Abstract
The biotransformation of nanoparticles plays a crucial role in determining their biological fate and responses. Although a few engineering strategies (e.g., surface functionalization and shape control) have been employed to regulate the fate of nanoparticles, the genetic control of nanoparticle biotransformation remains an unexplored avenue. Herein, we utilized a CRISPR-based genome-scale knockout approach to identify genes involved in the biotransformation of rare earth oxide (REO) nanoparticles. We found that the biotransformation of REOs in lysosomes could be genetically controlled via SMPD1. Specifically, suppression of SMPD1 inhibited the transformation of La2O3 into sea urchin-shaped structures, thereby protecting against lysosomal damage, proinflammatory cytokine release, pyroptosis and RE-induced pneumoconiosis. Overall, our study provides insight into how to control the biological fate of nanomaterials.
Similar content being viewed by others
Introduction
Engineered nanomaterials (ENMs) possess steric diameters similar to those of most viruses1,2. This resemblance can lead to confusion among immune cells, causing them to internalize ENMs via the same uptake pathway as viruses, namely, endocytosis for lysosomal digestion3. However, ENMs may have different cellular fates and induce diverse biological effects due to their distinct biotransformation in lysosomes, such as aggregation and agglomeration4, protein corona formation5, decomposition6, recrystallization7 and redox reactions8. For example, when silver (Ag) nanoparticles (NPs) are internalized by human T lymphocytes, they undergo a biotransformation process resulting in the formation of Ag2S and Ag thiolates9. This cellular detoxification pathway of Ag NPs highlights the critical role of biotransformation in determining the biological responses to ENMs. Consequently, considerable research interest has been directed toward regulating NP biotransformation to exert control over their biological fates10,11.
While it is well recognized that the physicochemical properties of ENMs, such as size, shape, surface charge and hydrophobicity, determine the nano-bio interactions10,12,13,14, researchers have explored innovative approaches to engineer specific functionalities on ENMs to achieve desired biological effects. For instance, lipid nanoparticles modified by SORT molecules have demonstrated increased bioavailability by overcoming the liver accumulation barrier15; sphingomyelin-modified camptothecin nanovesicles have shown improved pharmacokinetics and reduced systemic toxicity16; and grafting of functional groups on the surface of carbon dots has enhanced their efficiency in biomedical applications10. Nevertheless, the biotransformation of ENMs involves complex interactions with various biochemical species, and the constituents of the biological environment may also significantly impact the fate of ENMs8,17. For example, protons play a crucial role in the dissolution of ENMs such as ZnO, Fe2O3, and Al2O3 within lysosomes18,19. Since the biochemical constituents of lysosomes are regulated by certain genes (e.g., V-ATPase responsible for driving proton transport)20,21, we hypothesized that the biotransformation of ENMs could be genetically controllable.
In this study, we conducted a comprehensive screening of a metal oxide (MOx) nanoparticle library, and our investigation revealed significant lysosomal biotransformation of a representative rare earth oxide (REO), La2O3 NPs, along with associated adverse effects. To further understand the molecular basis of this biotransformation, we utilized genome-scale CRISPR knockout (GeCKO) screening in THP-1 cells to identify the specific genes involved in the transformation of La2O3 NPs (Fig. 1). Through this approach, we successfully identified SMPD1 as a key regulator of the lysosomal transformation process of La2O3 NPs. To ascertain the role of SMPD1 in this context, we examined the biological fate of La2O3 NPs as well as their adverse effects upon the suppression of SMPD1 both in vitro and in vivo. Overall, our findings offer an alternative approach for modulating the biological fates and effects of ENMs through genetic control of their transformation.
This schematic illustrates the methodology employed for genome-scale CRISPR knockout screens in THP-1 cells, including construction of CRISPR-Cas9 plasmid library and cell pool, La2O3 treatment, gDNA extraction, sgRNA amplification and sequencing. Some image elements were adapted from figures provided by Servier Medical Art, licensed under a Creative Commons Attribution 4.0 Unported License (https://creativecommons.org/licenses/by/4.0/).
Results
Biotransformation of La2O3 NPs in lysosomes
To study the biotransformation of NPs, we assembled a library of well-characterized MOx (Supplementary Table 1) and exposed them to phagolysosomal simulated fluid (PSF). Among the 60 tested MOx, we observed a significant morphology transformation of REO NPs, specifically La2O3, from rod or sphere shapes to sea urchin shapes, while TiO2, Au, Ag and Fe2O3 showed limited morphology changes (Fig. 2a, Supplementary Fig 1 and Supplementary Table 1). To delve deeper, La2O3 was selected as a representative rare earth-based nanoparticle for more explorations because it shares similar chemical characteristics with other REOs, such as Dy2O3, Gd2O3, and Eu2O3. Additionally, La2O3 is one of the most extensively studied REOs with controllable morphologies in the literatures2,22, providing a robust foundation of existing knowledge to build upon. Four MOx, including La2O3 spheres (La2O3-S), La2O3 rods (La2O3-R), Fe2O3 spheres (Fe2O3-S) and Fe2O3 rods (Fe2O3-R), were therefore selected. The transformation of La2O3 was confirmed by X-ray powder diffraction (XRD), where the characteristic peaks of hexagonal LaPO4 were detected (Supplementary Fig 2). These results suggest that phosphates are the key constituents dictating the biotransformation in PSF. Notably, reactive oxygen species (ROS) and proteins are two additional significant factors present in lysosomes23. To assess the impacts of ROS and proteins on the biotransformation of nanoparticles, La2O3-S was exposed to PSF containing 100 μM H2O2 or 1 mg/mL bovine serum albumin (BSA) for 24 h. Interestingly, the presence of H2O2 or proteins had minimal effects on the biotransformation of La2O3 (Supplementary Fig 3).
a Transmission electron microscopy (TEM) images displaying the transformation of representative nanoparticles (NPs) after exposure to deionized (DI) water or phagolysosomal simulated fluid (PSF). Treated NPs were suspended in DI water and observed via TEM. b Confocal microscopy imaging of La2O3 in cells. Cells incubated with 12.5 μg/mL FITC-labeled La2O3 NPs for 12 h were fixed, stained with Hoechst 33342 (blue) and Alexa Fluor 594-conjugated LAMP1 antibody (red) for visualizing nuclei and lysosomes, respectively. c TEM visualization of La2O3 biotransformation in lysosomes. THP-1 cells treated with 30 μg/mL La2O3-S or Fe2O3-S for 24 h were fixed for TEM observation. Blue arrows show the representative nanoparticles in cells. Energy dispersive X-ray (EDX) was used for particle composition analysis. d Confocal microscopy imaging of cathepsin B release in cells. THP-1 cells treated with La2O3 or Fe2O3 NPs were stained with a cathepsin B fluorogenic substrate (red) and Hoechst 33342 (blue) for confocal imaging. e IL-1β quantification in THP-1 cell supernatants. Supernatants from NP-treated THP-1 cells underwent IL-1β quantification via ELISA. Data are means with SDs (n = 4 biologically independent cell cultures). ***p < 0.001 compared with control (Ctrl) treatment by one-way ANOVA test. f Dose-response curves of THP-1 cell viability against La2O3 and Fe2O3 NPs, data are means with SDs (n = 3 biologically independent cell cultures). For (a–d), the representative images were selected from 3 independent experiments with similar results. Source data are provided as a Source Data file.
In addition to PSF, we also examined the morphology and composition of La2O3 after treatment in endosomal simulated fluid (ESF)24, since nanoparticles are initially encapsulated into endosomes during endocytosis. Specifically, La2O3-S was exposed to ESF and PSF for 3, 6, and 12 h, followed by transmission electron microscopy (TEM) examination of morphology changes and inductively coupled plasma optical emission spectrometry (ICP-OES) quantification of the ratio of phosphorus to lanthanum elements (P/La). While La2O3 fully transformed into sea urchin-like structures with 80% conversion in PSF at 12 h, only a few whiskers were observed in ESF-treated La2O3, resulting in a mere 11% transformation (Supplementary Fig 4). These results indicate that the biotransformation of La2O3 primarily occurs in lysosomes.
After labeling MOx with fluorescein isothiocyanate (FITC), we examined the cellular distribution of MOx in THP-1 cells, a macrophage-like cell line that serves as the first line of defense in biological systems against particulate stimuli. Confocal microscopy revealed significant colocalization of FITC-labeled MOx with Alexa Fluor 594-labeled LAMP1-positive compartments, i.e., lysosomes, with a high degree of overlap ranging from 76.7% to 92.4% (Fig. 2b and Supplementary Fig 5), as confirmed by TEM images of MOx-treated cell sections (Fig. 2c and Supplementary Fig 6). Notably, while Fe2O3-S and Fe2O3-R maintained their original morphologies in lysosomes, La2O3-S and La2O3-R transformed into sea-urchin-like structures, accumulating substantial phosphorus elements, which suggested that La2O3 interacted with phosphates in lysosomes and recrystallized into hexagonal LaPO4.
The biotransformation of La2O3 resulted in the impairment of lysosomal membranes, leading to the release of lysosomal cargos, e.g., cathepsin B, into the cytoplasm. This was evidenced by the diffuse red fluorescence of cathepsin B substrates in Fig. 2d. In contrast, these substrates appeared as dots in untreated or Fe2O3-treated cells, indicating that cathepsin B was confined to intact lysosomes. The released cathepsin B is known to contribute to the recruitment of the NLRP3 inflammasome, resulting in caspase-1 activation. This is evidenced by the intense green fluorescence of the fluorogenic caspase-1 substrate observed in the La2O3 treatments (Supplementary Fig 7). Consequently, caspase-1 catalyzes pro-IL-1β, leading to the release of up to 1500 pg/mL of IL-1β into THP-1 cell supernatants (Fig. 2e). Moreover, both La2O3-S and La2O3-R induced dose-dependent cytotoxicity (Fig. 2f), resulting in 30% and 40% cell death at 30 μg/mL, respectively. In contrast, neither Fe2O3-S nor Fe2O3-R induced a decrease in cellular viability. This cell death could be attenuated by a pyroptosis inhibitor, VX-765, whereas other cell death inhibitors targeting apoptosis, necrosis and ferroptosis had no effect on La2O3-induced cell death (Supplementary Fig 8). These results indicate that excessive caspase-1 activation can also proteolyze the maturation of the pore-forming protein gasdermin D (GSDMD), ultimately causing pyroptotic cell death.
We examined the cellular uptake of La2O3 and Fe2O3 nanoparticles to determine whether it contributes to the hazardous effects of La2O3. Specifically, the cellular uptake of La2O3-R, La2O3-S, Fe2O3-R, and Fe2O3-S was quantified using ICP-OES to measure Fe or La elements within THP-1 cells. As illustrated in Supplementary Fig 9a, rod-shaped nanoparticles exhibited relatively higher cellular uptake (0.36–0.37 mM/g protein) compared to spherical nanoparticles. However, Fe2O3 and La2O3 nanoparticles of the same shape demonstrated similar levels of cellular uptake. Furthermore, we normalized the IL-1β levels by the cellular uptake levels of the nanoparticles. Both La2O3-S and La2O3-R elicited similar levels of IL-1β in the supernatants of THP-1 cells, while Fe2O3-S and Fe2O3-R had minimal effects (Supplementary Fig 9b). These findings reinforce our assertion that biotransformation, rather than cellular uptake, is responsible for the observed adverse effects of REOs. To explore the relationship between the cytotoxicity and biotransformation, La2O3 was dispersed in ethylenediamine tetra (methylenephosphonic acid) (EDTMP) solution to form a thin coating layer, preventing La3+ leakage, precipitation with phosphate, and biotransformation. As shown in Supplementary Fig 10a, the coated nanoparticle (EDTMP- La2O3) did not show significant morphology change in PSF and maintained the spherical shape, unlike the sea urchin structure of transformed particles (T- La2O3). We then compared the toxicity of La2O3, T- La2O3 and EDTMP- La2O3 in THP-1 cells. Interestingly, while La2O3 elicited significant cell death, T- La2O3 and EDTMP- La2O3 had no effects on the viability of THP-1 cells (Supplementary Fig 10b). These results indicate that the transformed sea-urchin shape is not responsible for cytotoxicity. Instead, the transformation process itself, specifically the interaction with phosphates, appears to be responsible for the observed cytotoxicity.
GeCKO screening to identify genes involved in the transformation of La2O3
To identify the genes implicated in the cytotoxicity induced by La2O3 NPs, we conducted a GeCKO screening in THP-1 cells (Fig. 1). Initially, we generated a plasmid library consisting of 64,751 sgRNA plasmids targeting 18,080 human genes, with an average of 3–4 sgRNAs per gene, and established a GeCKO THP-1 cell pool by lentivirus transduction. Before transduction, we validated the plasmid library through deep sequencing (Supplementary Fig 11). Following transduction, over 99.4% of genes (17,972 genes) were successfully knocked out in the cell pool (Supplementary Fig 12a). The remaining 108 genes were associated with fundamental cellular functions (Supplementary Fig 12b), implying that their loss of function would directly induce THP-1 cell death and, therefore, were not relevant to our study.
Subsequently, GeCKO THP-1 cells were exposed to La2O3 NPs and subjected to deep sequencing. We compared the gene representation (sgRNA abundance) quantitatively between La2O3 treatment and vehicle control to identify genes implicated in the cytotoxicity of La2O3 NPs. Overrepresented genes were the candidates significantly contributing to cytotoxicity. We identified 194 and 185 overrepresented genes (p value < 0.01, Supplementary Tables 2 and 3) in GeCKO THP-1 cells exposed to La2O3-S and La2O3-R, respectively. Notably, twenty genes were common to both groups (Fig. 3a). Among the twenty genes, SMPD1 stood out as the most significantly overrepresented one, belonging to the lysosomal phospholipid hydrolase family (Supplementary Fig 13a, b, and Supplementary Tables 4 and 5). SMPD1 specifically codes for the protein of acid sphingomyelinase (ASM), a lysosomal acid sphingomyelinase. To validate the involvement of SMPD1, we examined the expression of ASM in La2O3-treated THP-1 cells using Western blotting (Fig. 3b). As expected, exposure to La2O3 induced 12.0–15.3-fold overexpression of ASM in cell lysates. Since ASM functions by hydrolyzing sphingomyelin, a phospholipid presents in lysosomal membranes, into ceramide and phosphorylcholine (Fig. 3c), we further investigated the role of ASM by imaging and quantifying ceramide production in THP-1 cells (Fig. 3b). As shown in Fig. 3d, THP-1 cells treated with La2O3 exhibited strong red fluorescence upon staining with ceramide antibody, indicative of increased ceramide levels, while the control cells displayed only faint emissions, suggesting limited ceramide production. Quantification results demonstrated a 6.3–8.1-fold increase in ceramide in THP-1 cells upon La2O3 treatment (Fig. 3e). These results suggested that ASM-mediated hydrolysis of sphingomyelin could be activated by La2O3 treatment. STRING network analysis revealed that SMPD1 was closely associated with GSDMD, a critical hallmark in pyroptosis (Supplementary Fig 13c). All these findings strongly validated the identified gene, SMPD1, and its involvement in La2O3 treatment.
a GeCKO screening for La2O3-induced gene representation changes. p-values were calculated based on gene representation in La2O3 and vehicle treatments using RNAi Gene Enrichment Ranking (RIGER). b Western blot analysis of acid sphingomyelinase (ASM) coded by SMPD1, the representative images were selected from 3 independent experiments with similar results. c Schematic illustrating ASM-catalyzed hydrolysis reaction, converting sphingomyelin into ceramide (Cer) and phosphorylcholine. d Detection of Cer production in THP-1 cells via fluorescence imaging. Wild-type and SMPD1 knockout (SMPD1−/−) cells exposed to La2O3 NPs were stained with ceramide monoclonal antibody (MID 15B4) and DAPI to visualize Cer (red) distribution and nuclei (blue), respectively (scale bar: 60 μm). Left panel: representative microscopy images were selected from 4 independent experiments with similar results; right (e): quantification results based on mean integrated optical density per unit area (MIOD) of Cer fluorescence. Bar plots represent means with SDs (n = 4 independent experiments), ***p < 0.001 compared with wild-type cells (THP-1) by two-way ANOVA test followed by Bonferroni post-tests. Source data are provided as a Source Data file.
SMPD1 facilitates the biotransformation and biological effects of La2O3 NPs
The transformation of La2O3 in lysosomes relies on an adequate supply of phosphates, and SMPD1 plays a pivotal role in providing these phosphates within lysosomes. Based on this premise, we hypothesized that SMPD1 is likely responsible for the biotransformation and biological effects of La2O3 NPs. To test this hypothesis, La2O3-S and La2O3-R were incubated with phosphorylcholine (ChoP), a degradation product of sphingomyelin catalyzed by SMPD1, in PSF to replace phosphates. As shown in Supplementary Fig 14, similar sea urchin-like structures were observed on both types of La2O3 nanoparticles, displaying the hexagonal LaPO4 phase. These results suggest that the biotransformation of both La2O3-S and La2O3-R in PSF can be replicated in acidic ChoP fluid. Additionally, we investigated the regulatory role of SMPD1 in the lysosomal transformation of La2O3 NPs in both wild-type (WT) and SMPD1 knockout (SMPD1−/−) THP-1 cells. As expected, SMPD1−/− cells exhibited limited ceramide production upon La2O3 treatments (Fig. 3d, e). This likely resulted in insufficient lysosomal phosphate supply, consequently hindering La2O3 transformation. To further validate this, we compared the biotransformation of La2O3 NPs in both WT and SMPD1−/− cells. In stark contrast to the significant transformation of La2O3 into needle-like structures observed in the lysosomes of WT cells, we did not observe significant morphological changes in La2O3 in the lysosomes of SMPD1−/− cells (Fig. 4a and Supplementary Fig 15). Additionally, we attempted to quantify the ratios of phosphorus (P) and lanthanum (La) elements in lysosomal internalized La2O3 in both cell types. For this purpose, we constructed a core-shell nanostructure (La2O3 @ Fe3O4), comprising La2O3 layer and Fe3O4 core (Supplementary Fig 16) for incubation with WT and SMPD1−/− cells. The La2O3@Fe3O4 NPs internalized into lysosomes were then extracted under a magnetic field and subjected to ICP‒OES analysis for P and La element quantification. As illustrated in Fig. 4b and Supplementary Fig 17, the element ratio of P vs. La in La2O3@Fe3O4 NPs extracted from the lysosomes of WT cells was twofold higher than that from SMPD1−/− cells, and significant difference in the transformation of the nanoparticles was also observed. Moreover, we examined the transformation dynamics of La2O3 in THP-1 cells by conducting ICP-OES analysis of P and La elements in lysosome-internalized La2O3@Fe3O4 at various incubation time points. As illustrated in Supplementary Fig 18, La2O3 exhibited rapid transformation within the initial 6 h, reaching a plateau after 12 h. These results well demonstrate that SMPD1 regulates the biotransformation of La2O3 NPs in lysosomes.
a TEM imaging of La2O3-S in wild-type and SMPD1−/− THP-1 cells. THP-1 cells were treated with 30 μg/mL La2O3 spheres for 24 h, followed by fixation and staining for TEM observation. Blue arrows show the representative nanoparticles in cells. b Quantification of lanthanum (La) and phosphorus (P) in La2O3 extracted from wild-type and SMPD1−/− cells. NPs were extracted from the lysosomes of THP-1 cells treated with 50 μg/mL La2O3@Fe3O4 for 24 h under magnetic field. The extracted NPs were subjected to ICP-OES analysis of La and P elements. Bar plots represent means with SDs (n = 3 biologically independent cell cultures), **p < 0.01 compared with wild-type cells (THP-1) by one-way ANOVA test. c Imaging of cathepsin B release upon La2O3 treatment. Cells treated with La2O3 were stained with a Magic Red® cathepsin-B assay kit for visualizing cathepsin B distribution. d Comparison of caspase-1 activity in wild-type or SMPD1−/− cells following La2O3 treatments. THP-1 cells exposed to La2O3 were stained with a fluorogenic probe for fluorescence intensity measurement. e IL-1β quantification in cell supernatants. f Cell viability of wild-type and SMPD1−/− THP-1 cells following La2O3 treatment. THP-1 cells were exposed to 30 μg/mL La2O3 NPs for 72 h. For (a and c), the representative images were selected from 3 independent experiments with similar results. For (d, e, and f) data are means ± SDs (n = 5 biologically independent cell cultures), ***p < 0.001 compared with wild-type (THP-1) cells by two-way ANOVA test followed by Bonferroni post-tests. Source data are provided as a Source Data file.
Furthermore, we investigated whether SMPD1 impacted the adverse effects of La2O3 by comparing the responses of WT and SMPD1−/− cells to La2O3 exposure. We examined La2O3-induced cathepsin B release, caspase-1 activation, inflammatory cytokine release, and cell death in both types of cells. The fluorescent signals of cathepsin B substrate were rarely detected in the cytoplasm of La2O3-treated SMPD1−/− cells (Fig. 4c), indicating limited cathepsin B release. Consequently, the levels of La2O3-activated caspase-1 decreased by 3.3–4.9-fold in SMPD1−/− cells (Fig. 4d), and IL-1β in supernatants decreased from 1200 pg/mL to a basal level of 100 pg/mL (Fig. 4e). Additionally, the viability of THP-1 cells exposed to La2O3 increased by 30–40% after SMPD1 knockout (Fig. 4f). These compelling findings strongly suggested that SMPD1 regulated the adverse effects of La2O3 NPs.
In addition to THP-1 cells, we investigated whether the key findings could be replicated in other macrophage-like cells. Specifically, a mouse macrophage cell line, RAW 264.7, was exposed to La2O3, Eu2O3, and Gd2O3 to examine the morphology changes of nanoparticles, IL-1β release, and cytotoxicity. As depicted in Supplementary Fig 19a, these particles underwent significant morphology changes into mesh- or urchin-like structures in the lysosomes of RAW 264.7 cells. Consequently, these REOs elicited IL-1β release in the supernatants and induced cell death (Supplementary Fig 19b, c). Notably, these adverse effects were significantly attenuated in desipramine (DPM)-treated cells, where the activity of SMPD1 is inhibited.
Validation of the regulatory roles of SMPD1 in vivo
To further investigate the regulatory role of SMPD1 in vivo, we compared the pulmonary toxicity of La2O3 NPs in normal and DPM-treated mice. After 48 h of exposure, we collected alveolar macrophages from the mice lungs for further examination. As illustrated in Fig. 5a, both La2O3-S and La2O3-R elicited high levels of ASM in alveolar macrophages, and DPM significantly inhibited the expression of ASM in various treatments. Subsequently, the alveolar macrophages were subjected to TEM observation. Consistent with the in vitro results, suppression of ASM inhibited the biotransformation of La2O3 NPs in the lysosomes of primary alveolar macrophages (Fig. 5b and Supplementary Fig 20).
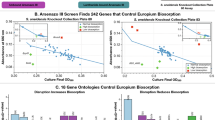
a Western blot analysis of ASM expression, the representative images were selected from 3 independent experiments with similar results. b TEM visualization of La2O3 in alveolar macrophages from bronchoalveolar lavage fluid (BALF). Blue arrows show the representative nanoparticles in cells. Quantification of (c) IL-1β and (d) TGF-β1 in BALF, bar plots represent means with SDs (n = 6 biologically independent mice), ***p < 0.001 compared with the mice without treatment of desipramine (DPM) by one-way ANOVA test. e H&E staining and (f) Masson trichrome staining of lung tissues. Mice pretreated with or without DPM were exposed to La2O3 via oropharyngeal aspiration (n = 6 mice for each treatment). Alveolar macrophages were obtained from BALF. Scale bars in (e and f) represent 150 μm. Source data are provided as a Source Data file.
In addition, we examined the biodistribution of La2O3 in lung tissues using confocal microscopy and its biotransformation using TEM. FITC-labeled La2O3-S nanoparticles with green fluorescence were instilled into animal lungs to visualize their interactions with lung cells, including macrophages and epithelial cells (Supplementary Fig 21a). Confocal images revealed a significant overlap between FITC-labeled La2O3-S and lung macrophages, indicating that these nanoparticles were primarily processed by lung macrophages. In contrast, the colocalization between La2O3-S and epithelial cells was relatively low. Notably, La2O3-S nanoparticles were surrounded by IL-1β+ cells, suggesting that La2O3-S recruits more IL-1β+ cells and elicits focal inflammation in lung tissues. Moreover, La2O3-treated lung tissues were fixed, stained, and sectioned for TEM visualization (Supplementary Fig 21b). Although similar endosomal/lysosomal internalization patterns were observed for La2O3 in lung tissue cells, the number of internalized nanoparticles in lung tissue cells was fewer than that in THP-1 cells. However, significant morphological changes from spherical shapes into mesh-like structures could still be clearly differentiated in lung cells. These in vivo findings provide robust evidence for the biotransformation and biodistribution of La2O3 nanoparticles, highlighting their interaction with lung macrophages and their role in eliciting inflammatory responses.
After 14 d of exposure, the mice were sacrificed to examine cytokine production in bronchoalveolar lavage fluid (BALF) and pathogenic changes in lung tissue. We found elevated levels of the proinflammatory cytokine IL-1β (Fig. 5c) and the profibrogenic growth factor TGF-β1 (Fig. 5d) in the BALF of mice exposed to La2O3, indicating that inhalation exposure to La2O3 may cause lung inflammation or fibrosis. Additionally, we examined lung sections using hematoxylin and eosin (H&E) and trichrome staining. The intensity of focal pulmonary infiltrates in H&E staining reflected the inflammatory response in the BALF (Fig. 5e), while collagen deposition around small airways in trichrome-stained lung sections confirmed the profibrogenic effect (Fig. 5f). Notably, suppression of ASM by DMP significantly attenuated La2O3-induced inflammation and profibrogenic effects in the lungs of the animals (Fig. 5). These in vivo results further corroborate the crucial regulatory roles of SMPD1 in the biotransformation and adverse effects of La2O3 NPs.
Overall, our findings provide valuable insights into the regulatory mechanism governing the biotransformation of La2O3 NPs (Fig. 6). Upon cellular uptake, La2O3 NPs are directed to lysosomes through endocytosis. Within the acidic lysosomal environment, La2O3 undergoes dissolution, leading to the release of La3+ ions, which subsequently precipitate with phosphates to form LaPO4. The pivotal role played by SMPD1-encoded ASM becomes evident in this process, as it catalyzes the hydrolysis of sphingomyelin into ceramide and phosphorylcholine, thereby supplying the needed phosphates for efficient biotransformation. As a consequence of biotransformation, lysosomal membrane permeabilization occurs, followed by the release of cathepsin B into the cytoplasm. This triggers the recruitment of the NLRP3 inflammasome and caspase-1 activation, culminating in the release of the proinflammatory cytokine IL-1β and pyroptotic cell death.
REO NPs, once internalized into cell lysosomes, release Re3+ ions in the acidic environment. These released Re3+ ions can then precipitate with phosphates, forming sea-urchin-shaped RePO4 NPs. The enzyme ASM, encoded by SMPD1, catalyzes the hydrolysis of sphingomyelin, supplementing the phosphate pool during the biotransformation of REO NPs. This molecular process leads to lysosomal damage, release of cathepsin B, activation of the NLRP3 inflammasome, and initiation of pyroptosis. These events culminate in lung inflammation and fibrosis in animals. Some image elements were adapted from figures provided by Servier Medical Art, licensed under a Creative Commons Attribution 4.0 Unported License (https://creativecommons.org/licenses/by/4.0/).
Discussion
The biotransformation of ENMs is a crucial factor that influences their biological efficacy and potential health implications2,11. Therefore, understanding and controlling the biotransformation of ENMs are important for harnessing their beneficial properties while mitigating potential risks25,26. For instance, the biotransformation of La2O3 has been reported in cucumber roots, where it forms needle-like LaPO4 nanoclusters that result in phytotoxicity27. Similarly, Zheng et al. observed comparable transformation behavior on bacterial cell membranes, which led to pulmonary microbiota imbalance under respiratory exposure conditions28. In this study, we focused on investigating the lysosomal biotransformation of REO NPs and its association with lysosomal damage and pyroptosis in macrophages. Notably, our results revealed the significant role of SMPD1 in regulating the biotransformation and adverse effects of La2O3 NPs. By genetically muting SMPD1 or chemically inhibiting ASM encoded by SMPD1, we successfully prevented the lysosomal biotransformation of La2O3 NPs and attenuated their adverse health effects. These findings highlight SMPD1 as a potential key target for controlling the biotransformation of REO NPs.
We employed a novel approach, GeCKO screening to uncover genes involved in the biotransformation of REO NPs. The biotransformation of REOs often involves specific biochemical reactions with biomolecules, such as phosphates or phospholipids2,11,26,27,29, catalyzed by corresponding enzymes29. However, identifying the causal associations between enzyme-encoding genes and biological effects is challenging30. While conventional omics techniques, such as transcriptomics and proteomics, provide potential toxicological mechanisms by identifying changes in gene product abundance, they may not establish causal relationships between genes and toxicological phenotypes30,31. In contrast, GeCKO screening in human macrophages successfully revealed causal associations between genes and the biological effects induced by REO NPs. This paradigm offers a promising tool to investigate other biotransformable ENMs, helping to identify key regulatory genes and signaling pathways in their biotransformation. Beyond SMPD1, among the other 19 genes (Supplementary Table 6), three genes (CASP1, CASP3, and GSDMD) are known to directly regulate pyroptosis, which is the endpoint event of the adverse outcome pathway of REOs. Additionally, MAX and KBTBD4 are also involved in the cell death caused by REOs. Eleven genes (SGIP1, TBX1, KRTAP1-5, MYBPC1, EPB41L2, POC5, MINDY1, ILF3, TMEM184B, RYR3, and NEB) regulate cell motility by acting on the membrane system or cytoskeleton, and may play critical roles in the endocytosis of REOs. Three genes (GIGYF2, ZG16, and OPRM1) function to regulate cellular protein trafficking by acting on the Golgi apparatus or plasma membrane.
REO-induced pulmonary diseases, such as rare earth pneumoconiosis32,33, have emerged as significant challenges for the sustainable development of RE-based nanotechnology. The identification of SMPD1 and its associated signaling pathways involved in REO-induced toxicity opens avenues for developing novel preventive and therapeutic strategies. For instance, our findings suggest that DPM, a FDA-approved medicine used to treat depression32, may effectively prevent REO toxicity by inhibiting ASM activity. This exciting finding may guide the design and development of targeted interventions for pneumoconiosis, offering new hope for the treatment of this chronic respiratory disease. Presently, pneumoconiosis affects over half a million patients globally34, but effective medications are lacking. The potential repurposing of DPM for pneumoconiosis treatment holds significant promise and warrants further investigation.
In our study, the sea urchin-like transformation into insoluble metal phosphates was exclusively observed in rare earth (RE)-based nanoparticles. Although some transition metals, such as Zn²+35,36, Cu²+, and Ti³+, can also form precipitates with phosphates, we did not identify transformed crystal metal phosphates in lysosomes. This can be attributed to the stringent criteria required for nanoparticle recrystallization in lysosomes: (i) an appropriate dissolution rate (neither too fast nor too slow) to facilitate the crystallization process, and (ii) a strong binding affinity with phosphate (with a solubility product constant, ksp ~10²6) to drive the dephosphorylation of biomolecules in lysosomes. In this study, we deliberately compared the adverse effects of two La2O3 with rod and sphere shapes. La2O3-S nanoparticles, with their large surface area, have a higher probability of reacting with the surrounding media, potentially enhancing their interaction with lysosomal components. On the other hand, La2O3-R nanoparticles may exhibit more surface defects, which could increase the release of La³+ ions, influencing their biotransformation dynamics. However, given that both the surface area and defects play crucial roles in the dissolution process, La2O3-S and La2O3-R displayed similar transformation behaviors in cells, as evidenced by the formation of mesh- or urchin-like structures in lysosomes. Consequently, both shapes elicited comparable levels of IL-1β release and cell death.
This study has provided a distinct approach for regulating the biological fates and effects of ENMs. We have demonstrated that the intracellular biotransformation of ENMs is genetically controllable. SMPD1 was identified as a key target for regulating the lysosomal biotransformation of REO NPs and impacting downstream toxic signaling cascades, including lysosomal impairment, cathepsin B release, caspase-1 activation, IL-1β production, pyroptotic cell death, inflammation and profibrogenic effects. The identification of this key regulatory gene provides insights for the prevention of adverse outcomes of rare earth-based nanotechnology.
Methods
Materials
We acquired a library of metal oxide (MOx) nanoparticles (NPs) through in-house synthesis37,38 or from US-Nano (Houston, TX, USA). Phagolysosomal simulated fluid (PSF; 142 mg/L Na2HPO4, 6.65 g/L NaCl, 62 mg/L Na2SO4, 29 mg/L CaCl2 ·3 H2O, 250 mg/L glycine, 8.09 g/L potassium phthalate, pH 4.5) was prepared7. Ethylenediamine tetra (methylenephosphonic acid) coated La2O3 (EDTMP-La2O3) was prepared by dispersing La2O3 in EDTMP solution to form a thin coating layer, preventing La3+ leakage, precipitation with phosphate, and biotransformation. Transformed La2O3 (T-La2O3) was prepared by incubating La2O3 in PSF for 24 h to transform into sea urchin-like structures. Magic Red Cathepsin B Assay Kit was purchased from Immunochemistry (Bloomington, MN, USA). Hoechst 33342, DAPI and acid sphingomyelinase (ASM) polyclonal antibody (PA5-95730) were purchased from Thermo Scientific (Waltham, MA, USA). Alexa fluor 594-conjugated LAMP1 antibody (ab302684), anti-ASM antibody (ab272729), goat anti-rabbit IgG H&L (HRP) (ab6721), Anti-GAPDH antibody (ab9485), goat anti-mouse IgM mu chain (Alexa Fluor 647) antibody (ab150123) were purchased from Abcam (Cambridge, UK). Ceramide monoclonal antibody (MID 15B4, ALX-804-196-T050) was purchased from Enzo Life Science (Raamsdonksveer, The Netherlands). Mouse F4/80 antibody (ab300421), mouse prosurfactant protein c (SPC) antibody (ab211326), and mouse IL-1β antibody (ab315084) were purchased from Abcam. ELISA kits for detection of human or mouse IL-1β (VAL101 or VAL601), mouse TGF-β1 (VAL611) were purchased from R&D Systems (Minneapolis, MN, USA). MTS assay kit (G3580) and Caspase-Glo 1 Inflammasome Assay (G9951) were purchased from Promega (Madison, WI, USA). Pierce BCA kit (23252, Thermo Scientific) was used to quantify protein concentration. Phosphorylcholine (P0378) and bovine serum albumin (BSA, 2905-OP) were purchased from Sigma-Aldrich. Other chemicals, unless stated otherwise, were purchased from Sigma-Aldrich (St. Louis, MO, USA).
Characterization of Mox
The morphology of MOx was observed by transmission electron microscopy (TEM) on a JEOL 1200 EX instrument (accelerating voltage: 80 kV) or a FEI Tecnai F20 instrument. Primary sizes and surface charges in DI water were assessed using dynamic light scattering (DLS) coupled with Zeta Potential Analyzer (ZPA) from Brookhaven Instruments Corporation, Holtsville, NY, USA. X-ray diffraction (XRD) analysis was conducted on a Philips X’Pert Pro diffractometer equipped with CuKα radiation. Quantitative analysis of metal and phosphorus elements was performed using inductively coupled plasma optical emission spectrometry (ICP-OES) on an ICAP7200 instrument by Thermo, America.
MOx transformation in PSF
MOx NPs were dissolved in 200 mL of PSF buffer at a concentration of 100 μg/mL and subjected to sonication at 32 W for 15 s. Following an incubation period of 24 h at 37 °C, the MOx NPs were harvested through centrifugation at 32479 × g for 10 min. The obtained pellets were washed with deionized water (DI H2O) and subsequently dried at 60 °C overnight. The dried nanomaterial was then pulverized for XRD analysis and TEM observation. Meanwhile, the supernatants were collected to quantify ion release using ICP-OES.
Cell culture, subcellular localization, and NP biotransformation in lysosomes
THP-1 cells (ATCC, Manassas, VA) and SMPD1−/− THP-1 cells were cultured in Roswell Park Memorial Institute 1640 medium (RPMI1640, Gibco), supplemented with 10% fetal bovine serum (FBS, Gibco), within a humidified environment containing 5% CO2 at 37 °C. RAW264.7 cells (ATCC, Manassas, VA) were maintained in Dulbecco’s Modified Eagle’s Medium (DMEM, Gibco), supplemented with 10% FBS (Gibco). The subcellular distribution of MOx within THP-1 cells was visualized through confocal microscopy (Leica confocal SP2 1 P/FCS microscopes) and TEM. For confocal imaging, cells were exposed to 12.5 μg/mL FITC-labeled MOx for 12 h. Subsequently, the treated cells were rinsed with PBS and then fixed in 4% paraformaldehyde. Nuclei were stained using Hoechst 33342 dye, while lysosomes were highlighted using Alexa Fluor 594-conjugated LAMP1 antibody (1:50). Regarding TEM observation, THP-1 cells were treated with La2O3 or Fe2O3 NPs (30 μg/mL spheres or 15 μg/mL rods) for a duration of 24 h. Post-treatment, the cells were washed, fixed, and stained, and TEM grids were prepared according to established protocols7,39. EDX analysis was employed to characterize the elemental composition of internalized NPs.
Cell viability, cathepsin B release, caspase-1 activity, and IL-1β release analysis
THP-1 cells were seeded at a density of 3 × 104 per well in 96-well plates and primed with 1 μg/mL phorbol 12-myristate acetate (PMA) overnight. The cells were subsequently exposed to particle suspensions at desired concentrations in complete RPMI1640 medium for an additional 72 h. For assessing cellular viability, 20 μL of Cell Titer 96 Aqueous One Solution Reagent (Promega) was added to each well, followed by an incubation of 1 h at 37 °C. The optical density of each well was then measured at 490 nm using a microplate reader (BioTek). Cathepsin B release was determined by treating THP-1 cells with La2O3 or Fe2O3 NPs (8.0 μg/mL spheres or 4.0 μg/mL rods) for 72 h. Following treatment, cells were washed with PBS, stained with Magic Red for 1 h, and fixed with 4% paraformaldehyde for 15 min. After an additional PBS wash, nuclei were stained with Hoechst 33342 for 10 min. High-content screening (Operetta) was utilized for cell visualization. To evaluate caspase-1 activity and IL-1β release, THP-1 cells were treated with La2O3 or Fe2O3 NPs (8.0 μg/mL spheres or 4.0 μg/mL rods) for 72 h. Caspase-Glo 1 Inflammasome Assay (Promega) and FAM-FLICA-labeled caspase-1 probe were employed to measure caspase-1 activity within THP-1 cells, and the stained cells were imaged by confocal microscope (FV 1200, Olympus, Japan). IL-1β ELISA kit (VAL101) was used to measure IL-1β concentrations in the culture supernatants. RAW264.7 cells were seeded at a density of 3 × 104 per well in 96-well plates, after attachment overnight, cells were treated with 30 μM desipramine or DMEM for 2 h, and then cells were exposed to 50 μg/mL La2O3-S, La2O3-R, Eu2O3 and Gd2O3 for 24 h, respectively. IL-1β concentrations in the culture supernatants were measured by IL-1β ELISA kit (VAL601), and the cellular viability was measured with the above method.
Effects of necroptosis, apoptosis, pyroptosis and ferroptosis inhibitors on La2O3-S and La2O3-R induced cytotoxicity
THP-1 cells were pretreated with 50 μM Nec-1, 20 μM Z-VAD-FMK, 5 μM VX-765 or 5 μM Ferr-1 for 1 h, respectively. Then the cells were exposed to 50 ng/mL TNF-α + 1 μg/mL cycloheximide (CHX), 1 μg/mL doxorubicin (DOX), 5 μg/mL nigericin, 5 μg/mL RSL3, 100 μg/mL La2O3-S or 100 μg/mL La2O3-R for 72 h, respectively. Finally, the cell viability was measured by MTS assay (same as the above).
Construction of GeCKO THP-1 cell pool
The construction of the GeCKO THP-1 cell pool was conducted by transfection using lentivirus31,40. In brief, the plasmids were sourced from the GeCKO version 1 library (Broad Institute of MIT and Harvard, Cambridge, MA), encompassing 64,751 single guide RNA (sgRNA) plasmids targeting 18,080 human genes, with an average of 3–4 sgRNAs per gene. These plasmids were incorporated into lentiviral vectors for transfection into THP-1 cells. Lentiviral packaging was performed through cotransfection of GeCKO library plasmids alongside two viral packaging plasmids, pVSVg and psPAX2 (Addgene), into HEK293T cells (ATCC, Manassas, VA) at 80–90% confluency. The plasmids were introduced into HEK293T cells using Opti-MEM (Thermo Scientific), Lipofectamine 2000 (Thermo Scientific), and PLUS reagent (Thermo Scientific) in accordance with the manufacturer’s protocol. After 6 h, the media were replaced with fresh D10 media (DMEM supplemented with 10% FBS). After 60 h post-transfection, the media were harvested and centrifuged at 83146 × g for 2 h at 4 °C to collect lentiviruses. The titer of the virus was determined to be 2.2 × 108 IFU/mL using the Lenti-X p24 Rapid Titer Kit (TaKaRa). For cell transduction, 40 μL of lentivirus and 8 μg/mL polybrene (Sigma) were added to 3 × 106 THP-1 cells per well of four 12-well plates. The volume of virus was optimized to ensure a multiplicity of infection (MOI) of 0.3, ensuring most cells received at most one genetic perturbation. The four plates were centrifuged at 149 × g for 4 h at 33 °C. Following a 24 h incubation, the medium was changed, and the cells were transferred to 225 cm2 flasks to eliminate untransfected cells. This was achieved through puromycin treatment (0.4 μg/mL) for 10 days. Subsequently, a total of 3 × 107 cells were harvested for genomic DNA extraction (Blood and Cell Culture DNA Midi Kit, Qiagen) and analysis. This step served as a quality control measure for the GeCKO library transduction efficiency, which was assessed through deep sequencing using HiSeq X-Ten as described below.
GeCKO screens in THP-1 cell pool exposed to La2O3 NPs
For the GeCKO screens, a total of 5 × 107 GeCKO THP-1 cells were seeded into two 225 cm2 flasks. These cells were primed with 1 μg/mL PMA overnight, after which the media were replaced. Complete RPMI1640 media containing La2O3 NPs (8.0 μg/mL spheres or 4.0 μg/mL rods) were introduced, and a negative control was concurrently performed. The exposure period lasted for 72 h. Subsequent to the exposure, the remaining cells were subjected to genomic DNA extraction using the Blood and Cell Culture DNA Midi Kit (Qiagen). The concentrations of genomic DNA were quantified using a Qubit dsDNA assay (Thermo Scientific). Amplification of sgRNA inserts was achieved through nested PCR31, the first PCR amplification was carried out with 130 μg of genomic DNA per sample, equally separated into 13 reactions for 100 μL system, using Herculase II Fusion DNA Polymerase Kit (Agilent Technologies, 600679). The products from the 13 reactions were pooled together, and then 5 μL of the pooled products were used as templates in each reaction of the second PCR amplification. Barcodes of sequencing were added to the second PCR, and products of the second PCR were pooled, purified, and sequenced on a HiSeq X-Ten (Illumina). Primers (GeneScript) and condition of the nested PCR are given in Supplementary Table 7.
Construction of SMPD1− /− THP-1 cell line
The generation of SMPD1 knockout THP-1 cells followed established procedures31,41. Oligonucleotide pairs (GeneScript) targeting SMPD1 and non-targeting sgRNAs were employed, and their details are accessible in Supplementary Table 8. These oligonucleotide pairs were annealed and linked to the pGK1.1 linear plasmid (Genloci) using T4 DNA ligase (Thermo Scientific). The resultant recombinant vectors were transiently electro-transfected into THP-1 cells through an electro-porator (Celetrix). Post-transfection, THP-1 cells were exposed to puromycin at a concentration of 1.0 μg/mL for 48 h to eliminate cells with negative transfection outcomes. The remaining cells were subjected to limited dilution and were seeded into wells of ten 96-well plates to facilitate the selection of 20 individual single-cell clones. Confirmation of SMPD1 knockout in THP-1 cells was carried out through Western blotting and single clone sequencing methods.
Western blotting
For total protein extraction, THP-1 cells or cell pellets from bronchoalveolar lavage fluids (BALFs) of mice were lysed on ice for 20 min using RIPA buffer (Thermo Scientific) supplemented with protease inhibitor cocktail (Sigma) and phosphatase inhibitor cocktail (Sigma). A total of 30 μg of total protein from each sample was loaded into SDS-PAGE. Primary antibodies (ab9485 for GAPDH; PA5-95730 for mouse SMPD1; ab272729 for human SMPD1) were employed at a dilution ratio of 1:1000. The secondary antibody (ab6721, Abcam) was used at a dilution ratio of 1:10000. Incubations with primary and secondary antibodies were conducted overnight at 4 °C and for 1 h at room temperature, respectively. After each incubation, the blots were washed for five times. Subsequently, an advanced digital imaging system (Tanon-4600) was utilized to capture images following incubation with ECL (Thermo Scientific).
Detection of ceramide in THP-1 cells
A total of 3 × 105 THP-1 cells were seeded into each well of a 12-well plate and primed with 1 μg/mL PMA overnight. The cells were subsequently treated with La2O3 NPs (8.0 μg/mL spheres or 4.0 μg/mL rods) for a 72-h duration. Following treatment, the cells were washed with PBS and then fixed with 4% paraformaldehyde for 15 min. They were subsequently permeabilized using 0.3% Triton X-100 for 5 min on ice. Blocking of the cells was performed using 1% BSA in PBST for 30 min. For primary staining, the cells were incubated with ceramide primary antibody (ALX-804-196-T050, 1:10) at 4 °C overnight. A secondary antibody (ab150123, 1:1000) was employed for further staining at room temperature for 1 h in the dark. The coverslips were mounted using ProLong Gold Antifade Reagent with DAPI (Invitrogen), and fluorescence signals were analyzed using a Zeiss laser scanning confocal microscope (Zeiss LSM 880 with Airyscan).
Detection of phosphorus and lanthanum in transformed NPs
A total of 2 × 107 THP-1 cells were seeded and primed with 1 μg/mL PMA overnight. These cells were then treated with 50 μg/mL La2O3@ Fe3O4 for a period of 24 h. Then the cells were thoroughly washed with saline thrice. Subsequently, the cells were collected using cell scrapers. The lysosomes of the collected cells were isolated by a Lysosome Enrichment Kit (Thermo Scientific, 89839). The extracted lysosomes were lysed using ultrasonication to collect the transformed NPs under magnetic field. After sufficient washing by DI H2O, the collected nanoparticles were suspended in 0.5 mL H2O, and digested in 2 mL mixture of concentrated nitric acid hydrogen peroxide (30%) at 100 °C for 1 h. Quantitative analysis of the phosphorus (P) and lanthanum (La) concentrations within these particles was conducted using ICP-OES.
Time-dependent kinetics of NPs transformation in THP-1 cells and in PSF/ endosomal simulated fluid
THP-1 cells were exposed to 50 μg/mL La2O3@Fe3O4 nanoparticles for 0, 1, 2, 3, 6, 9, 12, 18, and 24 h, respectively. The nanoparticles in cellular lysosomes were collected by using a Lysosome Enrichment Kit (Thermo Scientific, 89839) under a magnetic field. The ratio of phosphorus to lanthanum elements (P/La) in the nanoparticles collected form lysosomes was quantified by using ICP-OES. Endosomal simulated fluid (ESF; 0.128 g/L CaCl2 ·2 H2O, 20.8 g/L citric acid, 0.059 g/L glycerol, 0.050 g/L MgCl2, 0.077 g/L sodium citrate dihydrate, 3.21 g/L NaCl, 0.179 g/L sodium phosphate heptahydrate, 0.085 g/L sodium lactate, 6.0 g/L NaOH, 0.086 g/L sodium pyruvate, 0.039 g/L Na2SO4, 0.09 g/L sodium tartrate dihydrate; pH 5.5) was produced24. 100 μg/mL La2O3-S nanoparticles were treated with ESF or PSF for 3, 6, and 12 h. The nanoparticles were collected and underwent ICP-OES for quantification of P/La, and TEM analysis for morphological changes.
Cellular uptake of nanoparticles in THP-1 cells
A total of 3 × 107 THP-1 cells were seeded and primed with 1 μg/mL PMA overnight. These cells were then treated with 50 μg/mL La2O3-S, La2O3-R, Fe2O3-S and Fe2O3-R for a period of 24 h, respectively. The culture supernatants were collected for quantification of IL-1β by using ELISA kit. The cells were collected, half of the cells were lysed for quantification of metal elements by using ICP-OES, the remaining cells were lysed for quantification of protein by using BCA kit (23252, Thermo Scientific). To assess cellular uptake, we employed the following formula:
Here, CmNP and CmCTR represent the concentrations of metal elements in the lysates of NP-treated and non-treated cells, respectively, while CpNP and CpCTR denote the concentrations of proteins in the lysates of NP-treated and non-treated cells, respectively. The IL-1β concentrations were divided by the cellular uptake of metal element to perform normalization.
Animal experiments
C57BL/6 male mice aged 8–9 weeks and weighing between 23.1–26.4 g were procured from Cyagen Biosciences. All animal-related procedures were conducted in strict adherence to the regulations set by the Animal Care Committee of the Laboratory Animals at Soochow University (No. 202108A0134). The experimental conditions ensured standard animal housing parameters, including a controlled room temperature of 23 ± 2 °C, 60% relative humidity, and a balanced light-dark cycle of 12 h each. The animals were provided autoclaved food and acidified water within filter-topped cages. Animal exposure to NPs was carried out through oropharyngeal aspiration7,39. In brief, the mice were divided into two groups (n = 36 per group): mice receiving desipramine or PBS treatment. Anesthesia was conducted by intraperitoneal injection of a combination of ketamine (100 mg/kg) and xylazine (10 mg/kg) in a total volume of 100 μL for each mouse. The anesthetized mice were positioned vertically. Aliquots of 50 μL desipramine solution (2 mg/kg) or PBS were instilled into the back of the tongue 24 h prior to NPs exposure. Then La2O3 NPs (4 mg/kg spheres or 2 mg/kg rods) or PBS solutions were instilled into the lung of treated mice by oropharyngeal aspiration. Post 48 h of NPs exposure, half of the mice from each group were euthanized and sacrificed to collect BALF in 1 mL PBS. Primary alveolar macrophages were isolated from the BALF for Western blotting analysis of SMPD1 protein and for observing the transformation of La2O3. The lung tissues were collected, and the lung sections were fixed by 2.5% glutaraldehyde for staining and TEM imaging. After a 14-d interval following NPs exposure, the remaining mice were sacrificed to obtain both BALF and lung tissues. The collected BALF was used for measuring IL-1β and TGF-β1 levels, while the lung tissues underwent Hematoxylin-Eosin (H&E) and Masson staining for further examinations. Another 12 male mice were treated with FITC-labeled La2O3 spheres (4 mg/kg, n = 6) or PBS solutions by oropharyngeal aspiration. Post 48 h of NPs exposure, the mice were euthanized and sacrificed to collect the lung tissues, the lung sections were fixed by paraformaldehyde, and were stained with F4/80 antibody (1:50) for macrophages, Prosurfactant Protein C (SPC) antibody (1:2000) for alveolar epithelial cells, and IL-1β antibody (1:50), DAPI was used to stain nuclei. The immunofluorescence confocal images were obtained by using confocal microscope (Nikon Eclipse CI with Nikon digital sight DS-FI2).
Statistical analysis
In the GeCKO screening, if a sgRNA knocked out a gene mediated the cytotoxicity of La2O3 NPs, the gene-knockout cells would be more resistant to La2O3 than wild-type cells. The sgRNAs for that gene would be overrepresented in surviving cells compared with their representations in untreated cells, and that gene was termed overrepresented gene. The overrepresented genes in GeCKO screening were identified by using RNAi Gene Enrichment Ranking (RIGER) at the options of “Second Best Rank” and “Signal to Noise”31,42. Gene Ontology pathway enrichment of human genes was performed by using g:Profiler software43 in accordance with the recommendation of g:Profiler, and “All known genes” was selected for statistical domain scope. A cutoff of that adjusted p value was less than 0.01 was applied. Bar plots represent means with SDs. The method for statistical significance analysis is indicated in each figure legend (GraphPad Prism 8.0). The difference was considered statistically significant when the p value was less than 0.05.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The source data for Figs. 2–5, and Supplementary Figs. 3, 4, 8–13, 18, 19 are provided as a Source Data file with this paper. DNA sequencing data (fastq) for the GeCKO screens are available via NCBI (BioSample accessionss SAMN42410414, SAMN42410415, SAMN42410416, SAMN42410417, SAMN42410418, SAMN42410419, SAMN42410420 and bioproject PRJNA1134364 [https://www.ncbi.nlm.nih.gov/bioproject/1134364]). All other data that support the findings of this study are available from the corresponding author upon request. Source data are provided with this paper.
References
Palika, A. et al. An antiviral trap made of protein nanofibrils and iron oxyhydroxide nanoparticles. Nat. Nanotechnol. 16, 918–925 (2021).
Cai, X. et al. Molecular mechanisms, characterization methods, and utilities of nanoparticle biotransformation in nanosafety assessments. Small 16, 1907663 (2020).
Gao, J. et al. Determining the cytotoxicity of rare earth element nanoparticles in macrophages and the involvement of membrane damage. Environ. Sci. Technol. 51, 13938–13948 (2017).
Borkowska, M. et al. Targeted crystallization of mixed-charge nanoparticles in lysosomes induces selective death of cancer cells. Nat. Nanotechnol. 15, 331–341 (2020).
Mosquera, J., Garcia, I. & Liz-Marzan, L. M. Cellular uptake of nanoparticles versus small molecules: a matter of size. Acc. Chem. Res. 51, 2305–2313 (2018).
Kim, S. E. et al. Ultrasmall nanoparticles induce ferroptosis in nutrient-deprived cancer cells and suppress tumour growth. Nat. Nanotechnol. 11, 977–985 (2016).
Li, R. et al. Surface interactions with compartmentalized cellular phosphates explain rare earth oxide nanoparticle hazard and provide opportunities for safer design. Acs Nano 8, 1771–1783 (2014).
Cao, M. et al. Molybdenum derived from nanomaterials incorporates into molybdenum enzymes and affects their activities in vivo. Nat. Nanotechnol. 16, 708–716 (2021).
Malysheva, A., Ivask, A., Doolette, C. L., Voelcker, N. H. & Lombi, E. Cellular binding, uptake and biotransformation of silver nanoparticles in human T lymphocytes. Nat. Nanotechnol. 16, 926–932 (2021).
Dordevic, L., Arcudi, F., Cacioppo, M. & Prato, M. A multifunctional chemical toolbox to engineer carbon dots for biomedical and energy applications. Nat. Nanotechnol. 17, 112–130 (2022).
Cai, R. et al. Dynamic intracellular exchange of nanomaterials’ protein corona perturbs proteostasis and remodels cell metabolism. Proc. Natl. Acad. Sci. USA 119, e2200363119 (2022).
Nel, A. E. et al. Understanding biophysicochemical interactions at the nano-bio interface. Nat. Mater. 8, 543–557 (2009).
Leong, H. S. et al. On the issue of transparency and reproducibility in nanomedicine. Nat. Nanotechnol. 14, 629–635 (2019).
Oh, E. et al. Meta-analysis of cellular toxicity for cadmium-containing quantum dots. Nat. Nanotechnol. 11, 479–486 (2016).
Dilliard, S. A., Cheng, Q. & Siegwart, D. J. On the mechanism of tissue-specific mrna delivery by selective organ targeting nanoparticles. Proc. Natl. Acad. Sci. USA 118, e2109256118 (2021).
Wang, Z. et al. Immunogenic camptothesome nanovesicles comprising sphingomyelin-derived camptothecin bilayers for safe and synergistic cancer immunochemotherapy. Nat. Nanotechnol. 16, 1130–1140 (2021).
Liu, M. D. et al. A transistor-like Ph-sensitive nanodetergent for selective cancer therapy. Nat. Nanotechnol. 17, 541–551 (2022).
Spurgeon, D. J., Lahive, E. & Schultz, C. L. Nanomaterial transformations in the environment: effects of changing exposure forms on bioaccumulation and toxicity. Small 16, 202000618 (2020).
Curtis, B. J. et al. Cross-species transcriptomic signatures identify mechanisms related to species sensitivity and common responses to nanomaterials. Nat. Nanotechnol. 17, 661–669 (2022).
Zoncu, R. et al. Mtorc1 senses lysosomal amino acids through an inside-out mechanism that requires the vacuolar H+-Atpase. Science 334, 678–683 (2011).
Van Veen, S. et al. Atp13a2 deficiency disrupts lysosomal polyamine export. Nature 578, 419–424 (2020).
Chen, Y. et al. The origin of exceptionally large ductility in molybdenum alloys dispersed with irregular-shaped La2o3 nanoparticles. Nat. Commun. 15, 4105 (2024).
Qi, Y.-T. et al. Homeostasis inside single activated phagolysosomes: quantitative and selective measurements of submillisecond dynamics of reactive oxygen and nitrogen species production with a nanoelectrochemical sensor. J. Am. Chem. Soc. 144, 9723–9733 (2022).
Rabel, M. et al. Simulation of the long-term fate of superparamagnetic iron oxide-based nanoparticles using simulated biological fluids. Nanomedicine 14, 1681–1706 (2019).
Jiang, J. et al. Intracellular dehydrogenation catalysis leads to reductive stress and immunosuppression. Nat. Nanotechnol. https://doi.org/10.1038/s41565-025-01870-y (2025).
Ma, Y. et al. Origin of the different phytotoxicity and biotransformation of cerium and lanthanum oxide nanoparticles in cucumber. Nanotoxicology 9, 262–270 (2015).
Ma, Y. et al. Phytotoxicity and biotransformation of La2o3 nanoparticles in a terrestrial plant cucumber (Cucumis Sativus). Nanotoxicology 5, 743–753 (2011).
Zheng, H. et al. Biotransformation of rare earth oxide nanoparticles eliciting microbiota imbalance. Part. Fibre Toxicol. 18, 17 (2021).
Li, R. et al. Interference in autophagosome fusion by rare earth nanoparticles disrupts autophagic flux and regulation of an interleukin-1 beta producing inflammasome. Acs Nano 8, 10280–10292 (2014).
Fadeel, B. et al. Advanced tools for the safety assessment of nanomaterials. Nat. Nanotechnol. 13, 537–543 (2018).
Tian, M. et al. Toxicological mechanism of individual susceptibility to formaldehyde-induced respiratory effects. Environ. Sci. Technol. 56, 6511–6524 (2022).
Becker, K. A. et al. Acid sphingomyelinase inhibitors normalize pulmonary ceramide and inflammation in cystic fibrosis. Am. J. Respir. Cell Mol. Biol. 42, 716–724 (2010).
Teichgraber, V. et al. Ceramide accumulation mediates inflammation, cell death and infection susceptibility in cystic fibrosis. Nat. Med. 14, 382–391 (2008).
Qi, X. M. et al. Pneumoconiosis: current status and future prospects. Chin. Med. J. 134, 898–907 (2021).
Lv, J. et al. Dissolution and microstructural transformation of Zno nanoparticles under the influence of phosphate. Environ. Sci. Technol. 46, 7215–7221 (2012).
David, C. A., Galceran, J., Quattrini, F., Puy, J. & Rey-Castro, C. Dissolution and phosphate-induced transformation of Zno nanoparticles in synthetic saliva probed by agnes without previous solid-liquid separation. Comparison with Uf-Icp-Ms. Environ. Sci. Technol. 53, 3823–3831 (2019).
Li, Z. et al. Direct hydrothermal synthesis of single-crystalline hematite nanorods assisted by 1,2-Propanediamine. Nanotechnology 20, 245603 (2009).
Chen, L. et al. Continuous shape- and spectroscopy-tuning of hematite nanocrystals. Inorg. Chem. 49, 8411–8420 (2010).
Cai, X. et al. Multi-hierarchical profiling the structure-activity relationships of engineered nanomaterials at nano-bio interfaces. Nat. Commun. 9, 4416 (2018).
Joung, J. et al. Genome-scale crispr-Cas9 knockout and transcriptional activation screening. Nat. Protoc. 12, 828–863 (2017).
Tian, M. et al. Crispr screen identified that Ugt1a9 Was required for bisphenols-induced mitochondria dyshomeostasis. Environ. Res. 205, 112427 (2022).
Zhang, Y. et al. Functional genomic screen of human stem cell differentiation reveals pathways involved in neurodevelopment and neurodegeneration. Proc. Natl. Acad. Sci. USA 110, 12361–12366 (2013).
Raudvere, U. et al. G:Profiler: a web server for functional enrichment analysis and conversions of gene lists (2019 Update). Nucleic Acids Res. 47, W191–W198 (2019).
Acknowledgements
This work is supported by the National Natural Science Foundation of China (Grant No. 42307365), and National Key Research and Development Program of China (2022YFC3202101), the ‘100 Talent Program’ from Government of Jiangsu Province, China (SBX2019010069). X.Z. was supported by the Talents Research Fund of Yunnan University (No. CZ22623101, CY22623101). M.T. was supported by the Fundamental Research Funds for the Central Universities (No. 3132023160). R.L. was supported by the European Union under the Refresh-Research Excellence for Region Sustainability and High-tech Industries Project (CZ.10.03.01/00/22_003/0000048) via the Operational Programme Just Transition.
Author information
Authors and Affiliations
Contributions
M.T., X.Z. and R.L. conceived the study and designed the experiments. Most of the experiments were conducted by M.T. and D.W., and animal experiments were conducted by M.T. and X.G. This paper was written by M.T., X.Z. and R.L.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
All animal-related procedures were conducted in strict adherence to the regulations set by the Animal Care Committee of the Laboratory Animals at Soochow University (No. 202108A0134).
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Tian, M., Wu, D., Gou, X. et al. Genetic modulation of rare earth nanoparticle biotransformation shapes biological outcomes. Nat Commun 16, 3429 (2025). https://doi.org/10.1038/s41467-025-58520-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-58520-z
This article is cited by
-
Microbial aggregates and functional materials for mitigating soil nitrogen loss: a review
Reviews in Environmental Science and Bio/Technology (2026)