Abstract
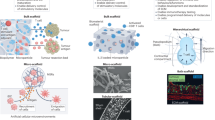
Immunotherapies are beneficial for a considerable proportion of cancer patients, but ineffective in others. In vitro modelling of the complex interactions between cancer cells and their microenvironment could provide a path to understanding immune therapy sensitivity and resistance. Here we develop MIRO, a fully humanised in vitro platform to model the spatial organisation of the tumour/stroma interface and its interaction with immune cells. We find that stromal barriers are associated with immune exclusion and protect cancer cells from antibody-dependent cellular cytotoxicity, elicited by targeted therapy. We demonstrate that IL2-driven immunomodulation increases immune cell velocity and spreading to overcome stromal immunosuppression and restores anti-cancer response in refractory tumours. Collectively, our study underscores the translational value of MIRO as a powerful tool for exploring how the spatial organisation of the tumour microenvironment shapes the immune landscape and influences the responses to immunomodulating therapies.
Similar content being viewed by others
Introduction
The presence of tumour-infiltrating lymphocytes (TIL) plays a determinant role in cancer progression1, and abundant infiltration of Natural Killer (NK) cells correlates with better patient outcomes2. In this context, immunotherapies designed to enhance immunity against cancer cells have considerably improved patient treatment and yielded durable therapeutic responses in many cancer subtypes3. Yet only 20–40% of patients respond to immune-modulating therapies4.
Growing evidence underscores the fundamental role of the tumour microenvironment (TME) in enhancing immunosuppressive signalling and interfering with cytotoxic immune cell infiltration inside the tumour mass5,6. The TME is a complex array of cellular and non-cellular components that include cancer-associated fibroblasts (CAF), blood vessels, immune cells, and the extracellular matrix (ECM), interacting chemically and physically with the cancer cells. CAFs are a major component of the TME in many solid tumours, supporting cancer cell survival, therapy resistance as well as metastatic dissemination7,8. CAFs can acquire a pro-inflammatory phenotype characterised by the production of immunomodulatory molecules and chemokines (e.g. TGF-β, PD-L1) that promote an immunosuppressive environment9. In addition, CAFs produce large quantities of collagens and fibronectins that organise into a highly dense meshwork constituting the ECM10. The density, alignment, and composition of the TME surrounding the tumour can dictate favourable local regions to initiate tumour invasion but also hinder immune cells and molecules from infiltrating the tumour, thus limiting the efficacy of anti-cancer therapies11,12,13. The high density of collagens in the tumour surrounding has been shown to reduce the ability of T-cells to penetrate the tumour, thus supporting the notion of stromal cells acting as a physical barrier against immune infiltration14,15,16,17. Yet, the impact of TME composition and organisation on immune cell trafficking in the tumour remains largely unexplored.
The development of bioengineered models allowed significant breakthroughs in understanding the impact of TME physical properties, including matrix architecture, stiffness, and mechanical plasticity, on immune cell motility8,18,19,20. In this line, important technical and conceptual efforts are currently being made to improve three-dimensional (3D) models for pre-clinical drug testing or for basic research purposes21. However, the extensive use of collagen originating from rat tails, Matrigel from mouse sarcoma cells or hydrogels made of synthetic polymers strongly reduces the physiological relevance of these assays22. Alternatively, advances in microengineering applied to biology have led to the development of microfluidic (on-chip) platforms allowing the in vitro replication of the complex cell organisations found within tissues and organs23. In oncology, microfluidics has been applied to the study of cell confinement and multicellular arrangement during tumour angiogenesis24,25 and metastasis26,27. More recently, microfluidic models enabled the monitoring of cancer-immune cell interaction and the evaluation of immunotherapy efficiency28,29,30,31. Yet, current approaches based on microscale chips often fall short in emulating human tumour spatial organisation and the multicellular interactions of the TME. High-resolution tracking of diverse cell populations and precise multicellular structure manipulation within a single platform present significant technical obstacles, particularly in 3D multicellular models22. Overcoming these challenges will contribute to obtaining more accurate on-chip tumour models, ultimately advancing our understanding of cancer and improving treatment strategies32.
To overcome some of these limitations, we develop MIRO (Micro Immune Response On-chip), a fully humanised immunocompetent model of TME of HER2+ breast cancer. In MIRO, cell co-culture is optimised in 2.5D to generate a well-defined cancer/stroma interface, where CAFs, ECM and immune cells adopt a spatial distribution frequently observed in solid tumours. Dynamic imaging of immune cell trafficking in MIRO reveals that the presence of a stromal barrier plays a critical role in immune cell guidance and exclusion from the tumour. As a proof of concept, we use MIRO to show that the anti-cancer immunity induced by anti-HER2 mAbs (monoclonal antibodies; Trastuzumab) can be restored in resistant/immune-excluded HER2+ breast cancer by IL2 administration. Our results indicate that the significant gain in efficacy obtained from the combination of anti-HER2 mAbs and IL2 is mainly driven by changes in immune cell motility and cytotoxic capacity, rather than by the modulation of the stromal barrier permeability.
Results
On-chip reconstruction of the spatial organisation of a cancer/stroma boundary
MIRO is an open fluidic device that allows the self-organisation of cancer cell spheroids, CAFs, and ECM deposition. The co-culture in the inlet of MIRO enables the establishment of cancer cell/CAF boundaries, while the channels connecting the inlet and the outlet mimic the migration route of immune cells through the CAFs and ECM in the stroma before reaching the tumour/stroma interface. MIRO is based on a microfabricated PDMS (polydimethylsiloxane) hollow cylinder (inner radius = 3 mm; outer radius = 5 mm) created by soft lithography and comprising an array of 30 radial channels (200 μm width, 50 μm height) that connect inner and outer compartments (Fig. 1a). The device was sealed onto a glass-bottom dish through plasma treatment. GFP-labelled and immortalised CAFs from human breast tumours (CAF#1-GFP) were cultured in the inner compartment until fully populating the inner surface and the channels connecting to the outer compartment. mCherry-labelled HER2+ breast cancer cell (BCC; HCC1954) spheroids were then seeded on top of the CAF layer at the proximity of the channel entry (Fig. 1b). During the following 48 h, BCC spheroids seeded on the CAF monolayer self-organised to generate a 2.5D interface (EDGE) segregating cancer cell compartments (IN) from the stromal compartment (EDGE + OUT) (Fig. 1c–g). Immune infiltration through the channels could then be monitored by fluorescence live confocal microscopy starting from 24 to 72 h after the addition of immune cells in the outlet (corresponding to day 7 and day 10 after CAFs seeding) (Fig. 1b). By comparing the cellular architecture in MIRO and in patient tumour samples by immunodetection of CAFs (α-SMA, FAP), cancer (E-Cadherin) and immune cells (CD45) we show that the cellular organisation in MIRO shares similarities with breast tumour samples (Fig. 1c, d and Supplementary Fig. 1).
a Illustration of the MIRO device composed of a 2 mm thick wall PDMS ring with a central 6 mm hole (inlet) and 30 channels (length: 2 mm, width: 200 μm, height: 50 μm) that connect the outer to the inner compartment. b Schematic of the co-culture timeline in MIRO. At day 0, CAFs are seeded in the inlet. At day 4, (HCC1954-mCherry red) BCC spheroids are seeded on the CAF layer. At day 6, immune cells are seeded in the outer compartment and imaging is performed at day 7 or day 10. c Left: MIRO co-culture on day 10 showing CAFs (green), tumour spheroid (visualised with DAPI, red) and immune cells (cyan). Scale bar: 2 mm. Right: Representative E-Cadherin (red), CD45 (blue) and alpha-SMA (green) immunofluorescence in tissue sections from a BC patient. Scale bar: 100 μm. d Left: Cellular organisation at the tumour/stroma barrier in MIRO CAF#1-GFP (green), BCC spheroid (red), immune cells (cyan). Scale bar: 100 μm. Right: zoomed areas relative to (c) (dotted white squares), scale bar: 25 μm. e 3D schematic representation of the cellular organisation in MIRO. f 3D rendering reconstituted from z-stack confocal images CAF#1-GFP (green), BCC spheroid, immune cells (blue). Scale bars: y-axis: 30 μm, z-axis: 30 μm. g Average fluorescence intensity profile of CAF#1-GFP (green), BCC spheroid (red) and immune cells (cyan) measured over z-stack image. h Relative CAF#1 fluorescent intensity profile and orientation displaying the EDGE, IN and OUT region delimitation (black dashed lines) relative to the tumour boundary (red dashed line). Lines indicate mean values, coloured areas SD (profile averaged on z-projection of n = 55 images, from at least 25 spheroids, 6 independent experiments). i Left: Representative fluorescence image of CAF#1-GFP distribution at the tumour/stroma boundary. Scale bar: 100 μm. Right: Representative map of the relative orientation of the CAFs meshwork (CAF#1) respective to the tumour/stroma boundary. The EDGE region is marked with red and black dashed lines. Scale bar: 100 μm. Source data are provided as a Source Data file.
We next characterised the geometry of the EDGE. Intensity profile analyses of GFP-tagged CAF#1 (Fig. 1h) indicated that this region has a width of 120 μm ± 33 μm and height of 24 μm ± 5 μm (mean ± SD), equivalent to the thickness of 4 ± 1 CAFs (Supplementary Fig. 2a, b). To test the versatility of MIRO, we co-cultured CAF#1-GFP with a distinct mCherry-labelled HER2+ BCC line (HCC1419). The seeding protocol was adapted to enable optimal CAF#1/HCC1419 self-organisation (see “Methods” section). We observed a lower peak in the GFP signal at the CAF#1/HCC1419 EDGE compared to CAF#1/HCC1954, suggesting reduced CAF accumulation in the region (Supplementary Fig. 3a, b). However, cell distribution and orientation were similar in the EDGE + OUT areas of both CAF#1/HCC1419 and CAF#1/HCC1954 co-cultures (Supplementary Fig. 3a–c), indicating that self-organisation can be achieved with distinct cancer types.
CAF fluorescence at the EDGE suggests distinct cell orientation and increased cellular compactness near the cancer cell compartment (Fig. 1h, i). This conformation together with the increased F-actin signal in this area suggests that the EDGE could bear higher tension than the OUT region, as proposed elsewhere33,34 (Supplementary Fig. 4a). To test this hypothesis, we performed local disruptions in both the EDGE and OUT regions with a custom-made laser ablation system (Supplementary Fig. 4b–g; Supplementary Movies 1 and 2). Our data indicate that the EDGE region recoils to a greater extent than the OUT region (Supplementary Fig. 4e–g) as shown by a higher retraction of the cut edges (Supplementary Fig. 4g). When a ROCK inhibitor (Y27632) was added in the medium, the amplitude of the recoil in both regions decreased but to a lesser extent in the proximity of the boundary, suggesting a higher tensional state in this area (Supplementary Fig. 4g). These observations align with previous studies suggesting that CAF accumulation and actomyosin contractility influence the mechanical and structural properties of the stroma surrounding the cancer cells33,34,35.
We next evaluated the potential of MIRO as a model to investigate immune cell distribution in the tumour and its microenvironment. 48 h after seeding BCC spheroids on top of the CAF monolayer, fluorescent-labelled naïve primary immune cells collected from the peripheral blood of healthy donors (peripheral blood mononuclear cells, PBMC) or natural killer cells (NK cells) derived from PBMCs, were inoculated in the outer compartment of MIRO (Fig. 1b). Time-lapse imaging in the channels and the EDGE region allowed us to track immune cells migrating through the CAF layers and CAF-derived ECM proteins, as shown by fibronectin immunostainings, when they entered the inlet compartment and reached the cancer/stroma boundary (Supplementary Fig. 5 and Supplementary Movies 3–6).
In conclusion, the spatial organisation of fibroblasts, fibroblast-derived ECM and immune cell distribution in MIRO can modelize the cancer/stroma interface with an immune-excluded phenotype, allowing quantitative studies of immune cell distribution and dynamics upon treatments.
MIRO captures spatial features of the tumour/stroma boundaries found in breast tumours
We aimed at further characterising similarities between fibroblasts and ECM repartition in MIRO and in patient tumours. Comparison between paraffin-embedded formalin-fixed samples of patient tumours and MIRO revealed a similar distribution of immune cells, largely excluded from the epithelial cancer cell compartment and aligning with the cancer/stroma interface (Fig. 2a, b; Supplementary Fig. 6a). We next carried out a comparative histopathological analysis of breast tumour samples and MIRO. For this, we performed immunohistochemical detection of CAF markers (fibroblast activation protein, FAP; vimentin, VIM), an epithelial marker (E-cadherin, E-cadh) and matrix proteins commonly found in solid tumours (collagens, Picrosirius red staining; collagen IV, COLIV; collagen I, COLI; fibronectin, FN) (Fig. 2c; Supplementary Fig. 6b-f). Fluorescent intensity signals from immunostainings show that patient tumours and MIRO share a similar distribution of CAFs (FAP and VIM) and ECM proteins (COLIV, FN) (Fig. 2c; Supplementary Fig. 6b). Quantitative analysis of fluorescent intensity profiles of CAF-GFP signal in MIRO and FAP, COLIV and FN expression in patient tumours revealed similar distribution of CAFs and ECM fibres with higher density at the EDGE compared to distal regions (Fig. 2d, e; Supplementary Fig. 6b). Furthermore, we observed that CAFs and ECM fibres similarly aligned along the edge of cancer cell compartments both in MIRO and in patient tumours (Fig. 2d, e; Supplementary Fig. 6d, h). Collectively, these data indicate that the TME in MIRO replicates key features of the aberrant production and architecture of ECM observed at the cancer/stroma interface in patient tumours16,36.
a Immune cell distribution in breast tumour samples (CD45 staining), and b in MIRO co-culture (PBMCs, fire-LUT), 3 independent experiments. Scale bars: 50 μm. c Immunohistochemistry and immunofluorescent staining in breast tumour samples targeting CAFs (FAP), tumour cells (E-cadherin (E-cadh)) and ECM components: COLIV, FN and COLI. Scale bars: 100 μm. d Relative fluorescent intensity profiles and orientation of CAF#1 (cytosolic GFP marker in MIRO, FAP in tumour samples), COLIV and FN in both breast tumour samples and MIRO. Lines indicate the mean values, coloured areas the SD. The “0” value refers to the tumour/stroma boundary location, set as reference (n = images, CAF (FAP) n = 27, COLIV n = 30, FN n = 27; from 3 patients samples, and in MIRO CAF (GFP) n = 18, COLIV n = 18, FN n = 17, from at least 3 independent experiments). e Heatmap of correlation coefficients comparing intensity profiles and fibre angles of tumour samples and MIRO (displayed in d) for CAF, COLIV and FN (Pearson correlation coefficient (two-tailed, 95% CI) with p-values < 0.0001. Source data are provided as a Source Data file.
MIRO co-culture enables the tumour/stroma boundary formation with diverse fibroblast lines
We tested the versatility of MIRO by using additional CAFs and normal fibroblasts. We used additional GFP-labelled immortalised CAF (CAF#2) and normal fibroblasts (BF#1, BF#2), as well as primary CAFs (pCAF#1, pCAF#2), with distinct mesenchymal and ECM-related gene expression levels37,38 (Fig. 3; Supplementary Fig. 7a, b). We evaluated fibroblast accumulation after 6 days of co-culture with BCC spheroids (HCC1954) by averaging GFP intensity profiles where applicable and measuring COLIV and FN expression for each combination. Furthermore, we analysed the fibroblasts, COLIV and FN orientation at the EDGE and OUT regions (Fig. 3a–e, Supplementary Fig. 8a and Supplementary Fig. 9). For primary fibroblasts, the tumour/stroma boundary was delineated using the autofluorescent signal emitted by the cells in the 488 nm excitation channel (Supplementary Fig. 8b). We observed increased fibroblast accumulation at the EDGE compared to OUT regions for all combinations (Fig. 3a–e). We also validate that the architecture of the tumour boundary using the additional fibroblasts shared similarities with the one observed in breast patient samples. Pearson correlation analysis performed on the CAF, COLIV and FN intensity curves and angle orientations for each fibroblast in MIRO compared to the breast tumour samples shown in Fig. 2e indicates a strong correlation of the marker expression patterns between the samples (Fig. 2e and Fig. 3f). Additionally, despite some significant differences between cell types, fibroblasts, COLIV and FN meshworks aligned along the cancer cell compartment with angles ranging between 10° and 40° relative to the EDGE, regardless of the fibroblast origin, and oriented randomly in the OUT regions (Fig. 3g, h and Supplementary Fig. 9). ECM and fibroblasts exhibit similar orientation and accumulation in the EDGE/OUT regions, indicating that the formation of intratumoral spatial architectures is independent of the fibroblast type seeded in MIRO. Altogether, these data demonstrate that reproducible models of cancer/stroma boundaries can be achieved in MIRO using distinct types of fibroblasts.
a–e Left column panels: representative images of CAF#2 (a), BF#1 (b) and BF#2 (c) (GFP signal), COLIV and FN co-cultured with HCC1954 BCC spheroids. For primary fibroblasts pCAF#1 (d) and pCAF#2 (e), no GFP signal is present. White dashed lines indicate the cancer cells' spheroid boundary. Scale bar: 100 μm. Right column panels: relative fluorescent intensity profiles and orientation of the fibroblast (GFP signal), COLIV and FN (immunostaining) relative to the tumour/stroma boundary. Lines indicate the mean values, coloured areas around the means the SD. The “0” value refers to the tumour/stroma boundary location set as reference (n = images; CAF#2: n = 20, COLIV n = 12, FN n = 8; BF#1: n = 23, COLIV n = 12, FN n = 11; BF#2: n = 23, COLIV n = 12, FN n = 11; pCAF#1: n = 22, COLIV n = 10, FN n = 12; pCAF#2: n = 20, COLIV n = 8, FN n = 12; 3 independent experiments). f Heatmap of correlation coefficients comparing intensity profiles and fibre angles of CAF#2, BF#1, BF#2, pCAF#1 and pCAF#2 (displayed in a–e) and BC tumour samples (Fig. 2d) for COLIV, FN and CAF (when possible). Pearson correlation (two-tailed, 95% CI), with p-values < 0.0001. g Fluorescence intensity ratio of fibroblast GFP signal, COLIV and FN immunostainings in the EDGE and OUT regions (ratio EDGE/OUT). One-way ANOVA (COLIV) and Kruskal–Wallis (GFP, FN) tests were performed with p-value *p = 0.0295. h Relative orientation of fibroblasts (major elongation axis), COLIV and FN respective to the tumour/stroma boundary Kruskal–Wallis test was performed p-values: CAF#1 vs CAF#2 (GFP) **p = 0.0073, (COLIV) *p = 0.0434; CAF#1 vs BF#1 (GFP) *p = 0.0187, (COLIV) * = 0.0286, (FN) *p = 0.0456; pCAF#1 vs BF#1 (FN) *p = 0.0303. In all graphs, ns indicates non-significant, non-shown comparisons are non-significant (n = measurements, n(CAF) = 20; n(COLIV, FN) = 8, from 3 independent experiments). Source data are provided as a Source Data file.
Stroma drives tumour immune exclusion in MIRO
Previous observations indicated that the stromal barrier formed by CAFs/ECM, and surrounding epithelial cancer cell compartments, may restrict immune cell trafficking into tumours14,15. We then assessed whether immune cells seeded in the MIRO outer compartment could reach and infiltrate the cancer cell compartments in the inner chamber (Fig. 4a). About 30 h after seeding, immune cells were detectable in the vicinity of the cancer cell compartments both in the presence or absence of CAFs (Supplementary Fig. 10a, b). Notably, immune cells infiltrated readily the cancer cell compartments in the absence of CAFs and ECM (Fig. 4b, c). By contrast, we observed a 4-fold reduction of immune infiltration into cancer compartments when co-cultured with CAFs (Fig. 4c). These findings indicate that the presence of CAFs and CAF-secreted ECM does not hinder the migration of immune cells from the outer to the inner compartment of MIRO. However, cancer/stroma boundaries are sufficient to drive consistent immune exclusion from the cancer cell compartments.
a Co-culture timeline in MIRO in the absence or presence of stroma. b Time projection of time-lapse imaging of immune cells reaching MIRO inlet and interacting with the tumour spheroid. For each time point, the fluorescence image of the immune cells was binarized and overlaid with the tumour cell fluorescent signal. Immune cells overlapping with tumour cells over time are displayed in yellow, the others are shown in cyan, and BCC are in red. Scale bar: 100 μm. c Quantification of immune cells infiltrated in the tumour compartments in the absence (without CAF#1 black dots) and presence (with CAF#1, green dots) of CAF#1. The dotted line represents the mean value of the immune cell counted over time; SD is shown as a filled coloured area (n = measures, with stroma n = 8; without stroma n = 6, from at least 3 independent experiments;), unpaired t-test over the last 10 time points (two-tailed) was performed, ****p < 0.0001. Source data are provided as a Source Data file.
Live visualisation of intraepithelial immune infiltration induced by Trastuzumab and Interleukin 2 combined regimen
Next, we assessed the potential of MIRO for drug testing with Trastuzumab (Ttz), an anti-HER2 monoclonal antibody (mAb) commonly used in the treatment of HER2+ breast cancer39. In addition to its direct cytotoxic effects on HER2+ breast cancer cells39, anti-HER2 mAbs have been shown to significantly enhance anti-cancer immunity through antibody-dependent cellular cytotoxicity (ADCC)40,41, particularly when combined with interleukin 2 (IL2)42, hereafter referred to as IL2+Ttz (Fig. 5a). Importantly, HCC1954 BCCs are intrinsically resistant to the cytotoxic effects of anti-HER2 mAbs43, allowing us to focus exclusively on the immunomodulatory effects of IL2+Ttz in our system. Based on previous studies, we anticipated an optimal anti-cancer response 72–96 h after starting treatment42. We observed that immune cells reached the tumour edge approximately 30 h after seeding in both treated and control conditions. However, in the absence of IL2+Ttz, immune cells remained mostly excluded from the cancer cell compartment (Fig. 5b, c; Supplementary Movie 7), mimicking the immune exclusion observed in refractory tumours.
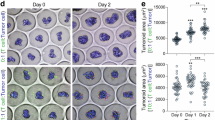
a Timeline of MIRO co-culture in the presence or absence of IL2+Ttz treatment. b Time-lapse imaging of MIRO inlet showing the effect of IL2+Ttz treatment on immune cell infiltration in tumour compartments (HCC1954-mCherry BCC spheroid; CAF#1-GFP). Scale bars: 200 μm; zooms: 100 μm. c Quantification of immune cells infiltrated in tumour compartments 24 h after immune cell seeding in the absence (control, black dots) or presence of IL2+Ttz treatment (pink dots). The dotted line represents the mean value of the immune cell counted over time, SD is shown as a filled coloured area (CT n = 8; IL2+Ttz n = 6 videos, from at least 3 independent experiments;), unpaired t-test over the last 10 time points (two-tailed) was performed, ****p < 0.0001. Immune cell counting in the tumour compartment (d) and in the stroma (e) for different fibroblasts (CAF#1, BF#1, pCAF#1 and pCAF#2) co-cultured with HCC1954 BCC spheroids and in presence or absence of IL2+Ttz treatment. t-tests (two-tailed) were performed (n = measurements, CT CAF#1 n = 26, IL2+Ttz CAF#1 n = 29 with **p = 0.0046; CT BF#1 n = 31, IL2+Ttz BF#1 n = 37 with **p = 0.0013; CT pCAF#1 n = 28, IL2+Ttz pCAF#1 n = 25 with ***p = 0.0003; CT pCAF#2 n = 17, IL2+Ttz pCAF#2 n = 24 with *p = 0.0225, 3 independent experiments). In all graphs, ns indicates non-significant. Source data are provided as a Source Data file.
In contrast, IL2+Ttz treatment significantly increased intraepithelial immune cell abundance regardless of the type of fibroblasts used in MIRO (Fig. 5d; Supplementary Movie 7). We assessed whether this increase in immune infiltration correlated with enhanced overall immune cell presence in the tumour vicinity. We quantified the immune cells in the EDGE + OUT regions and found no significant increase in immune cell abundance upon treatment (Fig. 5e). Additionally, treatment did not affect immune cell proliferation rates, as no significant differences were observed across conditions (Supplementary Fig. 11). Taken together, these findings indicate that the stromal barrier can be breached by infiltrating immune cells upon specific cell targeting and effector cell activation.
Trastuzumab and Interleukin 2 combined treatment effect on stromal barrier architecture and composition
Preclinical studies, although limited, have suggested the potential for Trastuzumab -based regimens to modulate CAF activity and ECM properties10. Hence, we asked whether IL2+Ttz treatment could affect the architectural and molecular characteristics of the TME, and therefore the permeability of the stromal barrier. We first quantified CAF apoptosis upon IL2+Ttz treatment by performing cell death assays in the presence or absence of PBMCs. We observed no significant differences in CAF death rate in any of the tested conditions (Fig. 6a). Furthermore, fibroblast repartition and orientation at the EDGE remained unaffected by IL2+Ttz treatment (Fig. 6b, c). We then hypothesised that treatment could modulate immune cell trafficking by altering ECM organisation2,12. Quantitative analysis of FN and COLIV immunodetection showed no significant changes in the distribution and orientation of ECM upon four-day IL2+Ttz treatment (Fig. 6d) compared to control conditions (Fig. 3d). Consistently, the expression levels of FN, COLIV, and COLI remained unchanged in CAFs upon IL2+Ttz treatment (Fig. 6e). These data suggest that IL2+Ttz treatment does not enhance anti-cancer immunity by modulating the composition and organisation of the fibroblasts and fibroblast-derived ECM in MIRO.
a Quantification of propidium iodide fluorescent signal on CAF#1 (cell death quantification) in the absence or presence of IL2+Ttz drug treatment and of immune cells (PBMCs) (n = measurements, CT no PBMCs n = 10; CT PBMCs n = 15; IL2+Ttz no PBMCs n = 11; IL2+Ttz PBMCs n = 18; 3 independent experiments). One-way ANOVA with multiple comparison tests was performed. b Quantification of the height of the EDGE region of fixed samples based on CAF#1-GFP fluorescent signal in the presence or absence of IL2+Ttz treatment (n = 30 measurements, 3 independent experiments). Unpaired t-test (two-tailed) was performed, p = 0.2217. Box plots in (a) and (b) show the median and bars the minimum to maximum values. c, d Relative fluorescence intensity profiles and relative orientation of CAF#1 (GFP signal) (c), COLIV and FN (d) respective to the tumour/stroma boundary in MIRO. Lines indicate the mean values and coloured areas the SD. The “0” value refers to the tumour/stroma boundary location set as a reference. Unpaired multiple t-tests performed shows no significant differences in the accumulation, distribution and orientation of CAFs, FN and COLIV upon IL2+Ttz drug treatment compared to CT (shown in Fig. 2d) except for intensity x = 0 CAF#1 CT vs CAF#1 IL2+Ttz, p < 0.0001 (n = measurements, CAF (GFP) n = 18, COLIV n = 17; FN n = 18, 3 independent experiments). e RT-qPCR quantification of FN (FN1), COLIV (COL4A1) and COLI (COL1A) expression in CAF#1 monoculture in the presence or absence of drug treatments. Values are mean ± SD (n = 3 independent experiments), ratio paired t-test (two-tailed) was performed, COL1A p = 0.8574, COL4A1 p = 0.0938, FN1 p = 0.7685. In all graphs, ns indicates non-significant. Source data are provided as a Source Data file.
Immune cell motility and spreading at the cancer/stroma interface upon treatment
We next examined the impact of treatments on immune cell activity in the presence of CAFs. We focussed on NK cells, which represent the main immune subtype mediating ADCC7,8,12. NK cell-directed immunotherapy stands as an emerging strategy attracting increasing attention11. Indeed, numerous clinical trials exploring diverse avenues of NK cell-focused immunotherapy, including those involving IL2+Ttz-based regimen (e.g. NCT05207722, NCT05395052, NCT05385705), are already in phase I/II stages11. Gene expression analyses showed that the migration markers ITGAM and ITGAL44,45 were upregulated in NK cells upon IL2+Ttz (Supplementary Fig. 12a), thus suggesting an increased immune migratory phenotype. Prompted by these findings, we assessed the capacity of IL2+Ttz regimen to modulate NK cell trafficking through the CAFs barrier in MIRO (Fig. 7a, b). Time-lapse imaging of single focal plans (~1 frame/min, for 90 min) enabled the tracking of a large majority of NK cells in the vicinity of the epithelial cancer cell compartment (Fig. 7c and Supplementary Movies 8 and 9). This approach allowed us to monitor the orientation, velocity, and morphology of individual NK cells trafficking into the tumour stroma. As previous reports highlighted the scaffolding role of the stroma in guiding immune cells46,47, we analysed the orientation of the NK cell migratory paths with the CAF major axis in our model. The comparative analysis of these patterns revealed a mild but significant correlation between NK cell trajectories and CAF orientation (Fig. 7d, e). Consistent with these results, previous studies showed that collagen alignment guides T-cell migration around the tumour islets14,15. Considering that IL2+Ttz treatment was not altering CAF, FN and COLIV architecture in our model (Fig. 6a–d), we then assessed the effect of treatments on NK cell velocity in the EDGE region. We observed a significant increase in NK cells average speed upon IL2 or IL2+Ttz treatment (55% and 21% respectively) compared to untreated control and to Trastuzumab (Ttz) monotherapy (Fig. 7f). These data suggest that IL2 is mainly responsible for the modulation of the migratory behaviour of NK cells. IL2 has been described to enhance integrin and F-actin expression, as well as lytic cellular behaviour48,49. Hence, we assessed the effect of treatment on the morphology of migrating NK cells. We performed time-lapse imaging using z-stacks at the cancer/stroma interface (1.5 min/frame, for 30 min). Z-projection of time-lapse images captured remarkable deformation of NK cells moving between cancer cells and CAFs (Supplementary Movies 10 and 11). We quantified the maximum spreading area of each NK cell tracked in the different treatment conditions masking each cell based on their fluorescent signal (cytosolic cell tracker) (Fig. 7g, h and Supplementary Fig. 12b). Our results showed that IL2 and IL2+Ttz treatments increase NK cell area by 85% and 59% respectively, compared to control condition. While not reaching significance, an increase was also noted in the presence of Trastuzumab alone (36%). Interestingly, we observed a similar although non-statistically significant effect of IL2 and Il2+Ttz treatment on the average speed of unsorted PBMCs despite the heterogeneity of the immune cell populations in these samples (Supplementary Fig. 12c). Consistent with prior studies13,14, our data indicate that IL2 enhances the migratory characteristics of NK cells in MIRO. This includes an increase in their velocity, integrin expression levels, and spreading capacity in the stromal and cancer compartments. Taken together, these data demonstrate that MIRO enables the assessment of treatment effects on individual cell behaviour within the TME, facilitating the characterisation of immune cell trafficking associated with anti-cancer response/resistance.
a Illustration of the co-culture timeline in MIRO with NK cells seeding. b Schematic of the experimental set-up using MIRO devices using 12-well plates. The multiwell configuration allows performing multidrug screening and triplicate of each condition per experiment. c Representative fluorescence image of CAF#1-GFP (green), HCC1954-mCherry BCC spheroids (red) (left panel) and NK cell tracks monitored over 90 min time-lapse acquisition (right panel, white lines) in the tumour/stroma boundary region. Scale bars: 100 μm. d Orientation of CAFs#1 and NK cell tracks relative to the tumour/stroma boundary (thick dashed red line). Scale bars: 100 μm. e Correlation analysis (correlation coefficient calculation, cc) of CAF#1 and NK cell track orientations in the EDGE region showing a positive correlation (cc (mean ± SD) = 0.16 ± 0.08) compared to randomised data correlation analysis (cc (mean ± SD) = 0 ± 0.09) (n = 1260 measurements, 4 independent experiments) Mann–Whitney test (two-tailed) was performed, ****p < 0.0001. f Frequency distribution (in %) of NK cell mean velocity in the tumour EDGE region upon IL2 treatment (n = measurements, CT n = 25, IL2 n = 119, Ttz n = 50, IL2+Ttz n = 95, at least 3 independent experiments); Kruskal–Wallis tests were performed. g Representative time-lapse binarized images of single NK cells migrating in the EDGE region in the presence or absence of treatment. Each montage depicts the dynamic of morphological changes of NK cells over 9 min (time between frame 1 min, n = 10 cells per condition). Scale bars: 5 μm. h Maximum NK cell spreading area (m) of NK cells tracked in the tumour EDGE in MIRO in the absence of treatment, CT m = 84.3 ± 31.04 μm2, or in the presence of IL2 m = 155.9 ± 78.49 μm2, Ttz m = 114.9 ± 45.64 μm2, or IL2+Ttz m = 134.3 ± 56.73 μm2. n = measurements, CT n = 24, IL2 n = 87, Ttz n = 49, IL2+Ttz n = 79, from at least 3 independent experiments). Kruskal–Wallis test (two-tailed) was performed, ****p < 0.0001, *p = 0.0409. In all graphs, non-shown comparisons are non-significant. Source data are provided as a Source Data file.
Immuno-infiltration and ADCC quantification upon treatment
To assess whether higher immune infiltration correlated with increased cancer cell death, we quantified NK cell abundance in the cancer cell and stroma compartments. Cell counting revealed that treatment with IL2 or IL2+Ttz increased NK cell abundance in the cancer cell compartments by ~4-fold compared to control (Fig. 8a, b and Supplementary Fig. 13) without significant increase in the number of NK cells in the stromal compartment. Similar results using unsorted PBMCs showed equivalent effects of IL2 and IL2+Ttz treatments on immune infiltration in the cancer cell compartment and higher infiltration of PBMCs in the stroma upon IL2 treatment (Supplementary Fig. 14b–d). We then quantified the cytotoxic activity of NK cells in the different treatment conditions using a fluorescent reporter of cell death (NucGreen® Dead reagent, Invitrogen). It is worth noting that while MIRO is an allogeneic model, no significant cancer cell death is induced within standard four-day experimental setting in presence of immune cells and absence of treatment (Fig. 8c and Supplementary Fig. 14a). Four days after immune cell seeding, we observed that IL2 monotherapy increased immune cell abundance in the cancer cell compartments without enhancing anti-cancer cell immunity in the absence of Trastuzumab (Fig. 8c and Supplementary Fig. 14b–d). These observations suggest that short-term co-culture with naïve NK or whole immune cells (PBMCs) does not trigger major histocompatibility complex class I-dependent immune or antigen-non-specific T-cell receptor-independent responses in MIRO, thus enabling the study of anti-cancer-immune activation upon therapy. Consistently, we observed a substantial increase in apoptosis among intrinsically resistant to Trastuzumab HER2+ BCCs following treatment with IL2+Ttz and to a lesser extent with Trastuzumab alone (Fig. 8c and Supplementary Fig. 14a). This strongly suggests a mechanism of ADCC relying on both IL2 ability to overcome the immunosuppressive effects of the tumour/stroma barrier and the recognition of Trastuzumab-labelled cancer cells by immune cells. Altogether, our data suggest that IL2 enhances immune cell activation and NK cell migration through the CAF/ECM barrier. The resulting increase of intraepithelial immune cells in turn favours Trastuzumab-induced ADCC and cancer cell eradication. Accordingly, expression analysis of ADCC markers in NK cells co-cultured with cancer cells shows significant upregulation of Perforin 1 and Granzyme B upon IL2 or IL2+Ttz treatment, as also observed with PBMCs (Fig. 8d and Supplementary Fig. 14e). These data suggest a higher cytolytic activity, cytokine secretion and degranulation of infiltrating immune cells upon IL2 administration.
a Representative images of tumour/stroma boundary region in MIRO in the presence or absence of treatment. White circles show NK cells in the tumour compartment (EDGE boundaries (dashed line), CAF#1-GFP (green); BCC spheroid, (red)). Scale bars: 100 μm. b NK cells counting in the tumour (IN) and in the stroma compartments in the control condition (CT) and upon treatment (n = images; CT n = 19, IL2 n = 26, Ttz n = 22, IL2+Ttz n = 29; 5 independent experiments), Kruskal–Wallis multiple comparison test was performed, CT vs IL2 **p = 0.002, CT vs IL2+Ttz p = 0.0045. c BCC death quantification of in the absence or presence of drug treatments and of NK cells (n = measurements; no NK cells: CT n = 15; IL2 n = 13; Ttz n = 12; IL2+Ttz n = 10; NK cells: CT n = 30, IL2 n = 17; Ttz n = 29; IL2+Ttz n = 27; at least 4 independent experiments). Box plots show the medians, bars the minimum to maximum values. One-way ANOVA was performed, **p = 0.0051, ***p = 0.0006 (CT vs Ttz) and p = 0.0001 (Ttz vs IL2+Ttz), ****p < 0.0001. d, e RT-qPCR quantification of Perforin 1 (PRF1), Granzyme B (GZMB) and ITGAL expression level in NK cells co-cultured with HCC1954 BCC in the absence (CT) or presence of Ttz and/or IL2 treatment (d n = 5 and e n = 3 independent experiments). Ratio paired t-test (two-tailed) was performed, **p = 0.0093; * PRF1 p = 0.0365 (CT vs IL2), p = 0.0272 (CT vs IL2+Ttz); GNZB p = 0.0252 (CT vs IL2), ITGAL p = 0.0431 (CT vs IL2+Ttz). f Left: Time-lapse imaging sequence of BCC death (BCC nuclear contour: dashed white line) triggered by an NK cell. Frame interval = 1 min. Scale bars: 5 μm. Right: Illustration of NK cell/cancer interaction leading to cancer cell death. g Frequency distribution of the time to lytic hit (left) and to death (right) induced by NK cell upon IL2+Ttz treatment (n = 48 measurements, 2 independent experiments). In all graphs, ns indicates non-significant; non-shown comparisons are non-significant. Source data are provided as a Source Data file.
ADCC is a local mechanism that implies transient immune cell-cancer cell physical contacts leading to targeted cellular membrane perforation50. In line with this, NK cells showed significant upregulation of ITGAL in the presence of cancer cells and IL2+Ttz regimen (Fig. 8e). Since ITGAL has also been involved in cell-cell adhesion during immunological synapses45, we speculated that IL2 and Trastuzumab treatments could also affect NK cell motility and the ADCC response. We then quantified these transient events upon IL2+Ttz administration. For this, we performed time-lapse imaging upon the addition of a fluorescent cell death reagent in MIRO upon IL2+Ttz treatment (Fig. 8f, g). With this approach, we could observe ADCC events mediated by NK cells upon IL2+Ttz treatment (Fig. 8g). Remarkably, 67% of immune/cancer cell contacts were under 6-min duration and led to 62,5% of total cancer cell death in the first 20 min following treatment administration (Fig. 8f, g and Supplementary Movies 12 and 13). In addition, NK cell and PBMC tracking in the IN region indicated that the average speed upon IL2, Trastuzumab and IL2+Ttz treatments was significantly increased compared to control, suggesting a global gain in the scanning behaviour of intraepithelial NK cells (Supplementary Fig. 14f and Supplementary Fig. 15a). IL2 and IL2+Ttz treatments also significantly increased NK cell maximum spreading area respectively by 44% and 32% in the cancer cell compartments, while in contrast Trastuzumab alone induced only marginal NK cell spreading (13%) (Supplementary Fig. 15b). These values are consistent with previous studies assessing NK cell-driven ADCC in 3D matrices51. Collectively, these data indicate that IL2 stimulates immune cell trafficking in tumours and restores Trastuzumab-induced ADCC in immunosuppressed tumours.
Discussion
Increasing evidence indicates that ECM and stromal cells encapsulating epithelial cancer cell islets hinder immune infiltration in a subset of tumours, thus favouring cancer progression and resistance to therapy21,52. However, the mechanisms underlying immune exclusion by the TME remain mostly unknown. Hence, there is an urgent need for approaches allowing the study of tumour immunity on a patient-by-patient basis and the identification of personalised treatments to overcome immune evasion. In this context, microfluidic models have the potential to recapitulate the complex cellular interactions occurring within the TME.
Here, we described MIRO (Micro Immuno-Response On-Chip), an in vitro platform creating a CAF-ECM stromal barrier that recapitulates key features observed in cancer patients. As proof of concept, we used MIRO to recreate cellular traits of a HER2+ breast cancer subtype resistant to therapy. We show that MIRO enables the formation of reproducible 2.5D cancer/stroma interface (EDGE) surrounding cancer cell compartments (IN), and the establishment of a meshwork of ECM proteins that shares molecular and structural similarities with the TME of patient tumours. In MIRO, fibroblasts, fibronectin and collagens align along the tumour edge in the vicinity of the cancer/stroma interface. We show that this organisation is reminiscent of the histopathology of cancer/stroma interface organisation commonly found in solid tumours52,53. In addition, CAF alignment, TME compactness and recoil speed in MIRO suggest an elevated tensional state at the tumour/stroma interface. Compressive forces in the vicinity of the cancer islets have been previously related to a reduction of the matrix hydraulic permeability34,35, favouring tumour encapsulation. However, the role of the stromal tension and mechanical properties driving TME architecture and impacting cancer treatment remains elusive as the current in vitro models that enable addressing those questions are sparse. In this context, MIRO offers a powerful approach to quantitatively assess the effect of diverse mechanical perturbations (e.g. laser ablation, targeting ECM/CAF-related genes and immune cells migration) on the immune infiltration state of the cancer cell compartment.
As proof of concept, we leveraged MIRO to recreate cellular traits of HER2+ breast cancer unresponsive to therapy using distinct types of fibroblasts and cancer cells. We monitored the immune infiltration upon anti-HER2 mAbs therapy (Trastuzumab) and demonstrated that stroma-encapsulated HER2+ BCCs are resistant to Trastuzumab-induced ADCC. IL2-based compounds have recently drawn growing interest for the treatment of cancers including HER2+ breast cancers refractory to therapy42,54,55,56. Over the past decade, approximately 800 clinical trials explored the benefits of IL2, with around 200 of these trials currently active and recruiting participants (as listed on http://clinicaltrials.gov). This extensive clinical activity underscores the widespread interest in investigating IL2 in cancer treatment, including in association with Trastuzumab (e.g. NCT05395052, NCT05385705, NCT05207722). In this line, we show that MIRO recapitulates the synergistic effect of Trastuzumab and IL2 on restoring anti-cancer immunity. Single-cell tracking of NK cells and PBMCs migrating within CAF-derived ECM and cancer cell compartments indicates that IL2 in combination with Trastuzumab increases the intraepithelial abundance of immune cells. Furthermore, we show that treatment-induced anti-cancer immunity is associated with increased immune cell motility as well as changes in spreading morphology both at the tumour margin (EDGE) and in the cancer cell compartments (IN). These observations are consistent with independent studies reporting similar speeds of NK cells in a 3D collagen matrix, ranging between 2 and 5 µm/min14,49. Importantly, our data suggest that IL2 stimulation overcomes immune exclusion by increasing the migratory capacity of immune cells, rather than altering the stromal barrier. In addition, higher cytolytic activity, cytokine secretion and degranulation of infiltrating immune cells were also observed upon IL2 administration.
On a technical level, it is essential to acknowledge the limits of our present model. Notably, MIRO lacks blood and lymphatic vessels, as well as neural components. The absence of vascularisation prevents the full replication of nutrient and oxygen gradients, as well as immune cell circulation. Additionally, the exclusion of neural networks neglects the influence of neuro-immune crosstalk on tumour behaviour and response to therapies57. Addressing these limitations in future models may substantially improve their physiological relevance, enabling even more accurate studies of cancer-stroma-immune interactions, treatment responses, and metastatic progression. Additionally, although the stability of the co-culture in this study was demonstrated over the culture period, it is important to point out that other cell types might behave differently, leading to diverse spatial arrangements. It is also important to mention that cancer cells need to maintain a minimal but critical level of intercellular cohesion, in addition to a certain density of fibroblasts to allow the formation and the stability of the tumour/stroma boundary in MIRO. Despite its inherent limitations, our model offers several substantial advantages. Its design provides controlled conditions for studying specific interactions and responses, facilitating the identification of key mechanisms in complex biological processes. One of the main advantages lies in the incorporation within a confined space of CAFs and ECM elements that are playing a pivotal role in forming barriers to immune cell infiltration and exerting immunosuppressive functions within the TME58,59. The outcome is the creation of a 2.5D cancer/stroma interface which is associated with immune exclusion and resembles the structure of the one observed in the human breast tumour margin. This configuration strikes a balance between 2D and 3D cultures, enabling the study of 3D-related effects while maintaining experimental flexibility and reproducibility, thus offering an opportunity for studying the underlying drivers and consequences of cancer cell-CAF-driven immunosuppressive barriers. In contrast, enrichment of these stromal proteins is mostly absent in currently available 3D assays and on-chip models. Alternatively, models based on patient tumour fragments may offer an enhanced representation of TME with patient-specific ECM and cellular complexity60,61,62. However, inherent sample variability can hinder reproducibility, and the practical and ethical challenges of obtaining and maintaining clinical fragments limit their widespread use. Compared to standard 3D environments and in vivo animal models, MIRO coupled with confocal microscopy and laser ablation enables high-resolution visualisation of cellular processes and precise manipulation of cellular structures. For instance, MIRO allows for the identification of multiple co-cultured cell subtypes through fluorescence labelling, tracking of individual cell behaviour including motility, migration, and spreading, monitoring immune cell trafficking, and real-time observation of single-cell apoptosis. These combined advantages provide powerful tools for gaining insights into dynamic cellular processes and interactions. Another benefit resides in the compatibility of the MIRO design with standard multiwell plates for cell culture. This multiplexing allows to easily test in parallel multidrug conditions and donor-to-donor variability of the immune cell population49,57,63. Hence, MIRO represents an adaptable platform for investigating anti-cancer immunity, which takes into account the organisation of BCC, stroma and ECM components.
While fully recapitulating the physiological TME remains a long-term goal, MIRO provides substantial advancements in replicating the spatial arrangement of human tumours. Given the potential of immunotherapy in oncology, MIRO offers a valuable ex vivo approach to identify key players mediating tumour immune exclusion in a controlled yet biologically relevant setting. This encompasses a wide range of uses, from tracking T-cell activation and migratory trends to investigating the mechanics of macrophage-tumour interactions. Potential applications of MIRO could include chemotaxis and the investigation of cytokine gradients. Furthermore, it provides a framework for comprehensive examinations of immunosuppressive pathways involving myeloid-derived suppressor cells and regulatory T-cells, facilitating a deeper understanding of how tumours evade immune surveillance. Considering the compatibility of MIRO with autologous cellular models, we envision that our approach may also offer a powerful platform for precision oncology. In the clinical setting, it may aid in personalised treatment selection by providing insights into tumour interactions with the microenvironment and immune cells. This could inform the choice of therapy targeting both cancer cells and the TME, facilitate biomarker discovery for treatment response, and optimise combination therapies while understanding resistance mechanisms. In pharmaceutical research, this platform may likely enhance drug development by providing insights into compound interactions in complex microenvironments, improving testing accuracy and candidate identification. Hence, MIRO holds the potential to guide targeted therapy development and inform immunotherapy design, advancing precision oncology by comprehensively addressing the interplay among cancer cells, stromal components, and immune responses.
Methods
Cell culture
HER2+ breast cancer cell lines (HCC1954, ATCC: CRL-2338; HCC1419, ATCC: CRL-2326) were provided by the American Type Culture Collection and were stably transfected via Lentivirus with a luciferase reporter coupled to a mCherry cytosolic marker (HCC1954 red; HCC1419 red). Breast fibroblasts were kindly provided by P. Gascon/P. Bragado laboratory (IDIBAPS, Barcelona) (CAF#1, CAF#2, BF#1, BF#2) and derived at Hospital del Mar Research Institute (pCAF#1 and pCAF#2) and are characterised elsewhere64. In brief, fibroblasts were expanded in culture from digested pieces of fresh human mammary tissue or HER2 + BC tissue. Differential trypsinization was performed to further isolate the fibroblast population. Fibroblast phenotypes were characterised by measuring the expression of α-SMA and FAP, in addition to other biomarkers described in Supplementary Fig. 7, by qPCR. A retroviral vector pMIG (MSCV-IRES-GFP) expressing hTERT and GFP was used to immortalise and label CAF#1, CAF#2, BF#1, BF#2. HER2+ breast cancer cell lines were cultured in RPMI (Thermo Fisher) medium and fibroblasts in DMEM (Gibco) both at 10% decomplemented foetal bovine serum (FBS; Gibco), penicillin (100 μg/ml) and streptomycin (100 μg/ml; Gibco), and 29.2 mg/ml glutamine (Invitrogen). Cells were incubated at 37 °C with 5% CO2 and passaged when confluency reached 80–90%. Peripheral blood mononucleated cells (PBMCs) were purified from blood samples of healthy donors with Lymphoprep™. Untouched NK cells were purified from PBMCs with the NK Cell Isolation Kit (Miltenyi Biotec) following the manufacturers’ instructions. PBMCs and NK cells were stained with CellTracker™ Deep Red Dye (Invitrogen) following manufacturers’ protocols as previously described (Rivas et al.42) and cultured in RPMI medium supplemented with 0.2% decomplemented foetal bovine serum (FBS), penicillin (100 μg/ml) and streptomycin (100 μg/ml). For cell culture in MIRO, at day 0 a suspension of 5000 fibroblasts were seeded in the inlet of the MIRO device, while for pCAF#1 and pCAF#1 a suspension of 10,000 cells was seeded, to account for their difference of proliferation rate. Breast cancer cell spheroids were formed by seeding 2500 HCC1954 or HCC1419 red cells in a 96-well Ultra-Low Attachment Microplate (Corning) in RPMI 10%. On day 4, 4 HCC1954 BCC spheroids were manually seeded inside the inlet on top of the fibroblast monolayer spaced from each other and away from the channel exit. On day 6, the co-culture was washed with 0.2% FBS RPMI medium, and was treated with 20 mg/ml of Trastuzumab, and/or 100 ng/ml of Interleukin 2 (IL2, Peprotech) corresponding to a specific activity of ≥107 units/mg. On the same day, 900,000 PBMCs or 150,000 NK cells were seeded in the outlet of the device. For HCC1419/CAF#1, on day 4 cancer cell spheroids were first seeded in MIRO to promote strong attachment to the substrate. 10,000 CAFs#1 were then seeded 4 h later, thus preventing their proliferation beneath the cancer spheroids, which could complicate boundary delineation. The spheroids positioned in a range of 200 μm distance from the channel exit were selected for imaging. The co-culture was imaged from 24 to 96 h after immune cell seeding, to achieve optimal response to the treatment 72–96 h after the immune cell and drug incubation, as previously described42.
MIRO device fabrication
A mixture of polydimethylsiloxane (PDMS) and curing agents (SYLGARD 184 Silicone Elastomer Kit, Dow) was mixed in a 10:1 weight-to-weight ratio and poured on a resin stamp, degassed in a vacuum desiccator, and solidified at 80 °C for 2 h. Then, the PDMS was peeled off from the mask. A custom-made puncher with 10 mm and 6 mm cutting rings was used to form the outlet and inlet of the device. Each PDMS device was activated using oxygen plasma for 40 s and sealed on a glass-bottom 12-well plate or 35 mm glass-bottom single dish. The devices were made hydrophilic with corona treatment and sterilised with exposure to ultraviolet light for 15 min.
Confocal microscopy image acquisition
Unless stated otherwise, the described co-culture was imaged in fluorescence on day 10 after CAF seeding at 37 °C in a CO2 humidity-controlled chamber using a confocal microscope Nikon C1Si with a spinning-disc confocal unit (CSU-W1, Yokogawa), and a Zyla sCMOS camera (Andor) controlled by the Micro-Manager software65 using 4×, 10×, 20×, or 40× air objectives.
IN, EDGE OUT areas characterisation
The delineation of the EDGE, OUT, and IN regions was extracted from the maximum fluorescent intensity z-projections of the confocal images of the fixed and immunostained co-culture using a custom-made MATLAB code (MATLAB4STROMA). For each field of view, the tumour/stroma boundary was manually outlined using the CAF/BF-GFP fluorescent signal. For the primary fibroblasts, the boundary was manually outlined using the autofluorescence signal from the 488 nm excitation channel (Supplementary Fig. 8b). The averaged fluorescent intensity profiles of the CAF-GFP, Fibronectin and Collagen IV signals were plotted in function of the distance to the tumour/stroma boundary and the CAF-GFP fluorescent profile was used to determine the EDGE width based on the findchangepts Matlab function66. The average width was estimated for the HCC1954 and CAF#1 (mean width = 120 ± 33 µm values; n = 51 images, at least 17 spheroids from at least 4 independent experiments), and the “EDGE region” was defined as the stromal area between 0 and 120 µm distance from the tumour/stroma boundary. The “OUT region” was defined as the stromal area corresponding to twice the distance away from the calculated edge width average value (for the CAF#1, the “OUT region” starts 240 μm away from the tumour/stroma boundary). The “IN region” refers to the region occupied by the tumour spheroid.
The EDGE height was estimated by counting the number of z-stack images from the acquisition of GFP-labelled CAF#1 on fixed samples (20x objective, 1 μm z-steps). To estimate the number of CAFs#1 stacked at the EDGE region (Nfib), the samples were incubated with Hoechst 33342 for DAPI staining. The height of individual CAFs was first calculated by counting the number of z-steps (with 1 μm step resolution) for which the nuclear fluorescent signal (Hoechst 33342) was distinguishable (n = 19 images, 19 spheroids, 6 independent experiments). With this method, the average CAF#1 height (Hfib) was estimated to be Hfib= 5 ± 1 μm. To obtain the Nfib values, each value of the EDGE height was then divided by Hfib. The EDGE and CAF height values were corrected with a correction factor (α) as previously described67 (α = 1.3765 for live samples).
Immune cell tracking and morphology analysis
Immune cell tracking was performed through time-lapse confocal acquisition (1 frame/min, 90 min) on day 4 after immune cell seeding. NK cell morphology analyses were performed using a combination of the TrackMate Fiji plugin and a custom MATLAB script (MATLAB4Trackmate). For each track, the maximal spreading area of the selected cell was calculated using the binarized fluorescent image (647 nm channel). The mean values of maximal spreading area were obtained by averaging all values of a given track. Immune cell mean velocity was calculated for each track by averaging the instant velocities corresponding to distances between time point (t) and (t + 4). For these analyses, tracks shorter than 6-time point lengths were discarded.
Stroma orientation and immune cell correlation analysis
The orientation of ECM fibres and fibroblast towards the tumour/stroma boundary was calculated using ctFIRE programme68 and a MATLAB custom script (MATLAB4Stroma). Briefly, the ctFIRE programme was run on the max z-projection fluorescent images of the MIRO stained samples and the TMA samples to detect the CAF and ECM fibre meshwork orientations and then analysed using a Matlab custom-made script. For each pixel of a fibre detected, the segment formed by pixels (i) and (i + 4) was used to calculate its angle respective to the tumour/stroma boundary. For clarity, the CAF and ECM angles distribution for the IN region are not plotted, as CAF and ECM mostly give background signals in this area.
To correlate the orientation of fibroblasts and NK cell trajectories, NK cell tracks were divided into segments from time point (t) to (t + 4) and the orientation of each segment relative to the tumour/stroma boundary was then calculated similarly as the CAF and ECM orientation. To analyse the correlation between NK cell tracks and CAFs, the orientation of each segment corresponding to NK cell tracks and its nearest neighbour fibroblast vector (θtrack and θCAF) were saved as pairs (n = 1260 pairs, 4 independent experiments). To measure the Pearson correlation coefficient with 95% confidence interval we performed a bootstrap analysis. Specifically, 10% of the NK cell track-CAF-pairs were randomly drawn without repetitions and the correlation coefficient was calculated. The operation was then repeated 1000 times and the mean value with 95% confidence interval was computed from the resulting distribution. To further test the null hypothesis, the same bootstrap analysis was performed by using the same algorithm and data set but drawing a random θCAF for each selected θtrack.
Immune cell infiltration quantification
For PBMC and NK cells count in the stroma (EDGE + OUT) and infiltrated in the tumour (IN) cells were counted using the Trackmate Fiji plugin69 based on the fluorescent signal of the cells using confocal images acquired with 20× objective. For long-term acquisition of PBMCs infiltrating the tumour compartment, time-lapse acquisitions were performed at the rate of 1 acquisition/15 min for 62.5 h starting on day 7 using a 10× objective. The quantification of immune cell infiltration over time was performed using binarized fluorescent images of PBMCs and cancer cells counting for each image the number of particles corresponding to PBMCs overlapping the tumour compartment.
To measure the signal from PBMCs exiting the channels, binarized fluorescent images of PBMCs were generated at each time point. For each acquisition, a region of interest (ROI) (100 µm length × 200 µm width) was positioned at the exit of one channel facing a tumour spheroid. The average value of the fluorescent signal over time was retrieved in the ROI and the mean value over all the acquisitions was calculated for each condition. The relative percentage increase of the signal over time was then calculated.
Cell death assay
To assess cancer cell death, on day 10 after CAF seeding the MIRO co-culture was incubated at 37 °C for 4 h with the cell viability imaging kit following the manufacturer’s protocol (ReadyProbes Cell Viability Imaging Kit, Thermo Fisher, R37609). NucGreen® Dead reagent stains only the nuclei of cells with compromised plasma membrane integrity. Quantification of the cancer cell death was performed by measuring the mean intensity of the NucGreen® Dead reagent signal in the IN region for the different drug treatments. Values were normalised to the mean value of the control conditions in the presence or absence of NK cells and PBMCs respectively. For the CAF cell death assay, a similar procedure was performed using propidium iodide from ReadyProbes Cell Viability Imaging Kit (Thermo Fisher, R37610). The mean intensity of the CAF death signal was measured in regions populated by the CAF layer, choosing an area in the inlet at the vicinity of the channels. Each value was normalised to the mean value of fluorescence in the absence of treatment.
To quantify ADCC lytic events mediated by NK cells, the cell viability reporter was added to the cell media 4 h before the acquisition (ReadyProbes Cell Viability Imaging Kit, Thermo Fisher, R37609). To determine the number of lytic events, the increase of fluorescent green signal in the nucleus of cancer cells was monitored through confocal time-lapse acquisition (1 frame/4 min). Cell death events with previous contact (NK cells arrest within 0–3 µm from the nucleus of the target cell) with NK cells were manually counted. To discriminate non-ADCC-mediated cell death or non-traceable ADCC events, only events of cell death that correlated with the presence of NK cells within 0–3 µm from the target cell were quantified. To determine the cancer cell time to death, for each ADCC event counted, the time interval between the first visible contact of NK and the first frame of green signal appearance in the nucleus was calculated.
Immunofluorescence, immunohistochemistry and picrosirius red staining
Cells were fixed in 4% paraformaldehyde (PFA; Electron Microscopy Sciences) for 10 min at room temperature (RT) and permeabilized with 0.1% Triton X-100 (Sigma-Aldrich) for 10 min at RT. Cells were blocked in 1% BSA (Signa-Aldrich) for 1 h at RT before being incubated for 1 h at RT with primary antibodies (see Table 1). After incubation for 1 h at RT with secondary antibodies (see Table 1) or phalloidin, nuclei were stained (Hoechst 33342, Invitrogen) at RT for 20 min. The samples were then washed and mounted with MOWIOL reagent (Merck Life Science).
Immunohistochemistry was carried out using 4 μm tissue sections according to standard procedures70. Briefly, after antigen retrieval, samples were blocked with Peroxidase-Blocking Solution (Dako, S202386) for 10 min at RT. Primary antibodies (see Table 1) were incubated overnight at 4 °C. Slides were washed with EnVision™ FLEX Wash Buffer (Dako, K800721) and the corresponding secondary antibody was incubated with the sample for 45 min at RT. Samples were developed using 3,3′-diaminobenzidine, counterstained with hematoxylin and mounted. Staining evaluation and acquisition were performed with QuPath software.
For Picrosirius Red staining, samples were stained with Thiosemicarbazide 99% (TSC) (Sigma, T33405) for 10 min and washed in distilled water. Next, samples were stained with 0.5% direct Red 80 (365548, Sigma) solution in Piric Acid Solution 1.3% (P6744-1GA, Sigma) for 90 min and rinsed with 1% acetic acid (Sigma, 320099) for 1 min. Samples were dehydrated and mounted with Mounting Medium, Toluene-Free (CS705, Dako, Agilent) using a Dako CoverStainer.
Laser ablation live imaging experiments
A laser ablation system was implemented as previously described71. In brief, a 470 picosecond-pulsed laser (Nd: YAG at 355 nm, “High-Power Chip”, Teem Photonics, France) was focused through a 40×/1.2 W objective lens to achieve plasma-induced ablation of several cells inducing cell death and removal from the ablated area. In the current hardware configuration, the beam is fixed, and the stage of the microscope enables the ablation laser to raster a user-defined spline-line shape, distributing laser pulses at about 5 pulses/μm at a frequency of about 100 Hz. Images were acquired with an ORCA Flash4 v2 SCMOS camera (Hamamatsu, Japan) and a lens-less spinning disk enables confocal imaging. Fluorescence is achieved with 488 and 561 solid-state lasers (Omicron Laserage Gmbh, Germany) and fluorescence filters (525/50, 609/54, Chroma). For the ablation of the EDGE regions, one line is scanned two times across the sample to achieve ablation. For the OUT regions, one scan was performed to achieve ablation. Up to three individual scans are achieved, each of them performed in separate planes, distant 10 μm axially, with increased laser frequency up to 500 Hz and increased scanning speed to maintain the laser pulse density and to minimise the time between pre-and post-ablation. The energy per pulse used in all experiments ranges from 400 nJ to 800 nJ.
To quantify the recoil time after laser ablation in the EDGE and OUT regions, each time-lapse acquisition (1 frame/0.63 to 0.68 s, total time analysed: 200 s) was converted into kymographs using the ImageJ Plugin Kymographbuilder. In brief, a line was drawn along the ablation path and positioned perpendicularly to the cut at the central region of the ablated region. Detection of the recoil edge on kymographs was performed with a semi-automatic ImageJ macro, using Wand and Lasso graphical tools to create a mask of the recoil gap. Recoil values and statistics were analysed in Igor Pro (Wavemetrics, v6.04). Curve fitting was done on each recoil record with a double exponential model that precisely recapitulates the recoil dynamics, yielding 5 parameters: y0 (recoil at t = ∞), A1 and Tau1 (slow exponential recoil), A2 and Tau2 (fast exponential recoil). (y0-A1) expresses the fast recoil amplitude in microns happening within a few seconds after ablation, while A1 expresses the amplitude in microns of the durable recoil happening within Tau1 (beyond 100 s). Comparisons of Tau1, (y0-A1) and y0 quantities were performed with the Wilcoxon test for non-normally distributed data, yielding a reported P-value below alpha = 0.05 when distributions were significantly different.
Immune cell proliferation assay
Immune cells were seeded in RPMI media containing 0.2% FBS and 1% L-glutamine and were treated with Trastuzumab (20 ng/mL), IL2 (20 ng/mL), or a combination of both. Cells were cultured at 37 °C with 5% CO2, 20% O2, and a relative humidity of 95% for 96 h. At specified time points, cell counts were conducted using a Neubauer chamber under a light microscope to assess the cell expansion rate over time excluding apoptotic and necrotic cells using trypan blue dye (Thermo Fisher Scientific).
Real-time reverse transcription quantitative PCR (RT-qPCR)
RNA was extracted using Trizol Reagent (Invitrogen) and isolated with the RNeasy Micro Kit (QIAGEN, Hilfen, Germany) following the manufacturers’ handbook. Reverse transcription was performed using High Capacity cDNA Reverse Transcription Kit (Applied Biosystems, Thermo Fisher Scientific, Pleasanton, CA). Quantitative real-time PCR was performed using TaqMan assays (Applied Biosystems) (PRF1, GZMB, FN1, COL1A and COL4A1) or SYBR Green (Thermo Fisher Scientific) (Table 2) following manufacturers’ instructions in a 7900HT Fast Real-Time System (Applied Biosystems). PPIA was used as a housekeeping gene.
To evaluate the expression of ADCC markers, NK cells or PBMCs extracted from healthy donors were inoculated with 0.2% FBS RPMI medium to a monoculture of HCC1954 (after its fourth day of growth) and treated with 20 mg/ml of Trastuzumab, and/or 100 ng/ml of Interleukin 2 (IL2). After 24 h, the medium containing the immune cells was collected and PRF1 and GZMB expression was quantified. To assess the expression of integrins in NK cells, HCC1954 cells were grown first. After expansion, the medium was changed to 0.2% FBS RPMI, and the conditioned medium (CM) was collected after 24 h. After two days, they were treated with 20 mg/ml of Trastuzumab, and/or 100 ng/ml of IL2 with 0.2% FBS RPMI medium and the CM (treated CM) was collected 24 h later. CAFs were grown on a different plate. On days 4 and 5, the medium was changed for 24 h CM. On day 6, NK cells or PBMCs extracted from a healthy donor were inoculated together with the treated CM. After 24 h, the medium (containing the immune cells) was collected and the expression of ITGAM and ITGAL genes was analysed. Finally, FN1, COL1A and COL4A1 were analysed on CAFs exposed to 20 mg/ml of Trastuzumab, and/or 100 ng/ml of IL2 with 0.2% FBS DMEM medium for 4 days. To evaluate the genetic profile of all the fibroblast cell lines, fibroblasts were seeded in 2D and cultured in 10% FBS for 3 days before harvesting.
Clinical material
Biological samples were obtained from 12 BC patients from Parc de Salut MAR Biobank (MARBiobanc, Barcelona) under informed consent and approval of the Bank Tumour Committee according to Ethical regulations. International standards of Ethical Principles for Medical Research Involving Human subjects (code of ethics, Declaration of Helsinki, Fortaleza, Brazil, October 2013) were followed under legal regulations on data confidentiality (Organic Law 3/2018 -December the 5th- on the Protection of Personal Data and Digital Rights Guarantee) and on biomedical research (Law 14/2007 -July the 3rd-).
Statistics and reproducibility
Statistical analysis was performed using GraphPad Prism (v.10.0.2) for Windows, GraphPad Software, Boston, Massachusetts USA, www.graphpad.com. Normality and lognormality tests were used to establish the appropriate significance test, followed by a statistical test to compare the means. To compare the means of two groups, unless specified otherwise, a Student’s t-test was performed. For data with more than one variable, unless otherwise specified analysis of variance tests (ANOVA) or mixed effects analysis followed by multiple comparisons tests were applied. Unless otherwise specified, all error bars shown are mean ± standard deviation and representative images refer to at least three independent samples. No statistical method was used to predetermine sample size. No data were excluded from the analyses and unless otherwise specified experiments were not randomised. The investigators were not blinded to allocation during experiments and outcome assessment. Differences were considered statistically significant at *p < 0.05, **p < 0.01, ***p < 0.001 and ****p < 0.0001.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Data are available within the Article, Supplementary Information or Source Data file. The source data underlying Figs. 1g, h, 2d, e, 3a–h, 4c, 5c–e, 6a–e, 7e, f, h, 8b–e, g and Supplementary Figs. 2a, b, 3b, 4d, f, g, 6d–f, 7a, b, 9, 10b, 11, 12a, c, 13, 14a, c–f, 15a, b are provided as a Source Data file. Source data are provided with this paper.
Code availability
The codes generated for this study can be found at: https://github.com/xt-prc-lab/Perucca_et_al_2024_Nature_Communications.git.
References
Denkert, C. et al. Tumour-infiltrating lymphocytes and prognosis in different subtypes of breast cancer: a pooled analysis of 3771 patients treated with neoadjuvant therapy. Lancet Oncol. 19, 40–50 (2018).
Lundgren, S. et al. The prognostic impact of NK/NKT cell density in periampullary adenocarcinoma differs by morphological type and adjuvant treatment. PLoS ONE 11, e0156497 (2016).
Galon, J. & Bruni, D. Approaches to treat immune hot, altered and cold tumours with combination immunotherapies. Nat. Rev. Drug Discov. 18, 197–218 (2019).
Subbiah, I. M. & Gonzalez-Angulo, A. M. Advances and future directions in the targeting of HER2-positive breast cancer: implications for the future. Curr. Treat. Options Oncol. 15, 41–54 (2014).
Giraldo, N. A. et al. The clinical role of the TME in solid cancer. Br. J. Cancer 120, 45–53 (2018).
Raskov, H., Orhan, A., Christensen, J. P. & Gögenur, I. Cytotoxic CD8+ T cells in cancer and cancer immunotherapy. Br. J. Cancer 124, 359–367 (2021).
Vizoso, M. et al. Aberrant DNA methylation in non-small cell lung cancer-associated fibroblasts. Carcinogenesis 36, 1453–1463 (2015).
Labernadie, A. et al. A mechanically active heterotypic E-cadherin/N-cadherin adhesion enables fibroblasts to drive cancer cell invasion. Nat. Cell Biol. 19, 224–237 (2017).
Takahashi, H. et al. Cancer-associated fibroblasts promote an immunosuppressive microenvironment through the induction and accumulation of protumoral macrophages. Oncotarget 8, 8633–8647 (2017).
Linares, J., Marín-Jiménez, J. A., Badia-Ramentol, J. & Calon, A. Determinants and functions of CAFs secretome during cancer progression and therapy. Front. Cell Dev. Biol. 8, 621070 (2021).
Cox, T. R. The matrix in cancer. Nat. Rev. Cancer 21, 217–238 (2021).
Park, D. et al. Extracellular matrix anisotropy is determined by TFAP2C-dependent regulation of cell collisions. Nat. Mater. 19, 227–238 (2019).
Ray, A. & Provenzano, P. P. Aligned forces: origins and mechanisms of cancer dissemination guided by extracellular matrix architecture. Curr. Opin. Cell Biol. 72, 63–71 (2021).
Salmon, H. et al. Matrix architecture defines the preferential localization and migration of T cells into the stroma of human lung tumors. J. Clin. Invest. 122, 899–910 (2012).
Sun, X. et al. Tumour DDR1 promotes collagen fibre alignment to instigate immune exclusion. Nat 599, 673–678 (2021).
Grout, J. A. et al. Spatial positioning and matrix programs of cancer-associated fibroblasts promote T cell exclusion in human lung tumors. Cancer Discov. 12, 2606–2625 (2022).
Enfield, K. S. S. et al. Spatial architecture of myeloid and T cells orchestrates immune evasion and clinical outcome in lung cancer. Cancer Discov. 14, 1018–1047 (2024).
Infanger, D. W., Lynch, M. E. & Fischbach, C. Engineered culture models for studies of tumor-microenvironment interactions. Annu Rev. Biomed. Eng. 15, 29–53 (2013).
Sunyer, R. et al. Collective cell durotaxis emerges from long-range intercellular force transmission. Science https://doi.org/10.1126/science.aaf7119 (2016).
Conti, S. et al. CAFs and cancer cells co-migration in 3D spheroid invasion assay. In Methods in Molecular Biology, Vol. 2179, 243–256 (Humana Press Inc., 2020).
Yamada, K. M., Doyle, A. D. & Lu, J. Cell–3D matrix interactions: recent advances and opportunities. Trends Cell Biol. 32, 883–895 (2022).
Rodrigues, J., Heinrich, M. A., Teixeira, L. M. & Prakash, J. 3D in vitro model (r)evolution: unveiling tumor-stroma interactions. Trends cancer 7, 249–264 (2021).
Sontheimer-Phelps, A., Hassell, B. A. & Ingber, D. E. Modelling cancer in microfluidic human organs-on-chips. Nat. Rev. Cancer 19, 65–81 (2019).
Moya, M. L., Hsu, Y. H., Lee, A. P, Christopher, C. W. H. & George, S. C. In vitro perfused human capillary networks. Tissue Eng. Part C Methods 19, 730–737 (2013).
Jeon, J. S. et al. Human 3D vascularized organotypic microfluidic assays to study breast cancer cell extravasation. Proc. Natl. Acad. Sci. USA 112, 214–219 (2015).
Choi, Y. et al. A microengineered pathophysiological model of early-stage breast cancer. Lab Chip15, 3350–3357 (2015).
Chen, M. B. et al. On-chip human microvasculature assay for visualization and quantification of tumor cell extravasation dynamics. Nat. Protoc. 12, 865–880 (2017). 125. 2017.
Parlato, S. et al. 3D Microfluidic model for evaluating immunotherapy efficacy by tracking dendritic cell behaviour toward tumor cells. Sci. Rep. 7, 1093 (2017).
Nguyen, M. et al. Dissecting effects of anti-cancer drugs and cancer-associated fibroblasts by on-chip reconstitution of immunocompetent tumor microenvironments. Cell Rep. 25, 3884–3893.e3 (2018).
Lee, S. W. L. et al. Characterizing the role of monocytes in T cell cancer immunotherapy using a 3d microfluidic model. Front Immunol. 9, 416 (2018).
Ayuso, J. M. et al. Microfluidic tumor-on-a-chip model to evaluate the role of tumor environmental stress on NK cell exhaustion. Sci Adv. https://doi.org/10.1126/SCIADV.ABC2331 (2021).
Brett, E. A., Sauter, M. A., Machens, H.-G. & Duscher, D. Tumor-associated collagen signatures: pushing tumor boundaries. Cancer Metab. 8, 1–5 (2020).
Kollmannsberger, P., Bidan, C. M., Dunlop, J. W. C., Fratzl, P. & Vogel, V. Tensile forces drive a reversible fibroblast-to-myofibroblast transition during tissue growth in engineered clefts. Sci. Adv. 4, eaao4881 (2018).
Ferruzzi, J. et al. Compressive remodeling alters fluid transport properties of collagen networks – implications for tumor growth. Sci. Rep. 9, 1–16 (2019).
Barbazan, J. et al. Cancer-associated fibroblasts actively compress cancer cells and modulate mechanotransduction. Nat Commun. https://doi.org/10.1038/s41467-023-42382-4 (2023).
Gomes, R. N., Manuel, F. & Nascimento, D. S. The bright side of fibroblasts: molecular signature and regenerative cues in major organs. npj Regen. Med. 6, 1–12 (2021).
Joshi, R. S. et al. The role of cancer-associated fibroblasts in tumor progression. Cancers 13, 1–27 (2021).
Cords, L. et al. Cancer-associated fibroblast classification in single-cell and spatial proteomics data. Nat Commun. https://doi.org/10.1038/s41467-023-39762-1 (2023).
Hudis, C. A. Trastuzumab - mechanism of action and use in clinical practice. N. Engl. J. Med. 357, 39–51 (2007).
Ahmed, S., Sami, A. & Xiang, J. HER2-directed therapy: current treatment options for HER2-positive breast cancer. Breast Cancer 22, 101–116 (2015).
Force, J. et al. Early stage HER2-positive breast cancers not achieving a pCR from neoadjuvant trastuzumab- or pertuzumab-based regimens have an immunosuppressive phenotype. Clin. Breast Cancer 18, 410–417 (2018).
Rivas, E. I. et al. Targeted immunotherapy against distinct cancer-associated fibroblasts overcomes treatment resistance in refractory HER2+ breast tumors. Nat Commun. https://doi.org/10.1038/s41467-022-32782-3 (2022).
Mullard, A. Restoring IL-2 to its cancer immunotherapy glory. Nat. Rev. Drug Discov. 20, 163–165 (2021).
Gismondi, A. & Santoni, A. Migration of NK cells. In Lymphocyte Trafficking in Health and Disorder 95–112 https://doi.org/10.1007/3-7643-7442-x_6 (2006).
Hornung, A. et al. A bistable mechanism mediated by integrins controls mechanotaxis of leukocytes. Biophys. J. 118, 565 (2020).
Wolf, K., Müller, R., Borgmann, S., Bröcker, E. B. & Friedl, P. Amoeboid shape change and contact guidance: T-lymphocyte crawling through fibrillar collagen is independent of matrix remodeling by MMPs and other proteases. Blood 102, 3262–3269 (2003).
Bhattacharjee, O., Ayyangar, U., Kurbet, A. S., Ashok, D. & Raghavan, S. Unraveling the ECM-immune cell crosstalk in skin diseases. Front. Cell Dev. Biol. 7, 68 (2019).
Orange, J. S. et al. IL-2 induces a WAVE2-dependent pathway for actin reorganization that enables WASp-independent human NK cell function. J. Clin. Invest. 121, 1535–1548 (2011).
Olofsson, P. E. et al. Distinct migration and contact dynamics of resting and IL-2-activated human natural killer cells. Front Immunol. 5, 80 (2014).
Lo Nigro, C. et al. NK-mediated antibody-dependent cell-mediated cytotoxicity in solid tumors: biological evidence and clinical perspectives. Ann. Transl. Med. 7, 105–105 (2019).
Olofsson, P. E. et al. A collagen-based microwell migration assay to study NK-target cell interactions. Sci. Rep. 9, 1–10 (2019).
Menon, A. G. et al. A basal membrane-like structure surrounding tumour nodules may prevent intraepithelial leucocyte infiltration in colorectal cancer. Cancer Immunol. Immunother. 52, 121–126 (2003).
Ioachim, E. et al. Immunohistochemical expression of extracellular matrix components tenascin, fibronectin, collagen type IV and laminin in breast cancer: their prognostic value and role in tumour invasion and progression. Eur. J. Cancer 38, 2362–2370 (2002).
Diab, A. et al. Bempegaldesleukin (NKTR-214) plus nivolumab in patients with advanced solid tumors: Phase I dose-escalation study of safety, efficacy, and immune activation (PIVOT-02). Cancer Discov. 10, 1158–1173 (2020).
Ptacin, J. L. et al. An engineered IL-2 reprogrammed for anti-tumor therapy using a semi-synthetic organism. Nat. Commun. 12, 1–14 (2021).
Quayle, S. N. et al. Cue-101, a novel E7-pHLA-IL2-Fc fusion protein, enhances tumor antigen-specific T-cell activation for the treatment of HPV16-driven malignancies. Clin. Cancer Res. 26, 1953–1964 (2020).
Vanherberghen, B. et al. Classification of human natural killer cells based on migration behavior and cytotoxic response. Blood 121, 1326–1334 (2013).
Kalluri, R. The biology and function of fibroblasts in cancer. Nat. Rev. Cancer 16, 582–598 (2016).
Melssen, M. M., Sheybani, N. D., Leick, K. M. & Slingluff C. L. Barriers to immune cell infiltration in tumors. J, Immunother, Cancer https://doi.org/10.1136/jitc-2022-006401 (2023).
Voabil, P. et al. An ex vivo tumor fragment platform to dissect response to PD-1 blockade in cancer. Nat. Med. 27, 1250–1261 (2021).
Lo Re, D. et al. Increased immune cell infiltration in patient-derived tumor explants treated with Traniplatin: an original Pt(iv) pro-drug based on Cisplatin and Tranilast. Chem. Commun. 54, 8324–8327 (2018.
Idrisova, K. F., Simon, H. U. & Gomzikova, M. O. Role of patient-derived models of cancer in translational oncology. Cancers https://doi.org/10.3390/cancers15010139 (2023).
Yang, C. et al. Heterogeneity of human bone marrow and blood natural killer cells defined by single-cell transcriptome. Nat. Commun. 10, 1–16 (2019).
Fernández-Nogueira, P. et al. Tumor-associated fibroblasts promote HER2-targeted therapy resistance through FGFR2 activation. Clin. Cancer Res. 26, 1432–1448 (2020).
Edelstein, A., Amodaj, N., Hoover, K., Vale, R. & Stuurman, N. Computer control of microscopes using µManager. Curr. Protoc. Mol. Biol. 92, 14.20.1–14.20.17 (2010).
MathWorks. MATLAB version R2021b, Natick, Massachusetts: The MathWorks Inc. (2021).
Diel, E. E., Lichtman, J. W. & Richardson, D. S. Tutorial: avoiding and correcting sample-induced spherical aberration artifacts in 3D fluorescence microscopy. Nat. Protoc. 15, 2773–2784 (2020).
Bredfeldt, J. S. et al. Automated quantification of aligned collagen for human breast carcinoma prognosis. J. Pathol. Inform. 5, 28 (2014).
Tinevez, J. Y. et al. TrackMate: an open and extensible platform for single-particle tracking. Methods 115, 80–90 (2017).
Kim, S. W., Roh, J. & Park, C. S. Immunohistochemistry for pathologists: protocols, pitfalls, and tips. J. Pathol. Transl. Med. 50, 411–418 (2016).
Colombelli, J. et al. Erratum: Mechanosensing in actin stress fibers revealed by a close correlation between force and protein localization (Journal of Cell Science vol. 122 (1665-1679)). J. Cell Sci. 122, 1928 (2009).
Acknowledgements
We thank S. Usieto, S. Menendez Romero, D. Zafra Puerta, A. Menendez, D. Izquierdo Garcia, C. Rols, J. Durand for their technical assistance. A.L. acknowledges funding from “La Caixa” Banking Foundation (LCF/BQ/PR18/11640001) and from the Spanish Ministry for Science and Innovation MICIU/AEI /10.13039/501100011033 and “ERDF-EU A way of making Europe”, (PID2021-128635NB-I00), and “ESF Investing in your future”, (RYC2020/029736/I). This work was also supported by grants from Instituto de Salud Carlos III (ISCIII) co-funded by the European Union (EU) (PI17/00211, CP16/00151, PI20/00011; Spanish Ministry of Economy and Competitiveness) to A.C. A.C. is the recipient of Miguel Servet research contracts from Instituto de Salud Carlos III co-funded by the European Union (MS16/00151, CPII21/00012). J.A. acknowledges funding from the Generalitat de Catalunya (AGAUR 2021 SGR 00776), the CIBERONC (CB16/12/00241), and the ISCIII co-funded by the EU (PI21/00002). This work was also supported by the Plan de Recuperación, Transformación y Resiliencia (CNS2023-145267) co-funded by the European Union to A.C. P.R.-C. acknowledges funding from the Spanish Ministry of Science and Innovation (PID2022-142672NB-I00), the European Research Council (AdG 101097753), the Generalitat de Catalunya (2017-SGR-1602), and the prize ‘ICREA Academia’ for excellence in research. X.T. acknowledges funding from the Generalitat de Catalunya (AGAUR SGR-2017-01602), the CERCA Programme, the Spanish Ministry for Science and Innovation MICCINN/FEDER (PID2021-128635NB-I00 MCIN/AEI/ 10.13039/501100011033 and “ERDF-EU A way of making Europe”), European Research Council (Adv-883739), Fundació la Marató de TV3 (project 201903-30-31-32), European Commission (H2020-FETPROACT-01-2016-731957), La Caixa Foundation (LCF/PR/HR20/52400004 and ID 100010434 under the agreement LCF/PR/HR20/52400004); IBEC and IRB are recipient of a Severo Ochoa Award of Excellence from the MINECO. This work was also supported by Grant PT20/00023, funded by Instituto de Salud Carlos III (ISCIII) and co-funded by the European Union, and the “Xarxa de Bancs de tumors” sponsored by Pla Director d’Oncologia de Catalunya (XBTC).
Author information
Authors and Affiliations
Contributions
Conceptualisation study: A.L., A.C. Conceptualisation and realisation of MIRO: A.L.; MIRO experiment realisation: A.P., A.L., O.M.B., C.H., E.D.; MIRO puncher realisation: X.B., J.P.; MIRO photolithography: Ju.L., M.C., T.G.; MIRO formal analysis: A.P., A.L., O.M.B., C.H., T.G.; Computational macro design: C.H., T.G.; Laser ablation experiment realisation: J.C.; Laser ablation analysis: J.C., A.L.; Primary fibroblasts derivation: J.A., A.R.; Fibroblast cells genetic engineering: P.F.N.; Immune cell preparation and in vitro experimentation: A.G.L., Je.L., E.R., A.R.B.; Cancer cells genetic engineering: Je.L., E.R.; Patient samples collection: L.C., J.A.; Patient sample preparation: A.G.L.; Immunohistochemistry: M.Ga., A.S.A.; Patient sample imaging: A.G.L.; RT-qPCR and analyses: A.G.L., A.R.B.; Funding: A.L., A.C., X.T.; Resources: A.L., J.A., L.C., A.C., P.R.C., X.T.; Writing – Original Draft: A.L., A.P.; Editing: A.L., A.P., A.C., A.G.L., X.T.
Corresponding authors
Ethics declarations
Competing interests
J.A. has received consulting fees and honoraria from Seagen, Pfizer, AstraZeneca, Lilly, Merck, Roche, Gilead, Novartis and Daiichi-Sankyo, receives royalties from a licensed patent to Biocartis (EP11382270.4; Mutations in the epidermal growth factor receptor gene) and holds stock options from Inbiomotion. L.C. reports receiving personal fees from Roche, MSD, AstraZeneca and Diaceutics, non-financial support from Roche, MSD, AstraZeneca and Phillips and held advisory roles for Roche and AstraZeneca. Other authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Feng Guo, Roger Kamm and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Perucca, A., Llonín, A.G., Benach, O.M. et al. Micro Immune Response On-chip (MIRO) models the tumour-stroma interface for immunotherapy testing. Nat Commun 16, 1279 (2025). https://doi.org/10.1038/s41467-025-56275-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-56275-1