Abstract
Chimeric antigen receptor-modified T cell (CAR-T) immunotherapy has revolutionised blood cancer treatment. Parsing the genetic underpinnings of T cell quality and CAR-T efficacy is challenging. Transcriptomics inform CAR-T state, but the nature of dynamic transcription during activation hinders identification of transiently or minimally expressed genes, such as transcription factors, and over-emphasises effector and metabolism genes. Here we explore whether analyses of transcriptionally repressive and permissive histone methylation marks describe CAR-T cell functional states and therapeutic potential beyond transcriptomic analyses. Histone mark analyses improve identification of differences between naïve, central memory, and effector memory CD8 + T cell subsets of human origin, and CAR-T derived from these subsets. We find important differences between CAR-T manufactured from central memory cells of healthy donors and of patients. By examining CAR-T products from a clinical trial in lymphoma (NCT01865617), we find a novel association between the activity of the transcription factor KLF7 with in vivo CAR-T accumulation in patients and demonstrate that over-expression of KLF7 increases in vitro CAR-T proliferation and IL-2 production. In conclusion, histone marks provide a rich dataset for identification of functionally relevant genes not apparent by transcriptomics.
Similar content being viewed by others
Introduction
In human blood, naïve, central memory (CM) and effector memory (EM) CD8 + T cells can be identified by distinct patterns of expression of surface antigens such as CCR7, CD45RA and CD45RO. These subsets differ in homing, proliferative and effector function, as well as in their propensity to dysfunction, hence their utilities as starting T cell products for manufacturing adoptive cellular immunotherapy1. Chimeric antigen receptor (CAR)-modified T cell (CAR-T) therapy is effective for a range of haematologic malignancies2,3,4. Being a “living drug” presents the opportunity to identify genetic factors in precursor T cells and CAR-T infusion products (IP) that predict product quality and patient outcomes. Pre-clinical and clinical trial data highlight the importance of the differentiation state of the T cell subset-of-origin from which CAR-T are manufactured. In immunodeficient, tumour-bearing xenografted mice treatment with human CAR-T manufactured from naïve or CM CD8 + T cells improved survival when compared to CAR-T manufactured from EM cells5. Furthermore, in clinical trials of CD19 CAR-T immunotherapies in adults and children, higher proportions of CM and naïve cells in apheresis products were associated with improved persistence and efficacy3,4.
Differentiation of T cell subsets is epigenetically programmed and perpetuated in progeny cells by chromatin states and DNA methylation1. T cell chromatin is dynamic, and during gene activation transitions from a condensed, heterochromatin structure, including trimethylation of lysine 27 of histone H3 (H3K27me3), to an active chromatin structure marked by dimethylation of lysine 4 of histone H3 (H3K4me2). Histone marks are central to T cell biology. For example, H3K27me3 marks repression of memory and pro-survival transcriptions factors (TFs) in terminal effector cells6; and permissive H3K4me2 marks master TF regulators of effector T cell differentiation and function6. To date, there have been no high-throughput, genome-wide, detailed analyses of histone marks in human CAR-T.
Gene expression signatures arising from T cell activation during CAR-T manufacturing engender stochastic, statistical transcriptional noise that can obscure differential expression7. This limits the capacity to use transcriptomics approaches to identify prognostic factors for CAR-T therapy3. Furthermore, transcriptomic approaches typically highlight effector and metabolism genes8 while underemphasising low or transiently expressed genes, like transcription factors, which are crucial for cell potential and fate9. Sparsity inherent in single-cell methods also limits the identification of cell fate-associated10. The limitations of transcriptomic approaches are even apparent in the longitudinal tracking of T cell clonotypes11.
Complementary and orthogonal epigenomic approaches provide a relatively narrow dynamic range that may address transcriptomic limitations and thus provide novel insights into T cell and CAR-T potential. Indeed, histone marks and analyses of open chromatin using Assay for Transposase-Accessible Chromatin with Sequencing (ATAC-Seq) better differentiate T cell subsets when compared to RNA-seq6,12. Recent work suggests that histone marks may better reflect stable, inherited cell potential than open DNA as identified by ATAC-seq13. Chromatin ImmunoPrecipitation with Sequencing (ChIP-Seq) of protein bound DNA is the most widely employed method for analysis of histone marks; however, it is limited by inter- and intra-experimental variation, a low signal-to-noise ratio, low throughput and requirement for large numbers of cells14. These experimental requirements are a major impediment to assessment of limited clinical samples. In contrast, Cleavage Under Targets and Release Using Nuclease (CUT&RUN) is a high throughput, low cost, high signal-to-noise ratio technology that allows deep sequencing of DNA bound to histone marks with relatively few cells and is less subject to methodological variation compared to ChIP-seq13.
To understand factors contributing to the different capabilities of distinct CAR-T IPs, we present paired analyses of the transcriptome using RNA-seq with analyses of H3K4me2 and H3K27me3 marks using CUT&RUN on human CD8 + T cell subsets and CAR-T manufactured from these subsets. We show how supplementation of transcriptome analyses with novel histone mark analyses improves the discriminative capacity of differentially expressed genes and enables better identification of T cell-fate determining genes in human T cell subsets. Furthermore, we demonstrate that histone mark analyses distinguish CAR-T derived from distinct T cell subsets from healthy donors and lymphoma patients. This novel platform was used to identify an association between a transcription factor, KLF7, and CAR-T proliferation in vivo in lymphoma patients; and to show that in vitro transgenic KLF7 overexpression is associated with an increased CCR7 + /CD45RA- phenotype and IL-2 production. These data demonstrate that histone mark analyses provide deep insights into the potential of T cell subsets and CAR-T immunotherapy products and define further avenues for the creation of next generation CAR-T cell products.
Results
High dimensional analyses of histone marks distinguish distinct human CD8 + T cell subsets
We set out to compare transcriptomic and epigenomic differences in T cell precursors and CAR-T products that might govern therapeutic potential. CD8 + T cells were sorted from healthy donors (HD) into CD62L+ CD45RAhi CD45RO- (naïve immunophenotype), CD62L + CD45RA- CD45RO+ (central memory immunophenotype; CM), and CD62L- CD45RA- (effector memory immunophenotype; EM) subsets (Supplementary Fig. 1A). The sorted naïve, CM, and EM subsets were of high purity and demonstrated expression of canonical immunophenotypic markers of naïve, CM, and EM cells (Supplementary Figs. 1B, C)1. To investigate the relationship of transcriptomic output and histone marks in CD8 + T cell subset, we performed RNA-seq analyses and CUT&RUN analyses of transcriptionally repressive H3K27me3 and permissive H3K4me2 histone marks on each subset (Supplementary Fig. 2). Quality check (QC) metrics were satisfied for RNA-seq samples and data (6 HD, 3 T cell subsets/HD; Supplementary Fig. 3) and CUT&RUN data (4 HD, 3 T cell subsets/HD; Supplementary Fig. 4). Across donors, consistent distributions of up- and down-regulated genes were observed (Supplementary Fig. 5) with differential signals being unaffected by small variations (Supplementary Fig. 6), indicating statistical robustness of these datasets. The high quality of RNA-seq data was confirmed by the observed correlation between genes differentially expressed in CM vs EM cells in our study with recently published human CD8 + T cell subset data (P = 4.85 × 10−13, odds ratio = 31.54 by Fisher’s exact test)15. We observed that H3K27me3 marks were mostly located within gene bodies and in intergenic regions, whereas a greater proportion of H3K4me2 marks were associated with promoter regions (Supplementary Fig. 7A-B), confirming the specificity of our CUT&RUN profiles. Genomic tracks coding for TFs that have been typically associated with naïve and CM subsets were marked by low H3K27me3 in naïve and CM subsets and low H3K4me2 in EM subsets; and vice-versa for TFs that have been associated with effector differentiation (Supplementary Fig. 8)16,17,18,19,20.
Next, a genome-wide peak calling strategy was implemented to identify regions enriched for H3K27me3 and H3K4me2 marks. To allow direct comparison of epigenomic and transcriptomic data, we focused on gene units, assigning H3K27me3 and H3K4me2 marks within ± 1 kb of the transcriptional start site (TSS) to an equivalent transcript, as previously described (Fig. 1A)13. In keeping with the transcriptionally permissive and repressive nature of H3K4me2 and H3K27me3 marks, respectively13, the most highly expressed genes had the highest proportion of H3K4me2 marks and the lowest proportion of H3K27me3 marks within ±1 kb of the TSS in each subset (Fig. 1A). When comparing the numbers of genes statistically differentially expressed (DEx) by RNA-seq or differentially enriched (DEn) by histone marks, larger differences were seen between naïve and EM subsets than between naïve and CM, and the smallest differences were observed between CM and EM subsets (Fig. 1B, Supplementary Fig. 9 & Supplementary Data 1), consistent with known relationships between these subsets1. When comparing subsets, more DEn genes were identified by CUT&RUN than DEx genes recognised by RNA-seq, including more DEn TFs between subsets (Fig. 1B, C). Principal Component Analyses (PCA) and Uniform Manifold Approximation and Projection (UMAP) more clearly distinguished T cell subsets using histone mark data compared to RNA-seq data (Fig. 1D, E). These analyses demonstrate that, even when comparing equivalent gene units, CUT&RUN profiling of the histone modifications H3K4me2 and H3K27me3 detects a greater variance and uncovers more differences between resting HD CD8 + T cell subsets than RNA-seq.
A Histogram (top row) of average log-normalised count of sequenced DNA bound to H3K4me2 (left column for each subset) or H3K27me3 (right column for each subset) within ±1 kb of the transcriptional start site (TSS). Tornado plots (second row) of CD8 + T cell subsets (naïve, N, left; central memory, CM, middle; effector memory, EM, right) depict the relationship between transcriptome abundance and histone marks. Each line represents a genomic region within ±1 kb of the TSS for an annotated gene. The genes in rows are ordered by transcript abundance by RNA-seq from high (top) to low (bottom). The heatmap is coloured by the intensity of H3K4me2 or H3K27me3 marks (histone marks) assessed by Cleavage Under Targets and Restriction Using Nuclease (CUT&RUN). H3K4me2 and H3K27me3 marks show a direct and inverse relationship with transcript abundance, respectively. B, C. Numbers of differentially expressed (by RNA-seq) and differentially enriched (by CUT&RUN) genes (B) and transcription factors (C) when comparing CD8 + T cell subsets. Genes assigned to peaks ± 1 kb of the transcriptional start site (TSS), each NCBI annotated gene counted only once (adjusted P value < 0.05 and Log2 fold change (LogFC) ≥ 1). D Principal component analyses (PCA) and (E) Uniform Manifold Approximation and Projection (UMAP) of RNA-seq (left) and H3K27me3 (middle left), H3K4me2 (middle right), and H3K27me3 and H3K4me2 (right) histone mark analyses show a distinction between CD8 + T cell subsets by histone marks that is less apparent by RNA-seq.
Genes marked by different combinations of H3K4me2 and H3K27me3 exhibit different mean levels of transcript abundance in CD8 + T cell subsets
The greater distinction between CD8 + T cell subsets by histone marks compared to RNA-seq led us to examine the associations between histone marks and transcript abundance. We defined genes that were positive and negative for H3K27me3 and H3K4me2 using a two-component Gaussian mixture model to fit histone mark distribution within ±1 kb of the TSS (Fig. 2A, B & Supplementary Fig. 10). This approach allowed us to visualise the enrichment of histone marks around the TSS and classify all genes as either H3K4me2 or H3K27me3 single-marked, bivalent, or unmarked (Fig. 2C). When integrated with RNA-seq, most genes with abundant transcripts were marked with H3K4me2 and not H3K27me3, as expected (Fig. 2D). However, transcripts were also detected in numerous genes that were H3K4me2-H3K27me3-, H3K4me2+H3K27me3+ or H3K4me2-H3K27me3 + . We identified a bimodal pattern of transcript abundance, exemplified by genes exhibiting low or undetected transcript normalised to sequencing depths (RNAlo; below 2.5 transcript copies/sample) and genes with more abundant transcript (RNAhi; above 2.5 transcript copies/sample) (Supplementary Fig. 11). The mean transcript abundance increased in a linear trend with changes in histone marks from H3K4me2-H3K27me3+ to H3K4me2-H3K27me3- to H3K4me2+H3K27me3+ to H3K4me2+H3K27me3- (Fig. 2G–I). Combining the bimodal distribution of RNA transcript (RNAhi and RNAlo) and the four patterns of histone marks (H3K4me2-H3K27me3 + ; H3K4me2-H3K27me3-; H3K4me2+H3K27me3 + ; H3K4me2+H3K27me3-) allowed categorisation of genes into eight groups (Fig. 2J & Supplementary Data 2). In each CD8 + T cell subset, the most frequently observed gene categories were those that aligned with known correlations of H3K4me2 with transcriptional permission and H3K27me3 with transcriptional repression (RNAhi H3K4me2+H3K27me3- genes, 26.93% in naïve, 26.72% in CM, 25.94% in EM; RNAlo H3K4me2-H3K27me3+ genes, 33.24% in naïve, 35.76% in CM and 35.22% in EM; Fig. 2K). However, numerous genes were identified in each subset that did not correspond to the most common histone marks and RNA-seq patterns.
A, B Histogram of two-component Gaussian mixture model (GMM) to characterise distribution of H3K27me3- and H3K4me2-bound sequenced DNA reads from naïve CD8 + T cells within 10 kb of transcriptional start site (TSS). A bimodal distribution of read intensity is noted and modelled by GMM with negative reads depicted in red and positive reads depicted in blue. The nadir of the distributions is represented by the dashed line, providing a positive and negative cut-off used to integrate epigenomic and transcriptomics analyses in subsequent figures. Identical cut-offs were seen with central memory (CM) and effector memory (EM) cells (see Supplementary Fig. 10). C Histogram (top row) of average log-normalised counts of genes that are deemed positive, negative and bivalent for H3K4me2- and H3K27me3-specific reads across subsets. Individual genes, single- or bivalent for each subset, are shown in tornado plots below, ordered and coloured by intensity of counts. D–F Three-dimensional scatter plots of naïve, CM, and EM CD8 + T cells. Each plot shows H3K27me3 (y-axis) and H3K4me2 (x-axis) histone mark enrichment signals for single annotated genetic regions, each region being represented by a single point; transcript abundance (by RNA-seq) is denoted by the colour of each point, as shown in the legend. Quadrants are constructed using the nadir of bimodal histograms of each histone mark as per A. G–I Mean total transcript level +/- SD for all genes (overlayed as data points) with different patterns of H3K27me3 (K27m3) and H3K4me2 (K4m2) histone marks in CD8 + T cell subsets. ****P value < 0.0001 by one-way ANOVA with a significant linear trend. J Four patterns of H3K4me2 and H3K27me3 histone marks in both low and high transcript abundance states are shown. K Percent annotated genes in CD8 + T cell subsets within each RNA-seq/histone methylation mark (HMM) pattern.
Patterns of combined H3K4me2 and H3K27me3 marks differ in the absence of differential gene expression when comparing distinct CD8 + T cell subsets
Statistical noise in high transcript abundance genes and sparsity of sequencing depth in low transcript abundance genes complicates the discriminative capacity of RNA-seq7. Because we noted that the mean transcript abundance for genes was found in an ordinal pattern (Fig. 2G–I), we considered whether relative differences in combinatorial histone mark patterns might inform differences between cell types. We constructed heatmaps to visualise the number of genes that differ in histone mark patterns when comparing pairs of CD8 + T cell subsets using genesets comprising highly (RNAhi) and lowly (RNAlo) expressed genes (Fig. 3A). More genes showed changes in histone mark patterns within the same transcript abundance category (i.e., RNAlo vs RNAlo; RNAhi vs RNAhi) then across transcript abundance categories, intimating that histone mark patterns might provide additional insight into differences between two cell subsets, even amongst non-DEx genes (Fig. 3A). To further investigate this possibility, we grouped RNAhi or RNAlo genes into DEx and non-DEx groups in each T cell subset comparison. We found that within each of these groups, there were numerous genes that differed in histone mark patterns (Fig. 3B, C & Supplementary Fig. 12A). Within genes that were DEx, we identified a moderate number of genes with changes in histone mark pattern between subsets, especially between naïve and memory subsets (i.e., naïve vs CM and naïve vs EM; Fig. 3B, C & Fig. 12A). However, in both RNAhi and RNAlo subsets there were more genes with histone mark pattern changes that were not DEx compared to those that were DEx, a difference most marked when comparing CM and EM subsets (Fig. 3B, C, Supplementary Fig. 12A). These data show that histone mark patterns differ between CD8 + T cell subsets in numerous highly and lowly expressed genes, even if those genes are not DEx between T cell subsets.
A Heatmaps summarising the number of genes that occur within each histone mark pattern (H3K4me2 = K4m2; H3K27me3 = K27m3) when comparing a less differentiated cell type (x axis) to a more differentiated cell type (y axis) for naïve vs central memory (CM), CM vs effector memory (EM), and naïve vs EM. Most genes tend to maintain RNAhi or RNAlo status and the same histone mark pattern between subsets. The number of genes is indicated by the colour-coding (maximum depicted colour-coded count, 500 genes). B, C. Sub-analyses of the heatmap from the top left corner of each comparative heatmap in A comparing a less differentiated cell type to a more differentiated cell type for naïve vs CM, CM vs EM and naïve vs EM. These analyses include RNAhi genes that do not (B) or do (C) show a significant change in transcript expression by limma (LogFC > 1, adjusted P < 0.05 by Benjamini-Hochberg procedure) between compared subsets. Genes on the diagonal (top left to bottom right of each plot) do not change histone mark pattern between subsets. Genes within red boxes show changes in histone mark patterns that indicate enrichment in the more differentiated cell types. Genes within white boxes show changes in histone mark patterns that indicate enrichment in the less differentiated cell types. Most genes that change histone mark patterns between subsets are not identified by differential gene expression analyses of RNA-seq data (compare B to C).
Directionality can be ascribed to genes that differ in patterns of combined H3K4me2 and H3K27me3 marks when comparing distinct CD8 + T cell subsets
Our finding that patterns of histone marks identified numerous differences between subsets that were not identified by RNA-seq prompted us to determine if directionality could be assigned to genes with changing histone mark patterns between subsets. This was feasible for genes that changed from a repressive H3K4me2-H3K27me3+ to permissive H3K4me2+H3K27me3- pattern, but the relevance of changes involving bivalent (H3K4me2+H3K27me3 + ) and unmarked (H3K4me2-H3K27me3-) patterns was less clear. Genes with bivalent patterns were common: approximately 3000–3300 bivalent genes, representing 8.9%−9.8% of all promoters investigated. The importance of bivalent patterns was emphasised when we evaluated RNAhi genes in which high fractions of DEx genes were found to be bivalently marked (Fig. 4A). Furthermore, when RNAhi genes were categorised according to histone mark patterns, the clearest distinctions between T cell subsets were observed in PCAs of RNA-seq data from bivalent genes (Fig. 4Bcf Fig. 1D). PCAs of RNA-seq data in transcriptionally permissive H3K4me2+H3K27me3- marked genes did not effectively distinguish T cell subsets. These data indicate that classifying genes into distinct patterns of H3K4me2 and H3K27me3 marks enhances the discriminatory capacity of RNA-seq in distinguishing T cell subsets, possibly by highlighting genes in states of active transcriptional flux.
A Stacked bar plots of the percent of all RNAhi genes within each histone mark pattern (H3K4me2 = K4m2; H3K27me3 = K27m3) in which relative transcript level is either increased in a more differentiated cell-type (“Up”), increased in a less differentiated cell-type (“Down”) or not differentially expressed (“notDE”) by limma. B PCA plots of transcript from RNAhi genes subsetted by the histone mark pattern. The plots demonstrate that transcript differences in the H3K4me2+H3K27me3+ pattern identify differences between resting naïve, central memory (CM) and effector memory (CM) cells that are not apparent with RNA-seq alone (compare to Fig. 1D). C Median LogFC in transcript in RNAhi genes (when the number of genes in each pattern ≥ 3) between histone mark patterns when comparing a less differentiated cell type (x axis) to a more differentiated cell type (y axis) for naïve vs CM, CM vs EM and naïve vs EM. Red boxes represent changes in histone mark patterns that represent a shift towards a more differentiated cell type as evidenced by positive (“Up”) median LogFC in transcript. White boxes represent changes in histone mark patterns that represent a shift towards a less differentiated cell type as evidenced by negative (“Down”) median LogFC in transcript. Grey indicates the number of genes in an individual pattern is < 3. D MA plot of genes with a CM to EM histone mark pattern (red box, CM vs EM heatmap, Fig. 3B) and genes DEx by RNA-seq (in red). Genes with a known association with effector-directed differentiation are named. E. MA plot of genes with an EM to CM histone mark pattern (white box, CM vs EM heatmap, Fig. 3B) and genes DEx by RNA-seq (in red). Genes with a known association with memory-directed differentiation and quiescence are named.
When we analysed DEx RNAhi genes, a characteristic association emerged between changes in histone mark pattern and the fold change in transcript abundance between subsets (Fig. 4C). Genes marked with bivalent or unmarked patterns showed a median fold change in transcript abundance that was intermediate between that of genes marked by repressive H3K4me2-H3K27me3+ in one subset and permissive H3K4me2+H3K27me3- in another. Therefore, directionality could be ascribed to genes exhibiting distinct histone mark changes between subsets: either towards a more differentiated cell type (positive fold change, red box, Fig. 4C) or a less differentiated cell type (negative fold change, white box, Fig. 4C). This allowed us to identify other genes associated with differentiation towards EM or dedifferentiation towards CM that were not DEx by RNA-seq. We identified non-DEx RNAhi and RNAlo genes between T cell subsets that exhibited changes in histone mark patterns towards CM (Fig. 3B & Supplementary Fig. 12A, red boxes) or towards EM (Fig. 3B & Supplementary Fig. 12A, white boxes). Such histone mark pattern analyses uncovered additional and non-overlapping genes that differed between subsets compared to those discovered by single H3K4me2 analyses and single H3K27me3 analyses (Supplementary Fig. 13). The data indicate that analyses of changes in histone mark patterns can identify genes associated with differentiation even in the absence of differential gene expression, further evidencing the capabilities of epigenomic analyses.
Comparison of patterns of combined H3K4me2 and H3K27me3 marks in CM and EM CD8 + T cells identifies T cell differentiation-associated genes not found by transcriptomics
Being able to identify potentially subtle differences between CM and EM CD8 + T cells is of importance given that preclinical and clinical datasets show the superiority of the CAR-T manufactured from the former when compared to those manufactured from the latter3,4,5. To determine if histone mark patterns could identify differences between CM and EM cells beyond that apparent by RNA-seq, we compared changes in the patterns of histone marks between CM and EM in all RNAhi genes (Fig. 3B, middle column). This approach uncovered numerous genes associated with T cell differentiation in genes with both low and high mean transcript abundance that was not DEx by RNA-seq (Fig. 4D, E & Supplementary Fig. 12B-C). We identified statistically significant overlap between these genes and published gene lists from comparisons of human CM and EM CD8 + T cells by scRNA-seq and bulk RNA-seq15,21 (Supplementary Fig. 14A), indicating the relevance of the RNAhi genes discovered by histone mark patterns. Although significant, the overlaps were small (Supplementary Fig. 14B), demonstrating the capacity to discover additional RNAhi genes by analyses of histone mark patterns. We used a similar approach to compare RNAlo genes between CM and EM CD8 + T cells. As expected, no overlap was identified between histone mark patterns of RNAlo genes and published RNA-seq gene sets (Supplementary Fig. 14C-D) – because RNAlo gene sets comprise genes with transcripts at or below the limit of detection that would not be seen in published RNA-seq analyses. However, investigating these RNAlo genes individually, we did identify differences in the patterns of histone marks between CM and EM subsets in known regulatory genes of T cell differentiation and effector function (e.g., LMO2, IDO2, IL23R, IL1R, SIGLEC7)22,23,24,25,26 (Supplementary Fig. 12B), as well as quiescence and memory states (e.g., GATA2, CDKN1C, KLF7)27,28,29 (Supplementary Fig. 12C). These data demonstrate that analyses of differences in patterns of histone marks between CD8 + T cell subsets identify genes associated with T cell differentiation that were not identified by RNA-seq alone or DEn of single histone marks.
Histone mark analyses identify biologically relevant differences between CAR-T manufactured from distinct sources that are not apparent by RNA-seq
Due to in vitro activation and expansion during CAR-T manufacture, the transcriptome of infused CAR-T products is dominated by activation- and cytotoxicity-associated genes that do not necessarily reflect genes governing cell fate11,30. Because we established that profiling of histone marks provides complementary and orthogonal information to RNA-seq, we next sought to use these approaches to investigate whether analysing histone marks could succeed in a challenging task: identification of differences between CAR-T products manufactured from similar, yet distinct, CD8 + T cell sources. This is clinically important, because the presence of less differentiated cells in leukaphereses or CAR-T infusion products has been associated with improved outcomes31,32,33. We manufactured CAR-T from naïve, CM or EM CD8 + T cells from HD, then isolated CAR-T by flow sorting for the transduction marker truncated epidermal growth factor receptor (EGFRt)2. PCA analyses of RNA-seq data did not distinguish naïve-, CM- and EM-derived CAR-T (Fig. 5A), consistent with the finding that few genes were DEx in pairwise comparisons between CAR-T manufactured from each subset (naïve vs CM; naïve vs EM; CM vs EM; Fig. 5B). In contrast, numerous DEn genes were identified in pairwise comparisons between CAR-T manufactured from each subset (Fig. 5B; Supplementary Data 3).
A PCA plots of RNA-seq, H3K27me3, or H3K4me2 of CAR-T manufactured from healthy donors (HD) CD8+ naïve-, central memory (CM)- or effector memory (EM)-derived CAR-T. B Venn diagram showing numbers of differentially expressed (DEx by RNA-seq) or differentially enriched (DEn by H3K4me2 or H3K27me3) genes when comparing naïve-, CM- and EM-derived CAR-T. C Volcano plot of -log(adjusted P value by Benjamini-Hochberg procedure) of transcript (left panel), H3K4me2 (middle panel) or H3K27me3 (right panel) vs logFC. DEx/DEn genes are denoted purple for expressed/enriched in CM-derived CAR-T, or green for expressed/enriched in EM-derived CAR-T. Labelled genes are those DEx from a recently published analysis comparing healthy donor ex vivo isolated CM and EM cells by RNA-seq15. D, E Mean scaled H3K4me2 (D) and H3K27me3 (E) +/- SD signal +1 of transcription factors (TF) genes differentially enriched in CM- and EM-derived CAR-T. Shown is adjusted P value by Benjamini-Hochberg procedure.
We focused on the differences between CM- and EM-derived CAR-T to enable subtle but potential, clinically important differences between similar subsets to emerge. Only three genes were DEx by RNA-seq between the EM- and CM-derived products (Fig. 5B, C, Supplementary Data 3). No changes in the numbers of DEx genes were found when we repeated the analyses with modified fold-change and P value thresholds (Supplementary Fig. 15). In contrast, histone marks distinguished CM- and EM-derived CAR-T, with 99 and 131 unique genes being DEn by single H3K4me2 and H3K27me3 analyses, respectively (Fig. 5B, C). Given that TFs are often expressed transiently and at low levels, we examined whether we could identify DEn histone marking among non-DEx TFs in CAR-T products. No DEx TFs were identified by RNA-seq, whereas two and seven TFs were identified by H3K4me2 and H3K27me3 histone mark analyses, respectively. All of these were concordantly validated by data from T cells in a manner consistent with EM-derived CAR-T displaying a more differentiated/exhausted phenotype than CM-derived CAR-T (Supplementary Data 4, Fig. 5D, E). For example, EM-derived CAR-T showed increased repressive H3K27me3 marks on LEF1 – a TF that can induce T cell quiescence/memory34–consistent with a more differentiated and less “stem-like” functionality of CAR-T derived from this subset.
We then assessed changes in H3K4me2 and H3K27me3 patterns when comparing CM- and EM-derived CAR-T (Fig. 6). Because only 3 genes were DEx by RNA-seq between subsets, we included all RNAhi and RNAlo genes in analyses to identify genes with histone mark patterns indicative of EM-derived (red box, Fig. 6A, B) and CM-derived CAR-T (white box, Fig. 6A, B). Gene ontology (GO) analysis of RNAhi genes showed pathways associated with cell migration and killing in EM-derived histone mark patterns (Fig. 6C) and immune and cytokine signalling within CM-derived histone mark patterns (Fig. 6D). Exhaustion-associated T cell surface marker genes (HAVCR2, CD160, CD96, CXCR4, CD55, KLRC1, KIR2DL4, TNFRSF935,36,37) were identified in multiple gene sets associated with an EM-derived histone mark pattern (WP4494, WP4298, GO:0031341, R-HSA-1280215, GO:0050778, R-HSA-6798695, GO:0002727) (Fig. 6C, Supplementary Data 5). Conversely, the gene encoding the apoptosis inhibitor Bcl-xL, BCL2L1, that increases therapeutic potency when overexpressed in CAR-T38, was identified within multiple gene sets (R-HSA-1280215, R-HSA-982446, GO:0051607, GO:0071214) associated with a CM-derived histone mark pattern (Fig. 6D, Supplementary Data 6).
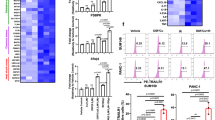
A, B Heatmaps of the numbers of genes within each histone mark pattern (H3K4me2 = K4m2; H3K27me3 = K27m3) when comparing central memory (CM)- and effector memory (EM)-derived CAR-T within RNAhi (A) and RNAlo (B) genes. C, D Comprehensive gene ontology analysis by Metascape.org of RNAhi genes with histone mark patterns indicative of EM-derived CAR-T (C, red box in A) or histone mark patterns indicative of CM-derived CAR-T (D, white box in A). E–H TRRUST analysis from Metascape.org denoting transcription factors and DNA binding proteins that regulate RNAhi (E & F) and RNAlo (G & H) genes within histone mark patterns indicative of EM-derived CAR-T (E & G) or CM-derived CAR-T (F & H). Displayed P value adjusted by Benjamini-Hochberg procedure.
We used TRRUST, a literature-curated reference database of transcriptional regulators and targets, to identify upstream regulators of genes that differ in histone mark patterns between CM- and EM-derived CAR-T. We found that RNAhi genes that show an EM-derived histone mark pattern were regulated by exhaustion-associated genes (NFKB1, RELA, MYC, HDAC329, 39,40,41) (Fig. 6E). In contrast, STAT1 and E2F1–transcriptional regulators that prevent T cell dysfunction and maintain T cell homoeostasis, respectively42,43–were upstream of genes associated with a CM-derived histone marks pattern (Fig. 6F). GO analyses of histone mark patterns in RNAlo genes showed pathways associated with development as opposed to T cell function (Supplementary Fig. 16, Supplementary Data 7 & 8); yet TRRUST analysis of these same genes revealed regulating factors known to impact T cell exhaustion and effector maturation (CEBPA, TAL1, RUNX1, CTCF44,45,46,47, Fig. 6G). RNAlo genes with a CM-derived histone mark pattern were associated with the regulator of lipid metabolism, SREBF2–a critical TF gene in memory T cell proliferation and metabolic programming48 (Fig. 6H).
To further challenge the capacity of histone mark analyses, we compared sorted CAR-T manufactured from two different starting populations of CM CD8 + T cells. One product was manufactured from CM cells sorted to high purity from HD. The other was manufactured from CM cells enriched by two-step CliniMACS immunomagnetic selection from large B cell lymphoma (LBCL) PTs on a phase 1 clinical trial (NCT01865617). Despite both CAR-T products being manufactured from CM cells, we expected the starting CM cells immunomagnetically enriched from LBCL PTs would be of lower purity but might also exhibit evidence of increased differentiation or exhaustion associated with malignancy and/or previous chemotherapy treatment. PCA using RNA-seq and histone marks distinguished patient CM-derived CAR-T from HD CM-derived CAR-T (Fig. 7A); however, H3K4me2 and H3K27me3 histone mark analyses identified 4.4-fold and 5.7-fold more DEn genes than RNA-seq (Fig. 7B, C, Supplementary Data 9), including more CAR-T exhaustion-associated TFs49. The DEn genes were histone marked in a manner concordant with patient CM-derived CAR-T displaying a more exhausted phenotype than HD CM-derived CAR-T (Fig. 7D–F). Together, these data show that analyses of histone marks, but not the transcriptome, uncovered key genes associated with T cell homoeostasis, function, metabolism, differentiation, and exhaustion that differed between CAR-T products manufactured from similar, yet distinct, sources.
A PCA plots of RNA-seq, H3K27me3 or H3K4me2 marks of CAR-T manufactured from healthy donor (HD) central memory (CM)-derived and patient (PT) CM-derived CD8 + T cells. B Numbers of differentially expressed (DEx by RNA-seq) or enriched (DEn by H3K4me2 or H3K27me3) genes when HD CM-derived CAR-T are compared to patient CM-derived CAR-T. C Volcano plot of -log(adjusted P value by Benjamini-Hochberg procedure) in transcript (left panel) or H3K4me2 (middle panel) or H3K27me3 (right panel) signal vs log(fold-change, FC). DEx/DEn genes are denoted purple for HD CM-derived CAR-T, or peach for patient CM-enriched CAR-T. Named genes are transcription factors (TFs) that are DEx/DEn in each analysis. D, E Mean scaled RNA transcript (D), H3K4me2 (E) and H3K27me3 (F) +/- SD signal +1 of exhaustion resistance-associated TF genes differentially expressed or enriched in patient and HD CM-derived CAR-T. Shown is adjusted P value by Benjamini-Hochberg procedure.
Identification of a novel transcription factor in CAR-T products that associates with robust in vivo CAR-T accumulation in LBCL patients
The capacity of histone mark analyses to differentiate CAR-T manufactured from distinct sources led us to ask if such analyses might uncover genes in IP that associate with outcomes of CAR-T infusion. We first embarked on analyses of IP from 30 LBCL patients (Supplementary Data 10) treated on a phase 1 clinical trial of CD19 CAR-T immunotherapy (NCT01865617). Transcriptomic and epigenomic analyses did not distinguish the IP of patients who had a complete response (CR) from those who did not have a CR (non-CR; Supplementary Fig. 17A). This was also the case when we compared IP from patients who received a CAR-T product manufactured from the same subset (CM-enriched CD8 + T cells; Supplementary Fig. 17B), likely due in part to the complexities of the LBCL tumour microenvironment (TME).
The in vivo CAR-T count in blood is a major determinant of clinical outcomes after CAR-T immunotherapy2; however, identification of factors in IP associated with in vivo CAR-T accumulation has been limited by the effects of activation and in vitro culture during manufacturing4,30. To minimise the confounding impact of the TME, we studied patients who achieved CR after CAR-T infusion; had similar pre-treatment tumour burden; received the same lymphodepletion (LD) regimen; received CAR-T manufactured from the same starting CM CD8 + T cell subset; and received the same CAR-T dose (n = 8, Supplementary Data 10). We compared IP from high expanders (maximal CD8 + CAR-T count in blood after infusion was above the median for the group) with low expanders (maximal CD8 + CAR-T count below the median) (Supplementary Fig. 18A). There were no differences between the cohorts in CD62L + CD45RA- composition in the IP; or in concentrations of serum LDH or cytokines associated with CAR-T accumulation in patients (Supplementary Fig. 18B-D).
PCA of histone marks, but not RNA-seq data, distinguished CAR-T IP from high and low expanding cohorts (Fig. 8A). DEn of H3K4me2 and H3K27me3 marks identified 30-fold and 10-fold more genes, respectively, than DEx genes by RNA-seq (Fig. 8B, C, Supplementary Data 11). Five DEn genes overlapped with previously identified genes from RNA-seq studies of CAR-T exhaustion and tumour-infiltrating lymphocytes (Supplementary Data 12). One of these genes, SLAMF7, has a known association with T cell exhaustion; it was H3K4me2 marked in low expanding CAR-T products50. These data demonstrate shared but not identical programming in CAR-T cell functions such as expansion and exhaustion identified by histone modification analyses when compared to those found by RNA-seq. We found two TFs – KLF7 and ZFP57 – were identified by histone mark analyses and none by RNA-seq (Fig. 8C, green boxes). In previous analyses (Fig. 5E & 7F), we found that Kruppel-like transcription factors, including KLF7, were associated with less-differentiated cell subsets and CAR-T products. KLF7 has also been associated with increased cell proliferation in other cell types51. When comparing high- and low-expanding infused CAR-T, and we found KLF7 was more H3K27me3-marked (transcriptionally repressed) in low expanding CAR-T IP (Fig. 8D). To further support a role for KLF7 in high-expanding CAR-T IP, we identified seven targets of KLF7 from Marbach et al.52. that were DEn in our histone mark analyses (Fig. 8E, F). All proposed targets of KLF7 showed an association with proliferation of various cell types from the literature in keeping with the directionality of association with CAR-T expansion in our analysis (Supplementary Data 13). In contrast, recently published analyses of highly expanding CAR T cell clonotypes by scRNA-seq in patients treated for B cell malignancies did not identify KLF7 or other transcription factors associated with expansion53.
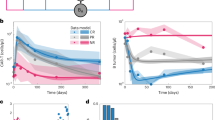
A PCA plots of RNA-seq, H3K27me3 or H3K4me2 histone marks of CAR-T manufactured from central memory (CM)-enriched CD8 + T cells from LBCL patients who achieved CR after CD19-targeted CAR-T immunotherapy (NCT01865617) and showed peak of expansion below the median (red, Low_Expander) or above the median (blue, High_Expander). B Number of differentially expressed (DEx, by limma, LogFC >1, adjusted P < 0.1 by Benjamini-Hochberg procedure) or enriched (DEn, by H3K4me2 and H3K27me3 histone marks, LogFC > 1, adjusted P < 0.1 by Benjamini-Hochberg procedure) genes when comparing Low_Expander and High_Expander. C Volcano plot of -log(adjusted P value by Benjamini-Hochberg procedure) in transcript (left panel) or H3K4me2 (middle panel) or H3K27me3 (right panel) signal vs logFC. Red points indicate genes enriched in low expanding CAR-T and blue points indicate genes enriched in high expanding CAR-T. D-F. Mean scaled KLF7 (D), H3K27me3 (E) and H3K4me2 (F) +/- SD signal +1 of KLF7 target genes in high- vs low-expanding CM-derived patient CAR-T. Shown is adjusted P value by Benjamini-Hochberg procedure. G Mean fold change +/- SD in accumulation of KLF7_P2A_GFP- and control P2A_GFP-lentiviral transduced T cells at increasing viral volumes. n = 5 donors; P values by two-way ANOVA with post-hoc Tukey correction. H Mean change +/- SD in geometric mean fluorescence intensity (geoMFI) of CellTrace Violet (CTV) as an inverse measure of CAR-T proliferation with and without KLF7 co-transduction when exposed to CD19-negative K562 cells. n = 2 donors in technical duplicate, P value by two-way ANOVA with post-hoc Tukey correction. I Mean frequency and 95% confidence intervals (dotted lines) of CD45RA- CCR7+ phenotype cells in CD19-directed CAR-T with and without KLF7 co-transduction. n = 3 donors performed as technical duplicate, P value by two-tailed t test. J Promoter region of IL2 in the human genome and predicted upstream KLF7 binding site. K Mean frequency and 95% confidence intervals (dotted lines) of IL-2 expressing CAR-T by intracellular cytokine staining with and without KLF7 co-transduction when exposed to CD19-negative K562 cells. n = 3 donors performed as technical duplicate, P value by two-tailed t test.
To determine whether KLF7 contributes to increased accumulation of CAR-T, we assessed cell counts, proliferation, phenotype, and cytokine secretion in T cells and/or CAR-T that were transduced to express GFP with or without KLF7. We found that KLF7-transduced T cells showed a dose-dependent increase in accumulation in culture compared to T cells transduced to only express GFP (Fig. 8G). We then transduced T cells to express a CD19-directed CAR with or without KLF7 and demonstrated enhanced proliferation of KLF7-transduced CAR-T (Fig. 8H). KLF7-dependent increased accumulation was only observed in the absence of stimulation through the CAR (Supplementary Fig. 19A). A higher percentage of CCR7 + CD45RA- CM phenotype cells were identified in KLF7 + CD8 + CAR-T, consistent with our finding of decreased KLF7 repression in CD8 + CM-derived CAR-T (Fig. 8I & 5E). Autocrine IL-2 production has been shown to enhance memory CD8 + T cell expansion54. To investigate the mechanism of enhanced proliferation in KLF7 + T cells, we interrogated the IL2 promoter and found a proposed KLF7 binding site (Fig. 8J). Validating this in silico finding, we found higher IL-2 production by CD8 + KLF7 + CAR-T that was only observed in the absence of stimulation through the CAR (Fig. 8K & Supplementary Fig. 19B). These data demonstrate that histone mark analyses identified a novel TF and epigenomic regulome in CAR-T IP that is associated with proliferative competence in T cells and CAR- T.
Discussion
We set out to expand our understanding of factors associated with T cell and CAR-T potential beyond what is afforded by transcriptomic approaches. To accomplish this, we employed the biology of the epigenome and focused on transcriptionally permissive H3K4me2 and transcriptionally repressive H3K27me3 marks, which are stable and heritable in T cell progeny and determinative of differentiation55. Our analyses show that single H3K4me2 and H3K27me3 marks, and changes in patterns of combined H3Kme2 and H3K27me3 marks, readily distinguish human naïve, CM and EM CD8 + T cells, and CAR-T manufactured using similar, yet distinct, starting CD8 + T cell populations. This study is focused on CD8 + T cells and CAR-T because of the association of CD8 + CAR-T counts with response to CAR-T immunotherapy2,56,57.
Analyses of histone marks were complementary to transcriptomic analyses. We identified DEn genes by histone marks that parse the identity of T cell subsets and CAR-T with different potentials. The fact that not all genes uncovered by histone mark analyses overlapped with those identified by RNA-seq demonstrates the orthogonal nature of epigenomic analyses12. This is likely because the transcriptome is not necessarily a heritable and transient composite output of histone modifications and methylation at any particular point in time58. Accordingly, the complementary and orthogonal view of T cell and CAR-T genomic activity inferred by histone mark analysis may be a consequence of therapeutic T cell manufacturing, in which activation-associated genes tend to be over-represented by transcriptional analyses11. Compared to transcripts, epigenomic marks are more stable in T cell progeny and not as subject to activation1. The identification of TFs is particularly demonstrative in this regard. Despite the importance in governing fate and function, transcription factors are classically lowly and transiently expressed, thus complicating identification by RNA-seq9,10. Assessing stable, heritable epigenomic binary marks in the cell circumvents the low dynamic range and ephemeral nature of TF transcripts. It follows that the narrow dynamic range of epigenomic data is less susceptible to the effects of highly expressed genes that overshadow the identification of genes with lower transcript abundance. Statistical noise afforded by high transcript dominance is not solved by single-cell RNA-seq (scRNA-seq); furthermore, the sparsity of transcript in scRNA-seq also hinders identification of key TFs, so much so that recent publications turn to pseudo-bulk approaches11,30. We are hopeful that future approaches may enable single-cell analysis of histone marks to further our understanding of CAR-T potential; however, these methods are also currently hampered by signal sparsity59.
Our analyses identified numerous genes associated with cell potential that differed between distinct CAR-T products. Most of these genes were uncovered using histone mark analyses, but not transcriptomic approaches, underscoring the power of histone mark analyses to overcome limitations of RNA-seq in identifying genes in activated cells. We progressively subjected the analytical platform to more challenging tasks, culminating in the identification of genes associated with T cell exhaustion and expansion in comparisons between CAR-T products that were manufactured from phenotypically similar or even identical sources. Previous studies have shown decreased in vivo efficacy of EM-derived CAR-T in comparison to N- and CM-derived CD8 + CAR-T cells5. Our RNA-seq data supports this observation by showing increased expression in EM-derived CAR-T cells of genes that prevent quiescence and stemness in T cells (e.g., LEF1-AS1)60. Our epigenetic studies both complement and expand upon transcriptomics by further highlighting the programmatic differences between subset-derived CAR-T cells that may not be detectable by RNA-seq due to the widespread transcriptional activity subsequent to activation and expansion during manufacturing.
Finally, we undertook an analysis of patient derived CAR-T cells. Our initial analysis did not show differences by either RNA-seq or CUT&RUN between patients who had a response versus no response. Similar outcomes have been noted after analyses of single cell RNA-seq data, in which only a small subset of genes and no cell clusters were associated with responses to CAR-T cell therapy in LBCL – likely a consequence of factors, such as the tumour microenvironment, that confound analyses61. To account for the complexities imposed by the tumour microenvironment, we focused on CM-derived CAR-T from patients who had achieved CR after CAR-T cell infusion. We identified H3K4me2 and H3K27me3 DEn in genes encoding the KLF7 TF and its targets that are concordantly associated with robust in vivo accumulation of CAR-T after infusion – a critical parameter for efficacy of CAR-T immunotherapy for B cell malignancies2. Our data showed epigenomic suppression of KLF7 in EM-derived CAR-T in comparison to CM-derived CAR-T, and in low proliferating products from CM-derived CAR-T. Together with previous studies showing associations of KLF7 with early murine thymocyte differentiation and naïve human T cells62,63, this posits a continuum of function of KLF7 in maintaining a less differentiated CD8 + T cell state64. Furthermore, the in silico predicted binding of KLF7 upstream of the IL-2 gene may also enable autocrine-driven expansion, thereby leading to higher CD8 + CAR-T counts in patients. Thus, transduction of KLF7 during manufacturing to generate less-differentiated CAR-T cells with enhanced proliferative potential is an attractive strategy that could be investigated to improve CAR-T efficacy. Chronic proliferation due to persistent antigen stimulation or tonic signalling through the CAR can induce CAR-T exhaustion; however, the mechanisms by which KLF7 support CAR-T proliferation are likely different from those induced by chronic signalling through the CAR, the latter often being a consequence of CAR design65,66. More studies are needed to determine if KLF7 augmentation can improve CAR-T proliferation and durable survival without loss of function.
The reported analytical platform is a high throughput, high sensitivity, high efficiency approach to identifying differences in T cell and CAR-T products that can be performed with few cells–an important problem in the world of limited clinical samples. This approach opens a suite of possibilities to compare differences in CAR designs, CAR-T manufacturing methods, and treatment regimens, all of which may have heritable effects on CAR-T progeny. It follows that data from analyses of histone marks in T cells and CAR-T will open a rich vein of potential genes for further investigation to modify to improve outcomes of CAR-T immunotherapy.
Methods
Study design
All studies were approved by the Institutional Review Board of the Fred Hutchinson Cancer Centre and the University of Sydney Human Research Ethics Committee. All participants provided written consent. Sample size was determined by known variation in CUT&RUN and RNA-seq of healthy donor T cells. All data was included except biological replicates that failed QC assessments. Sex/gender was self-reported. There was no bias in sex/gender inclusion in the reported clinical data.
Flow cytometry and sorting
Cryopreserved healthy human donor peripheral blood mononuclear cells were thawed, negatively selected using the CD8 + T cell isolation kit, human (Miltenyi Biotec) and sorted into CD8 + T cell subsets using the antibodies in Supplementary Data 14 with gating as outlined in Supplementary Fig. 1A on a BD FACSAria. Flow cytometry was performed using antibodies listed in Supplementary Data 14 at manufacturers’ recommendations on a BD FACSymphony or Cytek Aurora (for KLF7 studies) and analysed on FlowJo. CAR-T were similarly thawed and sorted by truncated EGFRt.
CAR-T manufacturing and retroviral transduction
patient CM-derived CAR-T were manufactured from CM-enriched CD8 + T cells isolated from PTs participating in a phase 1/2 clinical trial, as described previously2. HD CM- and EM-derived CAR-T were manufactured from CM or EM CD8 + T cells isolated from apheresis products, as described above, then stimulated and transduced as previously described2. Human TM-LCL and parental K562 cell lines and K62 transduced with CD19 used for generation and testing of CAR-T cells were a gift from the Riddell Laboratory at the Fred Hutchinson Cancer Center.
CUT&RUN assay
CUT&RUN was performed as previously described13. Briefly, live sorted cells were incubated with concavalin A beads, then permeabilized and incubated overnight at 4 C with anti-H3K27me3, anti-H3K4me2 or IgG isotype control with rabbit-anti-mouse as an adaptor (Supplementary Data 14). Samples were then processed on a Beckman Biomek FX liquid-handling robot as described13. Briefly, pA-MNAse is added to samples, DNA cleaved and released, and chromatin targets purified, followed by adaptor ligation, Ampure cleanup and PCR amplification of libraries. QC of amplified DNA was assessed using an Agilent 4200 TapeStation and quantified on a QuBit Fluoremeter (ThermoFisher). Samples were then pooled and submitted for paired-end sequencing on an Illumina HiSeq 2500.
CAR-T counts
CAR-T peak expansion was quantified as the highest absolute CD8 + CAR-T count, determined as the absolute lymphocyte count multiplied by the percentage of CD8+ EGFRt+ cells in a CD45+ vs SSC lymphocyte gate as described67.
CUT&RUN and RNA-seq data processing and analysis
Supplementary Data 16 lists packages and references used for analysis of differential detection, gene ontology and geneset enrichment analysis. CUT&RUN data processing and analysis follow the pipeline published in detail on protocols.io (https://www.protocols.io/view/cut-amp-tag-data-processing-and-analysis-tutorial-e6nvw93x7gmk/v1) and as summarised in Supplementary Fig. 2. Peaks were called using SEACR and we defined epigenomic enriched peaks simultaneously following two criteria: peaks that are among the top 10% enriched peak regions and peaks identified above background noise (profiled by IgG isotype control). To generate the heatmap and summary plot of histone modifications at each TSS region (from 1 kb downstream to 1 kb upstream), the countMatrix and plotHeatmap functions from deepTools version 3.3.1 was used to sum the total signals. RNA-seq raw data that passed the FASTQC quality check were processed by RSEM using STAR as the aligner. The expected count is used as the gene expression raw count. Log-transformed count per million (logCPM) was used to avoid sequencing depth biases unless otherwise specified. UCSC genome browser is employed to visualise the epigenomic landscape around the genes or regions of interest. When integrating RNA-seq and CUT&RUN gene lists, a single DEx or DEn was counted only once by selecting the transcript or peak with the highest logFC. KLF7 binding sequence was derived from MEME suite.
KLF7 transduction and functional assays
T cells were immunomagnetically selected using the Pan T cell isolation kit (Miltenyi) as per the manufacturer’s instructions, activated with Transact (Miltenyi), and transduced with different volumes of an epHIV7 lentiviral vector containing either the KLF7 open reading frame (NM_001270942) with a ribosomal skip sequence and GFP (KLF7_P2A_GFP) or control P2A_GFP. The cell counts of KLF7_P2A_GFP- and P2A_GFP-transduced cells were quantitated with acridine orange/propidium iodine dyes counted on a Cellaca PLX instrument (Revvity). KLF7 and GFP transcripts were quantitated by RT-qPCR relative to the housekeeping gene ACTB, using primers listed in Supplementary Data 15. Fold expansions of KLF7_P2A_GFP- and P2A_GFP-transduced cells were calculated by normalising to GFP transcript level. For functional CAR-T assays, T cells were transduced with lentivirus encoding a CD19-directed CAR with or without KLF7_P2A_GFP. Transduced cells were then sorted on co-expression of GFP and a CAR transduction marker (EGFRt) to high purity and then cultured for 10 days. Immunophenotyping of CD45RA, CD45RO and CCR7 was performed as previously described68. To assess the effects of KLF7 transduction on functions of CAR-T cells with and without antigen stimulation, untransduced, control P2A_GFP-transduced, and KLF7_P2A_GFP-transduced CAR-T cells were co-cultured with CD19-transduced K562 cells (K19), control K562 cells, or with media alone. CAR-T cell proliferation was assessed by Cell Trace Violet dilution at 48, 72 and 96 h after antigen engagement. Intracellular IL-2 was assessed 24 h after antigen-engagement by flow cytometry, as previously described2,69
Statistical analyses
Supplementary Data 16 includes statistical packages and references used for analysis. Data processing and analysis were conducted in R, Python and Linux. The Limma R package was used for differential detection analyses for both CUT&RUN and RNA-seq data, HD or PTs were considered as random effect variables to remove donor effect. Benjamini-Hochberg procedure was used to control for false discovery rate70. Principal Component Analyses (PCA) and Uniform Manifold Approximation and Projections (UMAP) provided visualisations of high-dimensional data in two-dimensional space. Two-component GMM was used to characterise the distribution of reads mapping in a 2 kb window surrounding the TSS for each gene and was fitted using the normalmixEM function from mixtools library in R. Distinct thresholds at 4.5 for H3K4me2 and 2.5 for H3K27me3 were inferred from the intersection point of the GMM distribution and separated genes into positive and negative groups per histone marker. All statistical tests were two-sided unless otherwise specified.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Source data that was used to generate the corresponding figures and Supplementary Figs. are provided as Source Data files. All raw data and workflow are available as an open-source resource. The raw sequencing data and the processed normalised bedGraph files generated in this study have been deposited in the NCBI GEO database under https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE254807 the CUT&RUN data and for the https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE254808 for the matching RNA-seq data. Both are available for the public without any restrictions. Source data are provided with this paper.
Code availability
CUT&RUN data processing and analysis is detailed on protocols.io available from the following address https://www.protocols.io/view/cut-amp-tag-data-processing-and-analysis-tutorial-e6nvw93x7gmk/v1.
References
Henning, A. N., Roychoudhuri, R. & Restifo, N. P. Epigenetic control of CD8(+) T cell differentiation. Nat. Rev. Immunol. 18, 340–356 (2018).
Gauthier, J. et al. Feasibility and efficacy of CD19-targeted CAR T cells with concurrent ibrutinib for CLL after ibrutinib failure. Blood 135, 1650–1660 (2020).
Chen, G. M. et al. Integrative bulk and single-cell profiling of premanufacture T-cell populations reveals factors mediating long-term persistence of CAR T-cell therapy. Cancer Discov. 11, 2186–2199 (2021).
Fraietta, J. A. et al. Determinants of response and resistance to CD19 chimeric antigen receptor (CAR) T cell therapy of chronic lymphocytic leukemia. Nat. Med. 24, 563–571 (2018).
Sommermeyer, D. et al. Chimeric antigen receptor-modified T cells derived from defined CD8+ and CD4+ subsets confer superior antitumor reactivity in vivo. Leukemia 30, 492–500 (2016).
Russ, B. E. et al. Regulation of H3K4me3 at Transcriptional enhancers characterizes acquisition of virus-specific CD8(+) T cell-lineage-specific function. Cell Rep. 21, 3624–3636 (2017).
Sheerin, D., O’Connor, D., Pollard, A.J. & Mohorianu I. Effects of technical noise on bulk RNA-seq differential gene expression inference. Preprint at bioRxiv https://doi.org/10.1101/843789 (2019).
Haradhvala, N. J. et al. Distinct cellular dynamics associated with response to CAR-T therapy for refractory B cell lymphoma. Nat. Med 28, 1848–1859 (2022).
Vallejo, A. F. et al. snPATHO-seq: unlocking the FFPE archives for single nucleus RNA profiling. Preprint at bioRxiv, https://doi.org/10.1101/2022.08.23.505054 (2022).
Weinreb, C., Rodriguez-Fraticelli, A., Camargo, F. D. & Klein, A. M. Lineage tracing on transcriptional landscapes links state to fate during differentiation. Science 367, eaaw3381 (2020).
Sheih, A. et al. Clonal kinetics and single-cell transcriptional profiling of CAR-T cells in patients undergoing CD19 CAR-T immunotherapy. Nat. Commun. 11, 219 (2020).
Roels, J. et al. Distinct and temporary-restricted epigenetic mechanisms regulate human alphabeta and gammadelta T cell development. Nat. Immunol. 21, 1280–1292 (2020).
Janssens, D. H. et al. Automated CUT&Tag profiling of chromatin heterogeneity in mixed-lineage leukemia. Nat. Genet. 53, 1586–1596 (2021).
Meyer, C. A. & Liu, X. S. Identifying and mitigating bias in next-generation sequencing methods for chromatin biology. Nat. Rev. Genet. 15, 709–721 (2014).
Rose, J. R. et al. Distinct transcriptomic and epigenomic modalities underpin human memory T cell subsets and their activation potential. Commun. Biol. 6, 363 (2023).
Chan, J. D. et al. FOXO1 enhances CAR T cell stemness, metabolic fitness and efficacy. Nature 629, 201–210 (2024).
Doan, A. E. et al. FOXO1 is a master regulator of memory programming in CAR T cells. Nature 629, 211–218 (2024).
Wei, H. et al. Cutting edge: Foxp1 controls naive CD8+ T cell quiescence by simultaneously repressing key pathways in cellular metabolism and cell cycle progression. J. Immunol. 196, 3537–3541 (2016).
Jung, I. Y. et al. BLIMP1 and NR4A3 transcription factors reciprocally regulate antitumor CAR T cell stemness and exhaustion. Sci. Transl. Med 14, eabn7336 (2022).
Akondy, R. S. et al. Origin and differentiation of human memory CD8 T cells after vaccination. Nature 552, 362–367 (2017).
Giles, J. R. et al. Human epigenetic and transcriptional T cell differentiation atlas for identifying functional T cell-specific enhancers. Immunity 55, 557–574.e557 (2022).
Wiekmeijer, A. S. et al. Overexpression of LMO2 causes aberrant human T-Cell development in vivo by three potentially distinct cellular mechanisms. Exp. Hematol. 44, 838–849.e839 (2016).
Li, Y., Wang, H., Lu, H. & Hua, S. Regulation of Memory T Cells by Interleukin-23. Int. Arch. Allergy Immunol. 169, 157–162 (2016).
Qin, R. et al. Tryptophan potentiates CD8(+) T cells against cancer cells by TRIP12 tryptophanylation and surface PD-1 downregulation. J. Immunother. Cancer 9, e002840 (2021).
Jain, A., Song, R., Wakeland, E. K. & Pasare, C. T cell-intrinsic IL-1R signaling licenses effector cytokine production by memory CD4 T cells. Nat. Commun. 9, 3185 (2018).
Haas, Q. et al. Siglec-7 represents a glyco-immune checkpoint for non-exhausted effector memory CD8+ T cells with high functional and metabolic capacities. Front. Immunol. 13, 996746 (2022).
Rowell, E. A. & Wells, A. D. The role of cyclin-dependent kinases in T-cell development, proliferation, and function. Crit. Rev. Immunol. 26, 189–212 (2006).
Ruiz-García, R. et al. Acquired senescent T-cell phenotype correlates with clinical severity in GATA binding protein 2-deficient patients. Front Immunol. 8, 802 (2017).
Lu, C. et al. NF-kB1 induces T cell dysfunction to promote colon tumorigenesis. J. Immunol. 204, 165.132 (2020).
Deng, Q. et al. Characteristics of anti-CD19 CAR T cell infusion products associated with efficacy and toxicity in patients with large B cell lymphomas. Nat. Med 26, 1878–1887 (2020).
Dickinson, M. J. et al. A novel autologous CAR-T THERapy, YTB323, with preserved T-cell stemness shows enhanced CAR T-cell efficacy in preclinical and early clinical development. Cancer Discov. 13, 1982–1997 (2023).
Filosto, S. et al. Product attributes of CAR T-cell therapy differentially associate with efficacy and toxicity in second-line large B-cell lymphoma (ZUMA-7). Blood Cancer Discov. 5, 21–33 (2024).
Arcangeli, S. et al. CAR T cell manufacturing from naive/stem memory T lymphocytes enhances antitumor responses while curtailing cytokine release syndrome. J. Clin. Invest 132, e150807 (2022).
Shan, Q. et al. Tcf1 and Lef1 provide constant supervision to mature CD8(+) T cell identity and function by organizing genomic architecture. Nat. Commun. 12, 5863 (2021).
Ramonell, K. M. et al. CXCR4 blockade decreases CD4+ T cell exhaustion and improves survival in a murine model of polymicrobial sepsis. PLoS One 12, e0188882 (2017).
Detsika, M. G. et al. Upregulation of CD55 complement regulator in distinct PBMC subpopulations of COVID-19 patients is associated with suppression of interferon responses. preprint available at bioRxiv. https://doi.org/10.1101/2022.10.07.510750 (2022).
Chen, X. et al. Differential expression of inhibitory receptor NKG2A distinguishes disease-specific exhausted CD8(+) T cells. MedComm (2020) 3, e111 (2022).
Maity, R. et al. A BCL2L1 armoured BCMA targeting CAR T cell to overcome exhaustion and enhance persistence in multiple myeloma. Blood 138, 327–327 (2021).
Krishna, S. et al. Chronic activation of the kinase IKKbeta impairs T cell function and survival. J. Immunol. 189, 1209–1219 (2012).
Toumi, R., Deo, R., Bhise, S., Sarkar, S. & Kalia, V. Exploring the role of c-Myc in defining the fate of exhausted CD8 T cells. J. Immunol. 208, 182.132–182.132 (2022).
McCaw, T. R. et al. Histone deacetylase inhibition promotes intratumoral CD8(+) T-cell responses, sensitizing murine breast tumors to anti-PD1. Cancer Immunol. Immunother. 68, 2081–2094 (2019).
DeRyckere, D. & DeGregori, J. E2F1 and E2F2 are differentially required for homeostasis-driven and antigen-induced T cell proliferation in vivo. J. Immunol. 175, 647–655 (2005).
Ryan, N. et al. STAT1 inhibits T-cell exhaustion and myeloid derived suppressor cell accumulation to promote antitumor immune responses in head and neck squamous cell carcinoma. Int J. Cancer 146, 1717–1729 (2020).
Tanaka, S. et al. CCAAT/enhancer-binding protein alpha negatively regulates IFN-gamma expression in T cells. J. Immunol. 193, 6152–6160 (2014).
Liang, C., Huang, S., Zhao, Y., Chen, S. & Li, Y. TOX as a potential target for immunotherapy in lymphocytic malignancies. Biomark. Res 9, 20 (2021).
Liu, J. et al. CTCF mediates CD8+ effector differentiation through dynamic redistribution and genomic reorganization. J. Exp. Med. 220, e20221288 (2023).
Hsu, F. C. et al. An essential role for the transcription factor Runx1 in T cell maturation. Sci. Rep. 6, 23533 (2016).
Kidani, Y. et al. Sterol regulatory element-binding proteins are essential for the metabolic programming of effector T cells and adaptive immunity. Nat. Immunol. 14, 489–499 (2013).
Good, C. R. et al. An NK-like CAR T cell transition in CAR T cell dysfunction. Cell 184, 6081–6100 e6026 (2021).
Awwad, M. H. S. et al. Selective elimination of immunosuppressive T cells in patients with multiple myeloma. Leukemia 35, 2602–2615 (2021).
Jia, Z. et al. KLF7 promotes preadipocyte proliferation via activation of the Akt signaling pathway by Cis-regulating CDKN3. Acta Biochim. Biophys. Sin. (Shanghai) 54, 1486–1496 (2022).
Marbach, D. et al. Tissue-specific regulatory circuits reveal variable modular perturbations across complex diseases. Nat. Methods 13, 366–370 (2016).
Wilson, T. L. et al. Common trajectories of highly effective CD19-Specific CAR T cells identified by endogenous T-cell receptor lineages. Cancer Discov. 12, 2098–2119 (2022).
Toumi, R. et al. Autocrine and paracrine IL-2 signals collaborate to regulate distinct phases of CD8 T cell memory. Cell Rep. 39, 110632 (2022).
Araki, Y. et al. Genome-wide analysis of histone methylation reveals chromatin state-based regulation of gene transcription and function of memory CD8+ T cells. Immunity 30, 912–925 (2009).
Galli, E. et al. The CD4/CD8 ratio of infused CD19-CAR-T is a prognostic factor for efficacy and toxicity. Br. J. Haematol. 203, 564–570 (2023).
Locke, F. L. et al. Tumor burden, inflammation, and product attributes determine outcomes of axicabtagene ciloleucel in large B-cell lymphoma. Blood Adv. 4, 4898–4911 (2020).
Talbert, P. B. & Henikoff, S. The Yin and Yang of histone marks in transcription. Annu. Rev. Genomics Hum. Genet 22, 147–170 (2021).
Stuart, T. et al. Nanobody-tethered transposition enables multifactorial chromatin profiling at single-cell resolution. Nat. Biotechnol. 41, 806–812 (2023).
Zheng, C. et al. Landscape of infiltrating T cells in liver cancer revealed by single-cell sequencing. Cell 169, 1342–1356.e1316 (2017).
Li, X. et al. A single-cell atlas of CD19 chimeric antigen receptor T cells. Cancer Cell 41, 1835–1837 (2023).
Mingueneau, M. et al. The transcriptional landscape of alphabeta T cell differentiation. Nat. Immunol. 14, 619–632 (2013).
Willinger, T. et al. Human naive CD8 T cells down-regulate expression of the WNT pathway transcription factors lymphoid enhancer binding factor 1 and transcription factor 7 (T Cell Factor-1) following antigen encounter in vitro and in vivo1. J. Immunol. 176, 1439–1446 (2006).
Geginat, J., Lanzavecchia, A. & Sallusto, F. Proliferation and differentiation potential of human CD8+ memory T-cell subsets in response to antigen or homeostatic cytokines. Blood 101, 4260–4266 (2003).
Selli, M. E. et al. Costimulatory domains direct distinct fates of CAR-driven T-cell dysfunction. Blood 141, 3153–3165 (2023).
Qiu, S. et al. CAR-Toner: an AI-driven approach for CAR tonic signaling prediction and optimization. Cell Res. 34, 386–388 (2024).
Gauthier, J. et al. Factors associated with outcomes after a second CD19-targeted CAR T-cell infusion for refractory B-cell malignancies. Blood 137, 323–335 (2021).
Hirayama, A. V. et al. A novel polymer-conjugated Human IL-15 improves efficacy of CD19-targeted CAR-T cell immunotherapy. Blood Adv. 7, 2479–2493 (2023).
Lynn, R. C. et al. c-Jun overexpression in CAR T cells induces exhaustion resistance. Nature 576, 293–300 (2019).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B (Methodol.) 57, 289–300 (1995).
Acknowledgements
The authors thank the staff of the Fred Hutchinson Cancer Center (Fred Hutch) Cell Processing Facility, Fred Hutch Cell Therapy Laboratory, the Fred Hutch Integrated Immunotherapy Research Center, and the Fred Hutch Bezos Family Immunotherapy Clinic. The authors would also like to thank Matthew Fitzgibbon, Dr Laila Hanafi, Reed Hawkins and Rachel Steinmetz for technical assistance. This work was supported in part by the Fred Hutch Immunotherapy Integrated Research Center, the CLEARBridge Foundation and a sponsored research agreement with Juno Therapeutics, a BMS Company. This work was also supported by the U.S. National Institutes of Health grant, HG012797, to Y.Z; the Haematology Society of Australia and New Zealand Clinical Fellowship to S.F., and a Research Training Programme Stipend from the Australian Government Department of Education to T.B.
Author information
Authors and Affiliations
Contributions
S.F. co-wrote and edited the manuscript; performed the experiments; curated, formally analysed and visualised the data, undertook project administration; Y.Z. co-wrote and edited the manuscript; curated, formally analyzed and visualised the data; J.P., T.J.B., A.S.S., D.K., and T.D.P performed the experiments; J.S. edited the manuscript; undertook formal data analysis; supervised and conceptualised the project; D.H.J. edited the manuscript and supervised the project; E.L.K. edited the manuscript; A.V.H. and J.G. curated clinical data; S.R.R. and D.G.M. administered the project; acquired funding; curated clinical data and supervised the project; Q.W., R.G., and J.Y.H.Y. supervised formal data analysis; S.H. edited the manuscript; oversaw analysis; and acquired funding and resources; C.J.T. co-wrote and edited the manuscript; conceptualized the project; acquired funding and resources; oversaw analysis; curated clinical data; and supervised the project.
Corresponding author
Ethics declarations
Competing interests
S.F. has filed and received license fees on patents for optimising CAR T cell function, has received research laboratory grants from Bristol Myers Squibb, and has consulted ad hoc for Prescient Therapeutics. E.L.K. receives research support from Juno Therapeutics, a BMS company. A.S.S. is a current employee of Umoja Biopharma. A.V.H. has received research funding from Juno Therapeutics, a Bristol Myers Squibb Company, and Nektar Therapeutics; it has received honoraria from Bristol Myers Squibb. J.G. reports honoraria from EUSA Pharma, JMP, Larvol, and Multerra Bio; serves on scientific advisory boards for Legend Biotech, Janssen, Kite (a Gilead company), and MorphoSys; and receives research funding from Sobi, Juno Therapeutics, Celgene (a BMS company), and Angiocrine Bioscience. S.R.R. is a cofounder and adviser to Lyell Immunopharma; has research funding from and intellectual property licensed to Lyell Immunopharma; was a cofounder of Juno Therapeutics; is an inventor of patents licensed to Juno Therapeutics; and served as an adviser to Juno Therapeutics and Adaptive Biotechnologies. D.G.M. has received research funding from Juno Therapeutics, a Bristol Myers Squibb Company, Celgene, and Kite Pharma, a Gilead company; has served on ad hoc advisory board meetings from Amgen, BMS, Genentech, Gilead, Incyte, Janssen, Legend Biotech, Mustang Bio, MorphoSys, Novartis, Pharmacyclics, and Umoja; has rights to receive royalties from Fred Hutch for patents licensed to Juno Therapeutics; serves on scientific advisory board with stock options and compensations for A2 Biotherapeutics and Navan Technologies. S.H. has filed patent applications on related work. C.J.T. received research funding from Juno Therapeutics/BMS, Nektar Therapeutics, and Nanostring; serves on Scientific Advisory Boards for Caribou Biosciences, T-CURX, Myeloid Therapeutics, ArsenalBio, Cargo Therapeutics, Celgene/BMS Cell Therapy, Differentia Bio, eGlint, and Advesya; is a DSMB member for Kyverna; has served on ad hoc advisory roles/consulting (last 12 months) for Prescient Therapeutics, Century Therapeutics, IGM Biosciences, Abbvie, Boxer Capital, Novartis; holds stock options for Eureka Therapeutics, Caribou Biosciences, Myeloid Therapeutics, ArsenalBio, Cargo Therapeutics, and eGlint; and has the right to receive payment from Fred Hutch Cancer Center as an inventor on patents related to CAR T-cell therapy. The remaining authors declare no competing interest.
Peer review
Peer review information
Nature Communications thanks Dorina Avram and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Fiorenza, S., Zheng, Y., Purushe, J. et al. Histone marks identify novel transcription factors that parse CAR-T subset-of-origin, clinical potential and expansion. Nat Commun 15, 8309 (2024). https://doi.org/10.1038/s41467-024-52503-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-024-52503-2
This article is cited by
-
Outcome correlates of approved CD19-targeted CAR T cells for large B cell lymphoma
Nature Reviews Clinical Oncology (2025)