Abstract
High doses of ionizing radiation (IR) cause severe damage to the hematopoietic system. However, the heterogeneity of hematopoietic stem and progenitor cells (HSPCs) in response to IR stress remains largely uncharacterized. Here, we present a dynamic single cell transcriptomic landscape and elucidate the complex crosstalk between HSPCs and the bone marrow (BM) microenvironment during IR-induced regeneration process. We reveal that BMP4 signaling in HSPCs confers IR resistance, and a single administration of BMP4 or SB4 can rescue mice from the IR-induced mortality. Furthermore, we identify BMPR2+ HSCs as a radiation resistant subset, displaying distinct epigenetic landscapes from BMPR2− HSCs under radiation stress. BMPR2+ HSCs sustain a strong self-renewal capacity primarily by reducing the H3K27me3 modification on the Nrf2 gene in response to radiation stress. In Nrf2 knockout mice, we demonstrate that Nrf2 is a critical downstream functional gene for BMP4-BMPR2 signaling on HSCs to resist IR-induced damage. Collectively, we provide insights into the molecular intricacies underlying HSPC heterogeneity and BM niche after radiation exposure, and we uncover that BMP4-BMPR2 signaling may serve as a promising target for developing innovative and effective intervention strategies to mitigate IR-induced hematopoietic injury.
Similar content being viewed by others
Introduction
Hematopoietic stem and progenitor cells (HSPCs) are the key seed cells for maintaining and regenerating the entire blood system throughout the lifetime1,2,3. The numbers and functionality of HSPCs are prone to be affected by various stress stimuli, including ionizing radiation (IR), chemotherapy or inflammatory microenvironment4. IR can cause direct DNA damage, oxidative stress, cell apoptosis, senescence, or death of HSPCs5. IR also induces bone marrow (BM) niche damage and disrupts the steady-state microenvironment. The bystander effects of IR, including inflammatory reactions and increased reactive oxidative species (ROS), further impair the functionality of HSPCs6,7,8. Despite significant progress in understanding the critical molecular mechanisms underlying the radiation-induced hematopoietic repopulation, the processes governing how HSPCs respond to IR and regenerate the entire hematopoietic system remain incompletely understood9. Therefore, further studies are imperative to investigate the in vivo dynamics of HSPC populations and the alterations of BM niche signals that influence HSPC fate under IR stress. It will be beneficial for developing promising radiation countermeasures to protect HSPCs or mitigate radiation-induced hematopoietic injury.
Hematopoietic stem cells (HSCs) reside at the apex of a hematopoietic hierarchy. Increasing evidence indicates that the rare adult HSCs in the BM are a functionally heterogeneous population and exhibit different responsiveness to diverse stresses10,11,12. Recent lineage-tracing studies uncovered that specific subsets of stem cells with radiotolerant properties reside in diverse tissues, such as intestine and muscle13,14,15. It is plausible that a specific radioresistant subpopulation of HSCs may also exist in the BM, considering that very few HSCs can survive and reconstitute all blood cell lineages in mice after exposure to high doses of IR16,17. Several studies have reported that HSCs have intrinsic stress-resistant gene clusters, including Bmi1, forkhead box protein O, and nuclear factor erythroid 2-related factor 2 (Nrf2) genes18,19,20,21. However, it remains a significant challenge to provide precise insights into the identity and regulatory mechanisms of the rare HSC subpopulation responsible for the reconstruction of the radiation-damaged hematopoietic system.
Here, we employed the single-cell RNA sequencing (scRNA-seq) technique to dissect the in vivo dynamics of murine HSPCs and the BM microenvironment during IR-stressed hematopoietic regeneration. We discovered that BMP4 signaling plays a crucial role in mitigating hematopoietic injury induced by IR. More importantly, we identified a rare population of HSCs exhibiting robust radioresistance and self-renewal properties. Furthermore, we elucidated the epigenetic regulatory mechanisms in HSCs governed by BMP4 and its receptor in response to radiation stress.
Results
A single-cell atlas of BM during IR-induced hematopoietic stress reveals a rapid exhaustion of the stem cell pool and an enhanced GMP skewing
To explore the dynamic transcriptomes of heterogeneous cell populations during IR-induced BM injury, we conducted scRNA-seq on BM Lin– cells isolated from irradiated mice at day 1 (D1), 3 (D3), 7 (D7), 14 (D14), and 21 (D21), as well as non-irradiated mice (D0), respectively (Fig. 1a). We identified a total of 13 major cell types, including a mixed population of HSCs and multipotent progenitors (MPPs), common lymphoid progenitors (CLPs), megakaryocyte and erythroid progenitors (MEPs), granulocyte-macrophage progenitors (GMPs), as well as precursor or mature cell types (Fig. 1b, Supplementary Fig. 1, Supplementary Data 1). The mixed HSCs/MPPs population was further segregated into the long-term HSCs (LT-HSCs), short-term HSCs or MPP1 (ST-HSCs/MPP1), MPP2, MPP3, and MPP4 by a second round of cell clustering. Annotation of the cell clusters was supported by a set of previously reported canonical marker genes of HSPCs (Supplementary Fig. 2a-b, Supplementary Data 2). We identified three branched differentiation trajectories, originating from LT-HSCs and ending in MEPs, GMPs, and CLPs, respectively, passing through MPP2, MPP3, and MPP4 (Fig. 1c). The ordering of these three differentiation paths was further demonstrated by signature gene expression profiles, delineating varying exit dynamics from the initial HSCs, marked by Hlf and Procr genes (Fig. 1d).
a Experimental design for studying the dynamic transcriptome changes of BM cells following IR-induced BM injury. The cartoon illustrations of a mouse and a bone were obtained from Figdraw. b UMAP visualization of different BM cell types identified by scRNA-seq. c UMAP visualization and differentiation trajectories among re-clustered HSPCs in BM. d UMAP visualization of pseudotime scores and expression profiles of signature genes along different pseudotime trajectories. e MELD plots (top) quantifying the distribution and stacked bar plots (bottom). f Differential signature scores of three differentiation trajectories and proliferation, comparing HSPC subpopulations after IR exposure at different time points with those at D0. The P values are calculated using a permutation-based approach in GSEA. g Expression profiles of representative genes of the GMP trajectory (Cebpe and Mt1) and proliferation (Mki67 and Ccnb2) along the pseudotime trajectories. Fitting curves of gene expression in two groups was shown with 95% confidence intervals. h Differentially activated transcription factors between MPP3 at D3 and those at D0. The differential expression analysis was performed by fitting an empirical Bayes linear model. Source data are provided as a Source Data file.
We then investigated the dynamic profile of HSPCs during the BM injury and regeneration process. A relative likelihood estimation algorithm, MELD was adopted to quantify the distribution of HSC subpopulations at different time points (Fig. 1e, Supplementary Fig. 2c–e)22. The analysis revealed a substantial increase in the proportion of LT-HSCs within the HSPC population at D1 compared to D0 (Fig. 1e, Supplementary Fig. 2c, d, Supplementary Data 3), indicating their higher radioresistance than MPPs. Strikingly, the proportion of LT-HSCs dropped sharply from D3 to D21 post-irradiation, indicating a rapid and continuous depletion of the stem cell pool following IR. Notably, LT-HSCs exhibited a remarkably skewed in the MEP differentiation path at D1 (Fig. 1e, f). Moreover, we found that the proportion of GMPs dramatically increased at D3 and remained at a higher level until D21 (Fig. 1e, Supplementary Fig. 2f), indicating an augmented GMP fate commitment and/or enhanced proliferation capacity along the GMP trajectory under IR stress. Indeed, ST-HSCs and MPP3 showed consistently elevated expression of GMP signature and proliferation-related gene sets from D3 to D21 after IR as compared to those in D0 (Fig. 1f, Supplementary Fig. 3), supporting the concurrent enhancement of differentiation and proliferation along the GMP path. In line with this data, the expression levels of marker genes of GMP signature (Cebpe and Mt1) and proliferation (Mki67 and Ccnb2) were upregulated along the GMP trajectory at D3 compared to D0 (Fig. 1g). Consistently, the transcription factor (TF) activity analysis using SCENIC further demonstrated that TFs associated with cell proliferation (Ybx1, Tfdp1, E2f1, and E2f4) and GMP signature (Cebpz) were upregulated significantly in MPP3 at D3 after irradiation compared to those at homeostasis (Fig. 1h, Supplementary Data 4)23. Together, these data revealed a short-term increase in the LT-HSC ratio in the irradiated BM, accompanied by a subsequent rapid exhaustion of the HSC pool. This phenomenon may be attributed to the skewing of the GMP differentiation path of HSCs under IR stress.
LT-HSCs exhibit reduced lymphoid differentiation signature under IR-induced regeneration stress
Given that HSC/MPP subsets are critical seed cells for hematopoietic regeneration, we further dissect the dynamic transcriptomes of these subpopulations before and after IR. A weighted gene co-expression network was constructed and grouped into six gene modules, among which Modules 2 and 6 exhibited the strongest yet opposing associations with both time-series and HSC/MPP subpopulations (Fig. 2a, b, Supplementary Data 5)24. LT-HSC subpopulation exhibited the highest Module 2 score and lowest Module 6 score (Fig. 2c). Notably, genes in Module 2 were enriched with “low-output”, “megakaryocyte-biased” and “HSC” signatures (Fig. 2d). Specially, the overlapped genes in “low-output” signatures included Hlf, Mycn, Procr, Mllt3, Hoxb8, and Cdkn1c (Fig. 2e)25,26,27,28,29. Functional enrichment analysis showed that both Module 2 and the “low-output” signature are enriched in pathways associated with “HSC homeostasis” and “regulation of hematopoiesis”, further supporting the low-output HSC feature of Module 2 (Fig. 2f). Interestingly, the gene expression profile in Module 2 of LT-HSC subpopulation at D0 resembled that at D1 post-irradiation but exhibited significant differences from those at D3 to D21 (Fig. 2a), indicating the maintenance of HSC homeostasis in the short term after IR stress.
a Hierarchical clustering and heatmap showing the distinct expression patterns of six gene modules in HSPCs identified by weighted gene co-expression network analysis (WGCNA). b Associations between the six gene modules and the temporal course and cell types of HSPCs. DPI, days post-irradiation. c Distribution of scores for Module 2 and Module 6 among HSPC subpopulations calculated by GSVA. d Heatmap showing the overlap ratios between different signature scores across the six gene modules. e Venn plot showing the shared and specific genes of Module 2 and the low-output signature. f Functional enrichment network showing the shared and specific GO terms related to Module 2 and the low-output signature. g Heatmap showing the dynamic expression profiles of the four sub-modules within Module 2. h Enriched GO terms and activated TFs in the four sub-modules. Significances of enrichment analyses were calculated using the Chi-square test. i GSEA analysis showing the high MK-biased HSC score at D1 and the low lymphocyte score at D3 in LT-HSCs compared to those at D0. Source data are provided as a Source Data file.
We next explored the gene expression heterogeneity of Module 2 in LT-HSCs during IR-induced injury and regeneration. Unsupervised clustering analyses revealed four sub-modules exhibiting different temporal gene expression patterns at multiple time points after IR (Fig. 2g, Supplementary Data 6). Sub-module 1 displayed a sharp increase in expression level at D1, returning to baseline level at D3, and was associated with “positive regulation of defense response” and “regulation of response to external stimulus”. Interestingly, this module was enriched with megakaryocytic genes, such as Pf4, Thbs1, Vwf, and Gp9 (Fig. 2g, h). Consistently, gene set enrichment analysis (GSEA) data showed a significant upregulation of MK-biased HSC signature at D1 post-irradiation compared to D0 (Fig. 2i). Sub-module 2 showed elevated expressions at D1 and D3, and was enriched with genes including Bmpr2, Hes1, and Smad7. Notably, the GO term “regulation of protein kinase B signaling” was enriched in sub-module 2, which is involved in the increase of neutrophil and monocyte fate choice of HSCs30. Sub-module 3 showed elevated expression levels only at D3 and was enriched with genes Tnf and Sulf2 and GO terms “regulation of leukocyte differentiation” and “regulation of hematopoiesis”. Notably, sub-module 4 showed gradually downregulated gene expression patterns at D1 and D3 after IR. Genes including Flt3, Rora, Stab1, and Ctla4 presented the strongest expression levels at D0, consistent with the enriched GO terms “T cell differentiation” and “lymphocyte differentiation” in this module. GSEA results further demonstrated a significant downregulation of the lymphocyte signature in D3 post-irradiation compared to D0 (Fig. 2i), indicating a reduced lymphoid differentiation ability of LT-HSCs in the early stage after IR. Consistently, the proportion of MPP4 in HSPCs reduced remarkably at D1 and recovered slowly until D21 (Fig. 1e, Supplementary Fig. 2c, d). Collectively, these data illustrate the complex and dynamic gene expression programming in HSPC populations during IR-induced injury and regeneration in the hematopoietic system.
Characterization of cellular interactions between the BM microenvironment and HSCs/MPPs during stress hematopoiesis
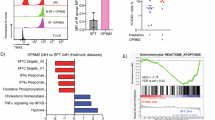
The normal and stress hematopoiesis is tightly modulated by BM niche cells and various cytokines31. To dissect the extracellular regulatory mechanisms of hematopoietic programming, we analyzed the gene expression profile of cytokines in various BM cells before and after IR stress. BM stromal cells were further divided into four distinct types of cells (Supplementary Fig. 4a–c). Of note, BM stromal cells and various blood cells, especially basophils and T/NKs, in the BM exhibited high cytokine scores (Fig. 3a–c). Specifically, basophils showed significantly increased expression levels of Osm, Hgf, and Il6 at D3 after IR (Fig. 3d), indicating a myeloid differentiation bias of HSPCs at the early stage of IR-induced BM injury32,33,34,35. Additionally, various chemokines, such as Ccl6 in basophils, exhibited highly induced expressions after IR (Fig. 3d), suggesting the activation of inflammatory reactions in BM basophils upon IR36. Notably, the Ccl5 gene was activated in T/NKs at D3 after IR (Fig. 3d), which was consistent with that Ccl5 enhances hematopoietic regeneration after IR37. We further confirmed by intracellular cytokine staining and flow cytometric analysis that, compared to D0, the protein expression levels of OSM and IL6 in basophils and CCL5 in T cells were significantly elevated on D3 after irradiation, supporting the results of the scRNA-seq analysis (Fig. 3e, Supplementary Fig. 4d).
a UMAP visualization of cytokine scores for each cell type in BM cells. b Violin plots showing the distribution of cytokine scores across different cell types in BM cells. c Dotplot showing the expression profiles of cytokines across different cell types. Dot color indicates the average expression levels, and dot size indicates the fraction of expressing cells. d Violin plots showing representative cytokine expression changes in basophils and T/NKs at D0 and at D3 after 6.5 Gy of IR. e Fold change in OSM and IL6 expression levels in basophils, and fold change in CCL5 expression levels in T cells, at D0 and at D3 after 6.5 Gy of IR. n = 3 or 4 biological replicates per group. Data shown are mean ± SEM. f Dotplot showing the enriched LR pairs between HSPC subpopulations and niche cells. Dot color indicates the fold change (FC) of enrichment, and dot size indicates the P value of enrichment. g Two clusters of LR interactions with distinct temporal intensity patterns (left) were observed between different HSPC subpopulations and niche cells (right). h Heatmap showing the intensities of specific LR pairs across different time points after IR. “Cell to” indicates the cell type highly expressing the receptor in the corresponding ligand-receptor. “Cell from” indicates the cell type highly expressing the ligand in the corresponding LR. i Fold change in BMP4 expression levels in CD45−VE-CAD⁺ endothelial cells and CD45−LEPR⁺ stromal cells at D0 and D1 after 6.5 Gy of IR. j Fold change in pSMAD1/5 expression levels in HSCs at D0 and D1 after 6.5 Gy irradiation. n = 4 biological repeats per group. Data shown are mean ± SEM. Statistical analyses were conducted using unpaired two-sided Student’s t tests (e, i, j). Source data are provided as a Source Data file.
We next investigated the cell-cell interactions mediated by ligand-receptor (LR) pairs between niche cells and HSPCs in the BM. Notably, a considerable number of interactions were identified between Lepr+ stromal cells and HSPCs (Fig. 3f, Supplementary Data 7). All LR interactions between niche cells and HSPCs were categorized into two clusters, each exhibiting distinct temporal profiles (Fig. 3g). LR pairs in Cluster 1 displayed a substantial increase in intensity at D1 post-irradiation, predominantly occurring between LT-HSCs and Lepr+ stromal cells. However, LR pairs in Cluster 2, with relatively high intensity at homeostasis, were significantly reduced shortly after irradiation and then gradually recovered at D21 post-irradiation. Notably, the Cxcl12-Cxcr4 interaction in Cluster 1 was remarkably increased between LT-HSCs and Lepr⁺ cells or VE-Cad⁺ cells at D1 after irradiation (Fig. 3h). This may be attributed to the increased expression of the CXCL12 gene and protein in these BM niche cells (Supplementary Fig. 4e–g), suggesting that HSC quiescence and niche retention were maintained and even enhanced by niche cells at D138,39,40,41. Moreover, we observed that both Lepr+ cells and VE-Cad+ cells showed interactions with LT-HSCs through the Bmp4-Bmpr2 LR, which was markedly enhanced at D1 in Cluster 1 (Fig. 3h). Consistently, the expression of the BMP4 gene and protein was significantly elevated in these BM niche cells, and the phosphorylation level of pSMAD1/5 was increased in HSCs on D1 (Fig. 3i, j, Supplementary Figs. 4e, 5a), which suggests that BMP4 signaling may play a role in mitigating radiation-induced damage to HSCs.
BMP4 protects HSCs against radiation-induced damage
To investigate whether the activation of BMP4 signaling is beneficial for reducing radiation-induced HSC injury, we conducted a series of in vitro experiments using the freshly isolated BM cells. Comet assays demonstrated that the addition of recombinant BMP4 proteins significantly reduced both the comet tail moment and DNA content of LSKs under IR stress (Fig. 4a, Supplementary Fig. 5b). BMP4 supplementation also remarkably decreased the formation of γH2AX foci in HSCs, represented by LSKCD48−CD150+ cells, at 2 hours post-IR (Fig. 4b, Supplementary Fig. 5c). In addition, ROS detection results showed that BMP4 treatment led to significantly reduced ROS levels in LSKs and HSCs after IR exposure (Fig. 4c, Supplementary Fig. 5d), indicating the positive role of BMP4 on mitigating radiation-induced oxidative stress damage. Correspondingly, we observed a significantly reduced apoptosis rate and increased colony-forming ability of irradiated LSKs in the presence of BMP4 (Fig. 4d, Supplementary Fig. 5e). Collectively, these data suggest that BMP4 mitigates the radiation-induced damage to HSPCs.
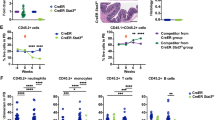
a Comet assay of LSK cells. n = 100 cells per group. b Representative images of γH2AX foci and γH2AX foci counts per HSC across treatment conditions. n = 100 cells per group. c Histograms of ROS levels measured by CellROX in HSCs and the relative CellROX MFI normalized to the non-IR group. n = 4 biological repeats per group. d Number of hematopoietic colonies formed by LSK cells in each group. n = 3 biological repeats per group. e Percent survival of mice after receiving 8.0 Gy of IR and different dosages of BMP4. n = 8 mice per group. f WBC, PLT, and RBC counts in the PB. n = 5 mice per group. BM morphology and the cellularity of BM (g, h n = 8 mice per group) on D10 after exposure to 6.5 Gy of IR. Frequencies of LSKs and HSCs (i), the total number of HSCs (j) and the number of CFU (k) in mice treated with vehicle or BMP4 on D10 following 6.5 Gy of IR. n = 8 mice per group (i, j). n = 6 biological repeats per group (k). l Competitive transplantation strategy. The cartoon illustrations of mice and bones were obtained from Figdraw. The percentages (m, p) and the lineage distribution (n) of donor-derived cells in the PB of recipient mice. The engraftment of donor HSCs was assessed in recipient mice at 16 weeks (o, q). n = 6 mice per group. r Percentage survival of mice after receiving 8.0 Gy of IR and different dosages of SB4. n = 6 mice for Veh group, n = 5 mice for each SB4-treated group. s WBC, PLT, and RBC counts in the PB. n = 5 mice per group. Data shown are mean ± SEM. Statistical analyses were conducted using two-way ANOVA with multiple comparisons by Tukey’s test (a–d), two-sided log-rank test (e, r), and unpaired two-sided Student’s t test (f, h–q, s). Source data are provided as a Source Data file.
We then sought to explore whether BMP4 has a radioprotective role on the hematopoietic system in vivo. Strikingly, we observed that a single injection of BMP4 at a dose of 100 μg/kg offered radiation-mitigating benefits when administered 2 h prior to IR exposure or up to 12 hours after IR exposure (Supplementary Fig. 5f). One single injection of BMP4 at dose of 100 μg/kg 2 hours prior to IR was highly effective in improving the survival rate of the mice exposed to lethal doses of radiation (8.0 Gy), resulting in a 100% survival rate without body weight loss (Fig. 4e, Supplementary Fig. 5g), suggesting that BMP4 might effectively protect the hematopoietic system against radiation injury. To gain insight into the radioprotective effect of BMP4, we performed hemogram analyses of the irradiated mice with BMP4 pre-treatment. The results showed that a single dose of BMP4 administration significantly increased the numbers of peripheral blood (PB) white blood cell (WBC), red blood cell (RBC), and platelet (PLT) counts from D7 to D21 following IR (Fig. 4f), indicating enhanced hematopoietic recovery by BMP4 injection. The radioprotective role of BMP4 administration was further evident by the decreased apoptosis rate of LSKs in BM at D1 post-IR (Supplementary Fig. 5h). Additionally, BMP4 administration remarkably increased BM cellularity, HSPC numbers and colony-forming unit (CFU) numbers compared to the vehicle IR group at D1 and D10 following IR (Fig. 4g–k, Supplementary Fig. 5i, j).
Furthermore, BM cells from the BMP4-pretreated irradiated mice at D10 post-IR contributed significantly to long-term trilineage and HSC pool reconstitution (Fig. 4l–q, Supplementary Fig. 5k–q). We also tested the radioprotective effect of the BMP4 signaling agonist SB4. Strikingly, a single dose of SB4 at 5 mg/kg rescued mice exposed to lethal doses of radiation (8.0 Gy), achieving a 100% survival rate (Fig. 4r). Additionally, it significantly enhanced the recovery of blood cells in mice subjected to sub-lethal irradiation (6.5 Gy) (Fig. 4s). Taken together, these findings suggest that BMP4 signaling plays a crucial role in enhancing the radiation resistance of HSCs and accelerating hematopoietic regeneration.
BMPR2+ HSCs are radioresistant stem cell population
Considering that BMP4 signaling depends on its receptors to regulate HSCs, we analyzed the heterogeneity of HSCs based on the expression levels of the receptor genes. Intriguingly, scRNA-seq data revealed that HSCs can be distinguished as Bmpr2+ and Bmpr2− cells, with the BMP4 signaling pathway being significantly activated in Bmpr2+ HSCs (Fig. 5a, b, Supplementary Data 8 and 9), which is confirmed by increased levels of phosphorylated SMAD1/5. Notably, the cell number and percentage of Bmpr2− HSCs dropped sharply at D1 following IR exposure compared to those at D0 (Fig. 5c, d). By contrast, Bmpr2+ HSCs became the major cell subpopulation at D1 in HSC population following IR exposure (Fig. 5c, d). In line with scRNA-seq data, flow cytometry detection data confirmed that the percentage and number of BMPR2+ HSCs and BMPR2− HSCs in the BM are at similar levels at homeostasis; while a much higher percentage and number of BMPR2+ HSCs were detected compared to those of BMPR2− HSCs at D1 after irradiation (Fig. 5e, f). Together, these results indicate that a great number of BMPR2+ HSCs survive at D1 after IR, suggesting that BMPR2+ HSCs may be more radioresistant than BMPR2− HSCs.
a UMAP visualization of Bmpr2+ and Bmpr2− LT-HSC. b Barplot showing the enriched GO terms in upregulated genes in Bmpr2⁺ HSCs compared to Bmpr2⁻ HSCs. Significances of enrichment analyses were calculated using Chi-square test. c Cell counts of Bmpr2⁺ and Bmpr2⁻ LT-HSCs at homeostasis (D0) and D1 after 6.5 Gy of IR. d UMAP visualization of Bmpr2⁺ and Bmpr2⁻ LT-HSCs at D0 and D1 after 6.5 Gy of IR. e, f Percentage and quantification of BMPR2⁺ HSCs at D0 and D1 after 6.5 Gy of IR. n = 8 mice per group. g The number of γH2AX foci in BMPR2⁺ and BMPR2⁻ HSCs at 2 hours post-irradiation. Representative plots of γH2AX foci and the number of γH2AX foci per HSC. n = 200 cells per group. h Hallmark pathways significantly upregulated in Bmpr2⁺ and Bmpr2⁻ LT-HSCs, respectively. Significances of enrichment analyses were calculated using Chi-square test. i Analysis of the ROS pathways in Bmpr2⁺ and Bmpr2⁻ LT-HSCs using Gene Set Enrichment Analysis (GSEA). j Representative analyses of CellROX levels in BMPR2⁺ and BMPR2⁻ HSCs at D0 and D1 after 6.5 Gy of IR, along with the fold change. n = 4 biological repeats per group. k The relative activity of Caspase 3/7 normalized to the non-IR BMPR2⁻ HSCs group. n = 4 biological repeats per group. Data shown are mean ± SEM. Statistical analyses were conducted using two-way ANOVA with multiple comparisons by Tukey’s test (e–g, j, k). Source data are provided as a Source Data file.
To further evaluate the difference in radiation responsiveness between these two HSC subpopulations, the DNA damage degree of these stem cells from the BM of irradiated mice was detected according to the quantification of γH2AX foci. Imaging cytometry analysis showed that fewer γH2AX-staining foci are observed in BMPR2+ HSCs than in BMPR2− HSCs at 2 hours post-IR (Fig. 5g). Consistently, BMPR2+ HSCs presented fewer γH2AX foci than BMPR2− HSCs after IR in vitro (Supplementary Fig. 6a, b). In addition, scRNA-seq data showed that Bmpr2− HSCs exhibit significant enrichment of the “ROS pathway” and “oxidative phosphorylation”, while Bmpr2+ HSCs do not (Fig. 5h, i). We further tested the intracellular ROS levels of these two HSC subgroups from mice with or without receiving IR. Both BMPR2+ and BMPR2− HSCs presented extremely low ROS levels under homeostasis (Fig. 5j). However, under in vivo or in vitro IR-stressed conditions, BMPR2+ HSCs presented significantly lower ROS levels than BMPR2− HSCs (Fig. 5j, Supplementary Fig. 6c), indicating that BMPR2+ HSCs had reduced oxidative stress injury induced by IR. Supporting this, Caspase 3/7 activity detection results showed lower apoptosis levels in BMPR2+ HSCs than in BMPR2− HSCs at D1 after IR exposure (Fig. 5k). Together, these data suggest that BMPR2+ HSCs are more radioresistant than BMPR2− HSCs.
BMPR2+ HSCs have robust self-renewal capacity
Given the relatively enhanced radioresistant characteristics of BMPR2+ HSCs compared to BMPR2− HSCs, we postulate that these two stem cell subpopulations might have different self-renewal capacity. We first employed the CFU assays to analyze the functionality of BMPR2+ and BMPR2− HSCs. Regardless of whether the mice are irradiated or not, the BMPR2+ HSCs isolated from them showed a stronger hematopoietic colony-forming ability than the BMPR2− HSCs (Fig. 6a). Recent studies have indicated that a low mitochondrial membrane potential (MMP) is essential for maintaining the self-renewal of HSCs42,43. We thus evaluated the MMP of these two HSC subgroups before and after IR exposure by testing the Tetramethylrhodamine, methyl ester (TMRM) signals. Indeed, BMPR2+ HSCs exhibited significantly lower MMP than BMPR2− HSCs at homeostasis and at D1 after IR exposure (Fig. 6b), indicating a greater self-renewal capacity of BMPR2+ HSCs compared to BMPR2− HSCs.
a Number of CFUs formed by BMPR2⁺ and BMPR2⁻ HSCs at homeostasis (D0) and D1 after 6.5 Gy of IR. n = 3 biological repeats per group. b MFI of TMRM and MitoTracker in BMPR2⁺ and BMPR2⁻ HSCs at D0 and D1 after 6.5 Gy of IR. n = 4 biological repeats per group. c–m Competitive transplantation. The cartoon illustrations of mice and bones were obtained from FigDraw. Donor-derived chimerism in the PB of recipients at 16 weeks using 100 BMPR2⁺ HSCs or BMPR2⁻ HSCs (d, g) and using 2000 BMPR2⁺ HSCs or BMPR2⁻ HSCs from 6.5 Gy-irradiated donor mice (i, l) after primary and secondary transplantation. The lineage distribution of donor-derived cells in the PB of recipient mice (e, j) and the engraftment of donor HSCs were assessed in recipient mice (f, h, k, m). n = 8 mice per group. n WGCNA Module 2 (stemness maintenance) and low-output signatures in Bmpr2⁺ and Bmpr2⁻ LT-HSCs using GSEA. o Violin plot showing the upregulation of stemness maintenance-related genes in Bmpr2⁺ LT-HSCs (n = 88) compared to Bmpr2− LT-HSCs (n = 86). Boxplot within the violin showing the 0.25, 0.5 and 0.75 quartiles of gene expression. Statistical analyses were conducted using the Wilcox rank test. Data shown are mean ± SEM. Source data are provided as a Source Data file.
We then performed the in vivo competitive transplantation assays to evaluate the hematopoietic function of these two HSC subgroups. The two stem cell subpopulations from non-irradiated or irradiated CD45.2 mice were transplanted with competitor cells into the lethally irradiated CD45.1 recipients (Fig. 6c). We observed that the reconstitution capacity of BMPR2+ HSCs isolated from non-irradiated mice is remarkably higher than that of BMPR2− HSCs (Fig. 6d). More donor-derived blood lineage cells were observed in PB, spleen, and thymus in those mice receiving BMPR2+ HSCs from non-irradiated mice (Fig. 6e, Supplementary Fig. 7a, b). Encouragingly, the donor HSCs frequency in the BM of recipients receiving BMPR2+ HSCs from non-irradiated mice was much higher than that of the recipients with BMPR2− HSC transplantation (Fig. 6f). The secondary transplantation experiments further demonstrated the remarkably enhanced chimerism of donor-derived blood lineage cells in PB and increased donor LSKs and HSCs in the BM of recipients (Fig. 6g, h, Supplementary Fig. 7c–e). Of note, very similar results were observed for higher donor blood lineage cells, LSK and HSC incorporation percentages in the first and second recipient mice that received BMPR2+ HSCs than those that received BMPR2− HSCs from the irradiated mice (Fig. 6i–m, Supplementary Fig. 7f–j). These results strongly indicate that BMPR2+ HSCs possess a greater self-renewal capacity than BMPR2− HSCs. Our scRNA-seq data further supported the notion that Bmpr2+ HSCs exhibit higher stemness scores compared to Bmpr2− HSCs (Fig. 6n). Consistently, Bmpr2+ HSCs demonstrated elevated expression levels of self-renewal-related genes, including Hlf, Mllt3, Runx1, Gata2, and Meis1 (Fig. 6o). Collectively, these findings suggest that BMPR2+ HSCs have a more robust self-renewal potential than BMPR2− HSCs, even following IR exposure.
BMPR2+ HSCs possess different chromatin landscapes from BMPR2– HSCs, with increased chromatin accessibility on Nrf2 gene under IR stress
Next, we sought to investigate the underlying mechanisms of radioresistance in BMPR2+ HSCs. Increasing evidence suggests that the BMP signaling pathway may modulate the epigenetic states in various types of cells44,45. We then investigated the chromatin accessibility alterations in BMPR2+ and BMPR2− HSCs obtained from the non-irradiated and irradiated mice using the assay for transposase-accessible chromatin with high-throughput sequencing (ATAC-seq). Globally, BMPR2+ HSCs exhibited lower intensity of ATAC peaks around gene transcription start sites (TSSs) than BMPR2− HSCs under homeostasis or IR stress (Fig. 7a, Supplementary Data 10), suggesting higher stemness with lower transcription activities in BMPR2+ HSCs. Of note, the genes with open TSS regions in BMPR2+ HSCs were enriched in pathways associated with “cellular response to oxidative stress”, “regulation of Wnt signaling pathway”, and “regulation of response to external stimulus” (Fig. 7b). Motif enrichment analysis showed that in BMPR2+ HSCs, the accessible regions are recognized by HSC-featured TFs, including RUNX1, ERG, ETV2, ETS1, ETV1, and FLI1, indicating a robust stem cell signature of BMPR2+ HSCs (Fig. 7c). Additionally, the binding motifs for stress-related TFs, such as NRF2, which is a key regulatory factor responding to oxidative stress and radiation, were remarkably enriched in BMPR2+ HSCs (Fig. 7c), suggesting the existence of stress-response regulation in this subgroup of HSCs. Consistently, TF enrichment analysis from scRNA-seq dataset also showed a significant activation of NRF2 in BMPR2+ HSCs as compared to BMPR2– HSCs (Fig. 7d), supporting the stress-resistant potential of BMPR2+ HSCs.
a Centered ATAC-seq peaks in BMPR2⁺ and BMPR2⁻ HSCs at D0 and D1 after 6.5 Gy of IR (top), and their distributions across different genomic regions. b Functional enrichment of genes with open chromatin specifically in BMPR2⁺ HSCs. c Top enriched TFBS motifs across the open chromatin regions specifically in BMPR2⁺ HSCs. d Top enriched TFs in BMPR2⁺ HSCs inferred from scRNA-seq data. e Chromatin accessibility landscapes across Nrf2, Hmox1, Runx1, and Etv2. f TF-gene networks showing the top enriched TFs and their target genes in BMPR2⁺ HSCs identified by ATAC-seq. Significances of enrichment analysis in b–d were calculated using the Chi-square test. Source data are provided as a Source Data file.
To further explore the intrinsic mechanism of function difference between BMPR2+ and BMPR2− HSCs, we analyzed the chromatin status of key TFs related to HSC signature and stress response. Indeed, the master TF genes involved in HSC stemness, including Runx1, Etv2, Erg, Ets1, Id1, and Id2, exhibited profoundly enhanced chromatin opening signals in BMPR2+ HSCs from both non-irradiated and irradiated mice (Fig. 7e and Source data). Consistently, higher activities of these TFs in BMPR2+ HSCs was observed than those in BMPR2− HSCs (Fig. 7f). Of interest, stress response-related TFs, such as Nrf2 and its downstream gene Hmox1, showed partly enhanced or reduced opening signals in BMPR2+ HSCs in non-irradiated mice, while their chromatin accessibility signals were remarkably increased in BMPR2+ HSCs from irradiated mice (Fig. 7e), indicating a dynamic and complex chromatin modification of Nrf2 gene loci in BMPR2+ HSCs under different conditions. The enhanced chromatin accessibility of the Nrf2 and Hmox1 genes in BMPR2+ HSCs under IR stress may contribute to the radioresistance of this cell population.
Nrf2 gene displays bivalent and dynamic epigenetic modification and plays a key role in radioresistance in BMPR2+ HSCs
We further investigate the underlying mechanisms that contribute to the differences in chromatin accessibility of genes between BMPR2+ and BMPR2− HSCs. Increasing evidence indicates that the BMP4 signal pathway can regulate histone modification, including the trimethylation of lysine 4 (H3K4me3) and lysine 27 (H3K27me3) on histone H346. To further elucidate the epigenetic status of these two stem cell subpopulations, we performed CUT&Tag assay for genome-wide mapping of H3K4me3 and H3K27me3 in BMPR2+ and BMPR2– HSCs before and after IR exposure. Under homeostasis, BMPR2+ HSCs exhibited lower intensity of H3K4me3 than BMPR2– HSCs (Fig. 8a), in line with their low transcriptional activities as shown by ATAC-seq. Upon IR, BMPR2+ HSCs also exhibited lower intensity of H3K27me3 than BMPR2– HSCs (Fig. 8a). Based on the distributions of H3K4me3 and H3K27me3 signals, we then classified all genes into four categories: H3K4me3-only, H3K27me3-only, bivalent domains (co-localized with both H3K4me3 and H3K27me3), and none-marked. Interestingly, we found that a collection of genes with bivalent domains in BMPR2– HSCs were switched to active (H3K4me3-only) domains in BMPR2+ HSCs upon IR (Fig. 8b), which were associated with “homeostasis of number of cells”, “response to oxidative stress and radiation” and “regulation of hematopoiesis” (Fig. 8c). Among these genes, Nrf2 exhibited similar bivalent modifications in both BMPR2+ and BMPR2− HSCs under homeostasis, while exhibited markedly reduced repressive H3K27me3 modifications with no obvious alteration of active H3K4me3 marks in BMPR2+ HSCs compared with BMPR2− HSCs after IR (Fig. 8d). These data implied that the bivalent marked gene Nrf2 is primed to be activated under the stimulation of IR. Encouragingly, the qRT-PCR assay demonstrated that under IR stress, the expression levels of Nrf2, Hmox1, and Gpx1 in BMPR2+ HSCs were significantly upregulated compared to BMPR2− HSCs. However, there was no significant difference in their expression between the two cell subgroups in the absence of IR exposure (Fig. 8e). Additionally, CUT&Tag results revealed a significant enrichment of NRF2 binding signals in the Hmox1 and Gsta1 genes in BMPR2⁺ HSCs following IR stimulation (Supplementary Fig. 8a). Together, these results suggest that Nrf2 may play a critical role in the radioresistance ability of BMPR2+ HSCs.
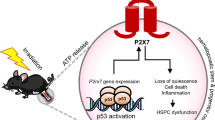
a Centered CUT&Tag peaks of H3K4me3 (±2 Kb of TSS) and H3K27me3 (gene body) in BMPR2⁺ and BMPR2− HSCs at homeostasis (D0) and D1 after 6.5 Gy of IR. b Circos plot showing the switching events across different histone modification (HM) statuses between BMPR2⁺ and BMPR2− HSCs. c Significantly enriched GO terms of genes with switched HM status from bivalent to H3K4me3-only. d CUT&Tag signals of H3K4me3 and H3K27me3 across Nrf2, showing its switched HM status from bivalent to H3K4me3-only. e Fold changes in gene expression levels in BMPR2⁺ and BMPR2⁻ HSCs at D0 and D1 after 6.5 Gy of IR. n = 4 biological repeats per group. f Fold change in CellROX levels in BMPR2⁺ and BMPR2⁻ HSCs derived from Nrf2⁺/⁺ and Nrf2−/− mice at D1 after 6.5 Gy of IR. n = 4 biological repeats per group. g, h Percentage and number of BMPR2⁺ and BMPR2⁻ HSCs from Nrf2−/− mice at D0 and D1 after 6.5 Gy of IR. n = 8 mice per group. i Number of CFUs formed by BMPR2⁺ and BMPR2− HSCs from Nrf2−/− mice at D0 and D1 after 6.5 Gy of IR. n = 3 biological repeats per group. j Schematic diagram of the molecular mechanisms by which the BMP4-BMPR2-NRF2 pathway regulates the radiation resistance of BM HSCs. Data shown are mean ± SEM. Statistical analyses were conducted using two-way ANOVA with multiple comparisons by Tukey’s test (e, g–i) and unpaired two-sided Student’s t test (f). Source data are provided as a Source Data file.
To test this hypothesis, we then employed Nrf2 knockout (Nrf2−/−) mice to explore whether Nrf2 is responsible for radioresistant characteristics of BMPR2+ HSCs (Supplementary Fig. 8b). We observed that the deficiency of Nrf2 leads to no significant suppression of ROS levels and MMP in BMPR2+ HSCs compared with those in BMPR2− HSCs upon IR exposure (Fig. 8f, Supplementary Fig. 8c). Consistently, when Nrf2 gene was deleted, BMPR2+ HSCs showed no obvious increase in their percentage, number, and colony-forming ability compared with those in BMPR2− HSCs after IR stress (Fig. 8g–i), supporting the idea that Nrf2 is the key functional gene for BMPR2+ HSC radioresistance. Furthermore, Nrf2 deficiency or the injection of NRF2 inhibitor Nrf-IN-1 abolished the beneficial roles of BMP4 in the improvement of the survival rate of lethally irradiated mice and hematopoietic colony-forming ability of the in vivo irradiated BM cells (Supplementary Fig. 8d–f). Collectively, these data suggest that Nrf2 is a crucial downstream gene in the BMP4-BMPR2 signaling pathway, which protects HSCs from radiation damage (Fig. 8j).
Discussion
HSPCs are prone to be injured by various stress stimuli, including IR4,47. However, how residual HSPCs respond to high doses of IR and reconstitute the whole hematopoietic system remains less well elucidated. Additionally, it is still lacking the precise molecule-level analysis of the fate of HSPCs upon radiation stress. By utilizing scRNA-seq, we have elucidated the previously inaccessible molecular kinetics and transcriptional differentiation fate skewing that orchestrate the complex in vivo interactions between HSPCs and the BM niche cells following radiation exposure. We found that HSCs exhibit rapid and continuous exhaustion and strong GMP path skewing under the IR-stressed regeneration demand. We further elucidated the intrinsic gene network transitions and extrinsic factors affecting the fate of GMP differentiation after radiation. Our findings provide previously unrecognized insights into the molecular kinetics of radiation stress-conditioned hematopoiesis and offer clues for identifying unexplored targets for protecting HSPCs and rapidly restoring hematopoiesis after radiation.
The need to find previously molecular pathways for protecting HSCs from radiation-stressed damage has yet to be fully realized. Accumulated evidence has shown that BMP4 signaling is indispensable for embryonic hematopoietic development48,49,50,51. BMP4 also exhibits various functions in the maintenance and homing of adult HSCs52,53,54. However, its role in radiation-induced HSC damage has not been investigated. Here, through the combined use of scRNA-seq technology as well as a series of in vitro and in vivo experiments, we discovered a previously unrecognized role of the BMP4 signaling axis that bestows HSCs with radiation-resistant abilities. Encouragingly, we found that a single injection of BMP6 was effective in inhibiting HSPC apoptosis and rescuing mice from radiation-induced mortality (Supplementary Fig. 9). In contrast, BMP4 demonstrated a more potent protective effect against hematopoietic radiation damage. To date, there is a lack of new radioprotectors and radiomitigators approved by the USA Food and Drug Administration55. The promising effect of the BMP4 signaling on hematopoietic radiation protection indicates that this cytokine or its signaling agonist has the potential to become a candidate drug for the treatment of hematopoietic injury.
BMP4 functions through its receptors BMPR1 and BMPR2. However, we observed that the expression level of Bmpr1 in murine HSCs is very low, while some HSCs exhibit high expression of the Bmpr2 gene, as supported by several previous reports50,52. Furthermore, scRNA-seq and flow cytometry detection results revealed that HSCs can be distinguished into Bmpr2+ and Bmpr2− HSCs. More importantly, BMPR2+ HSCs showed higher radioresistance characteristics than BMPR2− HSCs, manifested by lower DNA damage levels, lower oxidative stress injury and higher CFU numbers post-IR. Further transplantation experiments also provided solid evidence for the robust self-renewal and radiation resistance of BMPR2+ HSCs.
Accumulated evidence indicates that HSPCs present different epigenetic programming under various stress stimuli56,57. By using ATAC-seq, we found that BMPR2+ HSCs exhibit a different chromatin accessibility landscape compared to BMPR2− HSCs, particularly on the Nrf2 gene, where open chromatin accessibility was observed after IR stress. In combination with Cut&Tag technique, we further revealed that the Nrf2 gene is bivalently and dynamically epigenetically modified and plays a key role in radioresistance in BMPR2+ HSCs. Nrf2 is a well-known redox modulator, and the increased expression of this gene has been shown to decrease cell apoptosis against IR damage21,58,59,60. In addition, Nrf2 has also been shown to regulate HSC quiescence, survival and regeneration following irradiation21,58,59,60. Here, our experimental data with Nrf2−/− mice further revealed that Nrf2 plays a crucial role in the resistance capacity of BMPR2+ HSCs against IR stress. Taken together, these findings support that BMPR2+ HSCs play a stronger hematopoietic reconstruction function mainly via upregulation of Nrf2 gene expression against IR stress.
In conclusion, this study reveals a dynamic transcriptomic landscape of radiation-induced hematopoietic injury and identifies BMPR2+ HSCs as a crucial cell population for radiation-stressed hematopoietic reconstruction. The BMP4-BMPR2-NRF2 signaling axis is shown to play a crucial role in maintaining the stemness program of HSCs and in providing resistance to radiation injury by regulating the epigenetic program of BMPR2+ HSCs. Encouragingly, BMP4 or its agonist SB4 may have the potential to develop into a radiation protectant for clinical use. These findings provide valuable insights for identifying potential interfering signaling pathways to protect HSPCs and effectively restore hematopoiesis following IR or other stress-related injury.
Methods
Mice
C57BL/6-Ly5.2 (CD45.2) mice were purchased from the Vital River Laboratory Animal Technology (Beijing, China). C57BL/6-Ly5.1 (CD45.1) mice were purchased from the animal facility of the Institute of Hematology at Tianjin (Tianjin, China). Nrf2−/− mice, based on the CD45.2 genetic background, were kindly provided by Professor Yu ZY (Beijing Institute of Radiation Medicine, Beijing, China). All mice were housed in a specific pathogen-free (SPF) barrier environment and were continuously provided with sterilized food, water, and bedding. All animal experiments in this study were approved by the Animal Care and Use Committee of Beijing Institute of Radiation Medicine (Beijing, China).
Radiation and treatment
To establish an in vivo radiation-induced BM injury model, mice were subjected to total-body irradiation using a ⁶⁰Co irradiator at a dose of 6.5 Gy. For the in vitro BM injury model, an X-ray apparatus was used to irradiate mouse BM cells with a dose of 3.0 Gy.
To evaluate the radiation-protective effects of BMP4 or SB4, mice were administered injections of 100 μg/kg BMP4 (R&D Systems), 1 mg/kg SB4 (MedChemExpress), or vehicle control 2 hours prior to irradiation. For survival experiments, mice were subcutaneously injected with various doses of BMP4 (5, 25, 50, 100, and 200 μg/kg) or intraperitoneally injected with various doses of SB4 (1, 2.5, and 5 mg/kg), or vehicle, 2 hours before exposure to 8.0 Gy irradiation. After irradiation, the mice were immediately returned to a SPF barrier environment. Daily observations and recordings were made of the mice’s survival status and weight changes. BMP4 and SB4 were prepared into corresponding solutions according to the manufacturer’s instructions.
Flow cytometric analyses and fluorescence-activated cell sorting (FACS)
For analyses of hematopoietic cell composition, BM cells were obtained by crushing femur and tibia bones in IMDM supplemented with 2% fetal bovine serum (FBS) and 1% Penicillin/Streptomycin. Cells were then stained with an antibody cocktail diluted in FACS buffer on ice. The cells were stained with the following antibodies conjugated to FITC, PE, PE-Cy7, APC, APC-eFluor780, or eFluor450: anti-Lineage Cocktail (17A2, RA3-6B2, M1/70, TER-119, RB6-8C5; eBioscience), anti-c-Kit (2B8; eBioscience), anti-Sca-1 (D7; eBioscience), anti-CD48 (HM48-1; eBioscience), anti-CD150 (mShad150; eBioscience), anti-CD45.1 (A20; eBioscience), anti-CD45.2 (104; eBioscience), anti-CD3e (145-2C11; eBioscience), anti-CD4 (RM4-5; eBioscience), anti-CD8a (53-6.7; eBioscience), anti-Gr-1 (RB6-8C5; eBioscience), anti-CD11b (M1/70; eBioscience), and anti-B220 (RA3-6B2; eBioscience). Dead cells were excluded using 7-Amino-Actinomycin-D (7AAD) staining. To further detect ROS or mitochondrial activity, the following fluorescent probes were used: CellROX™, MitoSOX™, TMRM, and MitoTracker® Deep Red FM (Thermo Fisher Scientific). Cells were analyzed using a BD Aria II (BD Biosciences). For detecting and quantifying foci of DNA damage, cells pre-labeled with surface marker antibodies were fixed and permeabilized using a Cytofix/Cytoperm™ Fixation/Permeabilization Kit (BD Biosciences). They were then stained with antibodies specific for γH2AX (Cell Signaling Technology) overnight at 4 °C, followed by washing and resuspension in PBS. Cells were analyzed using an Amnis ImageStream-X Mk II imaging flow cytometer (Luminex).
For the enrichment of c-Kit+ or Lin− cells, BM cells were stained with c-Kit-biotin or lineage-biotin antibodies, followed by magnetic bead enrichment (Miltenyi Biotec). For sorting LSKCD48⁻CD150⁺ hematopoietic stem cells (HSCs), c-Kit+ or Lin− cells were stained with combinations of antibodies against the following surface markers: anti-Sca-1, anti-c-Kit, anti-CD150, anti-CD48, and anti-Lineage Cocktail. Dead cells were excluded using 7AAD. The cells were isolated using a BD Influx (BD Biosciences) or BD Aria SORP Cell Sorter (BD Biosciences). Additional details regarding the flow cytometry reagents are provided in Supplementary Data 11.
HSC transplantation
Recipient mice (CD45.1) were subjected to whole-body lethal irradiation using a ⁶⁰Co irradiator, receiving two doses of 4.5 Gy separated by 1 hour. Subsequently, the mice were reconstituted via intravenous injection with either 100 BMPR2⁺ or BMPR2⁻ HSCs from CD45.2 mice, or 2000 BMPR2⁺ or BMPR2⁻ HSCs from CD45.2 mice that had undergone 6.5 Gy irradiation. Each transplant also included 2 × 10⁴ BM cells from CD45.1 mice as competitors. For long-term reconstitution analysis, 2 × 10⁶ chimeric BM cells were injected into the recipients, and chimerism was assessed every 4 weeks following reconstitution in both primary and secondary transplantations.
Colony-forming cell (CFC) assay
The CFC assay was performed with slight modifications to the method previously reported61. In summary, BM cells, LSK cells, or HSCs were plated at densities of 1 × 10⁴, 1000, and 100 cells per dish, respectively, from the BM of non-irradiated mice, and at densities of 1 × 10⁵, 3000, and 1000 cells per dish, respectively, from the BM of irradiated mice. The cells were cultured in MethoCult™ GF M3434 Medium (STEMCELL). After a 7-day incubation period at 37 °C in an atmosphere of 5% CO₂ and 5% O₂, colonies were identified and counted under a microscope.
Comet assay
The Comet assay was performed using a Comet Assay Kit (Trevigen, MD, USA). Cells were resuspended in Comet Assay Low-Melting-Point Agarose and spread onto CometSlides. The slides were then immersed in lysis solution and incubated overnight at 4 °C. After lysis, electrophoresis was conducted to assess DNA damage. Following electrophoresis, the slides were immersed in DNA precipitation solution for 30 minutes, then transferred to a 70% ethanol solution for an additional 30 minutes. Once dried, the slides were stained with SYBR Gold Nucleic Acid Gel Stain (Thermo Fisher Scientific) to visualize DNA. The presence of comet tails, indicative of DNA damage, was examined using a fluorescence microscope (Nikon, Japan). The analysis of the comets was performed using Comet Analysis Software (Trevigen).
Quantitative RT-PCR
BMPR2⁺ or BMPR2⁻HSCs were directly sorted into lysis buffer, and total RNA was isolated using the RNeasy® Mini Kit (QIAGEN). The cycle threshold (CT) values were averaged, and the relative expression levels compared to the housekeeping gene Hprt were calculated using the 2−ΔCT formula. Primer sequences utilized in this assay were sourced from PrimerBank (https://pga.mgh.harvard.edu/primerbank/), and are detailed in Supplementary Data 12. RT-PCR reactions were performed using the CFX Connect Real-Time PCR Detection System (Bio-Rad).
Apoptosis assay
For the Caspase 3/7 activity assay, 1 × 10⁴ LSK cells or 2000 HSCs were re-suspended in 100 μL of cytokine-free StemSpan II medium, and 100 μL of Caspase-Glo® 3/7 reagent (Promega) was added directly to each well. For the Annexin V activity assay, 1 × 10⁴ LSK cells were re-suspended in 100 μL of cytokine-free StemSpan II medium, and 100 μL of the 2× Detection Reagent from the Real-Time-GloTM Annexin V Apoptosis and Necrosis Assay (Promega) was added directly to each well. After incubation for 1 hour, luminescence was measured using a microplate reader (Molecular Devices), and blank control values were subtracted from the readings of each well.
Single-cell RNA sequencing
At various time points post-irradiation (D1, D3, D7, D14, and D21), the irradiated mice were humanely euthanized, and their Lin⁻ cells were isolated. The enriched Lin⁻ cells were converted into barcoded scRNA-seq libraries using the Chromium Single-Cell 3’ Library & Gel Bead Kit, Multiplex Kit, and Chip Kit (10× Genomics, CA, USA), following the manufacturer’s instructions, with an estimated target of 1 × 10⁴ cells per library. The samples were processed using the V3.1 barcoding chemistry kits from 10× Genomics. Single samples were always processed in a single well of a PCR plate, ensuring that all cells from a sample were treated with the same master mix and in the same reaction vessel. All samples were processed in parallel in the same thermal cycler. The generated scRNA-seq libraries were sequenced on a NovaSeq sequencer (Illumina) at CapitalBio (Beijing, China).
Gene expression quantification, quality control and batch correction for scRNA-seq
The Cell Ranger software (version 4.0.0; 10× Genomics) was used for sample demultiplexing, barcode processing, and single-cell 3’ counting. The fastq files for each sample were processed using the count function in Cell Ranger, which aligned the reads to the mouse genome (build mm10) and quantified gene expression levels in single cells. To filter out low-quality cells, cells with fewer than 500 or >5000 expressed genes were removed62. To exclude dead or dying cells, cells with over 20% unique molecular identifiers (UMIs) derived from the mitochondrial genome were further removed. Gene expression in single cells was normalized using the SCTransform function in the R package Seurat (v4.0). The anchor-based batch correction method was employed to merge samples from different groups63.
Cells clustering and hematopoietic cell clusters annotation of scRNA-seq
Principal component analysis (PCA) was performed for dimensionality reduction, and 30 principal components (PCs) were selected for further analysis. UMAP visualization was used to embed the neighborhood graph and display the topology of the data. For cell clustering, the FindClusters function in Seurat (v4.0) was employed, which implements a shared nearest neighbor (SNN) modularity optimization-based clustering algorithm. The Scrublet package was used to identify doublets (two cells encapsulated in a single droplet) from the scRNA-seq data using default parameters. Cell clusters with extremely low nUMI counts and a high proportion of doublets were designated as low-quality clusters and were excluded from further analyses (Supplementary Fig. 1c, d). Major cell types were annotated using the canonical marker genes of hematopoietic cell types (Supplementary Fig. 1e, f). The mixed HSCs/MPPs cluster was re-clustered in a second round using the same procedure described above, and cluster annotation was performed using a set of previously reported canonical marker genes for HSPCs (Supplementary Fig. 2a, b).
Identification of marker genes for cell clusters
To identify marker genes for major cell types and cell subpopulations (clusters), we compared cells from a major type or subpopulation to all other cells using the FindAllMarkers function in Seurat (v4.0). This function identifies differentially expressed genes between two groups of cells using a Wilcoxon rank-sum test. P values were then corrected using the Bonferroni correction based on the total number of genes in the dataset. Marker genes were selected based on the following criteria: an adjusted P value < 0.01, an average expression level in that cluster at least two-fold higher than the average expression level in other clusters, and a detection rate higher than 25% in that cluster.
Lineage score calculation
GSEA was performed to identify up- and downregulated lineage-specific gene modules in HSPC subpopulations of mice at different time points before and after radiation64. Gene modules with an enrichment score (ES) > 0 and a false discovery rate (FDR) < 0.05 were defined as upregulated modules, while those with ES < 0 and FDR < 0.05 were defined as downregulated modules. Lineage signature gene sets, including erythroid, megakaryocytic, myeloid, lymphoid, and proliferation, were obtained from a previous study65.
Trajectory analysis and gene expression trend fitting
To reconstruct the differential trajectories among HSPCs, we derived HSCs, MPPs, CLPs, MEPs, and GMPs from the global scRNA-seq dataset and performed UMAP visualization on these cells. We used the partition-based graph abstraction (PAGA) method66 to construct the trajectory tree for all groups of cells, which indicates the distinct lineage differentiation potential for each group. PAGA analysis was implemented using the scanpy.tl.paga function in the Python package Scanpy. The root cell was defined as the cell expressing the highest level of Hlf in the LT-HSC cluster. Diffusion Pseudotime, introduced by Haghverdi et al.67, was used to reconstruct the progression of a biological process from snapshot data. This analysis was implemented using the scanpy.tl.dpt function in the Scanpy package. The gene expression trend versus pseudotime was fitted using the Local Polynomial Regression model implemented in the stats R package.
Weighted correlation network analysis
Weighted correlation network analysis (WGCNA) was performed using the WGCNA algorithm68. Based on the transcriptome-wide gene expression profiles of five HSPC subpopulations (LT-HSC, ST-HSC/MPP1, MPP2, MPP3, and MPP4), we constructed a weighted correlation network, which revealed six gene modules. The relationships between these gene modules and time points after irradiation or HSPC subpopulations were assessed using Pearson’s correlation coefficients.
Gene set variation analysis (GSVA)
To assess the activity scores of pathways or gene modules at the single-cell level, we performed GSVA using the GSVA package (version 1.32.0)69. The 50 hallmark gene sets (version 6.2), representing specific, well-defined biological states or processes, were obtained from the Molecular Signatures Database (MSigDB)70. GSVA was conducted using the gsva function with standard settings. We then fitted a linear model to the output gene set-by-cell pathway enrichment matrix using the lmFit function from the limma package (version 3.40.6) to detect differentially enriched pathways.
To calculate a cytokine score for each cell, we obtained the human cytokine gene list from a previously published paper71 and converted it to their mouse orthologs. GSVA was then performed using this cytokine gene set to assign a cytokine score to each cell based on its expression profile.
Cell-cell interaction analysis
The cell-cell interaction analysis was based on the expression of specific ligands (Ls) and receptors (Rs). A total of 1642 literature-supported and manually curated LR interactions, collected by the Fantom5 and CellPhoneDB databases72, were used for this analysis. As described in our previous paper73, we established the potential interaction between two cell clusters mediated by a specific LR pair by calculating the product of the average expression levels of the ligand in one cell cluster and the corresponding receptor in the other cluster.
To examine the statistical significance of the estimated interaction intensity, permutations were applied to the cell cluster labels of individual cells 1000 times, and the P value was estimated by the number of permutations that had interaction intensity greater than the real value. The adjusted P value, calculated using Bonferroni correction, was used for multiple testing correction across the hundreds of LR pairs. An LR pair was defined as a potential molecular axis mediating interactions between two cell clusters if it had an interaction intensity greater than 1 and an adjusted P value <0.01. For a given LR pair, cell clusters with the average expression level of either the ligand or the receptor less than 1 (log2(Normalized Counts) < 1) were excluded from the analysis.
The cellular communication intensity between two cell types was defined as the number of significant LR interactions between them, weighted by the number of cells in the corresponding tissue. LR interactions were classified into clusters based on their temporal profiles of weighted interaction intensity across six time points. This clustering was performed using the C-means unsupervised clustering algorithm with the timeclust function in the TCseq R package (version 1.10.0). The optimal number of clusters was determined using the fviz_nbclust function in the factoextra R package (version 1.0.7).
TF activity calculation
SCENIC analysis (version 0.9.11) was used to evaluate the activity levels of TFs. Two TF ranking databases, namely the TSS ± 10 kilobase pairs (kb) and the 500bpUp100Dw databases, were employed for the inference of co-expression modules and the identification of direct targets. A regulon-by-cell matrix was then calculated to measure the enrichment of each regulon as the area under the recovery curve of the genes defining the regulon. Differential analysis was implemented using the lmFit function from the limma package (version 3.40.6).
ATAC-seq
Library preparation for the assay for transposase-accessible chromatin with high-throughput sequencing (ATAC-seq) was performed on BMPR2⁻LSKCD48⁻CD150⁺ HSCs and BMPR2⁺LSKCD48⁻CD150⁺ HSCs from unirradiated and irradiated mice using the Nextera DNA Sample Preparation Kit (Illumina, CA, USA), as described previously74. Four ATAC-seq libraries were sequenced per lane on a HiSeq 2500 System (Illumina) to generate paired-end 150-bp reads at Jiayin Biotechnology (Shanghai, China). Reads were mapped against the mm10 mouse reference genome using BWA with default parameters. MACS2 (version 2.0.10) was used to call peaks with a uniform extension size of 150 bp. BAM files were converted to bigwig format using deeptools with the command bamCoverage -binSize 20 -normalizeUsing CPM -smoothLength 60. Heatmaps were created using the computeMatrix and plotHeatmap functions in deeptools. Motif enrichment was performed using HOMER against a catalog of all peaks called in any one sample. This catalog was produced by merging all called peaks that overlapped by at least one base pair using bedtools.
CUT&Tag assay
BMPR2⁻ and BMPR2⁺ HSCs from unirradiated and irradiated mice were collected as inputs. Libraries were prepared using the CUT&Tag-ITTM Assay Kit (Vazyme) according to the manufacturer’s protocol. The following antibodies were used for the reaction: anti-H3K4me3 (CST), anti-H3K27me3 (CST), and anti-NRF2 (CST). Sequencing was performed on a HiSeq 4000 (Illumina) with a paired-end read length of 150 bp at Annoroad Gene Technology (Beijing, China). Sequences were aligned to the mouse genome (mm10) using Bowtie2 with default settings. For peak calling and data normalization, MACS2 and deeptools were utilized as described in the ATAC-seq section.
To define active, bivalent, and inactive enhancers, we assessed the overlap of CUT&Tag peaks of H3K4me3 and H3K27me3. A gene promoter region was considered active if it overlapped with H3K4me3 but not with H3K27me3; bivalent if it overlapped with both H3K4me3 and H3K27me3; and inactive if it overlapped with H3K27me3 but not with H3K4me3. Sequencing reads were visualized using the IGV software.
Statistical analyses
Statistical analyses were performed using GraphPad Prism. Data are presented as mean ± standard error of the mean. Statistical significance was determined using one-way or two-way ANOVA with Tukey’s multiple comparisons test, two-sided log-rank test, or unpaired two-tailed Student’s t test. A P value of <0.05 was considered statistically significant. All experiments were independently repeated two or three times.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The raw sequence data generated in this paper have been deposited in the Genome Sequence Archive75 in the National Genomics Data Center76, China National Center for Bioinformation/Beijing Institute of Genomics, Chinese Academy of Sciences under the accession code PRJCA028965. Source data are provided with this paper.
Code availability
All the scripts and bioinformatic codes used in this paper were available in our public website (https://www.omic.tech/pubfiles/source_code_BM_scRNA-seq.zip).
References
Till, J. E. & McCULLOCH, E. A. A direct measurement of the radiation sensitivity of normal mouse bone marrow cells. Radiat. Res. 14, 213–222 (1961).
Laurenti, E. & Göttgens, B. From haematopoietic stem cells to complex differentiation landscapes. Nature 553, 418–426 (2018).
Kondo, M. et al. Biology of hematopoietic stem cells and progenitors: implications for clinical application. Annu. Rev. Immunol. 21, 759–806 (2003).
Singh, S., Jakubison, B. & Keller, J. R. Protection of hematopoietic stem cells from stress-induced exhaustion and aging. Curr. Opin. Hematol. 27, 225–231 (2020).
Shao, L., Luo, Y. & Zhou, D. Hematopoietic stem cell injury induced by ionizing radiation. Antioxid. Redox Signal. 20, 1447–1462 (2014).
Hu, L. et al. Radiation-induced bystander effects impair transplanted human hematopoietic stem cells via oxidative DNA damage. Blood 137, 3339–3350 (2021).
Shen, H. et al. An acute negative bystander effect of γ-irradiated recipients on transplanted hematopoietic stem cells. Blood 119, 3629–3637 (2012).
Lorimore, S. A. & Wright, E. G. Radiation-induced genomic instability and bystander effects: related inflammatory-type responses to radiation-induced stress and injury? A review. Int. J. Radiat. Biol. 79, 15–25 (2003).
Termini, C. M. & Chute, J. P. Hematopoietic stem cell stress and regeneration. Curr. Stem Cell Rep. 6, 134–143 (2020).
Kaufmann, K. B. et al. A latent subset of human hematopoietic stem cells resists regenerative stress to preserve stemness. Nat. Immunol. 22, 723–734 (2021).
Gulati, G. S. et al. Neogenin-1 distinguishes between myeloid-biased and balanced Hoxb5+ mouse long-term hematopoietic stem cells. Proc. Natl. Acad. Sci. USA 116, 25115–25125 (2019).
Mann, M. et al. Heterogeneous responses of hematopoietic stem cells to inflammatory stimuli are altered with age. Cell Rep. 25, 2992–3005.e5 (2018).
Scaramozza, A. et al. Lineage tracing reveals a subset of reserve muscle stem cells capable of clonal expansion under stress. Cell Stem Cell 24, 944–957.e5 (2019).
Metcalfe, C., Kljavin, N. M., Ybarra, R. & de Sauvage, F. J. Lgr5+ stem cells are indispensable for radiation-induced intestinal regeneration. Cell Stem Cell 14, 149–159 (2014).
Roche, K. C. et al. SOX9 maintains reserve stem cells and preserves radioresistance in mouse small intestine. Gastroenterology 149, 1553–1563.e10 (2015).
Velardi, E. et al. Suppression of luteinizing hormone enhances HSC recovery after hematopoietic injury. Nat. Med. 24, 239–246 (2018).
Patterson, A. M. et al. A single radioprotective dose of Prostaglandin E2 blocks irradiation-induced apoptotic signaling and early cycling of hematopoietic stem cells. Stem Cell Rep. 15, 358–373 (2020).
Nakamura, S. et al. Bmi1 confers resistance to oxidative stress on hematopoietic stem cells. PLoS ONE7, e36209 (2012).
Tothova, Z. et al. FoxOs are critical mediators of hematopoietic stem cell resistance to physiologic oxidative stress. Cell 128, 325–339 (2007).
Hu, L., Zhang, Y., Miao, W. & Cheng, T. Reactive oxygen species and Nrf2: functional and transcriptional regulators of hematopoiesis. Oxid. Med. Cell Longev. 2019, 5153268 (2019).
Tsai, J. J. et al. Nrf2 regulates haematopoietic stem cell function. Nat. Cell Biol. 15, 309–316 (2013).
Burkhardt, D. B. et al. Quantifying the effect of experimental perturbations at single-cell resolution. Nat. Biotechnol. 39, 619–629 (2021).
Aibar, S. et al. SCENIC: single-cell regulatory network inference and clustering. Nat. Methods 14, 1083–1086 (2017).
Langfelder, P. & Horvath, S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinf. 9, 559 (2008).
Rodriguez-Fraticelli, A. E. et al. Single-cell lineage tracing unveils a role for TCF15 in haematopoiesis. Nature 583, 585–589 (2020).
Laurenti, E. et al. Hematopoietic stem cell function and survival depend on c-Myc and N-Myc activity. Cell Stem Cell 3, 611–624 (2008).
Balazs, A. B., Fabian, A. J., Esmon, C. T. & Mulligan, R. C. Endothelial protein C receptor (CD201) explicitly identifies hematopoietic stem cells in murine bone marrow. Blood 107, 2317–2321 (2006).
Matsumoto, A. et al. p57 is required for quiescence and maintenance of adult hematopoietic stem cells. Cell Stem Cell 9, 262–271 (2011).
Calvanese, V. et al. MLLT3 governs human haematopoietic stem-cell self-renewal and engraftment. Nature 576, 281–286 (2019).
Buitenhuis, M. et al. Protein kinase B (c-akt) regulates hematopoietic lineage choice decisions during myelopoiesis. Blood 111, 112–121 (2008).
Pietras, E. M. Inflammation: a key regulator of hematopoietic stem cell fate in health and disease. Blood 130, 1693–1698 (2017).
Zhao, J. L. et al. Conversion of danger signals into cytokine signals by hematopoietic stem and progenitor cells for regulation of stress-induced hematopoiesis. Cell Stem Cell 14, 445–459 (2014).
Mossadegh-Keller, N. et al. M-CSF instructs myeloid lineage fate in single haematopoietic stem cells. Nature 497, 239–243 (2013).
Gonda, T. J., Macmillan, E. M., Townsend, P. V. & Hapel, A. J. Differentiation state and responses to hematopoietic growth factors of murine myeloid cells transformed by myb. Blood 82, 2813–2822 (1993).
Bruce, A. G., Hoggatt, I. H. & Rose, T. M. Oncostatin M is a differentiation factor for myeloid leukemia cells. J. Immunol. 149, 1271–1275 (1992).
Wang, L., Jiang, J., Chen, Y., Jia, Q. & Chu, Q. The roles of CC chemokines in response to radiation. Radiat. Oncol. 17, 63 (2022).
Piryani, S. O., Kam, A. Y. F., Vu, U. T., Chao, N. J. & Doan, P. L. CCR5 signaling promotes murine and human hematopoietic regeneration following ionizing radiation. Stem Cell Rep. 13, 76–90 (2019).
Nie, Y., Han, Y.-C. & Zou, Y.-R. CXCR4 is required for the quiescence of primitive hematopoietic cells. J. Exp. Med. 205, 777–783 (2008).
Wang, W. et al. Notch2 blockade enhances hematopoietic stem cell mobilization and homing. Haematologica 102, 1785–1795 (2017).
Kara, N. et al. Endothelial and leptin receptor+ cells promote the maintenance of stem cells and hematopoiesis in early postnatal murine bone marrow. Dev. Cell 58, 348–360.e6 (2023).
Ding, L., Saunders, T. L., Enikolopov, G. & Morrison, S. J. Endothelial and perivascular cells maintain haematopoietic stem cells. Nature 481, 457–462 (2012).
Ghaffari, S. Haematopoietic stem cell quiescence exposed using mitochondrial membrane potential. Curr. Opin. Hematol. 30, 1–3 (2023).
Qiu, J. et al. Using mitochondrial activity to select for potent human hematopoietic stem cells. Blood Adv. 5, 1605–1616 (2021).
Yu, S. et al. BMP4 resets mouse epiblast stem cells to naive pluripotency through ZBTB7A/B-mediated chromatin remodelling. Nat. Cell Biol. 22, 651–662 (2020).
Sun, Y. et al. Context-dependent tumor-suppressive BMP signaling in diffuse intrinsic pontine glioma regulates stemness through epigenetic regulation of CXXC5. Nat. Cancer 3, 1105–1122 (2022).
Wei, Y. et al. Efficient derivation of human trophoblast stem cells from primed pluripotent stem cells. Sci. Adv. 7, eabf4416 (2021).
Giladi, A. et al. Single-cell characterization of haematopoietic progenitors and their trajectories in homeostasis and perturbed haematopoiesis. Nat. Cell Biol. 20, 836–846 (2018).
Gupta, S., Zhu, H., Zon, L. I. & Evans, T. BMP signaling restricts hemato-vascular development from lateral mesoderm during somitogenesis. Development 133, 2177–2187 (2006).
Bhatia, M. et al. Bone morphogenetic proteins regulate the developmental program of human hematopoietic stem cells. J. Exp. Med. 189, 1139–1148 (1999).
Crisan, M. et al. BMP signalling differentially regulates distinct haematopoietic stem cell types. Nat. Commun. 6, 8040 (2015).
Zhang, B. et al. Prostaglandin E2 is required for BMP4-induced mesoderm differentiation of human embryonic stem cells. Stem Cell Rep. 10, 905–919 (2018).
Warsi, S. et al. BMP signaling is required for postnatal murine hematopoietic stem cell self-renewal. Haematologica 106, 2203–2214 (2021).
Goldman, D. C. et al. BMP4 regulates the hematopoietic stem cell niche. Blood 114, 4393–4401 (2009).
Khurana, S. et al. A novel role of BMP4 in adult hematopoietic stem and progenitor cell homing via Smad independent regulation of integrin-α4 expression. Blood 121, 781–790 (2013).
Obrador, E. et al. Radioprotection and radiomitigation: from the bench to clinical practice. Biomedicines 8, 461 (2020).
Martin, E. W. et al. Chromatin accessibility maps provide evidence of multilineage gene priming in hematopoietic stem cells. Epigenet. Chromatin 14, 2 (2021).
Itokawa N. et al. Epigenetic traits inscribed in chromatin accessibility in aged hematopoietic stem cells. Nat. Commun. 13, 2691 (2022).
Kim, J.-H. et al. NRF2-mediated Notch pathway activation enhances hematopoietic reconstitution following myelosuppressive radiation. J. Clin. Invest. 124, 730–741 (2014).
Chute, J. P. NRF2 mitigates radiation-induced hematopoietic death. J. Clin. Invest. 124, 960–961 (2014).
Dai, X. et al. Nrf2: redox and metabolic regulator of stem cell state and function. Trends Mol. Med. 26, 185–200 (2020).
Wang, C. et al. Recombinant human thrombopoietin promotes hematopoietic reconstruction after severe whole body irradiation. Sci. Rep. 5, 12993 (2015).
Wolock, S. L., Lopez, R. & Klein, A. M. Scrublet: computational identification of cell doublets in single-cell transcriptomic data. Cell Syst. 8, 281–291.e9 (2019).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 102, 15545–15550 (2005).
Dong, F. et al. Differentiation of transplanted haematopoietic stem cells tracked by single-cell transcriptomic analysis. Nat. Cell Biol. 22, 630–639 (2020).
Wolf, F. A. et al. PAGA: graph abstraction reconciles clustering with trajectory inference through a topology preserving map of single cells. Genome Biol. 20, 59 (2019).
Haghverdi, L., Büttner, M., Wolf, F. A., Buettner, F. & Theis, F. J. Diffusion pseudotime robustly reconstructs lineage branching. Nat. Methods 13, 845–848 (2016).
Zhang, B. & Horvath, S. A general framework for weighted gene co-expression network analysis. Stat. Appl. Genet. Mol. Biol. 4, Article17 (2005).
Hänzelmann, S., Castelo, R. & Guinney, J. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinf. 14, 7 (2013).
Castanza, A. S. et al. Extending support for mouse data in the Molecular Signatures Database (MSigDB). Nat. Methods 20, 1619–1620 (2023).
Ren, X. et al. COVID-19 immune features revealed by a large-scale single-cell transcriptome atlas. Cell 184, 1895–1913.e19 (2021).
Efremova, M., Vento-Tormo, M., Teichmann, S. A. & Vento-Tormo, R. CellPhoneDB: inferring cell-cell communication from combined expression of multi-subunit ligand-receptor complexes. Nat. Protoc. 15, 1484–1506 (2020).
Lu, Y. et al. A single-cell atlas of the multicellular ecosystem of primary and metastatic hepatocellular carcinoma. Nat. Commun. 13, 4594 (2022).
Qin, J. et al. Direct chemical reprogramming of human cord blood erythroblasts to induced megakaryocytes that produce platelets. Cell Stem Cell 29, 1229–1245.e7 (2022).
Chen, T. et al. The genome sequence archive family: toward explosive data growth and diverse data types. Genom. Proteom. Bioinf. 19, 578–583 (2021).
CNCB-NGDC Members and Partners Database resources of the national genomics data center, China national center for bioinformation in 2024. Nucleic Acids Res. 52, D18–D32 (2024).
Acknowledgements
We thank Dr. Dongmei Wang for her helpful discussions and language editing; Mrs. Bin Yu and Mr. Yan Liu for their technical support. This work was supported by the National Natural Science Foundation of China (No: 82270132, Y.H.L., 82200122, Y.Q.L., 32270714, Y.M.L., and 32200511, C.Q). Cartoon elements used in schematic diagrams in Figs. 1a, 4l, 6c, and Supplementary Fig. 6a were created using Figdraw (Copyright Code: YISTTdd82d).
Author information
Authors and Affiliations
Contributions
Y.H.L., Y.M.L., X.T.P., and G.Q.Z. conceived the study, designed the experiments, interpreted the results, and wrote the paper. Y.H.L., Y.X.L., B.W.Z., J.H.Z. conducted the experiments, analyzed the data and prepared the draft manuscript. J.H.Q., Y.Q.L., Y.Z.C., J.S.L., K.Y.C., and Y.L. assisted with mouse experiments and flow cytometry analysis. Y.M.L., Y.H.L., S.A.J., T.F., H.L., and C.Q. analyzed the single-cell sequencing data. Z.Y.Y. provided Nrf2−/− mouse model. All authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Toshio Suda, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Li, Y., Li, Y., Zhang, B. et al. Single-cell transcriptomics reveals BMP4-BMPR2 signaling promotes radiation resistance in hematopoietic stem cells following injury. Nat Commun 16, 5470 (2025). https://doi.org/10.1038/s41467-025-60557-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-60557-z