Abstract
The precise regulation of the RIG-I-like receptors (RLRs)-mediated type I interferon (IFN-I) activation is crucial in antiviral immunity and maintaining host immune homeostasis in the meantime. Here, we identify an E3 ubiquitin ligase, namely RNF167, as a negative regulator of RLR-triggered IFN signaling. Mechanistically, RNF167 facilitates both atypical K6- and K11-linked polyubiquitination of RIG-I/MDA5 within CARD and CTD domains, respectively, which leads to degradation of the viral RNA sensors through dual proteolytic pathways. RIG-I/MDA5 conjugated with K6-linked ubiquitin chains in CARD domains is recognized by the autophagy cargo adaptor p62, that delivers the substrates to autolysosomes for selective autophagic degradation. In contrast, K11-linked polyubiquitination in CTD domains leads to proteasome-dependent degradation of RLRs. Thus, our study clarifies a function of atypical K6- and K11-linked polyubiquitination in the regulation of RLR signaling. We also unveil an elaborate synergistic effect of dual proteolysis systems to control amplitude and duration of IFN-I activation, hereby providing insights into physiological roles of the cross-talk between these two protein quality control pathways.
Similar content being viewed by others
Introduction
The innate immunity serves as the first line of host defense against microbial invasion1. After viral infection, the viral components are predominantly recognized by pattern recognition receptors (PRRs) such as Toll-like receptors (TLRs)2, NOD-like receptors (NLRs), and RIG-I-like receptors (RLRs), including RIG-I, MDA5 and LGP23,4,5,6. Subsequently, the mitochondrial antiviral signaling protein (MAVS) is recruited to RLRs through interactions within the caspase activation and recruitment domains (CARDs) in the N-terminal region7,8,9. Then, the kinase TBK1 is activated, resulting in phosphorylation of the transcription factors IRF3 and IRF7 to induce the production of IFN-I and IFN-stimulated genes (ISGs), and ultimately resulting in viral clearance10.
The activation of IFN-I response is pivotal for viral elimination. However, prolonged IFN activation may cause autoimmune diseases, like systemic lupus erythematosus (SLE)11. Therefore, the PRRs, adaptors and the other proteins in type I IFN signaling pathway must be elaborately regulated. Post-translational modifications (PTMs), such as phosphorylation, dephosphorylation, SUMOylation, ISGylation and ubiquitination have received much attention12,13,14. Among them, the modification of ubiquitination is most extensively studied. Ubiquitin, a 76-amino acid polypeptide with 7 lysine residues (K6, K11, K27, K29, K33, K48 and K63), is conjugated to the substrate proteins15. K27-, K48- and K63-linked polyubiquitinations are commonly identified in the regulation of IFN-I pathway, whereas the other ubiquitination types are scarcely reported16,17. After the modification by ubiquitination, the substrate proteins are recognized and degraded through the ubiquitin-proteasome system (UPS) or the autophagy-lysosome pathway (ALP), the two primary pathways governing protein turnover in eukaryotes. There is accumulating evidence that the UPS and autophagy are simultaneously affected and may mutually affect each other in a variety of diseases, especially in disorders of the central nervous system18. Nevertheless, the interplay between UPS and ALP in innate immunity still remains largely unexplored.
RING finger protein 167 (RNF167) is a type I transmembrane protein that belongs to the family of the protease-associated (PA) domain, the transmembrane domain (TM) and the RING-H2 finger (Really Interesting New Gene, RING) domain (PA-TM-RING) E3 ligases, which also includes RNF128, RNF133, RNF148, RNF149, RNF150, RNF13 and RNF20419. RNF167 has been implicated in neurotransmission, endosomal trafficking and lysosome positioning20,21,22,23,24,25. Recent studies have also elucidated its role in modulating mTOR signaling in tumorigenesis26,27 and suppressing the TNF-α signaling through promoting the ubiquitination of Tollip28. However, its involvement in regulation of the other cellular signaling pathways, particularly antiviral innate immune responses, remains unknown.
Here, we show that viral infection induces the expression of RNF167; the latter can interact with RIG-I/MDA5. RNF167 catalyzes the atypical K6- and K11-linked polyubiquitination of RIG-I/MDA5, and subsequently directs them for degradation through dual proteolytic pathways of both UPS and ALP. It represents a more efficient mechanism to avoid the excessive activation of type I IFN signaling. Hereby, our results reveal a role of atypical K6- and K11-linked polyubiquitination in the regulation of RLR-triggered IFN-I activation, providing insights into the synergistic effect of UPS and ALP in antiviral immune responses. Our study shows that one E3 ligase can direct the substrate for degradation through both proteasome and lysosome pathways simultaneously.
Results
Genome-wide CRISPR/Cas9 screening identified RNF167 as a negative regulator of IFN-I signaling pathway
To identify cellular regulators of type I IFN signaling pathway, we conducted a genome-wide CRISPR library screen in our previous study29. Further validations confirmed the efficacy of our screen and highlighted several host factors critical for IFN regulation, including CCDC50, ASB1, and RNF16730. In order to investigate the biological roles of RNF167 in innate immune responses, we initially explored whether its expression was regulated in response to IFN activation. Human monocytic leukemia (THP-1) cells were treated with RNA mimics poly(I:C), or infected with Sendai virus (SeV), encephalomyocarditis virus (EMCV), the type I herpes simplex virus (HSV-1) or stimulated with the human recombinant IFN-β. The results showed that the expression of RNF167 protein was upregulated, with the level varying over time during treatment (Fig. 1a and Supplementary Fig. 1a). Notably, RNF167 is known to be glycosylated, resulting in a diffuse migration pattern with bands larger than the predicted molecular weight21. Our results displayed the presence of two bands at approximately 43 kDa and 55 kDa via RNF167 ectopic expression (Supplementary Fig. 1b). We then isolated mouse primary bone-marrow derived macrophages (BMDMs) and further confirmed the induced expression of RNF167 during viral infection and IFN activation (Fig. 1b and Supplementary Fig. 1c, d).
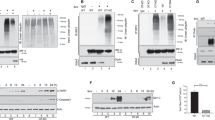
a, b Immunoblot analysis of RNF167 expression in THP-1 cells (a) and BMDM cells (b) treated with poly(I:C), SeV and recombinant IFN-β for indicated time points. The relative band density of RNF167 was measured by Quantity one and normalized to GAPDH, representing the protein expression levels of RNF167 (a, n = 3). c Immunoblot analysis of RNF167 expression in BMDMs treated with IFNAR1 antibody or IgG as control for 3 h and then left uninfected or infected with SeV for another 8 h. d Dual-luciferase assays in HEK293 cells that were transfected for 24 h with vector or RNF167-HA and then stimulated with SeV for 8 h. Luciferase values are relative to the values in uninfected mock cells (n = 3); western blots were probed with anti-HA and anti-GAPDH antibodies. e Dual-luciferase assays showing activities of indicated promoters in HEK293 cells transfected with siRNAs targeting RNF167 or the negative control non-targeting siRNA (siCtrl) for 24 h and then infected with SeV for another 8 h (n = 3). The expression of RNF167 was verified by immunoblot. Data are representative of three independent experiments and are shown as mean with SD (a, d, e). The unpaired two-tailed t-test was applied (ns, not significant). GAPDH or Actin was used as a loading control. Source data are provided as a Source Data file.
We then used the IFNAR1 (interferon alpha/beta receptor subunit 1) antibody to block the IFN-I pathway, and then detect the expression of RNF167. It showed that SeV infection could induce RNF167 expression, whereas the treatment with IFNAR1 antibody significantly blocked SeV-triggered RNF167 expression, suggesting that expression of RNF167 is IFN-dependent and it represents an uncharacterized ISG (Fig. 1c). These results proved that the IFN-I activation induces the expression of RNF167, indicating that RNF167 may play a role in antiviral innate immunity.
To characterize the potential role of RNF167 in antiviral IFN signaling, we performed IFN-β-promoter-based dual-luciferase assays with the RNF167 constructs. Ectopic expression of RNF167 significantly suppressed SeV-triggered activation of IFN-β, PRDI-III and NF-κB reporters in HEK293 cells (Fig. 1d). In contrast, RNF167 knockdown reinforced the activities of the promoters, suggesting that RNF167 inhibits the activation of IFN-1 signaling pathway (Fig. 1e). Altogether, viral infection and IFN-I treatment induce the expression of RNF167, that in turn suppresses the IFN-I signaling activity, indicating that RNF167 functions as a negative feedback regulator of IFN-I response.
RNF167 deficiency enhances the expression of antiviral genes and suppresses viral replication
To further illustrate the function of RNF167 in type I IFN-mediated antiviral responses, we generated the RNF167 gene knockout (KO) HEK293 cells using CRISPR/Cas9 system, and developed two distinct RNF167-deficient single cell clones, which exhibit similar growth rates to the wild-type cells (Fig. 2a and Supplementary Fig. 2a). As shown in Fig. 2b, the mRNA levels of IFNB1, downstream antiviral ISGs including CXCL10 and IFIT1 and proinflammatory genes CXCL1 and IL-6 were significantly higher in RNF167-deleted cells compared to those in wild-type control cells. To exclude the virus- or cell-type-specific phenomenon and ensure that RNF167 selectively inhibits the RLRs signaling pathway, we designed small interfering RNAs (siRNAs) targeting the human RNF167 (Fig. 2c) and mouse Rnf167 (Supplementary Fig. 2b), respectively. We then transfected siRNA into THP-1 cells, and verified that the expression of endogenous RNF167 was downregulated (Fig. 2c). Similarly, the siRNA-mediated knockdown of RNF167 promoted the transcription of IFNB1, ISGs and proinflammatory genes in THP-1 cells infected with VSV or transfected with poly(I:C) (Fig. 2d and Supplementary Fig. 2c), but had no effect in BMDM cells treated with exogenous poly(I:C) (Supplementary Fig. 2d), suggesting that RNF167 selectively inhibits the RLRs signaling pathway and its functions in regulation of antiviral responses are conserved. Consistently, in Raw264.7 cells infected with SeV, we observed the consistent result that RNF167 deficiency significantly upregulated mRNA expression of Ifnb1, Cxcl10, Ifit1, and Cxcl1. (Supplementary Fig. 2e).
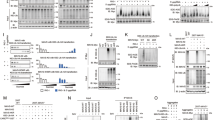
a qPCR and immunoblot analysis of RNF167 expression levels in wild type (WT) and RNF167-knockout HEK293 cells (n = 3). b qPCR analysis of mRNA expression levels of IFNB, the related ISGs and proinflammatory cytokines in wild-type and RNF167-deleted cells infected with SeV for 8 h (n = 3). c qPCR and immunoblot analysis of RNF167 expression levels in THP-1 cells transfected with siRNAs targeting RNF167 or the negative control non-targeting siRNA (siCtrl) (n = 3). d qPCR analysis of the mRNA expression levels of IFNB, the related ISGs and proinflammatory cytokines in THP-1 cells transfected with these siRNAs for 48 h and then infected with VSV for another 8 h (n = 3). e Dual-luciferase assays of the indicated promotor reporters in WT and KO (RNF167-deletion) cells transfected with RNF167-HA expression plasmid or its empty vector as control for 24 h and then infected with SeV for another 8 h. The expression of RNF167 was verified by immunoblot (n = 3). f Representative images of microscopy analysis in wild-type and RNF167-deleted cells infected with VSV-GFP for 24 h at the MOI of 0.1 (scale bars, 50 µm), and the VSV-GFP replication levels were presented in a graph by counting the infected cells (n = 3). g Virus titer detection by plaque assay of the cell supernatant in WT and RNF167-deletion cells infected with VSV-GFP in the MOI of 0.1 for 24 h (n = 3). Data are representative of three independent experiments with similar results, and are shown as mean with SD (a–g). The unpaired two-tailed t-test was applied. GAPDH or Actin were used as a loading control. Source data are provided as a Source Data file.
To rule out off-target effects, we reconstitute the expression of RNF167 in RNF167-KO HEK293 cells with an RNF167-expressing plasmid that is resistant to Cas9 cleavage. We found that RNF167 deletion significantly strengthened SeV-induced activation of IFN-β, PRDIII-I, and NF-κB reporters, whereas the enhancement effect of RNF167 ablation on the activation of promoters was reversed with the reconstitution of RNF167 (Fig. 2e). These results demonstrated that the deficiency of RNF167 enhanced viral infection-triggered anti-viral gene expression.
We then evaluated the effect of RNF167 on the replication of virus. As is shown in Fig. 2f, g and Supplementary Fig. 2f-h, RNF167 disruption attenuated VSV infection and replication in both HEK293 cells and mouse L929 cells. The fluorescence and the titer were significantly reduced in RNF167-deficient cells compared to those in wild-type cells. Taken together, RNF167 deficiency promotes viral infection-induced innate immunity and antiviral gene expression, thereby constraining the replication of the viruses.
RNF167 suppresses antiviral innate immune responses in mouse primary immune cells
Next, to examine the biological significance of RNF167, we generated Rnf167−/− mice on a C57BL/6J background using CRISPR/Cas9-based technology (Supplementary Fig. 3a). After verifying that the body sizes and weights of Rnf167-/- mice were similar to their wild-type littermates (Supplementary Fig. 3b), we concluded that the absence of RNF167 does not result in any significant developmental or growth differences under normal conditions. Thus, we prepared the BMDMs and BMDCs from the Rnf167−/− mice and their wild-type littermates, and confirmed the deletion of RNF167 (Supplementary Fig. 3c). Consistently, BMDMs and BMDCs isolated from Rnf167−/− mice showed significantly increased mRNA levels of Ifnb1, Cxcl10, Ifit1, Cxcl1 and Il6 after infection with SeV and VSV compared to those from the Rnf167+/+ counterparts (Fig. 3a, b). Moreover, the production and secretion of IFN-β protein was also higher in Rnf167−/− BMDMs and BMDCs infected by SeV and VSV than in Rnf167+/+ cells (Fig. 3c). In addition, we also treated the BMDMs with RNA mimics via transfection of high-molecular-weight and low-molecular-weight poly(I:C) into cells. We observed that the expression level of Ifnb1, Cxcl10 and Cxcl1 in RNF167−/− BMDMs was higher than in their wild-type counterparts (Fig. 3d). The time-course study also showed the increased expression of Ifnb1 in Rnf167−/− BMDMs than in Rnf167+/+ BMDMs at the indicated time points following SeV infection (Fig. 3e).
a, b qPCR analysis of the Ifnb1, Cxcl10, Ifit1, Cxcl1, and Il6 mRNA expressions levels in Rnf167+/+ and Rnf167-/- BMDMs/BMDCs infected with SeV and VSV for 8 h (n = 3). c ELISA quantification of IFN-β secretion in Rnf167+/+ and Rnf167-/- BMDMs/BMDCs infected with SeV and VSV for 12 h (n = 3). d qPCR analysis of the Ifnb1, Cxcl10, and Cxcl1 mRNA expression levels in WT/KO BMDMs transfected with the poly(I:C) of high molecular weight (HMW, -H) and low molecular weight (LMW, -L) for 12 h (n = 3). e qPCR analysis of the Ifnb1 induction by SeV infection in Rnf167+/+ and Rnf167-/- BMDMs at the indicated time points (n = 3). f, g Immunoblot analysis of the phosphorylated TBK1 and IRF3 in BMDMs/ BMDCs stimulated by SeV/VSV infection for 8 h, and poly(I:C)-H/L transfection for 12 h. Data are representative of three independent experiments with similar results and are shown as mean with SD (a–e). The unpaired two-tailed t-test was applied. GAPDH was used as a loading control. Source data are provided as a Source Data file.
We further detected the phosphorylation of the downstream molecules in signal transduction. Notably, the knockout of RNF167 increased the phosphorylation levels of TBK1 and IRF3 in both BMDMs and BMDCs upon treatment with SeV, VSV and poly(I:C) (Fig. 3f, g). Collectively, these results demonstrate that RNF167 inhibits the production of IFN-β and IFN-I-mediated antiviral immune response in mouse primary immune cells.
Rnf167-deficient mice exhibit higher viral resistance and improved survival from viral infection
To explore the physiological role of RNF167 in antiviral responses in vivo, we conducted viral infection experiments in a mouse model. The Rnf167−/− mice and wild-type Rnf167+/+ littermates were injected with VSV intravenously at a non-lethal dose (6 × 107 plaque-forming units per mouse). Rnf167−/− mice displayed enhanced expression of Ifnb1, Cxcl10, Ifit1, and Il6 in the liver, spleen, and lung in comparison with the wild-type control mice (Fig. 4a–c). The results of enzyme-linked immunosorbent assay (ELISA) also showed elevated levels of secreted IFN-β in serum from Rnf167−/− mice compared to in that of Rnf167+/+ mice (Fig. 4d). In contrast, the replication of the virus was significantly inhibited in different organs of Rnf167−/− mice (Fig. 4e–g). Furthermore, Rnf167−/− mice exhibited the increased ability of viral clearance, leading to attenuated tissue damage and inflammatory cell infiltration in the lungs after VSV infection (Fig. 4h). Similarly, Rnf167 deficiency protected mice from viral infection and, therefore, Rnf167−/− mice showed improved overall survival compared to their wild-type littermates when infected with a lethal dose of VSV (Fig. 4i). Altogether, these results demonstrated that RNF167 deficiency improved the clearance of virus and protects the mice by enhancing antiviral innate immune responses.
a–c qPCR analysis of Ifnb1, Cxcl10, Ifit1 and Il6 mRNA expression levels in the liver, spleen, and lung of the 6-week-old Rnf167+/+ and Rnf167-/- mice infected with VSV (6 × 107 PFU per mouse) by intravenous injection for 16 h (n = 3 per group). d IFN-β production in sera from Rnf167+/+ and Rnf167-/- mice treated as in (a–c) (n = 3 each group). e–g qPCR analysis of VSV replication levels in the liver, spleen, and lung from Rnf167+/+ and Rnf167-/- mice treated as in (a–c) (n = 3 each group). h Hematoxylin-eosin staining of lung sections from Rnf167+/+ and Rnf167-/- mice injected with VSV treated as in (a–c) (n = 3 per group). Scale bars, 50 µm. i Survival curves of 6-week-old WT and KO mice by intravenous injection (1.2 × 108 PFU per mouse, n = 9 each group). Data are representative of three independent experiments with similar results and shown as mean with SD (a–g), and the unpaired two-tailed t-test was applied. Data of survival curves was analyzed using Gehan-Breslow-Wilcoxon test in a two-sided manner (i). Source data are provided as a Source Data file.
RNF167 specifically targets RIG-I/MDA5/MAVS complex
We next explored the molecular mechanisms by which RNF167 suppressed type I IFN signaling. The exogenous expression of RNF167 significantly attenuated the activities of RIG-I-, MDA5-, and MAVS-triggered IFN-β, PRD-I-III, and NF-κB promoters, whereas RNF167 had no obvious effect on the TBK1-, IKKβ-, p65- and constitutively active phosphorylation mimetic IRF3-5D-induced reporter activation (Fig. 5a). In contrast, siRNA-mediated RNF167 knockdown markedly enhanced the activation of these signaling proteins-triggered IFN-β promotor (Fig. 5b and Supplementary Fig. 4a). These data suggested that the effective step of RNF167 was upstream of TBK1. Then, the association between RNF167 and RIG-I/MDA5/MAVS was verified by using immunoprecipitation assays. The result showed that RNF167 could interact with RIG-I, MDA5 and MAVS (Supplementary Fig. 4b). Moreover, endogenous immunoprecipitation assays showed that the interaction between RNF167 and RIG-I/MDA5 was increased upon viral infection and varied in a time-dependent manner (Fig. 5c). We also observed the interaction between RNF167 and MAVS with SeV infection, whereas RNF167 did not exhibit any affinity for TBK1 or IRF3 (Fig. 5d). Then we went to identify the domain of RNF167 responsible for the interaction with RIG-I/MDA5. RNF167 functions as an E3 ligase, which contains an N-terminal signal peptide (SP), a protease-associated (PA) domain, a transmembrane domain (TM), a really interesting new gene (RING) domain, and the C terminal domain (C). Four truncated mutants with PA, RING, C terminal domain or both RING and C terminal domains deletion of RNF167 were constructed (Fig. 5e). Co-immunoprecipitation experiments revealed that RNF167-ΔRING-ΔC lost its capability to interact with RIG-I and MDA5 and RNF167-ΔRING had strong while RNF167-ΔC had the weak interactions with RIG-I/MDA5, suggesting that the both RING domain and C terminal domain were used for the binding while C domain was in predominant (Fig. 5f and Supplementary Fig. 4c). Both RIG-I and MDA5 are composed of N-terminal CARDs, helicase and C-terminal domains (CTDs) (Fig. 5g and Supplementary Fig. 4d). Interestingly, each of the three domains could interact with RNF167 (Fig. 5h and Supplementary Fig. 4e). Altogether, these results demonstrated that RNF167 interacted with RIG-I, MDA5 and MAVS and specially interacted with RIG-I and MDA5 through its RING and C terminal domains.
a Dual-luciferase assays of the indicated promotor activities in HEK293 cells transfected with the empty vector (as control), RIG-I, MDA5, MAVS, TBK1, or IRF3-5D (TRAF6, IKK-β, and p65 for NF-κB-luc) expression plasmids together with RNF167-HA or its empty vector as control for 24 h (n = 3). b Dual-luciferase assays of IFN-β promotor activities in HEK293 cells transfected with negative control (siCtrl) or the siRNA targeting RNF167 (siRNF167) for 24 h, and then transfected with the transfection of empty vector (control), RIG-I, MDA5, MAVS, TBK1, or IRF3-5D expression plasmids for another 24 h (n = 3). c Immunoprecipitation and immunoblot analysis of endogenous RNF167 with RIG-I and MDA5 in HEK293 cells infected with SeV for the indicated time, and the IgG was set as control. d Immunoprecipitation and immunoblot analysis of endogenous RNF167 with IgG (control), MAVS, TBK1, and IRF3 in HEK293 cells infected with SeV for indicated time points. e Schematic structures of human RNF167. SP, signal peptide; PA, protease-associated domain; TM, transmembrane domain, RING domain, and the C terminal domain. f Coimmunoprecipitation and immunoblot analysis of IgG control, Flag-tagged empty vector or Flag-RIG-I with RNF167-HA or its mutants ΔPA/ΔRING/ΔC /ΔR-ΔC (deletion of PA, RING, C, or both RING-C domains) plasmids transfected into HEK293 cells. g Schematic structures of human RIG-I. h Coimmunoprecipitation and immunoblot analysis of IgG control, Flag-tagged empty vector, Flag-RIG-I-CARDs/Helicase/CTD or -ΔCARDs/ΔHelicase/ΔCTD (domains remained and deleted mutants) with RNF167-HA transfected into HEK293 cells. Data are representative of three independent experiments and are shown as mean with SD (a, b). The unpaired two-tailed t-test was applied (ns, not significant). GAPDH was used as the loading control. Source data are provided as a Source Data file.
RNF167 decreased the protein stability of RIG-I and MDA5
As shown in Fig. 6a, the endogenous protein levels of RIG-I and MDA5 but not MAVS were decreased gradually with the increased expression of RNF167. Consistently, the phosphorylation of TBK1 and IRF3, which modulates the activity of type I IFN signaling, was also reduced (Fig. 6a). We speculated that RNF167 might regulate the turnover of RIG-I/MDA5 protein but not MAVS. To prove this hypothesis, we found that the expression of MAVS was not affected by the increasing expression of RNF167 (Fig. 6b). We then constructed the HEK293 cell lines stably expressing RIG-I/MDA5 by using lentiviruses and observed a gradual decrease in the expression levels of RIG-I and MDA5 with increased expression of RNF167 (Supplementary Fig. 5a and Fig. 6c). Consistently, using siRNAs targeting RNF167, we found that the non-induced RIG-I and MDA5 levels were significant higher in groups with si-RNF167-1/2 compared to the control group (Fig. 6d). We then detected the SeV-induced expression of RIG-I and MDA5 in the presence or absence of RNF167, and it showed that the protein levels of RIG-I and MDA5 were increased in RNF167-KO cells compared to those in RNF167-WT cells, whereas the restoration of RNF167 inhibited RIG-I/MDA5 expression (Fig. 6e). Similar results were obtained by the exogenous expression of RIG-I/MDA5 in RNF167-WT, RNF167-KO and RNF167-reconstituted cells (Fig. 6f). Additionally, we performed cycloheximide (CHX) chase assay, and found that the ectopic expression of RNF167 shortened the half-life and significantly reduced the protein level of both non-induced and SeV-induced RIG-I/MDA5 (Fig. 6g, h). We also knocked down RNF167 in HEK293 cells and then infected cells with virus, and found that siRNA-mediated knockdown of RNF167 increased the protein level of RIG-I and MDA5, hereby decreasing the replication of virus (Supplementary Fig. 5b).
a Immunoblot analysis of indicated proteins in HEK293 cells transfected with empty vector or an increasing amount of RNF167 for 24 h and then infected with SeV for another 8 h. b Immunoblot analysis of Flag-MAVS in HEK293 cells transfected with an increasing amount of RNF167 for 24 h. c, d Immunoblot analysis of RIG-I-Flag/ MDA5-Flag in their stable expressing HEK293 cells transfected with an increasing amount of RNF167 for 24 h (c) or transfected with siRNAs targeting RNF167 for 48 h (d). e Immunoblot analysis of RIG-I/MDA5 in RNF167 WT and KO HEK293 cells transfected with RNF167-HA or its control for 24 h and then infected with SeV for 8 h. f Immunoblot analysis of Flag-RIG-I/Flag-MDA5 in RNF167 WT and KO HEK293 cells transfected with Flag-RIG-I/Flag-MDA5 and RNF167-HA or empty vector for 24 h. g Immunoblot analysis of RIG-I-Flag/MDA5-Flag in their stable expressing HEK293 cells transfected with RNF167-HA or empty vector for 24 h and then treated with CHX (50 μM) for the indicated time. (h) Immunoblot analysis of HEK293 cells transfected with RNF167-HA or empty vector for 16 h, infected with SeV for 8 h and then treated with CHX (50 μM) for the indicated time. i Dual-luciferase assays of IFN-β, PRDIII-I and NF-κB promotors in HEK293 cells transfected with RNF167 or its mutations for 24 h and then infected with SeV for 8 h (n = 3). The mutations were illustrated on the top panel. j Immunoblot analysis of HEK293 cells transfected with expression plasmids for RNF167-HA or its mutants V98G, C233S, ΔPA, and ΔRING-ΔC for 24 h, and then infected with SeV for 8 h. Data are representative of three independent experiments and are shown as mean with SD (g–i). The relative band density was measured by Quantity one and normalized to GAPDH/Actin (n = 3, g, h). The unpaired two-tailed t-test was applied (ns, not significant). GAPDH/Actin were used as loading controls. Source data are provided as a Source Data file.
As we illustrated above, RNF167 functions as an E3 ligase and contains two key domains, PA domain and RING domain. It was reported that V98 site was crucial for the function of PA domain and C233 and C268 sites were crucial for the catalyzing activity of RING domain20,23. We next went to clarify the functional domain of RNF167. To address this, we constructed V98G (valine to glycine), C233S (cysteine to serine), C268R (cysteine to arginine) together with the above ΔPA (PA domain deletion), ΔRING (RING domain deletion), ΔC (C terminal domain deletion) and ΔRING-ΔC (both RING and C domains deletion) mutants of RNF167. The IFN-β promoter-based dual-luciferase assays showed that the mutations of V98G and ΔPA showed negligible difference with wild-type RNF167 and the ΔC mutant partially lost its suppression ability, due to its primarily binding function as mentioned above, whereas C233S, C268R, ΔRING and ΔRING-ΔC mutants totally lost their suppression abilities (Fig. 6i and Supplementary Fig. 5c, d). Similar experiments showed that RNF167-C233S lost the ability to inhibit the activities of PRDIII-I and NF-κB reporters induced by SeV infection either (Fig. 6i). In consistence with above results, RNF167-C233S and RNF167-ΔRING-ΔC lost the suppression effect on the stability of RIG-I/MDA5, while RNF167-V98G and RNF167-ΔPA showed robust inhibition effect, akin to the full-length RNF167 protein (Fig. 6j). These data indicate that the E3 ligase activity and RING domain of RNF167 is indispensable for the regulation of the protein turnover of RIG-I/MDA5.
RNF167 catalyzes Lys6- and Lys11-linked polyubiquitination of RIG-I and MDA5
Our results showed that the RING domain of RNF167 mediated the interaction with RIG-I and MDA5 and was critical for the regulation of RIG-I/MDA5, indicating that the function of RNF167 in the regulation of RIG-I and MDA5 was dependent on its E3 ligase activity. To verify this, we co-transfected the plasmids expressing RIG-I/MDA5/MAVS, RNF167/RNF167-C233S, and ubiquitin to assess RNF167-mediated ubiquitination of RIG-I/MDA5. The results showed that RNF167 remarkably increased the polyubiquitination of RIG-I and MDA5, with no discernible effect on the ubiquitination of MAVS. However, the ligase-dead mutant of RNF167-C233S could not increase the polyubiquitination of RIG-I/MDA5 (Fig. 7a and Supplementary Fig. 6a, b). Therefore, the above results demonstrated that RNF167 has a significant effect on the polyubiquitination and stability of RIG-I/MDA5, but not on MAVS (Fig. 6a, b and Supplementary Fig. 6a). This indicates that RNF167 specially targets RIG-I/MDA5 with an indirect effect on MAVS-triggered activity to some extent.
a Coimmunoprecipitation analysis of RIG-I polyubiquitination in HEK293 cells transfected with Flag-RIG-I, His-ubiquitin along with empty vector, RNF167-HA, or its mutant RNF167-HA-C233S expression plasmids for 24 h. b Coimmunoprecipitation analysis of RIG-I polyubiquitination in HEK293 cells transfected with expression plasmids for Flag-RIG-I and His-ubiquitin-WT or its mutants (only a single lysine residue was retained) in presence of empty vector or RNF167-HA for 24 h. c Coimmunoprecipitation analysis of RIG-I polyubiquitination in HEK293 cells transfected with expression plasmids for Flag-RIG-I, His-ubiquitin-WT or its mutants (only the Lys residue 6/11 was retained/mutated to Arg or both Lys 6 and 11 were mutated to Arg), along with RNF167-HA for 24 h. d Coimmunoprecipitation analysis of RIG-I total Ub, K6- or K11-linked ubiquitination in WT or RNF167-deleted HEK293 cells transfected with Flag-RIG-I expression plasmids for 24 h, and then infected with SeV for 8 h or not. e Immunoblot analysis in HEK293 cells transfected with expression plasmids for Flag-RIG-I-CARDs, helicase, and CTD domains with RNF167-HA or the empty vector as control for 24 h. f Coimmunoprecipitation analysis of polyubiquitination of the RIG-I domains in HEK293 cells transfected with expression plasmids for Flag-RIG-I-CARDs or CTD domain with His tagged K6- or K11-ubiquitin in presence of RNF167-HA or its empty vector as control for 24 h. g Overview of identification of lysine sites in RIG-I ubiquitinated by RNF167 through mass spectrometry analysis; the diagram was created with Medpeer. h Immunoblot analysis in HEK293 cells transfected with expression plasmids of indicated Flag-RIG-I-CARDs/CTD mutants and RNF167-HA or its empty vector as control for 24 h. i Coimmunoprecipitation analysis of polyubiquitination in HEK293 cells transfected with His tagged K6- or K11-ubiquitin and the indicated Flag-RIG-I-CARDs/CTD mutants in presence of RNF167-HA or its empty vector as control for 24 h. j Immunoblot analysis of RIG-I-Flag or RIG-I-6KR-Flag (six lysine ubiquitination sites in RIG-I) in their stable expressing HEK293 cells transfected with a siRNAs targeting RNF167 for 48 h. The experiments were repeated at least once. GAPDH/Actin/Tubulin were used as loading controls. Source data are provided as a Source Data file.
We then explored which type of ubiquitin linkage that was promoted by RNF167 by employing vectors expressing His-tagged ubiquitin and the mutants of ubiquitin (K6, K11, K27, K29, K33, K48 and K63), which retain a single lysine residue at the specific position. As is shown in Fig. 7b and Supplementary Fig. 6c, the increased level of polyubiquitination of RIG-I and MDA5 by RNF167-HA was detected in the presence of WT ubiquitin, and K6- and K11- mutants. This result indicated that RNF167 catalyzed K6- and K11-linked polyubiquitination of RIG-I/MDA5. Therefore, we further performed ubiquitination assays using the ubiquitin mutants K6R, K11R and K6R/K11R, featuring lysine-to-arginine substitutions at position K6 or K11, or both K6 and K11 positions. The level of RIG-I/MDA5 polyubiquitination mediated by RNF167 was decreased in the presence K6R and K11R, but completely vanished in the presence of K6R/K11R (Fig. 7c and Supplementary Fig. 6d). Next, we examined the endogenous ubiquitination of RIG-I/MDA5 catalyzed by RNF167. As shown in Fig. 7d, the endogenous levels of total ubiquitination, as well as K6- and K11-linked ubiquitination of RIG-I, were significantly decreased in RNF167-deleted cells compared to that in WT cells. The result further confirmed that RNF167 catalyzes the K6- and K11-linked polyubiquitination of RIG-I/MDA5.
To identify which lysine residues of RIG-I/MDA5 were modified by RNF167-mediated ubiquitination, we used the functional domain truncations of RIG-I/MDA5. We found that RFN167 induced the degradation of RIG-I/MDA5-CARDs and RIG-I/MDA5-CTD but had no degradation effect on the RIG-I/MDA5-helicase domains, suggesting that RNF167 functions on CARDs and CTD domains of RIG-I/MDA5 (Fig. 7e, Supplementary Fig. 6e). As expected, polyubiquitination analyses showed that the RIG-I/MDA5-CARDs contained the primary sites of K6-linked ubiquitination, whereas the RIG-I/MDA5-CTD included the main sites of K11-linked ubiquitination (Fig. 7f, Supplementary Fig. 6f).
Next, we sought to identify RNF167-catalyzed ubiquitinated peptides of RIG-I using WT or RNF167-KO HEK293 cells transfected with Flag-RIG-I and RNF167-HA expression plasmid or its empty vector, and subsequentially, the immunoprecipitated RIG-I were analyzed by mass spectrometry (MS) with a final RIG-I peptide coverage up to 87% (Fig. 7g). We excluded the RIG-I peptides with the same ubiquitinated lysine sites in CARDs and CTD domains within both WT cells and the RNF167-KO cells. After analysis, the results showed that peptides with K45 and K203 in CARDs domain and K888 in CTD domain were distinguished in WT group from the RNF167-KO control, and peptides with K95, K96, K172 and K812 were not covered by every MS analysis (Supplementary Fig. 6g). Thus, we generated a series of RIG-I-CARDs and CTD mutants with the potential unique lysine sites identified in WT group replaced by arginine residues (including the lysine sites in uncovered peptides). As shown in Fig. 7h, i, these results demonstrated that only K95R, K96R, K172R, and K203R mutants of RIG-I-CARDs, as well as K812R and K888R mutants of RIG-I-CTD were partially promoted to degradation, while K4R mutants of RIG-I-CARDs and K2R mutants of RIG-I-CTD were barely ubiquitinated by RNF167. Collectively, these results indicate that RNF167 promotes K6-linked ubiquitination of RIG-I/MDA5 at CARDs domains with the K95-96, K172 and K203 sites ubiquitinated in RIG-I-CARDs, and K11-linked ubiquitination at CTD domains with the K812 and K888 sites ubiquitinated in RIG-I-CTD.
Furthermore, we constructed HEK293 cells stably expressing RIG-I-WT or RIG-I-6KR, in which six lysine sites of K95, K96, K172, K203, K812 and K888 were mutated to arginine (Supplementary Fig. 6h). As demonstrated in Supplementary Fig. 6i and Fig. 7j, RNF167 expression promoted the degradation of RIG-I, while knockdown of RNF167 increased RIG-I expression. However, RNF167 had no effect on the protein level of RIG-I-6KR. Next, we determined the activities of IFNβ promoters in cells stably expressing RIG-I-WT and RIG-I-6KR following ectopic expression of RNF167-WT or the RNF167-C233S mutant. The results indicated that RNF167-WT suppresses IFNβ promoter driven-luciferase activity in RIG-I-WT cells but not in RIG-I-6KR stably expressed cells. The RNF167-C233S mutant failed to suppress the activity in HEK293-RIG-I-WT cells and in RIG-I-6KR cells (Supplementary Fig. 6j). We obtained consistent results by using ectopic expression of Flag-RIG-I or Flag-RIG-I-6KR with RNF167-WT and RNF167-WT-C233S (Supplementary Fig. 6k). Consistently, we further detected that the endogenous ubiquitination level of RIG-I, and observed the higher endogenous levels of total Ub, K6- and K11-linked ubiquitination in the presence of RNF167, whereas RNF167 showed no effect on the ubiquitination of RIG-I-6KR (Supplementary Fig. 6l). Taken together, these data demonstrate that RNF167 promotes K6- and K11-linked ubiquitination of RIG-I at K95, K96, K172, K203 and K812 and K888, respectively.
RNF167 promotes the degradation of RIG-I and MDA5 in dual proteolytic pathways
The ubiquitin-proteasome system (UPS) and autophagy-lysosome pathway (ALP) are two major mechanisms governing proteolysis in eukaryotic cells31. Then, we determined the fate of RIG-I/MDA5 after polyubiquitin conjugation by RNF167. We used different inhibitors, including the proteasome inhibitor MG132 and lysosome inhibitor chloroquine (CQ), to block the degradation pathways. The result showed that RNF167 promoted the degradation of RIG-I/MDA5, and single treatment with MG132 or CQ failed to block the degradation (Supplementary Fig. 7a). Intriguingly, the combination of MG132 and CQ effectively blocked the degradation and rescued the expression of RIG-I and MDA5 (Fig. 8a and Supplementary Fig. 7b). 26S proteasome non-ATPase regulatory subunit 10 (PSMD10) is one crucial component of the 26S proteasome32. The expression level of RIG-I/MDA5 was completely restored with RNF167 expression in PSMD10-KO HCT116 cells treated with CQ, although either the UPS deficiency or autophagy blockage alone could not rescue RIG-I/MDA5 to the level of control cells (Fig. 8b). These data indicated that RNF167-mediated RIG-I/MDA5 degradation may involve both the UPS and ALP.
a Immunoblot analysis of HEK293 cells transfected with expression plasmids of Flag-RIG-I and RNF167-HA or its empty vector for 18 h, and then treated with DMSO, MG132 (10 μM), CQ (25 μM), or MG132 plus CQ for another 6 h; the relative band density of Flag-RIG-I was measured by Quantity one and normalized to GAPDH (n = 3). b Immunoblot analysis of RIG-I/MDA5 in PSMD10 WT and KO HCT116 cells transfected with RNF167-HA or empty vector for 24 h, infected with SeV for 8 h and then treated with DMSO or CQ for another 6 h. c, d Coimmunoprecipitation and immunoblot analysis of HEK293 cells transfected with the indicated plasmid combinations for 24 h. e Immunofluorescence analysis of HeLa cells transfected with indicated plasmids for 24 h (n = 5). Scale bars, 10 μm. f Immunoblot analysis of HEK293 cells transfected with plasmids for Flag-RIG-I-CARDs and CTD with RNF167-HA or empty vector for 24 h and then treated with DMSO, MG132 or CQ for another 6 h. g Immunoblot analysis of HeLa cells transfected with expression plasmids for Flag-RIG-I-4KR (K95R-K96R-K172R-K203R) or 2KR (K812R-K888R) together with RNF167-HA or empty vector as control for 24 h and then treated with MG132 or CQ for another 6 h. h Immunofluorescence analysis of HeLa cells transfected with indicated plasmids for 24 h (n = 5). Scale bars, 10 μm. i Schematic model of RNF167-mediated ubiquitination of RLRs in regulation of IFN-I signaling pathway; the diagram was generated with BioRender. Data are representative of three independent experiments and are shown as mean with SD (a, e, h). The fluorescence intensity of the indicated area (Red line) was shown and the Mander’s overlap coefficient (MOC) of the indicated proteins in cells were measured for overlapping with Flag/HA-RIG-I; cells with weak fluorescence intensity of indicated proteins were used as the background (e, h). The unpaired two-tailed t-test was applied (ns, not significant). GAPDH/Actin/Tubulin were used as loading controls (a, b, c, d, f, g). Source data are provided as a Source Data file.
To further verify our observations, we went to figure out which autophagic cargo receptor was involved in the ALP process. Co-immunoprecipitation assays displayed that RIG-I/MDA5 had interaction with OPTN, p62 and CCDC50 autophagy receptors, while RNF167 specifically interacted with p62 (Fig. 8c, d and Supplementary Fig. 7c). Our previous study revealed that CCDC50 showed a strong preference for K63-linkages and recognized K63-linked polyubiquitinated substrates, but CCDC50 aided the function of p62. Therefore, it could be envisaged that p62 was directly involved in the RNF167-mediated autophagy cargo degradation.
In further support of our findings, we performed confocal microscopy analyses to examine the association of RIG-I/MDA5, RNF167 and canonical autophagy machinery proteins. The images of confocal microscopy revealed clear colocalization between RIG-I/MDA5, RNF167 and p62 /LC3/LAMP1, indicating that RNF167 facilitated the interaction between RIG-I/MDA5 and the cargo receptor p62, or other autophagy-associated compartments (Fig. 8e and Supplementary Fig. 7d).
The data showed that RNF167 could colocalize with RIG-I/MDA5 and p62 and LC3B (Fig. 8e and Supplementary Fig. 7d), suggesting that the polyubiquitination of RIG-I/MDA5 recruited p62 and then linked them to LC3-anchored autophagosomes. We found that the treatment with CQ blocked the degradation of CARDs domain of RIG-I/MDA5, whereas MG132 treatment rescued the expression of RIG-I/MDA5-CTD (Fig. 8f and Supplementary Fig. 7e). The data suggested that RNF167-mediated K6-linked polyubiquitination of RIG-I/MDA5 in CARDs domain recruited p62 and served as the cargo of p62. Next, we generated the RIG-I mutants with the lysine residues (identified ubiquitination sites in Fig. 7f-j) replaced with arginine residues in CARDs (RIG-I-4KR) or CTD (RIG-I-2KR) domains. As shown in Fig. 8g, RIG-I-4KR and 2KR mutants were promoted to degradation by RNF167, while it was blocked by treatment with MG132 or CQ, respectively. Indeed, immunofluorescence assays showed the colocalization of RIG-I/MDA5, K6-Ub, and p62/LC3B, demonstrating that RNF167-mediated K6-ubiquitin-conjugation of RIG-I/MDA5 can be targeted for autophagosomes (Fig. 8h and Supplementary Fig. 7f). Collectively, these results indicate that RNF167 catalyzes the K6- and K11-linked polyubiquitination of RIG-I/MDA5, subsequently assigning these substrates to UPS and ALP for sequential degradation.
In summary, we propose a unique regulatory model: As is shown in Fig. 8i, after activation of type I IFN in response to virus infection or other stimuli, RNF167 is induced and bind to RIG-I/MDA5. Following K6- and K11-linked polyubiquitination catalyzed by RNF167, the soluble K11-polyubiquitinated RLRs are delivered for degradation through UPS; simultaneously, the excessively activated RIG-I/MDA5 coupled with K6-linked ubiquitination could oligomerize to form aggregates, are recognized by p62 and then shuttled to ALP for degradation. Our results provide insights into the cross-talk and synergistic effect between proteasomal and lysosomal degradation pathways.
Discussion
Although various E3 ligases have been identified for regulating viral infection-induced IFN-I responses, the functions of RNF167 and the mechanisms of dual-ubiquitination modification remain not yet reported. In this work, we present a previously unknown mechanism by which RNF167 can modify its substrate by two types of atypical polyubiquitination and direct the substrates for degradation through two distinct proteolytical pathways.
It has been reported that RNF167 interacts with multiple E2 enzymes and promotes its own self-ubiquitination, a process where the RING domain plays a crucial role22. This study suggests that self-ubiquitination may be important for the stability and catalytic activity of RNF167. Therefore, we speculate that self-ubiquitination may enhance the interaction between RNF167 and E2 enzymes and substrates, or improve their ubiquitin transfer efficiency. On the other hand, RNF167 is inducible and gradually degraded like an ISG as the pathway diminishes (Fig. 1 and Supplementary Fig. 1). Thus, we could infer that IFN-β may mediate RNF167 self-ubiquitination to maintain its appropriate protein levels. In summary, while type I IFN might be involved in RNF167 expression, further studies are needed to determine if IFN regulates RNF167 self-ubiquitination.
Although there are no known negative regulators for RNF167-mediated ubiquitination of RIG-I and MDA5, it has been reported that RNF167 and a deubiquitinase STAMBPL1 that function in concert to control the polyubiquitination level of Sestrin2 to modulate mTOR and cancer progression27. Therefore, it could be promising to identify a deubiquitylation enzyme that can inhibit RNF167-mediated degradation of RLRs.
Different types of ubiquitination play distinct roles in determining the fate of protein substrates. K48-linked ubiquitination often leads to proteasome-mediated degradation of the substrates, whereas K63-linked polyubiquitination is commonly associated with signaling activation33. These two types of polyubiquitination were thoroughly studied in regulation of RIG-I/MDA5 stability and activity34. However, the roles of other types of polyubiquitination are less reported17,35. Especially, the physiological functions of atypical K6-linked polyubiquitination are largely unexplored36,37,38. RNF144A was reported to promote K6-linked ubiquitination of STING to enhance DNA viral infection-induced IFN-I responses39. Several adaptor proteins, but not the RLRs, have been reported to undergo K11-linked ubiquitination by different E3 ligases in the regulation of IFN-I pathway40,41,42,43,44. Here, our study reveals that the E3 ligase RNF167 mediates the atypical K6- and K11-linked polyubiquitination of RIG-I/MDA5, leading to both proteasomal and lysosomal degradation.
The UPS primarily serves as the degradation pathway for misfolded, soluble and short-lived proteins, whereas larger protein aggregates are targeted to autophagosomes, with p62 at the crossroad of these two pathways45,46. As the early IFN response genes47, RIG-I and MDA5 are short-lived and primarily targeted for degradation through UPS by E3 ligases like RNF12548, RNF57 (c-Cbl)49, RNF12250, and RNF15351. However, when oligomerized RIG-I/MDA5 are recruit to MAVS through CARDs domains, they form into larger protein aggregates that are shuttled for degradation via ALP by autophagy receptors such as CCDC5029. As a unique endosome-lysosome located E3 ligase25,52, this feature may contribute to the dual-proteolysis mechanisms of RNF167.
The mechanisms of selectivity and cross-talk of UPS and ALP remain elusive. Although both degradation pathways are initiated by the ubiquitylation of the substrates, the branch of the adaptive protein quality control pathways is determined by the oligomerization potentials of the substrates but not their ubiquitin-binding properties53. Our results demonstrate that K6-linked polyubiquitination of RIG-I/MDA5 in their CARD domains that recruits p62 for selective autophagic degradation, while K11-linked polyubiquitination of RIG-I/MDA5 in CTD domains is targeted for UPS-dependent degradation (Fig. 8e-i and Supplementary Fig. 7c-f). Thus, we hypothesize that the oligomerization potential of RIG-I/MDA5 through CARDs domains is enhanced by K6-linked polyubiquitination catalyzed by RNF167, and the RLRs are sorted for ALP for degradation. In contrast, RNF167 mediated-K11-linked polyubiquitylation in the CTD domains of RLRs represents soluble proteins and are directed to the proteasome. The ubiquitylation modification of substrates are shared by proteasomal and autophagic protein quality pathways. Which degradation pathway to assign is not decided by ubiquitylation but by the oligomerization potential of substrates. Therefore, our results contribute to a better understanding of the substrate distribution mechanisms of these two degradation pathways for protein quality control.
Recent studies have reported that several neurodegenerative diseases-associated proteins can be degraded through both of the two proteolytic pathways, such as α-Synuclein, Rab21, and CNTNAP254,55,56,57. However, the specific mechanism and the involved E3 ligase also remain unknown. Our findings prove that RNF167 catalyzes the two non-canonical ubiquitination types and promotes subsequent degradation of RIG-I/MDA5 through both proteolytic pathways, underscoring the critical role of RNF167 in maintaining protein homeostasis, especially for the immune regulation. Additionally, our results underline the significance of cooperation of UPS and ALP in protein quality control.
Methods
All research complies with the Declaration of Sun-Yat Sen University, and the study protocols were approved by the Guangzhou Medical University Review Board.
Antibodies and reagents
RIG-I (3743S), MDA5 (5321S), MAVS (24930S), TBK1 (3013S), Phospho-IRF-3 (Ser396) (4947S), IRF-3 (4302S), Phospho-TBK1 (Ser172) (5483S) and GFP (2956 T) were purchased from Cell Signaling Technology. Ubiquitin (10201-2-AP), His (10001-0-AP), LAMP1 (55273-1-AP), Myc (60003-2-Ig), p62 (18420-1-AP), PSMD10 (12342-2-AP), β-actin (66009‐1‐Ig) and GAPDH (60004-1-Ig) were purchased from Proteintech. IFNAR1 (sc-53591, Santa Cruz), RNF167 (Santa Cruz, sc-515405 and Proteintech, 24618-1-AP), β-tubulin (Arigo, ARG62347), Ub-K11 Polyclonal Antibody (Invitrogen, PA5-120621), Flag (Sigma, clone M2, F1804), HA (Sigma, clone HA-7, H9658) and more detailed information about these primary antibodies were shown in supplementary table 1. Poly(I:C) (LMW and HMW) were purchased from InvivoGen and used at a final concentration of 1 μg /mL for transfection and 10 µg/mL for adding to supernatant of the cell culture for endogenous and exogenous stimulation, respectively. Cyclohexamide (CHX, 239764), (R)-MG132 (M8699), NH4Cl (09718), chloroquine (CQ, C6628), and Anti-Ubiquitin Lys6 specific Affimer reagent GFP/His tag (MABS1918) was purchased from Sigma, while 3-Methyladenine (3-MA, S2767) was purchased from Selleck. Human IFN-β (AF-300-02B) was purchased from PeproTech and mouse IFN-β (HY-P73130) was purchased from MCE (used at final concentration of 10 ng/mL).
Cells and virus
RNF167-deletion HEK293 cells were generated by using the CRISPR/Cas9 system with single short guide RNAs (sgRNAs) targeting specific sequences in the RNF167 genome (TGTGGAGGCTCACCCAGACA), and the primer sequences was on supplementary table 2. The sgRNAs were synthesized and constructed into lentiCRISPRV2 plasmids, and then were transfected into HEK293T cells with the packaging plasmids pMD2.G and psPAX2 by using the Polyethyleneimine (PEI) transfection reagent for collecting the 48 h and 72 h cell culture medium. The medium containing the lentivirus was filtered with the 0.45 μm filter, and used for infection of the HEK293 cells for 6 h, then cultured with fresh medium for 48 h, treated with puromycin for 4 days, and selected for the RNF167-/- single clone cell lines. We used PCR followed by sequencing, qPCR and immunoblotting to determine the knockout efficiency. The RIG-I-WT/6KR and MDA5-Flag stably expressing HEK293 cell lines were generated by constructing the RIG-I-WT/6KR or MDA5-Flag into the pHAGE-lentivirus plasmids. The plasmids were transfected into HEK293T cells together with the packaging plasmids pMD2.G and psPAX2 using the Polyethyleneimine (PEI) transfection reagent. The supernatants were collected 48 h and 72 h post transfection. Then the following processes were similar as above for constructing RNF167-deletion cell lines.
PSMD10-deletion HCT116 cells were gifted from Dr. Hong Peng (Sun Yat-Sen University). Bone marrow-derived macrophages (BMDMs) and bone marrow-derived dendritic cells (BMDCs) were isolated from the tibia and femur of the WT and KO mice. For the preparation of bone marrow–derived macrophages, bone marrow cells were cultured for 3–5 d in medium containing 30% supernatants of L929 mouse fibroblasts containing macrophage-stimulating factor. For the preparation of BMDCs with the cytokine GM-CSF, bone marrow cells were cultured for 7–9 d in medium containing mouse GM-CSF (50 ng/ml; Peprotech). THP-1 cells were maintained in 1640 medium (Gibco) supplemented with 10% heat-inactivated fetal bovine serum (FBS, Gibco) and cultured at 37 °C in a 5% CO2 incubator, and the other cells like HEK293, RAW264.7 and L929 cells, etc. were maintained in DMEM medium (Gibco) supplemented with 10% heat-inactivated fetal bovine serum (FBS, Gibco) and cultured at 37 °C in a 5% CO2 incubator. All the cell lines used have been tested the mycoplasma-free. HSV-1, SeV, EMCV, VSV-GFP, and VSV wildtype were used as described58 and also indicated in the figure legends.
Constructs
NF-κB-, interferon-stimulated response element, and IFN-β promoter luciferase reporter plasmids, and the mammalian expression plasmids about type I interferon like pEF-Flag-RIG-I, pEF-Flag-MDA5, pEF-Flag-MAVS, pEF-Flag-TBK1, pEF-Flag-IRF3, pEF-Flag-IKK-β and pEF-Flag-IRF3, etc. were described previously58. The RNF167-HA plasmid was amplified from cDNA from HEK293 genome and constructed into vector pCAGGS. All the truncations and site-directed mutants were constructed according to the manufacturer’s standard procedures. Flag-p62, Flag-OPTN, GFP-LC3 were from Dr. Hong Peng (Sun Yat-Sen University), and Flag-CCDC50 was described in the previous study of our groups29.
Mice
Rnf167-deficient mice were generated on a C57BL/6 background by the Laboratory of Genetic Regulators in the Immune System in Xinxiang Medical University through CRISPR/Cas9-mediated gene editing. The age (6-8 weeks old) and sex-matched wild type and Rnf167-/- mice were used in our experiments. All mice were maintained in a specific pathogen-free animal facility in Sun Yat-sen University. And all mice experiments were conducted in the supervisor of the Institutional Animal Care and Use Committee of Sun Yat-sen University.
Transfection and dual-luciferase reporter assay
HEK293 cells were seeded on 24-well plates (2 × 105 cells per well) and then cotransfected with 50 ng of luciferase reporter plasmid (IFN-β-luc, PRDI-III-luc or NF-κB-luc), 0.5 μg of RNF167 expression plasmid, its mutant expression plasmids, or the empty vector, together with the 20 ng of pRL-TK Renilla luciferase reporter plasmid via Polyethyleneimine (PEI) for 24 h, then infected with SeV for 8 h. Luciferase activity in total cell lysates was measured with a dual-specific luciferase reporter assay system (Promega), and the cell lysates left were collected for western blot analysis.
RNAi
The small interfering RNAs were designed and purchased from Ribobio, and the sequences were listed in Supplementary Table 2. The siRNAs were transfected to the different cell lines by using the Lipofectamine™ RNAiMAX Transfection Reagent (Invitrogen) for 48 h.
RNA quantification
Total RNA was extracted with TRIzol reagent according to the manufacturer’s instructions (Invitrogen), and then was reversed-transcribed with a Reverse Transcription System (Promega) before quantitative RT-PCR analysis. ABI Q5 Detection System (Applied Biosystems) and the SYBR Green PCR Master Mix (Roche) were used to detect the cDNA products for analysis with the primers listed in Supplementary Table 3.
Co-immunoprecipitation, ubiquitination and immunoblot analysis
For immunoblot analysis, after transfection, the cells were collected and lysed in RIPA lysis buffer containing 50 mM Tris-HCl, pH 8.0, 150 mM NaCl, 1.0% (v/v) Triton X-100, 1.0% sodium deoxycholate and 0.1% SDS with protease inhibitors. The samples were boiled and loaded on SDS-PAGE, transferred onto PVDF membranes and then blotted with indicated antibodies. The uncropped and unprocessed blots are provided in the Source Data file. For co-immunoprecipitation, HEK293 cells were seeded and transfected with the appropriate expression plasmids or empty vectors for the indicated time, cells were lysed in lysis buffer (50 mM Tris-HCl, pH 7.6, 150 mM NaCl, 0.5% Triton-X-100, 2 mM EGTA, 10 mM NaF, 1 mM Na3VO4 and 2 mM DTT) containing protease inhibitors, then the cell lysates were centrifuged and the supernatants were incubated with anti-Flag M2 affinity gel or anti-HA affinity gel (beads with rabbit or mouse IgG as control) for at least 4 h at 4 °C. For the endogenous protein, pre-clear whole cell extracts by adding protein A/G Sepharose beads and 2 µg of rabbit or mouse IgG control antibody for 2 h. After centrifuging the samples, the supernatants were incubated with rabbit or mouse IgG control antibody or the relevant antibody together with protein A/G Sepharose beads overnight at 4 °C. The beads were washed with the same lysis buffer four times by rotating and eluted in 2 × SDS sample loading buffer by boiling for 10 min, then loaded to SDS-PAGE for immunoblot analysis. For ubiquitination assays, whole-cell extracts were boiled for 10 min at 95 °C with 1% SDS, then diluted with 9 volumes of lysis buffer to the final concentration of 0.1% SDS, and were immunoprecipitated with anti-Flag M2 affinity gel and analyzed by immunoblot.
Immunofluorescence labeling and confocal microscopy
HeLa cells were fixed for 10 min with 4% paraformaldehyde in PBS and permeabilized for 15 min at room temperature with 0.2% Triton X-100 in PBS. After incubation for blockade of nonspecific binding with 1% bovine serum albumin (BSA) in PBS for 30 min, indicated primary antibodies were added and incubated for 2 h at room temperature (or 4 °C overnight). Then samples were stained with fluorescent-dye-conjugated secondary antibodies (Thermo Fisher Scientific). Nuclei were stained with DAPI (Invitrogen). Fluorescence in cells was visualized and images were acquired with a Carl Zeiss laser-scanning confocal microscope.
ELISA assay
The production and secretion of mouse IFN-β in cell supernatants and mouse serum were measured with the mouse IFN-β ELISA kit (Lianke Bio).
Viral infection and plaque assay
After infection, supernatants or sera-containing virus were collected and diluted to infect Vero cells plated on 24-well plates at 90% confluence. At 2 h after infection, supernatants were removed and cells were washed with PBS, then the medium containing methylcellulose was overlaid onto the cells. 2 days post infection, the cells were stained with crystal violet (0.2%). Plaques were counted, and results were averaged and multiplied by the dilution factor for calculation (PFU/mL).
Virus infection in vivo
For in vivo virus infection studies, age and sex-matched WT and Rnf167-/- mice were in intravenous injection of VSV 6 × 107 PFU per mouse. IFN-β production in the sera was measured by ELISA; Mice were monitored for survival after VSV infection for the survival experiments (1.2 × 108 PFU per mouse); Lungs from control or the VSV infected mice were dissected, fixed in 10% phosphate-buffered formalin, embedded into paraffin, sectioned, stained with hematoxylin–eosin solution and examined by light microscopy for histological changes; After 16 h of VSV infection, Lungs, spleens, and livers from WT and Rnf167-/- mice were also acquired to detect replication of VSV and the mRNA transcription of Ifnb1, Isgs and inflammatory factors.
HPLC-MS analysis
To identify RNF167-catalyzed ubiquitinated peptides of RIG-I, the wild-type HEK293 cells were transfected with Flag-RIG-I and RNF167-HA expression plasmids, while RNF167-KO cells transfected with an empty vector served as controls. 24 h post transfection, cell samples were collected and lysed in lysis buffer (50 mM Tris-HCl, pH 7.6, 150 mM NaCl, 0.5% Triton-X-100, 2 mM EGTA, 10 mM NaF, 1 mM Na3VO4 and 2 mM DTT) containing protease inhibitors. The cell lysates were centrifuged and the supernatants were boiled for 10 min at 95 °C with 1% SDS, then diluted with 9 volumes of lysis buffer to the final concentration of 0.1% SDS. These were immunoprecipitated with anti-Flag M2 affinity gel overnight (4 °C). The beads were washed with the same lysis buffer by rotating for four times (10 min for each time) and eluted in 2 × SDS sample loading buffer by boiling for 10 min. All samples were then loaded onto SDS-PAGE. Following electrophoresis, the gel sections containing the RIG-I bands and regions above them were collected for trypsin digestion. Due to the limited peptide matching rate with one enzyme, a separate set of samples was similarly prepared for chymotrypsin digestion.
Peptides were then dissolved in 0.1% FA and 2% ACN, directly loaded onto a reversed-phase analytical column (75 μm i.d. × 150 mm, packed with Acclaim PepMap RSLC C18, 2 μm, 100 Å, nanoViper). The gradient was comprised of an increase from 5% to 50% solvent B (0.1% FA in 80% ACN) over 40 min, and climbing to 90% in 5 min, then holding at 90% for the 5 min. All at a constant flow rate of 300 nl/min. The MS analysis was performed on Q Exactive hybrid quadrupole-Orbitrap mass spectrometer (ThermoFisher Scientific).
The peptides were subjected to NSI source followed by tandem mass spectrometry (MS/MS) in Q ExactiveTM (Thermo) coupled online to the UPLC. Intact peptides were detected in the Orbitrap at a resolution of 70,000. Peptides were selected for MS/MS using NCE setting as 27; ion fragments were detected in the Orbitrap at a resolution of 17,500. A data-dependent procedure that alternated between one MS scan followed by 20 MS/MS scans was applied for the top 20 precursor ions above a threshold ion count of 1E4 in the MS survey scan with 30.0 s dynamic exclusion. The electrospray voltage applied was 2.0 kV. Automatic gain control (AGC) was used to prevent overfilling of the ion trap; 1E5 ions were accumulated for generation of MS/MS spectra. For MS scans, the m/z scan range was 350 to 1800 m/z. Fixed first mass was set as 100 m/z.
Protein identifications were performed with MASCOT software by searching Uniprot_Homo sapiens. We excluded the RIG-I peptides with the same ubiquitinated lysine sites in CARDs and CTD domains within both WT cells and the RNF167-KO cells for each enzyme digestion group. The potential ubiquitinated lysine sites on unique peptides from wild-type cells, along with those uncovered in each MS analysis, were selected for further identification.
Image quantification
For western blot analysis, the relative band density of the proteins in the same blot was measured by Quantity one and normalized to their corresponding loading controls GAPDH/Actin/Tubulin, representing the protein expression levels. For immunofluorescence image analysis, the fluorescence intensity of the indicated area was measured by the ZEN 2 software of Carl Zeiss, and the data was collected and made into graphs by using GraphPad Prism 7.0; the images were imported into ImageJ and the Mander’s overlap coefficient (MOC) of the indicated proteins in the same cells were measured by using the plugin-JACoP.
Statistical analysis
All data are presented as mean SD (SEM) of the representative of 3 or more experiments. Statistical significance between different groups was calculated by two-tailed unpaired Student’s t-test. Mouse survival data were plotted as Kaplan–Meier curves and compared by Gehan-Breslow-Wilcoxon test. Data were analyzed with GraphPad Prism 7.0 Software. *P < 0.05 was considered statistically significant.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The raw and processed deep sequencing data are deposited into the NCBI Gene Expression Omnibus (GEO) with accession number GSE143467. The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE59 partner repository with the dataset identifier PXD060570. All of the other data supporting this research are included in the manuscript and supplementary information. Source data are provided with this paper.
References
Brubaker, S. W., Bonham, K. S., Zanoni, I. & Kagan, J. C. Innate immune pattern recognition: a cell biological perspective. Annu Rev. Immunol. 33, 257–290 (2015).
Alexopoulou, L., Holt, A. C., Medzhitov, R. & Flavell, R. A. Recognition of double-stranded RNA and activation of NF-kappaB by Toll-like receptor 3. Nature 413, 732–738 (2001).
Yoneyama, M. et al. The RNA helicase RIG-I has an essential function in double-stranded RNA-induced innate antiviral responses. Nat. Immunol. 5, 730–737 (2004).
Hornung, V. et al. 5’-Triphosphate RNA is the ligand for RIG-I. Science 314, 994–997 (2006).
Kato, H. et al. Differential roles of MDA5 and RIG-I helicases in the recognition of RNA viruses. Nature 441, 101–105 (2006).
Pichlmair, A. et al. RIG-I-mediated antiviral responses to single-stranded RNA bearing 5’-phosphates. Science 314, 997–1001 (2006).
Xu, L. G. et al. VISA is an adapter protein required for virus-triggered IFN-beta signaling. Mol. Cell 19, 727–740 (2005).
Kawai, T. et al. IPS-1, an adaptor triggering RIG-I- and Mda5-mediated type I interferon induction. Nat. Immunol. 6, 981–988 (2005).
Meylan, E. et al. Cardif is an adaptor protein in the RIG-I antiviral pathway and is targeted by hepatitis C virus. Nature 437, 1167–1172 (2005).
Tan, X., Sun, L., Chen, J. & Chen, Z. J. Detection of Microbial Infections Through Innate Immune Sensing of Nucleic Acids. Annu Rev. Microbiol 72, 447–478 (2018).
Shrivastav, M. & Niewold, T. B. Nucleic Acid sensors and type I interferon production in systemic lupus erythematosus. Front Immunol. 4, 319 (2013).
Zhou, Y., He, C., Wang, L. & Ge, B. Post-translational regulation of antiviral innate signaling. Eur. J. Immunol. 47, 1414–1426 (2017).
Liu, J., Qian, C. & Cao, X. Post-Translational Modification Control of Innate Immunity. Immunity 45, 15–30 (2016).
Chiang, C. & Gack, M. U. Post-translational Control of Intracellular Pathogen Sensing Pathways. Trends Immunol. 38, 39–52 (2017).
Park, Y., Jin, H. S., Aki, D., Lee, J. & Liu, Y. C. The ubiquitin system in immune regulation. Adv. Immunol. 124, 17–66 (2014).
van Gent, M., Sparrer, K. M. J. & Gack, M. U. TRIM Proteins and Their Roles in Antiviral Host Defenses. Annu Rev. Virol. 5, 385–405 (2018).
Cai, C., Tang, Y.-D., Zhai, J. & Zheng, C. The RING finger protein family in health and disease. Signal Transduct. Target. Ther. 7, 300 (2022).
Popovic, D., Vucic, D. & Dikic, I. Ubiquitination in disease pathogenesis and treatment. Nat. Med. 20, 1242–1253 (2014).
Nakamura, N. The Role of the Transmembrane RING Finger Proteins in Cellular and Organelle Function. Membr. (Basel) 1, 354–393 (2011).
van Dijk, J. R., Yamazaki, Y. & Palmer, R. H. Tumour-associated mutations of PA-TM-RING ubiquitin ligases RNF167/RNF13 identify the PA domain as a determinant for endosomal localization. Biochem J. 459, 27–36 (2014).
Lussier, M. P. et al. Ubiquitin ligase RNF167 regulates AMPA receptor-mediated synaptic transmission. Proc. Natl Acad. Sci. USA 109, 19426–19431 (2012).
Ghilarducci, K. et al. Functional interaction of ubiquitin ligase RNF167 with UBE2D1 and UBE2N promotes ubiquitination of AMPA receptor. FEBS J. 288, 4849–4868 (2021).
Deshar, R. et al. RNF167 targets Arl8B for degradation to regulate lysosome positioning and endocytic trafficking. FEBS J. 283, 4583–4599 (2016).
Yamazaki, Y. et al. Goliath family E3 ligases regulate the recycling endosome pathway via VAMP3 ubiquitylation. EMBO J. 32, 524–537 (2013).
Nair, S. V., Narendradev, N. D., Nambiar, R. P., Kumar, R. & Srinivasula, S. M. Naturally occurring and tumor-associated variants of RNF167 promote lysosomal exocytosis and plasma membrane resealing. J. Cell Sci. 133, jcs239335 (2020).
Li, T. et al. RNF167 activates mTORC1 and promotes tumorigenesis by targeting CASTOR1 for ubiquitination and degradation. Nat. Commun. 12, 1055 (2021).
Wang, D. et al. E3 ligase RNF167 and deubiquitinase STAMBPL1 modulate mTOR and cancer progression. Mol. Cell 82, 770–784.e779 (2022).
Yan, Z. et al. RNF167-mediated ubiquitination of Tollip inhibits TNF-α-triggered NF-κB and MAPK activation. Faseb j. 37, e23089 (2023).
Hou, P. et al. A novel selective autophagy receptor, CCDC50, delivers K63 polyubiquitination-activated RIG-I/MDA5 for degradation during viral infection. Cell Res. 31, 62–79 (2021).
Hou, P. et al. An unconventional role of an ASB family protein in NF-kappaB activation and inflammatory response during microbial infection and colitis. Proc. Natl Acad. Sci. USA 118, e2015416118 (2021).
Kwon, Y. T. & Ciechanover, A. The Ubiquitin Code in the Ubiquitin-Proteasome System and Autophagy. Trends Biochem Sci. 42, 873–886 (2017).
Hori, T. et al. cDNA cloning and functional analysis of p28 (Nas6p) and p40.5 (Nas7p), two novel regulatory subunits of the 26S proteasome. Gene 216, 113–122 (1998).
Cabana, V. C. & Lussier, M. P. From Drosophila to Human: Biological Function of E3 Ligase Godzilla and Its Role in Disease. Cells 11, 380 (2022).
Rehwinkel, J. & Gack, M. U. RIG-I-like receptors: their regulation and roles in RNA sensing. Nat. Rev. Immunol. 20, 537–551 (2020).
van Huizen, M. & Kikkert, M. The Role of Atypical Ubiquitin Chains in the Regulation of the Antiviral Innate Immune Response. Front. Cell Developmental Biol. 7, 392 (2020).
Durcan, T. M. & Fon, E. A. USP8 and PARK2/parkin-mediated mitophagy. Autophagy 11, 428–429 (2015).
Suryo Rahmanto, A. et al. K6-linked ubiquitylation marks formaldehyde-induced RNA-protein crosslinks for resolution. Mol. Cell 83, 4272–4289.e10 (2023).
Zhao, S. et al. RNF14-dependent atypical ubiquitylation promotes translation-coupled resolution of RNA-protein crosslinks. Mol. Cell 83, 4290–4303.e9 (2023).
Yang, B. et al. RNF144A promotes antiviral responses by modulating STING ubiquitination. EMBO Rep. 24, e57528 (2023).
Zhao, J. et al. TRIM26 positively regulates the inflammatory immune response through K11-linked ubiquitination of TAB1. Cell Death Differ. 28, 3077–3091 (2021).
Dynek, J. N. et al. c-IAP1 and UbcH5 promote K11-linked polyubiquitination of RIP1 in TNF signalling. Embo j. 29, 4198–4209 (2010).
Xing, J., Zhang, A., Minze, L. J., Li, X. C. & Zhang, Z. TRIM29 Negatively Regulates the Type I IFN Production in Response to RNA Virus. J. Immunol. 201, 183–192 (2018).
Qin, Y. et al. RNF26 temporally regulates virus-triggered type I interferon induction by two distinct mechanisms. PLoS Pathog. 10, e1004358 (2014).
Jin, S. et al. USP19 modulates autophagy and antiviral immune responses by deubiquitinating Beclin-1. Embo j. 35, 866–880 (2016).
Liebl, M. P. & Hoppe, T. It’s all about talking: two-way communication between proteasomal and lysosomal degradation pathways via ubiquitin. Am. J. Physiol. Cell Physiol. 311, C166–C178 (2016).
Pohl, C. & Dikic, I. Cellular quality control by the ubiquitin-proteasome system and autophagy. Science 366, 818–822 (2019).
Kang, D. C. et al. mda-5: An interferon-inducible putative RNA helicase with double-stranded RNA-dependent ATPase activity and melanoma growth-suppressive properties. Proc. Natl Acad. Sci. USA 99, 637–642 (2002).
Arimoto, K. et al. Negative regulation of the RIG-I signaling by the ubiquitin ligase RNF125. Proc. Natl Acad. Sci. USA 104, 7500–7505 (2007).
Chen, W. et al. Induction of Siglec-G by RNA viruses inhibits the innate immune response by promoting RIG-I degradation. Cell 152, 467–478 (2013).
Wang, W. et al. RNF122 suppresses antiviral type I interferon production by targeting RIG-I CARDs to mediate RIG-I degradation. Proc. Natl Acad. Sci. USA 113, 9581–9586 (2016).
Park, Y. J. et al. Dual targeting of RIG-I and MAVS by MARCH5 mitochondria ubiquitin ligase in innate immunity. Cell Signal 67, 109520 (2020).
Tam, W. Y., Jiang, L. & Kwan, K. M. Transmembrane 6 superfamily 1 (Tm6sf1) is a novel lysosomal transmembrane protein. Protoplasma 252, 977–983 (2015).
Lu, K., den Brave, F. & Jentsch, S. Receptor oligomerization guides pathway choice between proteasomal and autophagic degradation. Nat. Cell Biol. 19, 732–739 (2017).
Webb, J. L., Ravikumar, B., Atkins, J., Skepper, J. N. & Rubinsztein, D. C. Alpha-Synuclein is degraded by both autophagy and the proteasome. J. Biol. Chem. 278, 25009–25013 (2003).
Yan, J. Q., Yuan, Y. H., Chu, S. F., Li, G. H. & Chen, N. H. E46K Mutant α-Synuclein Is Degraded by Both Proteasome and Macroautophagy Pathway. Molecules 23, 2839 (2018).
Liu, P. et al. Rab21 Protein Is Degraded by Both the Ubiquitin-Proteasome Pathway and the Autophagy-Lysosome Pathway. Int. J. Mol. Sci. 23, 1131 (2022).
Zhang, Q. et al. CNTNAP2 Protein Is Degraded by the Ubiquitin-Proteasome System and the Macroautophagy-Lysosome Pathway. Mol. Neurobiol. 60, 2455–2469 (2023).
Li, S. et al. The tumor suppressor PTEN has a critical role in antiviral innate immunity. Nat. Immunol. 17, 241–249 (2016).
Perez-Riverol, Y. et al. The PRIDE database at 20 years: 2025 update. Nucleic Acids Res. 53, D543–D553 (2025).
Acknowledgements
This study was supported by the National Natural Science Foundation of China (#82230075 to D.G., #32370152 to P.H., and #32470160 to C.L.), and the National Key Research and Development Program of China (#2021YFC2300100 and #2021YFC2300102 to C.L.). This study was also supported by Major Project of Guangzhou National Laboratory (GZNL2024A01016 to P.H.), Shenzhen Science and Technology Program (#JCYJ20200109142201695 to D.G.) and the 111 Program (#D20036 to Y.L.). Figure 7g was created with Medpeer (License No.: 181f5yildjdpyzixe21738569418) and Fig. 8i was generated with BioRender software (https://biorender.com/f67q662).
Author information
Authors and Affiliations
Contributions
P.H., C.L., and D.G. conceived and supervised the research; P.H., C.L., and M.H. designed the experiments, analyzed the data and wrote the manuscript. M.H., Z.Y., L.X., J.C., S.L., Z.L., B.Z., Y.Y., B.L., J.X., Y.Z., P.J., and Q.L. performed the biochemical, cell biological and in vitro experiments; Y.L. and L.L. generated the RNF167-deficient mice; M.H., L.X., J.C., and Z.L. performed the animal experiments and viral infections.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
He, M., Yang, Z., Xie, L. et al. RNF167 mediates atypical ubiquitylation and degradation of RLRs via two distinct proteolytic pathways. Nat Commun 16, 1920 (2025). https://doi.org/10.1038/s41467-025-57245-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-57245-3