molecular docking between HDAC2 and G3-ME. e Schematic diagram of HDAC2 regulating the PI3K-AKT signaling pathway. f, g The protein expression levels of p-HDAC2 and HDAC2 in H9c2, hESC-CMs and cardiac tissues of wild-type (WT) or Myh6R404Q mice. n = 4 per group. h Drug affinity reaction target stability (DARTs) assay was performed in vitro and in vivo on H9c2. i DARTs assay was performed in vitro and in vivo on hESC-CMs. j Cellular thermal shift assay was performed on H9c2 and hESC-CMs, respectively. k The results of surface plasmon resonance indicated a direct binding interaction between G3-ME and HDAC2, with a binding affinity of 3.36 μM. The values are mean ± SEM. *P < 0.05.
The available data indicate that HDAC2 is activated mainly through phosphorylation at Ser394, which subsequently stimulates an intracellular signaling cascade downstream of the PI3K-AKT pathway, ultimately resulting in cardiac hypertrophy (Fig. 7e) [15, 22,23,24]. We detected the activity of HDAC2 in Ang II-induced hypertrophic cardiomyocytes and in the hearts of HCM mice. Specifically, Ang II increased HDAC2 phosphorylation levels in both H9c2 cells and hESC-CMs, whereas G3-ME reversed the effects of Ang II. As anticipated, HDAC2 phosphorylation levels were also elevated in the hearts of HCM mice, and G3-ME intervention effectively reduced HDAC2 phosphorylation in these hearts (Fig. 7f, g).
We further examined whether G3-ME directly binds to HDAC2. Both in vitro and in vivo DARTs experiments demonstrated that G3-ME effectively prevented the protease-induced degradation of HDAC2 in both rat cardiomyocytes and human cardiomyocytes, suggesting that G3-ME enhanced the structural stability of HDAC2 (Fig. 7h, i). Additionally, CETSA experiments revealed that as the temperature increased, the degradation of HDAC2 became more pronounced in both H9c2 cells and hESC-CMs. However, treatment with G3-ME significantly increased the stability of HDAC2 (Fig. 7j and Fig. S3). More importantly, the results of SPR, a ligand competition assay, indicated a direct binding interaction between G3-ME and HDAC2, with a binding affinity of 3.36 μM (Fig. 7k). Taken together, these findings indicate that G3-ME inhibits the phosphorylation of HDAC2 by directly binding to it, thereby negatively regulating the PI3K-AKT signaling pathway.
G3-ME reduces the activity of HDAC2 by decreasing its phosphorylation at Ser394
Phosphorylation at Ser394 of HDAC2 is crucial for its activation, which mediates the onset and progression of pathological cardiac hypertrophy [15, 20]. Available data indicate that casein kinase 2α (CK2α), PIM1, and G protein-coupled receptor kinase 3 can regulate the phosphorylation of HDAC2, with CK2α being the most significant regulator at Ser394 [25]. Additionally, the protein phosphatase 2 catalytic subunit alpha (PP2CA) and protein phosphatase 1 regulatory subunits are associated with the dephosphorylation of HDAC2, with PP2CA primarily responsible for dephosphorylation at Ser394 [50]. We thus speculated that CK2α or PP2CA could regulate HDAC2 protein phosphorylation through direct interactions with HDAC2. To this end, endogenous HDAC2 was immunoprecipitated and subjected to immunoblotting, and CK2α expression was found to be significantly decreased in G3-ME-treated precipitates, whereas the interaction between PP2CA and HDAC2 was not affected by G3-ME (Fig. S4a). Endogenous CK2α or PP2CA was further immunoprecipitated and subjected to immunoblotting to detect HDAC2. The results revealed that HDAC2 expression was also markedly reduced in G3-ME-treated CK2α-captured precipitates, but not in PP2CA-captured precipitates (Fig. S4b, c). Taken together, these results suggest that G3-ME directly interacts with HDAC2, preventing its interaction with CK2α, but not affecting its binding with PP2CA, ultimately leading to the inhibition of HDAC phosphorylation.
Interference with HDAC2 expression influences the cardioprotective effect of G3-ME on PI3K-AKT signaling and cardiac hypertrophy
To investigate the cardioprotective effects of G3-ME against cardiac hypertrophy through the targeting of HDAC2, we established stable cell lines with HDAC2 knockdown in H9c2 cells. As shown in Fig. 8a, the efficiency of HDAC2 knockdown was approximately 75%. The simple knockdown of HDAC2 expression significantly inhibited the mRNA expression of Nppa and Nppb. Notably, HDAC2 knockdown effectively abolished the Ang II-induced pathological hypertrophic effects, including the increased expression of Nppa and Nppb and the enlargement of the cardiomyocyte area. Additionally, the inhibitory effects of G3-ME on Ang II were also nullified by the knockdown of HDAC2 (Fig. 8b–e).
a The efficiency of HDAC2 knockdown (HDAC2-KD) in H9c2. b, c The mRNA expression levels of Nppa and Nppb in normal control (NC) or HDAC2-KD cells. n = 4 per group. d Representative images of NC or HDAC2-KD cell morphology were obtained through immunofluorescent staining of cTNT (red). e The cardiomyocyte size was analyzed from 8 confocal dishes, with all cardiomyocytes in the different groups being 95, 106, 101, 101, 102, and 95, respectively. f The protein expression levels of p-P85, P85, p-AKT and AKT in NC or HDAC2-KD cells. g The efficiency of HDAC2 overexpression (HDAC2-OE) in H9c2 cells. h, i The mRNA expression levels of Nppa and Nppb in control vector (Ctrl) or HDAC2-OE cells. n = 4 per group. j Representative images of Ctrl or HDAC2-OE cell morphology were obtained through immunofluorescent staining of cTNT (green). k The cardiomyocyte size was analyzed from 8 confocal dishes, with all cardiomyocytes in the different groups being 98, 84, 93, 81, 76, and 77, respectively. l The protein expression levels of p-P85, P85, p-AKT and AKT in Ctrl or HDAC2-OE cells. The values are mean ± SEM. *P < 0.05.
We further constructed an HDAC2 stable overexpression plasmid, and the efficiency of HDAC2 overexpression surpassed 90% in H9c2 cells (Fig. 8g). As anticipated, the overexpression of HDAC2 led to increased expression levels of Nppa and Nppb, as well as an enlargement of the cardiomyocyte area, which was further exacerbated by Ang II treatment. Although G3-ME partially alleviated the hypertrophic effects induced by Ang II in the context of HDAC2 overexpression, its beneficial effect was significantly diminished compared with that in the group with normal HDAC2 expression (Fig. 8h–k).
The effects of PI3K-AKT signaling were further investigated in G3-ME-treated HDAC2 knockdown or overexpression cardiomyocytes. HDAC2 knockdown inhibited the activity of the PI3K-AKT signaling pathway, regardless of treatment with Ang II or G3-ME (Fig. 8f). In contrast, HDAC2 overexpression activated the PI3K-AKT signaling pathway, with Ang II further enhancing the activation effect of HDAC2 overexpression. Consistent with these findings, under conditions of HDAC2 overexpression, G3-ME partially suppressed the Ang II-induced activation of the PI3K-AKT pathway. However, compared with that in the group with normal HDAC2 expression, its inhibitory effect was significantly reduced (Fig. 8l).
Existing evidence indicates that HDAC2 leads to the constitutive activation of AKT signaling by suppressing the expression of INPP5F [15]. We hypothesized that G3-ME inhibited HDAC2, which in turn activated INPP5F, resulting in the inactivation of AKT signaling and thereby the suppression of pathological cardiac hypertrophy. We first examined the effects of HDAC2 overexpression or knockdown on INPP5F expression and found that HDAC2 overexpression inhibited INPP5F expression, whereas HDAC2 knockdown promoted INPP5F expression (Fig. S5a, b). We further established a stable cardiomyocyte cell line with INPP5F knockdown and treated it with Ang II or G3-ME. The results demonstrated that INPP5F knockdown increased AKT phosphorylation, which was amplified by Ang II, whereas G3-ME treatment effectively inhibited this phosphorylation (Fig. S5c). These findings suggest that G3-ME plays a crucial role in regulating the progression of cardiac hypertrophy mediated by the HDAC2-INPP5F axis.
In summary, G3-ME inhibits pathological cardiac hypertrophy by directly targeting HDAC2 and inactivating PI3K-AKT signaling. The combined application of G3-ME and HDAC2 inhibitors is promising for alleviating cardiac hypertrophy and the consequent heart failure.
Discussion
Pathological cardiac hypertrophy is the adaptive cardiac response to various types of stress or pathological conditions [2]. This sustained abnormal response results in the enlargement and fibrosis of myocardial cells and tissues, which gradually diminishes cardiac pumping ability and may ultimately lead to heart failure [51, 52]. A significant new discovery of this study is that G3-ME effectively inhibited the progression of pathological cardiac hypertrophy. Specifically, G3-ME not only alleviates cardiac hypertrophy induced by stress stimulation, as represented by Ang II, but also effectively delays the disease progression of hereditary HCM. Mechanistically, G3-ME directly targets HDAC2, and the inactivation of HDAC2 mediated by G3-ME negatively influences the PI3K-AKT signaling pathway, which is vital for hindering the progression of pathological cardiac hypertrophy (Fig. 9). These results suggest that G3-ME may serve as an important target for therapeutic intervention in patients with both hereditary HCM and pathological cardiac hypertrophy.
Galangin 3-methyl ether (G3-ME) alleviates pathological cardiac hypertrophy both in vivo and in vitro by targeting HDAC2 and subsequently deactivating the PI3K-AKT signaling pathway.
G3-ME (also named 3-O-methylgalangin) is a natural flavonoid extracted from galangal. Galangal and its extracts exhibit distinct pharmacological effects, including strong antioxidant, anti-inflammatory, hypoglycemic, and anticancer effects [53]. The principal active constituents of galangal include galangin, curcumin, quercetin, kaempferide, and G3-ME, among others. Previous research has predominantly concentrated on galangin and curcumin. Evidence from multiple studies has indicated that galangin and curcumin have significant cardioprotective effects, including against cardiac hypertrophy [30, 54]. However, we were surprised to discover a lack of research on G3-ME in the context of heart diseases. Galangin undergoes methylation at the third hydroxyl position to form G3-ME (Fig. 1a). This methylation process reduces the polarity of G3-ME while increasing its lipophilicity, thereby facilitating its cellular entry and regulatory effects. Furthermore, the methylation of the hydroxyl group enhances the stability of G3-ME against oxidative stress and metabolic degradation, contributing to its sustained efficacy, which includes significant anti-inflammatory and antitumor effects. This research not only enhances the pharmacological capabilities of galangal and its active ingredients but also lays the groundwork for the development of G3-ME-based treatments for pathological cardiac hypertrophy and inherited HCM.
Pathological cardiac hypertrophy is a common pathophysiological feature of numerous heart conditions, such as inherited HCM [55]. While HCM is linked mainly to mutations in genes that encode sarcomere proteins, the key pathological effects include a reduction in ventricular chamber size and the presence of cardiac hypertrophy. This condition is the foremost contributor to unexpected cardiac fatalities in younger populations [56]. However, there is currently no effective clinical method to completely reverse the progression of HCM. Pharmacologic therapy remains the primary treatment approach for HCM, but no drug has been proven to provide a cure. In 2016, mavacamten (MYK-461), a targeted sarcomere power inhibitor, was identified and gained widespread attention [57]. Recent clinical research has indicated that mavacamten is effective at diminishing left ventricular outflow tract (LVOT) obstruction and enhancing exercise capacity among individuals diagnosed with obstructive HCM [58, 59]. However, considering its severe side effects of inducing heart failure, as well as its high treatment costs, mavacamten is primarily recommended for patients with severe LVOT obstructive HCM [60]. Consequently, identifying novel therapeutic strategies and targets for treating HCM is critical. In the present study, G3-ME, a natural flavonoid extracted from the rhizome of galangal, was demonstrated to improve pathological cardiac hypertrophy and ventricular remodeling associated with HCM caused by point mutations in various genes. Additionally, lower doses of G3-ME resulted in superior therapeutic effects in in vitro experiments, providing a solid foundation for the development of novel targeted treatments for HCMs based on G3-ME.
Most of the previous cellular research on the role of galangin, curcumin, or other natural products in heart diseases has been restricted to mouse- or rat-derived cardiomyocytes. The physiological, genetic, and mechanistic differences between small animal models and humans are substantial and cannot be overlooked [61]. To address this limitation, in this study, human cardiomyocytes were derived from differentiating hESCs to investigate the inhibitory effects of G3-ME on Ang II-induced pathological cardiac hypertrophy in vitro. This study revealed that Ang II enhances the expression of hypertrophy marker genes and increases the area of hESC-CMs. However, these effects were effectively reversed by G3-ME treatment. Notably, abnormal enlargement of the cardiomyocyte area is a fundamental aspect of the progression of cardiac hypertrophy and HCM. This study provides direct and compelling evidence that G3-ME may be beneficial for inhibiting the progression of pathological cardiac hypertrophy or HCM in humans.
A significant and noteworthy finding of this study is that HDAC2 was proven to be the direct binding target of G3-ME, and G3-ME effectively prevented the phosphorylation of HDAC2 through its binding. While class II HDACs are known to act as negative regulators of cardiac hypertrophy, HDAC2 has been demonstrated to function as a pro-hypertrophic molecule [15,16,17,18,19]. The development of cardiac hypertrophy is completely blocked by a nonselective HDAC inhibitor or an inhibitor that specifically targets class I HDACs [19]. Our findings indicate that in HDAC2 knockdown cardiomyocytes, the pathogenic effects of hypertrophic stimuli, such as Ang II, are completely abolished, which aligns with the findings of previous studies. The pro-hypertrophic effect of Ang II is further intensified in cardiomyocytes with stable HDAC2 overexpression, which is attenuated by treatment with G3-ME. We hypothesize that G3-ME may function as a natural targeted inhibitor of HDAC2, thereby preventing HDAC2-mediated pathological cardiac hypertrophy. Although the effects of G3-ME on cardiac hypertrophy were not observed in HDAC2-specific knockout or overexpression animals in this study, previous studies have indicated that the overexpression of HDAC2 alone can promote cardiac hypertrophy. In contrast, the deletion of HDAC2 impedes the progression of cardiac hypertrophy in TAC mice. On the basis of these data and our findings from HDAC2 knockdown and overexpression cell lines, we speculate that G3-ME may have therapeutic potential for addressing pathological cardiac hypertrophy resulting from HDAC2 overexpression in animal models or patients, which requires further investigation. Another significant finding of the present study is that G3-ME inhibits the interaction between CK2α and HDAC2, thereby reducing the phosphorylation level of HDAC2. We hypothesize that the binding of G3-ME to HDAC2 induces a conformational change that obstructs the interaction of CK2α with HDAC2. We are actively investigating this intriguing and significant question.
In summary, G3-ME alleviates pathological cardiac hypertrophy both in vivo and in vitro by targeting HDAC2 and subsequently deactivating the PI3K-AKT signaling pathway (Fig. 9). Thus, G3-ME is a promising option for treating pathological hypertrophy.
References
Gong R, Jiang Z, Zagidullin N, Liu T, Cai B. Regulation of cardiomyocyte fate plasticity: a key strategy for cardiac regeneration. Signal Transduct Target Ther. 2021;6:31.
Nakamura M, Sadoshima J. Mechanisms of physiological and pathological cardiac hypertrophy. Nat Rev Cardiol. 2018;15:387–407.
Wang L, Feng J, Feng X, Meng D, Zhao X, Wang J, et al. Exercise-induced circular RNA circUtrn is required for cardiac physiological hypertrophy and prevents myocardial ischaemia-reperfusion injury. Cardiovasc Res. 2023;119:2638–52.
Claessen G, De Bosscher R, Janssens K, Young P, Dausin C, Claeys M, et al. Reduced ejection fraction in elite endurance athletes: clinical and genetic overlap with dilated cardiomyopathy. Circulation. 2024;149:1405–15.
Zhao D, Xu R, Zhou Y, Wu J, Zhang X, Lin H, et al. ORP5 promotes cardiac hypertrophy by regulating the activation of mTORC1 on lysosome. J Adv Res. 2024;S2090-1232(24)00591–5.
Lan F, Lee AS, Liang P, Sanchez-Freire V, Nguyen PK, Wang L, et al. Abnormal calcium handling properties underlie familial hypertrophic cardiomyopathy pathology in patient-specific induced pluripotent stem cells. Cell Stem Cell. 2013;12:101–13.
Masri A, Cardoso RN, Abraham TP, Claggett BL, Coats CJ, Hegde SM, et al. Effect of Aficamten on cardiac structure and function in obstructive hypertrophic cardiomyopathy: SEQUOIA-HCM CMR substudy. J Am Coll Cardiol. 2024;84:1806–17.
Abdellatif M, Madeo F, Kroemer G, Sedej S. Spermidine overrides INSR (insulin receptor)-IGF1R (insulin-like growth factor 1 receptor)-mediated inhibition of autophagy in the aging heart. Autophagy. 2022;18:2500–2.
Li J, Yousefi K, Ding W, Singh J, Shehadeh LA. Osteopontin RNA aptamer can prevent and reverse pressure overload-induced heart failure. Cardiovasc Res. 2017;113:633–43.
Marat AL, Wallroth A, Lo WT, Muller R, Norata GD, Falasca M, et al. mTORC1 activity repression by late endosomal phosphatidylinositol 3,4-bisphosphate. Science. 2017;356:968–72.
Stephens L, Anderson K, Stokoe D, Erdjument-Bromage H, Painter GF, Holmes AB, et al. Protein kinase B kinases that mediate phosphatidylinositol 3,4,5-trisphosphate-dependent activation of protein kinase B. Science. 1998;279:710–4.
Chou CH, Hung CS, Liao CW, Wei LH, Chen CW, Shun CT, et al. IL-6 trans-signalling contributes to aldosterone-induced cardiac fibrosis. Cardiovasc Res. 2018;114:690–702.
Izumo S, Nadal-Ginard B, Mahdavi V. Protooncogene induction and reprogramming of cardiac gene expression produced by pressure overload. Proc Natl Acad Sci USA. 1988;85:339–43.
Warren JS, Tracy CM, Miller MR, Makaju A, Szulik MW, Oka SI, et al. Histone methyltransferase Smyd1 regulates mitochondrial energetics in the heart. Proc Natl Acad Sci USA. 2018;115:E7871–E7880.
Trivedi CM, Luo Y, Yin Z, Zhang M, Zhu W, Wang T, et al. Hdac2 regulates the cardiac hypertrophic response by modulating Gsk3 beta activity. Nat Med. 2007;13:324–31.
Wolffe AP. Histone deacetylase: a regulator of transcription. Science. 1996;272:371–2.
Ekwall K. Genome-wide analysis of HDAC function. Trends Genet. 2005;21:608–15.
Trivedi CM, Zhu W, Wang Q, Jia C, Kee HJ, Li L, et al. Hopx and Hdac2 interact to modulate Gata4 acetylation and embryonic cardiac myocyte proliferation. Dev Cell. 2010;19:450–9.
Eom GH, Nam YS, Oh JG, Choe N, Min HK, Yoo EK, et al. Regulation of acetylation of histone deacetylase 2 by p300/CBP-associated factor/histone deacetylase 5 in the development of cardiac hypertrophy. Circ Res. 2014;114:1133–43.
Yoon S, Kim M, Lee H, Kang G, Bedi K, Margulies KB, et al. S-Nitrosylation of histone deacetylase 2 by neuronal nitric oxide synthase as a mechanism of diastolic dysfunction. Circulation. 2021;143:1912–25.
Zhang CL, McKinsey TA, Chang S, Antos CL, Hill JA, Olson EN. Class II histone deacetylases act as signal-responsive repressors of cardiac hypertrophy. Cell. 2002;110:479–88.
Kee HJ, Eom GH, Joung H, Shin S, Kim JR, Cho YK, et al. Activation of histone deacetylase 2 by inducible heat shock protein 70 in cardiac hypertrophy. Circ Res. 2008;103:1259–69.
Lin H, Zhu Y, Zheng C, Hu D, Ma S, Chen L, et al. Antihypertrophic memory after regression of exercise-induced physiological myocardial hypertrophy is mediated by the long noncoding RNA Mhrt779. Circulation. 2021;143:2277–92.
Shang L, Pin L, Zhu S, Zhong X, Zhang Y, Shun M, et al. Plantamajoside attenuates isoproterenol-induced cardiac hypertrophy associated with the HDAC2 and AKT/ GSK-3beta signaling pathway. Chem Biol Interact. 2019;307:21–28.
Eom GH, Cho YK, Ko JH, Shin S, Choe N, Kim Y, et al. Casein kinase-2alpha1 induces hypertrophic response by phosphorylation of histone deacetylase 2 S394 and its activation in the heart. Circulation. 2011;123:2392–403.
Yang T, Liu H, Yang C, Mo H, Wang X, Song X, et al. Galangin attenuates myocardial ischemic reperfusion-induced ferroptosis by targeting Nrf2/Gpx4 signaling pathway. Drug Des Devel Ther. 2023;17:2495–511.
Zhang J, Hu S, Gao Y, Wei X, Qu Y, Gao R, et al. Galangin alleviated myocardial ischemia-reperfusion injury by enhancing autophagic flux and inhibiting inflammation. Eur J Pharmacol. 2023;945:175621.
Thapa R, Afzal O, Alfawaz Altamimi AS, Goyal A, Almalki WH, Alzarea SI, et al. Galangin as an inflammatory response modulator: an updated overview and therapeutic potential. Chem Biol Interact. 2023;378:110482.
Abukhalil MH, Althunibat OY, Aladaileh SH, Al-Amarat W, Obeidat HM, Al-Khawalde AAA, et al. Galangin attenuates diabetic cardiomyopathy through modulating oxidative stress, inflammation and apoptosis in rats. Biomed Pharmacother. 2021;138:111410.
Thangaiyan R, Arjunan S, Govindasamy K, Khan HA, Alhomida AS, Prasad NR. Galangin attenuates isoproterenol-induced inflammation and fibrosis in the cardiac tissue of Albino Wistar rats. Front Pharmacol. 2020;11:585163.
Fang G, Li X, Yang F, Huang T, Qiu C, Peng K, et al. Galangin attenuates doxorubicin-induced cardiotoxicity via activating nuclear factor erythroid 2-related factor 2/heme oxygenase 1 signaling pathway to suppress oxidative stress and inflammation. Phytother Res. 2023;37:5854–70.
Shu G, Chen K, Li J, Liu B, Chen X, Wang J, et al. Galangin alleviated Doxorubicin-induced cardiotoxicity by inhibiting ferroptosis through GSTP1/JNK pathway. Phytomedicine. 2024;134:155989.
Zhang F, Zhou H, Xue J, Zhang Y, Zhou L, Leng J, et al. Deficiency of transcription factor Sp1 contributes to hypertrophic cardiomyopathy. Circ Res. 2024;134:290–306.
Belus A, Piroddi N, Scellini B, Tesi C, D’Amati G, Girolami F, et al. The familial hypertrophic cardiomyopathy-associated myosin mutation R403Q accelerates tension generation and relaxation of human cardiac myofibrils. J Physiol. 2008;586:3639–44.
Nag S, Sommese RF, Ujfalusi Z, Combs A, Langer S, Sutton S, et al. Contractility parameters of human beta-cardiac myosin with the hypertrophic cardiomyopathy mutation R403Q show loss of motor function. Sci Adv. 2015;1:e1500511.
Tyska MJ, Hayes E, Giewat M, Seidman CE, Seidman JG, Warshaw DM. Single-molecule mechanics of R403Q cardiac myosin isolated from the mouse model of familial hypertrophic cardiomyopathy. Circ Res. 2000;86:737–44.
Lorenzini M, Norrish G, Field E, Ochoa JP, Cicerchia M, Akhtar MM, et al. Penetrance of hypertrophic cardiomyopathy in sarcomere protein mutation carriers. J Am Coll Cardiol. 2020;76:550–9.
Maass AH, Ikeda K, Oberdorf-Maass S, Maier SK, Leinwand LA. Hypertrophy, fibrosis, and sudden cardiac death in response to pathological stimuli in mice with mutations in cardiac troponin T. Circulation. 2004;110:2102–9.
Sparrow AJ, Sievert K, Patel S, Chang YF, Broyles CN, Brook FA, et al. Measurement of myofilament-localized calcium dynamics in adult cardiomyocytes and the effect of hypertrophic cardiomyopathy mutations. Circ Res. 2019;124:1228–39.
Tardiff JC, Hewett TE, Palmer BM, Olsson C, Factor SM, Moore RL, et al. Cardiac troponin T mutations result in allele-specific phenotypes in a mouse model for hypertrophic cardiomyopathy. J Clin Invest. 1999;104:469–81.
Li B, Yang H, Wang X, Zhan Y, Sheng W, Cai H, et al. Engineering human ventricular heart muscles based on a highly efficient system for purification of human pluripotent stem cell-derived ventricular cardiomyocytes. Stem Cell Res Ther. 2017;8:202.
Huang J, Liu Y, Chen JX, Lu XY, Zhu WJ, Qin L, et al. Harmine is an effective therapeutic small molecule for the treatment of cardiac hypertrophy. Acta Pharmacol Sin. 2022;43:50–63.
Ye C, Zheng F, Xu T, Wu N, Tong Y, Xiong XQ, et al. Norepinephrine acting on adventitial fibroblasts stimulates vascular smooth muscle cell proliferation via promoting small extracellular vesicle release. Theranostics. 2022;12:4718–33.
Ye C, Geng Z, Zhang LL, Zheng F, Zhou YB, Zhu GQ, et al. Chronic infusion of ELABELA alleviates vascular remodeling in spontaneously hypertensive rats via anti-inflammatory, anti-oxidative and anti-proliferative effects. Acta Pharmacol Sin. 2022;43:2573–84.
Ye C, Tong Y, Wu N, Wan GW, Zheng F, Chen JY, et al. Inhibition of miR-135a-5p attenuates vascular smooth muscle cell proliferation and vascular remodeling in hypertensive rats. Acta Pharmacol Sin. 2021;42:1798–807.
Davoodi M, Segal S, Kirschner Peretz N, Kamoun D, Yaniv Y. Semi-automated program for analysis of local Ca2+ spark release with application for classification of heart cell type. Cell Calcium. 2017;64:83–90.
Coats CJ, Gallagher MJ, Foley M, O’Mahony C, Critoph C, Gimeno J, et al. Relation between serum N-terminal pro-brain natriuretic peptide and prognosis in patients with hypertrophic cardiomyopathy. Eur Heart J. 2013;34:2529–37.
Zou Y, Komuro I, Yamazaki T, Kudoh S, Aikawa R, Zhu W, et al. Cell type-specific angiotensin II-evoked signal transduction pathways: critical roles of Gbetagamma subunit, Src family, and Ras in cardiac fibroblasts. Circ Res. 1998;82:337–45.
Nishikimi T, Maeda N, Matsuoka H. The role of natriuretic peptides in cardioprotection. Cardiovasc Res. 2006;69:318–28.
Yoon S, Kim M, Min HK, Lee YU, Kwon DH, Lee M, et al. Inhibition of heat shock protein 70 blocks the development of cardiac hypertrophy by modulating the phosphorylation of histone deacetylase 2. Cardiovasc Res. 2019;115:1850–60.
Hou J, Kang YJ. Regression of pathological cardiac hypertrophy: signaling pathways and therapeutic targets. Pharmacol Ther. 2012;135:337–54.
Lu QB, Ding Y, Liu Y, Wang ZC, Wu YJ, Niu KM, et al. Metrnl ameliorates diabetic cardiomyopathy via inactivation of cGAS/STING signaling dependent on LKB1/AMPK/ULK1-mediated autophagy. J Adv Res. 2023;51:161–79.
Wu Y, Luo J, Xu B. Insights into the anticancer effects of galangal and galangin: a comprehensive review. Phytomedicine. 2024;135:156085.
Li H, Sureda A, Devkota HP, Pittala V, Barreca D, Silva AS, et al. Curcumin, the golden spice in treating cardiovascular diseases. Biotechnol Adv. 2020;38:107343.
Maron BJ. Hypertrophic cardiomyopathy. Lancet. 1997;350:127–33.
Polovina M, Tschope C, Rosano G, Metra M, Crea F, Mullens W, et al. Incidence, risk assessment and prevention of sudden cardiac death in cardiomyopathies. Eur J Heart Fail. 2023;25:2144–63.
Green EM, Wakimoto H, Anderson RL, Evanchik MJ, Gorham JM, Harrison BC, et al. A small-molecule inhibitor of sarcomere contractility suppresses hypertrophic cardiomyopathy in mice. Science. 2016;351:617–21.
Olivotto I, Oreziak A, Barriales-Villa R, Abraham TP, Masri A, Garcia-Pavia P, et al. Mavacamten for treatment of symptomatic obstructive hypertrophic cardiomyopathy (EXPLORER-HCM): a randomised, double-blind, placebo-controlled, phase 3 trial. Lancet. 2020;396:759–69.
Tian Z, Li L, Li X, Wang J, Zhang Q, Li Z, et al. Effect of mavacamten on Chinese patients with symptomatic obstructive hypertrophic cardiomyopathy: the EXPLORER-CN randomized clinical trial. JAMA Cardiol. 2023;8:957–65.
Ho CY, Mealiffe ME, Bach RG, Bhattacharya M, Choudhury L, Edelberg JM, et al. Evaluation of mavacamten in symptomatic patients with nonobstructive hypertrophic cardiomyopathy. J Am Coll Cardiol. 2020;75:2649–60.
Ramachandra CJA, Chua J, Cong S, Kp MMJ, Shim W, Wu JC, et al. Human-induced pluripotent stem cells for modelling metabolic perturbations and impaired bioenergetics underlying cardiomyopathies. Cardiovasc Res. 2021;117:694–711.
Acknowledgements
We are deeply grateful to Professor Yael Yaniv’s team for developing the program for the analysis of Ca2+ spark release. We would like to express our gratitude to Jian-zhen Lei (co-author) for their selfless assistance and guidance during the revision and proofreading of this article.
Funding
This work was supported by Young Scientists Fund of the National Natural Science Foundation of China (No.82300393), General Program of the National Natural Science Foundation of China (No. 82470364), National Science and Technology Major Project of the Ministry of Science and Technology of China (No.2023ZD0504601), Natural Science Foundation of Jiangsu Province (BK20231048), Jiangsu Funding Program for Excellent Postdoctoral Talent (2023ZB341), Wuxi Science and Technology Development Fund Project (K20221019), and Scientific Research and Practice Innovation Program of Jiangsu Province (KYCX24_2636).
Author information
Authors and Affiliations
Contributions
KZ, TX, NS, HY, and CY designed the experiments. KZ, TX, XYH, JZL, DSH, SHH, MQH, LP, YYW, and CY conducted the experiments. KZ, TX, JZL, HY, and CY performed data and statistical analyses. KZ, TX, NS, HY, and CY wrote the manuscript. KZ, TX, JZL, and CY revised the study and wrote the review & editing. NS, HY, and CY supervised the study. KZ, NS, HY, and CY provided funding.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhang, K., Xu, T., Heng, Xy. et al. Galangin 3-methyl ether alleviates mouse hypertrophic cardiomyopathy via targeting HDAC2 and subsequently inactivating the PI3K-AKT signaling pathway. Acta Pharmacol Sin 47, 636–651 (2026). https://doi.org/10.1038/s41401-025-01676-7
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41401-025-01676-7