Abstract
Incidence of colorectal cancer (CRC) is increasing likely due to different mechanisms driving initiation and progression. The initial model proposed by Fearon and Vogelstein posits it as a multi-hit neoplasia, originating from adenomatous-polyps induced by WNT activation, ultimately progressing to aggressiveness through p53 loss. Integrating human data with mouse genetics, we redefine this paradigm, highlighting pivotal roles of MYC, oncogenic URI and p53 degradation to initiate CRC. Early APC loss activates MYC to transcriptionally upregulate URI, which modulates MDM2 activity, triggering p53 proteasomal degradation, essential for tumour initiation and mutation burden accrual in CRC mice. Remarkably, reinstating p53 levels via genetic URI depletion or p53 super-expression in CRC mice with WNT pathway activation prevents tumour initiation and extends lifespan. Our data reveal a “two-hit” genetic model central to APC loss-driven CRC initiation, wherein MYC/URI axis intricately controls p53 degradation, offering mechanistic insights into transitional mutation acquisition essential for CRC progression.
Similar content being viewed by others
Introduction
Comprehending the molecular mechanisms that underlie tumour initiation and progression is essential to identify targets for both cancer prevention and treatment. CRC, which stands as the third leading cause of cancer-related deaths and the third most frequently diagnosed cancer type as of 2022 (GLOBOCAN), has benefited from genetic studies involving patients with the hereditary form of the disease, enabling the identification of key events involved in tumour progression. The validation of these findings in genetic mouse models, which to some extent recapitulated the condition in humans, confirmed the involvement of key signalling pathways in CRC, contributing to the establishment of the step-wise model for tumour development, as proposed by Fearon and Vogelstein1 and still supported today (Supplementary Fig. 1a).
According to this model, a first step for tumour formation in CRC requires the aberrant activation of the WNT pathway, which typically results from mutations or loss of the tumour suppressor adenomatous polyposis coli (APC). APC protein is part of a multiprotein, destruction complex for β-catenin, serving as a crucial binding partner which under normal, steady-state conditions, assists in the phosphorylation of β-catenin and its subsequent ubiquitination, targeting it for degradation. Thus, upon APC mutations or loss, β-catenin degradation is impaired, and the protein is stabilized and translocated into the nucleus, where it activates the expression of WNT target genes, including MYC. Aberrant MYC expression promotes CRC by increasing overall proliferation and reducing differentiation. Given that most CRC tumours exhibit loss or mutations in APC, and loss of APC expression is sufficient to induce the formation of polyps in the intestine, the consensus model agrees that APC loss is critical for CRC initiation (Supplementary Fig. 1a).
Following the initial transformation, polyps undergo a series of sequential genetic and epigenetic mutations for their progression to more aggressive and malignant CRC. This progression involves acquiring mutations in the KRAS oncogene2, resulting in a constitutively active form of the protein that enhances cell proliferation and survival. In the ultimate stages, adenoma progression involves the loss of the tumour suppressor p53, believed to be a crucial factor in the transformation from adenoma to adenocarcinoma1,3. Yet, as individuals grow older, the function of p53 can become impaired due to the accumulation of mutations over time or to age-related decline in the cellular environment that affects p53 activity. This impairment allows damaged cells to survive and proliferate, increasing the likelihood of cancer development as the body ages4,5.
While the Fearon and Vogelstein’s model significantly contributes to understanding CRC progression1, it falls short in explaining the complex relationships among altered pathways in tumours, especially during tumorigenesis initiation and the transition to the development of aggressive tumours where the acquisition of molecular alterations in different genes and proteins is critical. Key unanswered questions in CRC research include identifying the trigger for the initial transformation and understanding the factors determining the transitional mutational burden essential for polyp progression into a malignant carcinoma.
In this study, we show that at early stages of APC loss-driven CRC, MYC overexpression transcriptionally upregulates the oncogene URI, which directly binds and enhances MDM2 activity, leading to p53 degradation via the proteasome. This degradation of p53 is crucial for CRC initiation and provides mechanistic insights into the acquisition of mutations necessary for CRC progression. Preventing these events inhibits tumour formation, suggesting that URI may serve as a therapeutic target in early CRC initiation.
Results
URI is overexpressed in APC-loss driven human CRC, correlating with poor prognosis
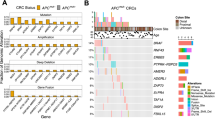
Searching for initiation factors driving WNT activation-based CRC, the oncogenic protein URI emerged as a compelling target that could have an impact in CRC development6. URI, encoded by URI1 gene, is an unconventional member of the prefoldin protein family and a key component of the URI prefoldin-like complex7,8. URI has been reported to exhibit oncogenic activities and seems to play a crucial role in the initiation of various cancers6,9. However, its role in the initiation of CRC remains to be addressed. Although analysis of genomic alterations in The Cancer Genome Atlas (TCGA) dataset revealed that URI1 was not frequently amplified in CRC and was neither mutated nor lost (Supplementary Fig. 1b), analysis of differential mRNA expression showed an upregulation of URI1 in CRC patients, as well as in other cancer types (Fig. 1a). Notably, URI1 was overexpressed in tumours compared with normal adjacent tissue (Fig. 1b and Supplementary Fig 1c). Accordingly, URI1 mRNA correlated negatively with patient survival in CRC samples from TCGA (Fig. 1c and Supplementary Fig. 1d). Additionally, immunohistochemistry analysis of URI protein levels in a tissue microarray of 230 human CRC samples confirmed its high expression in CRC tissue. Moreover, URI expression positively correlated with tumor grade and other disease progression markers of WNT activation-based CRC such as Ki67 and AXIN2, a target gene of β-catenin (Fig. 1d, e and Supplementary Fig. 1e). Analysis of genomic alterations in somatic driver genes associated with CRC revealed a correlation between URI1 mRNA levels and genetic aberrations in the most frequently mutated genes, including APC, TP53, and KRAS (Supplementary Fig. 1f), showing a significant positive association between high URI1 mRNA levels and APC loss or mutations, and with TP53 loss or mutations, but surprisingly not with KRAS mutations (Fig. 1f), suggesting that URI upregulation certainly plays a critical role in human WNT activation-based CRC development, possibly independently of KRAS mutations.
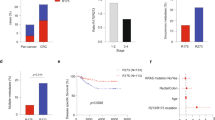
a URI1 mRNA levels in normal tissue (NT) and tumours from cancer patients, and fold change (FC) of expression in tumours compared to NT from TCGA samples. Stars denote the FDR-adjusted p. value. b URI1 mRNA in CRC tumours compared to matched normal tissue (n = 32). c Survival time in colorectal tumours expressing either high (n = 135) or low (n = 103) URI1 mRNA. Data are presented as mean ± SEM. d URI protein expression score from human CRC samples (n = 125), and its distribution according to tumour grade, Ki67 and AXIN2 scores, measured by IHC. e Association between Ki67 (left) and AXIN2 (right) expression scores with URI expression score. f Co-occurrences between high URI1 mRNA levels and APC mutations/loss, TP53 mutations/loss, KRAS mutations/gain in CRC samples from TCGA. g URI1 mRNA expression according to the consensus molecular subtype of CRC patients from TCGA. Odds ratio (OR) and p. value were calculated by fisher’s test. h Correlation between URI1 mRNA and patient survival in CMS2 patients. i Correlation between URI1 and MYC mRNA in CRC patients from TCGA. j Correlation between URI1 and CTNNB1 (upper) and AXIN2 (bottom) mRNA, in CRC patients from TCGA. k URI1 mRNA in human adenomas, serrated dysplasia and unclassified polyps (n = 28, 16, 11). l H&E and URI IHC of human adenomatous CRC polyps. m URI expression score in human colon pre-cancerous polyps and the adjacent non-polypoid mucosa (n = 75). n Detail of URI IHC of human adenomatous CRC polyp presenting non transformed and transformed cells with elongated nuclei. Dots represent individual patients. Statistical relevance of the correlations was assessed using Pearson correlation test. OR odds ratio, PCC Pearson correlation coefficient, p p. value. (***) = p. value < 0.005; (**) = p. value < 0.01. All statistical tests were two-sided. In linear correlations, grey shadow represents the 95% confidence interval of the linear regression fit. Boxplots represent Q1–Q3. Source data are provided as a Source Data file. Pictures of stainings are representative of the results obtained when staining and analysing all the samples available.
To corroborate these findings, we analysed URI1 mRNA expression across the different human consensus molecular subtypes (CMS) in which CRC patients have been classified, according to their gene expression patterns, and linked to either DNA mismatch repair deficiency and microsatellite instability (CMS1, MSI immune), WNT/MYC activation (CMS2, canonical), KRAS mutations (CMS3, metabolic), or TGF-β signalling (CMS4, mesenchymal)10. Consistent with our previous findings, using TCGA dataset, we showed that URI1 was specifically over-expressed in the CMS2 of CRC (Fig. 1g), and negatively correlated with patient survival in this specific subset but not in other CMSs (Fig. 1h and Supplementary Fig. 1g). URI could thus be an oncogene transcriptionally upregulated in CMS2, correlating with the progression of the disease.
Considering that URI expression correlated with CMS2 of CRC (WNT/MYC activation) and that MYC is a crucial transcription factor driving WNT activation-based CRC, we investigated whether a correlation exists between URI and MYC expression. Interestingly, human URI1 mRNA levels were positively correlated with high MYC and MYC-related genes in CRC from TCGA dataset (Fig. 1i, j and Supplementary Fig. 1h), suggesting that URI exhibits oncogenic activities in a human CMS2 model, likely upregulated by MYC expression, where MYC plays a critical role in its initial development11.
Accordingly, elevated URI1 expression was observed in the early stages of CRC, specifically in human adenomatous polyps as compared to serrated or unclassified dysplasia, based on analysis of publicly available RNA-sequencing (RNA-seq) data12 (Fig. 1k). These findings were further corroborated by immunohistochemistry (IHC) performed on human samples, which confirmed significantly higher URI expression in precancerous colon polyps (n = 75) compared to adjacent non-polypoid mucosa (Fig. 1l, m). Interestingly, URI was markedly increased in colonocytes displaying polarized, pseudostratified, elongated, and hyperchromatic nuclei, organized within simple, non-complex crypts; hallmarks of dysplastic epithelium and an early event in colon dysplasia13,14 (Fig. 1n). Moreover, URI was significantly upregulated in patients with ulcerative colitis (UC), a condition that precedes CRC15, (Supplementary Fig. 1i), as evidenced by RNA-seq analysis using the inflammatory bowel disease Transcriptome and Metatranscriptome Meta-Analysis (TAMMA) platform, which integrates data from 26 independent studies16. Notably, this upregulation was absent in patients with Crohn’s disease. Consistently, IHC revealed significantly higher URI expression in UC patients (n = 21), but not in Crohn’s disease, compared to healthy individuals (n = 10) (Supplementary Fig. 1j, k). Thus, URI expression may represent an early oncogenic event in CRC.
MYC transcriptionally upregulates URI
The above findings prompted us to investigate whether URI could be a direct target gene of MYC. Analysis of CpG islands, cis-regulatory elements from ENCODE, DNAse I hypersensitive clusters and H3K4Me3 regions in human URI1 locus revealed a promoter region, a potential upstream enhancer, and another upstream regulatory element (Fig. 2a). Unbiased analysis of available chromatin immunoprecipitation (ChIP)-seq data in ReMap database17 highlighted that MYC binds to both the promoter and the potential enhancer region of human URI1 (Fig. 2a), being the third most-frequently found transcription factor of these regions only following BRD4 and CTCF (Fig. 2b). This analysis also unveiled that the other upstream regulatory element is likely to be a CTCF binding region (Fig. 2b), being CTCF a general factor involved in chromatin remodelling18. Notably, the MYC partner MAX was also found among the top transcription factor hits able to bind to human URI1 promoter and enhancer regions, as well as n-MYC (Fig. 2b), another member of the MYC family of transcription factors mainly expressed in neurones and related to neuroblastoma19. Furthermore, two MYC binding motifs (CACGTG, also called E-box20,21) were found in human URI1 promoter, and another two in URI1 enhancer (Fig. 2c). A similar analysis of the Uri1 locus in mice unveiled a promoter region, a CTCF binding region and a potential enhancer (Supplementary Fig. 2a), and MYC was found among the top transcription factors binding to Uri1 promoter in mice (Supplementary Fig. 2b). ChIP followed by RT-qPCR analysis confirmed that MYC could bind to the URI1 promoter in human RKO CRC cells (Fig. 2d), and to the enhancer region (Fig. 2e). Expression of the URI1 promoter luciferase reporter, generated in our lab22, exhibited a 1.5-fold increase in luciferase signal following MYC overexpression in these cells (Fig. 2f), compared to control cells without MYC overexpression, suggesting that MYC directly binds to the URI1 promoter to regulate its expression. Consistently, silencing MYC with siRNA in human RKO CRC cells led to a decrease in URI protein levels (Fig. 2g). Altogether, these results demonstrate that URI expression, directly regulated by the proto-oncogene MYC, may exhibit oncogenic activity in the initiation of CRC.
a Genomic locus of human URI1 gene according to GENCODE v44. CpG islands, tri-methylation of histone H3 at lysine 4 (H3K4Me3) mark, DNAse I hypersensitivity peak clusters and ReMap density for transcription factors binding sites are showed, highlighting the genomic regions predicted to be promoters and enhancers of URI1. b Top 20 transcription factors found to bind to URI1 promoter and enhancer regions in ChIP-seq experiments from RefMap database. c MYC binding motifs found in URI1 promoter and enhancer regions. ChIP-qPCR analysis of MYC in URI1 promoter (d) and enhancer (e) regions in RKO CRC cell line (n = 5). Paired t-test analysis was used to analyse the statistical significance of the results. f Luciferase-reporter assay representing the luciferase normalized to the renilla activity in RKO human CRC cells transfected with either empty vector (pGL4.10) or URI1 promoter-luciferase cloned vector, and co-transfected or not with human MYC-expressing plasmid (n = 3). One-way ANOVA test and post-hoc tests with Tukey’s correction was used to compare the statistical differences among the means of different groups. g WB of MYC and URI protein levels in RKO human CRC cells after silencing of MYC with siRNA at increasing doses (20–80 nM). Vinculin is used as a protein loading control. Dots represent independent experiments. Bars represent the mean. Error bars represent the standard error of the mean (SEM). All statistical tests were two-sided. Source data are provided as a Source Data file.
APC loss induces CRC through MYC-mediated URI upregulation
To comprehensively investigate whether URI upregulation by MYC plays a central role in driving CRC pathogenesis, we used the Apc15lox mouse genetic model, where two loxP sites flank exon 15 of APC23. The flanking exon was recombined in 8 week-old mice by the tamoxifen-induced Cre-ERT2 recombinase expressed under the intestine-specific villin promoter (vil-Cre-ERT2 mouse model)24 (Fig. 3a, b), leading to a truncated form of the APC protein that does not fill its function23, which mimics intestinal APC loss in humans. Heterozygous Apc(+/15Δ)vil mice developed pre-neoplastic lesions, or small dysplastic areas, as early as 17 weeks of age, which progressed to visible and countable adenomas and adenocarcinomas in both small intestine and colon by 25 weeks of age (Fig. 3c, d and Supplementary Fig. 3a–c). Further characterization indicated that the 25-week-old mice exhibited significantly body weight loss and had splenomegaly and anaemia (Fig. 3e–h), with a spleen weight that correlated with the number of tumours (Fig. 3i). Thus, Apc(+/15Δ)vil mice mimic initial features of human CRC and represent a noteworthy CMS2-like model to study the initiation and the progression of the disease.
a Scheme of the vil-Cre-ERT2; Apc15lox mouse model. b Experimental design for tumorigenesis in this model. c Representative pictures of small intestine and colon of Apc(+/15Δ)vil mice harbouring tumours. d H&E stainings of mouse intestinal tissue showing representative adenoma low grade or grade 2, adenoma high grade or grade 3, and adenocarcinoma or grade 4. Body weight (e), spleen weight (f), spleen weight normalized to body weight (g), red blood cells, haematocrit, and haemoglobin levels at time of sacrifice, as measures of the anaemia levels (h) of Apc(+/15Δ)vilmice compared to Apc(+/+)vil mice (n = 22, 16). i Correlation between spleen weight and number of tumours in the Apc(+/15Δ)vil mice. Blue line represents the linear regression fit and grey shadow represents the 95% confidence interval of the linear regression fit. j IF staining showing URI expression in tumours and small dysplastic areas from Apc(+/15Δ)vil mice. k WB of URI and MYC protein levels in the intestines of Apc(+/15Δ)vil mice compared to Apc(+/+)vil mice. l ChIP-qPCR analysis of MYC in Uri1 promoter and enhancer regions in tumours from Apc(+/15Δ)vil mice (n = 3, 4, 4, 3). Bars represent the mean. Error bars represent the standard error of the mean. Dots represent individual mice. T. test was used to compare the statistical differences between means of two different groups. p p. value. (***) = p. value < 0.005. All statistical tests were two-sided. Source data are provided as a Source Data file.
Immunofluorescence staining demonstrated that URI was elevated in preneoplastic lesions and early tumours derived from Apc(+/15Δ)vil mice (Fig. 3j and Supplementary Fig. 3d, e). Moreover, Western blot analysis showed that URI was upregulated at later stages in the intestinal tissue of 25-week-old Apc(+/15Δ)vil mice, together with the expected increase in MYC protein levels (Fig. 3k), a critical mediator in the early stages of neoplasia following APC loss11. These results confirm the causal relationship between MYC and URI expression during early tumorigenesis.
Importantly, URI was detected in mouse intestinal crypts, as shown by immunofluorescence (Supplementary Fig. 3e), following a similar expression pattern to MYC11. Co-immunostaining further revealed that MYC and URI were co-expressed in mouse intestinal crypts (Supplementary Fig. 3f), confirming their spatial and causal relationship in vivo. Genetic depletion of MYC in the mouse intestinal epithelium using the floxed Myc mouse model25 crossed with vil-Cre-ERT2 mice under tamoxifen treatment (Supplementary Fig. 3g, h) caused a decrease in URI expression (Supplementary Fig. 3i). Consistently, ChIP-qPCR analysis demonstrated that MYC could bind to Uri1 promoter and enhancer regions in tumours from Apc(+/15Δ)vil mice (Fig. 3l). Thus, oncogenic MYC mediates URI overexpression in human and mouse APC loss-driven CRC.
To corroborate that MYC upregulation is a critical mediator of CRC in our model, we genetically depleted MYC in Apc(+/15Δ)vil mice by crossing them with the floxed Myc model25 (Supplementary Fig. 3j, k). Expectedly, MYC depletion diminished the number of tumours (Supplementary Fig. 3l, m) and prevented body weight loss of Apc(+/15Δ)vil mice (Supplementary Fig. 3n), although mice still presented splenomegaly and anaemia (Supplementary Fig. 3o–q), as previously reported11. Taken together, our data demonstrate that the initiation of CRC is driven by oncogenic MYC-mediated URI expression.
Oncogenic URI modulates p53 protein stabilization
Prompted by these findings and to further study the role of URI in the initiation steps of CRC, we have ablated URI specifically in intestinal epithelium by crossing the conditional URI knockout mice (URIlox mice)26 with vil-Cre-ERT2 mice (Fig. 4a, b). Notably, complete URI loss in intestinal epithelium led to mouse death within a few days after activation of the Cre-ERT2 recombinase by tamoxifen in 8 weeks-old mice26. We thus evaluated the phenotype of URI heterozygous mice, referred to as Uri(+/Δ)vil mice. Checking for molecular pathways that could be affected by URI loss, we directed our attention to the tumor suppressor p53, given that the association between URI and p53 has been previously confirmed in the intestinal epithelium of Urivil mice26 and in human data (Fig. 1). At 25 weeks of age, Uri(+/Δ)vil mice exhibited elevated p53 at protein but not mRNA levels in intestinal crypts when compared to their littermate controls Uri(+/+)vil mice, as shown by Western blot and IHC analysis (Fig. 4c–e). Co-IHC staining demonstrated that URI and p53 co-localized in the mouse intestinal crypts (Fig. 4f). Importantly, the protein levels of the E3 ubiquitin-protein ligase Mouse double minute 2 homolog (MDM2), a key negative regulator of p53 responsible for its ubiquitination and subsequent degradation, remained unchanged upon URI loss in mouse intestine (Fig. 4c). Notably, the increase in p53 protein levels came together with enhanced cell cycle arrest markers, including increased p21 levels (Fig. 4c, g), while other p53-related pathways such as apoptosis, senescence or DNA damage repair where not increased but rather diminished in intestines of 25 week-old Uri(+/Δ)vil mice as shown by Western blot, immunohistochemistry and RT-qPCR analysis (Supplementary Fig. 4a–c). Thus, cell proliferation arrest rather than increased apoptosis may result from enhanced p53 upon URI loss. Additionally, in aged mice, approximately 20% of mice, regardless of their genotypes, developed spontaneous tumours in their intestine, identified as intestinal adenomas based on histopathological analysis (Supplementary Fig. 4d–g).
a Scheme of the vil-Cre-ERT2; URIlox mouse model. b Experimental design used in these mice. c WB analysis of p53, p21 and MDM2 protein levels in the small intestine from URI(+/+)vil and URI(+/Δ)vil mice. d Relative Tp53 mRNA levels in URI(+/Δ)vil mice intestines (n = 13) compared to URI(+/+)vil mice (n = 11). e IHC staining of p53 in mouse intestinal tissue from URI(+/+)vil and URI(+/Δ)vil mice. f Co-IHC staining of URI and p53 in mouse intestinal tissue. g Relative mRNA levels of the p53-target gene 14-3-3-σ involved in cell-cycle regulation (n = 6, 9). h Scheme of the villin-rtTA; ColA1-tetO-hURI mouse model for human URI overexpression in the intestinal epithelium. i Experimental design for URI overexpression in these mice. j WB analysis of p53, p21 and MDM2 protein levels in the small intestine from hURI(+/+)vil and hURI(+/KI)vil mice. k Relative Tp53 mRNA levels in hURI(+/KI)vil mice intestines compared to hURI(+/+)vil mice (n = 7, 8). l IHC staining for DNA damage marker γ–H2AX in mouse intestinal tissue from hURI(+/+)vil and hURI(+/KI)vil mice (n = 74, 155 crypts). In WB analysis, vinculin was used as a protein loading control. In qPCR analysis, Tbp was used as a housekeeping gene to normalize the values. Bars represent the mean. Error bars represent the standard error of the mean. Dots represent individual mice. T. test was used to compare the statistical differences between means of two different groups. p p. value. (*) = p. value < 0.05; (***) = p. value < 0.005. All statistical tests were two-sided. Source data are provided as a Source Data file.
These observations were mirrored in vivo using a mouse model for intestine-specific human URI overexpression upon doxycycline treatment22,26, refereed here as hURI(+/KI)vil mice (Fig. 4h, i and Supplementary Fig. 4h). hURI(+/KI)vil mice exhibited substantially lower levels of p53 protein, but not mRNA levels, with no changes in MDM2 levels, in their intestinal epithelium compared to their littermate control hURI(+/+)vil mice (Fig. 4j, k). This decrease in p53 protein levels came along with a decrease in different p53 targets, in particular the ones regulating cell cycle (Fig. 4j and Supplementary Fig. 4i, j) and, consistently, with increased DNA damage in the intestinal epithelium (Fig. 4l). Interestingly, no spontaneous tumours were detected in aged hURI(+/KI)vil mice (Supplementary Fig. 4k, l), suggesting that URI overexpression alone might not be sufficient to cause tumour formation in the intestine.
Oncogenic URI regulates p53 stabilization by modulating MDM2 activity
Building on these observations, next we explored mechanisms regulating p53 levels in the absence of URI. Given that tp53 mRNA levels and MDM2 protein levels remained unchanged after URI loss, we examined whether URI regulated p53 at the protein level in CRC cell lines. Interestingly, overexpression of hemagglutinin (HA) tagged-URI in human CRC cells resulted in a decrease in p53 protein levels, which was restored when proteasome activity was inhibited with MG132 (Fig. 5a), but not when inhibiting autophagy with chloroquine (Supplementary Fig. 5a), suggesting that URI regulates p53 protein levels by modulating its proteasomal degradation. Accordingly, overexpression of HA-URI reduced the half-life of p53, as demonstrated by time-dependent cycloheximide treatment in human CRC cells (Fig. 5b, c). Importantly, immunoprecipitation of p53 followed by ubiquitination blotting revealed that overexpression of HA-URI in human CRC cells increased p53 ubiquitination, whereas silencing of URI by siRNA decreased it (Fig. 5d). Thus, URI regulates p53 protein levels by promoting its ubiquitination.
a WB analysis of p53 protein levels and HA-tagged URI upon either empty pcDNA3 vector or HA-URI transfection in p53 wild-type LIM1215 CRC cells, treated with the proteasome inhibitor MG132 or with DMSO as control. b WB analysis of p53, vinculin and HA-URI levels in HA-URI overexpressing cells and pcDNA3 control HCT-116 CRC cells over time after treatment with the translation inhibitor cycloheximide (CHX). c Quantification of p53 protein levels normalized to vinculin over time after CHX treatment in the indicated cells. Dashed line represents the polynomic model of p53 degradation over time. n = 4 independent experiments. Two-way ANOVA was used to assess the statistical significance of the results. d WB analysis of p53 ubiquitination in control, HA-URI overexpressing and siURI-mediated silenced RKO CRC cells. e Co-immunoprecipitation (Co-IP) followed by WB analysis of URI, p53 and MDM2 in HCT-116 CRC cells. f MDM2-mediated p53 ubiquitination in vitro assay with increased concentrations of GST-tagged URI (5, 10 and 30 µL). s.e short exposure, l.e. long exposure. g Co-IP analysis of URI and proteasome subunit PSMB4 and p53 proteins performed in RKO CRC cells. h Co-IP analysis of PSMB4 and p53 HCT-116 CRC cells. WB are representative of three independent experiments. Source data are provided as a Source Data file.
Further experiments to investigate the underlying mechanisms revealed that URI, p53, and MDM2 reciprocally co-immunoprecipitated in CRC cells (Fig. 5e), suggesting the existence of a trimeric complex independently of TRIM28, which was previously demonstrated to promote p53 ubiquitination in HCC cell lines27 (Supplementary Fig. 5b). Direct URI binding to p53 and MDM2 was therefore tested by incubating bacterially-expressed glutathione S-transferase (GST)-tagged URI and URI fragments (Supplementary Fig. 5c, d) with recombinant p53 and MDM2 proteins. Pulldown assays revealed that URI interacts with MDM2 at its N-terminal region, where the prefoldin domain is located, while p53 binds to the middle portion of URI, specifically between the amino acids 158 and 390 (Supplementary Fig. 5e–g). Downregulation of URI did not disrupt the interaction between p53 and MDM2 (Supplementary Fig. 5h). Moreover, MDM2 protein levels remained unchanged upon URI loss or overexpression in human CRC cell lines (Supplementary Fig. 5i), suggesting that URI promotes p53 ubiquitination independently of modulation of MDM2 protein or mRNA levels. Altogether, these data suggest that URI binds to both MDM2 and p53, forming a functional trimeric complex that potentially regulates p53 ubiquitination through modulation of MDM2 activity. We thus investigated whether p53 ubiquitination could instead be attributed to the regulation of MDM2 activity by URI. To this end, we established an in vitro MDM2 ubiquitination activity assay using recombinant Histidin-tagged MDM2 (full-length), and which incubation with increased concentrations of recombinant GST-URI (full-length) caused a marked increase of p53 ubiquitination in vitro in a dose-dependent-manner (Fig. 5f). Thus, URI promotes p53 ubiquitination by directly binding and enhancing MDM2 activity, likely targeting it to proteasomal degradation.
Consistent with these findings and considering that MG132 restored p53 levels upon URI overexpression, URI was found to co-immunoprecipitate with the proteasomal subunit PSMB4 (Fig. 5g), which also co-immunoprecipitated with other proteasomal subunits like PSMD11 and PSMB2 (Supplementary Fig. 5j) as well as with p53 (Fig. 5h), further supporting the relationship between URI, MDM2 and proteasomal-mediated p53 protein degradation. Together, these data suggest that URI overexpression increases MDM2 activity, triggering p53 ubiquitination and degradation via the proteasome, potentially leading to CRC initiation.
URI depletion prevents APC loss-driven CRC initiation in a p53-dependent manner
Given the relationship between URI expression and p53 degradation, next we checked p53 protein levels in CMS2-like mice with high MYC and URI levels. Strikingly, a significant reduction in p53 levels was detected in intestines and colons of 25-week-old Apc(+/15Δ)vil mice (Fig. 6a and Supplementary Fig. 6a). Downregulation of p53 was accompanied by a reduction in p21 levels but consistently not MDM2 protein levels (Fig. 6a). To assess whether p53 loss is responsible for CRC initiation, we restored p53 downregulation by breeding Apc(+/15Δ)vil mice with Super p53 mice, which carry additional copies of the p53 gene and exhibit tumour resistance28 (Fig. 6b, c). Super-expression of p53 in Apc(+/15Δ)vil mice significantly reduced tumour burden, including decreased tumour number and area, together with a reduced splenomegaly (Fig. 6d–g), confirming that abolishing p53 protein downregulation suppresses tumour initiation.
a WB of p53, p21 and MDM2 in intestines from indicated mice. b Scheme of vil-Cre-ERT2; Apc15lox; Super-p53 mouse model. c Experimental design for tumorigenesis. d H&E of tumours in Apc(+/15Δ)vil; Super p53(+/+) and Apc(+/15Δ)vil; Super p53(+/Tg) mice. e Number of intestinal tumours in these mice (n = 12, 14). f Area of each individual tumour. g Spleen weight of these mice. h Scheme of vil-Cre-ERT2; Apc15lox; URIlox; p53lox/lox mouse models. i Experimental design for tumorigenesis. j WB of p53, p21 and MDM2 in intestines from Apc(+/15Δ)vil; URI(+/+)vil and Apc(+/15Δ)vil; URI(+/Δ)vil mice. k H&E of intestinal tumours in the indicated mice. l Number of intestinal tumours in the indicated mice (n = 27, 20, 7, 13). m Total tumour area per mouse in these models. n Distribution of areas of the tumours. o Maximum tumour grade reached per mouse. p Overall looking of the mice. q Spleen weight of the mentioned mice. r Kaplan-Meier curve indicating survival time of the corresponding mice. s H&E of small dysplastic areas, indicated with (*), from Apc(+/15Δ)vil; p53(Δ/Δ)vil mice. t Number of small dysplastic areas found in the indicated mice. u Scheme of vil-Cre-ERT2; Apc15lox; villin-rtTA; ColA1-tetO-hURI mouse model. v Experimental design for tumorigenesis. w H&E of the tumours in Apc(+/15Δ)vil; hURI(+/+)vil and Apc(+/15Δ)vil; hURI(+/KI)vil mice. x Number of intestinal tumours found in these mice (n = 34, 19). y Area of tumours in the indicated mice. z WB of p53, p21 and MDM2 in intestines from Apc(+/15Δ)vil; hURI(+/+)vil and Apc(+/15Δ)vil; hURI(+/KI)vil mice. In WB analysis, vinculin was used as a protein loading control. Bars represent the mean ± SEM. Dots represent individual mice, and individual tumours (g, n, y). Boxplots represent Q1–Q3. One-way ANOVA test and post-hoc tests with Tukey’s correction was used to compare the statistical differences among the means of different groups. Two-sided t. test was used to compare the statistical differences between means of two different groups. p p. value. (*) = p. value < 0.05; (**) = p. value < 0.01; (***) = p. value < 0.005. Source data are provided as a Source Data file.
Similarly, we checked if URI downregulation in Apc(+/15Δ)vil mice restored p53 protein levels. To this end, we bred Apc15lox and URIlox mice together with vil-Cre-ERT2 mice (Fig. 6h, i), generating Apc(+/15Δ)vil; Uri(+/Δ)vil mice. Remarkably, genetic decrease of URI in Apc(+/15Δ)vil mice reinstated p53 and p21 protein levels, but as expected not MDM2, in their intestines and colons when compared to Apc(+/15Δ)vil; Uri(+/+)vil littermate controls (Fig. 6j and Supplementary Fig. 6b). Accordingly, Apc(+/15Δ)vil; Uri(+/Δ)vil mice displayed significantly reduced tumour number, size, aggressiveness, and splenomegaly; and their probability of survival was improved compared to Apc(+/15Δ)vil; Uri(+/+)vil mice (Fig. 6k–r). These data indicate that p53 downregulation occurs in the early steps of tumour formation, likely driven by increased MYC-URI axis. Counteracting this event by downregulating oncogenic URI efficiently prevents APC loss-driven CRC development.
To evaluate whether the beneficial effects of URI downregulation on tumorigenesis initiation were dependent on p53, we genetically eliminated p53 in the mouse intestinal epithelium of Apc15lox and URIlox mice by crossing them with conditional p53 knockout (p53lox) mice29 and vil-Cre-ERT2 mice (Fig. 6h, i). The beneficial effects of URI downregulation observed in Apc(+/15Δ)vil; Uri(+/Δ)vil mice, including reducing tumour burden and splenomegaly, were completely nullified in the absence of p53 (Fig. 6k–r). This indicates that the prevention of APC loss-driven CRC initiation mediated by URI downregulation is p53 dependent. Notably, homozygous p53 depletion in the intestinal epithelium of Apc(+/15Δ)vil mice led to the appearance of multiple small dysplastic areas (Fig. 6s, t), which further indicates that p53 downregulation occurs in the early stages of APC loss-driven tumour formation.
As expected, overexpression of URI in the intestinal epithelium of Apc(+/15Δ)vil by crossing them with hURI(+/KI)vil mice resulted in increased number of tumours (Fig. 6u–x) and in particular, of small dysplastic areas (Fig. 6y), resembling p53 depletion in this mouse model. Expectedly, p53 levels in Apc(+/15Δ)vil; hURI(+/KI)vil mice were further decreased compared to Apc(+/15Δ)vil; hURI(+/+)vil control mice, along with a decrease in p21 but not with MDM2 (Fig. 6z). Importantly, proliferation of intratumoral cells was similar across the different groups of mice (Supplementary Fig. 6c, d). However, an increase in p53 levels, resulting from both super-p53 expression or URI downregulation, tended to reduce cell proliferation, whereas URI overexpression and decreased p53 levels enhanced the percentage of Ki67 positive cells, in line with our previous findings (Supplementary Fig. 6c, d). Differences in tumour burden among the different groups of mice could not be attributed to apoptosis, since modulation of URI and p53 protein levels did not induce apoptosis in tumours from Apc(+/15Δ)vil mice (Supplementary Fig. 6c). These data further demonstrate that URI-mediated degradation of p53 impacts the initiation phase of tumorigenesis in the APC loss background by modulating cell proliferation. Hence, URI upregulation in APC loss-driven CRC mouse model promotes tumor initiation, in a p53-dependent manner.
p53 barrier prevents CRC in a β-catenin-dependent manner
To determine whether the p53 barrier against CRC is also established in the presence of activated β-catenin, we employed a conditional β-catenin–floxed mouse strain, Catnblox(ex3) mouse, in which exon 3 of the β-catenin gene is flanked by two loxP sites30. Recombination of these sites, facilitated by the tamoxifen-induced Cre-ERT2 recombinase, results in a mutated version of the β-catenin protein where the phosphorylation sites by GSK3β are eliminated, leading to the stabilization and activation of β-catenin, ultimately causing polypoid dysplasia30. Catnblox(ex3) mice were crossed with vil-Cre-ERT2 mice to specifically induce recombination in the intestinal epithelium of 8 weeks-old mice, generating Catnb+/Δ(ex3)vil mice (Supplementary Fig. 7a, b). As expected, the intestinal epithelium of these mice hyperproliferated, showing very evident signs of dysplasia especially in the small intestine (Supplementary Fig. 7c–e). Despite no significant body weight lost, Catnb+/Δ(ex3)vil mice presented splenomegaly and signs of anaemia (Supplementary Fig. 7f–i). Histopathological analysis of the tissue revealed that dysplastic tissue, both polypoid and non-polypoid, was found in these mice, which also showed high grade dysplasia (Supplementary Fig. 7j). Importantly, overactivation of β-catenin in intestinal epithelium caused a significant reduction in p53 and p21 protein levels (Fig. 7a), but not in MDM2, mirroring the Apc(+/15Δ)vil mice. Ectopic expression of p53 in Catnb+/Δ(ex3)vil mice by crossing them with Super p53 mice (Fig. 7b, c) successfully reduced the percentage of high-grade dysplastic tissue (Fig. 7d–f). These findings suggest that p53 plays a role in preventing tumorigenesis in the β-catenin overactivation model.
a WB of p53, p21 and MDM2 protein levels in intestines from Catnb+/Δ(ex3)vil mice compared to Catnb+/+(ex3)vil mice. b Scheme of the vil-Cre-ERT2; Catnblox(ex3); Super p53 mouse model. c Experimental design for tumorigenesis in these mice. d H&E staining showing the dysplastic areas in the intestine of Catnb+/Δ(ex3)vil; Super p53(+/+) mice compared to Catnb+/Δ(ex3)vil; Super p53(+/Tg) mice. e Quantification of the percentage of tissue with dysplasia and with polypoid dysplasia (n = 18, 10). f Percentage of high-grade dysplastic tissue in the indicated mice. g WB analysis of URI protein levels in Catnb+/Δ(ex3)vil mice compared to Catnb+/+(ex3)vil mice. h IF staining of URI expression in normal and dysplastic intestinal tissue from Catnb+/Δ(ex3)vil mice. i Scheme of vil-Cre-ERT2; Catnblox(ex3); URIlox; p53lox/lox mouse models. j Experimental design for tumorigenesis in these mice. k H&E staining of the intestinal architecture of the indicated mice. l Percentage of tissue with dysplasia and polypoid dysplasia (n = 50, 34, 5, 11). m Percentage of high-grade dysplastic tissue in the indicated mice. n Kaplan-Meier curve representing the survival time (in days) of the indicated mice. o WB analysis of p53, p21 and MDM2 protein levels in Catnb+/Δ (ex3)vil; URI(+/+)vil mice compared to Catnb+/Δ (ex3)vil; URI(+/Δ)vil mice. p Kaplan-Meier curve representing the survival time (in days) of the indicated mice. In WB analysis, vinculin was used as a protein loading control. Bars represent the mean. Error bars represent the standard error of the mean. Dots represent individual mice. Two-sided t. test was used to assess the statistical differences between the means of two groups. One-way ANOVA test and post-hoc tests with Tukey’s correction was used when comparing the statistical differences among the means of more than two groups. p p. value. (*) = p. value < 0.05. Source data are provided as a Source Data file.
As seen in Apc(+/15Δ)vil mice, URI protein levels were also upregulated in intestines of Catnb+/Δ(ex3)vil mice (Fig. 7g, h). Thus, we evaluated if p53 upregulation caused by URI downregulation could also prevent dysplasia in these mice (Fig. 7i, j). Accordingly, Catnb+/Δ(ex3)vil; Uri(+/Δ)vil mice showed a lower percentage of high-grade dysplastic tissue than their littermates control Catnb+/Δ(Ex3)vil; Uri(+/+)vil; with only a minor reduction in the percentage of polypoid and non-polypoid dysplasia (Fig. 7k–m). Moreover, decreasing URI levels in Catnb+/Δ(ex3)vil mice increased the survival probability of mice (Fig. 7n).
To investigate if the beneficial effects of URI downregulation were also dependent on p53 in Catnb+/Δ(Ex3)vil; URI(+/Δ)vil mice, as in the APC loss-driven CRC model previously discussed, we corroborated that the expression of p53 and p21 proteins, but consistently not MDM2, were retained in their intestinal epithelium, compared to their littermates Catnb+/Δ(ex3)vil; Uri(+/+)vil controls (Fig. 7o). We therefore genetically depleted p53 in Catnb+/Δ(ex3)vil; Uri(+/Δ)vil mice (Fig. 7i, j). In the absence of p53, URI downregulation failed to prevent the high-grade dysplasia (Fig. 7k–m) and to improve mouse survival (Fig. 7p) in the β-catenin overactivation model. Therefore, URI prevents initiation of high-grade intestinal dysplasia that arises from the overactivation of β-catenin in a manner dependent on p53 loss-mediated hyperproliferation.
KRAS activation occurs after p53 protein degradation contributing to aggressiveness
According to the current stepwise molecular model for CRC development, as proposed by Fearon and Vogelstein1, APC loss triggers MYC-dependent cell proliferation, followed by KRAS oncogene mutations, promoting the transition from adenomas to aggressive adenocarcinomas1,2 (see Supplementary Fig. 1a). To investigate the interplay between the APC-β-catenin-MYC pathway and KRAS, we crossed the conditional KRAS+/V12 mice31, with a commonly found mutation in CRC (Fig. 8a), with vil-Cre-ERT2 mice to induce oncogenic KRASV12 protein in intestinal epithelium after tamoxifen treatment (Fig. 8b, c). Oncogenic KRASV12 expression in the intestinal epithelium did not result in spontaneous tumour formation (Fig. 8d), consistent with previous studies32. However, mice with oncogenic KRASV12 expression in the APC loss background developed tumours in the intestinal epithelium (Fig. 8e, f), demonstrating that KRASV12 mutations occur after loss of APC in CRC development. Interestingly, oncogenic KRASV12 expression biased the phenotype towards the colon instead of the small intestine (Fig. 8g, h).
a Frequency of mutations in codons 12 and 13 of KRAS protein found in CRC patients (n = 223). b Scheme of the vil-Cre-ERT2; KRAS+/V12; Apc15lox; p53lox/lox; URIlox mouse model. c Experimental design for tumorigenesis in these mice. d H&E staining showing the intestinal tissue found in KRAS(+/V12)vil mice;; KRAS(+/V12)vil; URI(+/Δ)vil mice; KRAS(+/V12)vil; p53(Δ/Δ)vil mice; and KRAS(+/V12)vil; p53(Δ/Δ)vil; URI(+/Δ)vil mice. e H&E staining showing the intestinal tumours found in Apc(+/15Δ)vil; KRAS(+/V12)vil mice; Apc(+/15Δ)vil; KRAS(+/V12)vil; URI(+/Δ)vil mice; Apc(+/15Δ)vil; KRAS(+/V12)vil; p53(Δ/Δ)vil mice; and Apc(+/15Δ)vil; KRAS(+/V12)vil; p53(Δ/Δ)vil; URI(+/Δ)vil mice. f Number of tumours found in these mice (n = 15, 12, 25, 7). g Representative picture of a colon harbouring tumours from a Apc(+/15Δ)vil; KRAS(+/V12)vil mouse. Arrows indicate tumours. h Distribution of intestinal tumours in the small intestine and colon of Apc(+/15Δ)vil and Apc(+/15Δ)vil; KRAS(+/V12)vil mice. i Maximum tumour grade reached in the indicated mice. j Kaplan-Meier curve representing the survival time (in days) of the indicated mice. k H&E staining showing liver metastasis found in two Apc(+/15Δ)vil; KRAS(+/V12)vil; p53(Δ/Δ)vil mice. Bars represent the mean. Error bars represent the standard error of the mean (SEM). Dots represent individual mice. One-way ANOVA test and post-hoc tests with Tukey’s correction was used when comparing the statistical differences among the means of the three groups. p p. value. (***) = p. value < 0.005; (**) = p. value < 0.01; (*) = p. value < 0.05. Source data are provided as a Source Data file.
We then investigated if URI loss-mediated p53 upregulation could prevent tumour formation in this model by crossing these mice with URI(+/Δ)vil mice. Notably, Apc(+/15Δ)vil; KRAS(+/V12)vil; URI(+/Δ)vil mice showed a decreased number of tumours in the intestine, as previously shown in Apc(+/15Δ)vil mice, but the grade of the tumours and the survival of mice were not affected (Fig. 8a–e). These data support the model where URI upregulation followed by APC loss and MYC overexpression initiates CRC, and that mutations in KRAS occur later in the transformation process.
Additionally, genetic deletion of p53 in this model aggravated the observed phenotype, since mice died before the established time point, and some mice even developed liver metastases (Fig. 8a–k). Expectedly, reduction of URI levels failed to prevent tumorigenesis in these mice with genetic depletion of p53 in the intestinal epithelium (Fig. 8a–j). As expected, KRASV12 expression together only with p53 depletion was insufficient to induce tumorigenesis (Fig. 8d), further corroborating that MYC activation triggered by APC-β-catenin axis is the essential first event in CRC onset. Thus, while KRAS activation drives aggressiveness and metastasis in mice (Fig. 8), its combination with APC loss in mice better models advanced human disease stages compared to APC loss alone. Such combined models more accurately mimic the advanced stages of the disease.
URI depends on p53 to promote tumour initiation
After demonstrating that URI degrades p53 to initiate the transformation process in APC loss-driven CRC, we examined whether URI could also play an important role in the initiation process of intestinal tumours arising from p53 loss, independently of APC mutations. To that end, we depleted p53 in the intestinal epithelium by crossing p53lox mice29 with vil-Cre-ERT2 mice and induced recombination in 8 weeks-old mice, generating p53(Δ/Δ)vil mice (Fig. 9a, b). Specific homozygous p53 depletion in the intestinal epithelium gave rise to tumours in around 25% percent of mice at 50 weeks of age (Fig. 9c–e). Importantly, these tumours did not express URI (Fig. 9f), and depletion of MYC did not prevent tumorigenesis in this model (Fig. 9a–e), demonstrating that tumours arising in the p53 knockout background alone are independent of the MYC-URI axis. In line with these findings, reduction of URI levels in p53(Δ/Δ)vil mice failed to reduce tumorigenesis and to extend their lifespan (Fig. 9g–m). Similarly, genetic overexpression of URI in p53(Δ/Δ)vil mice by crossing them with hURI(+/KI)vil mice had no impact on tumour burden in these mice (Fig. 9n–r).
a Scheme of the vil-Cre-ERT2; p53lox/lox; Myclox/lox mouse models. b Experimental design for the tumorigenesis in these mice. c H&E of the intestinal tumours found in p53(Δ/Δ)vil; Myc(+/+)vil mice; and p53(Δ/Δ)vil; Myc(Δ/Δ)vil mice. d Percentage of mice presenting tumours in these models. e Number of tumours found in the indicated mice (n = 8, 17). f IF staining of URI in a tumour from p53(Δ/Δ)vil; Myc(+/+)vil mice. g Scheme of the vil-Cre-ERT2; p53lox/lox; URIlox mouse model. h Experimental design for tumorigenesis in these mice. i Pictures of H&E staining of intestinal tumours found in p53(Δ/Δ)vil; URI(+/+)vil and p53(Δ/Δ)vil; URI(+/Δ)vil mice. j Proportion of mice presenting intestinal tumours. k Number of intestinal tumours found in these mice (n = 8, 34). l Experimental design for ageing experiments in these mice. m Kaplan-Meier curve indicating survival probability and time (in days) of the corresponding mice. n Scheme of the vil-Cre-ERT2; p53lox/lox; villin-rtTA; ColA1-tetO-hURI mouse model. o Experimental design for the tumorigenesis in these mice. p Pictures of H&E staining of intestinal tumours found in p53(Δ/Δ)vil; hURI(+/+)vil mice and p53(Δ/Δ)vil; hURI(+/KI)vil mice. q Proportion of mice presenting intestinal tumours. r Number of intestinal tumours found in these mice (n = 24, 8). s Scheme of the hUb-CreERT2; p53lox/lox; URIlox mouse model. t Experimental design for tumorigenesis in these mice. u Kaplan-Meier curve indicating survival probability and time (in days) of p53(Δ/Δ)Ub; URI(+/+)Ub and p53(Δ/Δ)Ub; URI(+/Δ)Ub mice. v Percentage of mice presenting tumours in the indicated tissues. DIC = mouse dead in cage. Bars represent the mean, except in (v). Error bars represent the standard error of the mean (SEM). Dots represent individual mice. Two-sided t. test was used to compare the statistical differences between means of two different groups. Source data are provided as a Source Data file.
To further explore the relationship between URI and p53 in tumour initiation, we used a mouse model for carcinogenesis based on p53 depletion in the whole body. Homozygous deletion of p53, achieved by crossing p53lox29 mice with hUb-CreERT2 line33, combined with reduced URI expression through the crossing of URIlox mice26, resulted in the same tumour burden and survival incidence as observed with p53 deletion alone (Fig. 9s–v). Thus, modulating URI levels does not initiate tumorigenesis in tumours arising from a p53-depleted background.
URI exhibits oncogenic activity in CRC independently of p53 at later stages
We next investigated whether, beyond APC-loss-driven CRC initiation, URI could exert other oncogenic activities at later stages in tumorigenesis, in a p53-dependent manner. URI immunoprecipitated p53 in already transformed HT-29 CRC cells, which are tumour-derived and represent a model of already initiated CRC, and which overexpress a mutant version of p53 (Supplementary Fig. 8a). HA-URI overexpression increased p53 ubiquitination in these cells (Supplementary Fig. 8b) without altering total p53 protein levels, which remained unaffected by the proteasome inhibitor MG132, chloroquine, or cycloheximide (Supplementary Fig. 8c, d). Accordingly, p53 failed to interact with PSMB4 in these p53 mutant cells, as demonstrated by co-IP experiments (Supplementary Fig. 8e). These findings suggest that while URI can still bind mutant p53 in HT-29 CRC cells and enhance its ubiquitination, degradation does not occur, likely because mutant p53 fails to interact with the proteasome machinery. Nevertheless, HA-URI overexpression in these p53-mutated HT-29 CRC cells increased cell numbers (Supplementary Fig. 8f), while downregulation of URI with siRNA had the contrary effect, confirming that URI may exhibit oncogenic activity in CRC independently of p53 degradation.
Accordingly, patients with mutations or genomic alterations in TP53 gene presented higher URI levels than those with wild-type TP53 (Supplementary Fig. 8g); in line with the positive correlation of URI expression with the tumoral stage and different markers of the cancer process (Fig. 1). The previously discussed correlation between high URI levels and lower survival was also observed in patients with alterations in TP53 (Supplementary Fig. 8h), further supporting the idea that URI might present oncogenic activities in tumoral cells at later stages, independently of its role on degrading p53 to initiate tumorigenesis.
In summary, CRC develops as a result of the stepwise acquisition of mutations in the intestinal epithelial cells. The loss of APC typically represents the initial “first hit,” triggering the transformation process. By using human data and genetically engineered mouse models, we demonstrate that APC loss-induced downregulation of p53 protein is a critical early event in CRC development. Mechanistically, we identify URI as an oncogenic target of MYC, a transcription factor activated by β-catenin following APC loss. MYC-driven URI overexpression enhances MDM2 activity, promoting p53 ubiquitination and degradation via proteasomal pathways, triggering CRC development. The downregulation of p53 disrupts cell cycle regulation, allowing uncontrolled proliferation, DNA damage, predisposing cells to acquire additional mutations necessary for adenoma progression, such as mutations in KRAS or p53 gene loss. The genetic loss of Tp53 often occurs as a later event, facilitating the transition from adenomas to adenocarcinomas. We also show that genetic p53 loss can drive CRC independently of oncogenic URI, but p53 protein degradation remains an initiating event that depends exclusively on the MYC-URI axis during APC loss-driven CRC initiation. Therefore, URI is oncogenic by promoting p53 degradation, but it can also exert oncogenic activities at later stages of CRC that depend on URI targets other than p53 (Fig. 10).
CRC develops as a result of the stepwise acquisition of mutations in the intestinal epithelial cells. The loss of APC typically represents the initial “first hit,” triggering the transformation process. This loss leads to β-catenin activation, which in turn activates MYC. MYC drives the transcription of the oncogenic protein URI, which enhances MDM2 activity. Increased MDM2 activity results in p53 ubiquitination and proteasomal degradation, a critical step in initiating tumorigenesis. The downregulation of p53 disrupts cell cycle regulation, allowing uncontrolled proliferation, and predisposing cells to acquire additional mutations necessary for adenoma progression, such as mutations in KRAS or p53 gene loss. The genetic loss of Tp53 often occurs as a later event, facilitating the transition from adenomas to adenocarcinomas. We also show that genetic p53 loss can drive CRC independently of oncogenic URI, but p53 protein degradation remains an initiating event that depends exclusively on the MYC-URI axis during APC loss-driven CRC initiation. Therefore, URI is oncogenic by promoting p53 degradation, but it can also exert oncogenic activities at later stages of CRC that depend on URI targets other than p53.
Discussion
Although various genetic mutations associated with the onset and progression of CRC have been identified and extensively studied, the interplay between these mutations and their implication in cellular pathways is not fully characterized, and neither are the molecular events involved in tumour initiation and the acquisition of transitional mutations critical for the development of aggressive tumours. This lack of understanding may justify why CRC persists as the third leading cause of cancer-related deaths, despite global efforts to comprehend, diagnose, and treat this prevalent disease. In this context, we propose in humans and mice a revised model for CRC development. We demonstrate that APC loss mediated-MYC activation induces URI expression, which in turn triggers MDM2 activation, promoting p53 protein degradation as an early event that is essential in the initiation of CRC. Loss of p53 at the protein level contributes to tumour initiation possibly by allowing hyperproliferation of epithelial cells and triggering replicative stress and DNA damage accumulation, ultimately facilitating the acquisition of transitional mutations in genes such as KRAS, which are essential for aggressive tumour development, regardless of the CRC molecular subtypes. However, at later stages of CRC, URI appears to exert oncogenic activity in a p53-independent manner, suggesting that URI functionally regulates other targets that help sustain CRC tumours.
Aging is a major risk factor for cancer, including colorectal cancer, with the tumour suppressor protein p53 playing a crucial role in this relationship. p53 is key in promoting repair and survival of damaged cells. With age, p53 function becomes impaired and associated cellular processes decline, increasing the risk to initiate a tumoral process. However, according to the well-established model by Fearon and Vogelstein, loss of p53 is classically considered as a late event in CRC progression and aggressiveness (see Supplementary Fig. 1a), a perspective that contradicts the correlation between aging and tumorigenesis. Our data indicate that upregulating p53 protein levels effectively prevents the occurrence of colorectal tumours, supporting the view that aging is a risk factor for CRC. Conversely, loss of p53 increased tumour burden in mice with APC loss or β-catenin overactivation background, and notably, significantly elevated the number of small dysplastic areas. These findings demonstrate that loss of p53 at the protein level is required for tumour initiation after WNT/MYC activation, most likely by inducing aberrant cell cycle progression, driving replicative stress accumulation and increasing DNA damage.
These findings are supported by the fact that functional p53 expression has been linked to better prognosis, disease stage, and treatment response in CRC34,35. Moreover, increasing the copy number of Tp53 in mice effectively prevented CRC. Similarly, elephants have evolved protective mechanisms against cancer by possessing an increased copy number of the TP53 gene36. Interestingly, elephants are protected from age-related disorders37. Although the mechanistic intricacies are yet to be fully elucidated, our findings suggest that URI loss plays a crucial role in preventing p53 degradation via the proteasome, thereby providing protection against CRC initiation. Recent discoveries corroborate the importance of proteasomal degradation of p53 in tumour formation, as one truncated form of p53 protein escapes MDM2-mediated proteasomal degradation and this serves as a mechanisms of tumour defence in elephants38,39. Notably, the regulation of p53 protein levels appears to be an independent process from the genetic loss of p53. Also, genetic p53 loss can drive CRC independently of oncogenic URI. This genetic loss is obviously not universal in patients, typically occurring at later stages of tumorigenesis and often associated with metastases and poor prognosis.
Our findings also offer strong evidence for choosing the most appropriate mouse model depending on the specific stage of human CRC. We demonstrate that oncogenic activation of KRAS alone is insufficient for tumor initiation but is crucial for metastasis in the context of APC and p53 loss. This data confirms the significance of oncogenic KRAS mutations in CRC aggressiveness and metastases, serving as an indicator of poor prognosis2,40,41. Moreover, despite KRAS mutations characterized a CMS of CRC that is different from the CMS of WNT activation/MYC signaling, it appears that constitutive activation of KRAS in APC-depleted mice could lead to the development of polyps in the colon in these mice, indicating that this model could reticulate human CRC, where polyps are found in the colon rather than in the small intestine.
In conclusion, here we provide a revised and updated version of the stepwise progression model for CRC tumorigenesis during aging (Fig. 10). The initial molecular oncogenic event involves MYC activation though aberrant β-catenin overactivity mediated by APC loss. Subsequently, MYC promotes p53 degradation by activating its target oncogene URI, which degrades p53 protein via modulation of MDM2 activity, a crucial step for the initiation of CRC development by inducing hyperproliferation. p53 loss could also contribute to accumulate replicative stress, ultimately leading to the acquisition of a mutational burden essential for the development of aggressive CRC. Among the acquired mutations, activation of KRAS oncogenic protein is identified as a later event, increasing aggressiveness of adenomas and, in conjunction with genetic loss of p53, driving metastases in CRC. Therefore, preventing p53 degradation by targeting URI could be an effective therapy to prevent CRC in humans.
Methods
Our research complies with all relevant ethical regulations. All animal procedures regarding housing and treatments were approved by the corresponding ethics committee at CNIO-ISCIII (Instituto de Salud Carlos III) and CAM (Comunidad de Madrid) and followed the recommendations from the European Union guidelines.
Reagents and antibodies
All reagents and antibodies used are listed in Supplementary Data 1.
Mouse models
Generation of intestinal-specific lines was obtained by crossing the corresponding conditional lox mice, described below, with vil-Cre-ERT2 mice24.
APC15lox mouse23 was kindly provided by Dr. Ron Smits. Apc(+/15Δ)vil mice presenting tumours in both small intestine and colon were generated by crossing APC15lox mice with vil-Cre-ERT2. To investigate the involvement of MYC in the tumour formation in Apc(+/15Δ)vil mice, conditional Myc lox mice25 were crossed with APC15lox mice, generating Apc(+/+)vil; Myc(Δ/Δ)vil mice and Apc(+/15Δ)vil; Myc(Δ/Δ)vil mice. To decipher the role of URI downregulation in the tumour formation process in Apc(+/15Δ)vil mice, conditional Urilox mice targeting exon 4 of Uri126 were crossed with APC15lox and vil-Cre-ERT2 mice, generating Apc(+/15Δ)vil; Uri(+/Δ)vil mice. To investigate the role of KRAS oncogene activation in intestinal tumour formation, KRAS+/V12 mice31 were crossed with vil-Cre-ERT2 mice, generating KRAS(+/V12)vil mice. These mice were also bred with APC15lox and URIlox mice, generating Apc(+/15Δ)vil; KRAS(+/V12)vil and Apc(+/15Δ)vil;KRAS(+/V12)vil; Uri(+/Δ)vil mice.
Overactivation of β-catenin in intestinal epithelium by depletion of exon 3 of β-catenin, containing the phosphorylation sites required for its degradation, was achieved by crossing the conditional Catnblox(ex3) mice30 with vil-Cre-ERT2 mice, generating Catnb+/Δ(ex3)vil mice. Catnb+/Δ(ex3)vil mice were crossed with Urilox mice in order to investigate the role of URI downregulation in the tumour formation derived from β-catenin over activation, generating Catnb+/Δ(ex3)vil; Uri(+/Δ)vil mice.
In order to investigate the role of p53 in these tumour models, p53 downregulation was prevented by crossing Apc(+/15Δ)vil and Catnb+/Δ(ex3)vil mice with "Super p53” mice, harbouring supernumerary copies of the p53 gene28. To confirm the role of p53 in tumour initiation and in URI loss-mediated tumour prevention, the models described above were crossed with conditional p53lox mice29 generating (1) p53(Δ/Δ)vil mice; (2) Apc(+/15Δ)vil; p53(Δ/Δ)vil mice; (3) Uri(+/Δ)vil; p53(Δ/Δ)vil mice; (4) Apc(+/15Δ)vil; Uri(+/Δ)vil; p53(Δ/Δ)vil mice; (5) KRAS(+/V12)vil; p53(Δ/Δ)vil mice (6) Apc(+/15Δ)vil; KRAS(+/V12)vil; p53(Δ/Δ)vil mice (7) Apc(+/15Δ)vil; KRAS(+/V12)vil; Uri(+/Δ)vil; p53(Δ/Δ)vil mice; (8) Catnb+/Δ(ex3)vil; p53(Δ/Δ)vil mice; and (9) Catnb+/Δ(ex3)vil; Uri(+/Δ)vil; p53(Δ/Δ)vil mice.
To study the role of URI overexpression in the intestinal epithelium regarding modulation of p53 levels, inducible human URI overexpressing mice22 (hURI(+/KI)) were bred with intestinal epithelium-specific reverse tetracycline trans-activator rtTA2-M2 (Villin-rtTA2-M2) mice42, generating hURI(+/KI)vil mice26. To investigate the role of URI overexpression in the intestinal epithelium in the tumour formation process in Apc(+/15Δ)vil mice, hURI(+/KI) mice with Villin-rtTA2-M2 were crossed with APC15lox mice, generating Apc(+/15Δ)vil; hURI(+/+)vil and Apc(+/15Δ)vil; hURI(+/KI)vil mice.
To investigate the role of URI overexpression in the intestinal epithelium in the tumour formation process in p53(Δ/Δ)vil mice, hURI(+/KI) mice with Villin-rtTA2-M2 were crossed with p53lox mice, generating p53(Δ/Δ)vil; hURI(+/+)vil mice and p53(Δ/Δ)vil; hURI(+/KI) mice.
To investigate the involvement of MYC in the tumorigenesis process in p53(Δ/Δ)vil mice, conditional Myc lox mice were crossed with p53lox mice, generating p53(Δ/Δ)vil; Myc(Δ/Δ)vil mice.
To model the tumorigenesis process initiated by loss of p53 in the whole body, and the role of URI in this process, p53lox mice were crossed with hUb-CreERT2 mice provided by CNIO animal facility and with URIlox mice, generating p53(Δ/Δ)Ub; URI(+/+)Ub mice and p53(Δ/Δ)Ub; URI(+/Δ)Ub mice.
All strains were backcrossed to a C57BL/6 for at least seven generations, and background was periodically refreshed by crossing the mice from the different mouse strains with C57BL/6 mice provided by CNIO animal facility. Siblings were never set together for mating. All experiments were performed by comparing littermates housed in the same cage. Both males and females were used for this study. Same Cre recombinase levels across mice were assured by always crossing a wild-type mouse with a Cre positive mouse, ensuring heterozygosity for the mice used in the experimental sets.
Mouse conditions and housing
All mice were housed at CNIO animal facility in pathogen-free conditions. They were exposed to a 12 hours light and dark cycle between 8:00 and 20:00. Room temperature was controlled and maintained at 23 ± 1 °C. Mice were fed ad libitum a chow diet (Harlan Laboratories, 2018S), unless otherwise indicated. Mice were sacrificed at the specified time points according to the mouse model, or when reaching the human end point criteria as established by the above mentioned corresponding ethical committees. For the ageing experiments, time of death was considered when either mice died naturally or when they had to be sacrificed because of reaching the human end point, including reaching a tumour size of 1 cm. That maximum tumour size was never exceeded, and mice were sacrificed immediately when reaching that size. Mice that died because of natural causes not related with this study were excluded from the analysis.
Mouse diets and treatments
To induce the Cre-ERT2-mediated recombination of the different conditional lox alleles, mice were fed a tamoxifen diet at a concentration of 400 mg/kg (mouse body weight) for two weeks, starting at 8 weeks old (TAM400/ CreER, TD.55125, ENVIGO).
To induce hURI expression in the mouse intestinal epithelium of hURI(+/KI)vil mice, they were fed a 0.625 g/kg doxycycline diet (AIN93G 0.625g_kg Doxycycline Hyclate, U8978 Version 59, SAFE Custom Diets).
Mouse genotyping
Genotyping of mice was evaluated by PCR from new-born’s tissue (fingers or ears) and confirmed from tissue obtained when sacrificing the mice (tail). Tissue was digested with proteinase K at 400 µg/mL in digestion buffer (1% SDS, 100 mM NaCl, 100 mM EDTA, 50 mM Tris pH 8) overnight at 56 °C. Supernatant was recovered after centrifugation of samples with 5 M NaCl, and DNA was precipitated with 0.7 volumes of isopropanol after a 30 minutes centrifugation at 4 °C. Pellet was washed with 75% ethanol and fully dried before resuspension in 500 µL of Milli-Q water. 2 µL of sample were used to run standard PCR using a commercial PCR master mix from NZYTech and the primers listed in Supplementary Table 1. A standard PCR program following manufacturer’s recommendations was used, adjusting the annealing temperature for the corresponding primers used, as specified in Supplementary Table 1. PCR products were analysed in 2.5% agarose gels in TAE buffer and DNA bands were visualized with Green Safe (NZYTech).
Mouse phenotyping
Overall looking of the mice at the time of sacrifice was pathologically analysed as follows: “sick” was considered when mice presented obvious signs of pathology (pale skin, loss of body weight), and “very sick” was considered when mice reached the human end point at the time of sacrifice (more than 20% loss of body weight, very pale skin, apathy and signs of suffering).
Mouse samples collection
Mice were euthanized for necropsy analysis and tissues collection at the indicated time points using a CO2 chamber. To determine anaemia markers, blood was collected from the heart of euthanized mice and kept in EDTA-containing vials. Hemogram analysis was conducted using Abacus Junior Vet analyser (Diatrom). Intestine and colon were harvested quickly, length was measured using a ruler and they were opened longitudinally and cleaned for macroscopic tumour analysis. A piece of tumour-free tissue was immediately frozen and kept at −80 °C for further analysis. The rest of the tissue was “Swiss-rolled”, kept in 10% formalin overnight and processed for paraffin embedding. Spleen and liver were extracted from mice, weight was assessed using an analytical balance and they were as well kept in 10% formalin overnight before processing for paraffin embedding.
Mouse tumour grade and size quantification
Paraffin blocks containing mouse tissues were cut to obtain 3 µm sections, deparaffinised and rehydrated. Samples were stained for haematoxylin and eosin following standard methods, and afterwards mounted in xylene. Tumour grade was assessed by histopathological analysis by a specialized pathologist following previously described methods for tumour grade classification43. GIN (gastrointestinal intraepithelial neoplasia) was scored as grade 1, low grade adenoma was scored as grade 2, high grade adenoma was scored as grade 3 and adenocarcinoma as grade 4. Tumour size was quantified by analytical measurement of all tumour area with a bright field microscope.
Immunostaining and immunofluorescence
Immunostaining and immunofluorescence have been performed as previously reported15,26,44. Briefly, sections containing mouse tissue were deparaffinised and rehydrated. Antigen retrieval was performed by heating samples in 1 M sodium citrate buffer pH 6.5. Then, endogenous peroxidase activity was blocked by 10 minutes incubation with 3% H2O2. Tissue was permeabilized by incubating the samples with 0.2% Triton X-100 diluted in PBS for 10 min. Samples were blocked for unspecific antibody binding by a 1-hour incubation with 5% BSA diluted in PBS with Triton X-100. Incubation with primary antibody was performed overnight at 4 °C with the corresponding antibodies (Supplementary Data 1), diluted 1:200 in 5% BSA diluted in PBS with Triton X-100. Samples were washed in PBS with Triton X-100, and secondary antibody incubation was performed using Vectastain ABC kit following manufacturer’s indications. Signal was detected using DAB (33-diaminobenzidinetetracloride), and samples were counterstained with haematoxylin and mounted with xylene.
For co-immunohistochemistry for two proteins, slides were sequentially incubated with two primary antibodies from different host species, and detected by DAB and fast-red following the protocol described above.
For immunofluorescence, after the primary antibody blocking step described above, endogenous avidin and biotin binding from mouse samples was blocked using biotin and avidin solutions, respectively. Primary antibody against URI was incubated overnight at 4 °C, 1:200. Samples were washed, incubated with a secondary antibody coupled with HRP (1:500) during 1 hour at room temperature, washed again, and then signal was amplified using TSA Plus Biotin Kit, following manufacturer’s guidelines. Samples were washed, incubated with streptavidin Alexa Fluor 488 conjugate, washed again and incubated with DAPI (1:10000) for 5 min at room temperature. Samples were mounted in Mowiol 4-88.
Protein expression and purification
Rosetta (DE3) competent cells were transformed with GST fusion-protein plasmids in pGEX4T-1 backbone (Supplementary Data 1) for URI full length protein and fragments, and cultured in 200 mL of LB media until reaching an optical density (OD) at 600 nm of 0.4. Induction of protein expression was performed with 400 µM of isopropyl β-d-1-thiogalactopyranoside (IPTG) for 16 hours at 30 °C. Bacteria were pelleted and washed with PBS before lysis with 50 mM Tris-HCl (pH 8), 1000 mM NaCl2, 1% NP-40, 2 mM EDTA, 7,5% glycerol, 1 mM PMSF, lysozyme (100 µg/mL), and DNAse I (10 µg/mL). Lysates were sonicated three times for 10 s, with 15 s pause between pulses, at 70% amplitude and clarified by centrifugation at 6000 rcf for 15 min at 4 °C. Proteins were pulled-down by adding 100 µL of glutathione sepharose beads slurry and centrifugation after 2 hours of incubation. Pellets were washed 5 times with PBS, resuspended in 50% glycerol in PBS and stored. Quality of samples was assessed by acrylamide gel resolving followed by Coomassie blue staining following standard protocols.
In vitro protein binding assays
30 ng of human recombinant p53 or MDM2 (Supplementary Data 1) were incubated with 10 µL of bacterially expressed and purified GST-URI protein and fragment constructs, or with GST alone, bound to glutathione sepharose beads, during 2 hours at 4 °C, in 250 mM Tris-HCl pH 8, 150 mM NaCl, 5% glycerol, supplemented with 1 mM EDTA, 0.5 mM PMSF and cOmpleteTM. Pull-downs were collected by 1 min centrifugation at 5000 g and washed 5 times before Western blot analysis.
Ubiquitination in vitro assay
MDM2 ubiquitination activity of p53 was performed as previously reported45 by combining a commercially available ubiquitination assay kit (Supplementary Data 1) with human recombinant MDM2 (0.35 µM), human recombinant p53 (0.25 µM), human recombinant UBE2D1 E2-ligase (0.85 µM), and increasing concentrations of purified GST-URI (5, 10 and 30 µL). Reactions were performed as reported45 and following manufacturer’s instructions. Results were analysed by Western blotting after 1 hour of incubation at 37 °C.
Cell culture conditions and treatments
HCT-116 and HT-29 human colorectal cancer cell lines were cultured in Dulbecco’s modified minimum eagle media (DMEM). RKO human colorectal cancer cell line was cultured in minimum eagle media (MEM). LIM1215 human colorectal cancer cell line was cultured in RPMI media. All cultured media were supplemented with 10% fetal bovine serum (FBS), 1% penicillin-streptomycin, and sodium pyruvate. All cell lines were cultured at 37 °C and 5% CO2. Passages were done 1/10 using trypsin when reaching 80% of confluence.
Proteasome inhibition was performed by treating confluent cells with 10 μM MG132, diluted in DMSO, in cell culture media for 6 hours. DMSO was used in control cells.
Inhibition of translation was performed by treating confluent cells with cycloheximide (CHX) at a final concentration of 10 ng/µL, diluted in DMSO, in cell cultured media for up to 6 hours.
Inhibition of autophagy was achieved by treating confluent cells with of chloroquine (CHQ) at a final concentration of 10 µM, diluted in DMSO, in cell cultured media for 6 hours.
Cell numbers were studied by trypsinizing cells followed by trypan blue staining to differentiate dead and alive cells.
Cell plasmid and siRNA transfection
Plasmids (Supplementary Data 1) were prepared using Miniprep kits. 300000 cells per well were plated in 6 wells plates. Transfection was performed in sub-confluent cells, which were washed and treated with 300 µL of Optimem containing 2500 ng of the corresponding plasmid (Supplementary Data 1) or siRNA at a final concentration of 10 nM (Supplementary Data 1), and 2 µL of Lipofectamine 2000, as previously reported46. Four hours later, 2 mL of media were added per well. Cells were harvested for protein analysis after 48 hours.
URI promoter reporter assays
RKO cells were co-transfected with Renilla-encoding plasmid and with either a pGL4.10-Luc reporter vector or a URI-reporter plasmid with the 440 base pairs regulatory sequence of URI1 opening reading frame (ORF) cloned as reported22. After two days of transfection, cells were lysed and luciferase and renilla activities were measured using the Dual-Luciferase Reporter Assay System following manufacturer’s instructions.
Immunoblotting
Approximately 70 mg of frozen mouse tissue were lysed in 750 µl of RIPA buffer (50 mM Tris pH 8.0, 150 mM NaCl, 2 mM MgCl, 2 mM CaCl2, 0.5% sodium deoxycholate, 0.1% SDS, 1% NP-40 alternative and 10% glycerol), supplemented with 2 mM EDTA, 1 mM Na3VO4, 1 mM NaF, 0.5 mM PMSF, 10 µg/mL aprotinin and 0.5 mM DTT. Samples were homogenised using Precellys 24 Bead homogenizer (15 ×2 s, 5500 w) and supernatant containing tissue’s proteins was collected after a 30 min centrifugation at 13000 g. Lysates from cells’ samples were obtained by scratching the corresponding wells with the supplemented RIPA lysis buffer above mentioned. Lysates were cleared by a 10-minute centrifugation at 13000 g. Protein concentration was measured with the colorimetric Bradford assay, using a range of BSA dilutions of known concentrations (0-500 mg) as a standard curve, to ensure equal amounts of protein loaded across all samples. Concentration was adjusted and samples were boiled in Laemmli buffer (2% SDS, 10% glycerol, 1% DTT, 0.0025% bromophenol blue in 250 mM Tris-HCl pH 6.8) at 95 °C for 10 minutes. 20 µg of protein per sample were loaded into SDS-PAGE gels prepared in house and proteins were transferred into nitrocellulose membranes, as previously described. Membranes were blocked for unspecific antibody binding by 1 hour incubation with 5% BSA in TBS-T (Tris Buffered Saline with 0.1% Tween-20). Membranes were incubated overnight at 4 °C with the corresponding primary antibodies (Supplementary Data 1), diluted 1:1000 in 5% BSA TBS-T with 0.05% sodium azide. After washing in TBS-T, membranes were incubated with secondary antibody diluted 1:5000 in TBS-T for 1 hour at room temperature and washed again. Membranes were developed after 1 minute incubation with homemade ECL 1:1 solutions A (1.25 mM luminol, 0.2 mM p-coumaric acid in Tris-HCl 0.1 M pH 8.5) and B (0.03% H2O2 in Tris-HCl 0.1 M pH 8.5), using X-ray films. Full scans of the blots are provided as Source Data files.
Chromatin immunoprecipitation (ChIP)
106 of RKO cells or 10 mg of mouse intestinal tumour tissue were used per ChIP experiment. Cells and tumoral tissue were cross-linked with 1% formaldehyde in serum-free DMEM (4.5 g/L glucose) media. After the reaction was quenched with 0.125 M glycine, the cells or tissue were washed three times with ice-cold PBS, collected by scraping and centrifuged for 5 min at 1000 g. Pellets were re-suspended in lysis buffer (2% Triton X-100, 1% SDS, 100 mM NaCl, 10 mM Tris-HCl pH 8.0, 1 mM EDTA supplemented with 1x cOmplete™ EDTA-free protease inhibitors) and incubated for 30 min on ice followed by sonication for 6 min for cells (duty factor 5%, peak power 140, cycles per burst 200). For mouse intestinal tumours, prior to sonication, tissue was homogenized by douncer (20-30 times) and sonicated for 2 min in a E220 sonicator. The chromatin was centrifuged at 16.000 g for 10 min, and supernatant was recovered and diluted with dilution buffer containing 50 mM HEPES pH 8.0, 140 mM NaCl, 1 mM EDTA, 0.75% Triton X-100, 0.1% Na-deoxycholate supplemented with 1x cOmplete™ EDTA-free protease inhibitor to obtain a final concentration of SDS 0.2% for ChIP. For each immunoprecipitation, 60 µL of Dynabeads protein G were used (30 µL for immunoprecipitation and 30 µL for pre-clearing). Beads were washed three times with working buffer (1:4 – lysis buffer:dilution buffer supplemented with 1x cOmplete™ EDTA-free protease inhibitors). After the final wash, beads were re-suspended in 60 µL of working buffer for each immunoprecipitation. 30 µL of washed beads were added to diluted chromatin and incubated for 2-3 h at 4 °C for pre-clearing. The rest of the beads were topped up with working buffer to a final volume of 1 mL where 50 µL of 10% BSA was added. These beads were blocked overnight. After 2-3 h of pre-clearing, chromatin was placed on a magnetic stand to remove beads and supernatant was transferred to a new protein-low binding tube. 5 µg of the MYC antibody together with 10% BSA final volume and 1% of tRNA (10 µg/µL stock concentration) was added for overnight incubation. The next day, blocked beads were placed on the magnet, re-suspended in 30 µL of working buffer, added to the chromatin-antibody tube, and incubated for 2 h at 4 °C. Then, chromatin-antibody bound beads were washed once for 5 min at 4 °C with 1 mL first with low salt immune complex buffer (1% Triton X-100, 150 mM NaCl, 20 mM Tris-HCl pH 8.0, 0.1% SDS and 2 mM EDTA), then with high salt immune complex buffer (1% Triton X-100, 500 mM NaCl, 20 mM Tris-HCl pH 8.0, 0.1% SDS and 2 mM EDTA), and finally with LiCl immune complex buffer (0.25 M LiCl, 1% deoxycholate sodium, 10 mM Tris-HCl pH 8.0, 1% NP-40 and 1 mM EDTA) followed by three short washes with ice-cold TE buffer. To elute bound chromatin, beads were incubated with rotation in 150 µL of elution buffer (1% SDS, 0.1 M NaHCO3) for 15 min, placed on a magnetic stand and clear supernatant was transferred to a new tube after which this step was repeated such that the final volume of the elute was 300 µL. 1.5 µL of RNAse A was added to each tube for incubation at 37 °C for 30 min followed by addition of 12 µL of 5 M NaCl and 4.5 µL of Proteinase K for overnight de-crosslinking at 65 °C overnight. The next day DNA was isolated by Phenol:CHCl3:isoamyl extraction and ethanol precipitation. This DNA was used for quantitative PCR (qPCR) as described below. Primers are listed in Supplementary Table 2.
Reverse transcription-quantitative polymerase chain reaction (RT-qPCR)
Approximately 70 mg of frozen tissue were lysed using Trizol to obtain total RNA. RNA concentration was determined using Nanodrop and 3 µg of RNA were used to synthetize first strand cDNA using M-MLV reverse transcriptase. Relative amount cDNA was measured by qPCR using GoTaq Real-Time qPCR mix and the corresponding primers at 3 mM (Supplementary Table 3). Results are reported as the 2-ΔΔCt values, obtained by subtracting the mean ΔCt value (Ct value of the gene of interest minus the Ct value of the house keeping gene, Actin or Tbp, as indicated) to the mean ΔCt value of the control group.
Co-immunoprecipitation
Sub-confluent cells were lysed in lysis buffer (250 mM Tris-HCl pH 8, 150 mM NaCl, 1% NP-40, 5% glycerol) supplemented with 1 mM EDTA, 1 mM NaF and cOmpleteTM. Immunoprecipitation of the indicated proteins was performed using 25 µl of Protein A Sepharose beads, previously washed and incubated with the corresponding antibody (Supplementary Data 1) for 1 hour at 4 °C, with 500 μL of the obtained lysate, at a final antibody concentration of 1 μg/mL, by rotation for 2 hours at 4 °C. After washing, samples were boiled in Laemmli buffer (2% SDS, 10% glycerol, 1% DTT, 0.0025% bromophenol blue in 250 mM Tris-HCl pH 6.8) at 75 °C for 15 min, and protein presence was analysed by immunoblotting as described above. Whole cell extract control refers to 5% of the total lysate.
Human samples
Human CRC samples were obtained from CNIO Biobank integrated in the Spanish National Biobanks Network and Biomodels Platform of the ISCIII (PT20/00045). Samples from patients with ulcerative colitis (UC) and Crohn’s disease (CD) were obtained and processed at the University Guadalajara Hospital. The human adenomatous polyps were obtained from Cristian Perna from the Ramon y Cajal Hospital. Human samples were processed following standard operating procedures with the appropriate approval of the Ethical and Scientific Committees. Informed consent was obtained from all subjects and samples were processed in paraffin-embedded blocks generating TMAs for CRC, UC and Crohn´s disease simples (tissue micro arrays). Paraffin-embedded tissues were cut into 3 µm sections, and they were stained for haematoxylin & eosin and immunostained as detailed above.
Human cancer samples analysis from The Cancer Genome Atlas
Genomic data referring to copy number variants and mutations in human cancer patient samples from The Cancer Genome Atlas Pan-Cancer study (mc3.v0.2.8.PUBLIC.maf.gz; syn5049520)47. Functional somatic mutation calls were collected for all premature truncation mutations (including splicing variants, frameshift indels, and nonsense variants) as well as for non-synonymous (missense mutations and single-residue substitutions, including in-frame indels). Using GISTIC 2.048, genomic regions exhibiting significant levels of copy number (CN) alterations and their target genes were identified, along with their respective q-values. Gene-level CN data were acquired from Synapse (syn5049520). High-level deletion for a gene was designated by a GISTIC threshold CN value of -2, while high-level amplification was indicated by a threshold CN value of +2. Broad-level deletion (loss) and broad-level amplification (gain) were determined based on GISTIC threshold CN values below -1 or above +1, respectively. Raw counts for the mRNA data from human patient samples from The Cancer Genome Atlas was downloaded from Firebrowse (http://firebrowse.org). Normalization of the raw annotated counts and differential expression analysis was done using DESeq2 package49 for R. High URI1 levels were considered when the fold increase compared to normal tissue was greater than 1.5. Stratification of the patients according to the molecular subtype was done according to previous publications10. A comprehensive collection of 235 cancer genes was curated by integrating predictions from eight cancer driver gene prediction tools (20/20 + , ActiveDriverDB, CompositeDriver, MuSiC, MutSig2CV, OncodriveCLUST, OncodriveFML, and e-Driver), along with manual literature review and mining individual TCGA studies featured in the Pan-Cancer Atlas47.
Human colorectal polyps, ulcerative colitis and Crohn’s disease samples analysis
URI1 mRNA levels in human colorectal polyps was analysed from transcriptomic data publicly available12. Polyps were classified according to their molecular type (adenomas, serrated neoplasias or unknown). Analysis of URI1 mRNA levels in ileum, colon and rectum from ulcerative colitis (UC) and Crohn’s disease (CD) patients was performed on publicly available transcriptomic analysis16.
Image analysis and quantifications of immunostainings
Stainings were analysed using bright field or fluorescence microscopes, accordingly. To quantify the number of positive cells, a minimum of 50 intestinal villi or crypts (as indicated) per mouse were considered, from randomized areas of the sample. Adobe Photoshop was used to adjust the contrast and white balance of representative pictures for aesthetic purposes.
Statistical analysis and data representation
Statistical assessments of the differences among the groups evaluated in this study was performed using R (R version 4.0.2, released on (2020−06−22), which was also used for data representation. Different packages for R were used to facilitate data analysis and representation and are listed in Supplementary Data 1. When comparing means between only 2 groups, two-tailed Student’s t-test with Welch correction was used. When comparing means among more than two groups, one-way ANOVA with post-test corrected by Tukey’s approach was used. Outliers were considered when observations were outside the range of the first quartile minus 1.5 times the interquartile range and the third quartile plus 1.5 times the interquartile range and removed. Individual dots in the graphs represent individual mice, patients, tumours or individual experiments, as indicated. Bars represent the mean within the group, and error bars represent the standard deviation of the mean (SEM). Boxplots represent Q1, Q2, and Q3.
Analysis of categorical variables and co-occurrences among different groups was assessed with contingency tables and Fisher’s test.
Correlation analyses were performed using Pearson correlation test.
For the survival analysis in mice, competing risk analysis was performed in order to also consider the mice within the cohort who died because of unknown or cancer unrelated reasons. Data was represented as the survival probability.
Figures display was arranged using Adobe Illustrator. Graphic schemes are original and were drawn with Adobe Illustrator.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Materials are available upon request to N.D. and the sharing of materials described in this work will be subject to standard material transfer agreements. All data are available within the Article, Supplementary Information or Source Data file. Source data are provided with this paper.
References
Fearon, E. R. & Vogelstein, B. A genetic model for colorectal tumorigenesis. Cell 61, 759–767 (1990).
Sansom, O. J. et al. Loss of Apc allows phenotypic manifestation of the transforming properties of an endogenous K-ras oncogene in vivo. Proc. Natl Acad. Sci. USA 103, 14122–14127 (2006).
Schmitt, M. & Greten, F. R. The inflammatory pathogenesis of colorectal cancer. Nat. Rev. Immunol. 21, 653–667 (2021).
Feng, Z. et al. Declining p53 function in the aging process: a possible mechanism for the increased tumor incidence in older populations. Proc. Natl Acad. Sci. USA 104, 16633–16638 (2007).
Wu, D. & Prives, C. Relevance of the p53-MDM2 axis to aging. Cell Death Differ. 25, 169–179 (2018).
Herranz-Montoya, I., Park, S. & Djouder, N. A comprehensive analysis of prefoldins and their implication in cancer. iScience 24, 103273 (2021).
Djouder, N. et al. S6K1-mediated disassembly of mitochondrial URI/PP1gamma complexes activates a negative feedback program that counters S6K1 survival signaling. Mol. Cell 28, 28–40 (2007).
Gstaiger, M. et al. Control of nutrient-sensitive transcription programs by the unconventional prefoldin URI. Science 302, 1208–1212 (2003).
Chaves-Pérez, A., Thompson, S. & Djouder, N. Roles and functions of the unconventional prefoldin URI. Adv. Exp. Med. Biol. 1106, 95–108 (2018).
Guinney, J. et al. The consensus molecular subtypes of colorectal cancer. Nat. Med. 21, 1350–1356 (2015).
Sansom, O. J. et al. Myc deletion rescues Apc deficiency in the small intestine. Nature 446, 676–679 (2007).
Chen, B. et al. Differential pre-malignant programs and microenvironment chart distinct paths to malignancy in human colorectal polyps. Cell 184, 6262–6280 (2021).
Gibson, J. A. & Odze, R. D. Pathology of premalignant colorectal neoplasia. Dig. Endosc. 28, 312–323 (2016).
Bressenot, A., Cahn, V., Danese, S. & Peyrin-Biroulet, L. Microscopic features of colorectal neoplasia in inflammatory bowel diseases. World J. Gastroenterol. 20, 3164–3172 (2014).
Brandt, M. et al. mTORC1 inactivation promotes colitis-induced colorectal cancer but protects from APC loss-dependent tumorigenesis. Cell Metab. 27, 118–135.e118 (2018).
Massimino, L. et al. The inflammatory bowel disease transcriptome and metatranscriptome meta-analysis (IBD TaMMA) framework. Nat. Comput. Sci. 1, 511–515 (2021).
Hammal, F., de Langen, P., Bergon, A., Lopez, F. & Ballester, B. ReMap 2022: a database of Human, Mouse, Drosophila and Arabidopsis regulatory regions from an integrative analysis of DNA-binding sequencing experiments. Nucleic Acids Res. 50, D316–D325 (2022).
Dehingia, B., Milewska, M., Janowski, M. & Pekowska, A. CTCF shapes chromatin structure and gene expression in health and disease. EMBO Rep. 23, e55146 (2022).
Rickman, D. S., Beltran, H., Demichelis, F. & Rubin, M. A. Biology and evolution of poorly differentiated neuroendocrine tumors. Nat. Med. 23, 1–10 (2017).
Blackwell, T. K. et al. Binding of myc proteins to canonical and noncanonical DNA sequences. Mol. Cell Biol. 13, 5216–5224 (1993).
Sabo, A. & Amati, B. Genome recognition by MYC. Cold Spring Harb. Perspect. Med. 4, a014191 (2014).
Tummala, K. S. et al. Inhibition of de novo NAD+ synthesis by oncogenic URI causes liver tumorigenesis through DNA damage. Cancer Cell 26, 826–839 (2014).
Robanus-Maandag, E. C. et al. A new conditional Apc-mutant mouse model for colorectal cancer. Carcinogenesis 31, 946–952 (2010).
el Marjou, F. et al. Tissue-specific and inducible Cre-mediated recombination in the gut epithelium. Genesis 39, 186–193 (2004).
de Alboran, I. M. et al. Analysis of C-MYC function in normal cells via conditional gene-targeted mutation. Immunity 14, 45–55 (2001).
Chaves-Perez, A., Yilmaz, M., Perna, C., de la Rosa, S. & Djouder, N. URI is required to maintain intestinal architecture during ionizing radiation. Science 364, eaaq1165 (2019).
Ding, Z. et al. URI alleviates tyrosine kinase inhibitors-induced ferroptosis by reprogramming lipid metabolism in p53 wild-type liver cancers. Nat. Commun. 14, 6269 (2023).
Garcia-Cao, I. et al. Super p53” mice exhibit enhanced DNA damage response, are tumor resistant and age normally. EMBO J. 21, 6225–6235 (2002).
Marino, S., Vooijs, M., van Der Gulden, H., Jonkers, J. & Berns, A. Induction of medulloblastomas in p53-null mutant mice by somatic inactivation of Rb in the external granular layer cells of the cerebellum. Genes Dev. 14, 994–1004 (2000).
Harada, N. et al. Intestinal polyposis in mice with a dominant stable mutation of the beta-catenin gene. EMBO J. 18, 5931–5942 (1999).
Guerra, C. et al. Tumor induction by an endogenous K-ras oncogene is highly dependent on cellular context. Cancer Cell 4, 111–120 (2003).
Zafra, M. P. et al. An in vivo kras allelic series reveals distinct phenotypes of common oncogenic variants. Cancer Discov. 10, 1654–1671 (2020).
Ruzankina, Y. et al. Deletion of the developmentally essential gene ATR in adult mice leads to age-related phenotypes and stem cell loss. Cell Stem Cell 1, 113–126 (2007).
Sarasqueta, A. F. et al. Integral analysis of p53 and its value as prognostic factor in sporadic colon cancer. BMC Cancer 13, 277 (2013).
Iacopetta, B. TP53 mutation in colorectal cancer. Hum. Mutat. 21, 271–276 (2003).
Sulak, M. et al. TP53 copy number expansion is associated with the evolution of increased body size and an enhanced DNA damage response in elephants. Elife 5, e11994 (2016).
Chusyd, D. E. et al. Aging: what we can learn from elephants. Front. Aging 2, 726714 (2021).
Preston, A. J. et al. Elephant TP53-RETROGENE 9 induces transcription-independent apoptosis at the mitochondria. Cell Death Discov. 9, 66 (2023).
Padariya, M. et al. The elephant evolved p53 isoforms that escape MDM2-mediated repression and cancer. Mol. Biol. Evol. 39, msac149 (2022).
Jones, R. P. et al. Specific mutations in KRAS codon 12 are associated with worse overall survival in patients with advanced and recurrent colorectal cancer. Br. J. Cancer 116, 923–929 (2017).
Huang, D. et al. Mutations of key driver genes in colorectal cancer progression and metastasis. Cancer Metastasis Rev. 37, 173–187 (2018).
Roth, S. et al. Generation of a tightly regulated doxycycline‐inducible model for studying mouse intestinal biology. genesis 47, 7–13 (2009).
Boivin, G. P. et al. Pathology of mouse models of intestinal cancer: Consensus report and recommendations. Gastroenterology 124, 762–777 (2003).
Chaves-Pérez, A. et al. Transit-amplifying cells control R-spondins in the mouse crypt to modulate intestinal stem cell proliferation. J Exp Med. 219, e20212405 (2022).
Giansanti, C. et al. MDM2 binds and ubiquitinates PARP1 to enhance DNA replication fork progression. Cell Rep. 39, 110879 (2022).
Burén, S. et al. Regulation of OGT by URI in Response to Glucose Confers c-MYC-Dependent Survival Mechanisms. Cancer Cell. 30, 290–307 (2016).
Bailey M.H. et al. Comprehensive Characterization of Cancer Driver Genes and Mutations. Cell. 173, 371–385.e18 (2018).
Mermel, C.H. et al. GISTIC2.0 facilitates sensitive and confident localization of the targets of focal somatic copy-number alteration in human cancers. Genome Biol. 12, R41 (2011).
Love, M.I., Huber, W. & Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Acknowledgements
We thank all mouse providers as described in the Methods. We are also thankful to the CNIO Mouse Genome Editing Core Unit and Animal Facility for the mouse re-derivation and maintenance, respectively. We would like to also acknowledge the patients and the CNIO-Biobank (PT20/00070) integrated in the Platform ISCIII Biobanks and Biomodels for providing us with the human samples. I.H.-M. and M.A.-A. are respectively recipients of FPU fellowship and Severo Ochoa-FPI fellowship from the Ministry of Science, Innovation and Universities. This work was funded by grants to N.D. supported by the State Research Agency (MICIU/AEI/10.13039/501100011033) from the Ministry of Science, Innovation and Universities (PID2021-122695OB-I00), also including the iDIFFER network of Excellence (RED2022-134792-T), co-funded by European Regional Development Fund (ERDF), by the BBVA Foundation grants for biomedicine (EIC21-1-243) and by the Asociación Española Contra el Cáncer (AECC) (PRYGN211184DJOU). This work was developed at the CNIO funded by the Health Institute Carlos III (ISCIII) and the Spanish Ministry of Science and Innovation.
Author information
Authors and Affiliations
Contributions
I.H.-M. designed most of the experiments and analysed the data with N.D. M.A.-A. performed some biochemical studies and stainings. C.P. histopathologically analyzed tissues from mice and patients. S.Z. performed ChIP assays. L.G.-J. and S.P. analysed bioinformatics data together with I.H.-M. I.H.-M. and N.D. wrote the manuscript. N.D. conceived the project, developed the study and secured funding.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Kevin Myant and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Herranz-Montoya, I., Angulo-Aguado, M., Perna, C. et al. p53 protein degradation redefines the initiation mechanisms and drives transitional mutations in colorectal cancer. Nat Commun 16, 3934 (2025). https://doi.org/10.1038/s41467-025-59282-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-59282-4
This article is cited by
-
Single-cell sequencing and organoids: applications in organ development and disease
Molecular Biomedicine (2025)
-
OTUD4 suppresses colorectal cancer progression through deubiquitinating p53
Discover Oncology (2025)