Abstract
Mitochondrial membrane dynamics control the shape, number, and distribution of mitochondria and regulate energy production and cell health. In a screen for yeast mutants with increased levels of templated insertions (~10-1000 bp) in the nuclear genome, we identified mitochondrial fusion deficient mutants (mgm1Δ, ugo1Δ, fzo1Δ). We found that fusion mutants activate the iron regulon, have decreased iron-sulfur clusters (ISCs), and increased DNA damage, suggesting a role of iron homeostasis in preventing insertions. Consistently, a secondary screen found mutants affecting iron-sulfur cluster production (yfh1Δ, grx5Δ), vacuolar iron storage (ccc1Δ) or general iron homeostasis (aft1Δ) to exhibit high insertion levels. Treatment with iron chelators or hydrogen peroxide also increased insertions. We propose that iron dysregulation leading to oxidative DNA damage and compromised DNA repair drives insertions. These studies suggest that severe iron imbalance, associated with many human diseases and pharmacological treatments, can trigger genome instability in the form of templated insertions.
Similar content being viewed by others
Introduction
Mitochondria are highly dynamic organelles that undergo fusion or fission to respond to changing needs in the cell. Fusion and fission can facilitate mixing of mitochondrial contents, preserve mtDNA, regulate ATP production, respond to oxidative or nutrient stress, regulate apoptosis, facilitate mitophagy, distribute mitochondria to daughter cells, and more1. Mitochondrial fusion in yeast is coordinated by a complex of three protein components: dynamin-like GTPase Mgm1 (human OPA1), responsible for fusion of inner membranes and formation of cristae structures2, mitofusin Fzo1 (human MFN1, MFN2), needed for outer membrane fusion3, and linking protein Ugo1 (human SLC25A43) needed for fusion of both membranes and formation of cristae structures4,5. Mitochondrial fission is primarily regulated by dynamin-related GTPase Dnm1 (human DRP1), fission protein Fis1 (human FIS1), Caf4, and Mdv1. In humans, impaired mitochondrial fusion or increased fission are associated with severe neurological disorders such as autosomal dominant optic atrophy (ADOA), Behr syndrome, Charcot-Marie-Tooth disease, Parkinson’s disease, Alzheimer’s disease, Huntington’s disease, dementia, metabolic and cardiovascular diseases, mitochondrial DNA (mtDNA) depletion syndromes, cancers and more1,6,7,8,9,10,11,12,13,14.
Here, we unexpectedly identified mitochondrial fusion mutants to have elevated nuclear genome instability. We used yeast that offers a simple and cost-effective model to screen for mutants with high levels of templated insertions, called hereafter “insertions,” and found all three mitochondrial fusion mutants: mgm1Δ, fzo1Δ and ugo1Δ. Insertions are DNA fragments copied from elsewhere in the genome, or fragments of retrotransposon cDNA, or mtDNA pasted at chromosomal double-strand breaks (DSBs). They typically range in size from ~10 bp to >1 kb and depend on classic Ku mediated NHEJ. Insertions can occur at CRISPR/Cas9 or endonuclease-induced breaks or programmed breaks at the V(D)J locus where they contribute to antibody diversification15,16,17,18,19. These types of copy number variations (CNVs) are among the most common in cancer genomes and contribute to genetic disorders20,21. The genetic conditions that predispose to insertion formation remain mostly unknown.
While investigating the possible reasons underlying mitochondrial fusion mutants’ predisposition to insertions, we found that these mutants show increased iron uptake, decreased iron-sulfur clusters (ISCs), increased spontaneous DNA damage, and sensitivity to reactive oxygen species (ROS). A secondary screen focused on various mitochondrial pathways revealed multiple mutants with high levels of insertions sharing one common feature: altered iron homeostasis. These include mutants of ISC biogenesis, iron uptake, recycling, storage, as well as other mitochondrial pathways indirectly linked to iron homeostasis. All these iron-related defects are linked to increased DNA damage, genome instability and many human diseases22,23,24. Evidence suggests that insertions in mitochondrial fusion mutants result at least partially from ROS damage. We demonstrate that poor growth, ISC deficiency, iron overload, and high insertions in fusion mutants can be suppressed by increasing mitochondrial membrane potential (MMP) by either genetic manipulation or a compound increasing MMP. This result provides a potential therapeutic strategy in patients affected by mitochondrial fusion mutations that currently lack any cure6. In summary, this work suggests that mutants affecting iron homeostasis, either directly or indirectly, have a broad effect on both spontaneous DNA damage and the ability of cells to repair this damage. We propose that iron imbalance typical for many cancers, rather than loss of individual DNA repair enzymes or pathways, can explain high incidence of insertions and likely other types of genome instability.
Results
Mitochondrial fusion machinery suppresses templated insertions
To identify additional mutants with an increased rate of >10 bp insertions, we screened ~40 mutants involved in different aspects of genome stability, DNA repair, recombination, DNA damage checkpoint response, and replication. We also tested mutants affecting chromatin, cell death, autophagy, mitophagy, retrotransposition, or mutants with mtDNA instability (Supplementary Data 1). The screen was done using a yeast strain carrying a galactose-inducible HO endonuclease that generates a single-per-genome DSB at the MATa locus. Wild-type and mutant cells were plated on galactose-containing plates to induce DSBs, and only cells that repaired the break by NHEJ and altered the HO cleavage site formed colonies. The majority of NHEJ products are represented by small indels described previously in detail and therefore not analyzed here17,25. To identify NHEJ products carrying insertions, we used the amplicon sequencing method Break-Ins (Fig. 1a)17. About 20,000–250,000 independent NHEJ repair products were analyzed per mutant strain. Only sequencing reads carrying >10 bp long insertions were subjected to analysis. Unexpectedly, nearly all mutants with severe defects in DNA repair, recombination, replication, or checkpoint showed marginal increases or no increase of insertions, despite previously documented high rates of spontaneous DNA damage (Supplementary Data 1). Also unexpectedly, we found that mgm1Δ mutant with compromised mitochondrial dynamics showed a high level of insertions. Mgm1 encodes a dynamin-related GTPase required for mitochondrial fusion and maintenance of a tubular mitochondrial network. mgm1Δ mutant displayed a large ~28-fold increase in insertions from across the nuclear genome compared to wild type (Fig. 1b). mgm1Δ mutant is respiration deficient and lacks mtDNA26; therefore, mtDNA insertions could not be followed. To ensure that defective mitochondrial fusion is responsible for increased insertions, we tested additional fusion mutants. Mgm1 forms a complex with Fzo1 and Ugo1 and all three proteins are essential for a tubular mitochondrial network. Both fzo1Δ and ugo1Δ showed increased insertions (Fig. 1b). Among the three mutants, ugo1Δ has the highest level of insertions. The reason for the stronger insertions phenotype in ugo1Δ is not known but could be related to larger, more aggregated and defective mitochondria27 that results in slower growth of ugo1Δ cells when compared to mgm1Δ or fzo1Δ (Supplementary Fig. 1a, b). About 2–6% of insertions were complex with 2–3 fragments joined together, and inserted DNA originated preferentially from fragile regions of the genome such as R-loops, tRNA, tandem repeats, and telomeres (Fig. 1c and Supplementary Data 1). Additionally, all mitochondrial fusion mutants displayed elevated insertions from retrotransposable elements. As expected, this increase depended on Ty1 transcription regulated by Spt3 (Supplementary Fig. 1c). Inserted cDNA originated from all over the Ty1 with an increased level from LTR as previously shown (e.g17,28.) (Supplementary Fig. 1d). All fusion mutants showed increased levels of Ty1 cDNA and transposition, suggesting that increased Ty1 replication may be responsible for increased insertions originating from cDNA (Supplementary Fig. 1e, f). Indeed, at a lower temperature (23 °C) where cDNA and transposition increase29, insertions from Ty1 elements increased compared to 30 °C both in wild type and in mgm1Δ (Supplementary Fig. 1c). However, in this work we mostly focus on insertions originating from the nuclear genome.
a Schematic of Break-Ins method to detect templated insertions at DSBs. b Frequency and types of inserted DNA at DSBs in mitochondrial dynamics mutants. P values were determined using two-tailed χ2 test. n – number of independent NHEJ products tested ranged from ~50,000 to ~150,000 and shown in Supplementary Data 1 and 2. c Features analysis of donor DNAs for templated insertions in mitochondrial fusion mutants. Proximity is defined as a sequence within 0.2 kb from the R-loop or tRNA, within 1 kb from telomere end, and the overlap with tandem repeats indicates at least 1 bp overlap. Empirical P values were calculated using two-sided permutation test by comparing the observed value to 10,000 random values generated by permutations. Error bars represent 2 × standard deviations of the permuted distributions as 95% confidence intervals. d Frequency of insertions in WT and mgm1Δ with increased treatment of 500 nM Concanamycin A (ConcA) or DMSO control. n – number of NHEJ products tested shown in Supplementary Data 3. e Frequency and types of inserted DNA at DSBs in mitochondrial dynamics point mutants. P values were determined using two-tailed χ2 test. n – number of independent NHEJ products and shown in Supplementary Data 1 and 2. Source data are provided as a Source Data file.
In contrast to fusion mutants, mitochondrial fission mutant dnm1Δ showed no increase of insertions (Fig. 1b). As mitochondrial fusion mutants lack mtDNA30, we examined insertions in wild-type cells that lost mtDNA (rho0) to check if merely the presence of mtDNA is required to suppress insertions. Wild-type rho0 showed only a minor increase in insertions compared to mitochondrial fusion mutants (Fig. 1b), suggesting that proper mitochondrial dynamics rather than mtDNA presence primarily suppresses genome instability. It was previously shown that cells treated with vacuolar-type ATPase (V-ATPase) inhibitor concanamycin A quickly degrade Fzo1 mitofusin and fragment mitochondria within hours31. Consistently, here we found that cells treated with concanamycin A for 2, 4, or 6 h showed an increased level of insertions similar and epistatic to mgm1Δ (Fig. 1d). Deficiencies in mitochondrial fusion genes are related to many human disorders including Parkinson’s disease, dementia, optic atrophy, neuropathies and others. Here we constructed leucine to proline mgm1-L273P mutant, equivalent to human OPA1-L331P mutant, one of the pathogenic variants resulting in autosomal dominant optic atrophy (ADOA) characterized by progressive loss of optic nerve and vision10,32, the most common hereditary optic neuropathy in humans. Again, we observed an increased level of insertions (Fig. 1e), albeit lower than in mgm1Δ null mutant. This could be due to a healthier mitochondrial network in the point mutant (Supplementary Fig. 1b), though an alternative explanation is discussed further below. We also constructed and tested several point mutants of FZO1. All three mutations (fzo1-K200A, fzo1-S201N and fzo1-V327T) target the GTPase domains, with the latter one corresponding to the human MFN2-I213T mutant, a pathogenic variant resulting in Charcot-Marie-Tooth disease3,33. These mutants showed higher levels of insertions from the nuclear genome compared to wild-type rho0 cells, but also lower when compared to the null mutant (Fig. 1e). This suggests that either Mgm1 and Fzo1 have additional functions apart from mediating mitochondrial fusion or that point mutants retain some activity. Finally, the elimination of Mdm30 or Psd1 that are important regulators of mitochondrial fusion34,35 show only very mild or no increase in insertions, respectively (Supplementary Data 2). Therefore, only mutants with the most severe loss of mitochondrial fusion show a high insertions phenotype.
Iron regulon upregulation and decreased ISC in mgm1Δ cells
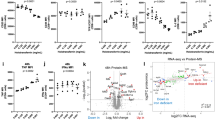
To gain insights into the possible reason for increased genome instability in fusion mutants, we performed bulk RNA-seq analysis in mitochondrial fusion mutants and found significant upregulation of iron regulon genes such as FIT2 or FIT3 (Fig. 2a). To further validate these results, we used a previously published iron regulon reporter, Aft1-responsive FIT2 promoter (FIT2pr) driving expression of GFP36. Aft1 is the major transcriptional activator of the iron regulon, controlling expression of genes involved in iron import, mobilization, and recycling37. When treated with iron chelator, bathophenanthrolinedisulfonate (BPS), to remove iron from the media, wild-type cells exhibited a strong GFP signal, confirming activation of the iron regulon (Supplementary Fig. 2a). As tested by fluorescence microscopy or flow cytometry, we observed a significant increase of GFP intensity in mgm1Δ cells, further confirming activation of the iron regulon. The control wild-type rho0 cells or aft1Δ showed only a slight increase or no signal, respectively (Fig. 2b, Supplementary Fig. 2a–c and Supplementary Data 4). Additionally, we assessed iron regulon activation by examining the fluorescence intensity of GFP-tagged iron regulon responsive genes FTR1, FET3 and ARN1 from their native promoters. Again, we observed increased GFP intensity in mgm1Δ cells, further confirming activation of the iron regulon (Fig. 2c). Interestingly, fluorescence microscopy analysis revealed that iron regulon upregulation in mgm1Δ mutant cells was not equal among all cells, with some cells carrying strong GFP signal and some weak signal. Such heterogeneity was particularly clear in cells expressing the FIT2pr-GFP reporter (Supplementary Fig. 2a). While the reason behind the heterogeneous response remains unknown, we note that unequal transcriptional responses to different stressors is common in all organisms (e.g refs. 38,39.).
a Examples of upregulated genes of the iron regulon in mitochondrial fusion-deficient mutants with three biological replicates of each strain, logFC - log fold-change. b FIT2pr-GFP fluorescence signal indicating iron regulon activation in arbitrary units (mean ± SD, n = 3 biological replicates of 10,000 events each, F = 39.53, W = 3612, df=3). P values were determined using two-tailed Brown-Forsythe and Welch ANOVA with Dunnett’s T3 multiple comparisons test. Exact cell counts are provided in Supplementary Data 4. c Intensity of iron uptake proteins (Ftr1-GFP, Fet3-GFP, and Arn1-GFP) in wild-type and mgm1Δ cells. Exposure time was the same in wild-type and mgm1Δ cells. Scale bars represent 5 µm. Experiments were repeated independently three times with similar results. d Percentage of cells containing nuclear Rps2-GFP foci and representative images from mgm1Δ cells. P values determined by two-tailed χ2 test, n = 1607, 1052, 1209, and 1000 total cells, respectively. Scale bar represents 5 µm. e Localization of ISC biogenesis proteins (Yfh1-GFP, Nfs1-GFP, Acp1-GFP, and Grx5-GFP) with MitoTracker Red mitochondrial staining in wild-type and mgm1Δ cells. Scale bar represents 5 µm. Experiments were repeated independently three times with similar results. f Percentage of cycling cells (excluding G0/G1 phase) with spontaneous Rad52-GFP foci and representative image from mgm1Δ cells. P values determined by two-tailed χ2 test, n = 1558, 1040, 1064, and 1058 total cycling cells, respectively. Scale bar represents 5 µm. Source data are provided as a Source Data file.
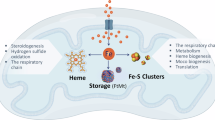
Aft1’s function in transcription regulation is not directly regulated by the levels of free iron but rather by ISC levels40. ISC biogenesis takes place in the mitochondria with many ISC-containing proteins being mitochondrial residents. Mitochondria are also required for production of cytosolic- and nuclear-targeted ISCs in yeast, via the export of a sulfur-containing precursor that is matured in the cytosol. Many of the proteins containing iron cofactors play essential roles in promoting DNA replication and protecting genome stability41,42,43,44,45,46. To test the sufficiency of nuclear ISC proteins, we used a previously established reporter assay in which the ribosomal protein Rps2 is tagged with GFP. Rps2 is exported from the nucleus by the essential protein Rli1, a process that requires Rli1 to bind an ISC. Therefore, accumulation of Rps2-GFP in the nucleus has been used as a measure of Rli1 activity and nuclear ISCs in general47,48. We found a 6-fold higher level of Rps2-GFP nuclear foci consistent with deficient ISCs in mgm1Δ mutant cells. Wild-type rho0 cells exhibited less than 3-fold more nuclear foci than wild-type rho + (Fig. 2d and Supplementary Fig. 3). Also, we observed a lower level of nuclear Rps2-GFP in mgm1-L273P compared to mgm1Δ null mutant, suggesting a possible explanation for the lower frequency of insertions in mgm1-L273P cells. Next, we compared the localization of mitochondrial ISC biogenesis proteins (Nfs1, Yfh1, Acp1, Grx5) fused with GFP in wild-type and mgm1Δ cells. All ISC biogenesis proteins were localized to fragmented and globular mitochondria in mgm1Δ. This result suggests that decreased ISC biogenesis does not result from altered localization of ISC proteins to mitochondria (Fig. 2e). Finally, we also analyzed the level of nuclear Rad52-GFP foci that indicates the level of spontaneous DNA damage. We found a significant ~3-fold increase in Rad52-GFP foci in mgm1Δ cells compared to wild-type cells. This effect was also specific to loss of fusion, as elimination of fission in dnm1Δ did not exhibit increased Rad52-GFP foci. (Fig. 2f).
It was previously shown that proper mitochondrial membrane potential (MMP) is important for ISC biogenesis49,50. Mitochondrial fusion mutants have decreased MMP in yeast and humans as previously shown (e.g ref. 51.) and measured here using 3,3′-dihexyloxacarbocyanine iodide (DiOC6) staining (Fig. 3a and Supplementary Data 4). It was previously shown that the atp1−111 allele of the α-subunit of the F1Fo-ATP synthase with increased ATP hydrolysis activity improves MMP in respiration-deficient rho0 cells and that this increased MMP correlates with better growth and better iron homeostasis50,52. To improve MMP of mgm1Δ cells, we constructed double mutant atp1-111 mgm1Δ. This double mutant showed better overall growth, partially suppressed insertions, lower iron regulon activation, and improved mitochondrial morphology (Fig. 3a–e, Supplementary Fig. 2a). A previous study identified several small compounds that improve MMP53. We found that one of these compounds, ST005213, improved MMP in both wild-type and mgm1Δ cells and decreased the number of globular mitochondria in mgm1Δ (Supplementary Fig. 4a-d and Supplementary Data 4). Higher MMP in wild-type cells increased insertions from Ty1 cDNA, suggesting an impact on retrotransposition. However, in mgm1Δ cells ST005213 significantly reduced insertions from the nuclear genome (Supplementary Fig. 4e). These results suggest that iron imbalance and genome instability in fusion mutants can partially stem from decreased MMP and provide a potential therapeutic strategy in patients affected by mitochondrial fusion mutations.
a DiOC6 fluorescence signal indicating MMP in arbitrary units (mean ± SD, n = 3 biological replicates of 10,000 events each, F = 26.35, W = 150.0, df = 6). P values were determined using two-tailed Brown-Forsythe and Welch ANOVA with Welch’s correction. Exact cell counts are provided in Supplementary Data 4. b Serial dilution on YPED media demonstrating improved growth rate in atp1-111 mgm1Δ double mutant compared to mgm1Δ single. The experiment was conducted three times with similar results. c Frequency of inserted DNA at DSBs in mgm1Δ mutants with improved mitochondrial membrane potential. P values were determined using two-tailed χ2 test. n – number of NHEJ products tested is shown in Supplementary Data 1–3. d Percentage of cells containing globular, fragmented, or tubular mitochondrial networks (n = 169, 193, 207, 181, 154, 198 total cells, respectively) and representative images of each morphology in strains carrying mitochondria-localized GFP. The scale bar represents 2 µm. e FIT2pr-GFP fluorescence signal indicating iron regulon activation (mean ± SD, n = 3 biological replicates of 10,000 events each, F = 27.10, W = 1935, df = 6). P values were determined using two-tailed Brown-Forsythe and Welch ANOVA with Dunnett’s T3 multiple comparisons test. Exact cell counts are provided in Supplementary Data 4. f Percentage of cells containing Rps2-GFP nuclear foci. P values determined by two-tailed χ2 test, n = 1607, 1052, 1209, 1510, and 1032 total cells, respectively. Source data are provided as a Source Data file.
It was also demonstrated that mitochondrial fusion mutant phenotypes can be suppressed by elimination of mitochondrial fission54,55. We therefore constructed a double mutant dnm1Δ mgm1Δ that has better overall mitochondrial morphology55,56 compared to single mgm1Δ mutant. In addition to drastically reducing insertions, the dnm1Δ mgm1Δ mutant also had a significant reduction in iron regulon activation and improved ISC formation; however, MMP was not significantly improved (Fig. 3a, c–f, Supplementary Figs. 2a and 3). In summary, our results indicate that globular, aggregated mitochondria result in iron dyshomeostasis that likely further promotes nuclear genome instability such as insertions.
Iron homeostasis mutants have increased templated insertions
To test whether mitochondrial iron homeostasis deficits or broadly many mitochondrial deficits can lead to nuclear genome instability, we tested ~50 additional mitochondrial mutants (Supplementary Data 2). All of these mutants were also tested for respiration proficiency by plating on media containing a non-fermentable carbon source (Supplementary Data 2). Among mutants related to iron homeostasis, we tested grx5Δ, a mutant of glutathione-dependent oxidoreductase responsible for transfer of [2Fe-2S] ISC from scaffold proteins to target proteins or for further ISC synthesis; frataxin mutant yfh1Δ, deficient in initial ISC assembly and related to human neurodegenerative disorder Friedreich’s ataxia; isu1Δ, involved in scaffolding for initial ISC formation; and isa1Δ and iba57Δ, partially deficient in transfer of [4Fe-4S] ISCs to mitochondrial target apoproteins. We also tested mutants of the ERMES complex (mdm10Δ, mdm12Δ, mdm34Δ, mmm1Δ) responsible for ER-mitochondria contact that previously showed iron regulon upregulation indicating low ISCs57. Nearly all these ISC biogenesis mutants showed increased levels of insertions (~ 5-to-60-fold increase of insertions from nuclear genome), suggesting that ISC formation is important to prevent insertions (Fig. 4a). The exception was ISC biogenesis mutant isu1Δ (rho+), which is likely due to functional compensation from its paralog, ISU2. Among other ISC biogenesis mutants, isa1Δ and iba57Δ showed the lowest increase of insertions. Isa1 and Iba57 are late-acting ISC assembly proteins needed for maturation of mitochondrial [4Fe-4S] proteins but do not impair cytosolic-nuclear Fe/S proteins58. Thus, the high insertions phenotype of grx5Δ or yfh1Δ likely reflects their importance in cytosolic-nuclear Fe/S proteins. Finally, in ISC biogenesis mutants, as in fusion mutants, insertions originated preferentially from fragile regions of the genome (Fig. 4b).
a Frequency and types of inserted DNA at DSBs in ISC defective mutants. P values were determined using two-tailed χ2 test. n – number of NHEJ products tested shown in Supplementary Data 1 and 2. b Features analysis of the enriched donor DNAs for templated insertions in ISC mutants. Empirical P values were calculated using two-sided permutation test by comparing the observed value to 10,000 random values generated by permutations. Error bars represent 2 × standard deviations of the permuted distributions as 95% confidence intervals. c FIT2pr-GFP fluorescence signal indicating iron regulon activation (mean ± SD, n = 3 biological replicates of 10,000 events each, t = 20.4, df = 4). P values were determined using two-tailed unpaired t-test. mmr1Δ cells are rho+ in this genetic background. Exact cell counts are provided in Supplementary Data 4. d Percentage of cells containing Rps2-GFP nuclear foci. P values determined by two-tailed χ2 test, n = 1607, 1052, 1357, 1172, and 1131 total cells, respectively. Source data are provided as a Source Data file.
The other >30 mitochondrial mutants tested here included mutants with deficits in organelle partitioning, inter-organellar communication, mitochondrial transcription/translation, mitochondrial genome maintenance, ion transport, and metabolism (Supplementary Data 2). Among all these mutants, the only one that showed a similar pattern of increased insertions to the fusion mutants was mitochondrial partitioning mutant mmr1Δ (Fig. 4a). Previous work documented iron imbalance in mmr1Δ cells59. Consistently, we found upregulation of the iron regulon via the FIT2pr-GFP reporter assay and by assessing Ftr1-GFP fluorescence signal (Fig. 4c, Supplementary Figs. 2a and 5). We also observed decreased ISC production in mmr1Δ, though not as strongly as in mutants directly related to ISC production like grx5Δ (Fig. 4d, Supplementary Fig. 3). In mmr1Δ cells, the insertion level is higher than in grx5Δ even though the nuclear ISC levels are not as diminished and the iron regulon is not as strongly activated (Supplementary Figs. 2a, d and 3). We cannot exclude the possibility that additional deficits in mmr1Δ, such as its role in mitochondrial partitioning60,61, can contribute to genome instability. In summary, mitochondrial mutants that have the highest level of insertions are all directly or indirectly related to iron homeostasis.
Very low or excess iron can result in increased templated insertions
Mutants with low ISC levels accumulate iron due to constitutive activation of Aft162,63,64,65. Excess labile iron is toxic to cells due to its ability to generate high levels of ROS through the Fenton reaction. These ROS can further damage ISCs and render ISC-containing proteins catalytically inactive66,67,68. Considering the multi-faceted roles of iron in the cell, we asked if the increase of insertions is related to the partial loss of ISC or excessive iron accumulation, or both. Toward this goal, we eliminated Aft1 responsible for iron uptake in response to low ISC in mgm1Δ or grx5Δ, or increased the iron load in mgm1Δ or grx5Δ cells by adding supplementary iron to the media. Unexpectedly, we found that simply decreased iron homeostasis (uptake, recycling, mobilization) in aft1Δ single mutant had a dramatic effect on insertions. This result was further confirmed by demonstrating a similar outcome in wild-type cells treated with iron chelator BPS (Fig. 5a). Supplemental iron in aft1Δ cells completely suppressed the high level of insertions, confirming that Aft1’s role in iron homeostasis is required to suppress insertions. We note that the high level of insertions in Aft1 deficient cells (rho+) is not related to respiration, as respiration deficient aft1Δ cells (rho−) still have high levels of insertions (Supplementary Data 2). Thus, basal homeostatic control of iron availability by Aft1 is essential for genome stability. This could be related to lower levels of iron cofactors in aft1Δ cells (Fig. 4d). Furthermore, we found that double mutants mgm1Δ aft1Δ and particularly grx5Δ aft1Δ showed higher levels of insertions from the nuclear genome when compared to single mutants (Fig. 5a). Therefore, decreased iron uptake in mutants with low ISC does not rescue high levels of insertions, but rather aggravates genome instability. This increase may result from an even greater reduction in ISC production in double mutants with both impaired ISC biosynthesis and reduced iron uptake. Next, we tested the impact of supplemental iron in wild-type, mgm1Δ, or grx5Δ cells and found increased levels of insertions in mutants that have a constantly active iron regulon (Fig. 5a), but not in wild type. Therefore, excess supplemental iron in cells with an active iron regulon and low iron cofactors stimulate genome instability. To further test it, we constructed ccc1Δ that fails to store excess iron in vacuoles and accumulates iron in cytosol and mitochondria69,70 but is respiration proficient. In ccc1Δ alone, there was a modest increase in insertion frequency compared to wild type. However, when treated with additional iron, ccc1Δ exhibited a ~31-fold increase of insertions compared to wild type, indicating that failure to appropriately sequester reactive iron in the vacuoles can also lead to genome instability (Fig. 5b). Furthermore, we assessed ccc1Δ cells for ISC sufficiency by counting nuclear-localized Rps2-GFP and found that while ccc1Δ cells alone had wild-type levels of Rps2-GFP nuclear foci, when excess iron was added to the media there was a large increase in foci in ccc1Δ (Fig. 5c, Supplementary Fig. 3). Thus, excess cytoplasmic iron alone is sufficient to promote insertions either by generating ROS or interfering with ISC biogenesis or both. We note that we have not tested ISC levels in mitochondria in ccc1Δ cells. In summary, these results indicate that diverse iron homeostasis deficits, including low iron availability, low ISC biogenesis, or overload of cytoplasmic iron, can result in genome instability manifested as insertions.
a, b Frequency and types of inserted DNA at DSBs. “+Fe” indicates treatment with 3 mM ammonium iron (II) sulfate for 6 h prior to plating on galactose. “+BPS” indicates treatment with 250 mM BPS iron chelator overnight prior to plating on galactose. P values were determined using two-tailed χ2 test. n – number of NHEJ products tested shown in Supplementary Data 1–3. c Percentage of cells containing Rps2-GFP nuclear foci. “+Fe” indicates treatment with 3 mM ammonium iron (II) sulfate for 6 h prior to imaging. P values determined by two-tailed χ2 test, n = 1607, 1120, 1009 total cells, respectively. d Frequency of NUMTs at DSBs. “+BPS” indicates treatment with 250 mM BPS iron chelator overnight prior to plating on galactose. P values were determined using two-tailed χ2 test. n – number of NHEJ products tested shown in Supplementary Data 1–3. e Distribution of insertion donor DNA in indicated mutants. n – number of independent insertion donor DNA. Source data are provided as a Source Data file.
Interestingly, a very high level of mtDNA insertions called NUMTs (nuclear mitochondrial segments) was observed in aft1Δ cells (>700-fold increase compared to wild type) and in cells treated with iron chelator BPS (>350-fold increase) (Fig. 5d). A total of ~600 mtDNA insertions were found in aft1Δ and ~300 NUMTs in BPS treated cells. Nuc1 (human EndoG) is responsible for most of the NUMTs in yeast stationary phase cells. Nuc1 degrades large mtDNA fragments and the remaining short products of this degradation are inserted at DSBs17. In aft1Δ nuc1Δ double mutant, NUMTs greatly decreased, confirming a major role of yeast EndoG in formation of NUMTs (Fig. 5d). Therefore, iron depletion, similar to prolonged starvation17, induces mtDNA release and its degradation by Nuc1 that results in NUMT formation. We note that many mitochondrial mutants tested above have completely lost mtDNA and therefore cannot be assessed for transfer of mtDNA to the nucleus. In addition to aft1Δ, a significant increase of NUMTs was observed in the ISC biosynthesis mutants isa1Δ and iba57Δ that showed >30-fold increase compared to wild type (~100 NUMTs together) and mitochondrial ribosomal protein mutant mrpl37Δ and mrs3Δ (>10-fold increase) (Fig. 5d). All these mutant cells are respiration deficient rho− cells, suggesting that mtDNA fragments transferred to the nucleus originate from a rearranged mitochondrial genome. It is possible that similar NUMTs can occur in human cells with low iron, cells treated with iron chelators, or in cells with compromised mitochondrial ISC biogenesis. The overall distribution of inserted donor DNA of different origins in mutants with high levels of insertions is shown in Fig. 5e.
ROS damage increases templated insertions
Iron is indispensable for enzymatic functions in the electron transport chain, for enzymes protecting cells from oxidative damage (catalase, peroxidases, superoxide dismutases), and for many enzymes involved in DNA replication and repair45,71. Excess iron also induces DNA damage through its high reactivity with hydrogen peroxide (H2O2). Consequently, cells with altered iron homeostasis, such as low or high iron levels or decreased ISC biogenesis, have increased spontaneous levels of oxidative DNA damage and are sensitive to oxidative DNA damage-inducing agents (e.g refs. 72,73.). Mitochondrial fusion mutants were also reported to have higher levels of spontaneous oxidative DNA damage74,75. Here we considered the possibility that increased insertions in fusion mutants and in iron homeostasis mutants in general could stem from ongoing spontaneous ROS damage and compromised DNA repair/replication enzymes that require iron cofactors. In support of this premise, we found that fusion mutants mgm1Δ or mgm1-L273P show decreased growth rate in the presence of H2O2. aft1Δ mutant is also sensitive to ROS damage (Fig. 6a). Second, we compared the effect of H2O2 on DNA damage Rad52-GFP foci formation in mgm1Δ and wild-type cells. When treated with H2O2, both wild type and mgm1Δ exhibit an increase in Rad52-GFP foci. However, when the H2O2 is washed out of the media and cells are allowed to recover for two hours, Rad52-GFP foci return nearly to baseline in wild type while remaining elevated in mgm1Δ above spontaneous levels (Fig. 6b). These findings confirm that mgm1Δ cells are more susceptible to ROS induced damage and are unable to repair it in a timely manner. Third, H2O2 treatment has a synergistic effect on insertions in iron homeostasis mutants (Fig. 6c). These results confirm that ROS damage stimulates insertions and may be responsible for insertions in iron homeostasis mutants. Consistently, treatment of mgm1Δ cells with antioxidant glutathione (GSH) mildly reduced insertions by ~1.5-fold (Fig. 6d). We also tested whether H2O2 treatment could increase insertions in mutants deficient in DNA damage response genes that promote resistance to H2O2 but do not strongly promote insertion formation on their own. We treated homologous recombination deficient rad51Δ mutant and DNA damage checkpoint deficient mec1Δ sml1Δ tel1Δ mutant cells with H2O2 and found very substantial increases of insertions in these mutants, indicating the importance of ROS damage repair in preventing insertions (Supplementary Fig. 6a). We further considered whether other DNA-damaging agents besides H2O2 could increase insertions in mutants with iron imbalance. We treated wild-type and mgm1Δ cells with camptothecin (CPT, Top1 inhibitor) and hydroxyurea (HU, reduces dNTPs by inhibiting ribonucleotide reductase - RNR). Wild-type cells treated with CPT exhibited a small increase in insertions, while HU treated cells showed a substantial 10-fold increase. In mgm1Δ cells, CPT treatment had no effect, but HU treatment resulted in >7-fold increase in insertions (Supplementary Fig. 6b). HU, apart from its effect on RNR, is also a weak iron chelator, activates the Aft1 iron regulon and Yap1-mediated ROS damage response pathway, and decreases the activity of ISC-containing enzymes76,77. These results suggest that not all DNA-damaging agents induce insertions; instead, insertions are preferentially triggered by agents that generate ROS damage and/or disrupt iron homeostasis. We also tested tsa1Δ mutant of peroxiredoxin that has very high levels of spontaneous ROS damage and increased genome instability78. Unlike iron homeostasis mutants, tsa1Δ did not exhibit increased insertions (Fig. 6d). We conclude that increased spontaneous ROS alone is insufficient to induce insertions; however, higher ROS stress coupled with compromised iron homeostasis promotes insertions.
a Doubling times (min) of cells with or without hydrogen peroxide treatment (mean ± SD; n = 3 biological replicates, F = 726.0, df=17). “+H2O2” indicates treatment with 0.5 mM hydrogen peroxide for 9 h. P values were determined using one-way ANOVA with Tukey’s multiple comparisons test. b Percentage of cycling cells (excluding G0/G1 phase) with Rad52-GFP foci indicating DSBs. “+H2O2” indicates treatment with 0.5 mM hydrogen peroxide for 4 h. “+Recovery” indicates 2 h washout with fresh media after H2O2 treatment. P values determined by two-tailed χ2 test, n = 1558, 1639, 1090, 1064, 1209, and 1026 total cycling cells, respectively. c Frequency of insertions with oxidative stress. “+H2O2” indicates treatment with 0.5 mM hydrogen peroxide for 4 h prior to plating on galactose. Fold-change of total insertions indicated. n – number of NHEJ products tested shown in Supplementary Data 1 and 3. d Frequency of insertions in indicated strains and conditions. “+GSH” indicates treatment with 10 mM glutathione for 6 h prior to plating on galactose. P values determined by two-tailed χ2 test n – number of NHEJ products tested shown in Supplementary Data 1 and 3. Source data are provided as a Source Data file.
ROS damage is repaired by base excision repair (BER). We therefore asked if major nucleases involved in repair of oxidative DNA damage, Apn1/Apn2 (apurinic/apyrimidinic endonucleases) or Ntg1/Ntg2 (apurinic/apyrimidinic lyases), are needed for the formation of insertions. Previous studies showed that the human homolog of Apn1, APE1 nuclease, can mediate fragmentation of chromosomes during chromothripsis79. Elimination of nucleases Apn1 and Apn2 or Ntg1 and Ntg2 showed only a slight decrease of insertions, suggesting redundancy or that additional enzymes may operate to process ROS damage leading to formation of insertions (Supplementary Fig. 6c). All insertions in mgm1Δ depend on key NHEJ enzyme Yku70 and show microhomologies at both ends typical for NHEJ (Supplementary Fig. 7a, b and Supplementary Data 3), similar to previously reported insertions (e.g refs. 15,16,18.). The percentage of inserted DNA originating from essential genes is proportional to the percentage of essential genes in yeast, demonstrating that insertions represent “copy-paste” and not “delete-paste” types of events as deletions within essential genes would be lethal in yeast haploid cells (Supplementary Fig. 7c). Together, we propose that insertions can arise from processing of spontaneous oxidative DNA damage in mgm1Δ and other iron imbalance mutants resulting in DNA fragments that are subsequently inserted at DSBs.
Discussion
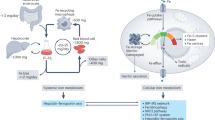
The goal of this work was to identify mutants that predispose cells to increased levels of insertions. Unexpectedly, mutants in DNA repair pathways, DNA damage checkpoint, or replication tested here have little to no impact on the level of insertions. Also, we observed modest or no impact on insertions after exposure of wild-type cells to high levels of DNA damage or in a mutant with very high levels of spontaneous ROS. Instead, we found mitochondrial fusion mutants to have high levels of insertions coinciding with iron imbalance and low ISC. The reason for lower ISC biogenesis in mitochondrial fusion mutants is not known but could be related to decreased contact and metabolite exchange with other organelles, such as the ER, decreased MMP, or other factors80. We cannot exclude the possibility that deficits unrelated to iron balance in mitochondrial fusion mutants, such as in energy homeostasis or other functions, could contribute to the high insertion phenotype. The reason we favor the idea of iron imbalance promoting insertions in mitochondrial fusion mutants is that many mutants directly affecting iron homeostasis exhibit a high insertion phenotype. Three types of mutants related to iron homeostasis showed increased insertions (Fig. 7): (i) mutants affecting ISC biogenesis that activate iron uptake and therefore also show an overload of iron, (ii) a mutant with decreased basic iron uptake and recycling, and (iii) a mutant that does not sequester iron in vacuoles and therefore overloads it in cytoplasm. In addition, cells treated with strong iron chelator, BPS, or with weak iron chelator that decreases the activity of ISC-carrying enzymes, HU77, also showed increased levels of insertions. These results suggest that there are unique features of iron dyshomeostasis that predispose cells to insertions.
Model is based on previous literature and our findings describing mechanism by which different iron dysregulation types may contribute to genome instability. Red arrows indicate direction of iron species transport. See main text for more details. a In mutants with decreased ISC production, iron regulon is upregulated leading to increased iron uptake and mitochondrial iron overload. Iron overload results in increased ROS, which can further interfere with ISC biosynthesis. Low ISC decreases activity of DNA repair and replication enzymes carrying ISC. Supplemental iron and ROS-damaging agent potentiate, while antioxidant suppresses, templated insertions in these mutants. b Excess of iron in ccc1Δ mutant lacking major iron storage and detoxification vacuolar pathway results in increased cytoplasmic iron, increased ROS, and decreased nuclear ISC. c In the aft1Δ mutant, which is generally defective in iron homeostasis, iron uptake, recycling, and mobilization are dysregulated, leading to decreased ISC production and increased ROS-induced damage. Supplemental iron rescues the phenotype of aft1Δ. d Cells grown in media containing strong iron chelator BPS have decreased ISC.
How does iron imbalance lead to insertions? First, proper iron levels are essential to maintain iron cofactors of many enzymes directly involved in generating nucleotides, and in DNA repair or replication. For example, major replicative polymerases with decreased ISC binding show lower stability46, and decreased levels of replicative polymerases induce genome instability with hotspots at positions described here81. Second, iron imbalance results in ROS induced DNA damage, and ISC themselves are intrinsically sensitive to oxidative stress. Many BER enzymes involved in repair of ROS damage contain ISC. Third, proper iron balance is needed for cell cycle progression, and low iron or low ISC results in prolonged G1 phase that can stimulate repair of DSBs by NHEJ, a pathway needed for insertions45,82. Therefore, a single mutation affecting iron balance has a broad impact on genome instability as it generates a situation in which DNA damage increases while the ability to repair this damage decreases at many levels.
All three types of iron homeostasis defects that increase insertions are observed in humans and are related to many diseases and cancer. Patients with iron deficiency anemia show increased levels of ROS and DNA damage22. Mutations in genes regulating iron uptake and storage result in hereditary hemochromatosis associated with excessive DNA damage and ~200-fold increase of liver cancer incidence83,84. Iron overload is a general hallmark of cancer24. Fast proliferating cancer cells require increased iron to meet the high demand for ATP production, oxygen transport, DNA replication, and other essential iron-dependent processes. While beneficial for fast proliferation, higher iron levels come at the cost of increased ROS and DNA damage. Also, proper storage of iron in lysosomes is essential for human health85. Downregulation of ISC biogenesis in humans, similar to yeast, can induce an iron starvation response, overload of iron, and ROS increase (e.g86.). For example, hypoxia in cancer cells leads to a decrease of ISC assembly proteins, increased iron uptake, and ROS (e.g87,88.). Finally, sublethal ferroptosis or pharmacological treatments involving iron chelators or supplementation may cause transient iron imbalance, potentially promoting genome instability. In summary, we propose that insertions and other types of CNVs observed across many cancer genomes may result from iron imbalance rather than solely from the loss of individual DNA repair genes. Furthermore, iron imbalance can modify and exacerbate genome instability associated with mutations in DNA repair genes.
Methods
Yeast strains
Yeast strains used to determine frequency of insertions at HO endonuclease-induced DSBs are derivatives of JKM139 strain, a gift from James Haber. Strains used for Rad52-GFP nuclear foci analyses, MMP measurements, and ROS sensitivity assays are also JKM139 derivatives. All strains used to study iron regulon activation via Aft1-mediated activation of GFP under the control of a FIT2 promoter are derivatives of AHY5005. yJL285 was generated by crossing yYY703 with yJL44. All strains used to study nuclear ISC sufficiency by Rps2-GFP nuclear foci are derivatives of KC481, a gift from Brian Wasko. All strains for microscopy analysis of GFP-tagged iron uptake and ISC biogenesis proteins are derivatives of BY4741. The mgm1-L273P, fzo1-K200A, fzo1-S201N, and fzo1-V327T point mutants were constructed with CRISPR/Cas9. DNA oligos corresponding to each target sequence were duplexed and ligated into plasmid pJH2972 (a gift from James Haber, derived from pbRA90 (Addgene plasmid #100951; http://n2t.net/addgene:100951; RRID:Addgene_100951) marked with URA3) digested with BplI. The template oligos was transformed into cells with the ligated plasmids to introduce each base substitution and confirmed by Sanger sequencing. The atp1-111 allele was introduced using the plasmid pCORE-UK, a gift from Michael Resnick and Francesca Storici (Addgene plasmid # 72238; http://n2t.net/addgene:72238; RRID:Addgene_72238). First, the KlURA3-kanMX double marker was amplified using primers Atp1-pCORE-Fw and Atp1-pCORE-Rw and integrated into the ATP1 locus. Then, a fragment of ATP1 with a point mutation (V111F) was amplified using primers ATP1-F5/ ATP1-R5 from the strain KWY99, a gift from Peter Thorsness, and transformed into cells to replace KlURA3-kanMX. The mutation was confirmed by Sanger sequencing. Cells were rendered rho0 by growing twice to saturation in synthetic complete media containing ethidium bromide and isolating clones that failed to grow when replica plated on non-fermentable glycerol-containing media. Mitochondrial morphology was assessed by transforming plasmid pYX142-mtGFP, a gift from Benedikt Westermann (Addgene plasmid #45050; http://n2t.net/addgene:45050; RRID:Addgene_45050). All strains used are listed in Supplementary Data 5. All primers and plasmids used are listed in Supplementary Tables 1 and 2, respectively.
Yeast growth, media, and DSB induction
To prepare samples for amplicon sequencing by Break-Ins analysis, cells were grown to saturation overnight in YEPD (1% yeast extract, 2% peptone, 2% dextrose), washed twice with YEP-Raffinose (1% yeast extract, 2% peptone, 2% raffinose), and inoculated in 12 mL YEP-Raffinose overnight. Once cultures reached a density of 2 × 107 cells/mL, 500 µL culture was plated on six 150 × 15 mm YEP-Gal plates (1% yeast extract, 2% peptone, 2.5% agar, 2% galactose. Galactose was filter sterilized and added to the autoclaved media) per strain. In addition, ~100 cells were plated on YEPD plates and replica plated to glycerol plates to determine the proportion of respiration-competent cells at the time of plating. Additional treatments with various additives were conducted using the timings and concentrations indicated in the figure legends. All treatments with H2O2 were conducted in the dark. For iron treatments, YEP-Raffinose media was treated overnight with 80 µM BPS to chelate iron before adding 3 mM ammonium iron (II) sulfate. Iron-treated cultures were grown overnight in standard YEP-Raffinose to log phase, then the media was replaced with the iron-treated media and grown for an additional four hours to 2 × 107 cells/mL. Cultures for expression analysis, flow cytometry, microscopy, and ROS sensitivity assays were grown to saturation overnight in YEPD, washed with YEP-raffinose, and grown to log phase in YEP-raffinose prior to each experiment. For mitochondrial morphology assessments, cells were grown to saturation in synthetic complete media lacking leucine prior to growth to log phase in YEP-Raffinose. Concentrations and durations of drug treatments are indicated in the figure legends. Incubations were carried out at 30 °C unless otherwise indicated.
Library preparation and Break-Ins analysis
Library preparation was conducted for Break-Ins analysis as previously described17. Briefly, after DSB induction on YEP-Gal plates, colonies were grown for 5–7 days at 30 °C. Colonies representing individual NHEJ repair products were counted, collected, and pooled for each strain. Genomic DNA from ~100 µL cells per sample was isolated according to a standard glass beads and phenol/chloroform extraction method and DNA was treated overnight with 0.1 mg/mL RNase A and normalized to 10 ng/µL in water. Library preparation was conducted according to an adapted Illumina amplicon sequencing protocol #15044223 Rev. B. The MATa locus was amplified with homemade barcoded primers added with a high-fidelity polymerase (KAPA HiFi HotStart ReadyMix (Roche, 7958927001)) to prevent index hopping prior to cleanup with AMPure XP beads (Beckman Coulter A63880) and the addition of barcoded Illumina primers (Nextera XT V2 Index, Illumina FC-131-2001) by high-fidelity PCR and final bead purification. The average size of the DNA was determined by TapeStation and barcoded samples were equally pooled in a 4 nM final library and sequenced with PhiX library (Illumina, FC-110-3001) on the Illumina MiSeq platform with MiSeq Reagent Kit v3 (600 cycle) (Illumina, MS-102-3003).
Fluorescence microscopy
Images were acquired with a BX51 fluorescence microscope (Olympus) equipped with an Axiocam 506 monochromatic camera (Carl Zeiss) and 100× oil immersion objective. Images were acquired and processed with ZEN software (Carl Zeiss). For all experiments, 800 µL of culture was pelleted then washed and resuspended in 1X PBS prior to imaging. Rps2-GFP nuclear foci were scored per total number of cells (n ≥ 1000) while Rad52-GFP foci were scored per total number of cycling cells, excluding those in G0/G1 (n ≥ 1000). For Rad52-GFP analyses with H2O2, cells were treated with 0.5 mM H2O2 for 4 h and either imaged or washed and resuspended in fresh YEP-Raffinose without H2O2 for an additional 2 h of recovery prior to imaging as described in figure legends. Mitochondrial morphologies were quantified per total number of mtGFP-expressing cells (n ≥ 150). Cells with reticular morphologies were designated “tubular,” those with several dispersed fragments were designated “fragmented,” and those with large aggregates were designated “globular,” as shown in Fig. 3d. Mitochondria were stained with 500 nM MitoTracker Red CMXRos (Cell Signaling Technology, 9082S) where indicated.
Flow cytometry
Cells were grown to log phase in YEP-Raffinose as described above, washed with 1X PBS, and resuspended in 1X PBS to ~1-3 million cells/mL. For MMP analyses, 0.1 µg/mL DiOC6 was added to log phase cells and incubated for 15 min in the dark. Cells were washed and resuspended in 1X PBS prior to analysis. GFP or DiOC6 fluorescence of 1 × 104 cells per sample in triplicate was analyzed on an Attune NxT flow cytometer (Thermo Fisher). Cells were differentiated from debris by forward and side scatter. Imaging cytometry using an Attune Cytpix flow cytometer (Thermo Fisher) was used to validate that doublet discrimination would be detrimental to downstream analysis. All cytometers were maintained by the Baylor College of Medicine Flow Cytometry Core Laboratory. Analyses were conducted in FlowJo (BD BioSciences). Gating strategies are shown in Supplementary Figs. 2a and 4c. Representative histograms are shown in Supplementary Figs. 2b and 4b.
Growth rate analysis
Strains were grown to saturation in 5 mL YEPD media overnight, washed with YEP-Raffinose, and diluted to an OD600 of 0.25 in 8 mL YEP-Raffinose media with or without 0.5 mM H2O2. Cells were incubated in the dark at 30 °C and OD600 was measured at 3, 6, and 9 h (n = 3 biological replicates). Doubling times were calculated using the following equation:
540 minutes* ln(2)/ln(Final OD600/Initial OD600).
Estimation of Ty1 transposition frequency
The rate of Ty1 retrotransposition was analyzed in strains carrying a Ty1-270his3-AI reporter. Cells were struck on YEPD plates for single colonies, and individual colonies were inoculated in 5 mL YEPD media incubated at 24 °C. Cells were diluted and grown to 1 × 107 cells/mL before plating 20–30 cells on YEPD and incubating at 24 °C for 6 days. 10-20 colonies per strain were diluted in water, spread on synthetic complete media lacking histidine, and incubated at 24 °C for 4 days before counting His+ colonies. Statistical comparisons were conducted using the Drake estimator and bootstrap resampling approach89.
Analysis of Ty1 cDNA amount
Cells were grown in 5 mL YEPD overnight at 24 °C. Cells were pelleted, washed with YEP-Raffinose, and inoculated in 25 mL fresh YEP-Raffinose and incubated for two days at 24 °C. Cells were collected and genomic DNA was harvested using glass bead disruption and standard phenol-chloroform extraction. DNA was treated overnight with RNaseA followed by digestion with PvuII-HF and separated on 0.8% agarose gel. Southern blotting and hybridization with radiolabeled DNA probes were conducted using a standard protocol. The Ty1 PvuII-SnaBI fragment of Ty1-H3 was prepared by PCR using primers PvuII-F1 (CTGTAAAAGCAGTAAAATCAATCAAAC) and SnaB1-R1 (GTATAGATTATTACCTGATACTTCATCTCT) and was used as a P32-labeled DNA probe. The intensities of bands corresponding to probed DNA fragments were analyzed using ImageQuant TL software (Amersham Biosciences) and normalized to four genomic Ty1 fragments.
Expression analysis
Cells were grown to 2 × 107 cells/mL as described in YEP-Raffinose and 500 µL were pelleted and stored at −80 °C. RNA was extracted using the miRNeasy Mini Kit (Qiagen, #217004) according to manufacturer instructions. Briefly, cells were lysed in a bead miller with acid-washed glass beads and Qiagen Buffer RLT. One volume of 70% ethanol was added to the homogenized lysate and the sample was purified using the RNeasy spin column and eluted in RNAse-free water. The cDNA library preparation and sequencing were conducted by Novogene using a paired-end 150 bp sequencing strategy on the Illumina NovaSeq 6000 platform. Paired-end reads were mapped to the yeast reference transcriptome (S288C) and counted using TopHat90 and HTseq-count91 with default parameters, respectively. A gene-count matrix was then processed using edgeR v3.40.292 for differential expression (DE) analysis in R v 4.2.2 and subsequently gene ontology enrichment analyses were performed using clusterProfiler v4.6.293.
Break-Ins analysis
Break-Ins analysis was described in detail17. In brief, first, PhiX reads were removed from paired-end raw sequencing reads with BBDuk v38.46 (https://sourceforge.net/projects/bbmap/) using default parameters. Subsequently, high-quality reads were retained with an average sequencing quality score (Q score) of each read greater than 25. Q scores were calculated by converting the quality symbol in the FASTQ files using the Quality Score Encoding table (http://support.illumina.com/help/BaseSpace_OLH_009008/Content/Source/Informatics/BS/QualityScoreEncoding_swBS.htm). Second, we detected reads with insertions of 10 bp or longer. The forward and reverse reads were merged with PEAR v0.9.1194 and then the merged reads were aligned to the yeast reference genome (S288C) using BLASTN v2.8.195. We kept merged reads that were aligned to the MATa sequence with at least 10 bp insertion at the break point as well as unmerged reads which had insertion portions at the 3’ ends and aligned to different loci ranging from 10 bp to 3 kb away from each other on the same chromosome. Third, we identified unique insertions by removing duplicates and contaminations due to either sequencing errors or short insertion length (<10 bp). Fourth, the locations of the donor sequences for the unique insertions were defined based on BLASTN mapping results. For insertion sequences mapped to more than one donor category, we selected the donor that covered the largest proportion of the insertion sequence. Finally, we identified microhomology sequences between MATa and insertion donor as having perfect match to both the terminal regions of the donor and the MATa sequences.
Features analysis
We analyzed genomic features either overlapped (≥1 bp overlap between insertions and genomic features) or located in the vicinity of the donor sequences using Bedtools v2.25396. The genomic features included in our analyses are confirmed origins of replication (ARSes; http://cerevisiae.oridb.org), R-loops, known tandem repeats (TRF; https://hgdownload.soe.ucsc.edu/goldenPath/sacCer3/bigZips/chromTrf.tar.gz), tRNA, telomere, and centromere (https://downloads.yeastgenome.org/). We performed two-sided permutation tests to determine the enrichment of the genomic features overlapping or in vicinity of the observed insertions by comparing to 10,000 genome-wide random controls generated with BEDTools shuffle96. For the enrichment analysis, empirical P values were calculated and reported as
P = (1 + (S ≥ S0))/(N + 1),
emp.P = 2 × min (P, 1 − P)
where S is the observed value and S0 is the random distribution generated by 10,000 sets of random controls, N is the total number of tests. For emp.P = 0 in our tests, we report emp. P < 0.0001.
Frequency of Ty1 and mtDNA insertions
The inserted sequences aligned to LTR retrotransposon (Ty1) were further realigned to the same Ty1 sequence (YPLWTy1-1) using BLASTN (-word_size 11 -evalue 0.01 -gapopen 5 -penalty −1 -perc_identity 60 -dust no -soft_masking false). We grouped the insertions as right LTR alignment when the Ty1 inserts map to both LTR regions. We further measured the frequency of nucleotide inserted per 100,000 independent NHEJ products at every Ty1 position, normalized by the number of colonies as 100,000*nt,i /n colonies, where nt,i is the number of times each Ty1 nucleotide was inserted and n colonies is the number of colonies. Frequency of mtDNA insertions were measured in the same way as for Ty1 but without the realignment step.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The Break-Ins and RNA-seq data has been archived at GEO under the accession numbers GSE277752 (https://urldefense.com/v3/__https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE277752 and GSE277766 (https://urldefense.com/v3/__https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE277766, respectively. Source data are provided with this paper.
Code availability
All original code has been deposited on GitHub. The code for insertion detection can be accessed at the following GitHub repository: https://github.com/gucascau/iDSBins.git. For short indel detection, the code can be accessed at this GitHub repository: https://github.com/gucascau/iDSBindel.git. The features related to insertion analysis have been uploaded to a dedicated GitHub repository, which can be found here: https://github.com/gucascau/LargeInsertionFeature.git.
References
Chan, D. C. Mitochondrial dynamics and its involvement in disease. Annu. Rev. Pathol. 15, 235–259 (2020).
Meeusen, S. et al. Mitochondrial inner-membrane fusion and crista maintenance requires the dynamin-related GTPase Mgm1. Cell 127, 383–395 (2006).
Hermann, G. J. et al. Mitochondrial fusion in yeast requires the transmembrane GTPase Fzo1p. J. Cell Biol. 143, 359–373 (1998).
Sesaki, H. & Jensen, R. E. Ugo1p links the Fzo1p and Mgm1p GTPases for mitochondrial fusion. J. Biol. Chem. 279, 28298–28303 (2004).
Harner, M. et al. The mitochondrial contact site complex, a determinant of mitochondrial architecture. EMBO J. 30, 4356–4370 (2011).
Chen, W., Zhao, H. & Li, Y. Mitochondrial dynamics in health and disease: mechanisms and potential targets. Signal Transduct. Target Ther. 8, 333 (2023).
Bonneau, D. et al. Early-onset Behr syndrome due to compound heterozygous mutations in OPA1. Brain 137, e301 (2014).
Spiegel, R. et al. Fatal infantile mitochondrial encephalomyopathy, hypertrophic cardiomyopathy and optic atrophy associated with a homozygous OPA1 mutation. J. Med. Genet. 53, 127–131 (2016).
Abrams, A. J. et al. Mutations in SLC25A46, encoding a UGO1-like protein, cause an optic atrophy spectrum disorder. Nat. Genet. 47, 926–932 (2015).
Delettre, C. et al. Nuclear gene OPA1, encoding a mitochondrial dynamin-related protein, is mutated in dominant optic atrophy. Nat. Genet 26, 207–210 (2000).
Alexander, C. et al. OPA1, encoding a dynamin-related GTPase, is mutated in autosomal dominant optic atrophy linked to chromosome 3q28. Nat. Genet. 26, 211–215 (2000).
Carelli, V. et al. Syndromic parkinsonism and dementia associated with OPA1 missense mutations. Ann. Neurol. 78, 21–38 (2015).
Wang, X. et al. Impaired balance of mitochondrial fission and fusion in Alzheimer’s disease. J. Neurosci. 29, 9090–9103 (2009).
Shirendeb, U. et al. Abnormal mitochondrial dynamics, mitochondrial loss and mutant huntingtin oligomers in Huntington’s disease: implications for selective neuronal damage. Hum. Mol. Genet 20, 1438–1455 (2011).
Yu, Y. et al. Dna2 nuclease deficiency results in large and complex DNA insertions at chromosomal breaks. Nature 564, 287–290 (2018).
Yu, Y. et al. RPA and Rad27 limit templated and inverted insertions at DNA breaks. Nucleic Acids Res. 53, https://doi.org/10.1093/nar/gkae1159 (2025).
Yu, Y. et al. Yeast EndoG prevents genome instability by degrading extranuclear DNA species. Nat. Commun. 15, 7653 (2024).
Min, J. et al. Mechanisms of insertions at a DNA double-strand break. Mol. Cell 83, 2434–2448.e2437 (2023).
Lebedin, M. et al. Different classes of genomic inserts contribute to human antibody diversity. Proc. Natl. Acad. Sci. USA 119, e2205470119 (2022).
Li, Y. et al. Patterns of somatic structural variation in human cancer genomes. Nature 578, 112–121 (2020).
Carvalho, C. M. & Lupski, J. R. Mechanisms underlying structural variant formation in genomic disorders. Nat. Rev. Genet. 17, 224–238 (2016).
Aslan, M. et al. Lymphocyte DNA damage and oxidative stress in patients with iron deficiency anemia. Mutat. Res. 601, 144–149 (2006).
Federico, G. et al. NCOA4 links iron bioavailability to DNA metabolism. Cell Rep. 40, 111207 (2022).
Torti, S. V., Manz, D. H., Paul, B. T., Blanchette-Farra, N. & Torti, F. M. Iron and Cancer. Annu Rev. Nutr. 38, 97–125 (2018).
Moore, J. K. & Haber, J. E. Cell cycle and genetic requirements of two pathways of nonhomologous end-joining repair of double-strand breaks in Saccharomyces cerevisiae. Mol. Cell Biol. 16, 2164–2173 (1996).
Rapaport, D., Brunner, M., Neupert, W. & Westermann, B. Fzo1p is a mitochondrial outer membrane protein essential for the biogenesis of functional mitochondria in Saccharomyces cerevisiae. J. Biol. Chem. 273, 20150–20155 (1998).
Sesaki, H. & Jensen, R. E. UGO1 encodes an outer membrane protein required for mitochondrial fusion. J. Cell Biol. 152, 1123–1134 (2001).
Yu, X. & Gabriel, A. Patching broken chromosomes with extranuclear cellular DNA. Mol. Cell 4, 873–881 (1999).
Paquin, C. E. & Williamson, V. M. Temperature effects on the rate of ty transposition. Science 226, 53–55 (1984).
Stenger, M., Le, D. T., Klecker, T. & Westermann, B. Systematic analysis of nuclear gene function in respiratory growth and expression of the mitochondrial genome in S. cerevisiae. Micro. Cell 7, 234–249 (2020).
Goodrum, J. M., Lever, A. R., Coody, T. K., Gottschling, D. E. & Hughes, A. L. Rsp5 and Mdm30 reshape the mitochondrial network in response to age-induced vacuole stress. Mol. Biol. Cell 30, 2141–2154 (2019).
Weisschuh, N. et al. Mutation spectrum of the OPA1 gene in a large cohort of patients with suspected dominant optic atrophy: Identification and classification of 48 novel variants. PLoS One 16, e0253987 (2021).
Amiott, E. A., Cohen, M. M., Saint-Georges, Y., Weissman, A. M. & Shaw, J. M. A mutation associated with CMT2A neuropathy causes defects in Fzo1 GTP hydrolysis, ubiquitylation, and protein turnover. Mol. Biol. Cell 20, 5026–5035 (2009).
Fritz, S., Weinbach, N. & Westermann, B. Mdm30 is an F-box protein required for maintenance of fusion-competent mitochondria in yeast. Mol. Biol. Cell 14, 2303–2313 (2003).
Chan, E. Y. & McQuibban, G. A. Phosphatidylserine decarboxylase 1 (Psd1) promotes mitochondrial fusion by regulating the biophysical properties of the mitochondrial membrane and alternative topogenesis of mitochondrial genome maintenance protein 1 (Mgm1). J. Biol. Chem. 287, 40131–40139 (2012).
Hughes, C. E. et al. Cysteine toxicity drives age-related mitochondrial decline by altering iron homeostasis. Cell 180, 296–310.e218 (2020).
Ramos-Alonso, L., Romero, A. M., Martinez-Pastor, M. T. & Puig, S. Iron regulatory mechanisms in Saccharomyces cerevisiae. Front Microbiol 11, 582830 (2020).
Gasch, A. P. et al. Single-cell RNA sequencing reveals intrinsic and extrinsic regulatory heterogeneity in yeast responding to stress. PLoS Biol. 15, e2004050 (2017).
Meng, X., Reed, A., Lai, S., Szavits-Nossan, J. & McCarthy, J. E. G. Stochastic scanning events on the GCN4 mRNA 5’ untranslated region generate cell-to-cell heterogeneity in the yeast nutritional stress response. Nucleic Acids Res. 51, 6609–6621 (2023).
Hausmann, A., Samans, B., Lill, R. & Muhlenhoff, U. Cellular and mitochondrial remodeling upon defects in iron-sulfur protein biogenesis. J. Biol. Chem. 283, 8318–8330 (2008).
Gari, K. et al. MMS19 links cytoplasmic iron-sulfur cluster assembly to DNA metabolism. Science 337, 243–245 (2012).
Stehling, O. et al. MMS19 assembles iron-sulfur proteins required for DNA metabolism and genomic integrity. Science 337, 195–199 (2012).
Lill, R. et al. Mechanisms of iron-sulfur protein maturation in mitochondria, cytosol and nucleus of eukaryotes. Biochim. Biophys. Acta 1763, 652–667 (2006).
Puig, S., Ramos-Alonso, L., Romero, A. M. & Martinez-Pastor, M. T. The elemental role of iron in DNA synthesis and repair. Metallomics 9, 1483–1500 (2017).
Zhang, C. Essential functions of iron-requiring proteins in DNA replication, repair and cell cycle control. Protein Cell 5, 750–760 (2014).
Netz, D. J. et al. Eukaryotic DNA polymerases require an iron-sulfur cluster for the formation of active complexes. Nat. Chem. Biol. 8, 125–132 (2011).
Kispal, G. et al. Biogenesis of cytosolic ribosomes requires the essential iron-sulphur protein Rli1p and mitochondria. EMBO J. 24, 589–598 (2005).
Chen, K. L. et al. Loss of vacuolar acidity results in iron-sulfur cluster defects and divergent homeostatic responses during aging in Saccharomyces cerevisiae. Geroscience 42, 749–764 (2020).
Kispal, G., Csere, P., Prohl, C. & Lill, R. The mitochondrial proteins Atm1p and Nfs1p are essential for biogenesis of cytosolic Fe/S proteins. EMBO J. 18, 3981–3989 (1999).
Veatch, J. R., McMurray, M. A., Nelson, Z. W. & Gottschling, D. E. Mitochondrial dysfunction leads to nuclear genome instability via an iron-sulfur cluster defect. Cell 137, 1247–1258 (2009).
Olichon, A. et al. Loss of OPA1 perturbates the mitochondrial inner membrane structure and integrity, leading to cytochrome c release and apoptosis. J. Biol. Chem. 278, 7743–7746 (2003).
Francis, B. R., White, K. H. & Thorsness, P. E. Mutations in the Atp1p and Atp3p subunits of yeast ATP synthase differentially affect respiration and fermentation in Saccharomyces cerevisiae. J. Bioenerg. Biomembr. 39, 127–144 (2007).
Montague, C. R., Fitzmaurice, A., Hover, B. M., Salazar, N. A. & Fey, J. P. Screen for small molecules increasing the mitochondrial membrane potential. J. Biomol. Screen. 19, 387–398 (2014).
Sesaki, H., Southard, S. M., Yaffe, M. P. & Jensen, R. E. Mgm1p, a dynamin-related GTPase, is essential for fusion of the mitochondrial outer membrane. Mol. Biol. Cell 14, 2342–2356 (2003).
Sesaki, H. & Jensen, R. E. Division versus fusion: Dnm1p and Fzo1p antagonistically regulate mitochondrial shape. J. Cell Biol. 147, 699–706 (1999).
Bernhardt, D., Muller, M., Reichert, A. S. & Osiewacz, H. D. Simultaneous impairment of mitochondrial fission and fusion reduces mitophagy and shortens replicative lifespan. Sci. Rep. 5, 7885 (2015).
Xue, Y. et al. Endoplasmic reticulum-mitochondria junction is required for iron homeostasis. J. Biol. Chem. 292, 13197–13204 (2017).
Muhlenhoff, U., Richter, N., Pines, O., Pierik, A. J. & Lill, R. Specialized function of yeast Isa1 and Isa2 proteins in the maturation of mitochondrial [4Fe-4S] proteins. J. Biol. Chem. 286, 41205–41216 (2011).
Lesuisse, E. et al. Genome-wide screen for genes with effects on distinct iron uptake activities in Saccharomyces cerevisiae. Genetics 169, 107–122 (2005).
Chelius, X. et al. Selective retention of dysfunctional mitochondria during asymmetric cell division in yeast. PLoS Biol. 21, e3002310 (2023).
Bockler, S. et al. Fusion, fission, and transport control asymmetric inheritance of mitochondria and protein aggregates. J. Cell Biol. 216, 2481–2498 (2017).
Muckenthaler, M. U., Galy, B. & Hentze, M. W. Systemic iron homeostasis and the iron-responsive element/iron-regulatory protein (IRE/IRP) regulatory network. Annu Rev. Nutr. 28, 197–213 (2008).
Miao, R., Holmes-Hampton, G. P. & Lindahl, P. A. Biophysical investigation of the iron in Aft1-1(up) and Gal-YAH1 Saccharomyces cerevisiae. Biochemistry 50, 2660–2671 (2011).
Yamaguchi-Iwai, Y., Dancis, A. & Klausner, R. D. AFT1: a mediator of iron regulated transcriptional control in Saccharomyces cerevisiae. EMBO J. 14, 1231–1239 (1995).
Babcock, M. et al. Regulation of mitochondrial iron accumulation by Yfh1p, a putative homolog of frataxin. Science 276, 1709–1712 (1997).
Imlay, J. A. The molecular mechanisms and physiological consequences of oxidative stress: lessons from a model bacterium. Nat. Rev. Microbiol. 11, 443–454 (2013).
Rodriguez-Manzaneque, M. T., Ros, J., Cabiscol, E., Sorribas, A. & Herrero, E. Grx5 glutaredoxin plays a central role in protection against protein oxidative damage in Saccharomyces cerevisiae. Mol. Cell Biol. 19, 8180–8190 (1999).
Rodriguez-Manzaneque, M. T., Tamarit, J., Belli, G., Ros, J. & Herrero, E. Grx5 is a mitochondrial glutaredoxin required for the activity of iron/sulfur enzymes. Mol. Biol. Cell 13, 1109–1121 (2002).
Li, L., Chen, O. S., McVey Ward, D. & Kaplan, J. CCC1 is a transporter that mediates vacuolar iron storage in yeast. J. Biol. Chem. 276, 29515–29519 (2001).
Lin, H., Li, L., Jia, X., Ward, D. M. & Kaplan, J. Genetic and biochemical analysis of high iron toxicity in yeast: iron toxicity is due to the accumulation of cytosolic iron and occurs under both aerobic and anaerobic conditions. J. Biol. Chem. 286, 3851–3862 (2011).
Paul, V. D. & Lill, R. Biogenesis of cytosolic and nuclear iron-sulfur proteins and their role in genome stability. Biochim Biophys. Acta 1853, 1528–1539 (2015).
Turrens, J. F. Mitochondrial formation of reactive oxygen species. J. Physiol. 552, 335–344 (2003).
Schilke, B., Voisine, C., Beinert, H. & Craig, E. Evidence for a conserved system for iron metabolism in the mitochondria of Saccharomyces cerevisiae. Proc. Natl. Acad. Sci. USA 96, 10206–10211 (1999).
Chen, L., Gong, Q., Stice, J. P. & Knowlton, A. A. Mitochondrial OPA1, apoptosis, and heart failure. Cardiovasc Res. 84, 91–99 (2009).
Iannielli, A. et al. Pharmacological Inhibition of Necroptosis Protects from Dopaminergic Neuronal Cell Death in Parkinson’s Disease Models. Cell Rep. 22, 2066–2079 (2018).
Dubacq, C. et al. Role of the iron mobilization and oxidative stress regulons in the genomic response of yeast to hydroxyurea. Mol. Genet. Genomics 275, 114–124 (2006).
Shaw, A. E. et al. Revised mechanism of hydroxyurea-induced cell cycle arrest and an improved alternative. Proc. Natl. Acad. Sci. USA 121, e2404470121 (2024).
Iraqui, I. et al. Peroxiredoxin Tsa1 is the key peroxidase suppressing genome instability and protecting against cell death in Saccharomyces cerevisiae. PLoS Genet. 5, e1000524 (2009).
Tang, S., Stokasimov, E., Cui, Y. & Pellman, D. Breakage of cytoplasmic chromosomes by pathological DNA base excision repair. Nature 606, 930–936 (2022).
de Brito, O. M. & Scorrano, L. Mitofusin 2 tethers endoplasmic reticulum to mitochondria. Nature 456, 605–610 (2008).
Zheng, D. Q. & Petes, T. D. Genome instability induced by low levels of replicative DNA polymerases in yeast. Genes 9, 539 (2018).
Ira, G. et al. DNA end resection, homologous recombination and DNA damage checkpoint activation require CDK1. Nature 431, 1011–1017 (2004).
Pantopoulos, K. Inherited Disorders of Iron Overload. Front. Nutr. 5, 103 (2018).
Kowdley, K. V. Iron, hemochromatosis, and hepatocellular carcinoma. Gastroenterology 127, S79–S86 (2004).
Rizzollo, F., More, S., Vangheluwe, P. & Agostinis, P. The lysosome as a master regulator of iron metabolism. Trends Biochem. Sci. 46, 960–975 (2021).
Terzi, E. M., Sviderskiy, V. O., Alvarez, S. W., Whiten, G. C. & Possemato, R. Iron-sulfur cluster deficiency can be sensed by IRP2 and regulates iron homeostasis and sensitivity to ferroptosis independent of IRP1 and FBXL5. Sci. Adv. 7, eabg4302 (2021).
Chan, S. Y. et al. MicroRNA-210 controls mitochondrial metabolism during hypoxia by repressing the iron-sulfur cluster assembly proteins ISCU1/2. Cell Metab. 10, 273–284 (2009).
Favaro, E. et al. MicroRNA-210 regulates mitochondrial free radical response to hypoxia and krebs cycle in cancer cells by targeting iron sulfur cluster protein ISCU. PLoS One 5, e10345 (2010).
Mayle, R. et al. DNA REPAIR. Mus81 and converging forks limit the mutagenicity of replication fork breakage. Science 349, 742–747 (2015).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Putri, G. H., Anders, S., Pyl, P. T., Pimanda, J. E. & Zanini, F. Analysing high-throughput sequencing data in Python with HTSeq 2.0. Bioinformatics 38, 2943–2945 (2022).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Yu, G., Wang, L. G., Han, Y. & He, Q. Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16, 284–287 (2012).
Zhang, J., Kobert, K., Flouri, T. & Stamatakis, A. PEAR: a fast and accurate Illumina Paired-End reAd mergeR. Bioinformatics 30, 614–620 (2014).
Ye, J., McGinnis, S. & Madden, T. L. BLAST: improvements for better sequence analysis. Nucleic Acids Res. 34, W6–W9 (2006).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Acknowledgements
We thank Dr. James Haber, Dr. Brain Wasko, Dr. Peter Thorsness, Dr. Francesca Storici, and Dr. Benedikt Westermann for the gift of plasmids or strains. This work was funded by grants from the National Institute of Health (GM080600 and GM125650 to G.I., and R01GM125632, R01GM138407, R01HL148338, R01HL133254, R01HL174928, and R01CA278832 to K.C., and R01AG061376 to A.H.). This project was also supported by the Cytometry and Cell Sorting Core at Baylor College of Medicine with funding from the CPRIT Core Facility Support Award (CPRIT-RP180672), the NIH (P30 CA125123 and S10 RR024574) and the expert assistance of Joel M. Sederstrom.
Author information
Authors and Affiliations
Contributions
J.F. constructed strains, designed experiments, and performed most of the experiments. Y.Y. (BCM) made substantial contributions to initiate this project. Y.Y. (HMS) and X.W. analyzed sequences using the Break-Ins pipelines and toolkits and analyzed expression patterns. Y.Y. (HMS) performed features analysis and mapping of Ty and mtDNA insertions. A.H. provided essential strains and provided expertise on iron-related experiments. C.F. constructed strains, conducted fluorescence microscopy, and assessed respiration capacity. Q.L. constructed strains and assessed respiration capacity. P.T. constructed fzo1 point mutants and assisted with revision of the manuscript. Y.W. analyzed microhomologies of insertions and organized raw files for GEO submission. G.I., J.F., X.W. and K.C. designed experiments, analyzed the data. G.I. and J.F. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Ludovic Enkler and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Fox, J., Yu, Y., Yu, Y. et al. Deficits in mitochondrial dynamics and iron balance result in templated insertions. Nat Commun 16, 5454 (2025). https://doi.org/10.1038/s41467-025-60546-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-60546-2