Abstract
Esophageal adenocarcinoma (EAC), the dominant subtype of esophageal cancer in developed countries, is a growing health problem, characterized by poor patient prognosis and dismal survival due to ineffective screening tools and a lack of efficacious options targeting the interception or treatment of EAC. Despite molecular advances, molecular targeting of EAC remains elusive, suggesting the need for identifying alternative targets with improved prognostic and therapeutic value. Herein, we performed RNA-sequencing analysis in EAC and Barrett’s Esophagus (BE) precursor lesions to identify isoform switching events significantly linked with all-cause and cancer-specific mortality. Patients were stratified based on histopathology alone or in combination with TP53 mutation status, the most commonly mutated gene in EAC. To gain mechanistic insight, we performed isoform-specific siRNA knockdown of two isoforms, TTLL12 and HM13, both linked to patient survival, and investigated mechanisms associated with isoform dysregulation and whether targeting specific isoforms in EAC acts synergistically to improve therapeutic potential. Isoform-specific knockdown of TTLL12 and HM13 significantly decreased the viability of two EAC cell lines, sensitized EAC cell lines to standard-of-care chemotherapy agents (paclitaxel and carboplatin) with synergy, and inhibited EAC cell migratory potential. Knockdown of the TTLL12 isoform led to activation of chaperone-mediated autophagy, which, in turn, decreased expression of CHK1 and TP53; whereas knockdown of the HM13 isoform activated the unfolded protein response and induced endoplasmic reticulum stress-induced autophagy and apoptosis. In addition, HM13 isoform knockdown increased the response to an anti-PD-L1 agent, avelumab, in EAC cells, suggesting a role for isoform switching in immunosuppression. Taken together, study results suggest that isoform switching may provide novel insight for the identification of prognostic markers and inform new potential therapeutic targets for EAC treatment or prevention.
Similar content being viewed by others
Introduction
Esophageal adenocarcinoma (EAC), the dominant subtype of esophageal cancer in developed countries such as the United States (US), represents a growing health problem characterized by rising incidence since the 1970s and continued poor prognosis [1]. It is the 6th leading cause of cancer-related mortality worldwide and the 7th among males in the US, with a five-year survival rate of 22% [2, 3]. Ineffective screening tools and late-stage diagnosis, coupled with the lack of efficacious treatment regimens, contribute to the abysmal survival rate of EAC. The only known precursor lesion to EAC is Barrett’s Esophagus (BE), a premalignant condition where the lining of the esophagus changes from a squamous epithelium to a columnar epithelium. BE with metaplasia or BE with low-grade dysplasia (LGD) has a lower risk of progressing to EAC, whereas BE with high-grade dysplasia (HGD) is considered to be at high risk of progression [4]. Although endoscopic surveillance is recommended for patients with BE, only 7.3% of EAC patients have a prior BE diagnosis [5], reinforcing that better risk-prediction models are needed. For patients diagnosed with EAC, the most common standard-of-care treatment is neoadjuvant chemoradiotherapy, followed by esophagectomy for surgery-eligible patients. Concurrent prescription of paclitaxel and carboplatin is commonly utilized to treat EAC [6]. However, only 23.1% of EAC patients achieve a pathological complete response with this treatment regimen, and even these patients are at risk for recurrence [7]. EAC is a heterogeneous cancer dominated by high mutation rates, increased copy number alterations, and large-scale genomic alterations compared to other cancer types [8, 9]. Although it is a cancer with a high mutational burden, it lacks dominant mutations that can be clinically targeted. Over 80% of EAC patients carry a TP53 mutation, yet mutation frequencies of multiple other genes are all lower than 40% [10]. For these reasons, the identification of alternative targets in BE and EAC is urgently needed for improving patient outcomes.
We previously performed RNA sequencing (RNA-seq) of 57 esophageal tissue samples derived from treatment-naïve HGD and EAC patients undergoing esophagectomy and investigated isoform switching (IS) events linked with EAC progression [11]. IS is defined as the change of dominant gene isoforms between two conditions. We identified 75 genes that are isoform switched between BE with LGD (BE.LGD) and BE with HGD (BE.HGD). The inclusion of TP53 mutation status in the analysis further increased the number to 135. Importantly, over 40% of identified isoform-switched genes are not differentially expressed at the gene level [11]. Herein, we extended our investigation to assess whether IS events are significantly linked to patient prognosis based on cancer-specific and all-cause mortality, followed by mechanistic studies targeting two isoforms linked to patient survival (TTLL12 and HM13 isoforms). Results show that siRNA knockdown of TTLL12 and HM13 isoforms significantly sensitized EAC cells to standard-of-care chemotherapeutic agents while having minimal effect on normal esophageal epithelial cells (Het-1A). Mechanisms implicated include activation of chaperone-mediated autophagy (CMA) for the TTLL12 isoform and unfolded protein response (UPR) and immunosuppression for the HM13 isoform. Moving forward, specific IS events may inform new potential therapeutic targets for EAC treatment and be developed as prognostic markers.
Results
Isoform switching analysis reveals isoforms significantly linked with patient survival
To identify isoforms linked with EAC patient survival, IS analysis was performed utilizing patient samples derived from BE.LGD compared to BE.HGD + EAC, alone or in combination with TP53 mutation status. As shown in Fig. 1A, 71 genes were isoform-switched compared to BE.LGD and BE.HGD + EAC patient samples. With the inclusion of TP53 mutation status, a total of 67 genes were isoform-switched. The Venn diagram revealed that 40 isoform-switched genes were shared between the two comparisons (Figs. 1A and S1).
A Venn diagram showing common and unique isoform-switched genes between BE.LGD vs. BE.HGD + EAC (n = 71) and BE.LGD TP53 WT vs. BE.HGD + EAC TP53 MUT (n = 67). Top 20 most significant isoform-switched genes in B BE.LGD vs. BE.HGD + EAC and C BE.LGD TP53 WT vs. BE.HGD + EAC TP53 MUT. FDR, false discovery rate. Isoform fractions determined using D qRT-PCR and E RNA-seq data. Significant differences in isoform fractions were determined by one-tailed two-sample t-tests. Bar plots represent the mean isoform fraction, and error bars indicate the standard error of the mean.
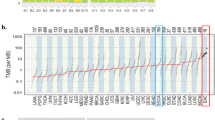
The top 20 most significant isoform-switched genes comparing BE.LGD and BE.HGD + EAC are shown in Fig. 1B. Eighty-five percent (17/20) of the top isoform-switched genes were protein-coding genes, including TPM4, KRAS, and TTLL12. Two RNA genes, MIR4458HG and LINC02826, and a pseudogene, H2BW4P, also showed significant IS. With the inclusion of TP53 mutation status, 85% (17/20) of the top isoform-switched genes remain protein-coding genes, including TPM4, HM13, and KRAS (Fig. 1C). Among the top 20 genes reported in the two comparisons, 70% (14/20) were shared. Isoform-specific primers were designed to validate specific IS events using qRT-PCR (Fig. 1D), as shown for ATP9B, HM13, KIF16B, KRAS, TRIM29, and TTLL12, with parallel results from the RNA-seq data (Fig. 1E).
Next, survival analysis was performed to identify IS significantly linked with patient survival based on individual isoform fraction (IF) determination. As shown in Fig. 2A, 42 isoforms are significantly linked with all-cause patient mortality comparing BE.LGD to BE.HGD + EAC. Isoforms with a positive hazard ratio represent isoforms with elevated IF in BE.HGD + EAC vs. BE.LGD groups that are associated with reduced patient survival, whereas isoforms with a negative hazard ratio are isoforms that when reduced, are correlated with worse patient survival. As shown in Fig. 2A, higher IF of SMIM6 isoform (ENST00000556126), C1orf54 isoform (ENST00000369098), CFDP1 isoform (ENST00000283882), TPM4 isoform (ENST00000653979), and TTLL12 isoform (ENST00000216129) are among the top isoforms with increased IF significantly correlated with poor patient survival. A similar number of isoforms (n = 39 isoforms) are significantly linked with patient survival when TP53 mutation status is included in the analysis (Fig. 2B). Isoforms including HM13 isoform (ENST00000340852), CFDP1 isoform (ENST00000283882), DANT2 isoform (ENST00000430756), and C1orf54 isoform (ENST00000369098) show higher IF in BE.HGD + EAC TP53 MUT group, which were also significantly linked with worse patient survival. Next, survival analysis was performed to identify IS significantly linked with cancer-specific patient mortality (Fig. 2C, D). Forty and twenty-six isoforms were significantly linked with patient survival in BE.LGD vs. BE.HGD + EAC and BE.LGD TP53 WT vs. BE.HGD + EAC TP53 MUT, respectively. Although the number of isoforms significantly linked with patient survival was similar between all-cause and cancer-specific mortalities in BE.LGD vs. BE.LGD + EAC, the number of isoforms significantly linked with cancer-specific mortality (n = 26 isoforms) was lower than the number of isoforms linked with all-cause mortality in BE.LGD TP53 WT vs. BE.HGD + EAC TP53 MUT (n = 39 isoforms). Venn diagrams further reveal survival-linked isoforms that are common and unique to each comparison group (Fig. S1).
Isoforms significantly linked with all-cause patient mortality in A BE.LGD vs. BE.HGD + EAC and B BE.LGD TP53 WT vs. BE.HGD + EAC TP53 MUT. Isoform significantly linked with cancer-specific mortality in C BE.LGD vs. BE.HGD + EAC and D BE.LGD TP53 WT vs. BE.HGD + EAC TP53 MUT. P-values were determined using the Cox proportional hazards model. Each bar represents a 95% confidence interval.
To gain mechanistic insight and investigate whether isoforms can be therapeutically targeted, two isoforms, TTLL12 (ENST00000216129) and HM13 (ENST00000340852), were chosen for the genetic knockdown studies in EAC cell lines. Both isoforms are the coding isoform of the gene and show significantly higher IF in BE.HGD + EAC group and significantly linked with reduced patient survival. Specifically, the TTLL12 isoform was among the top isoforms identified to be associated with all-cause patient mortality in BE.LGD vs BE.HGD + EAC group, whereas the HM13 isoform was among the top isoforms identified in BE.LGD TP53 WT vs. BE.HGD + EAC TP53 MUT (both all-cause and cancer-specific mortality). In addition, IS events of both genes were validated using qRT-PCR, as shown in Fig. 1D.
To further validate the potential oncogenic role of TTLL12-201 and HM13-201 in EAC, we performed RNA-seq analysis of esophageal samples collected from rats with reflux-driven EAC, as we previously published [12]. A significant upregulation of Ttll12-201 was observed in rats with reflux-driven EAC (Fig. S2A), compared to water controls. Similarly significant upregulation of the HM13 ortholog, Mcts2, was also observed, although the exact ortholog of HM13-201 in rats is unknown (Fig. S2B). We also performed RNA-seq analysis of EAC samples from a human cohort in the Cancer Genome Atlas Program (TCGA) [13], revealing significant upregulation in TTLL12-201 (Fig. S2C) and HM13-201 in EAC samples, compared to normal tissues (Fig. S2D).
Isoform-specific knockdown of TTLL12 inhibited EAC cell viability and migration via chaperone-mediated autophagy
IS of TTLL12 was observed between two of its isoforms, TTLL12-201 (ENST00000216129) and TTLL12-204 (ENST00000494035) (Fig. 3A). TTLL12-201 is the coding isoform of TTLL12, containing several tubulin tyrosine ligase (TTL) domains, whereas TTLL12-204 is a predicted non-coding isoform (Fig. 3A). TTLL12-201 has a significantly higher IF in BE.HGD + EAC group and TTLL12-204 showed a decreased IF in this group, supporting a significant IS (Fig. 3B). As shown in Figs. 3C and S3B, a higher IF of TTLL12-201 significantly decreases overall patient survival (44.4% survival probability versus 72.2%), linking TTLL12-201 to overall survival among EAC patients for the first time. In addition, gene-level expression of TTLL12 is significantly lower in BE.HGD + EAC compared to BE.LGD (Fig. S3A).
A Isoform switching of TTLL12. B Isoform fractions of TTLL12 comparing BE.LGD to BE.LGD + EAC. C Kaplan–Meier survival curve indicating significant survival differences based on the TTLL12-201 isoform fraction. D Immunoblot showing knockdown of TTLL12-201 in OE19 EAC cells using siRNA. E Immunoblot showing knockdown of TTLL12-201 in OE33 EAC cells using siRNA. F Viability results and representative fluorescent images (72 h) of OE19 cells treated with siRNA alone or in combination with chemotherapy agents (n = 4–6/group). G Migration assay results and representative microscopic images of OE19 cells treated with siRNA alone or in combination with chemotherapy agents (n = 10–20/group). H Viability results and representative fluorescent images (72 h) of OE33 cells treated with siRNA alone or in combination with chemotherapy agents (n = 4–6/group). I Protein interaction prediction network of TTLL12 determined by STRING analysis. J Biological processes result from enrichment analysis of TTLL12. Error bars represent the standard error of the mean. Significance was determined using ANOVA with Tukey’s post-hoc test for multiple comparisons between treatments. Within each time point, treatments were significantly different from a = NTC-VEH, b = siRNA#1–VEH, c = siRNA#2–VEH, d = NTC–Chemo, e = siRNA#1–Chemo, f = siRNA#2–Chemo. Pound sign (#) denotes synergy. Synergy was calculated using the Combination Index equation. IF isoform fraction, NTC non-targeting control, Scale bar, 200 μm; Chemo, combination of paclitaxel and carboplatin.
To investigate the therapeutic potential of targeting TTLL12-201, two EAC cell lines, OE19 and OE33, were treated with two isoform-specific siRNAs targeting TTLL12-201. As shown in Figs. 3D, E, and S4, isoform-specific knockdown of TTLL12-201 led to a 50–70% reduction in TTLL12-201 isoform and protein expression levels across cell lines over time, supporting successful knockdown using both siRNAs. Results of the viability assay in OE19 cells (Fig. 3F) reveal that treatment with either of the TTLL12-201 siRNAs led to significant inhibition of OE19 cell viability at 48 h (69–93%) and 72 h (65–87%), respectively. Additionally, synergy, reported as the combination index, was observed for OE19 cells treated with both siRNA#1 and chemotherapy agents at 24 h and 72 h. A migration assay performed in OE19 cells (Fig. 3G) reveals that siRNA knockdown alone (siRNA#1: 190 ± 73.3 μm and siRNA#2: 242 ± 35.4 μm) or in combination with chemotherapy agents (siRNA#1+Chemo: 333 ± 41.8 μm and siRNA#2+Chemo: 379 ± 28.5 μm) significantly inhibits cell migration compared to cells treated with non-targeting control (NTC) (107 ± 41.4 μm) or chemotherapy agents alone (283 ± 21.9 μm) at 72 h. Similar but more substantial effects were also observed in OE33 cells. As shown in Fig. 3H, TTLL12-201 isoform knockdown led to significant inhibition of OE33 cell viability at all three time points. At 72 h, siRNA treatment alone reduced OE33 cell viability to 34–74%. Synergistic effects were observed at all three time points for combination treatments of individual siRNAs and chemotherapy agents (Fig. 3H). At 72 h, the combination treatment decreased OE33 cell viability to 12–22%. TTLL12-201 knockdown also led to significant inhibition of OE33 cell migration (Fig. S5). The results indicate that TTLL12-201 knockdown sensitized EAC cells to chemotherapeutic agent-induced cell death.
Isoform-specific TTLL12 knockdown induces chaperone-mediated autophagy (CMA)
To investigate potential mechanisms linked with TTLL12 isoform knockdown, a STRING protein interaction prediction was performed (Fig. 3I, J). Biological process analysis of the TTLL12 protein interaction network reveals significant biological processes linked with protein polyglutamylation, regulation of blastocyst development, and regulation of CMA (Fig. 3J). Considering altered CMA is reported in esophageal squamous cell carcinoma (ESCC) [14], immunoblotting of two chaperone-mediated autophagy markers, LAMP2A and HSC70, was performed. Results show upregulation of LAMP2A and HSC70 upon TTLL12-201 knockdown in both OE19 and OE33 cells (Fig. 4A, B), with more substantial induction in OE19 cells. TTLL12 isoform knockdown also resulted in potent induction of autophagy-linked LC3-II in OE33 cells and minor LC3-II induction in OE19 cells, where LC3-I has low baseline expression levels, consistent with previous reports of differential capacity for autophagy induction in EAC cells [15]. Immunoblotting of CHK1, phospho-CHK1 (Ser345), and TP53 was also performed (Fig. 4C, D) because CMA targets both CHK1 and missense mutant TP53 for degradation [16, 17]. Results showed decreased expression of CHK1 in both OE19 and OE33 cells. Moreover, decreased expression of phospho-CHK1 and TP53 was observed in OE33 cells but less evident in OE19 cells, as they carry a frameshift mutation resulting in expression of only the truncated form of TP53 (45 kD).
Immunoblot of CMA markers, LAMP2A, and HSC70, in A OE19 and B OE33 EAC cell lines. Immunoblot of CHK1 and TP53 in C OE19 and D OE33 cells following TTLL12-201 siRNA knockdown alone or in combination with chemotherapy agent treatment. E Viability results and representative fluorescent images (48 h) of OE19 cells treated with APR-246. Immunoblot of CMA markers, CHK1, and TP53 in F OE19 and G OE33 cells following APR-246 treatment. NTC non-targeting control, Scale bar, 200 μm; ***, P < 0.001.
To further validate that targeting mutant TP53 leads to EAC cell death, OE19 cells were treated with APR-246, a pharmacological agent that targets mutant TP53 and induces autophagy in cancer cells [18]. As shown in Fig. 4E, treatment of APR-246 led to a significant decrease in cell viability at 24 h and 48 h in OE19 cells, and, as we previously reported, a significant decrease in cell viability in OE33 cells following APR-246 treatment [11]. Immunoblots of LAMP2A and HSC70 suggest that APR-246 treatment increased LAMP2A expression in OE19 cells and HSC70 expression in OE33 cells, suggesting activation of CMA (Fig. 4F, G). Similar to TTLL12-201 siRNA-induced protein modulation, decreased expression of TP53 and increased expression of autophagic LC3-II were observed in OE33 cells (Fig. 4G). Although modulation of CHK1 and/or phospho-CHK1 was not observed in either EAC cell line at 24 h post-treatment, inhibition was observed at 48 h post-treatment, with more substantial changes noted in OE33 cells (Fig. S6).
Isoform-specific knockdown of HM13 inhibited EAC cell viability and migration through the unfolded protein response pathway
HM13 IS was observed between BE.HGD + EAC and BE.LGD. Specifically, IS of HM13 was observed between HM13-201 (ENST00000340852) and HM13-220 (ENST00000493364) (Fig. 5A). HM13-201 is one of the coding isoforms of HM13, containing several signal peptide peptidase domains, whereas HM13-220 is one of the non-coding isoforms of HM13 (Fig. 5A). Similar to TTLL12-201, the HM13-201 isoform is significantly elevated in BE.HGD + EAC patient samples compared to HM13-220, which shows a significantly reduced IF (Fig. 5B). Higher levels of HM13-201 IF are significantly linked with reduced patient survival (40.0% survival probability vs. 90.0%, Figs. 2B, 5C, and S7B), supporting its role in EAC. Overall, gene-level expression of HM13 is also significantly lower in BE.HGD + EAC compared to BE.LGD (Fig. S7A).
A Isoform switching of HM13. B Isoform fractions of HM13 comparing BE.LGD to BE.LGD + EAC. C Kaplan-Meier survival curve indicating significant survival differences based on the HM13-201 isoform fraction. D Immunoblot showing knockdown of HM13-201 in OE19 cells using siRNA. E Immunoblot showing knockdown of HM13-201 in OE33 cells using siRNA. F Viability results and representative fluorescent images (72 h) of OE19 cells treated with siRNA alone or in combination with chemotherapy agents (n = 4-6/group). G Migration assay results and representative microscopic images of OE19 cells treated with siRNA alone or in combination with chemotherapy agents (n = 10–36/group). H Viability results and representative fluorescent images (72 h) of OE33 cells treated with siRNA alone or in combination with chemotherapy agents (n = 4-6/group). I Protein interaction prediction network of HM13 determined by STRING analysis. J Biological processes result from enrichment analysis of HM13. Error bars represent the standard error of the mean. Significance was determined using ANOVA with Tukey’s post-hoc test for multiple comparisons between treatments. Within each time point, treatments were significantly different from a = NTC-VEH, b = siRNA#1–VEH, c = siRNA#2–VEH, d = NTC–Chemo, e = siRNA#1–Chemo, f = siRNA#2–Chemo. Pound sign (#) denotes synergy. Synergy was calculated using the combination index equation. IF isoform fraction, NTC non-targeting control, Scale bar, 200 μm; Chemo, combination of paclitaxel and carboplatin.
OE19 and OE33 cells were similarly treated with isoform-specific siRNAs targeting HM13-201, and immunoblots confirmed the successful knockdown in both cell lines at 48 h and 72 h (Fig. 5D, E). As shown in Fig. 5F, HM13-201 knockdown significantly decreases OE19 cell viability at 24 h, 48 h, and 72 h. At 72 h, siRNA treatment alone reduced OE19 cell viability to 77–85%. Moreover, synergistic effects were observed for siRNA#1 at 48 h and 72 h and for siRNA#2 at 24–72 h. At 72 h, the combination treatment of siRNA#1 and chemo drugs reduced OE19 cell viability to 68%, and the combination treatment of siRNA#2 and chemo drugs decreased cell viability to 59%, supporting that HM13 isoform knockdown sensitizes OE19 cells to chemotherapy agents. Results of migration assays evaluating OE19 cells upon siRNA treatment, shown in Fig. 5G, reveal that HM13 isoform knockdown significantly inhibits OE19 cell migration compared to cells treated with NTC or chemotherapy agents alone. Similar results in cell viability were also observed in OE33 cells (Fig. 5H). HM13-201 knockdown significantly inhibited OE33 cell viability, with synergistic effects observed with both siRNA#1 and siRNA#2 treatments combined with chemotherapy agents. At 72 h, siRNA#1 combined with chemo drugs decreased OE33 cell viability to 13%, and siRNA#2 combined with chemo drugs decreased cell viability to 14%. Similar to results in OE19 cells, HM13-201 knockdown also significantly inhibited the migration capability of OE33 cells (Fig. S8).
Isoform-specific knockdown of HM13 activates unfolded protein response (UPR) and halts protein translation
A STRING protein interaction prediction reveals interacting proteins of HM13 and related biological processes (Fig. 5I, J), identifying multiple pathways associated with the endoplasmic reticulum (ER)-associated protein degradation (ERAD) pathway and ER UPR pathway. The UPR consists of three main branches: PERK, IRE-1α, and ATF-6 [19]. Immunoblotting of key UPR-related proteins was performed in both OE19 (Fig. 6A) and OE33 cells (Fig. 6B). In OE19 cells, increased expression of BIP, phosphorylation of PERK (as indicated by an upshift of the PERK band), and activation of downstream target proteins (GADD34, ATF-4, and phospho-eIF2α) were observed upon HM13-201 knockdown alone and in combination with chemotherapy agents, compared to NTC or chemotherapy treatment alone. Similar induction of IRE-1α and its downstream targets XBP1 and phospho-JNK, and ATF-6 was also observed. The UPR is also documented to induce autophagy and apoptosis [19]. Expression of autophagy markers (PINK1, NDP52, and LC3-II) and the apoptosis marker BAK and cleaved PARP were increased upon HM13-201 knockdown alone and in combination with chemotherapy agents, compared to NTC, with modest reductions with chemotherapy treatment alone. In contrast, in OE33 cells, the PERK branch was not activated. Expression of IRE-1α was increased upon HM13 knockdown, but not its downstream factor XBP1, and ATF-6 was increased upon HM13 knockdown. HM13 knockdown in OE33 cells also resulted in increased expression of the autophagy-related markers, PINK1, NDP52, and LC3-II, and the apoptosis marker, cleaved PARP. Interestingly, OE33 cells did not express BIP, phospho-eIF2A, or phospho-JNK in contrast to OE19 cells (data not shown). In addition, because studies of HM13 in other cancers suggest an association with an immunosuppressive microenvironment and PD-1 expression [20, 21], an antibody-dependent cell-mediated cytotoxicity (ADCC) assay was performed to evaluate whether HM13-201 knockdown increases response to the anti-PD-L1 agent, avelumab, in EAC cells. A significantly increased response to avelumab is observed in OE33 cells upon HM13 knockdown, compared to cells treated with NTC followed by avelumab (Fig. 6C). However, such a response was not observed in OE19 cells. Since the UPR also leads to translation arrest [19], SUnSET assays were performed to measure changes in protein translation. Both OE19 (Fig. 6D) and OE33 (Fig. 6E) cells show decreased protein translation upon HM13-201 knockdown. A stronger magnitude of effect was evident in cells treated with HM13 siRNA combined with chemotherapy agents.
Immunoblot of key UPR proteins in A OE19 and B OE33 cells. C HM13-201 knockdown increases response to avelumab in OE33 cells (n = 3/group). Error bars represent the standard error of the mean. SUnSET assay of D OE19 and E OE33 cells showing inhibition of protein translation. NTC non-targeting control, ND not detected, CL cleaved protein, ns non-significant, *, P < 0.05; ***, P < 0.001.
To further validate the causality that HM13-201 knockdown led to activation of UPR followed by EAC cell death induction, OE19 and OE33 cells were treated with pharmacological agents, thapsigarin and tunicamycin, that induce UPR [22, 23]. As shown in Fig. 7A, both thapsigarin and tunicamycin treatment significantly decreases the viability of OE19 cells at 24 h and 48 h. At 48 h, 2 μM thapsigarin treatment decreased OE19 cell viability to 24.16%, and 500 ng/mL tunicamycin treatment decreased OE19 cell viability to 41.86%. Similarly, thapsigarin and tunicamycin potently inhibited OE33 cell viability in a time and dose-dependent manner (Fig. 7B). At 48 h, 2 μM thapsigarin treatment decreased OE33 cell viability to 35.30%, and 500 ng/mL tunicamycin treatment decreased OE33 cell viability to 19.16%. Immunoblotting was also performed on key UPR proteins and their downstream factors. Pharmacological-agent-induced UPR shows a similar trend compared to UPR induced by HM13-201 siRNA treatment in both OE19 and OE33 cells. In OE19 cells (Fig. 7C), thapsigarin or tunicamycin treatment increases the expression of BIP, phosphorylation of PERK (based upon altered migration), and expression of downstream factors of the PERK branch (GADD34 and phospho-eIF1α). In addition, increased expression of IRE-1α and its downstream factors (XBP1 and phospho-JNK) was observed, as well as increased expression of activated or cleaved ATF-6 (50kD). Lastly, thapsigarin or tunicamycin treatment also led to increased expression of LC3-II and cleaved PARP, suggesting activation of programmed cell death and autophagy following agent treatment. In OE33 cells (Fig. 7D), thapsigarin treatment led to phosphorylation of PERK, as did high concentrations of tunicamycin. Similar to HM13-201 siRNA-induced UPR, increased expression of IRE-1α and its downstream factor XBP1 was observed, as well as induction of ATF-6, LC3-II, and cleaved PARP in OE33 cells following thapsigarin or tunicamycin treatment. Taken together, the results suggest that UPR, induced by either HM13-201 knockdown or pharmacological agents, can lead to cell death in EAC cells via apoptosis and autophagy.
Viability results and representative fluorescent images (48 h) of A OE19 and B OE33 cells treated with thapsigarin and tunicamycin. Error bars represent the standard error of the mean. Immunoblot of key UPR proteins and their downstream factors in C OE19 and D OE33 cells, following thapsigarin or tunicamycin treatment. Significance was determined using ANOVA with Tukey’s post-hoc test for multiple comparisons between treatments. Within each time point, treatments were significantly different from a = Vehicle, b = 1 μM thapsigarin, c = 2 μM thapsigarin, d = 50 ng/mL tunicamycin, e = 250 ng/mL tunicamycin, f = 500 ng/mL tunicamycin. CL cleaved protein. Scale bar, 500 μm.
Isoform-specific knockdown of TTLL12 and HM13 had minimal effect on normal esophageal epithelial cells
To investigate whether isoform-specific knockdown of TTLL12 and HM13 has cytotoxic effects in normal esophageal epithelial cells, Het-1A cells were treated with isoform-specific TTLL12 and HM13 siRNAs. Results from cell viability assays indicated that knockdown of TTLL12-201 led to a 2–4% decrease in Het-1A cell viability and knockdown of the HM13-201 led to a 3–7% decrease in Het-1A cell viability (Fig. S9), suggesting minimal cytotoxicity associated with targeting these isoforms in normal cells.
Discussion
This research sought to investigate IS events linked to EAC patient survival, reveal mechanisms associated with dysregulation of specific isoforms, and study whether isoforms may be targeted for improved therapeutic response in EAC. Specifically, IS analysis was performed using RNA-seq data from EAC and Barrett’s Esophagus (BE) precursor lesions to identify IS events during pathological progression that are significantly linked with patient survival, and in vitro studies were conducted to explore mechanisms implicated in IS in the context of EAC. We identified 42 and 40 IS significantly linked to all-cause and cancer-specific mortality, respectively. A number of isoforms involved in metabolic reprogramming and drug resistance were uniquely linked to cancer-specific mortality, including KRAS and SLC28A3 [24, 25]. Given the role of TP53 mutation in EAC progression, we further stratified results based on TP53 mutation status, identifying 39 and 26 isoforms associated with all-cause and cancer-specific mortality, respectively (Fig. 2).
Among the survival-linked isoforms we identified in this cohort, IS of TPM4, TTLL12, COL4A2, IGSF3, FSIP1, RPL22L1, and KRAS have been previously reported in other cancer tissues or cell lines [26,27,28,29,30,31,32,33]. In EAC, RNF128 is the only isoform-switched gene that has been broadly studied [34]. Herein, we selected two isoforms identified based on survival analysis, TTLL12-201 and HM13-201, and performed isoform-specific siRNA knockdown followed by a series of in vitro studies. The results showed that knockdown of TTLL12 and HM13 isoforms significantly inhibited the viability of two EAC cell lines, sensitized EAC cell lines to standard-of-care chemotherapy agents, and inhibited cell migratory potential. Importantly, the knockdown of both induced isoforms had minimal effects on a normal esophageal cell line (Het-1A), suggesting both isoforms may be therapeutically targeted in EAC with minimal side effects.
TTLL12, also known as tubulin tyrosine ligase 12, is one of the most understudied proteins in the TTL/TTLL family. Overexpression of gene-level TTLL12 is documented in various cancers, including gastric adenocarcinoma, lung adenocarcinoma, colorectal cancer (CRC), and prostate cancer [27, 35,36,37,38]. IS of TTLL12 is also observed in ESCC cell lines compared to normal esophageal Het-1A cells [27]. In CRC, an alternate promoter leads to IS of TTLL12 compared to normal tissues. In our study, we observe significantly higher isoform usage of a coding transcript of TTLL12 (TTLL12-201) in HGD and EAC compared to LGD tissues (Fig. 3). Isoform-specific knockdown of this isoform significantly decreased the viability and cell migration potential of two EAC cell lines. Additionally, isoform-specific knockdown sensitized both cells to standard-of-care chemotherapy agents. STRING analysis and immunoblot support that TTLL12-201 is linked with CMA. TTLL12 is speculated to be a negative regulator in inflammation-driven nitrosative stress, which inhibits nitrotyrosine-induced cytotoxicity in malignant cells. In non-cancerous cells, nitrotyrosine can induce cell death through multiple mechanisms, including apoptosis and autophagy. However, in cancer cells, TTLL12 decreases the cytotoxicity of nitrotyrosine, potentially by inhibiting nitrotyrosination of α-tubulin [39,40,41]. EAC is also an inflammation-driven cancer, and nitrosative stress is reportedly linked with EAC progression [42,43,44], raising the possibility that upregulation of the coding transcript of TTLL12 in HGD and EAC suppresses cell death. The role of CMA in EAC remains underexplored. In other cancers, CMA exerts both anti-tumor and pro-tumor activities [45]. The anti-tumor effect of CMA is largely achieved by targeted degradation of cancer promoters, genome quality control, and influencing glucose metabolism [45]. For example, activated CMA is known to target missense mutant TP53 for degradation and inhibit CHK1 to prevent DNA damage response [16, 17]. In EAC, over 80% of patients carry a TP53 mutation, and a CHK1/2 inhibitor, AZD7762, potently inhibits viability of an EAC cell line [8, 10]. EAC cell lines used in our study, OE19 and OE33, carry a frameshift and a missense TP53 mutation, respectively [46, 47]. The reason why activation of CMA fails to degrade TP53 in OE19 is likely because OE19 carries a frameshift TP53 mutation. We also treated both EAC cells with a pharmacological agent that targets mutant TP53 and induces autophagy, which revealed similar trends in loss of cell viability and protein modulation as observed in EAC cells treated with TTLL12-201 siRNA. Our results align with published studies reporting that activation of CMA leads to CHK1 inhibition and degradation of missense mutant TP53, further supporting the therapeutic potential of targeting TTLL12-201 in EAC.
HM13, also known as histocompatibility minor 13, is linked with poor prognosis and patient survival, metastasis, and cancer proliferation and migration in hepatocellular carcinoma, breast cancer, and CRC [20, 21, 48,49,50]. As an ER-resident protein, HM13 plays a role in the UPR [51]. ER stress induces activation of the UPR, which either leads to recovery of ER function or ER-stress-induced cell death if cells fail to overcome ER stress [19]. In cancer, two distinct routes have been proposed to target the UPR: to trigger UPR-mediated cell death or block ER recovery mechanisms [52]. In EAC, whether the UPR could be therapeutically targeted remains poorly understood. One study shows that EAC cell lines have higher expression levels of BIP, PERK, IRE-1α, and XBP1, as compared to normal esophageal cells, suggesting a vulnerability [53]. However, the latter study did not measure expression levels of downstream factors of UPR. Herein, we measured expression levels of downstream factors of UPR, including GADD34, phospho-JNK, PINK1, NDP52, LC3, PARP, and BAK, to determine whether isoform-specific knockdown of HM13-201 can trigger UPR-mediated cell death in EAC cell lines. In OE19 cells, elevated expression of phospho-JNK and BAK (UPR-mediated apoptosis markers [19]), and cleaved PARP was observed, as well as increased expression of LC3-II, PINK1, and NDP52, implying activation of mitophagy in OE19 cells. Mitophagy can trigger cell death, both dependent (via BAK) and independent of apoptosis [54]. However, additional studies are required to delineate the specific crosstalk between UPR and mitophagy. In contrast, multiple UPR makers were not expressed in OE33 cells, likely due to the molecular heterogeneity of EAC. However, activation of mitophagy and inhibition of global translation were observed in both EAC cell lines, suggesting that alternative UPR pathways likely exist in OE33 cells. We also treated both EAC cells with pharmacological agents, thapsigarin and tunicamycin, that induce UPR. The results further support our observation that activation of UPR can lead to EAC cell death. Moreover, one study in hepatocellular carcinoma showed that HM13 is a predictive biomarker for immunotherapy [20]. The ADCC assay result in our study suggests that isoform-specific HM13 knockdown increased the response to avelumab, an anti-PD-L1 agent, in OE33 cells, pointing to a potential linkage between HM13-201 and response to immunotherapy in EAC. This finding is of potential importance considering EAC has a low response rate to PD-1/PD-L1 immune checkpoint inhibitors [55]. OE19 has a significantly lower PD-L1 expression compared to OE33 and did not respond to avelumab treatment [56].
This research provides new insight regarding genes that are isoform switched with pathological progression to EAC and are associated with patient mortality, suggesting that specific isoforms may be targeted for enhanced therapeutic or prognostic value. Overall, genetic knockdown of two of the top mortality-linked isoforms (HM13 and TTLL12 isoforms) revealed that isoforms may be therapeutically targeted to sensitize EAC to chemotherapy agents or immunotherapy with minimal cytotoxicity in normal esophageal cells. Generalizability of the current study is unknown because, despite our comprehensive efforts to search the public and permission-based databases for external validation datasets, we could not identify any parallel datasets containing sufficient BE patient samples with stratification by grade of dysplasia and TP53 mutation status among EAC progressors. Additionally, most of the datasets we identified lacked the depth of sequencing required to perform isoform switching analysis and were further hindered by small sample sizes in subgroups (i.e., BE with dysplasia). Future studies will evaluate the potential of using small molecules or splice-switching antisense oligonucleotides to target isoforms in EAC [57, 58]. In addition, we plan to further identify specific IS driver events associated with EAC progression using multi-omics approaches, such as long-read RNA-seq, proteomics, and functional validation. Research in a larger cohort of EAC progressors with multiple biopsies acquired from patients over time, or comparing EAC progressors to non-progressors would be informative.
Material and methods
Isoform switching analysis of patient RNA-seq data
IS analysis was performed on patient RNA-seq data derived from esophageal tissues, as we previously reported [11]. Kallisto (version 0.46.2) was used to quantify transcripts in the transcriptomics data [59]. IsoformSwitchAnalyzeR (version 1.14.1) was applied in R (version 4.1.1) to characterize IS events [60, 61]. Briefly, IS events were determined by calculating isoform fractions (IFs) and differential isoform fractions (dIFs) between BE.LGD esophageal samples (n = 17) and BE.HGD + EAC esophageal samples (n = 28) alone or in combination with TP53 mutation status. IFs were calculated by dividing specific isoform expression by total gene expression for a given pathology, and dIFs were determined by calculating the difference of IFs between BE.LGD and BE.HGD + EAC alone or in combination with TP53 mutation status. Significant isoform switching events were determined using an absolute dIF cutoff of 0.1 and an FDR cutoff at 0.05, as determined by the Mann–Whitney U-test [60]. In addition, several external analyses were performed to identify significant isoform switching events with predicted functional consequences, including prediction of coding potential using CPAT, signal peptides using SignalP, intrinsically disorder regions using IUPred2A, coding domains using Pfam, and sensitivity to nonsense-mediated decay [62,63,64,65]. The Venn diagrams in Figs. 1A and S1 were generated using InteractiVenn, and dot plots summarizing isoform switching analysis results in Fig. 1B, C were generated using ggplot2 in R [66, 67]. Individual isoform switching plots (Figs. 3A, B and 5A, B) were generated from the IsoformSwitchAnalyzeR package. Protein interaction prediction (Figs. 3I and 5I) and Biological Process enrichment analysis (Figs. 3J and 5J) were performed using the STRING database [68].
Quantification of isoform expression using qRT-PCR
Quantification of isoform expression using qRT-PCR was performed using RNA extracted from the same patient cohort, as previously described [11]. Isoform-specific primers of ATP9B, HM13, KIF16B, TRIM29, and TTLL12 were designed using the NCBI Primer-BLAST [69]. Isoform-specific primer sequences of TRIM29 were previously published, and primer sequences of ATP9B, HM13, KIF16B, and TTLL12 isoforms or genes are reported in Table S1 [11]. Primers for qRT-PCR were synthesized by Eurofins Scientific (Louisville, KY, USA). qRT-PCR was performed using SsoAdvanced Universal SYBR Green Supermix with CFX Connect Thermo Cycler (Bio-Rad Laboratories, Hercules, CA, USA). Relative isoform expression was determined using GAPDH as the housekeeping gene, and IF was calculated similarly to the IsoformSwitchAnalyzeR package.
Patient survival analysis
To identify isoforms significantly linked with patient survival, IF of isoform-switched genes was calculated for each patient using TPM values, and patients were stratified based on IF into tertiles. Survival analysis was performed by comparing patients sorted into the high tertile isoform usage group to the low tertile isoform usage group using the Cox proportional hazards model (survival package, version 3.5-5) in R [70]. Cancer-specific mortality was determined by only including patients whose cause of death was related to esophageal cancer, with all other patients censored or not counted as endpoints, but removed from the at-risk group in the same manner that patients who are lost to follow-up are removed [71]. Selection bias was minimized by using an unbiased, genome-wide approach to identify IS events using isoformSwitchAnalyzeR, and survival analysis was performed for all isoforms that are significantly switched.
Cell culture
OE19 and OE33 EAC cell lines were cultured in RPMI 1640 medium with 2.0 mM L-glutamine, 104 units/mL penicillin, 104 μg/mL streptomycin, 1 mM sodium pyruvate, and 10% fetal bovine serum (FBS), as previously reported [72]. Cell culture reagents were acquired from ThermoFisher Scientific (Waltham, MA, USA). The cell lines were maintained as monolayers and incubated at 37 °C with 5% CO2. Both cell lines were acquired from the European Collection of Authenticated Cell Cultures (ECACC, Wiltshire, UK) and confirmed to be authentic and mycoplasma-free. Het-1A, a normal esophageal cell line, was cultured in BEGM BulletKit medium without GA-1000 (Lonza Bioscience, Walkersville, MD, USA) and were cultured on T-75 tissue culture flasks pre-coated with 0.01 mg/mL fibronectin, 0.03 mg/mL bovine collagen type I, and 0.01 mg/mL bovine serum albumin (all from ThermoFisher Scientific) dissolved in the medium. Het-1A was acquired from the American Type Culture Collection (ATCC, Manassas, VA, USA) and confirmed to be authentic mycoplasma-free.
siRNA knockdown of TTLL12 and HM13 isoforms and in vitro assays
TTLL12 and HM13 specific isoforms were selected for knockdown studies because both are among the top isoforms significantly linked with patient mortality. In addition, IS of both isoforms was validated via qRT-PCR, and it is feasible to design isoform-specific siRNA for these isoforms. Isoform-specific siRNAs of TTLL12 were synthesized by MilliporeSigma (Burlington, MA, USA) using previously published sequences, and pre-designed isoform-specific siRNAs of HM13 (SASI_Hs01_00100621 and SASI_Hs01_00100625) were acquired from MilliporeSigma [41]. Specificity of siRNA used in this study was confirmed using the NCBI BLAST [73]. Twenty-four hours prior to transfection, OE19 (140 000 cells/well), OE33 (4 000 cells/well), and Het-1A (200 000 cells/well) cells were seeded in black-walled, clear-bottom 96-well plates. On the day of transfection, Lipofectamine RNAiMAX Reagent (ThermoFisher Scientific) prepared in Opti-MEM medium was combined with 100 nM of siRNA prepared in Opti-MEM medium (ThermoFisher Scientific), followed by 20 minutes of incubation. Then, the siRNA-lipid complex was added to both cell lines and incubated for 72 h. Next, OE19 and OE33 cells were treated with either combination chemo drugs (1 μM paclitaxel and 5 μM carboplatin) or vehicle control (0.01% water and 0.1% DMSO; n = 6/condition). Both treatments and vehicle were prepared in RPMI 1640 medium with 2.0 mM L-glutamine, 104 units/mL penicillin, 104 μg/mL streptomycin, 1 mM sodium pyruvate, and 5% FBS. OE19 and OE33 EAC cells were stained using Calcein-AM (ThermoFisher Scientific) for in vitro viability measures at 24, 48, and 72 h post-treatment, as previously reported [15]. Het-1A normal esophageal cells were stained using Calcein-AM at 48 and 72 h post-siRNA-knockdown. Fluorescent images and readings were acquired using the SpectraMax MiniMax Imaging Cytometer (Molecular Devices, San Jose, CA, USA). Synergy was determined using CompuSyn (http://www.combosyn.com/).
In vitro migration assay of OE19 and OE33 EAC cells treated with siRNA
To assess alteration in migration, OE19 and OE33 cells were seeded in T-25 tissue culture flasks at 3E6 cells/flask and 1.5E6 cells/flask, respectively. Twenty-four hours after seeding, both cell lines were similarly transfected with siRNA and incubated for 48 h. Then, cells were trypsinized with TrypLE Express (ThermoFisher Scientific) and seeded in 12-well plates at 1.5E6 cells/well and 1.3E6 cells/well, respectively. Following an overnight incubation, transfected cells were similarly treated with either combination chemo drugs or vehicle as described above. At 24 h post-treatment, cells reached about 90–100% confluency, and a vertical scratch was created in each well using a P-200 pipette tip. Microscope images were taken immediately after creating the scratch (n = 7 images per well). Scratch closure was measured every 24 h for up to 96 h with microscopic images of the same fields initially scratched measured. Distance of the scratch was quantified using Fiji software with the wound healing size tool [74, 75].
In vitro antibody-dependent cell-mediated cytotoxicity (ADCC) assay
To measure whether isoform knockdown increases response to immunotherapy agents in EAC cells, the ADCC assay was performed using the ADCC Reporter Bioassay (Promega, Madison, WI, USA) following the manufacturer’s instructions. OE19 and OE33 cells were seeded in T-25 tissue culture flasks at 3E6 cells/flask and 1.5E6 cells/flask, respectively, and transfected with siRNA (NTC or pooled HM13 siRNA) and incubated for 72 h. Then, OE19 and OE33 cells were trypsinized and seeded in white flat-bottom 96-well plates at 25 000 cells/well and 12 500 cells/well, respectively. Following an overnight incubation, OE19 and OE33 cells were treated with either avelumab, an anti-PD-L1 agent (MedChemExpress, Monmouth Junction, NJ, USA), or Human IgG1 Isotype Control (ThermoFisher Scientific) at 2, 4, and 8 μg/mL (n = 3/treatment). Treatment concentrations of avelumab were determined based on previously published studies [76, 77]. Immediately following treatment, ADCC Bioassay Effector Cells were added to each well at 75 000 cells/well and incubated for 6 h. The effector-to-target cell (E:T) ratio was 3:1 and 6:1 for OE19 and OE33 cells, respectively. Bio-Glo Luciferase Assay Reagent was then added to each well, and luminescence was measured using the SpectraMax MiniMax Imaging Cytometer (Molecular Devices). Fold change of induction was calculated by dividing the Relative Light Unit (RLU) from each treated well with RLU from non-antibody control.
In vitro viability assay of OE19 and OE33 cells treated with thapsigarin, tunicamycin, and APR-246
OE19 and OE33 cells were seeded in black-walled, clear-bottom 96-well plates at 150 000 cells/well and 8 000 cells/well, respectively. Followed by overnight incubation, both cells were treated with thapsigarin (1 μM and 2 μM; MilliporeSigma), tunicamycin (50 ng/mL, 250 ng/mL, and 500 ng/mL; MilliporeSigma), APR-246 (30 μM and 60 μM; Cayman Chemical, Ann Arbor, MI, USA), or Vehicle (0.1% DMSO; n = 6/condition). Both treatments and vehicle were prepared in RPMI 1640 medium with 2.0 mM L-glutamine, 104 units/mL penicillin, 104 μg/mL streptomycin, 1 mM sodium pyruvate, and 5% FBS. Concentrations of thapsigarin, tunicamycin, and APR-246 treatment were determined from previously published studies [11, 22, 78]. In vitro cell viability was similarly measured using Calcein-AM at 24 and 48 h post-treatment. Fluorescent images and readings were acquired using the SpectraMax MiniMax Imaging Cytometer. Cell lysates were harvested using RPPA lysis buffer after viability measurement, as previously described [79].
Lysate collection, SUnSET assay, and immunoblotting
Both OE19 and OE33 cells were seeded in T25 tissue culture flasks, transfected with siRNA, and treated with chemo drugs, as described above. Cell lysates were harvested using RPPA lysis buffer. To measure changes in global protein translation, the SUnSET assay was performed. Cell lysates for the SUnSET assay were treated with 1 μM puromycin for 30 min prior to lysate collection [80]. Protein was quantified using the DC Protein Assay (Bio-Rad, Hercules, CA, USA). Immunoblotting was performed using precast Criterion TGX Gels (Bio-Rad) with commercially available antibodies from Abcam (Waltham, MA, USA), Novus Biologicals (Littleton, CO, USA), Cell Signaling Technology (Danvers, MA, USA), ThermoFisher Scientific, MilliporeSigma, Santa Cruz Biotechnology (Dallas, TX, USA), and Developmental Studies Hybridoma Bank (Iowa City, IA, USA) (Table S2). Protein expression was quantified using ImageLab software (Bio-Rad) and normalized to loading controls (GAPDH or HSP60). siRNA transfection efficiency was also validated using protein detection via Western blots. Full-length, uncropped original Western blots are available as a Supplementary File.
Statistical analysis
For cell-based experiments, all treatment groups had a similar confluency at experimental initiation to ensure unbiased results following treatments. Statistical analysis of in vitro viability, migration, and ADCC assays was performed using Prism (version 10.0.3, GraphPad Software, San Diego, CA, USA) via one-way ANOVA with Tukey’s post-hoc test. Isoform switches linked with patient survival were identified using the Cox proportional-hazards model via the Survival package (version 3.5-5) in R. P-values ≤ 0.05 were considered statistically significant.
Data availability
RNA-seq data analyzed in this study are available in the NCBI Gene Expression Omnibus (GEO: GSE193946).
References
Rodriguez GM, DePuy D, Aljehani M, Bien J, Lee JSH, Wang DH, et al. Trends in epidemiology of esophageal cancer in the US, 1975-2018. JAMA Netw Open. 2023;6:e2329497.
Uhlenhopp DJ, Then EO, Sunkara T, Gaduputi V. Epidemiology of esophageal cancer: update in global trends, etiology and risk factors. Clin J Gastroenterol. 2020;13:1010–21.
Siegel RL, Kratzer TB, Giaquinto AN, Sung H, Jemal A. Cancer statistics, 2025. CA Cancer J Clin. 2025;75:10–45.
Anaparthy R, Sharma P. Progression of Barrett oesophagus: role of endoscopic and histological predictors. Nat Rev Gastroenterol Hepatol. 2014;11:525–34.
Bhat SK, McManus DT, Coleman HG, Johnston BT, Cardwell CR, McMenamin U, et al. Oesophageal adenocarcinoma and prior diagnosis of Barrett’s oesophagus: a population-based study. Gut. 2015;64:20–5.
van Hagen P, Hulshof MC, van Lanschot JJ, Steyerberg EW, van Berge, Henegouwen MI, et al. Preoperative chemoradiotherapy for esophageal or junctional cancer. N Engl J Med. 2012;366:2074–84.
Gaber CE, Sarker J, Abdelaziz AI, Okpara E, Lee TA, Klempner SJ, et al. Pathologic complete response in patients with esophageal cancer receiving neoadjuvant chemotherapy or chemoradiation: a systematic review and meta-analysis. Cancer Med. 2024;13:e7076.
Secrier M, Li X, de Silva N, Eldridge MD, Contino G, Bornschein J, et al. Mutational signatures in esophageal adenocarcinoma define etiologically distinct subgroups with therapeutic relevance. Nat Genet. 2016;48:1131–41.
Dulak AM, Stojanov P, Peng S, Lawrence MS, Fox C, Stewart C, et al. Exome and whole-genome sequencing of esophageal adenocarcinoma identifies recurrent driver events and mutational complexity. Nat Genet. 2013;45:478–86.
Testa U, Castelli G, Pelosi E. Esophageal cancer: genomic and molecular characterization, stem cell compartment and clonal evolution. Medicines. 2017;4:67.
Zhang Y, Weh KM, Howard CL, Riethoven JJ, Clarke JL, Lagisetty KH, et al. Characterizing isoform switching events in esophageal adenocarcinoma. Mol Ther Nucleic Acids. 2022;29:749–68.
Weh KM, Howard CL, Zhang Y, Tripp BA, Clarke JL, Howell AB, et al. Prebiotic proanthocyanidins inhibit bile reflux-induced esophageal adenocarcinoma through reshaping the gut microbiome and esophageal metabolome. JCI Insight. 2024;9:e168112.
Cancer Genome Atlas Research N, Weinstein JN, Collisson EA, Mills GB, Shaw KR, Ozenberger BA, et al. The Cancer Genome Atlas Pan-Cancer Analysis Project. Nat Genet. 2013;45:1113–20.
Cao D, Shan D, Yan W, Zhang Z, Song Q, Jiang Y, et al. Chaperone-mediated autophagy affects tumor cell proliferation and cisplatin resistance in esophageal squamous cell carcinoma. Thorac Cancer. 2021;12:1048–57.
Weh KM, Howell AB, Kresty LA. Expression, modulation, and clinical correlates of the autophagy protein Beclin-1 in esophageal adenocarcinoma. Mol Carcinog. 2016;55:1876–85.
Park C, Suh Y, Cuervo AM. Regulated degradation of Chk1 by chaperone-mediated autophagy in response to DNA damage. Nat Commun. 2015;6:6823.
Vakifahmetoglu-Norberg H, Kim M, Xia HG, Iwanicki MP, Ofengeim D, Coloff JL, et al. Chaperone-mediated autophagy degrades mutant p53. Genes Dev. 2013;27:1718–30.
Russo D, Ottaggio L, Foggetti G, Masini M, Masiello P, Fronza G, et al. PRIMA-1 induces autophagy in cancer cells carrying mutant or wild type p53. Biochim Biophys Acta. 2013;1833:1904–13.
Hetz C. The unfolded protein response: controlling cell fate decisions under ER stress and beyond. Nat Rev Mol Cell Biol. 2012;13:89–102.
Zhang G, Lv X, Yang Q, Liu H. Identification of HM13 as a prognostic indicator and a predictive biomarker for immunotherapy in hepatocellular carcinoma. BMC Cancer. 2022;22:888.
Liu J, Li W, Wu L. Pan-cancer analysis suggests histocompatibility minor 13 is an unfavorable prognostic biomarker promoting cell proliferation, migration, and invasion in hepatocellular carcinoma. Front Pharmacol. 2022;13:950156.
Wu J, Chen S, Liu H, Zhang Z, Ni Z, Chen J, et al. Tunicamycin specifically aggravates ER stress and overcomes chemoresistance in multidrug-resistant gastric cancer cells by inhibiting N-glycosylation. J Exp Clin Cancer Res. 2018;37:272.
Lindner P, Christensen SB, Nissen P, Moller JV, Engedal N. Cell death induced by the ER stressor thapsigargin involves death receptor 5, a non-autophagic function of MAP1LC3B, and distinct contributions from unfolded protein response components. Cell Commun Signal. 2020;18:12.
Ma Q, Zhang W, Wu K, Shi L. The roles of KRAS in cancer metabolism, tumor microenvironment and clinical therapy. Mol Cancer. 2025;24:14.
Zhang Y, Weh KM, Tripp BA, Clarke JL, Howard CL, Sunilkumar S, et al. Cranberry proanthocyanidins mitigate reflux-induced transporter dysregulation in an esophageal adenocarcinoma model. Pharmaceuticals. 2023;16:1697.
Jeong S, Lim S, Schevzov G, Gunning PW, Helfman DM. Loss of Tpm4.1 leads to disruption of cell-cell adhesions and invasive behavior in breast epithelial cells via increased Rac1 signaling. Oncotarget. 2017;8:33544–59.
Wen R, Xiao Y, Zhang Y, Yang M, Lin Y, Tang J. Identification of a novel transcript isoform of the TTLL12 gene in human cancers. Oncol Rep. 2016;36:3172–80.
Xiong Y, Deng Y, Wang K, Zhou H, Zheng X, Si L, et al. Profiles of alternative splicing in colorectal cancer and their clinical significance: a study based on large-scale sequencing data. Ebiomedicine. 2018;36:183–95.
Wijeratne S, Gonzalez MEH, Roach K, Miller KE, Schieffer KM, Fitch JR, et al. Full-length isoform concatenation sequencing to resolve cancer transcriptome complexity. BMC Genomics. 2024;25:122.
Zhang B, Wu Q, Cheng S, Li W. Systematic profiling of mRNA splicing reveals the prognostic predictor and potential therapeutic target for glioblastoma multiforme. J Oncol. 2021;2021:4664955.
Zhao W, Hoadley KA, Parker JS, Perou CM. Identification of mRNA isoform switching in breast cancer. BMC Genomics. 2016;17:181.
Larionova TD, Bastola S, Aksinina TE, Anufrieva KS, Wang J, Shender VO, et al. Alternative RNA splicing modulates ribosomal composition and determines the spatial phenotype of glioblastoma cells. Nat Cell Biol. 2022;24:1541–57.
Yan Y, Ren Y, Bao Y, Wang Y. RNA splicing alterations in lung cancer pathogenesis and therapy. Cancer Pathog Ther. 2023;1:272–83.
Ray D, Ray P, Ferrer-Torres D, Wang Z, Nancarrow D, Yoon HW, et al. Isoforms of RNF128 regulate the stability of mutant P53 in Barrett’s esophageal cells. Gastroenterology. 2020;158:583–97.e1.
Sanada H, Seki N, Mizuno K, Misono S, Uchida A, Yamada Y, et al. Involvement of dual strands of miR-143 (miR-143-5p and miR-143-3p) and their target oncogenes in the molecular pathogenesis of lung adenocarcinoma. Int J Mol Sci. 2019;20:4482.
Liu J, Li H, Shen S, Sun L, Yuan Y, Xing C. Alternative splicing events implicated in carcinogenesis and prognosis of colorectal cancer. J Cancer. 2018;9:1754–64.
Massoner P, Lueking A, Goehler H, Hopfner A, Kowald A, Kugler KG, et al. Serum-autoantibodies for discovery of prostate cancer specific biomarkers. Prostate. 2012;72:427–36.
Heitor da Silva Maues J, Ferreira Ribeiro H, de Maria Maues Sacramento R, Maia de Sousa R, Pereira de Tommaso R, Dourado Kovacs Machado Costa B, et al. Downregulated genes by silencing MYC pathway identified with RNA-SEQ analysis as potential prognostic biomarkers in gastric adenocarcinoma. Aging. 2020;12:24651–70.
Wang F, Yuan Q, Chen F, Pang J, Pan C, Xu F, et al. Fundamental mechanisms of the cell death caused by nitrosative stress. Front Cell Dev Biol. 2021;9:742483.
Deshpande A, Brants J, Wasylyk C, van Hooij O, Verhaegh GW, Maas P, et al. TTLL12 has a potential oncogenic activity, suppression of ligation of nitrotyrosine to the C-terminus of detyrosinated alpha-tubulin, that can be overcome by molecules identified by screening a compound library. PLoS ONE. 2024;19:e0296960.
Xue L, Zeng Y, Fang C, Cheng W, Li Y. Effect of TTLL12 on tubulin tyrosine nitration as a novel target for screening anticancer drugs in vitro. Oncol Lett. 2020;20:340.
Abdel-Latif MM, Duggan S, Reynolds JV, Kelleher D. Inflammation and esophageal carcinogenesis. Curr Opin Pharmacol. 2009;9:396–404.
Suzuki H, Iijima K, Scobie G, Fyfe V, McColl KE. Nitrate and nitrosative chemistry within Barrett’s oesophagus during acid reflux. Gut. 2005;54:1527–35.
Iijima K, Grant J, McElroy K, Fyfe V, Preston T, McColl KE. Novel mechanism of nitrosative stress from dietary nitrate with relevance to gastro-oesophageal junction cancers. Carcinogenesis. 2003;24:1951–60.
Liu J, Wang L, He H, Liu Y, Jiang Y, Yang J. The complex role of chaperone-mediated autophagy in cancer diseases. Biomedicines. 2023;11:2050.
Fichter CD, Herz C, Munch C, Opitz OG, Werner M, Lassmann S. Occurrence of multipolar mitoses and association with Aurora-A/-B kinases and p53 mutations in aneuploid esophageal carcinoma cells. BMC Cell Biol. 2011;12:13.
Bairoch A. The Cellosaurus, a cell-line knowledge resource. J Biomol Tech. 2018;29:25–38.
Yang H, Li Z, Wang Z, Zhang X, Dai X, Zhou G, et al. Histocompatibility Minor 13 (HM13), targeted by miR-760, exerts oncogenic role in breast cancer by suppressing autophagy and activating PI3K-AKT-mTOR pathway. Cell Death Dis. 2022;13:728.
Ren Y, Mao Y, Yuan X. HM13 is a predictive biomarker of metastasis and neutrophil infiltration in colorectal cancer. Histol Histopathol. 2024:18857:1435–1446.
Jin X, Wang H, Wang Y. The role of HM13 expression and its relationship to PI3K/Akt and p53 signaling pathways in colorectal cancer. Tissue Cell. 2025;93:102702.
Acosta-Alvear D, Karagoz GE, Frohlich F, Li H, Walther TC, Walter P. The unfolded protein response and endoplasmic reticulum protein targeting machineries converge on the stress sensor IRE1. eLife. 2018;7:e43036.
Liang D, Khoonkari M, Avril T, Chevet E, Kruyt FAE. The unfolded protein response as regulator of cancer stemness and differentiation: mechanisms and implications for cancer therapy. Biochem Pharmacol. 2021;192:114737.
Lu H, Gomaa A, Wang-Bishop L, Ballout F, Hu T, McDonald O, et al. Unfolded protein response is activated by Aurora Kinase A in esophageal adenocarcinoma. Cancers. 2022;14:1401.
Yang M, Wei X, Yi X, Jiang DS. Mitophagy-related regulated cell death: molecular mechanisms and disease implications. Cell Death Dis. 2024;15:505.
Ge F, Huo Z, Cai X, Hu Q, Chen W, Lin G, et al. Evaluation of clinical and safety outcomes of neoadjuvant immunotherapy combined with chemotherapy for patients with resectable esophageal cancer: a systematic review and meta-analysis. JAMA Netw Open. 2022;5:e2239778.
Davern M, Fitzgerald MC, Buckley CE, Heeran AB, Donlon NE, McGrath J, et al. PD-1 and TIGIT blockade differentially affect tumour cell survival under hypoxia and glucose deprived conditions in oesophageal adenocarcinoma; implications for overcoming resistance to PD-1 blockade in hypoxic tumours. Transl Oncol. 2022;19:101381.
Cakan E, Lara OD, Szymanowska A, Bayraktar E, Chavez-Reyes A, Lopez-Berestein G, et al. Therapeutic antisense oligonucleotides in oncology: from bench to bedside. Cancers. 2024;16:2940.
Zhang Y, Qian J, Gu C, Yang Y. Alternative splicing and cancer: a systematic review. Signal Transduct Targeted Therapy. 2021;6:78.
Bray NL, Pimentel H, Melsted P, Pachter L. Near-optimal probabilistic RNA-seq quantification. Nat Biotechnol. 2016;34:525–7.
Vitting-Seerup K, Sandelin A. IsoformSwitchAnalyzeR: analysis of changes in genome-wide patterns of alternative splicing and its functional consequences. Bioinformatics. 2019;35:4469–71.
Vitting-Seerup K, Sandelin A. The landscape of isoform switches in human cancers. Mol Cancer Res. 2017;15:1206–20.
Wang L, Park HJ, Dasari S, Wang S, Kocher JP, Li W. CPAT: coding-potential assessment tool using an alignment-free logistic regression model. Nucleic Acids Res. 2013;41:e74.
Almagro Armenteros JJ, Tsirigos KD, Sonderby CK, Petersen TN, Winther O, Brunak S, et al. SignalP 5.0 improves signal peptide predictions using deep neural networks. Nat Biotechnol. 2019;37:420–3.
Mészáros B, Erdős G, Dosztányi Z. IUPred2A: context-dependent prediction of protein disorder as a function of redox state and protein binding. Nucleic Acids Res. 2018;46:W329–W37.
Punta M, Coggill PC, Eberhardt RY, Mistry J, Tate J, Boursnell C, et al. The Pfam protein families database. Nucleic Acids Res. 2011;40:D290–D301.
Heberle H, Meirelles GV, da Silva FR, Telles GP, Minghim R. InteractiVenn: a web-based tool for the analysis of sets through Venn diagrams. BMC Bioinformatics. 2015;16:169.
Wickham H. ggplot2: elegant graphics for data analysis. New York: Springer-Verlag; 2016.
Szklarczyk D, Franceschini A, Wyder S, Forslund K, Heller D, Huerta-Cepas J, et al. STRING v10: protein-protein interaction networks, integrated over the tree of life. Nucleic Acids Res. 2015;43:D447–52.
Ye J, Coulouris G, Zaretskaya I, Cutcutache I, Rozen S, Madden TL. Primer-BLAST: a tool to design target-specific primers for polymerase chain reaction. BMC Bioinformatics. 2012;13:134.
Li JCA. Modeling survival data: extending the Cox model. Sociol Method Res. 2003;32:117–20.
Mariotto AB, Noone AM, Howlader N, Cho H, Keel GE, Garshell J, et al. Cancer survival: an overview of measures, uses, and interpretation. J Natl Cancer Inst Monogr. 2014;2014:145–86.
Kresty LA, Weh KM, Zeyzus-Johns B, Perez LN, Howell AB. Cranberry proanthocyanidins inhibit esophageal adenocarcinoma in vitro and in vivo through pleiotropic cell death induction and PI3K/AKT/mTOR inactivation. Oncotarget. 2015;6:33438–55.
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. Basic local alignment search tool. J Mol Biol. 1990;215:403–10.
Suarez-Arnedo A, Torres Figueroa F, Clavijo C, Arbelaez P, Cruz JC, Munoz-Camargo C. An ImageJ plugin for the high throughput image analysis of in vitro scratch wound healing assays. PLoS ONE. 2020;15:e0232565.
Schindelin J, Arganda-Carreras I, Frise E, Kaynig V, Longair M, Pietzsch T, et al. Fiji: an open-source platform for biological-image analysis. Nat Methods. 2012;9:676–82.
Boyerinas B, Jochems C, Fantini M, Heery CR, Gulley JL, Tsang KY, et al. Antibody-dependent cellular cytotoxicity activity of a novel anti-PD-L1 antibody avelumab (MSB0010718C) on human tumor cells. Cancer Immunol Res. 2015;3:1148–57.
Giles AJ, Hao S, Padget M, Song H, Zhang W, Lynes J, et al. Efficient ADCC killing of meningioma by avelumab and a high-affinity natural killer cell line, haNK. JCI Insight. 2019;4:e130688.
Ma Z, Fan C, Yang Y, Di S, Hu W, Li T, et al. Thapsigargin sensitizes human esophageal cancer to TRAIL-induced apoptosis via AMPK activation. Sci Rep. 2016;6:35196.
Weh KM, Turgeon DK, Rubenstein JH, Clarke JL, Howell AB, Chang AC, et al. Proanthocyanidins mitigate bile acid-induced changes in GSTT2 levels in a panel of racially diverse patient-derived primary esophageal cell cultures. Mol Carcinog. 2022;61:281–7.
Ravi V, Jain A, Mishra S, Sundaresan NR. Measuring protein synthesis in cultured cells and mouse tissues using the non-radioactive SUnSET assay. Curr Protoc Mol Biol. 2020;133:e127.
Acknowledgements
We would like to thank all patients who provided tissue samples for this study.
Funding
We thank the National Institutes of Health and National Cancer Institute (R01CA158319, U54CA163059) and the University of Michigan (U057239) for supporting this study. This study was additionally supported by the John and Carla Klein Family Research Fund, awarded to L.A.K., and the University of Michigan Medical School Pandemic Research Recovery Fund, awarded to Y.Z.
Author information
Authors and Affiliations
Contributions
Conceptualization, Y.Z. and L.A.K.; methodology, Y.Z., J.R., J.L.C., A.D., M.A.S, and L.A.K.; software, Y.Z. and J.R.; validation, Y.Z., D.N., and L.A.K.; formal analysis, Y.Z., D.N., R.I., and B.V.; investigation, Y.Z., D.N., R.I., and B.V.; resources, S.B., K.H.L., J.L., R.M.R., A.C.C., D.D.O., and L.A.K.; data curation, Y.Z., J.R., and L.A.K.; writing—original draft preparation, Y.Z. and L.A.K.; writing—review and editing, Y.Z., D.N., R.I., B.V., S.B., J.R., J.L.C., K.H.L., J.L., R.M.R., A.C.C., D.D.O., A.D., M.A.S., and L.A.K.; visualization, Y.Z. and D.N.; supervision, L.A.K.; project administration, L.A.K.; funding acquisition, L.A.K.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics statement
The study is approved by the University of Michigan Institutional Review Board. Each patient provided written informed consent and the study was performed in accordance with the Declaration of Helsinki.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited By Professor Stephen Tait
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhang, Y., Ntsiful, D.A., Israel, R. et al. Identification of isoform switching events linked with esophageal adenocarcinoma patient survival informs novel prognostic and therapeutic targets. Cell Death Dis 17, 305 (2026). https://doi.org/10.1038/s41419-026-08542-2
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41419-026-08542-2