Abstract
How cells change shape is crucial for the development of tissues, organs and embryos. However, studying these shape changes in detail is challenging. Here we present a comprehensive real-time cellular map that covers over 95% of the cells formed during Caenorhabditis elegans embryogenesis, featuring nearly 400,000 3D cell regions. This map includes information on each cell’s identity, lineage, fate, shape, volume, surface area, contact area, and gene expression profiles, all accessible through our user-friendly software and website. Our map allows for detailed analysis of key developmental processes, including dorsal intercalation, intestinal formation, and muscle assembly. We show how Notch and Wnt signaling pathways, along with mechanical forces from cell interactions, regulate cell fate decisions and size asymmetries. Our findings suggest that repeated Notch signaling drives size disparities in the large excretory cell, which functions like a kidney. This work sets the stage for in-depth studies of the mechanisms controlling cell fate differentiation and morphogenesis.
Similar content being viewed by others
Introduction
The systematic tracking of key cellular behaviors in real time, including the dynamics of cell shape, volume, surface, and neighborhood, is crucial for understanding various developmental processes but remains technically challenging, especially during animal embryogenesis. This is because, in response to cell autonomous or non-autonomous regulation, embryonic cells undergo rapid division and migration along with cell fate specification and differentiation to ensure proper tissue formation and organogenesis1. On top of this, the precise delineation of cell lineage provides further insurance on each cell’s identity and developmental trajectory, enabling the dissection of the developmental process with precision and spatiotemporal and cellular resolution. However, achieving such a resolution is impractical in higher animals, especially in mammals, due to the excessive number of cells and difficulties in culturing such embryos in vitro for live cell imaging2,3,4,5. For example, cell shape changes dramatically during compaction of earlier embryogenesis6. Single-cell analyses in mice or humans usually focus on the very early stage of embryogenesis with inferred cell lineage that comes with inherent uncertainties7,8. In addition, all these single-cell analyses deduce a cell state mainly dictated by molecular signatures, but ignore important cellular features, i.e., cellular morphology, including the dynamics of cell shape, volume, surface, and neighborhood. Integration of quantitative data on these parameters with single-cell omics data is expected to significantly improve the power to draw more biologically relevant conclusions in terms of the progression of cell fate specification and differentiation.
Cellular morphology, such as cell shape, volume, surface area, and contact between neighboring cells, plays a fundamental role in morphogenesis and tissue formation in various species9,10,11,12,13,14. Moreover, the spatial distribution and fate specification of cells in animals rely on the mechanical and signaling interactions between neighboring cells. For example, in Caenorhabditis elegans (a hermaphrodite, which reproduces mostly through selfing although rare males are present and can mate with hermaphrodite; a well-established model animal cultured at laboratory over four decades), a change in the division geometry in the four-cell-stage embryo disrupts Notch signaling from the P2 to the AB cell, leading to the failure of fate specification15. In addition, such fate induction through Notch signaling is time-dependent, and thus, may have an opposite effect at different developmental times16. Most of these signaling interactions have been mapped either by genetic screening or by the expression of ligands and receptors of signaling interactions15,17,18,19. The identification of such interactions is becoming increasingly challenging over development due to the difficulties in resolving cell identity and the lack of expression profiles of ligands and receptors with cellular resolution, especially during late embryogenesis. In addition to cell shape, cell size is also vital for the proper development of tissues and organs. For example, in C. elegans, heterogeneous cell size is correlated with heterogeneous cell cycle length during embryogenesis, which helps to coordinate the migrations, positions, and contacts of all cells20,21,22,23,24,25,26. Unfortunately, information on cell shape, size, and contact is commonly neglected along with cell cycle length during studies of gene regulation of embryogenesis. Equipped with light-sheet microscopy and a newly developed algorithm, systematic mapping of cell shape and contact was performed in early-to-mid-stage ascidian embryos, thereby revealing the invariance of ascidian embryogenesis and that the contact areas between signaling and responding cells are indicative of the embryonic inductions required for fate specification27. However, a complete picture of the cellular morphology of ascidian embryos has yet to be obtained due to technical difficulties in the segmentation of late-stage embryos.
Control of cell size, including cell volume and surface area, is critical for proper tissue formation and organogenesis. Attempts have been made to systematically map the cellular morphology of a few species27,28,29. Numerous studies demonstrated the role of cell size accuracy in the control of development in both biological and physical contexts12,20,22,23. However, a comprehensive map of cellular morphology with resolved cell lineage and fate as well as other quantitative morphological features (e.g., cell volume, cell surface area, and cell-cell contact area) throughout embryogenesis has not been constructed in any species. The organism of choice for systematic mapping of cellular morphology is C. elegans, due to its invariant development and transparent body30, the well-established methods for automated cell lineage tracing31,32,33, and the abundance of genetic and molecular tools34,35. On top of this, the availability of lineal expression profiles of numerous genes (especially that of transcription factors36,37) enables the study of molecular and cellular controls of embryogenesis with precision. As such, numerous attempts have been made to reconstruct the cellular morphology of a developing C. elegans embryo with or without cell identity29,38,39. However, these methods can only produce cellular morphology during early embryogenesis in C. elegans, due to the high density and small size of the cells during late embryogenesis. For example, a C. elegans cell is roughly 100 times smaller than that of an ascidian when the number of cells is comparable between the embryos of the two species40. We have previously attempted to reconstruct the cellular morphology of C. elegans embryos using nucleus-based modeling41 or nucleus-independent deep learning based on cell membrane fluorescence (CShaper)12,42. However, both the modeling-based method and CShaper were only able to produce cellular morphology for the first half of embryogenesis (i.e., up to approximately the 350-cell stage), due to the difficulties in modeling or segmenting membranes of the highly crowded cells at a stage beyond the 350-cell stage. In addition, the expression intensity of the transgenic membrane markers becomes dimmer than in the early stage, making it impractical to reconstruct the cellular morphology map beyond the 350-cell stage. However, most embryonic cells at this stage have not completed their final round of embryonic division, and have not yet differentiated to their terminal fate, which prevents the study of the gene regulations that control the fate asymmetry for most embryonic cells.
In this study, we establish a platform that allows qualitative and quantitative analysis of three-dimensional (3D) cell shape, volume, surface area, and contact area as well as lineal expression of various genes with defined cell lineage in C. elegans embryos up to the 550-cell stage, when most embryonic cells complete their final round of division with terminal fate. We use the platform to generate a comprehensive cellular morphological map up to the comma stage of embryos at ~1.5-minute intervals, which consists of cell shape, size, and contact between neighboring cells for all cells with resolved cell lineage. We then demonstrate the power of the platform by analyzing the effect of Notch signaling on not only breaking the symmetry of cell fate, but also cell size, which shows that Notch signaling interaction between neighboring cells not only regulates fate asymmetry, but also controls the size asymmetry of the same cell pair in a division orientation-dependent manner, i.e., such interaction invariably enlarges the anterior daughter cell at the cost of the posterior daughter cell. To further demonstrate the power of the platform, we integrate lineal expression of Notch ligands and receptors with the morphological map. We show that in addition to an existing Notch interaction targeting the cell “ABplpapp” at around the 80-cell stage17,43,44, four more rounds of consecutive Notch interactions target itself, its daughter, and its granddaughter by different ligand-expressing cells, which drives asymmetric divisions in terms of both cell fate and size, leading to its final differentiation into the C. elegans excretory cell, an equivalent of the kidney, which has the largest size in the adult45,46,47. We finally make our data accessible both locally as standalone software or online through interactive query and vivid visualization.
Results
A method for automated segmentation of cell membranes labeled by fluorescent protein up to the 550-cell stage
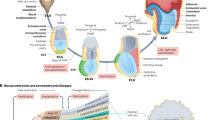
To enable systematic reconstruction of the cellular morphologies up to the 550-cell stage of C. elegans embryogenesis, we developed a fully automated pipeline, CMap, which shows a superior segmentation accuracy (Fig. 1A; Figs. S1–S3; Supplementary Data 1) and a reasonably high computational speed (Supplementary Data 2). The improved performance is partially contributed by a C. elegans transgenic strain that was built with biolistic bombardment to facilitate membrane segmentation. The strain showed a higher intensity of membrane fluorescence than the one we previously used12, especially at the late stage (350- to 550-cell stages) of the embryo (Fig. 1B; Figs. S4, S5). We used the pipeline to reconstruct cellular morphologies for nearly all embryonic cells up to the comma stage at ~1.5-min intervals for a total of eight wild-type embryos (“WT_Sample1” to “WT_Sample8”, all derived from the same experimental condition; Supplementary Data 3; “Methods”), along with the complete cell identities and lineages produced by StarryNite and AceTree30,32,33 (Supplementary Movie 1).
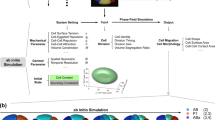
A The data processing pipeline of CMap. Time-lapse 3D (4D) images of GFP-labeled cell nuclei and mCherry-labeled cell membranes are used for cell lineage tracing and morphology segmentation respectively, with output of cell identity (with information on cell lineage and cell fate), cell shape, volume, surface area, and contact area over embryogenesis. The EDT-DMFNet segmentation model of cell membrane morphology is explained in detail in Fig. S6. B Top: 3D projections of a dually labeled embryo at representative developmental stages indicated above; middle: outputs of cell nucleus tracing; bottom: outputs of cell membrane segmentation. Nuclei and membranes are differentially colored based on their lineal origins as indicated.
CMap not only outputs cell shape, but also computes cell volume, surface area, and contact area (Fig. 1A). Nuclei were ubiquitously labeled with the green fluorescent protein (GFP) to enable cell lineage tracing32. The lineage tracing produced cell nucleus positions which were used as alternative seeds to facilitate CMap in reconstructing the morphology of individual cells, especially those at late-stage embryos where cells are densely packed with small size. This reconstruction was achieved by using an advanced adaptive deep convolutional neural network, denoted the Euclidean distance transform dilated multifiber network (EDT-DMFNet), for cell membrane recognition (Fig. S6). As a result, the EDT-DMFNet segments the fluorescently labeled cell membranes up to the 550-cell stage of the embryo with high quality. At this stage, cells are approximately half the size they are at the 350-cell stage. (Fig. 1B). The segmentation results up to the 550-cell stage serve as an input for downstream analyses whereby cell lineage- and fate-wise 3D cell objects and morphological features of individual cells are automatically extracted for an entire embryo (Supplementary Movies S2 and S3). On average, it takes CMap about 3 h to implement automatic cell segmentation for a C. elegans embryo from the four- to 550-cell stages (Supplementary Data 2), allowing high-throughput and efficient processing of wild-type embryos as well as embryos perturbed with genetic mutation, RNA interference (RNAi), mechanical compression, eggshell removal, or laser ablation25,41,48,49,50.
A quantitative morphological map with resolved lineage for nearly all cells produced during C. elegans embryogenesis
Equipped with the CMap, we established a comprehensive morphological map with resolved cell identity, cell lineage, and cell fate up to the 550-cell stage of C. elegans embryogenesis with both statistical reliability and data completeness. We took the following measures to ensure the comprehensiveness, reliability, and accuracy of our data. First, we collected time-lapse 3D images for eight wild-type C. elegans embryos with GFP-labeled cell nuclei and mCherry-labeled cell membranes from no later than the four-cell stage to no earlier than the 550-cell stage at ~1.5-minute intervals (Supplementary Data 3). Next, to minimize the deformation of cell shape, unlike the embryo mounting techniques we used previously, in which the embryo was immobilized by applying some pressure from the cover slide, leading to a severely deformed embryo shape12 (Fig. S7), we applied no pressure (without external mechanical compression) on the mounted embryos (“Methods”). Third, we performed automated cell lineage analysis for all embryos with manual curation up to the time point when all 48 progeny cells of the somatic founder cell “C” were present for at least five time points (more than 7 min); this time point marks the completion of hypodermal cell production30,51 (Fig. 2A, B).
A The embryonic cell lineage tree averaged over the eight C. elegans wild-type embryos up to the 550-cell stage. Cell fates are differentially color-coded as indicated. The excretory cell and the sole body-wall muscle cell derived from the AB lineage are indicated with black and gray arrowhead respectively. The cells with consistent failures in segmentation in all embryo samples are indicated with black dots. Developmental time is shown on the left, with the last time point of the four-cell stage set as the time zero. B Cell counts across developmental stages for the eight embryos, with the average cell numbers represented by dots (black for surviving cells and red for apoptotic ones) and their standard deviations by vertical lines. The duration of significant developmental landmarks is indicated by differential shading51. C, D Comparison of average cell volume (C) and cell surface area (D) with individual measurements from the eight C. elegans embryos. Data points represent individual cell comparisons, with the average across embryos on the horizontal axis and individual embryo measurements on the vertical axis. Cells present before and after the ~350-cell stage are color-coded in blue and yellow, respectively. Insets show the distribution of variation coefficients (defined as the ratio of standard deviation to average) for these comparisons, based on 322 and 267 unique cells before and after the ~350-cell stage respectively. Source data are provided as a Source Data file.
In the morphological map, over 95% of all cells in C. elegans embryogenesis were successfully segmented in at least one embryo sample52 (Supplementary Data 4). More specifically, we segmented the membranes of a total of 1292 unique cells in the eight embryos, including 1188 non-apoptotic cells and 104 apoptotic cells, with the latter representing 92.04% of all embryonic apoptotic cells. Moreover, 1190 unique cells were reproducibly segmented and recorded in all eight embryos, with 589 having a complete lifespan (cell cycle length) and 79 being apoptotic. This was almost twice the number of cells that were previously segmented by CShaper12 (Supplementary Data 5), enabling morphological characterization of embryonic cells from the four- to 550-cell stages. This is significant because at this point most embryonic cells complete their final round of division with terminal fate, allowing the study of genetic and molecular networks regulating development at cellular resolution with defined cell lineage and fate for nearly every cell throughout embryogenesis. Specifically, for a 3D cell region (defined as a unique compartment formed by cell membrane) or a so-called “data point” (c, j, T) denoted by cell identity c, embryo sample j, and time point T, a total of 395,741 cell regions (covering 99.99% out of all the existing cell nuclei recorded) were effectively segmented by CMap, with 5,905 containing two cell nuclei inside, which were usually observed at the time points immediately before cytokinesis (Supplementary Data 4). Subsequently, 10.89% of the cell regions are automatically filtered out due to their abnormal volume or shape (Figs. S8–S10; “Methods”). 69.10% of these regions are non-apoptotic cells that are presumed to have a nearly identical volume and space occupation over time. Thus, these were removed from our reported dataset. The remaining 13,320 regions are the apoptotic ones that exhibited a sudden decrease in volume over time, and therefore we retained them in our final dataset along with a subset manually checked and corrected53 (Figs. S11, S12). As a result, a comprehensive cellular morphological map of C. elegans embryo was established with a missing rate of less than 8% for total time-lapse 3D data points and with coverage for over 95% of all cells (recorded with at least one data point) in C. elegans embryogenesis (Supplementary Data 6 to 8), which provides an invaluable resource for studying cellular behaviors for nearly all cells with defined cell lineage and identity throughout C. elegans embryogenesis.
The overall cellular morphology, produced by automated cell segmentation followed by filtering with rigorous quality control, is consistent with that published by various independent labs. It also demonstrates a high accuracy based on manually annotated ground truth. Specifically, compared with cell volumes derived from five 7- to 8-celled embryos and 49 2- to 96-celled embryos segmented respectively in two previous studies39,54, the volumes of non-identical cells produced by CMap segmentation exhibit an average relative variation smaller than 14% (Fig. S13A, B). When compared with cell volumes using two embryos manually annotated at around the 100-, 200-, 300-, 400-, 500-, and 550-cell stages, the cell volumes produced by CMap segmentation exhibit an average relative variation smaller than 4% (Fig. S13C, D).
In summary, CMap produces the shape of nearly all cells from the four- to 550-cell stages with resolved cell identity, cell lineage, and cell fate, and with reproducibility and statistical support provided by the eight wild-type embryos (Fig. S14). It outputs three other quantitative morphological features: cell volume (\(V\)), cell surface area (\({A}_{{{{\rm{S}}}}}\)), and cell-cell contact area (\({A}_{{{{\rm{C}}}}}\)) (Fig. 1A). For each C. elegans wild-type embryo in our dataset, in addition to the 322 unique cells prior to the ~350-cell stage with a complete lifespan that were correctly segmented as we did before12, we segmented another 267 unique cells beyond the 350-cell stage with a complete lifespan; such requirement on a complete lifespan allows the study on the cellular morphological dynamics at the time scale within the cell cycle length (Fig. S14), for instance, the study on the consecutive rounding and elongation realized by cytoskeleton remodeling during cell division54,55,56. Statistical analyses of cell volume and surface area demonstrated that both cellular parameters are tightly controlled throughout C. elegans embryogenesis, similar to cell lineage and fate patterns, cell cycle lengths, and cell positions discovered before (Fig. 2C, D)57,58.
Characterization of morphogenesis and organogenesis with cellular resolution using the morphological map
Our morphological map with defined cell identity, lineage, and fate allowed us to reconstruct cell deformations and migrations for a specific tissue or organ as well as their lineal origin over embryogenesis (Fig. 3A; Fig. S15; Supplementary Movies S2 to S4). We used the following terms to describe cell fate during embryogenesis. Information on cell fate for labeling a specific tissue or organ (marked with “Neuron”, “Pharynx”, “Skin”, “Muscle”, “Intestine”, “Germline”, and “Others”) and apoptosis (marked with “Death”) is extracted from previous description37. The apoptotic fate was also manually checked30. If the two daughters of a cell have different cell fates, it will be marked with “Unspecified”; if the two daughters of a cell have same cell fate, it will be marked with its daughters’ cell fate; if the fate of a cell has been determined, its two daughters will be marked with the same fate as its mother’s one (Supplementary Data 4). For example, the EMS cell (the second somatic founder cell produced during the third cell division of C. elegans embryogenesis) is marked with “Unspecified” because its anterior daughter, MS (marked with “Unspecified”), derives pharynx and muscle among others, whereas its posterior daughter, E (marked with “Intestine”), derives only the intestine (Fig. 2A).
A Visualization of cell shapes within whole embryo or within specific tissues/organs from various perspectives. B–D Depiction of dynamic cell shape changes in different tissues during late embryogenesis: skin cells (hypodermis) during dorsal intercalation (B), intestine cells during intestinal twisting and elongation (C), and body-wall muscle cells during the ingression of the AB-derived body-wall muscle cell (ABprpppppaa), which is indicated by an arrow (D). Developmental time and stage are shown on the left, with the last time point of the four-cell stage set as the time zero. E–G Quantification of cell irregularity (η) for the processes shown in (B–D). Panel (E) presents the average (solid dot) and standard deviation (solid line) of the irregularity for 9 skin cells, as indicated by arrowheads in (B), during the developmental timeline (t) in an exemplary embryo with the last time point of the four-cell stage set as the time zero. Panel (F) presents similar data for all cells during the developmental timeline (t) in an exemplary embryo. Panel (G) presents the average (solid dot) and standard deviation (solid line) of cell shape irregularity (η) for the cell ABprpppppaa during the developmental timeline (t) in eight embryos. In (E, G), the maximums and minimums are denoted by green and pink triangles respectively, and the correlation coefficients for the monotonic \(\eta -t\) curves are shown at the top. Source data are provided as a Source Data file.
With the complete cell fate information over C. elegans embryogenesis, the morphology of cells forming various tissues and organs can be visualized vividly in 3D, exhibiting an apparent bilateral symmetry (Fig. 3A). The pharynx and intestine form a tube along the anterior-posterior axis of the embryo, where the intestinal tube bends precisely over its contact with the germline progenitors, i.e., the Z2 and Z3 cells, which may provide physical protection or produce a unique position for the engulfment and degradation of large lobes extended by the Z2 and Z3 cells during embryogenesis59 (Fig. 3A; Supplementary Movie S5). The cell-resolved morphological map enables the reconstruction and vivid visualization of both early and late critical morphogenetic events in C. elegans embryogenesis, including gastrulation60 (Supplementary Movie 6), dorsal intercalation51,61 (Fig. 3B), intestinal formation62,63 (Fig. 3C) and body-wall muscle assembly64,65 (Fig. 3D).
The dimensionless surface-to-volume ratio \(\left(\eta=\frac {\root {2} \of {{A}_{{{{\rm{S}}}}}}}{\root {3}\; \of{V}}\right)\) was used to evaluate the irregularity of cell shape, which revealed the roles played by cell migration and lineal origin in determining cell irregularity12,56. For example, during dorsal intercalation, two rows of skin cells, i.e., hypodermis, originally located on each dorsal side of the embryo, move toward the dorsal midline and eventually form a single row, with cell irregularity increasing gradually during squeezing and narrowing of the relevant cells (Fig. 3B, E; Supplementary Movie 7). Intestine cells also show an apparent low–high–low pattern in cell irregularity over the course of morphogenesis in the first transition punctuated with the intercalation (Fig. 3C, F; Supplementary Movie 8) and insertion of the E cells (the Ealpa, Earpa, Ealpp, and Earpp cells), and the latter transition corresponds to elongation and twisting (Fig. S16; Supplementary Movie 8). Body-wall muscle cells are derived from various lineal origins (i.e., the AB, MS, C, and D sublineages) and assembled into two bilaterally symmetric stripes (Fig. 3D; Supplementary Movie 9). This raises an interesting question, i.e., how cells with different lineal origins are able to “locate” each other and assemble into two coherent tubular stripes, particularly in the case of ABprpppppaa, which is the sole body-wall muscle cell derived from the AB sublineage. Intriguingly, ABprpppppaa is initially located outside the stripe formed by the remaining muscle cells but is eventually inserted into the stripe reproducibly in different embryos, suggesting that its insertion is genetically programmed (Supplementary Movie 10). Three reproducible peak–valley pairs are present in the ABprpppppaa cell irregularity curve as the cell ingresses from the periphery of the body-wall muscle stripe, during which its shape changes first from spherical to oblate and then to spherical (Fig. 3G; Fig. S17).
All three critical morphogenetic events above demonstrate an increase in cell irregularity, which is a common hallmark when cells drastically migrate over short distances or intercalate with their neighbors for positional and contact rearrangement (Fig. 3B–G; Figs. S16 and S17). Taken together, our 550-cell-stage morphological map allows qualitative and quantitative analyses of cell deformations and migrations during morphogenesis, permitting characterization of embryogenesis with exceptional spatiotemporal resolution in C. elegans embryos.
Characterization of cell lineage and fate differentiation with cellular resolution using the morphological map
Apart from the interplay between cell migration and cell irregularity (Fig. 3B–G; Figs. S16 and S17; Supplementary Movies S7 to S10), we further ask whether cell irregularity can characterize morphogenesis over embryogenesis. To this end, we chose the MS (mesoderm) and E (endoderm) sublineages derived from EMS, one of the first four founder cells (Fig. 2A). This choice was based on their well-characterized differential cell fates and roles in gastrulation, cell lineage patterns (cell cycle lengths), and fate induction by Wnt signaling from P2 to EMS30,62,66. The lineage differentiation can be visualized by binary expression of fate markers (mesodermal marker ceh-51 and endodermal marker elt-7) (Fig. S18), which is also characterized by asynchronous cell cycle lengths, with a G2 gap phase introduced in the E sublineage during the early maternal-zygotic transition and gastrulation60,67,68.
From the morphological perspective, the irregularity of both MS and E cells oscillates with their cell cycle, peaks at mid-lifespan, and decreases near cell division (Fig. S19, A and B), showing a modest positive correlation (Pearson correlation coefficient = 0.4353) with its variability among the eight wild-type embryos across all cells and time points (Fig. S19C). This pattern aligns with previous experimental and computational studies on cell stiffness and rounding, driven by the periodic transport of actin filaments between the cell cortex and cytosol55,56. In terms of differentiation, from the first cell cycle onward, the MS cell exhibits a significantly higher irregularity compared to the E cell (calculated at mid-lifespan and upon division respectively; one-sided Wilcoxon signed-rank test, \(p\le 0.05\)), consistent with the MS cell’s pivotal role in facilitating embryo rotation around the anterior-posterior axis through its elaborate lamellipodial protrusions, essential for left-right repatterning69,70 (Fig. S19A). The cell irregularities of the second cell cycle of MS and E cells begin to diverge with the introduction of the gap phase in the E2 cells, enabling specific gene expression for maternal-zygotic transition and gastrulation68. During the second and third cell cycles, MS cells exhibit smooth, large-amplitude oscillations in cell irregularity; meanwhile, E2 cells, undergoing approximately one cell cycle, show an initial increase in irregularity after division, and then the cell irregularity exhibits fast, small-amplitude fluctuations around a moderate value for most of their lifespans (Fig. S19A). This fast, small-amplitude fluctuation in cell irregularity reflects the unique cellular morphological characteristics during the internalization of E2 (initial gastrulation), distinguishing it from the normal cell shape dynamics observed throughout the lifespans of all MS and E cells (Fig. S19A). Such positional rearrangement will make all E progeny settle down and divide inside the embryo to produce the intestine (Fig. S19B). Starting with the next cell cycle of the E sublineage, E cells regain smooth oscillations in cell irregularity similar to those of MS cells; however, the extended duration of the E cell cycle results in a continuously longer oscillation period compared to MS cells (Fig. S19A).
Intriguingly, cell shape dynamics can be altered by gene mutation or perturbation. In two wee-1.1 mutant embryos (strain: RB66971; “MT_wee-1.1_1” and “MT_wee-1.1_2”, all derived from the same experimental condition; Supplementary Data 3; “Methods”), the E2 cell cycle lengths are known to be shortened (from 38.07 ± 2.01 min to 26.51 ± 0.67 min, defined by the cell nuclei separation72) but still sufficient for successful gastrulation and embryogenesis. As a result, the second duration of E cell irregularity oscillation also becomes shortened (from 45.76 min to 27.17 min, defined by the valleys of cell irregularity curve) (Fig. S20A). In two embryos (“RNAi_pop-1_1” and “RNAi_pop-1_2”, all derived from the same experimental condition; Supplementary Data 3; “Methods”) with RNAi against pop-1 (the terminal effector of the Wnt signaling pathway), the signaling induction on EMS was blocked so that the MS was converted into an E-like fate. As a result, the oscillation period and amplitude of MS and E sublineages become more similar (Fig. S20B), consistent with the fate transformation from MS to E. The results demonstrate that cell morphological dynamics indeed are correlated with their fate. Therefore, our publicly accessible morphological map can be utilized to address their questions of interest during development by those in the community.
Roles of Notch signaling in promoting the size asymmetry between anterior and posterior daughters of its target cell
Given the important roles of Notch signaling in inducing asymmetry in cell fates in C. elegans embryo73, we examined whether the Notch signaling that induced the fate asymmetry also produced asymmetry in the cell size or cell cycle length. Intriguingly, we observed that there is an overall negative correlation between cell size asymmetry and cell cycle length asymmetry25 (also referred to as division asynchrony) (Fig. S21; Supplementary Data 9), which is consistent with the previous finding that cell-cycle-related factors are positively correlated with cell-volume partitioning during cytokinesis22,74. Given the dependence between these two properties, we will henceforth exclusively investigate the effect of Notch signaling on cell size asymmetry and its coupling with cell fate asymmetry.
Here, we focused on the five rounds of Notch signaling events reported previously but utilizing actual cell size41. These signaling events target the ABp, ABalp, ABara, ABplaaa, and ABplpppp cells but not their sisters (ABa, ABala, ABarp, ABplaap, and ABplpppa) (Fig. 4A), except ABplpapp and its sister, ABplpapa. Previous studies proposed that both ABplpapa and ABplpapp receive Notch signaling, leading to differential gene expressions and fate assignments in their progeny75. Our data reveal that the total contact area between ABplpapp and the signaling cells (MSap sublineage) is approximately twice as large as that between ABplpapa and the signaling cells (Fig. S22), which suggests a substantially stronger impact of Notch signaling on ABplpapp. To analyze the six Notch target cells, where their sisters receive no effective signaling or substantially weaker signaling, we defined the cell size asymmetry as the ratio of net volume difference between the anterior and posterior daughter to their total volume, i.e., \(\frac{{V}_{{{{\rm{D}}}}1}\,-\,{V}_{{{{\rm{D}}}}2}}{{V}_{{{{\rm{D}}}}1}\,+\,{V}_{{{{\rm{D}}}}2}}\), where \({{{\rm{D}}}}1\) denotes the anterior daughter cell, \({{{\rm{D}}}}2\) the posterior daughter cell, and \(V\) the cell volume.
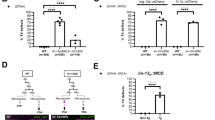
A Reconstructed 3D morphologies of contacting cell pairs engaged in the Notch signaling events during C. elegans embryogenesis identified previously. The relevant cell identities were described elsewhere41. Cells expressing Notch ligands are highlighted in green, while those expressing receptors are in red. B Plots showing the volume asymmetry ratio, calculated as the net volume difference over the combined volume of anterior and posterior daughter cells from six sister-cell pairs in eight wild-type embryos. Data for the six Notch target cells (ABp, ABalp, ABara, ABplaaa, ABplpapp, and ABplpppp) are shown in red, and data for their sisters receiving no effective signaling (ABa, ABala, ABarp, ABplaap, and ABplpppa) or substantially weaker signaling (ABplpapa) are shown in blue. The statistical significance is obtained by the one-sided Wilcoxon rank-sum test and is listed at the top. C A summary graph showing the alteration in cell volume asymmetry between the daughters of six Notch target cells (red) or between the daughters of their sisters receiving no effective signaling or substantially weaker signaling (blue). Cells are colored as in (B). D Comparison of volume (V) asymmetries of daughters of Notch target cells between wild-type (horizontal) and perturbed (vertical) embryos by RNAi against lag-1. The cell volume asymmetry between each pair of daughter cells is averaged over eight wild-type replicates and over two lag-1- replicates. The statistical significance is obtained by the one-sided Wilcoxon rank-sum test and is listed on the left, along with the data average (solid dot) and standard deviation (solid line) presented. E Comparison of morphological changes between the Notch-responsive ABplpapp cell (middle) that receives the fourth Notch signaling and its sibling (top), which receives substantially weaker or no effective Notch signaling, in a wild-type embryo, or the ABplpapp cell in a perturbed embryo with RNAi against lag-1 (bottom). Note the directional size asymmetry in the division of ABplpapp (middle) in contrast to its sister (top), and its perturbed state in embryos with RNAi against lag-1 (bottom). \({T}_{{{{\rm{C}}}}}\) denotes the last time point of cytokinesis. Source data are provided as a Source Data file.
Surprisingly, we found that in addition to the fate asymmetry between the daughter cells of Notch target cells, the Notch signaling always enlarges the anterior daughter at the cost of the posterior one (one-sided Wilcoxon rank-sum test, \(p\le 0.001\), for all sister-cell pairs), examined in eight wild-type embryos and without exception (Fig. 4B). Such a significant shift also occurs in cell surface area asymmetry (one-sided Wilcoxon rank-sum test, \(p\le 0.01\) for all sister-cell pairs), without any exception in the eight wild-type embryos (Fig. S23). As a result, the size asymmetry between the daughters of the cells receiving the first, second, and fourth Notch signals is completely reversed, whereas that of the third Notch signal is decreased, and the initial size asymmetry between the anterior and posterior daughters of cells receiving the fifth Notch signal is further increased (Fig. 4C). Notably, these shifts were reproducibly observed in the sister-cell pairs of all eight wild-type embryos, which indicates that Notch signaling not only induces fate asymmetry but also probably promotes directional change in cell size.
We functionally validated the effect of Notch signaling on the size asymmetry of its target cells by comparing the size asymmetry between the daughters of Notch target cells before and after the RNAi against lag-1 (“RNAi_lag-1_1” and “RNAi_lag-1_2”, all derived from the same experimental condition; Supplementary Data 3; “Methods”), which encodes the terminal effector of the Notch signaling pathway76. We found that the difference in the value of \(\frac{{V}_{{{{\rm{D}}}}1}-{V}_{{{{\rm{D}}}}2}}{{V}_{{{{\rm{D}}}}1}+{V}_{{{{\rm{D}}}}2}}\) between the anterior and posterior sister-cell pairs (i.e., \(\Delta \left[\frac{{V}_{{{{\rm{D}}}}1}-{V}_{{{{\rm{D}}}}2}}{{V}_{{{{\rm{D}}}}1}+{V}_{{{{\rm{D}}}}2}}\right]\), which is always positive in a wild-type animal), in which one receives Notch signaling and the other receives no effective signaling or substantially weaker signaling, is significantly reduced in four out of the six Notch signaling events (Fig. 4D). These results further suggest that Notch signaling probably plays a role in enlarging the anterior but shrinking the posterior daughter of its target cells or a combination of the two for most target cells, and suggest that Notch probably mediates its effect independent of lag-1 (Fig. 4E).
A comprehensive comparative scanning reveals that the cell-size asymmetry effect in both wild-type and lag-1- embryos does not occur frequently across all anterior and posterior sibling cells. For 106 sister-cell pairs whose daughter cells completed full cell cycles in all wild-type and lag-1- embryo samples, we calculated the difference in the value of \(\frac{{V}_{{{{\rm{D}}}}1}-{V}_{{{{\rm{D}}}}2}}{{V}_{{{{\rm{D}}}}1}+{V}_{{{{\rm{D}}}}2}}\) between the anterior and posterior sister-cell pairs (i.e., \({\Delta }_{{{{\rm{A}}}}-{{{\rm{P}}}}}\left[\frac{{V}_{{{{\rm{D}}}}1}-{V}_{{{{\rm{D}}}}2}}{{V}_{{{{\rm{D}}}}1}+{V}_{{{{\rm{D}}}}2}}\right]={\left[\frac{{V}_{{{{\rm{D}}}}1}-{V}_{{{{\rm{D}}}}2}}{{V}_{{{{\rm{D}}}}1}+{V}_{{{{\rm{D}}}}2}}\right]}_{{{{\rm{A}}}}}-{\left[\frac{{V}_{{{{\rm{D}}}}1}-{V}_{{{{\rm{D}}}}2}}{{V}_{{{{\rm{D}}}}1}+{V}_{{{{\rm{D}}}}2}}\right]}_{{{{\rm{P}}}}}\)) (Supplementary Data 10). First, there is no significant population-wide bias toward anterior (i.e., \({\Delta }_{{{{\rm{A}}}}-{{{\rm{P}}}}}\left[\frac{{V}_{{{{\rm{D}}}}1}-{V}_{{{{\rm{D}}}}2}}{{V}_{{{{\rm{D}}}}1}+{V}_{{{{\rm{D}}}}2}}\right] > 0\)) or posterior (i.e., \({\Delta }_{{{{\rm{A}}}}-{{{\rm{P}}}}}\left[\frac{{V}_{{{{\rm{D}}}}1}-{V}_{{{{\rm{D}}}}2}}{{V}_{{{{\rm{D}}}}1}+{V}_{{{{\rm{D}}}}2}}\right] < 0\)) in terms of cell size (sign test, \(p=0.2853\)), although such bias was previously reported in gene expression regulated by Wnt signaling66,77 (Fig. S24). Second, 28% out of the 106 sister-cell pairs exhibit a significant difference in daughter cell size asymmetry, which is significantly reduced after RNAi against lag-1 (one-sided Wilcoxon rank-sum test, \(p\le 0.1\)), including the four sister-cell pairs involved in the second, third, and fourth Notch signaling events. Further investigation is warranted to determine whether the remaining sister-cell pairs are affected by ancestral cells or interacting cells involved in Notch signaling events, unidentified Notch signaling41, Notch-independent functions of lag-148,78,79, or distinct mechanical conditions arising from altered positions and contacts with neighboring cells80.
Multiple rounds of asymmetric divisions leading to the disproportionately large size of the C. elegans excretory cell since its birth
The C. elegans excretory cell ABplpappaap is the largest cell in adults and plays a vital role in osmotic and ionic regulation, and waste elimination30,47,81,82. (Fig. S25). However, how this disproportionately large size is established during embryogenesis is unclear. Our data reveals that ABplpappaap is indeed one of the largest cells in the late-stage embryo (the largest among AB cells and slightly smaller than E cells) (Fig. 5A). Given that the cell ABplpapp, the great-grandparent cell of ABplpappaap, was reported to receive the fourth Notch signaling, which is essential for the terminal fate specification and size asymmetry giving rise to the excretory cell ABplpappaap43 (Fig. 4B, C), we wondered whether the unusual size of ABplpappaap is generated during the subsequent rounds of cell division. We explored this process by using our cell lineage and cell volume data from eight wild-type embryos to plot the volumes of ABplpapp and its progenies over its subsequent three rounds of cell division that lead to its terminal fate differentiation. We found that the disproportionally large size of ABplpappaap is caused not only by the fourth Notch signaling event that induces asymmetric division in ABplpapp (Fig. 4B, C), but also by the subsequent two rounds of asymmetric division (Fig. 5B; Supplementary Movies S11 and S12). Strikingly, both wild-type and Notch-blocked embryos support that the general role of Notch signaling in promoting directional volume asymmetry is still at work (Fig. 5B, C). More than 80% of the parental cell volume is allocated to the excretory cell parent ABplpappaa and itself during the second and third rounds of asymmetric division with the second round producing an apoptotic cell. These results from wild-type and Notch-blocked embryos suggest that multiple rounds of Notch-regulated asymmetric cell divisions establish the disproportionately large size of ABplpappaap during the embryonic stage, which may contribute to its terminal size and its role as the functional equivalent of a kidney in the adult.
A The “H”-shaped excretory cell labeled by GFP (left) or its merge with differential interference contrast microscopy (DIC) (right) in an adult. B Top: quantification of volume changes over embryogenesis for the excretory cell and all the progeny of its great-grandmother, ABplpapp. The graph shows the average cell volumes (line) and their standard deviations (shaded area) for the excretory cell and its progenitors from eight wild-type embryos in red, and for their sister cells in green. Bottom: quantification of volume change over embryogenesis for all cells derived from AB (blue) and E (gray) with that for the excretory cell and its progenitors (red, same data as in upper row). The time of ABplpapp’s birth is used as the reference point (time zero). C Comparison of morphological changes between the excretory cell’s grandmother ABplpappa with Notch signaling (middle) and its sibling with substantially weaker or no effective Notch signaling (top), in a wild-type embryo, or the ABplpappa cell in a perturbed embryo with RNAi against lag-1 (bottom). Note the directional size asymmetry in the division of ABplpappa (middle) in contrast to its sister (top), and its perturbed state in embryos with RNAi against lag-1 (bottom). \({T}_{{{{\rm{C}}}}}\) denotes the last time point of cytokinesis. Source data are provided as a Source Data file.
Identification of asymmetric divisions regulated by cell signaling
Availability of the comprehensive cell-cell contact map throughout C. elegans embryogenesis allows not only the confirmation of the existing signaling interactions (Fig. 4A), but also the inference of interactions that drive cell fate or size asymmetry if the map is integrated with lineal expression of the components of signaling pathways. For example, if a cell expressing a Notch receptor bears direct contact with its neighbor that expresses a Notch ligand, and the cell undergoes a more asymmetric division in the anterior-posterior direction than its sister, the Notch signaling interaction is likely responsible for breaking the division symmetry. To this end, we collected lineal expression profiles from a total of nine existing or newly generated C. elegans transgenic embryos that express Notch ligands, i.e., apx-1 (strain: ZZY0559; construct: Papx-1:HIS-24:GFP:his-72 UTR) and lag-2 (strain: ZZY0596; construct: Plag-2:HIS-24:GFP:his-72 UTR), or Notch receptors, i.e., lin-12 (strain: ZZY0541; Plin-12:HIS-24:GFP:his-72 UTR) and glp-1 (strain: ZZY0558; construct: Pglp-1:HIS-24:GFP:his-72 UTR)41 (Fig. 6A; Fig. S26; Supplementary Data 11). By superimposing cell pairs that demonstrate a reproducible cell-cell contact and expression of the ligands and receptors respectively in each cell, we were able to identify five additional putative Notch signaling events, involving the previously identified target cell ABplpapp, and the newly identified target cells ABplpappa, ABplpappaa, and ABprpapp (Fig. 6B). All these Notch-responsive cells belong to the contralateral ABplpapp and ABprpapp sublineages, which have substantial directional size amplification of their anterior daughters compared to the size asymmetry of their sisters’ daughters, ranking in the top 7% within all sister-cell pairs with daughter volume available. To verify the effect of the inferred Notch signaling interaction, we compared the size asymmetries of the daughters of newly identified Notch target cells ABplpappa and ABprpapp and their respective sisters. We observed a significantly decreased size asymmetry in their daughters in the perturbed embryos with RNAi against lag-1 (one-sided Wilcoxon rank-sum test, \(p\le 0.05\)) (Fig. 5C). It is worth noting that the remaining Notch target cell ABplpappaa ceases division in the perturbed embryos with RNAi against lag-1, possibly attributed to its significantly reduced size compared to its normal size in wild-type embryos (11.9158 ± 17.6402 μm3 versus 93.11 ± 1.86 μm³; one-sided Wilcoxon rank-sum test, \(p\le 0.05\)).
A Lineal expression (blueness on cell lineage tree) of two ligands, lag-1 and apx-1, and one receptor, lin-12, of the Notch signaling pathway, each derived from an exemplary wild-type embryo. The lag-2 expression in MSapap is deduced based on previous studies43,75,84,85. The color scale is shown on the right. Cells involved in the sixth-tenth signaling events are indicated with arrows. Apoptosis is marked with an “X”. B Reconstructed 3D morphologies of contacting cell pairs engaged in the newly identified Notch signaling events during C. elegans embryogenesis. Cells expressing Notch ligands are highlighted in green, while those expressing the receptor are in red. C Comparison of morphological changes between the Notch-responsive ABprpapp cell (2nd row) that is deduced to receivs the tenth Notch signaling and its sibling (1st row), which is deduced to receive substantially weaker or no effective Notch signaling, in a wild-type embryo, or the ABprpapp cell in a perturbed embryo by RNA against lag-1 (3rd row) or laser ablation on MSpp (4th row). Note the directional size asymmetry in the division of ABprpapp (2nd row) in contrast to its sister (1st row), or its perturbed counterpart in embryos with RNAi against lag-1 (3rd row) or laser ablation of MSpp (4th row). \({T}_{{{{\rm{C}}}}}\) denotes the last time point of cytokinesis. D Positions (illustrated with the cell nucleus positions) of the Notch-responsive cells (red), Notch-signaling cells (green), and others (semi-transparent gray) at the moments when the sixth-tenth signaling events take place. Source data are provided as a Source Data file.
Specifically, following the well-established fourth Notch signaling interaction that targets ABplpapp by MSapp73 (Fig. 4A), we observed that the ABplpapp cell continues to receive Notch signaling by both daughters of MSapp and MSapap, which we referred to as the sixth Notch signaling event as judged by their direct contact and the lineal expression profile of a Notch receptor, lin-12, and a ligand, lag-2 (Fig. 4E and Fig. 6A, B, D) in the two cells respectively. It should be noted that while the daughters of MSapp (MSappa and MSappp) exhibit notable lag-2 expression in our data, the MSapap cell displays dimer and more indiscernible expression (Fig. 6A). This is likely due to the absence of some regulatory elements in our promoter construct or caused by the expression intensity that was too dim to be seen83. Therefore, we assume the lag-2 expression in MSapap based on the previous evidences43,75,84,85.
Similarly, the anterior daughter of ABplpapp, i.e., the grandparent of the excretory cell, also receives Notch signaling from three cells, MSaapa, MSaapp, and MSappa, which we referred to as the seventh Notch signaling event; and then the same cell receives Notch signaling from another three cells, MSaapap, MSaappa, and MSaappp, which we referred to as the eighth Notch signaling event (Fig. 6A, B, D). Finally, the cell ABplpappaa, the parent of the excretory cell, also receives Notch signaling from another two cells, MSaappa and MSaappp, which we referred to as the ninth Notch signaling event (Fig. 6A, B, D).
In addition, we observed that the ABprpapp cell, the symmetric cell of the ABplpapp that receives the well-established fourth Notch signaling73,75 (classically considered to break left-right lineage symmetry to produce the excretory cell and other unilateral cells), is also targeted by Notch signaling from the MSpppp cell, which expresses another Notch ligand, apx-1, whereas itself shows an expression of a Notch receptor, lin-12 (Fig. 6A, B, D). We referred to this as the tenth Notch signaling event. To confirm that Notch signaling indeed targets this sublineage, we used a laser beam to specifically ablate the MSpp cell, the grandmother of the proposed MSpppp signaling cell, as described previously41 (Supplementary Data 3; “Methods”). We found that the difference in the value of \(\frac{{V}_{{{{\rm{D}}}}1}-{V}_{{{{\rm{D}}}}2}}{{V}_{{{{\rm{D}}}}1}+{V}_{{{{\rm{D}}}}2}}\) between the ABprpapp and ABprpapa sister-cell pairs (i.e., \(\Delta \left[\frac{{V}_{{{{\rm{D}}}}1}-{V}_{{{{\rm{D}}}}2}}{{V}_{{{{\rm{D}}}}1}+{V}_{{{{\rm{D}}}}2}}\right]\), which is always positive in a wild-type animal), is significantly reduced (one-sided Wilcoxon rank-sum test, \(p\le 0.05\)) in two individual embryos (“Abla_MSpp_1” and “Abla_MSpp_2”, all derived from the same experimental condition; Supplementary Data 3; “Methods”), resulting in an ABprpappa cell that was smaller than its wild-type counterpart (Fig. 6C). This result together with those size asymmetries in wild-type and lag-1- embryos further supports that the ABprpapp cell receives Notch signaling, which results in size asymmetry between its daughters’ cell size. The results suggest that the fourth Notch signal alone may not be sufficient to break the lineage symmetry, and the fourth interaction together with the newly identified seventh, eighth, and ninth downstream interactions might be all required for proper differentiation between the relevant left and right lineages.
Taken together, equipped with our comprehensive map of intercellular contact in combination with lineal expression of ligand and receptor of signaling pathways, we were able to not only confirm the existing signaling interactions, but also identify signaling interactions that are responsible for inducing size asymmetry that is often coupled with fate asymmetry.
Extensive regulation of cell size asymmetries by multiple mechanisms
Given that the fate and size asymmetries of a subset of cells can be regulated by the Notch signaling interactions (Figs. 4–6), we wonder whether other mechanisms are also involved in breaking the size symmetry of other cells. It is known that Wnt signaling contributes to the specification of C. elegans embryonic cell fate62,66,86, and the nuclear level of the T cell factor (TCF) protein POP-1 (the terminal effector of the Wnt signaling pathway) is lowered in all posterior cells77,87. Therefore, we did RNAi against pop-1 followed by cell size analysis to investigate whether Wnt signaling regulates division asymmetry in terms of cell size (Supplementary Data 3). Blocking the Wnt signaling pathway can alter cell division orientation, which hinders the unambiguous assignment of cell identities based on division axes. Therefore, we focused on only cell size asymmetry between sister cells, regardless of their position, by calculating the absolute value of cell volume asymmetry, i.e., \(\left|\frac{{V}_{{{{\rm{D}}}}1}-{V}_{{{{\rm{D}}}}2}}{{V}_{{{{\rm{D}}}}1}+{V}_{{{{\rm{D}}}}2}}\right|\). We analyzed 257 cell divisions that were present in both wild-type and RNAi-treated embryos, for which a complete lifespan for a mother cell and two daughter cells was available. There was a significant decrease in cell volume asymmetry in the RNAi-treated embryos, i.e., the average value decreased from 0.1194 (wild-type) to 0.0877 (RNAi-treated) (one-sided Wilcoxon rank-sum test, \(p=1.44\times {10}^{-4}\)) (Fig. 7A). When Wnt signaling was perturbed, the number of cells whose daughter cells exhibited low volume asymmetry ( < 0.1) increased substantially in the RNAi-treated embryos, and vice versa (Fig. 7A). These results indicate that Wnt signaling plays a role in breaking cell size symmetry during embryogenesis.
A The distribution of cell volume asymmetry between daughter cells without positional bias (\(\left|\frac{{V}_{{{{\rm{D}}}}1}\,-\,{V}_{{{{\rm{D}}}}2}}{{V}_{{{{\rm{D}}}}1}\,+\,{V}_{{{{\rm{D}}}}2}}\right|\)) in the wild-type and pop-1- (pop-1 RNAi) embryos. The cell volume asymmetry between each pair of daughter cells is averaged over eight wild-type replicates and over two pop-1- replicates. Based on 257 pairs of daughter cells present in all wild-type and pop-1- embryos, the statistical significance is obtained by the one-sided Wilcoxon rank-sum test and is listed in the top right corner, along with the data average (solid dot) and standard deviation (solid line) presented. B The negative correlation between the shift of cell volume asymmetry with (\({\delta }_{{{{\rm{U}}}}-{{{\rm{C}}}}}\left[\frac{{V}_{{{{\rm{D}}}}1}\,-\,{V}_{{{{\rm{D}}}}2}}{{V}_{{{{\rm{D}}}}1}\,+\,{V}_{{{{\rm{D}}}}2}}\right]\)) and without mechanical compression (\({\left[\frac{{V}_{{{{\rm{D}}}}1}-{V}_{{{{\rm{D}}}}2}}{{V}_{{{{\rm{D}}}}1}+{V}_{{{{\rm{D}}}}2}}\right]}_{{{{\rm{U}}}}}\)). The result of proportional fitting between \({\left[\frac{{V}_{{{{\rm{D}}}}1}-{V}_{{{{\rm{D}}}}2}}{{V}_{{{{\rm{D}}}}1}+{V}_{{{{\rm{D}}}}2}}\right]}_{{{{\rm{U}}}}}\) and \({\delta }_{{{{\rm{U}}}}-{{{\rm{C}}}}}\left[\frac{{V}_{{{{\rm{D}}}}1}\,-\,{V}_{{{{\rm{D}}}}2}}{{V}_{{{{\rm{D}}}}1}\,+\,{V}_{{{{\rm{D}}}}2}}\right]\) is shown with a solid line, with the proportional coefficient (\(K\)) and goodness of fit (\(G\)) listed in the top right corner. The cell volume asymmetry between each pair of daughter cells is averaged over eight wild-type uncompressed replicates and 17 wild-type compressed replicates. The statistical significance is obtained by the one-sided Wilcoxon rank-sum test and is listed in the bottom left corner. C The apoptotic cells (vertical) are mostly smaller in volume compared to their sisters (horizontal) upon their birth. Shown are average volumes of 93 non-apoptotic and apoptotic sister-cell pairs recorded in the eight wild-type embryos, 80 of which have a relatively smaller volume for the apoptotic cells. D The illustration for asymmetric divisions of three representative parents of apoptotic cells from the AB (left), MS (middle), and C (right) lineages. For each cell, only cellular morphology at the time points before and after cytokinesis is shown. Source data are provided as a Source Data file.
In C. elegans embryogenesis up to the 28-cell stage, a proportion of cell division events decreased cell volume asymmetry between daughter cells when the eggshell was removed50. We wondered whether this indicates that external mechanical compression, which is believed to increase internal pressure and change cell positions in an embryo, increased cell size asymmetry. We investigated this by using the 17 C. elegans wild-type embryos (“Sample04” to “Samples20”, all derived from the same experimental condition; published previously) under external mechanical compression (applied by cover slide), which deformed the originally ellipsoidal shape of the eggshell into an elliptical cylinder with an approximate width-to-height ratio of 1:2 (9.4675 ± 0.2693 μm to 18.2534 ± 0.0373 μm) in its cross-section parallel to the direction of imaging12 (Supplementary Data 12). These embryo samples were segmented for the first half of C. elegans embryogenesis (approximately from the four- to 350-cell stages), such that 285 cells whose daughter cells had a complete lifespan were analyzed (Supplementary Data 13). We regarded a change in cell volume asymmetry \(\left({\delta }_{{{{\rm{U}}}}-{{{\rm{C}}}}}\left[\frac{{V}_{{{{\rm{D}}}}1}-{V}_{{{{\rm{D}}}}2}}{{V}_{{{{\rm{D}}}}1}+{V}_{{{{\rm{D}}}}2}}\right]\right)\) relative to the default in the mechanically uncompressed state (\({\left[\frac{{V}_{{{{\rm{D}}}}1}-{V}_{{{{\rm{D}}}}2}}{{V}_{{{{\rm{D}}}}1}+{V}_{{{{\rm{D}}}}2}}\right]}_{{{{\rm{U}}}}}\)) as positive if the volume of the anterior/left/dorsal daughter cell (\({{{\rm{D}}}}1\)) decreased under external mechanical compression, and negative if this volume increased under external mechanical compression. Intriguingly, we found that there was a modest negative correlation (goodness of fit = 0.3171) between the two variables, with 151 cell division events (~50%) exhibiting significantly different cell volume asymmetry (one-sided Wilcoxon rank-sum test, \(p\le 0.1\)) and more than half being in the second quadrant (\({\left[\frac{{V}_{{{{\rm{D}}}}1}-{V}_{{{{\rm{D}}}}2}}{{V}_{{{{\rm{D}}}}1}+{V}_{{{{\rm{D}}}}2}}\right]}_{{{{\rm{U}}}}} < 0\), \({\delta }_{{{{\rm{U}}}}-{{{\rm{C}}}}}\left[\frac{{V}_{{{{\rm{D}}}}1}-{V}_{{{{\rm{D}}}}2}}{{V}_{{{{\rm{D}}}}1}+{V}_{{{{\rm{D}}}}2}}\right] > 0\)), which means a collective increased volume asymmetry under external mechanical compression (Fig. 7B; Supplementary Data 13). This is consistent with the results of a comparison of the daughter cells of ABpl between mechanically uncompressed and compressed embryos reported previously39. Overall, our evaluation of multiple cells in multiple generations indicates that external mechanical compression amplifies the effect of Wnt signaling on cell volume asymmetry, i.e., most cell pairs exhibited significantly higher asymmetry when they were under mechanical compression than when they were not (Fig. 7B). Apart from cell volume, we also observed a population-wide shift in cell irregularity when comparing uncompressed and compressed embryos. This conclusion was based on an analysis of 321 unique cells prior to the ~350-cell stage, each with a complete lifespan recorded and morphological data available across all embryo samples. In compressed embryos, cells with originally low average irregularity ( < 2.5) tended to become less regular (21 out of 23 cells, >90%, occupying 87.5% of the cells in the fourth and fifth generations after fertilization). In contrast, cells with originally high average irregularity ( > 2.5) tended to become more regular (282 out of 298 cells, >90%, occupying most cells in the sixth generation and beyond) (Fig. S27; Supplementary Data 14). Notably, under normal conditions (particularly in older or starved adult worms), embryos experience some degree of compression in utero due to their tightly packed arrangement, and previous studies reported changed cell movements versus those in well-fed animals49. Whether the simultaneous changes in cell volume and cell irregularity are causally related remains an interesting question that warrants further exploration.
Cell divisions resulting in an apoptotic daughter are frequently found to exhibit size asymmetry, regardless of the timing of apoptotic onset88,89,90. For example, MSpaapp proceeds to apoptosis immediately after its birth, whereas ABprppppppa is involved in the development of the spike tail and proceeds to apoptosis at a very late stage30, they are both smaller than their sister cells right after their birth (with an average volume proportion of 0.15 and 0.59 respectively), suggesting that the smaller daughter tends to be more likely to undergo apoptosis (Fig. S28). Here, we systematically explored whether all such divisions are correlated with an asymmetric division by comparing the cell sizes immediately after cytokinesis of 93 sister-cell pairs in our dataset that contained one apoptotic cell and one non-apoptotic cell. Most apoptotic cells (78 out of 93 sister-cell pairs, >80%) demonstrated an average smaller volume and surface area than their sister cells, regardless of their lineal origin (Fig. 7C, D; Fig. S29; Supplementary Data 15). The result shows that the size asymmetry between an apoptotic cell and its sister cell that has been reported previously in specific cell types88,89,90 (e.g., neuroblasts) is a global feature of cell division involving apoptosis. However, additional follow-up experiments are needed to determine whether the small cell size is a cause of apoptosis or merely a correlation, particularly given that a small subset of these cell divisions produce near-symmetric daughters in size or even produce an apoptotic daughter that is larger than its non-apoptotic sister83 (Fig. 7C, D; Fig. S29; Supplementary Data 15).
A customized software tool for the visualization and interactive analysis of embryonic cell morphologies
To facilitate access to our cell morphological data, we employed the public software ITK-SNAP to visualize a 3D image of both raw and segmented cell regions within three orthogonal cross-sections and generate rendered objects91, and built a new version of the software, named ITK-SNAP-CVE (where “CVE” stands for “C. elegans virtual embryogenesis”), which allows customized visualization and analysis of C. elegans cell morphological data from multiple embryo samples, including cell identities, shapes, and quantitative morphological features (i.e., cell volume, surface area, and contact area) over embryogenesis (Fig. 8A; Supplementary Movie 13). To facilitate the use of the software, we reformatted all the raw images and processed images of the eight uncompressed wild-type embryos, two uncompressed cell-ablated wild-type embryos, four uncompressed RNAi-treated embryos, and two uncompressed mutant embryos used in this study, and those from the 17 compressed wild-type embryos described previously12 (Supplementary Data 16). We also developed several display modes, namely, “Show all cells”, “Show master cells only”, “Show master cells and neighbors”, and “Show master cells and other cells” (Fig. 8B–E). Master cells can be arbitrarily selected by the user through inputting or selecting a cell name (cell identity) from cells with a certain lineage (Fig. 8F) or fate (Fig. 8G), and their neighboring cells as well as the remaining embryonic cells can be shown with tunable opacity. A main menu selection option (on the top right of the interface) and a submenu (on the bottom right of the interface) enable the quantitative morphological features of a cell to be tracked. Thus, ITK-SNAP-CVE is an integrative tool that allows researchers to navigate cell-resolved C. elegans embryonic morphological maps interactively, thereby facilitating comprehensive visualization and analysis of this informative data.
A The main graphical user interface of ITK-SNAP-CVE, showcasing the layout and available tools. B The visual representations of all cells within an embryo using the software’s “Show all cells” display mode, with 2D views (top) and 3D reconstructions (bottom). C–E The detailed visualization of a selected individual cell, i.e., the somatic founder cell “C”, within an embryo, as seen through different viewing options: Panel (C) “Show master cells only” display mode, highlighting the “C” cell alone. Panel (D) “Show master cells and neighbors” display mode, highlighting the “C” cell along with its immediate neighboring cells. Panel (E) “Show master cells and other cells” display mode, where the “C” cell is visible in the context of the entire cell population. F A comprehensive view of all cells derived from the same lineage, exemplified here by the MS sublineage, demonstrating the lineage-specific visualization capabilities of the software. G A display of all cells that are destined to become part of the same organ, in this case, the intestine, illustrating the software’s functionality to group cells by their developmental fate.
An interactive web platform for visualizing embryonic cell morphologies, intercellular contacts, and cell-resolved lineal gene expressions
Gene expression underpins cell morphology, bridging the molecular mechanism and the cellular morphology dynamics. The effect is mutual: in one way, activities of specific genes as well as their products control the cell morphology through intracellular to intercellular mechanics, including cytoskeleton remodeling, cell adhesion, and gap junction92,93,94; in the other way, intracellular and intercellular mechanics control the cell morphology, such as the cell-cell contact area, then the signaling transduction that regulates fate specification and division orientation would be affected73,95,96. Actually, this effect is even more complex, given the existence of various mechanosensitive pathways and gene interactions, making the comprehensive understanding of development a difficult task as it crosses multiple scales and dimensions that influence each other65,97,98,99.
To further facilitate access to our data, we developed a website, CMOS (where “CMOS” stands for “cellular morphology of C. elegans embryo”; https://bcc.ee.cityu.edu.hk/cmos), that allows interactive access to the cell morphological data for the eight wild-type embryos generated in this study, including cell shape, volume, surface area and contact area as well as cell identity, lineage, and fate of all cells from the four- to 550-cell stages (Fig. 9; Supplementary Movie 14). To enhance the usability of our data for gene-related research, we integrated our morphological data with the existing and newly generated lineal expression profiles of various genes, mostly consisting of transcription factors4,36,37,41,65,99, which involved expression profiles of 412 unique genes derived from 1,044 individual embryos (approximately 30% of expression profiles from 87 cells up to the 350-cell stage, 60% between the 350- to 450-cell stages, and 10% between the 450- to 550-cell stages) (Supplementary Data 17), including those of tads-1 and snfc-5 that were generated in this study (Fig. S30). A gene’s lineal expression intensity can be projected onto all embryonic cells or onto cells in tissue-, organ- or lineage-specific manner (Fig. 9A–C). This will permit in-depth interpretation of a gene’s function in the context of cells’ lineal history, fate, and position on top of quantitative morphological data, which is not practical in any other species.
A The lineage-specific expression of the transcription factor, ceh-36, over approximately four hours from the four-cell stage. The relationship between gene expression level and color is displayed on the right. B The 3D views of an exemplary embryo at specified developmental stages (t, imaging time) with an overlay of ceh-36 expression (color-coded as in (A)). The embryo is oriented in a dorsal view with the anterior to the left. C The 3D views of different tissues and organs with highlighted expression of corresponding specific cell fate markers (color-coded as in (A)). The embryo is oriented in a ventral view with the anterior to the left. D The comparative views of a cell-cell contact map in an over 200-minute-old embryo (as seen in (B)): a global (left) and a cell-centric perspective (right). Intercellular contacts can be further examined in detail via an interactive table that appears upon clicking on a cell of interest. The thickness of the connecting lines corresponds to the cell-cell contact area. Expression levels for ceh-36 are superimposed on relevant cells, consistent with the visualization in (B). E The visualization of intercellular contacts for the sixth-tenth Notch signaling events (Fig. 6B) through the website. Source data are provided as a Source Data file.
The website allows the visualization of cell-cell contact maps either systematically or by focusing on a cell of interest with detailed quantitative data on cell morphology and intercellular contact displayed (Fig. 9D, E). The details on all the data source of each embryo, name of profiled genes, time point subjected to manual curation, construct type, and terminal cell number with a complete record for all cells are listed in Supplementary Data 17 (“Methods”). Cell morphological data, including cell identities, shapes, and quantitative morphological features (i.e., cell volume, surface area, and contact area), can be visualized along with the expression profile of a gene of interest over embryogenesis through navigating different time points, permitting time-lapse monitoring of gene expression and cell morphology as exemplified by pha-4 for pharynx assembly, hlh-1 for body-wall muscle assembly, and pal-1 and end-1 for transient expression (Figs. S31, S32).
Discussion
Systematic and quantitative characterization of cellular morphology over development is critical for an in-depth understanding of regulatory control of embryogenesis. Here we developed a platform that allows the systematic reconstruction and quantification of the cellular morphology of C. elegans embryogenesis up to beyond the 550-cell stage, when most embryonic cells complete their final round of division and thus differentiate into their terminal fate (Figs. 1, 2). Specifically, it is the improvement in the following features that make this platform significantly outperform the previous one we built earlier12,42. The first is the development of a cell segmentation algorithm, CMap, which allows the segmentation of time-lapse 3D cell images up to beyond the 550-cell stage with a much higher accuracy. This was achieved by a combination of integration of cell nuclei information to guide segmentation and a rigorous quality control of data post segmentation. The second is the generation of a transgenic strain of C. elegans that shows bright and ubiquitous expression on embryonic cell membranes, especially in late-stage embryos, which contributes to improved segmentation accuracy during late embryogenesis. The third is the development of a worm mounting method that allows long-duration imaging without applying any pressure on the imaged embryo, preventing artificial embryonic or cellular shape deformation. The fourth is the development of customized standalone software, ITK-SNAP-CVE, and a customized online website, CMOS, which allows local and remote access to our cell morphological data, permitting interactive visualization of 3D cell shape, intercellular cell contact, their superimposition with lineal expression of genes, and so forth (Figs. 8, 9). Finally, our comprehensive morphological map covers over 95% of all cells that are present during C. elegans embryogenesis (Supplementary Data 4), including cell shape, volume, surface area, and contact area in qualitative and quantitative format, together with their cell identity, lineage, and fate as well as lineal expression of about 400 genes, forms an invaluable resource for the study of the regulation of embryogenesis with spatiotemporal resolution and depth. For example, our data allows the vivid examination of gene expression in specific tissues, organs, or cell lineages with various cell shapes and sizes throughout embryogenesis (Fig. 9; Figs. S30 and S31).
Through the integration with lineal expression of ligands and receptors, our morphological map not only allows the confirmation of existing Notch signaling interactions but also permits the identification of signaling interactions that drive asymmetry of cell size that is coupled with cell fate (Figs. 4–6 and Fig. 7A). It also enables the study of the regulation of cell size by external mechanical compression and apoptosis (Fig. 7B–D). Quantification of cell irregularity permits the characterization of major morphogenetic events such as dorsal intercalation, intestinal formation, and body-wall muscle assembly (Fig. 3). Furthermore, the map could be used to answer many other key questions. For example, why are some divisions involving apoptosis asymmetric, whereas others are symmetric in terms of cell size? Is an apoptotic body engulfed randomly by any of its neighbors or by a specific neighbor (Fig. S33)? Whether a division asynchrony is co-regulated with fate or size asymmetry? To what extent does a shift in cell size asymmetry caused by external mechanical compression lead to lethality? How does the asymmetry in cell fate or size correlate with gene expression at the cellular level, for example, by integrating with existing single-cell RNA sequencing data4? Answering these questions may reveal compensatory or fail-safe mechanisms that underpin C. elegans embryogenesis22,49. In summary, the morphological map of C. elegans embryonic cells and the associated methodology and tools developed in this study are expected to facilitate addressing these questions among many others, which would be difficult otherwise.
The cell-resolved morphological data introduce an understudied dimension to developmental biology, complementing the traditional 2D dimension or molecular dimension exemplified by RNA sequencing4. In doing so, many longstanding questions in developmental biology can be revisited: how do cells acquire different properties during embryogenesis? For example, what morphological features distinguish cells based on tissue type or developmental stage? Can different cell types be differentiated by their morphological characteristics? In this study, critical morphogenetic events (including dorsal intercalation, intestinal formation, and body-wall muscle assembly) demonstrate a clear correlation between the increase in cell irregularity and short-range cell migration (Fig. 3). In addition, analysis of the EMS cell lineage and its fate differentiation shows that the changes in cell irregularity correlate with diverse developmental properties, including cell fates, cell lineage patterns (cell cycle lengths), cell positions, gene expression profiles, and cell morphological variability (Figs. S19 and S20). A more holistic, systems-wide examination of morphological properties from cell- or lineage-specific cells to all embryonic cells holds promise for a deeper understanding of metazoan development. Achieving this goal urgently calls for the advancement of morphological analytical methods, ranging from techniques like spherical harmonics and explicit geometric shape descriptors to emerging artificial intelligence-based approaches56,100.
The biological discoveries integrating the morphological map with biochemical experiments are valuable but necessitate further validation. Taking the identification of Notch signaling (Fig. 6) as an example, gene expression profiling and RNAi treatment were utilized for validation.
1) Regarding gene expression profiling, the promoter fusion used in this study has inherent limitations when the target gene contains large introns and/or extensive intergenic regions. In such cases, the reporter may lack critical cis-regulatory elements, potentially resulting in incomplete or ectopic expression patterns in the real mRNA and protein levels83. However, while protein fusion labels the final translated protein, it may affect the function or stability of the tagged protein, leading to altered protein levels and subcellular localization. This is particularly problematic when ligands and receptors are localized on the cell membrane, making them unsuitable for detection using cell-nucleus-based tools like AceTree/AceBatch33,36,101. To address this, computational methods for expression profiling at subcellular resolution are needed. To overcome the limitations of specific experimental approaches, future studies should employ alternative gene expression profiling techniques, such as Notch biosensor systems or single-cell RNA sequencing4,102,103, or direct Notch signaling activity monitoring by tracing the nucleus import of the Notch intracellular domain.
2) Regarding RNAi treatment, it is important to note that RNAi typically induces global effects in the embryo and achieves knockdown rather than complete knockout, which may result in incomplete elimination of the target gene activities. Besides, although the RNAi effect was checked by counting the ratio of embryonic death (which reached 100% in embryos laid 12 h after injection in this study), this measure may not necessarily reflect RNAi effectiveness in all embryonic cells, but likely to be a portion of them. Mutant alleles could serve as alternatives to achieve total gene knockout. Additionally, cell- or lineage-specific RNAi or somatic knockout could focus on the effect of a specific cell/population of interest, minimizing unintended effects on other cells. Furthermore, testing alternative Notch signaling pathway genes may be necessary, as a specific gene may play pleiotropic roles in multiple processes other than signaling. For instance, lag-1 is known to have Notch-independent functions48,78,79.
Overall, achieving a comprehensive understanding of morphological and biochemical processes requires multiple parallel lines of evidence. The morphological map and experimental-computational methodology generated in this study will facilitate the detailed characterization of cellular and molecular control of embryogenesis with exceptional spatial and temporal resolution.
Despite the comprehensiveness of our morphological map, it still suffers from data missing for certain cells. For example, we were unable to recover data on cellular morphology for all embryonic cells that appear during C. elegans embryogenesis. The total data-point loss rate showed an increase gradually as embryogenesis proceeded (Supplementary Data 7), and the highest cell loss rate was seen for the cells at the surface of late-stage embryos, where skin cells exhibited an overall data-point loss rate of 17.3%, with 19.5% of the C-derived skins (a major contributor to skin formation) being lost (Fig. S34; Supplementary Data 8). This segmentation failure may be attributable to both experimental and computational factors. First, the boundaries of the cells located inside the embryo are mostly defined by two layers of cell membranes due to cell–cell contact between neighboring cells, whereas those of cells located on the surface of the embryo consist of a single layer of their fluorescently labeled cell membranes (Fig. S35). Thus, the cell membrane fluorescence intensity of the embryo surface would have been half that inside the embryo, leading to a higher segmentation failure rate. This would have occurred because the extraembryonic space is treated as a dilatable region by CMap and thus invades the embryo when there is an insufficient number of pixels of membrane fluorescence on the surface of the embryo. Second, attenuation of laser intensity through the z axis of the embryo causes the signal-to-noise ratio of cell membrane fluorescence images collected from the top focal planes to be less than that of the images from the bottom focal planes (Fig. S35). This is on top of the insufficient z-axis resolution relative to that of the x/y axis, i.e., 0.42 µm/pixel versus 0.09 µm/pixel. Third, the parallelism between a cell membrane and an image focal plane prevents sampling of the membrane image due to the low z-axis resolution. This is a problem even in the segmentation of an early embryo29. Fourth, apoptosis frequently leads to cell shrinkage, which leads to the aggregation of cell membrane fluorescence signals into a “solid” ball (Fig. S35), making it impractical to segment the membrane boundary. Finally, given that our platform demands long-duration (over six hours) live-cell imaging with highly frequent image sampling (92 focal planes for each channel at ~1.5-minute intervals), phototoxicity and photobleaching prevent us from obtaining high-quality fluorescence images. This necessitates a tradeoff between the laser intensity and animal viability to ensure the viability of imaged animals and minimize the bleaching of fluorescence signals, which means that the quality of our actual images used for segmentation is much lower than that of still, single-shot images (Fig. S36). Therefore, future work should focus on the development of a more robust segmentation algorithm to deal with image deficiencies, alternative imaging methods to compensate for signal loss over laser passages or insufficient z-axis resolution, and transgenic strains that demonstrate uniform expression of membrane signals with a higher signal-to-noise ratio.
Methods
Worm strains, maintenance, and transgenesis
All the Caenorhabditis elegans were maintained on nematode growth media (NGM) plates seeded with Escherichia coli OP50 at a constant ambient temperature of 20 °C. To build a transgenic strain that shows bright and ubiquitous expression on cell membrane especially beyond the 350-cell stage, a transgenic strain, ZZY0855, with an insertion allele zzyIs139 [(Phis-72::PH(PLC1delta1)::mCherry::pie-1 3’ UTR+unc-119(+))], was generated using biolistic bombardment104. The allele zzyI139 was backcrossed with the N2 strain for five generations and crossed into another transgenic strain ZZY0535 with insertion alleles of zuIs178 [Phis-72(1 kb 5’ UTR)::HIS-72::SRPVAT::GFP::his-72 (1KB 3’ UTR) + 5.7 kb XbaI - HindIII unc-119(+)] and stIs10024 [Ppie-1::H2B::GFP::pie-1 3’ UTR+unc-119(+)], ltIs44 [Ppie-1::mCherry::PH(PLC1delta1)+unc-119( + )]41. All alleles were rendered homozygous to generate the strain, ZZY0861, for all live-cell imaging in this study. Additionally, wee-1.1 mutant allele ok418 was crossed into the ZZY0861 background by following genotype using single-worm PCR and marker expression and rendered homozygous before imaging. The resulting strains show bright and ubiquitous expressions of two fusions. One is between histone (HIS-72) and GFP that is expressed in all cell nuclei for automated cell tracing and lineaging; the other is between a membrane-specific domain PH(PLC1delta1) and mCherry that is expressed in all cell membranes, enabling automated membrane segmentation. The transgenic strain, BC10210, dpy-5(e907), sIs10089[dpy-5(+)+rCes-Ppgp-12-GFP+pCes361]82, was used for imaging the C. elegans excretory cell at postembryonic stages.
Fluorescence microscopy for wild-type, mutant, RNAi-treated, and cell-ablated embryos subjected to cell membrane segmentation
Time-lapse 3D imaging is similar to the one in our previously established data collection protocol12, with the following modifications to eliminate mechanical compression applied to the embryos. Embryos were mounted on a polylysine-pretreated glass slide to immobilize them during imaging. Instead of using glass beads that have a slightly smaller diameter than that of embryos, Vaseline was dotted outside the four corners of the polylysine pad to support the coverslip without touching the embryos to be imaged. For wild-type, mutant, and cell-ablated embryos, one- to four-celled embryos were retrieved from young adults (hermaphrodite) using a mouth pipette for mounting. For RNAi-treated embryos, double-stranded RNAs for lag-1 and pop-1 were synthesized in the test tube and injected into both gonads of young adults (hermaphrodite). One- to four-celled embryos were retrieved from the injected animals 12–24 h after injection. 12 µL of Boyd’s buffer was added on the polylysine pad before embryo transfer. The coverslip was gently placed on top of the buffer and pushed to the proximity of embryos by applying gentle pressure on the Vaseline on each corner of the polylysine pad. The edges of the coverslip were sealed with melted Vaseline before imaging.
Imaging was performed with the SP8 confocal microscope (Leica) at a constant ambient temperature of 20 °C. Images were acquired from both GFP and mCherry channels with a frame size of 712×512 pixels (x/y-axis resolution: 0.09 μm/pixel) and a scanning speed of 8000 Hz, using a water immersion objective. The excitation laser beams used for GFP and mCherry were 488 nm and 594 nm, respectively. Fluorescence images from 92 focal planes were consecutively collected for four embryos per imaging session, with a z-axis resolution of 0.42 μm/pixel, at ~1.5-minute intervals (Supplementary Data 3). Images were continuously collected for at least a total of 240 time points, during which the cell count would reach up to 550 and 330 in wild-type and RNAi-treated embryos, respectively. The entire imaging duration was divided into five blocks to accommodate fluorescence signal variation over development (1–60, 61–130, 131–160, 161–200, and 201–240 time points). The z-axis compensation was 0.5-3% for the 488 nm laser and 20-95% for the 594 nm laser. The pinhole sizes for the five blocks were 2.3, 2.0, 1.6, 1.6, and 1.3 Airy Units, respectively. Prior to image analysis, all images were subjected to deconvolution and renaming for automated tracing and lineaging, and membrane segmentation.
Fluorescence microscopy for postembryonic animals
Micrographs were acquired using the SP5 confocal microscope (Leica) with tile scanning and merged using the software LAS X (Leica). Intact animals were mounted on an agarose pad in Boyd’s buffer/methyl cellulose containing 0.1 M sodium azide for imaging with a scanning speed of 200–400 Hz depending on the size of the animals31. For the acquisition of 3D stack images, imaging settings were the same as those used for the embryo, except that the z-axis resolution is 1 µm/pixel for the adult and 0.42 µm/pixel for the embryo.
Automated cell tracing and lineaging
Using GFP-labeled cell nuclei images as input, automated cell tracing and lineaging as well as lineage curation were performed as described previously31,32,33. The automated tracing and lineaging results were manually curated up to beyond the 550-cell stage when all the progeny of the founder cell “C” were born and present for at least five time points. The cell nucleus information, including spatial position and cell identity at each time point, was output as a separate file to be used as input by CMap.
Calculation of cell volume, cell surface area, and cell-cell contact area
For the segmented 3D cell objects in C. elegans embryos, the cell volume V was calculated by summing the corresponding cell object’s pixels. Moreover, the Alpha shape mesh algorithm105, which is commonly used to create a 3D triangular mesh from the surface of a 3D object (in voxels format), was employed to extract the surface area AS of a cell and the contact area AC between two cell objects. The detailed procedure is as follows: (1) Dilation with the thickness of a pixel added was executed on each cell object, followed by the generation of a 3D triangular mesh from the dilated surface; (2) The cell surface area was calculated by summing the areas of all the triangles on the mesh; (3) The pixels at the points of contact between any two cell objects were detected and recorded; (4) For each of a pair of contacting cells, the partial area of its surface mesh that contains those pixels was calculated; (5) Among two cells that contact each other, the larger value of the abovementioned partial area (shared pixels between two cells), was adopted as the contact area between them. The quantification of the three morphological properties was jointly validated by two evaluations on each cell object of the eight wild-type embryos presented in this study: (1) The dimensionless cell irregularity defined as \(\frac{\root {2} \of {{A}_{{{{\rm{S}}}}}}}{\root {3}\; \of{V}}\) is always larger than the theoretical minimum \({\root {3} \of{6}} \,\cdot {\root {6} \of{\pi}}\approx 2.1991\) in a perfect sphere12; (2) The sum of the contact areas with neighboring cells never exceeds the surface area of any contacting cell with a relative discrepancy of 20%.
Automated cell membrane segmentation
In this study, an integrated method, CMap, was proposed for automatically segmenting 3D fluorescence images, also known as cellular volumetric segmentation. The proposed method includes three successive parts: (1) Interpolation is performed for the 2D TIFF images (x/y-axis resolution: 0.09 μm/pixel) collected with a z-axis resolution of 0.42 μm/pixel, transforming them into 3D NIfTI images with designed sizes and equal resolutions in all the x, y, and z directions; (2) A deep 3D neural network, the Euclidean distance transform dilated multifiber network (EDT-DMFNet), predicts the probability that a single pixel belongs to cell membrane; (3) With the probability map, the segmented instances corresponding to multiple separate 3D cell objects are generated. The details of these three parts are elaborated below.
Image preprocessing
For every embryo, the fluorescence images at each time point were originally collected from xy focal planes along the z axis (i.e., the depth direction z is perpendicular to the focal plane, xy) as a stack of 2D images, namely, a 3D stack image. Its corresponding pixel number along x (the width of the focal plane) and y (the height of the focal plane) axes are 712 and 512, with a total of 92 focal planes, respectively. For each raw 3D image, the pixels whose fluorescence intensity is within the highest and lowest 1% of the global range were discarded. Then, the 3D image underwent spatial resampling with downsampling along the x and y axes from 712 × 512 to 356 × 256 and upsampling along the z axis from 92 to 214, using bilinear interpolation.
Euclidean distance map prediction
The volumes of the rescaled 3D image were processed by EDT-DMFNet, which follows the structure of U-Net106, and were transformed into a membrane-centered 3D Euclidean distance map. To increase the perceptive field, within which the communications between convoluted channels are made in our small-scale network (a tiny fully connected network), inspired by DMFNet107, a 3D dilated regression network EDT-DMFNet was deployed to incorporate the information across multiple neighboring images in different convoluted channels with high computational speed (Fig. S6; Supplementary Data 2). EDT-DMFNet utilized a weighted fully dilated convolution to summarize the features at different scales adaptively. While group convolution was used to implement channel-wise convolution for a small network, a multiple-stage multiplexer, which is composed of several 1×1×1 filters, was applied to route the information among groups.
EDT-DMFNet transforms the segmentation task from a pixel classification problem to a pixel probability map regression problem. The designed network output, namely the inference on the probability map of pixels belonging to cell membrane \(P\in {\left[{{\mathrm{0,1}}}\right]}^{W\times H\times D}\) according to the input image \({I}_{0}\in {\left[{{\mathrm{0,255}}}\right]}^{W\times H\times D}\), allows multiple cell objects segmentation with limited computational resources with high speed. Here, W = 356, H = 256, and D = 214 are the width (along x axis), height (along y axis), and depth (along z axis) of 3D stack images. Therefore, the Softmax activation function at the output layer was replaced with the Sigmoid activation function for probability prediction. An adaptive membrane-centered weighted loss \(L\) was used to measure the difference between the target map (the ground-truth data) \(I\) and the predicted probability map \(\hat{I}\). The loss \(L\) of a 3D volumetric image was calculated by taking the weighted average of the mean squared error (MSE) between the target map \(I\) and the probability map \(\hat{I}\). Here, the adaptive membrane-centered weighted mask \({W}_{{{{\rm{mask}}}}}\) is defined as \({W}_{{{{\rm{mask}}}}}=\mu \cdot I+{avg}\left\{I\right\}\), where \(\mu\) is a constant (0.2) that scales the probability map \(\hat{I}\), and \({avg}\left\{I\right\}\) is the average value of \(I\) over the entire 3D volumetric image. In training, the pixels of the predicted probability map \(\hat{I}\) nearby cell membranes become more significant with the weighted mean squared error. Thus, the weighted mask substantially enhanced the contributions of non-zero pixels in the loss function because the non-zero value represents the probability of a pixel belonging to cell membrane in the predicted probability map \(\hat{I}\). Meanwhile, the pixels of the target map \(I\) surrounding the cell membrane should accordingly generate larger values in the weighted mask \({W}_{{{{\rm{mask}}}}}\). As a result, the loss grows rapidly if the EDT-DMFNet makes the wrong prediction for the pixels near the cell membrane, effectively encouraging the network to focus on the cell membrane and its surrounding pixels. In training, the input image\(\,{I}_{0}\) was imposed with random noises, cropped into a 128×128×128 volume, and flipped, which realized the augmentation of the training data and ensured the robustness of the network. The Adam optimizer was employed to update the network with an initial learning rate of \(5\times {10}^{-3}\) and a weight decay rate of \(1\times {10}^{-5}\), using AMSGrad gradient descent optimization. The model was trained for 50 epochs with a batch size of eight on an NVIDIA 2080 Ti GPU. The output of EDT-DMFNet, \(\hat{I}\), is a probability map as well as a Euclidean distance map for a 3D volumetric image, which will be subjected to the subsequent multiple cell objects segmentation.
Cell region generation
Based on the nucleus marker-seeded watershed algorithm as described previously108, CMap pre-inserts the experimentally established positions of cell nuclei into the Euclidean distance map \(\hat{I}\). This strategy confers solid cell position information to cell membrane segmentation as guidance cues. The cell nucleus marker-based watershed algorithm successfully improved the segmentation performance of CMap for C. elegans embryos, avoiding under- and over-segmentation problems, especially at the stage with over 350 cells in the embryo when cells become smaller and more crowded with a deteriorated signal-to-noise ratio. Such panoramic cell segmentation was executed up to the last time point with manual curation of automated lineage in each embryo.
Training data augmentation
Due to the rapidly changing cell morphology during embryogenesis and the uneven distribution of fluorescence signal, the 3D images captured in the experiment often have unexpected noise and indistinct intensity, which are harmful to cell membrane recognition (semantic segmentation) correctness to varying degrees. Importantly, by adding the low convoluted feature images into the upsampling layers, the U-shaped network accepts spatially operated images as different training data, which allows the deep convolutional neural networks to run effectively on a small medical or biological training dataset106. Thus, data augmentation is critical for the training of EDT-DMFNet. Appropriate and valid augmentation acquired manually not only guarantees the robustness of the network but also improves its universality, avoiding the potential over-fitting and under-fitting. Because of the small amount of training data (54 3D stack images reconstructed with two embryos), the augmentation part helps to improve the extent to which the training dataset simulates real-world data12. By perturbing the pixels’ intensity and randomly flipping and cropping an image, the 54 3D stack images in the training dataset were augmented to 21,600 effective training 128 × 128 × 128 cube images, which significantly improves the robustness of the trained EDT-DMFNet for other wild-type embryonic images. The protocols for data augmentation in training are described below.
Random intensity scale change: the intensity of all pixels in each training image was scaled with a uniform distribution, the half-open interval \([1,\,1.1)\).
Random intensity shift change: the intensity of all pixels in each training image was shifted with a uniform distribution, the half-open interval \([0,\,0.1)\).
Random flip: each training image had a 50% chance to be flipped along the x, y, and z axes, respectively.
Random crop: each training cube image was cropped randomly as \([128,\,128,\,128]\) from the original size \([205,\,285,\,134]\).
Manual annotation
The ground truth of cell morphology is necessary for evaluating the performance of machine segmentation algorithms. In this study, the cell morphology of two embryos, WT_Sample1 and WT_Sample7 (Fig. S11), was annotated by ten well-trained experts based on the segmentation results from CMap and the raw fluorescence images. The segmentation results were strictly checked and corrected slice by slice and cell by cell, ensuring the correctness of the ground truth established. Specifically, we gained two sets of ground truth data in different dimensions. First, the middle slice at each imaging time points throughout embryogenesis (255 2D images for WT_Sample1 and 205 for WT_Sample7) was annotated for 2D comparison, providing the cross-section of 30,509 cell objects in total. Second, the whole 3D stack within 100 ± 5-, 200 ± 5-, 300 ± 5-, 400 ± 5-, 500 ± 5-, and 550 ± 5-cell stages (six 3D images each of WT_Sample1 or WT_Sample7) were annotated for 3D comparison, providing the full morphology of 4046 cell objects in total.
Performance comparison between CMap and other state-of-the-art algorithms
The convolutional-neural-network-based method proposed in this study, CMap (Fig. 1A), consists of two algorithmic advances for processing the denoised 3D fluorescence images: (1) the deep learning network with optimal loss function and network structure (Fig. S6); (2) manually curated cell nuclei of automated tracked cells that provide seeds for cell membrane recognition. To evaluate how close the cell morphology reconstructed by CMap is to the real one, we manually segmented the 3D images with fluorescently labeled cell membranes at six different time points (at around 100-, 200-, 300-, 400-, 500-, and 550-cell stages) in two embryos (WT_Sample1 and WT_Sample7). Besides CShaper, using those ground truths (4,046 cell objects in total) as a reference, we compared the performance between CMap and other four state-of-the-art cell segmentation algorithms: 3DCellSeg109, CellPose3D110,111, StarDist3D112,113, and VNet114, in order to study the advantages and disadvantages of each algorithm.
In comparison to the manually annotated ground truth, the segmentation performance is quantitatively evaluated by the Hausdorff distance (defined as the largest of all distances from a pixel in one region to the closest pixel in the other region) and Dice score (defined as the ratio between the overlapping region and the overall region) for each segmented 3D cell object. Among the other four general cell segmentation algorithms (3DCellSeg, CellPose3D, StarDist3D, and VNet), StarDist3D and VNet have a Hausdorff distance approaching the ones of CShaper and CMap at up to the ~300- and ~200-cell stage respectively, revealing their applicability for the first half of C. elegans embryogenesis even though it is not customized for this system (one-sided Student’s t test, p > 0.1). Nonetheless, CMap still exhibits a significantly better performance than all the other algorithms at the ~400-, ~500-, and ~550-cell stages, no matter for Hausdorff distance or Dice score (one-sided Student’s t test, p ≤ 0.05) (Fig. S1; Supplementary Data 1). Note that VNet is a semantic segmentation network so we integrated the instance segmentation part of CShaper into its binary output for subsequent performance evaluation. All six algorithms were trained through the same pipeline.
Each algorithm generated multiple cell objects for 12 3D images (WT_Sample1 at 90/123/132/166/178/185 time points with 101/201/300/400/495/551 cells respectively and WT_Sample7 at 78/114/123/157/172/181 time points with 100/203/304/400/505/552 cells respectively). Then the segmentation outputs were quantitatively evaluated with the manually annotated ground truth by two metrics, Hausdorff distance and Dice score (Fig. S1). To avoid the complications associated with extreme cases and achieve a fair comparison with the Hausdorff distance, we excluded the data values smaller than 1 and larger than 20. For the Dice score, we excluded the data values smaller than 0.1 and larger than 0.9. The detailed results of each 3D image and each algorithm are listed in Supplementary Data 1.
Quality control on embryo imaging and cell tracing and lineaging
Every C. elegans wild-type embryo sample was imaged for a duration with the following criteria:
-
1.
The last moment of the four-cell stage must be recorded.
-
2.
The whole C sublineage must be generated, namely, with 8 progenies in the Caa sublineage, 8 progenies in the Cpa sublineage, 16 progenies in the Cap sublineage, and 16 progenies in the Cpp sublineage. Lastly, there are 48 C progenies in total, each of which has at least five time points recorded.
-
3.
The final recognized cell number must be over 570 and the total edited time points must be no less than 185 ( ≈ 265 min).
For the time-lapse 3D images, all the non-apoptotic cells have to be tracked by StarryNite and AceTree till their division or the last edited time point, while the apoptotic cells are tracked to their last time point when they are still distinguishable by the naked eye31,32,33. The results of cell tracing and lineaging were subjected to manual editing according to the procedures:
-
1.
For each cell with a complete lifespan recorded in all embryo samples, its normalized cell cycle lengths in all embryos are noted as [L1, L2, …, L8]. The top 10% cells with the largest max([L1, L2, …, L8])/mean([L1, L2, …, L8]) and the bottom 10% cells with the smallest min([L1, L2, …, L8])/mean([L1, L2, …, L8]) were subjected to manual check.
-
2.
For each 3D region marked by cell identity c, embryo sample j, and time point T, we characterized its volume dynamics with three characteristics:
-
The relative difference in volume between daughter cells and their mother is defined as
$${CPR}\left({c}_{1},{c}_{2},j,T\right)=\frac{\left|V\left({c}_{1},j,T\right)+V\left({c}_{2},j,T\right)-\hat{V}\left({p}_{0},j\right)\right|}{\hat{V}\left({p}_{0},j\right)}$$(1)where \(V\left({c}_{1},j,t\right)\) and \(V\left({c}_{2},j,t\right)\) are the volumes of two daughter cells \({c}_{1}\) and \({c}_{2}\) at time point \(T\); \(\hat{V}\left({p}_{0},j\right)\) is the median volume of the mother cell \({p}_{0}\) throughout its lifespan.
-
The volume inconsistency of cell \(c\) at two consecutive time points \(T+1\) and \(T\) is defined as
$${VIR}\left(c,j,T\right)=\frac{\left|V\left(c,j,T+1\right)-V\left(c,j,T\right)\right|}{\hat{V}\left(c,j\right)}$$(2) -
The volume variation of cell \(c\) at a specific time point \(T\) is defined as
For an embryo, if a specific cell at a specific time point belongs to the top 1/30 ≈ 3.33% outliers in all three criteria, it was subjected to manual check.
RNA interference
Gene knockdown was performed by RNA interference (RNAi) through microinjection. Primers for the amplification of the double-stranded RNA (dsRNA) template were as described previously115,116. For dsRNA production, the T7 promoter was included at the 5′ ends of both forward and reverse. The primers are as follows: lag-1: forward: TAATACGACTCACTATAGGG TCAGTCTCTTGCAAACCACG; reverse: TAATACGACTCACTATAGGG ATGCTGCAATCGAGAATGA; pop-1: forward: TAATACGACTCACTATAGG TTCCCAGGAAAGTTAGGCAA; reverse: TAATACGACTCACTATAGG AAACCGACACCCGTATGAAG. Polymerase Chain Reaction (PCR) was performed using C. elegans N2 genomic DNA as a template in 20 μL volume using ExTaq DNA polymerase (Cat# RR001Q, TaKaRa). After checking the PCR product on 1% agarose gel, 1 μL of the PCR product was used as a template for dsRNA production with NEB HiScribe T7 Quick High Yield RNA Synthesis Kit (Supplementary Data 11) according to the manufacturer’s description. The reaction mixture was incubated at 75 °C for 15 min in a water bath followed by turning off the heating power and incubating overnight in the same water bath for annealing of dsRNAs. The dsRNA was diluted to a concentration of 300 ng/μL in TE buffer for microinjection. At last, the automated tracing and lineaging results were manually corrected up to the stage when embryos were arrested, i.e., up to the 500-cell stage for two embryo samples with RNAi against lag-1 (the nucleus effector of the Notch signaling pathway) and up to the 330-cell stage for two embryo samples with RNAi against pop-1 (the nucleus effector of the Wnt signaling pathway) (Supplementary Data 3).
Cell ablation
One- to four-celled embryos were selected for time-lapse 3D imaging up to the time point when the target cell for ablation was born, as described previously41. The target cell was traced manually under direct observation, aided by the lineage marker. Immediately after the target cell was born, the following procedures were conducted within 1.5 min. The imaging was halted, and the focus was adjusted to the midplane of the target cell. The bleaching point was selected at the center of the target cell in the focal plane. The bleaching time was set to 40 seconds, and shutters for all other wavelengths were temporarily closed, leaving only the 488 nm laser beam active. The laser intensity was adjusted to 50%, and then the bleaching process was initiated. Once bleaching was complete, the standard time-lapse 3D imaging resumed.
Data filtering based on abnormal cell volume and cell shape
As a subset of the 3D cell objects segmented by CMap were apparently wrong, we devised a pipeline to filter them out from the last time point of the four-cell stage to the last edited time point for every embryo sample. Six strategies were applied as listed below.
-
1.
The volume V of the cell objects: V satisfies a bimodal distribution, where the extremely small cell volume is shown to result from the cell region that fails to dilate toward the distinguishable cell membrane and is close to the area of the cell nucleus (Figs. S8A and S10). A cell region is filtered out if it has a volume on the left of the valley of the bimodal distribution.
-
2.
The relative difference in volume between daughter cells and their mother CPR: A cell region is filtered out if it has a value over 99% of cell objects in the first half of C. elegans embryogenesis recorded by the CShaper dataset12, considering its high quality validated (Figs. S8B and S9).
-
3.
The volume inconsistency VIR of cell \(c\) at two consecutive time points \(T+1\) and \(T\): A cell region is filtered out if it has a value over 99% of cell objects in the first half of C. elegans embryogenesis recorded by the CShaper dataset, considering its high quality validated before12 (Figs. S8C and S9).
-
4.
The volume variation VVR of cell \(c\) at a specific time point \(T\): A cell region is filtered out if it has a value over 99% of cell objects in the first half of C. elegans embryogenesis recorded by the CShaper dataset12, considering its high quality validated (Figs. S8D and S9).
-
5.
The existence of separate regions SR, which disobeys the space continuity of cell shape: A cell region is filtered out if it contains two cell nuclei inside but neither the middle point nor the two quarter points between them belong to the cell region (Fig. S10).
-
6.
The filter rate FRL of lifespan recorded: A cell region is filtered out if over half of the time points within the lifespan of this cell have been filtered out by the five criteria above (Figs. S8E and S10).
Systematic checks for data consistency
To ensure that the massive data brought to the public is in the correct format, we wrote independent code to check its different aspects, including but not limited to:
-
1.
Every cell recorded in “SegCell\<embryo sample > _<time point > _segCell.nii.gz” is included in “<embryo sample > _lifescycle.csv” and “TPCell\<embryo sample > _<time point > _cells.txt”.
-
2.
Every cell recorded in “SegCell\<embryo sample > _<time point > _segCell.nii.gz” has been recorded in “name_dictionary.csv” and has a volume recorded in “<embryo sample > _volume.csv” and a surface area recorded in “<embryo sample > _surface.csv”.
-
3.
Every cell recorded in “SegCell\<embryo sample > _<time point > _segCell.nii.gz” has a volume and a surface area recorded in “<embryo sample > _<time point > _nucLoc.csv”.
-
4.
Every cell with a volume recorded in “<embryo sample > _volume.csv” and a surface area recorded in “<embryo sample > _surface.csv” has a corresponding segmented region recorded in “SegCell\<embryo sample > _<time point > _segCell.nii.gz”.
-
5.
Every cell with a volume and a surface area recorded in “<embryo sample > _<time point > _nucLoc.csv” has a corresponding segmented region recorded in “SegCell\<embryo sample > _<time point > _segCell.nii.gz”. The cell in “<embryo sample > _<time point > _nucLoc.csv” without a volume and a surface area recorded has a nucleus recorded in “Tracing_<embryo sample > .csv”.
-
6.
Every cell nucleus at a specific time point recorded in “Tracing_<embryo sample > .csv” is also recorded in “<embryo sample > _<time point > _nucLoc.csv”.
-
7.
Every cell-cell contact recorded in “GuiNeighbor\<embryo sample > _<time point > _guiNeighbor.txt” has an area recorded in “<embryo sample > _Stat.csv”, while the contacting cells have corresponding segmented regions recorded in “SegCell\<embryo sample > _<time point > _segCell.nii.gz”.
-
8.
Every dividing cell recorded in “DivisionCell\<embryo sample > _<time point > _division.txt” has no nucleus recorded in “Tracing_<embryo sample > .csv” but the nuclei of its two daughter cells are recorded.
-
9.
Every dividing cell recorded in “DivisionCell\<embryo sample > _<time point > _division.txt” has a volume recorded in “<embryo sample > _volume.csv” and a surface area recorded in “<embryo sample > _surface.csv”. If its division ends before the terminal edited time point, there is a volume recorded in “<embryo sample > _volume.csv” and a surface area recorded in “<embryo sample > _surface.csv” for its two daughter cells.
-
10.
Every dividing cell recorded in “DivisionCell\<embryo sample > _<time point > _division.txt” is included in “<embryo sample > _lifescycle.csv” and “TPCell\<embryo sample > _<time point > _cells.txt”, and has a corresponding segmented region recorded in “SegCell\<embryo sample > _<time point > _segCell.nii.gz”.
Three-dimensional illustration for segmented cell morphologies
All the 3D illustrations, including both figures and movies, are rendered with the software Fiji and Blender117,118 or visualized with ITK-SNAP91, ITK-SNAP-CVE (Supplementary Movie S13), and CMOS (Supplementary Movie S14).
Data collection and processing for lineal gene expression
All the C. elegans wild-type embryos dually with a lineaging marker and a gene of interest were imaged starting from no later than the four-cell stage, where the lineal gene expression was profiled by either promoter fusion or protein fusion as described before101. The ones reused from previous literature end no earlier than the initiation of the five primary somatic lineages, namely, the appearance of five somatic founder cells: AB, MS, E, C, and D; meanwhile, the tracing and lineaging for all cells is unambiguous and continuous for consecutive time points4,36,37,41,65,99. The ones generated in this study end at beyond the 350-cell stage, covering the following cells – AB256 (progenies after eight rounds of division in the AB lineage), MS32 (progenies after five rounds of division in the MS lineage), E16 (progenies after four rounds of division in the MS lineage), C8 (progenies after three rounds of division in the C lineage), D8 (progenies after three rounds of division in the D lineage), and Z2 and Z3, the same as our previously established data collection protocol25,119 except that the embryos with a time resolution of 1.41 and 1.54 minute/interval were imaged with 41 and 30 focal planes respectively, where the z-axis resolution was 0.71 and 1.01 μm/pixel respectively. For the CMOS website (Supplementary Movie 14), the parallel gene expression profiles from different individual embryo samples with the same labeled gene, the same construct type, and the same literature source were averaged and linearly normalized to 0 ~ 1, which was then superimposed on the cell lineage and morphological data of the eight segmented C. elegans wild-type embryos at each time point by proportionally fitting the differential developmental paces; if there’s only one embryo sample with the same labeled gene, the same construct type, and the same literature source, the processing procedure for the gene expression profile is the same except that no averaging operation was executed.
Statistics and reproducibility
The data produced in this study is systematic and unbiased, based on a pipeline integrating time-lapse 3D fluorescence imaging, cell lineage tracing (based on cell nucleus fluorscence), cell morphology segmentation (based on cell membrane fluorescence), and data quality control (based on abnormal cell volume or cell shape) on C. elegans embryos (Supplementary Data 3; “Methods”). No data were excluded from the subsequent analyses.
Six experimental groups with cell lineage-resolved embryonic morphological map were used for statistics (Supplementary Data 3): uncompressed wild-type embryos (“WT_Sample1” to “WT_Sample8”), uncompressed wee-1.1 mutant embryos (“MT_wee-1.1_1” and “MT_wee-1.1_2”), uncompressed pop-1 RNAi-treated embryos (“RNAi_pop-1_1” and “RNAi_pop-1_2”), uncompressed lag-1 RNAi-treated embryos (“RNAi_lag-1_1” and “RNAi_lag-1_2”), uncompressed cell-ablated wild-type embryos (“Abla_MSpp_1” and “Abla_MSpp_2”), and compressed wild-type embryos (“Sample04” to “Samples20”)12. Within each group, all embryo samples were derived from the same experimental condition (“Methods”). Wherever statistics have been derived, data from all embryos (or units) were used together, directly, and equally.
All statistical results are based on at least two replicates and associated significance values. For establishing the cell lineage-resolved embryonic morphological map under the wild-type condition, a sample size of 8 was chosen, in line with the typical range (between 2 and 10) employed in morphological map statistical research in other animal and plant species27,120. For gene expression profiling (using a null signal as baseline) and perturbed phenotype identification (using the wild-type phenotype as baseline), sample sizes of 2 or 3 were chosen, in line with the typical range (between 2 and 3) employed in C. elegans systems and quantitative genetics research36,121.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The ITK-SNAP-CVE software (including Windows/Mac/Linux versions) constructed in this study has been deposited in the figshare database under accession code https://doi.org/10.6084/m9.figshare.24768921122. The CMOS website (including quantitative cell morphology and gene expression data) constructed in this study can be accessed at https://bcc.ee.cityu.edu.hk/cmos123. The raw images of eight uncompressed wild-type embryos and four uncompressed RNAi-treated embryos as well as their cell lineage tracing and morphology segmentation results (readable directly in ITK-SNAP-CVE) generated in this study have been deposited in the figshare database under accession code https://doi.org/10.6084/m9.figshare.24768921122; the raw images of two uncompressed cell-ablated wild-type embryos and two uncompressed mutant embryos as well as their cell lineage tracing and morphology segmentation results (readable directly in ITK-SNAP-CVE) generated in this study have been deposited in the figshare database under accession code https://doi.org/10.6084/m9.figshare.27872238124. The manually annotated ground truth for training, evaluation, and apoptotic cells have been deposited in the OneDrive database under accession code https://doi.org/10.6084/m9.figshare.24768921122. Source Data for each figure and supplementary data has been deposited in the figshare database under accession code https://doi.org/10.6084/m9.figshare.28467284125. The strain (ZZY0861) used for cell morphology reconstruction by CMap is available from the Zhao Lab. It can also be ordered through the Caenorhabditis Genetics Center (CGC).
Code availability
The code related to the CMap algorithm is deposited on GitHub, under accession codes https://doi.org/10.5281/zenodo.14916840 (cell segmentation) and https://doi.org/10.5281/zenodo.14920949 (performance evaluation)126,127. In this study, the data collection code was implemented using multiple platforms: cell tracing and lineaging with StarryNite31,32, AceTree31,128, and Matlab (R2018b)129; cell membrane segmentation with Python (3.7)130; data quality evaluation and control with Matlab (R2018b) and Python (3.7)129,130. The data analysis code was implemented using multiple platforms: segmentation algorithm performance evaluation with Python (3.7) 130; biological quantitative analysis with Matlab (R2018b)129.
References
Arendt, D. et al. The origin and evolution of cell types. Nat. Rev. Genet. 17, 744–757 (2016).
Domcke, S. & Shendure, J. A reference cell tree will serve science better than a reference cell atlas. Cell 186, 1103–1114 (2023).
Qiu, C. et al. Systematic reconstruction of cellular trajectories across mouse embryogenesis. Nat. Genet. 54, 328–341 (2022).
Packer, J. S. et al. A lineage-resolved molecular atlas of C. elegans embryogenesis at single-cell resolution. Science 365, eaax1971 (2019).
Qiu, C. et al. A single-cell time-lapse of mouse prenatal development from gastrula to birth. Nature 626, 1084–1093 (2024).
Firmin, J. et al. Mechanics of human embryo compaction. Nature 629, 646–651 (2024).
Mittnenzweig, M. et al. A single-embryo, single-cell time-resolved model for mouse gastrulation. Cell 184, 2825–2842 (2021).
Junyent, S. et al. The first two blastomeres contribute unequally to the human embryo. Cell 187, 2838–2854 (2024).
Rasmussen, J. P., Feldman, J. L., Reddy, S. S. & Priess, J. R. Cell interactions and patterned intercalations shape and link epithelial tubes in C. elegans. PLoS Genet. 9, e1003772 (2013).
Goldstein, B. & Nance, J. Caenorhabditis elegans gastrulation: A model for understanding how cells polarize, change shape, and journey toward the center of an embryo. Genetics 214, 265–277 (2020).
Chen, H., Qian, W. & Good, M. C. Integrating cellular dimensions with cell differentiation during early development. Curr. Opin. Cell Biol. 67, 109–117 (2020).
Cao, J. et al. Establishment of a morphological atlas of the Caenorhabditis elegans embryo using deep-learning-based 4D segmentation. Nat. Commun. 11, 6254 (2020).
Pentinmikko, N. et al. Cellular shape reinforces niche to stem cell signaling in the small intestine. Sci. Adv. 8, eabm1847 (2022).
Van Den Eynden, J. et al. Phosphoproteome and gene expression profiling of ALK inhibition in neuroblastoma cell lines reveals conserved oncogenic pathways. Sci. Signal. 11, eaar5680 (2018).
Mello, C. C., Draper, B. W. & Priess, J. R. The maternal genes apx-1 and glp-1 and establishment of dorsal-ventral polarity in the early C. elegans embryo. Cell 77, 95–106 (1994).
Shelton, C. A. & Bowerman, B. Time-dependent responses to glp-1-mediated inductions in early C. elegans embryos. Development 122, 2043–2050 (1996).
Neves, A. & Priess, J. R. The ref-1 family of bHLH transcription factors pattern C. elegans embryos through Notch-dependent and Notch-independent pathways. Dev. Cell 8, 867–879 (2005).
Lin, R., Thompson, S. & Priess, J. R. pop-1 encodes an HMG box protein required for the specification of a mesoderm precursor in early C. elegans embryos. Cell 83, 599–609 (1995).
Rasmussen, J. P., English, K., Tenlen, J. R. & Priess, J. R. Notch signaling and morphogenesis of single-cell tubes in the C. elegans digestive tract. Dev. Cell 14, 559–569 (2008).
Jankele, R. et al. Physically asymmetric division of the C. elegans zygote ensures invariably successful embryogenesis. eLife 10, e61714 (2021).
Arata, Y., Takagi, H., Sako, Y. & Sawa, H. Power law relationship between cell cycle duration and cell volume in the early embryonic development of Caenorhabditis elegans. Front. Physiol. 5, 529 (2015).
Fickentscher, R., Struntz, P. & Weiss, M. Setting the clock for fail-safe early embryogenesis. Phys. Rev. Lett. 117, 188101 (2016).
Guan, G., Wong, M. K., Zhao, Z., Tang, L.-H. & Tang, C. Volume segregation programming in a nematode’s early embryogenesis. Phys. Rev. E 104, 054409 (2021).
Kuang, X. et al. Computable early Caenorhabditis elegans embryo with a phase field model. PLoS Comput. Biol. 18, e1009755 (2022).
Ho, V. W. S. et al. Systems‐level quantification of division timing reveals a common genetic architecture controlling asynchrony and fate asymmetry. Mol. Syst. Biol. 11, 814 (2015).
Tian, B., Guan, G., Tang, L.-H. & Tang, C. Why and how the nematode’s early embryogenesis can be precise and robust: A mechanical perspective. Phys. Biol. 17, 026001 (2020).
Guignard, L. et al. Contact area–dependent cell communication and the morphological invariance of ascidian embryogenesis. Science 369, eaar5663 (2020).
Stegmaier, J. et al. Real-time three-dimensional cell segmentation in large-scale microscopy data of developing embryos. Dev. Cell 36, 225–240 (2016).
Azuma, Y. & Onami, S. Biologically constrained optimization-based cell membrane segmentation in C. elegans embryos. BMC Bioinform. 18, 307 (2017).
Sulston, J. E., Schierenberg, E., White, J. G. & Thomson, J. N. The embryonic cell lineage of the nematode Caenorhabditis elegans. Dev. Biol. 100, 64–119 (1983).
Murray, J. I., Bao, Z., Boyle, T. J. & Waterston, R. H. The lineaging of fluorescently-labeled Caenorhabditis elegans embryos with StarryNite and AceTree. Nat. Protoc. 1, 1468–1476 (2006).
Bao, Z. et al. Automated cell lineage tracing in Caenorhabditis elegans. Proc. Natl. Acad. Sci. USA 103, 2707–2712 (2006).
Katzman, B., Tang, D., Santella, A. & Bao, Z. AceTree: A major update and case study in the long-term maintenance of open-source scientific software. BMC Bioinform. 19, 1–7 (2018).
Nance, J. & Frøkjær-Jensen, C. The Caenorhabditis elegans transgenic toolbox. Genetics 212, 959–990 (2019).
Dickinson, D. J. & Goldstein, B. CRISPR-Based Methods for Caenorhabditis elegans genome engineering. Genetics 202, 885–901 (2016).
Murray, J. I. et al. Multidimensional regulation of gene expression in the C. elegans embryo. Genome Res. 22, 1282–1294 (2012).
Ma, X. et al. A 4D single-cell protein atlas of transcription factors delineates spatiotemporal patterning during embryogenesis. Nat. Methods 18, 893–902 (2021).
Cao, J., Wong, M. K., Zhao, Z. & Yan, H. 3DMMS: Robust 3D membrane morphological segmentation of C. elegans embryo. BMC Bioinform 20, 176 (2019).
Thiels, W., Smeets, B., Cuvelier, M., Caroti, F. & Jelier, R. spheresDT/Mpacts-PiCS: Cell tracking and shape retrieval in membrane-labeled embryos. Bioinformatics 37, 4851–4856 (2021).
Guan, G., Zhao, Z. & Tang, C. Delineating mechanisms and design principles of Caenorhabditis elegans embryogenesis using in toto high-resolution imaging data and computational modeling. Comput. Struct. Biotechnol. J. 20, 5500–5515 (2022).
Chen, L. et al. Establishment of signaling interactions with cellular resolution for every cell cycle of embryogenesis. Genetics 209, 37–49 (2018).
Cao, J. et al. CShaperApp: Segmenting and analyzing cellular morphologies of the developing Caenorhabditis elegans embryo. Quant. Biol. 12, 329–334 (2024).
Hutter, H. & Schnabel, R. Establishment of left-right asymmetry in the Caenorhabditis elegans embryo: A multistep process involving a series of inductive events. Development 121, 3417–3424 (1995).
Lambie, E. J. & Kimble, J. Two homologous regulatory genes, lin-12 and glp-1, have overlapping functions. Development 112, 231–240 (1991).
Altun, Z. F. & Hall, D. H. Excretory system. WormAtlas. https://doi.org/10.3908/wormatlas.1.17 (2009).
Fujita, M. et al. The role of the ELAV homologue exc-7 in the development of the Caenorhabditis elegans excretory canals. Dev. Biol. 256, 290–301 (2003).
Buechner, M., Hall, D. H., Bhatt, H. & Hedgecock, E. M. Cystic canal mutants in Caenorhabditis elegans are defective in the apical membrane domain of the renal (excretory) cell. Dev. Biol. 214, 227–241 (1999).
Maicas, M., Jimeno-Martín, Á., Millán-Trejo, A., Alkema, M. T. & Flames, N. The transcription factor LAG-1/CSL plays a Notch-independent role in controlling terminal differentiation, fate maintenance, and plasticity of serotonergic chemosensory neurons. PLoS Biol. 19, e3001334 (2021).
Jelier, R., Kruger, A., Swoger, J., Zimmermann, T. & Lehner, B. Compensatory cell movements confer robustness to mechanical deformation during embryonic development. Cell Syst. 3, 160–171 (2016).
Fickentscher, R. & Weiss, M. Physical determinants of asymmetric cell divisions in the early development of Caenorhabditis elegans. Sci. Rep. 7, 9369 (2017).
Chisholm, A. D. & Hardin, J. Epidermal morphogenesis. WormBook, 1–22. https://doi.org/10.1895/wormbook.1.35.1 (2005).
Sulston, J. E. & White, J. G. “Parts list”, in “The Nematode Caenorhabditis elegans” edited by Wood W. B. et al., Cold Spring Harbor Laboratory Press, 415–431. https://www.wormatlas.org/celllistsulston.htm (1988).
Driscoll, K., Stanfield, G. M., Droste, R. & Horvitz, H. R. Presumptive TRP channel ced-11 promotes cell volume decrease and facilitates degradation of apoptotic cells in Caenorhabditis elegans. Proc. Natl. Acad. Sci. USA 114, 8806–8811 (2017).
Azuma, Y., Okada, H. & Onami, S. Systematic analysis of cell morphodynamics in C. elegans early embryogenesis. Front. Bioinform. 3, 1082531 (2023).
Fujii, Y. et al. Spatiotemporal dynamics of single cell stiffness in the early developing ascidian chordate embryo. Commun. Biol. 4, 341 (2021).
Guan, G., Chen, Y., Wang, H., Ouyang, Q. & Tang, C. Characterizing cellular physiological states with three-dimensional shape descriptors for cell membranes. Membranes 14, 137 (2024).
Bao, Z., Zhao, Z., Boyle, T. J., Murray, J. I. & Waterston, R. H. Control of cell cycle timing during C. elegans embryogenesis. Dev. Biol. 318, 65–72 (2008).
Li, X. et al. Systems properties and spatiotemporal regulation of cell position variability during embryogenesis. Cell Rep. 26, 313–321.e317 (2019).
Abdu, Y., Maniscalco, C., Heddleston, J. M., Chew, T. L. & Nance, J. Developmentally programmed germ cell remodelling by endodermal cell cannibalism. Nat. Cell Biol. 18, 1302–1310 (2016).
Harrell, J. R. & Goldstein, B. Internalization of multiple cells during C. elegans gastrulation depends on common cytoskeletal mechanisms but different cell polarity and cell fate regulators. Dev. Biol. 350, 1–12 (2011).
Altun, Z. F. & Hall, D. H. Epithelial system, hypodermis. WormAtlas. https://doi.org/10.3908/wormatlas.1.13 (2009).
Thorpe, C. J., Schlesinger, A., Carter, J. C. & Bowerman, B. Wnt signaling polarizes an early C. elegans blastomere to distinguish endoderm from mesoderm. Cell 90, 695–705 (1997).
Asan, A., Raiders, S. A. & Priess, J. R. Morphogenesis of the C. elegans intestine involves axon guidance genes. PLoS Genet. 12, e1006077 (2016).
Altun, Z. F. & Hall, D. H. Muscle system, somatic muscle. WormAtlas. https://doi.org/10.3908/wormatlas.1.7 (2009).
Guan, G. et al. Multilevel regulation of muscle-specific transcription factor hlh-1 during Caenorhabditis elegans embryogenesis. Dev. Genes Evol. 230, 265–278 (2020).
Eisenmann D. M. Wnt signaling. WormBook. https://doi.org/10.1895/wormbook.1.7.1 (2005).
Sommermann, E. M., Strohmaier, K. R., Maduro, M. F. & Rothman, J. H. Endoderm development in Caenorhabditis elegans: The synergistic action of ELT-2 and -7 mediates the specification->differentiation transition. Dev. Biol. 347, 154–166 (2010).
Wong, M. K. et al. Timing of tissue-specific cell division requires a differential onset of zygotic transcription during metazoan embryogenesis. J. Biol. Chem. 291, 12501–12513 (2016).
Pohl, C. & Bao, Z. Chiral Forces Organize Left-right patterning in C. elegans by uncoupling midline and anteroposterior axis. Dev. Cell 19, 402–412 (2010).
Pohl, C. Left-right patterning in the C. elegans embryo. Commun. Integr. Biol. 4, 34–40 (2011).
The C. elegans Deletion Mutant Consortium. Large-scale screening for targeted knockouts in the Caenorhabditis elegans genome. G3: Genes, Genomes, Genetics 2, 1415–1425 (2012).
Robertson, S. M., Medina, J. & Lin, R. Uncoupling different characteristics of the C. elegans E lineage from differentiation of intestinal markers. PLoS ONE 9, e106309 (2014).
Priess, J. R. Notch signaling in the C. elegans embryo. WormBook. https://doi.org/10.1895/wormbook.1.4.1 (2005).
Fickentscher, R., Krauss, S. W. & Weiss, M. Anti-correlation of cell volumes and cell-cycle times during the embryogenesis of a simple model organism. New J. Phys. 20, 113001 (2018).
Bertrand, V., Bisso, P., Poole, R. J. & Hobert, O. Notch-dependent induction of left/right asymmetry in C. elegans interneurons and motoneurons. Curr. Biol. 21, 1225–1231 (2011).
Christensen, S., Kodoyianni, V., Bosenberg, M., Friedman, L. & Kimble, J. lag-1, a gene required for lin-12 and glp-1 signaling in Caenorhabditis elegans, is homologous to human CBF1 and Drosophila Su(H). Development 122, 1373–1383 (1996).
Zacharias, A. L., Walton, T., Preston, E. & Murray, J. I. Quantitative differences in nuclear β-catenin and TCF pattern embryonic cells in C. elegans. PLoS Genet. 11, e1005585 (2015).
Ghai, V. & Gaudet, J. The CSL transcription factor LAG-1 directly represses hlh-6 expression in C. elegans. Dev. Biol. 322, 334–344 (2008).
Johnson, J. E. & MacDonald, R. J. Notch-independent functions of CSL. Curr. Top. Dev. Biol. 97, 55–74 (2011).
Cuvelier, M. et al. Stability of asymmetric cell division: A deformable cell model of cytokinesis applied to C. elegans. Biophys. J. 122, 1858–1867 (2023).
Nelson, F. K. & Riddle, D. L. Functional study of the Caenorhabditis elegans secretory‐excretory system using laser microsurgery. J. Exp. Zool. 231, 45–56 (1984).
Zhao, Z. et al. Distinct regulatory elements mediate similar expression patterns in the excretory cell of Caenorhabditis. elegans. J. Biol. Chem. 280, 38787–38794 (2005).
Reece-Hoyes, J. S. et al. Insight into transcription factor gene duplication from Caenorhabditis elegans Promoterome-driven expression patterns. BMC Genomics 8, 27 (2007).
Moskowitz, I. P. & Rothman, J. H. lin-12 and glp-1 are required zygotically for early embryonic cellular interactions and are regulated by maternal GLP-1 signaling in Caenorhabditis elegans. Development 122, 4105–4117 (1996).
Hermann, G. J., Leung, B. & Priess, J. R. Left-right asymmetry in C. elegans intestine organogenesis involves a LIN-12/Notch signaling pathway. Development 127, 3429–3440 (2000).
Goldstein, B. Induction of gut in Caenorhabditis elegans embryos. Nature 357, 255–257 (1992).
Huang, S., Shetty, P., Robertson, S. M. & Lin, R. Binary cell fate specification during C. elegans embryogenesis driven by reiterated reciprocal asymmetry of TCF POP-1 and its coactivator β-catenin SYS-1. Development 134, 2685–2695 (2007).
Sethi, A. et al. A caspase‐RhoGEF axis contributes to the cell size threshold for apoptotic death in developing Caenorhabditis elegans. PLoS Biol. 20, e3001786 (2022).
Mishra, N., Wei, H. & Conradt, B. Caenorhabditis elegans ced-3 caspase is required for asymmetric divisions that generate cells programmed to die. Genetics 210, 983–998 (2018).
Cordes, S., Frank, C. A. & Garriga, G. The C. elegans MELK ortholog pig-1 regulates cell size asymmetry and daughter cell fate in asymmetric neuroblast divisions. Development 133, 2747–2756 (2006).
Yushkevich, P. A., Gao, Y. & Gerig, G. ITK-SNAP: An interactive tool for semi-automatic segmentation of multi-modality biomedical images. Conf. Proc. IEEE Eng. Med. Biol. Soc., 3342-3345 (2016).
Hall, D. H. Gap junctions in C. elegans: Their roles in behavior and development. Dev. Neurobiol. 77, 587–596 (2017).
Pocock, R., Bénard, C. Y., Shapiro, L. & Hobert, O. Functional dissection of the C. elegans cell adhesion molecule SAX-7, a homologue of human L1. Mol. Cell. Neurosci. 37, 56–68 (2008).
Chisholm, A. D. Cytoskeletal dynamics in Caenorhabditis elegans axon regeneration. Annu. Rev. Cell Dev. Biol. 29, 271–297 (2013).
Guan, G., Kuang, X., Tang, C. & Zhang, L. Comparison between phase-field model and coarse-grained model for characterizing cell-resolved morphological and mechanical properties in a multicellular system. Commun. Nonlinear Sci. Numer. Simul. 117, 106966 (2023).
Walston, T. et al. Multiple Wnt signaling pathways converge to orient the mitotic spindle in early C. elegans embryos. Dev. Cell 7, 831–841 (2004).
Martino, F., Perestrelo, A. R., Vinarský, V., Pagliari, S. & Forte, G. Cellular mechanotransduction: From tension to function. Front. Phys. 9, 824 (2018).
Burridge, K., Monaghan-Benson, E. & Graham, D. M. Mechanotransduction: From the cell surface to the nucleus via RhoA. Philos. Trans. R. Soc. B, Biol. Sci. 374, 20180229 (2019).
Huang, X. T., Zhu, Y., Chan, L. H. L., Zhao, Z. & Yan, H. Inference of cellular level signaling networks using single-cell gene expression data in Caenorhabditis elegans reveals mechanisms of cell fate specification. Bioinformatics 33, 1528–1535 (2017).
Bavel, C., Thiels, W. & Jelier, R. Cell shape characterization, alignment, and comparison using FlowShape. Bioinformatics 39, btad283 (2023).
Murray, J. I. et al. Automated analysis of embryonic gene expression with cellular resolution in C. elegans. Nat. Methods 5, 703–709 (2008).
Pani, A. M., Gibney, T. V., Medwig-Kinney, T. N., Matus, D. Q. & Goldstein, B. A new toolkit to visualize and perturb endogenous lin-12/Notch signaling in C. elegans. microPubl. Biol. https://doi.org/10.17912/micropub.biology.000603 (2022).
Shaffer, J. M. & Greenwald, I. SALSA, a genetically encoded biosensor for spatiotemporal quantification of Notch signal transduction in vivo. Dev. Cell 57, 930–944.e6 (2022).
Bi, Y. et al. A genome-wide hybrid incompatibility landscape between Caenorhabditis briggsae and C. nigoni. PLoS Genet. 11, e1004993 (2015).
Still, E. & Hosemann, P. Alpha shape analysis: Extracting composition, surface area, and volume post clustering. Microsc. Microanal. 26, 2080–2082 (2020).
Ronneberger, O. et al. U-Net: Convolutional networks for biomedical image segmentation. Med. Image Comput. Comput.-Assist. Interv. 9351, 234–241 (2015).
Chen, C., Liu, X., Ding, M., Zheng, J. & Li, J. 3D dilated multi-fiber network for real-time brain tumor segmentation in MRI. Med. Image Comput. Comput. Assist. Interv. 11766, 184–192 (2019).
Van Der Walt, S. et al. scikit-image: Image processing in Python. PeerJ 2, e453 (2014).
Wang, A. et al. A novel deep learning-based 3D cell segmentation framework for future image-based disease detection. Sci. Rep. 12, 342 (2022).
Stringer, C., Wang, T., Michaelos, M. & Pachitariu, M. Cellpose: A generalist algorithm for cellular segmentation. Nat. Methods 18, 100–106 (2021).
Eschweiler, D., Smith, R. S. & Stegmaier, J. Robust 3d cell segmentation: Extending the view of Cellpose. 2022 IEEE International Conference on Image Processing - ICIP, 191-195 (2022).
Schmidt, U., Weigert, M., Broaddus, C. & Myers, G. Cell detection with star-convex polygons. Med. Image Comput. Comput. Assist. Interv. 11071, 265–273 (2018).
Weigert, M., Schmidt, U., Haase, R., Sugawara, K. & Myers, G. Star-convex polyhedra for 3D object detection and segmentation in microscopy. Proceedings of the IEEE/CVF Winter Conference on Applications of Computer Vision - WACV, 3655-3662 (2020).
Milletari, F., Navab, N. & Ahmadi, S.-A. V-Net: Fully convolutional neural networks for volumetric medical image segmentation. 2016 Fourth International Conference on 3D Vision - 3DV, 565-571 (2016).
Green, R. A. et al. A high-resolution C. elegans essential gene network based on phenotypic profiling of a complex tissue. Cell 145, 470–482 (2011).
Sönnichsen, B. et al. Full-genome RNAi profiling of early embryogenesis in Caenorhabditis elegans. Nature 434, 462–469 (2005).
Community, B. O. Blender – A 3D modelling and rendering package. https://www.blender.org (2023).
Schindelin, J. et al. Fiji: An open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Guan, G. et al. System-level quantification and phenotyping of early embryonic morphogenesis of Caenorhabditis elegans. Preprint on bioRxiv. https://doi.org/10.1101/776062 (2019).
Fernandez, R. et al. Imaging plant growth in 4D: Robust tissue reconstruction and lineaging at cell resolution. Nat. Methods 7, 547–553 (2010).
Xiao, L., Fan, D., Qi, H., Cong, Y. & Du, Z. Defect-buffering cellular plasticity increases robustness of metazoan embryogenesis. Cell Syst. 13, 615–630.e9 (2022).
Guan, G. Quantitative cell morphology reveals fate-coupled regulation of cell size in C. elegans embryo. figshare. https://doi.org/10.6084/m9.figshare.24768921 (2023).
Ye, P., Guan, G. & Li, Z. An online database for visualization and analysis of cell morphologies and gene expressions throughout Caenorhabditis elegans embryogenesis. EE Dept. of CityUHK. https://bcc.ee.cityu.edu.hk/cmos (2024).
Li, Z. & Guan, G. CMap additional data/files. figshare. https://doi.org/10.6084/m9.figshare.27872238 (2025).
Li, Z. & Guan, G. Source data underlying all figures, supplementary data, and supplementary movies of Paper: Cell lineage-resolved embryonic morphological map reveals novel signaling associated with cell fate and size asymmetry. figshare. https://doi.org/10.6084/m9.figshare.28467284 (2025).
Li, Z. & Cao, J. chiellini/CMap: Release CMap code for Nature Communications paper (v1.0). Zenodo https://doi.org/10.5281/zenodo.14916840 (2025).
Li, Z. & Cao, J. chiellini/CMapEvaluation: Release CMap evaluation code for Nature Communications paper (v0.9). Zenodo https://doi.org/10.5281/zenodo.14920949 (2025).
Boyle, T. J. et al. AceTree: A tool for visual analysis of Caenorhabditis elegans embryogenesis. BMC Bioinform. 7, 275 (2006).
The MathWorks Inc. MATLAB version: 9.5.0 (R2018b). https://www.mathworks.com (2018).
Python Software Foundation. Python Language Reference version: 3.7. http://www.python.org (2018).
Acknowledgements
We thank all the members of Yan Lab, Tang Lab, and Zhao Lab for the fruitful discussion and constructive comments. We appreciate the assistance of Dr. David Smith, Yixuan Chen, Qianli Zhang, Tongshu Wen, Yiqing Liu, and Cunmin Zhao for improving the paper materials, and Ahjol Hyrat for the preliminary analysis of C. elegans developmental symmetry breaking, Kai Kang for the preliminary analysis of gene expression profile and 3D illustration, and Dr. Hung Chim for the assistance in building stable server and website. We are grateful to Prof. Zhuo Du, Dr. Xuehua Ma, and Dr. Zhiguang Zhao for providing the summarized information on C. elegans cell identity and cell fate. Gratitude is extended to Prof. Robert Hugh Waterston and Prof. John Isaac Murray for instructing the data collection from EPIC. The computation of the quantitative and statistical analyses was performed in part on the High-Performance Computing Platform at Peking University. The strains used for the newly generated 97 embryos for profiling gene expression were provided by the Caenorhabditis Genetics Center (CGC), which is funded by the National Institutes of Health (NIH) Office of Research Infrastructure Programs (P40 OD010440). This work was supported by the National Natural Science Foundation of China (12090053, 32088101) to C.T.; by the Hong Kong Innovation and Technology Commission (ITC) (InnoHK Project CIMDA), Hong Kong Research Grants Council (RGC) (11204821), and the Institute of Digital Medicine of City University of Hong Kong (9229503) to H.Y.; and by the Hong Kong Innovation and Technology Commission (ITC) (GHP/176/21SZ) and Hong Kong Research Grants Council (RGC) (HKBU12101520, HKBU12101522, and HKBU12101323) to Z.Z.
Author information
Authors and Affiliations
Contributions
H.Y, C.T., and Z.Z. conceived and supervised this project; Y.M., M.-K.W., V.W.S.H, and L.-Y.C. cultured C. elegans, took and curated the images, and performed automated cell lineage tracing; Z.L. and J.C. devised the CMap algorithm, performed cell segmentation, and extracted morphological features; G.G., Z.L., and J.C. coordinated the manual annotation for ground truth, performed data quality evaluation and control, and organized the datasets; G.G. carried out the quantitative and statistical analyses; Z.L., G.G., and P.Y. generated the 3D illustration; G.G., Z.L., Y.M., P.Y., J.C., M.-K.W., and L.-Y.C. designed and improved the CVE software; P.Y., G.G., and Z.L. designed and improved the CMOS website; G.G., Z.L., Y.M., and P.Y. wrote the manuscript; H.Y, C.T., and Z.Z. revised the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Xiao Liu, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Guan, G., Li, Z., Ma, Y. et al. Cell lineage-resolved embryonic morphological map reveals signaling associated with cell fate and size asymmetry. Nat Commun 16, 3700 (2025). https://doi.org/10.1038/s41467-025-58878-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-58878-0