Abstract
Recent studies have highlighted the importance of mitochondria in NP cells and articular chondrocyte health. Since the understanding of mechanisms governing mitochondrial dynamics in these tissues is lacking, we investigated the role of OPA1, a mitochondrial fusion protein, in their homeostasis. OPA1 knockdown in NP cells altered mitochondrial size and cristae shape and increased the oxygen consumption rate. OPA1 governed the morphology of multiple organelles, including peroxisomes, early endosomes and cis-Golgi and loss resulted in the dysregulation of autophagy. Metabolic profiling and 13C-flux analyses revealed TCA cycle anaplerosis and altered metabolism in OPA1-deficient NP cells. Noteworthy, Opa1AcanCreERT2 mice showed age-dependent disc degeneration, osteoarthritis, and vertebral osteopenia. RNA-Sequencing of Opa1cKO NP tissue revealed dysregulation of metabolism, autophagy, cytoskeletal reorganization, and extracellular matrix and shared strong thematic similarities with a subset of human degenerative NP samples. Our findings underscore that maintenance of mitochondrial dynamics and multi-organelle cross-talk is critical in preserving metabolic homeostasis of disc and cartilage.
Similar content being viewed by others
Introduction
Chronic low back pain, associated with intervertebral disc degeneration, and knee and hip joint pain, a common sequela of osteoarthritis, are the leading causes of disability in the aging population1. In a healthy state, these tissues provide the joint with flexibility and the ability to absorb applied forces. However, with the onset of degenerative disease, these properties are lost, and the balance between cell survival, autophagy, and apoptosis becomes dysregulated leading to altered extracellular matrix (ECM) production, and increased tissue catabolism2,3. Nucleus pulposus (NP) cells of the disc and articular chondrocytes reside in an avascular, hypoxic environment and express the transcription factor HIF-1α and exhibit limited replication and regenerative capacity4,5. Consequently, these primarily glycolytic cells were thought to rely minimally on mitochondria to meet their energetic and biosynthetic demands5,6,7. However, mounting evidence suggests that mitochondrial dysfunction promotes osteoarthritis development3. The deletion of mitochondrial superoxide dismutase 2 exacerbated, whereas overexpression of catalase or peroxiredoxin 3 reduced the severity of age-associated osteoarthritis in mice, implying that mitochondrial ROS generation plays a role in the pathogenesis of osteoarthritis8,9,10. Moreover, we have recently shown that functional mitochondrial networks exist in NP cells, and to meet metabolic requirements, NP cells evidence an active mitophagic flux governed by the HIF-1α-BNIP3 axis6. Notably, loss of BNIP3 in NP cells resulted in mitochondrial dysfunction, triggering meta-inflammation, and causing early onset of disc degeneration5 corroborating earlier studies suggesting association of mitochondrial dysfunction with disc degeneration11.
Mitochondria undergo fission and fusion to control their mass and numbers which is governed by the fission protein DRP1 and the outer mitochondrial membrane (OMM) fusion proteins MFN1, MFN2, and inner membrane (IMM) protein OPA112,13. During fusion, mitochondria control stress and energy needs by incorporating components of the damaged organelles to modulate apoptosis, and calcium signaling14. Notably, OPA1 regulates cristae remodeling and apoptosis, independent of mitochondria fusion15 and ablation results in cristae disorganization and release of cytochrome c16. Conversely, OPA1 overexpression facilitates the formation and stability of respiratory chain supercomplexes17. OPA1 also functions as a sensor of metabolic changes through interactions with IMM solute transporters, facilitating the adaptation of cristae shape and respiration to cellular energy needs. Despite, the increasingly important role of mitochondria in disc and cartilage the foundational understanding of mechanisms governing mitochondrial dynamics and function in these tissues is lacking.
Herein, we investigated the role of OPA1 in disc and cartilage health. For the first time, we report the importance of OPA1 in maintaining the autophagy, metabolism and morphology of several organelles in the NP cells and show that conditional deletion of OPA1 (Opa1AcanCreERT2) causes disc degeneration, osteoarthritis and osteopenia in aged mice. Our findings establish that OPA1 is critical in maintaining health and function of the disc and knee articular cartilage.
Results
OPA1 maintains mitochondrial and multi-organelle morphology in NP cells
We knocked down Opa1 in primary NP cells using lentivirally delivered ShRNAs (Fig. 1A, A’). OPA1-knockdown (OPA1-KD) NP cells showed increased mitochondrial fragmentation, evident from the smaller aspect ratio and form factor (Fig. 1A, A”). TEM imaging showed that mitochondria in knockdown NP cells were smaller, with aberrant cristae. Although some cristae were often detached, the tubular structure was preserved with narrow cristae width (Fig. 1B). Notably, OPA1-KD NP cells contained more mitochondria but without changes in mtDNA content, indicating that altered cristae and mitochondria morphology are an early consequence of OPA1 deletion18 (Fig. 1C, D). Moreover, the levels of MFN1, and MFN2 (Fig. S1 A, A’) and DRP1 and its receptors FIS1 and MFF were unaffected in OPA1-KD cells (Fig. S1 A, A’). Together these findings showed that OPA1 deficiency affects mitochondrial number, shape, and cristae morphology without compensatory changes in other fusion and/or fission proteins.
A Immunofluorescence staining for OPA1 and MitoTracker Red in primary NP cells transduced with lentivirally delivered control (ShCtrl) and Opa1 (ShOpa1 #1 and ShOpa1 #2) ShRNAs. The staining experiment was performed 2 times independently. Scale bar: Top rows- 15 μm, bottom row- 2 μm. A’ Western blot to confirm OPA1 knockdown in NP cells. A” Mitochondrial morphology and network analysis. B TEM images of ShCtrl and ShOpa1 transduced cells. Scale bar: Top row − 600 nm, bottom row − 200 nm C Mitochondrial number measurements from OPA1-deficient NP cells; 28-30 cells were measured from two independent experiments. D The mtDNA content in control and OPA1-deficient cells. Data are represented as box and whisker plots showing all data points with median and interquartile range and maximum and minimum values. Statistical significance was performed using One-way ANOVA or Kruskal-Wallis test with Sidak’s multiple comparisons test as appropriate (A’, C, D). E Staining for peroxisome marker PMP70, early endosome marker EEA1, and cis-Golgi marker GM130. Scale bar: 10 and 2 μm E’, E” Morphology analysis of PMP70, EEA1 shows altered morphology of peroxisomes and early endosomes in ShOpa1 cells. E”’ Quantification shows increased fragmented cis-Golgi in ShOpa1 transduced cells. 100-110 cells were measured for each of the organelle markers. Statistical significance was measured with a two-sided chi-square test. Source data are provided as a Source Data file.
We determined the impact of OPA1 deficiency on the morphology of other organelles considering mitochondrial interactions with many organelles. Unlike peroxisomes (PMP70 positive) in control cells with lengths between 0.1 to 1 µm, the OPA1-KD cells contained hyper-tubulated and hyper-branched peroxisomes ranging in size from 0.1 to 6 µm (Fig. 1E, E’). OPA1-KD NP cells also exhibited enlarged EEA1-positive early endosomes ranging in size from 0.5 - 2 µm (Fig. 1E, E”). Additionally, there was a pronounced fragmentation of cis-Golgi (GM130 positive) in OPA1-KD cells (Fig. 1E, E”’, Fig. S2A, B). In contrast, TGN46-stained trans-Golgi, as well as RAB7-stained late endosomes, and LAMP1-stained lysosomes were unaffected (Fig. S2C). These findings suggest that OPA1 is essential for sustaining the morphology of organelles that include peroxisomes, early endosomes, and the cis-Golgi in NP cells.
OPA1 deletion disrupts NP cell autophagy
We noted that OPA1-KD NP cells possess fewer LC3B-positive autophagosomes (Fig. 2A, A’) and a significantly lower LC3II level, suggesting either an overall decrease in the autophagy or autophagy happening at a much faster rate (Fig. 2B, B’). Since BNIP3 activation and mitochondrial translocation are required for hypoxia-induced mitophagy in NP cells6,19, we investigated whether BNIP3 mitochondrial localization is affected in OPA1-KD cells. Indeed, we observed BNIP3 sequestration in the nucleus of OPA1-KD cells, without alteration in NIX localization (Fig. S1B, B’) and protein levels (Fig. S1C, C’). The levels of autophagy-related proteins, p62, BECLIN1, and the ATG12-ATG5 complex remained unaffected, whereas those of the apoptotic transcription factor CHOP decreased suggesting OPA1-KD cells did not activate apoptosis (Fig. S1C, C’). Moreover, levels of ubiquitinated proteins (Fig. S1D, D’), PARK2, and its substrate phosphoubiquitin, were unaffected by OPA1-knockdown (Fig. 2B, B’, C, C’)20. To ascertain whether OPA1 deletion impacts autophagy initiation and/or rate of degradation, we treated cells with Bafilomycin A1 (BafA1), an inhibitor of autophagic degradation. This short-term treatment caused robust accumulation of LC3II in OPA1-KD cells, although the accumulated levels did not exceed levels seen in BafA1 treated ShC transduced cells (Fig. 2D, D’), suggesting that a) decreased LC3II under basal conditions were not due to increased autophagic flux, has it been the case, BafA1 treatment would have resulted in significantly higher levels compared to control b) Opa1-knockdown did not inhibit the acidification of autolysosomes or autophagosome-lysosome fusion and thus autophagic degradation of the cargo, implying that the Opa1-knockdown caused an overall decreased in autophagy. Moreover, a slightly less accumulation of LC3II in BafA1 treated control cells also suggests autophagic flux at a steady state, wherein fewer faulty organelles and proteins require degradation. These data indicate dysfunction in the initiation of the autophagic pathway in OPA1-KD NP cells.
A Immunofluorescence staining of LC3B; OPA1-deficient NP cells shows a stark absence of LC3B puncta. Scale bar: Top Row-15, Bottom Row - 2 μm. Staining was performed 3 times independently. B, B’ Representative Western blot showing data from 3 experiments and densitometric analysis of autophagy/mitophagy pathway markers LC3II, p62, LAMP1, and PARK2 in NP cells shOpa1. C, C’ Representative Western blot and densitometry of phospho-ubiquitin after OPA1 deletion. n = 6 independent experiments. D, D’ Western blot analysis of LC3B in ShCtrl and ShOpa1 transduced cells cultured under hypoxia with or without bafilomycin A1 (n = 4 experiments). Quantitative data represented as box and whisker plots showing all data points with median and interquartile range and maximum and minimum values. Statistical significance was performed using One-way ANOVA or Kruskal Wallis test with Sidaks’s multiple comparisons test as appropriate. Source data are provided as a Source Data file.
OPA1-deficient NP cells show dysregulated bioenergetics
We measured ATP production rates from glycolysis and oxidative metabolism in OPA1-KD cells by integrating ECAR and OCR measurements (Fig. 3A, B)5,21. Under basal conditions, OCR was significantly higher, while ECAR remained unchanged, suggesting OPA1-KD NP cells consumed more oxygen (Fig. 3A–C). Following the addition of glucose, OPA1-KD cells showed similar ECAR profiles to the control group, but the OCR remained significantly higher and similar to the levels observed under basal conditions (Fig. 3C). Further determination of glycolytic ATP generation showed a significant increase in OPA1-KD cells; however, oxidative ATP production was comparable between groups despite higher oxygen consumption by OPA1-KD cells. These data suggested uncoupling of OXPHOS in OPA1-KD NP cells, in conjunction with the aberrant cristae morphology. We also investigated the effect of OPA1-deficiency on the glycolytic capacity of NP cells5,22. The OPA1-KD cells showed no difference in EACR but demonstrated a prominent increase in OCR (Fig. 3E, F). However, the basal glycolytic rate, maximum glycolytic capacity, ATP demand-limited rate, and glycolytic reserve computed from H + /lactate production were unaffected (Fig. 3G). As a general observation, since NP cells rely primarily on glycolysis, when glucose was available, both ShC and OPA1-KD NP cells were noted to “switch” to more glycolytic metabolism, lowering oxidative metabolism and thereby decreasing oxygen consumption (Fig. 3B, F). Overall, these results suggest that OPA1-deficiency dysregulates the bioenergetics of NP cells.
A, B ECAR and oxygen OCR traces from NP cells transduced with ShCtrl and ShOpa1 in the absence of exogenous glucose and after sequential addition of 10 mM glucose followed by oligomycin and rotenone plus myxothiazol. C Mitochodrial OCR measurement between ShCtrl and ShOpa1 under basal, glucose, and oligomycin addition (D) ATP production rate from glycolytic and oxidative metabolism calculated from traces shown in (A) and (B). E, F ECAR and OCR traces in the absence of exogenous glucose and after sequential addition of 10 mM glucose, followed by rotenone plus myxothiazol, and finally monensin plus FCCP. G Proton production rate (PPR) of NP cells calculated from traces shown in (D) and (E). Data represent 3-4 independent experiments, each with four replicates/group. Data is shown mean ± SEM. t-test or Mann–Whitney two-tailed test or one-way ANOVA was used as appropriate (C, D, G). Source data are provided as a Source Data file.
OPA1-deficient NP cells evidence altered glucose and glutamine metabolism
To understand the metabolic implications of OPA1 deficiency, we performed metabolic profiling of OPA1-KD NP cells. A total of 416 metabolites (104 up and 60 down) (CV < 30%) were used for multivariate analysis. An unsupervised PCA and supervised PLS-DA analysis showed a separation between the OPA1-KD and Ctrl samples (Fig. 4A–C). In terms of altered metabolites, fructose-1,6-phosphate decreased, whereas pentose phosphate pathway metabolites 6-phosphogluconic acid, sedoheptulose-7-phosphate and TCA metabolites citrate, malate, and acylcarnitine increased. Similarly, the one-carbon metabolites homocysteine, s-adenosyl homocysteine and s-adenosyl methionine and 5’ hydroxylysine were elevated in OPA1-KD cells (Fig. S3A–J) along with NAD, NADP, ATP, and ADP, with a decrease in AMP, ADP/ATP and AMP/ATP (Fig S3K–Q). Metabolic pathway analysis of upregulated metabolites showed that sugar metabolism, amino acid metabolic pathways, lipid metabolic pathways, betaine, spermidine, and spermine biosynthesis, along with taurine and hypotaurine, pyrimidine, and urea cycle (Fig. 4D). Moreover, the metabolite set enrichment analysis (MSEA) showed highly enriched metabolites from 50 out of the 97 metabolic pathways in OPA1-KD cells (Fig. 4D’). In contrast, OPA1-KD negatively impacted phosphatidylethanolamine, catecholamine biosynthesis, β-oxidation of short-chain and very long-chain saturated fatty acid, phosphatidylcholine biosynthesis, propanoate metabolism, glycolysis, fructose, and mannose degradation (Fig. 4E) and MESA showed a significant selective enrichment of purine and thiamine pathway-related metabolites (Fig. 4E’). The untargeted metabolic profiling suggested a broader dysregulation of metabolism in OPA1-KD cells.
A Unsupervised PCA and B Supervised PLS-DA model of widely targeted small metabolites from NP cells transduced with ShCtrl and ShOpa1. n = 6 independent experiments C Heat map of normalized concentration of metabolites differentially present between ShCtrl and ShOpa1, FDR ≤ 0.05%. D, E Metabolic pathway analysis (MetPA) showing pathway impact and p-value of significantly upregulated and downregulated metabolites. D’, E’ Metabolite set enrichment analysis (MSEA) showing enrichment ratio with p-value of upregulated and downregulated metabolites (FDR < 0.05) from ShOpa1 vs ShCtrl transduced cells. Statistical significance was tested using a t-test or a Mann-Whitney two-tailed test as appropriate. Source data are provided as a Source Data file.
Furthermore, OPA1-KD NP cells were cultured with either [1,2]-13C-glucose or U13C-glutamine (Fig. 5A). Based on the lactate and glutamate enrichment in the medium we calculated the glycolytic, pentose cycle, PDH, PC, PDH + PC, and PHD/PC flux, as previously described5. OPA1-knockdown did not affect glucose flux through glycolysis or the pentose cycle (Fig. 5B, C). However, PDH flux was significantly decreased, without altering flux through PC; combined PDH + PC flux or the PDH/PC also remained unchanged (Fig. 5D–G). Analysis of extracted metabolites from the cell pellet showed a modest increase in sigma mean (∑ mn = 1 *M1 + 2*M2 + 3*M3, etc.) of alanine and a decrease in serine with minimal changes in lactate, citrate, and succinate. A small decrease in M1 glutamate with minimal change in palmitate as well as enhanced enrichment in stearic acid was also noted (Fig. 5H–O). When U13C-glutamine utilization was measured, OPA1-KD cells evidenced a reduction in TCA cycle intermediates citrate, succinate, fumarate, malate, and aspartate, without changes in lactate (Fig. 5P–U). Both hypoxia and ETC inhibition are shown to increase the levels of succinate23. Interestingly, however, our 13C-MFA data showed increased succinate ∑ mn compared to fumarate in both Ctrl and OPA1-KD cells, suggesting impairment of the TCA cycle. Furthermore, when we assessed succinate oxidation and fumarate reduction, no change in the oxidation, with a downward trend in fumarate reduction was noted (Fig. 5V, W). These studies lent support to the notion that OPA1 is required for optimal NP cell metabolism.
A Summation of flux results through glycolysis, pentose, and TCA cycle using [1,2]-13C-glucose and U13C-glutamine. B–G [1,2]-13C-glucose enrichment in the culture media from NP cells transduced with ShCtrl and ShOpa1 and cultured under hypoxia for 24 h. B Glycolysis flux as measured by enrichment of M2 lactate m/z 91, C Pentose cycle flux as measured by lactate (M1/M2)/ (3 + M1/M2) D PDH flux as measured glutamate m/z 103, E PC flux measured as glutamate m/z 104 F PC + PDH flux measured as glutamate m/z 131, G PDH/PC flux as measured by glutamate m/z 103-M1/104-M2. H–O [1,2]-13C-glucose tracing measured from the cell pellets into metabolites (H) alanine m/z 260, I serine m/z 390, J lactate m/z 261, K citrate m/z 591, L succinate m/z 289, M glutamate m/z 432, N palmitate (C16:0) m/z 313, O stearic acid (C18:0) m/z 341. P–W U13C-glutamine tracing measured from NP cells transduced with ShCtrl and ShOpa1 and cultured under hypoxia for 24 h P lactate m/z 261, Q citrate m/z 591, R succinate m/z 289, S fumarate m/z 287, T malate m/z 419, U aspartate m/z 418, V succinate oxidation and W fumarate reduction. Quantitative data are represented as box and whisker plots showing all data points with median and interquartile range and maximum and minimum values. Data points with negative values (undetectable ∑ mn) are not included. Statistical significance was computed using t-test or Mann-Whitney two-tailed test as appropriate. Source data are provided as a Source Data file.
Conditional deletion of Opa1 in IVD accelerates age-associated degeneration
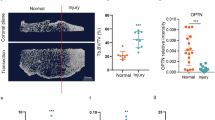
To gain a better understanding of how OPA1 impacts spinal health, we generated Opa1 conditional knockout mice AcanCreERT2Opa1fl/fl (Opa1cKO) and Opa1fl/fl (WT) mice (Fig. 6A, B)24. In adult mice, the Acan-CreERT2 driver is highly effective in targeting all three compartments of the disc, as well as the articular and growth plate cartilages25. The successful deletion of OPA1 in the disc was confirmed by mRNA and protein evaluation of the NP and AF tissues (Fig. 6C, D, D’). Quantitative histopathological analysis of disc morphology using the Modified Thompson grading on 7, and 12-month-old Opa1cKO mice (Fig. S4) indicated no noticeable disc degeneration in the lumbar (Fig. S4. A–A”, B–B”) or caudal regions of the spine (Fig. S4. C-C”, D-D”). At 12 months, however, caudal discs of Opa1cKO mice lost the notochordal hallmark of large cytoplasmic vacuoles, showing changes in NP cell morphology and AF hyperplasia (Fig. S4D”’). When Opa1cKO mice were evaluated at 20 months, a prominent NP and AF degenerative phenotype in caudal discs was evident (Fig. 6E). The distribution of grading scores showed a significantly higher proportion of Opa1cKO discs had NP and AF scores of 3 or 4, indicating severe degeneration (Fig. 6E’, E”). At 20 months, the degenerative phenotype was characterized by a diminished SafraninO stained NP matrix, a significant loss of NP cells with the remainder of cells acquiring a hypertrophic chondrocyte-like morphology with irregularities in AF lamellar organization (Fig. 6E). There was also a clear loss of NP-AF demarcation in Opa1cKO mice (Fig. 6E). Those discs that retained notochordal NP cells displayed morphological changes such as loss of cytosolic vacuoles (Fig. 6E”’). Additionally, Opa1cKO mice exhibited AF hyperplasia reflected in increased tissue area (Fig. 6F, F’).
A Schematic showing tamoxifen induced AcanCreERT2 mediated deletion of exons 10-13 of Opa1 to generate Opa1 null allele. B Tamoxifen treatment and analysis timeline of WT (Opa1fl/fl) and Opa1AcanERT2 (Opa1cKO) mice. C RT-PCR analysis shows Opa1 deletion in NP and AF tissues of Opa1cKO mice (n = 3 mice/genotype). D, D’ Western blotting and IHC of OPA1 from NP and AF tissues (n = 2 mice/genotype). E Representative Safranin O/Fast Green/Hematoxylin staining of 20-month-old WT and Opa1cKO caudal disc sections. Scale bar: Top row-500 μm, Bottom rows - 100 μm. E’, E” Modified Thompson Scores of NP and AF compartment of WT and Opa1cKO caudal discs. E”’ Distribution graph showing NP cell phenotype at 20-months. n = 9 WT (4 F, 5 M), 11 Opa1cKO (6 F, 5 M) mice; 3 caudal discs/animal, 27-33 discs/genotype. F, F’ AF tissue hyperplasia in Opa1cKO caudal discs, dotted lines demarcate AF compartment. scale bar = 100 μm. G Representative Picrosirius red stained polarized light images of caudal discs from 20-month-old mice showing NP fibrosis. Scale bar = 500 μm (G’) The distribution of discs based on %NP area occupied by collagen fibers. G” Thin, intermediate, and thick AF collagen fibril fraction. n = 9-11 mice/genotype; 3 caudal discs/animal, 27-33 discs/genotype. H, H’ Representative images of LC3B immunohistology and quantification of LC3B-stained autophagosomes. Scale bar = 10 μm, insert panels 2 and 4 = 20 μm, n = 6-9 mice/genotype: 1-3 caudal discs/animal, 12-24 discs/genotype. Significance for E’, E”’, G’ was determined using a two-sided chi-square test. The significance of the fiber percentage of AF collagen area (G”) was determined using Kruskal-Wallis with Dunn’s test. Quantitative Data represents violin (E”) or box and whiskers (F’, G”, H’) plot showing all data points with median and interquartile range and maximum and minimum values. Significance was determined using an unpaired t-test (F’) or Mann-Whitney two-tailed test (H’). Source data are provided as a Source Data file.
Picrosirius red staining coupled with polarized light imaging was used to ascertain alterations in collagen organization (Fig. 6G). At 20 months, Opa1cKO mice showed the presence of collagen fibers in the NP and exhibited a higher proportion of NP fibrous tissue area than a few discs in WT mice with NP fibrosis (Fig. 6G’). In the AF, Opa1cKO mice showed a higher proportion of thin collagen fibers suggesting increased turnover (Fig. 6G”). In agreement with OPA1-KD cells, we observed decreased LC3B-positive autophagosomes in Opa1cKO mice (Fig. 6H, H’). Notably, lumbar discs of 20-month-old mice showed a milder phenotype compared to caudal discs; there was a higher proportion of lumbar discs with NP and AF compartments scoring grade 4 but when scores of all discs were averaged it did not reach statistical significance (Fig. S5A, A’, A”). Similar to caudal discs, the NP of lumbar discs in 20-month-old Opa1cKO mice displayed a higher fibrosis underscoring degeneration (Fig. S5B, B’).
OPA1 deletion affects NP cell phenotype and alters the disc matrix composition
We assessed the NP cell phenotype in 20-month-old mice by measuring the abundance of the phenotypic markers, carbonic anhydrase (CA3), glucose transporter I (GLUT1), and keratin 19 (KRT19). Strikingly, expression of these markers was lost by some cells that persisted in the NP compartment of Opa1cKO mice, suggesting the native notochordal cells underwent a phenotypic switch (Fig. 7A, B, Fig. S6). When stained for COLX, a marker of hypertrophic chondrocytes, we noted a considerably higher signal (Fig. 7C). We stained for MMP13, a marker of chondrocyte hypertrophy and disc degeneration, and found that both NP cells and the degenerated disc expressed MMP13. (Fig. S6). We also noted lower AF abundance of COLI (Fig. 7D, D’) and COMP, an important non-collagenous matrix component (Fig. 7D, D”) in Opa1cKO mice. While ACAN staining revealed no differences (Fig. 7E, E’), the cells in the NP and AF of Opa1cKO showed a robust increase in the pericellular staining of an ACAN neoepitope formed by ADAMTS4/5 dependent cleavage (Fig. 7E, E”). Overall, these findings suggest that OPA1 is required for maintaining the disc cell phenotype and the ECM.
Immunohistological staining of 20-month WT and Opa1cKO caudal discs for A carbonic anhydrase 3 (CA3); B glucose transporter 1 (GLUT1) and C collagen 10 (COLX). Scale bar = 100 and 50 μm. Immunohistological staining for D, D’ collagen 1 (COL1); D, D” cartilage oligomeric matrix protein (COMP), E, E’ aggrecan (ACAN); E, E” aggrecan G1 neoepitope (ARGxx). Scale bar = 100 μm. (n = 9 WT (4 F, 5 M), 11 Opa1cKO (6 F, 5 M) mice/genotype, 1-3 discs/mouse, 13-14 discs/genotype/marker). Quantitative measurements are shown as Box and whisker plots showing all data points with median and interquartile range and maximum and minimum values. Significance was determined using an unpaired t-test (D”) or Mann-Whitney two-tailed test (D’, E’, E”, E”’), as appropriate.
Opa1cKO mice evidence alterations in vertebral bone health
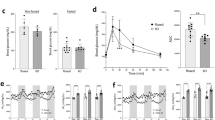
We performed μCT on the L3-6 vertebrae and found expected trabecular bone loss with aging in WT; however, Opa1cKO mice exhibited an accelerated reduction in vertebral trabecular bone architecture at 7- and 12-months (Fig. 8A). Thus, bone volume/trabecular volume (BV/TV), bone mineral density (BMD), trabecular thickness (Tb.Th), and trabecular number (Tb.N) was decreased and these changes persisted at 20 months implying early osteopenia in Opa1cKO (Fig. 8B–E). As expected, trabecular separation was also greater in 7-month-old Opa1cKO mice (Fig. 8F) and the structural model index (SMI), a parameter that is linked to bone strength and fracture risk, was considerably greater at 7 and 12 months (Fig. 8G). Interestingly, unlike early changes in the trabecular bone of Opa1cKO, changes in cortical bone structure were evident predominantly at 20-months, with an increase in mean total cross-sectional bone area (B.Ar.), tissue area (T.Ar.) without any alteration in the cross-sectional thickness (Cs.Th.) and tissue mineral density (TMD) (Fig. 8H–L). We also noted an increased vertebral length and disc height index (DHI) in Opa1cKO mice at 7- and 12-months (Fig. 8M–P), which are correlated with disc pathology26. These findings suggest that OPA1 plays an important role in the control of vertebral health.
A The representative 3D reconstruction of hemi-section and trabecular lumbar vertebral bone from 7-month, 12-month, and 20-month-old WT and Opa1cKO mice. Scale bars 500 μm. Vertebral trabecular bone parameters B Percent bone volume/ tissue volume (BV/TV) (%), C Bone mineral density (BMD) (g/cm3), D trabecular thickness (Tb.Th) (mm), E trabecular number (Tb.N) (1/mm), F trabecular separation (Tb.Sp) (mm), G structural model index (SMI) of WT and Opa1cKO mice. H Representative 3D reconstruction of transverse section through vertebrae of WT and Opa1cKO mice showing cortical shell geometry and corresponding quantitative parameters I mean total cross-sectional thickness bone area (B.Ar) (mm2), J mean total cross-sectional tissue area (T.Ar) (mm2), K cross-sectional thickness (Cs.Th) (mm), L Tissue mineral density (TMD) (g/cm3). Scale bars 500 μm. M Representative microCT reconstructions of hemi-sections through vertebrae of WT and Opa1cKO mice to determine N Vertebral length (VL) (mm), O disc height (DH) (mm), and P disc height index (DHI). Scale bars 1 mm. Quantitative data are shown as box and whisker plots showing all data points with median and interquartile range and maximum and minimum values. 7-month n = 10 WT (3 F, 7 M), 8 Opa1cKO (3 F, 5 M); 12-month n = 10 WT (6 F, 4 M), 10 Opa1cKO (4 F, 6 M); 20-month n = 9 WT (4 F, 5 M), 11 Opa1cKO (6 F, 5 M), 4 vertebrae and 3 discs/mouse were analyzed. Significance for B-G and I-P was determined using a t-test or a Mann-Whitney two-tailed test as appropriate. Source data are provided as a Source Data file.
NP tissue transcriptome of Opa1cKO mice highlights metabolic dysregulation and shares similarities to transcriptomes of degenerated human NP tissues
To delineate whether metabolic changes underline prominent disc degeneration seen in 20-month-old Opa1cKO mice, we performed RNA-sequencing at 12 months when structural degeneration is not evident. The PCA demonstrated distinct grouping between Opa1cKO and WT samples (Fig. 9A) and volcano plot and heat map were used to determine the distribution and hierarchical clustering of DEGs (FDR < 0.05) (Fig. 9B, C). CompBio tool was used for pathway-level analysis27 and to identify DEG enrichment into up- and down-regulated themes and related concepts (Fig. S7A, B) represented as ball and stick model (Fig. 9D). Down-regulated themes included biological processes associated with metabolism, autophagy/mitophagy, ECM-muscle, ubiquitination, posttranslational modification, cell signaling, endocytosis, and transporters (Fig. 9E–K; Fig. S8A–P). Upregulated themes comprised a biological process related to cytoskeletal reorganization, ECM/cartilage/bone biology, inflammatory mediators, and apoptosis (Fig. S9A–K). We used MitoPathways 3.028 to perform gene set enrichment analysis (GSEA) and found downregulation of genes related to ETC complex and branched-chain amino acid metabolism in Opa1cKO (Fig. 9L–N’; Fig. S10). When signature GSEA was used, we found 29 signatures set in upregulated DEGs and 6 in down regulated DEGs were significant with a recommended FDR < 0.25 (Fig. S11A-L’; S12A, B).
A WT and Opa1cKO transcriptomic clustering profiles using Principal Component Analysis (PCA). B, C Volcano plot and Hierarchical clustering of differentially expressed genes (DEGs) with FDR < 0.05%. D The CompBio biological map generated from downregulated DEGs FDR < 0.05%. The size of the sphere corresponds to its enrichment score, while the thickness of the lines connecting themes represents the number of genes shared between them. E–K Downregulated themes and related DEGs associated with hypoxia, glycolysis, lysophosphatidic acid (LPA), one-carbon metabolism, procollagen dioxygenase, and autophagy/mitophagy in Opa1cKO mice were shown with CompBio entity score. L–N’ The gene set enrichment analysis (GSEA) of mitochondrial pathways (MitoPathways 3.0) of Opa1cKO downregulated DEGs showing ETC complex IV subunits, OXPHOS and branched chain amino acids metabolism. MitoPathways3.0 was employed to calculate gene set enrichment using the log2 values for pre-ranking the genes and gene set size filters (min=15 and max = 500). A negative enrichment score (ES) indicates gene set enrichment at the bottom of the ranked list. Source data are provided as a Source Data file.
Next, we used the CompBio tool to determine whether the gene profile of Opa1cKO captures the biological signatures identified in human disc degeneration (GSE70362)11,29. As reported11, human samples were grouped into seven clusters, including a cluster consisting of only healthy samples, and degenerative clusters 1-5 were compared with Opa1cKO. For Opa1cKO upregulated DEGs, human clusters 1, 2, 3, and 5 showed significant global similarity and thematic level overlap (Fig. S13). Remarkably, human cluster 1 showed prominent overlap for the individual concepts of articular cartilage development, vertebra cartilage element, CTGF, FGF, SMAD/TGFβ, protein-tyrosine phosphatase signaling, and IL-11 binding. For downregulated DEGs, clusters 1, 3, and 5 showed a substantial global resemblance, while at the thematic level, clusters 1 and 5 exhibited an overall 15% and 12% overlap, respectively. (Fig. S14). These three clusters also showed between 9-11% overlap for the metabolic theme. Since NP tissue in mice and humans is reported to possess cellular heterogeneity, we ascertained if Opa1-deletion in aggrecan-expressing NP cells will affect the constituent cell subpopulations30,31,32,33. The analysis of the deposited scRNAseq data30,31,32,33, mouse NP cells robustly expressed notochordal Bonafide markers Brachyury/T, Keratin 18, and Keratin 19 (T, Krt18, and Krt19, respectively) at both P30 and 8 weeks old discs (Fig. S15). Moreover, in line with earlier findings, a robust and widespread Aggrecan expression in mouse, rat, and human NP cells (Fig. S16). Furthermore, regardless of the species, aggrecan-expressing NP cells also showed ubiquitous Opa1 expression (Fig. S16) suggesting that all the constituent subpopulations within mouse NP will exhibit Opa1 deletion and contribute to the observed phenotype. Overall, our findings imply that metabolic imbalance caused by OPA1 deletion is one of the key causes contributing to disc deterioration in aging mice and resembles underlying biological processes in a subset of human patient NP tissue samples.
Opa1cKO mice show increased severity of age-associated OA
Since the AcanCreERT2 allele efficiently targets articular cartilage, we investigated if OPA1 deletion has deleterious effects on knee joint health in mice. μCT imaging at 12 months did not evidence any changes in the knee joints; however, at 20-months Opa1cKO mice showed significant osteophyte formation in the joints (Fig. S17A, Fig. S17A). At 12-months, tibial subchondral BV/TV, Tb.Th, Tb.Sp, and subchondral bone plate thickness (SCBP) were comparable between genotypes (Fig. S17B–E). However, there was a significant increase in SCBP in the lateral compartment of the 20-month-old Opa1cKO mice (Fig. S18B–E); To study cartilage structure, we stained knee joints sectioned in the mid-coronal plane with H&E and toluidine blue34. There were no morphological and histomorphometric changes between genotypes at 12 months (Fig. S17F-M’), however, at 20 months, Opa1cKO mice showed severe OA with loss of articular cartilage on the lateral tibial plateaus and femoral condyle, but only minor degradation of articular cartilage in the medial compartment (Fig. S18F). Furthermore, H&E staining revealed osteophyte formation in the medial compartment of 20-month-old WT mice, and osteophyte formation in both the lateral and medial compartments of Opa1cKO animals (Fig. S18G). When these structural changes were quantified, Opa1cKO mice exhibited a significant increase in articular cartilage score (ACS) in the lateral knee compartment at 20 months. There were no deviations in toluidine blue scores and osteophyte scores between knee compartments or genotypes (Fig. S18H–J), but the cumulative osteophyte score in Opa1cKO mice was significantly higher (Fig. S18K).
Representative 4X images of the lateral and medial compartments of WT and Opa1cKO 20-month-old mouse joints demonstrate the profound effect of OPA1 loss on multiple tissues within the joint (Fig. 10A upper panels) which includes significant cartilage loss on the lateral side of Opa1cKO mice. Furthermore, 10X images of H&E staining showed synovial hyperplasia/ossification in the lateral and medial knee compartments of 20-month-old Opa1cKO mice (Fig. 10A lower panels), and we quantified changes with histomorphometric analysis. In the lateral compartment of 20-month Opa1cKO mice, there was a significant decrease in the area and thickness of articular cartilage and calcified cartilage (Fig. 10B, B’, C, C’). SCBP area and thickness were significantly increased in Opa1cKO mice when compared to controls, indicating enhanced subchondral bone sclerosis in OPA1 loss mice, a finding that aligns with our µCT data (Fig. 10D, D’). We also noted a considerable increase in the synovial hyperplasia/ossification score in the lateral joint compartment of 20-month-old Opa1cKO mice (Fig. 10E). It is interesting to note that the profound OA phenotype in Opa1cKO mice was primarily observed in the lateral compartment as opposed to the medial compartment which is typically more affected. These findings demonstrate that OPA1 loss in knee cartilage enhances the severity of age-associated OA in mice.
A Representative H&E images (top row 4X; scale bar - 250 μm; bottom row 10X; scale bar − 100 μm) of WT and Opa1cKO mouse joint sections showing medial and lateral compartments of the same joint. Profound synovial hyperplasia and/or ossification is noted in Opa1cKO mice in both the medial and lateral compartments (yellow asterisks), when compared to WT mice B–E Histomorphometric analyses of lateral and medial joint compartments conducted on midcoronal sections of WT and Opa1cKO mouse limbs. B, B’ articular cartilage (Art. cart) area and thickness, C, C’ calcified cartilage (Calc. cart) area and thickness, and D, D’ subchondral bone plate (SCBP) area and thickness E quantitative analysis of synovial hyperplasia. n = 10 WT (4 F, 6 M), n = 11 Opa1cKO (5 F, 6 M). Quantitative data is represented as box and whiskers B–D’ or violin E plots showing all data points with median and interquartile range and maximum and minimum values. Significance for B-E was determined using a t-test or a Mann-Whitney two-tailed test as appropriate. F Schematic showing the in vitro consequences of OPA1-deficiency on mitochondrial and organelle morphology and metabolic functions of NP cells and in vivo phenotypic manifestations of OPA-deletion on the spinal column and knee joint in Opa1cKO mice. Source data are provided as a Source Data file.
Discussion
This study for the first time establishes a causal relationship between sustaining mitochondrial dynamics through OPA1 and maintenance of the spine and knee joint health during aging in mice. The reliance on mitochondrial function was notable since hypoxic chondrocytes and in particular NP cells are primarily glycolytic5,6. Relevant to this finding, we have previously shown the existence of mitochondrial networks in NP cells and that mitochondrial activity is governed by the HIF-1α-BNIP3 axis controlling mitophagic flux6. We also demonstrated that BNIP3 deletion in NP cells caused mitochondrial dysfunction and early disc degeneration in mice5. Herein, we show that, OPA1 controls the morphology of mitochondria and cristae as well as organelles including peroxisomes, early endosomes, and cis-Golgi, and that OPA1-loss results in dysregulated autophagy in NP cells. Further, we demonstrate that Opa1AcanCreERT2 mice evidence accelerated age-dependent IVD degeneration, vertebral osteopenia, and severe OA of knee joints. Our findings underscore that dysregulation of mitochondrial dynamics affects the metabolism, organelle integrity, and the autophagic/mitophagic pathway causing disc and cartilage degeneration in mice.
OPA1 is required for IMM fusion, and maintenance of cristae shape, and its mutations are linked to pathologies including vision impairment35, developmental delay, muscle-related disorders, peripheral neuropathy, and cardiomyopathy36. Concerning axial skeleton, we have previously shown that changes in mitochondrial morphology impact NP cell metabolism and mitophagy5,6. Herein we observed that OPA1deletion resulted in a reduction in mitochondrial size but an increase in numbers, a change unlikely due to mitochondrial biogenesis since there was no difference in the mtDNA content18,37. OPA1 is required for IMM invagination and cristae development which is reflected in OPA1-KD NP cells lacking cristae or detached cristae. Mitochondria also interact with other organelles to maintain cellular homeostasis38, from this perspective, the impact on peroxisomes, endosomes, and cis-Golgi morphology suggested that OPA1 is critical for preserving multi-organelle morphology likely through governing their interactions with mitochondria. To the best of our knowledge, this is the first report on the involvement of OPA1 in sustaining the morphology of these organelles.
When mitochondrial quality control was assessed, a significant decrease in key autophagy-related protein LC3-II in OPA1-KD cells was noted without impairment in autophagic flux, suggesting impaired autophagosome formation. LC3-II targets ubiquitin-positive cargo to sequester them into developing autophagosomes39. Notably, these autophagy proteins are subjected to regulatory post-translational phosphorylation by mTORC1 (negative) and ULK1 and AMPK (positive) whereby AMPK promotes autophagy by phosphorylating ULK1 when the ADP/ATP or AMP/ATP ratio is elevated. Indeed, our metabolic profiling indicated that decreased AMP/ATP and ADP/ATP ratios would be associated with decreased phosphorylation, and, as a consequence, reduced autophagy. In contrast to previous studies demonstrating enhanced mitophagy in OPA1-defective cells18,40,41, we noted that OPA1-KD not only inhibited selective autophagy (mitophagy) but also macroautophagy, an event that influenced the morphology of multiple organelles, including the ER, endosomes, and Golgi42,43,44. For example, enlarged early endosomes and fragmented Golgi, together with autophagosome accumulation, is implicated in neurodegenerative diseases such as Down syndrome and Parkinson’s45,46. Importantly, changes in these organelles underscore the defects in autophagy and disruptions in endocytic and secretory pathways in OPA1-KD NP cells. Together, our data supports the hypothesis that OPA1 regulates organelle morphology and autophagic/endocytic/secretory pathways in NP cells. One important outcome of these findings is that autophagy and mitochondrial dysfunction are associated with aging, disc degeneration, and the pathogenesis of OA2,47,48.
It is known that in many cell types, which primarily rely on oxidative ATP generation, structural changes in mitochondria profoundly alter energy metabolism. Since NP cells are predominantly glycolytic, it was important to determine whether alterations in mitochondrial shape and cristae morphology influenced NP cell bioenergetics. We found that while fragmented mitochondria with aberrant cristae morphology consumed more oxygen, it did not manifest in increased oxidative ATP production rates. The number of ATP molecules generated for each dioxygen molecule consumed might vary depending on the mitochondrial efficacy49,50, suggesting mitochondrial dysfunction in the OPA1-KD NP cells. Our findings also differed from prior work showing lower OCR and oxidative ATP generation in OPA1-deficient cells51,52 but agreed with the increase in glycolytic ATP production rate noted in OPA1-deficient neutrophils53. Overall, the results of the current study underscored the observation that OPA1 deficiency influences the energy metabolism of NP cells.
Metabolic pathways most positively impacted in OPA-KD cells included spermidine and spermine biosynthesis, betaine metabolism along with amino acid metabolism, taurine, and hypotaurine synthesis, pyrimidine, and urea cycle metabolism. This contrasted a study in MEFs, where OPA1 deficiency decreased metabolites associated with spermidine and spermine and taurine and hypotaurine pathways54. Notably, spermidine and taurine levels are shown to decline with age, and mitochondrial dysfunction is one of the primary contributors to their deficiency55,56. Supplementation of spermidine and taurine has been proven to prolong longevity in mice via boosting autophagy/mitophagy, mitochondrial biogenesis, and mitochondrial respiration55,56. Furthermore, lower fructose-1,6-phosphate suggested a reduction in glycolysis, whereas higher malate and enhanced malate-aspartate shuttle activity, as well as increased pentose phosphate pathway metabolites sedoheptulose-7-phosphate, 6-phosphogluconic acid, and NADP, indicated oxidative stress in OPA1-KD cells. Remarkably, acetylcarnitine level was increased in OPA1-KD cells, which links mitochondrial metabolism to histone acetylation and lipogenesis57. In agreement with a report on OPA1 regulation of lipid metabolism, we observed reduced fatty acid metabolism58. OPA1-deficiency positively affected one-carbon metabolism in OPA1-KD cells, further supported by elevated pyrimidine metabolism, polyamines, and amino acid metabolism. In summary, the metabolite profiling studies revealed an underlying metabolic dysregulation in NP cells with OPA1 deficiency.
13C isotope labeling experiments utilizing two stable isotope tracers [1,2]-13C-glucose and U13C-glutamine shed further insights into the metabolic dysregulation of OPA1-KD NP cells. The use of [1,2]-13C-glucose MFA revealed a reduction in PDH flux despite an increase in alanine ∑ mn, indicating an elevated pyruvate pool. We also noted a decrease in intracellular M + 1 glutamate enrichment, an indirect measurement for alpha-ketoglutarate, suggesting pyruvate is prevented from entering the TCA cycle via the conventional pathway. Furthermore, since there was no change in PC flux, this supports the anaplerosis, i.e., pyruvate carboxylation replenishing TCA intermediates, allowing the Krebs cycle to continue, since the intermediates are essential for macromolecule synthesis as well as post-translational modifications, chromatin modification, and DNA methylation59. Interestingly, in solid hypoxic tumors, increased anaplerosis through PC is required for extracellular collagen production and aberrant fibrosis by tumor-associated fibroblasts, a phenotype we have noted in the discs of Opa1cKO mice60. Regarding glutamine utilization, the labeled glutamine entered the TCA via an anaplerotic reaction to alpha-ketoglutarate with M + 4 labeling of the forward TCA cycle intermediates succinate, fumarate, malate, and citrate. In contrast to glucose, glutamine labeling showed decreased TCA cycle intermediates. The Citrate M + 4 label signifies the oxidative (forward) TCA cycle while the M + 5 label is for the reductive (reverse) TCA cycle. We noted a substantial reduction in citrate M + 5 label ( ~ 2.5:1 M + 4:M + 5 citrate), indicating a modest reversal in TCA flux. Of interest, under physiological conditions, the majority of succinate is generated by oxidative or forward TCA cycle. Recent studies found that under hypoxia or when ETC was inhibited, a part of succinate was generated from fumarate, implying that fumarate reduction to succinate may act as a valve for surplus electrons from the ETC. Further, it was reported that fumarate was catalyzed explicitly by complex II, rather than passively collecting leaky electrons from the ETC23. Interestingly, our 13C MFA data showed increased succinate ∑ mn compared to fumarate, suggesting an oxidative TCA cycle in NP cells. Furthermore, we observed no changes in the oxidative process when we assessed succinate oxidation (fumarate M + 4/succinate M + 4) and fumarate reduction (succinate M + 3/fumarate M + 3). These findings suggested that NP cells do not experience metabolic stress under hypoxic conditions. Together, these studies suggested that mitochondrial morphology and cristae architecture are not only central to energy metabolism, but the integrity of these structures is critical for the proper functioning of mitochondria in NP cells. We acknowledge here that these mechanistic conclusions on how OPA1 contributes to NP cell metabolism are shown only in vitro and partially depend on a single shRNA clone.
Notably, we found that Opa1cKO mice evidenced enhanced caudal disc degeneration with aging. Previous studies have shown that caudal discs of mice are more prone to metabolic dysregulation and subsequent degeneration with aging61. Moreover, the caudal spine experiences relatively lower axial loading and different motions than the lumbar spine, it is therefore plausible that the unique interactions between genetics and environmental factors produce varying phenotypic outcomes across different spine regions62. Similarly, while the medial knee compartment is usually prone to cartilage degeneration63, the lateral compartment which experiences lower compressive loads exhibited more pronounced OA in Opa1cKO mice64. Consistent with previous studies that demonstrate exercise and/or loading in human and animal models partially restores the functionality of faulty mitochondria65,66, lack of substantial degeneration in lumbar discs and the medial compartment of the knee suggests a protective effect of loading on the joint tissues62,67. Concerning poor vertebral bone health noted in Opa1cKO animals, it has been shown that alterations in mitochondrial dynamics affect osteoblast and osteoclast activity, accelerating the development and progression of osteoporosis68,69.
In agreement with a previously reported analysis of human AF tissues that showed altered genes related to mitochondrial function during degeneration70, significant degenerative changes were noted in AF of Opa1cKO mice, underscoring the important contribution of mitochondrial activity to tissue health. Moreover, in degenerated AF tissues, ROS-related gene expression is dramatically altered, suggesting that mitochondrial dysfunction and ROS production promote AF degeneration70.
Notably, the NP transcriptomic signatures of Opa1cKO showed a significant impact of cell metabolism and thematic similarities with several human degenerative clusters highlighting commonalities between mouse and human pathology. It is important to emphasize that metabolic dysregulation is tightly correlated to larger transcriptome alterations observed in many organs, including skeleton71,72. Accordingly, we noted a significant downregulation in glycolysis, monocarboxylate transporter, 2-oxoglutarate-dependent dioxygenase activity along with LDL assembly, glutamate-glutamine uptake, and one-carbon metabolism. Furthermore, pathways related to mitochondrial iron transport activity, ETC complex, activation of NOXA, and mitochondrial translocation were significantly downregulated. The Mitopathways GESA analysis also showed suppression of ETC complex I and IV subunits, OXPHOS, and branched-chain amino acid metabolism in Opa1cKO mice. Notably, the downregulation of the mitophagy pathway also coincided with our in vitro results. Concerning upregulated DEGs, their enrichment into themes related to cytoskeletal organization, articular/vertebral cartilage, TGFβ, CTGF, and cell-matrix adhesion with varying degrees of similarity with human clusters supported hypertrophic chondrocyte-like differentiation noted in aged Opa1cKO mice. Likewise, an overall increase in themes related to proinflammatory signaling, including IL-11 binding, a cytokine recently associated with aging phenotypes in mice and humans, suggested a pro-fibrotic environment promoted by metabolic dysfunction73. Additionally, based on robust and ubiquitous Acan and Opa1 expression in mouse NP cells, AcanCreERT2 will efficiently target all constituent, mature, and progenitor-like subpopulations within NP, thereby ensuring their involvement in the phenotype development24,25,30,31. Furthermore, since Opa1 deletion affects not only NP and AF but also articular chondrocytes in mice, we hypothesize that all the NP subpopulations in humans, including those with fibrotic and chondrocytic properties, will be affected by OPA1-deficiency31,33. Overall, our data underscored that dysregulation of metabolism and mitochondrial dysfunction drives intervertebral disc degeneration in aged Opa1cKO mice.
Our studies suggest that the mouse disc, spinal column, and joint cartilage phenotypes are the outcome of metabolic dysregulation and cumulative degenerative processes driven by OPA1-deficiency. Moreover, in many soft tissues, age-related pathologies, and metabolic disorders due to the accumulation of faulty mitochondria have been demonstrated74,75. From this perspective, targeting and modifying the autophagic pathway and preserving mitochondrial function should be a major goal when designing therapeutics to treat diseases linked to degenerative musculoskeletal conditions.
Methods
Cell isolation and treatments
All rat cell isolation studies were carried out in conformity with the Institutional Animal Care and Use Committee (IACUC) of Thomas Jefferson University’s applicable rules and regulations. Primary NP cells were isolated from 10-week-old, male Sprague Dawley rats (Charles River) and cultured in antibiotic-supplemented low glucose (1 g/L) containing DMEM and 10% FBS76. Passage 2 NP cells were transduced with either lentiviral particles encoding ShCtrl or ShOpa1 (clone 1# TRCN0000091111, clone2 # TRCN0000348537) (Sigma, St. Louis, MO, USA) using 8 mg/ mL polybrene and after 3 days of transduction, cells were cultured in hypoxia workstation (Invivo2 400; Baker Ruskinn, Bridgend, UK) at 1% O2 for 24 h and on day 5 cells were harvested for protein extraction. Since both OPA1 shRNA clones showed similar effects on mitochondrial morphology and mitophagy, cells transduced with LV-ShOpa1 #1 (TRCN0000091111) were utilized for all metabolic investigations.
Immunocytochemistry
ShCtrl, ShOpa1 or DFP-treated NP cells were plated on glass coverslips and treated with 100 nM MitoTracker Red CMXRos (Thermo Fisher Scientific, Waltham, MA, USA; M7512) for 30 minutes after completion of the experimental treatments. Cells were then fixed with 4% PFA or ice-cold methanol for 15 minutes and permeabilized with 0.1% Triton X-100 for 10 minutes and blocked with 1% BSA for 1 h. Cells were incubated with anti-OPA1 (1:200, 612607) (BD Bioscience), anti-PMP70 (PA1-650), anti-GM130 (MA5-35107) (Fisher Scientific, Pittsburgh, PA, USA), anti-EEA1 (24115), anti-BNIP3 (3769S), anti-LC3A/B (12741), anti-LAMP1 (9091), anti-BNIP3L (12396) (Cell signaling, Denver, MA, USA), anti-TNG46 (ab16059), and anti-RAB7 (R8779) (EMD Millipore, Burlington, MA, USA), in blocking buffer at 1:100 to 200 at 4 oC overnight. After washing, cells were incubated with Alexa Fluor 488 and mounted with ProLong Gold Antifade Mountant with DAPI. The specificity of staining cells was confirmed using isotype mouse (7076P2) or rabbit (7074P2) IgG antibodies (Cell Signaling, Danvers, MA, USA). Stained cells were visualized using a Zeiss LSM 800 Axio Inverted confocal microscope (Plan-Apochromat 40x/1.3 oil or 63x/1.40 oil).
Organelles morphology analysis
Mitochondria, peroxisomes, endosomes, and Golgi number, branching and morphology were quantified in ImageJ5,6. Briefly, the confocal images were converted to binary by threshold and then converted to a skeleton that represented the features in the original image using a wireframe of lines one pixel wide. All pixels within a skeleton were then measured using analyze skeleton. The output will give the number of particles, which denotes the total number of mitochondria, the aspect ratio (AR) represents the “length to width ratio” and the form factor (FF), the complexity and branching aspect of mitochondria were calculated from circularity. For the endosome, Feret diameter was used to plot the graph.
Western blotting
ShCtrl, ShOpa1 transduced cells were lysed and 35 μg of protein was electroblotted to PVDF membranes (EMD Millipore, Burlington, MA, USA). The membranes were blocked and incubated overnight at 4oC with antibodies against BNIP3 (3769), BNIP3L (12396), LC3B (12741), LAMP1 (9091), SQSTM1/p62 (5114), CHOP (78063), Beclin1 (3738), MFF (84580), MFN2 (9482), FIS1 (32525), Ub (3933), pUb (62802), GAPDH (5174) (Cell Signaling), DRP1 (611113), OPA1 (612607) (BD Biosciences, San Jose, CA, USA), PARKIN (sc-32282), MFN1 (ab126575) (Abcam, Cambridge, MA, USA). Immunolabeling was detected using an ECL reagent (Azure biosystems 300, Dublin, CA, USA), and densitometric analysis was performed using ImageJ software. The uncropped scans of the most important blots are provided as a Source Data file.
Seahorse XF analysis
Maximum glycolytic capacity and ATP production rate using methods reported by Mookerjee and colleagues5,21,22. In brief, cells were plated in a 24-well Seahorse V7-PS test plate under hypoxia 24 h before the experiment. Cells were washed three times with 500 μl of KRPH (Krebs Ringer Phosphate HEPES) before being cultured for one hour at 37 oC in 100% air. For glycolytic capacity calculation, OCR and ECAR were determined in a Seahorse XFe24 analyzer (Agilent Technologies) by adding 10 mM glucose, 1 μM rotenone plus 1 μM myxothiazol, and 200 μM monensin plus 1 μM FCCP via ports A-C. OCR and ECAR were assessed by adding 10 mM glucose, 2 μg oligomycin, 1 μM rotenone plus 1 μM myxothiazol to determine ATP generation rate from oxidative and glycolytic pathways.
Widely targeted small metabolite measurements
Cells were transduced with ShCtrl and ShOpa1 viral particles as described above. On the third day, the medium was changed to DMEM without pyruvate, 10% dialyzed FBS (Sigma F0392), and the cells were grown under hypoxia for 24 h. Cells were washed and collected in ice-cold 80% methanol before being snap-frozen in liquid nitrogen and kept at -80 degrees Celsius until use. Prior to analyzing the metabolites, cell pellet samples were centrifuged and pipetted into a LC sampling vial. Each sample had internal standards. After drying under mild nitrogen flow, the samples were reconstituted in 150 μl of 80% methanol for injection. The samples were analyzed on an ABsciex 6500+ coupled with a Waters UPLC. Small metabolites were separated using the Ace PFP column and the iHILIC-p column (HILICON) and a pooled quality control (QC) sample was added to the sample list. The QCs sample was injected six times to calculate the coefficient of variation (CV) for data quality control. Metabolites with CVs lower than 30% used for the quantification. MetaboAnalyst 5.0 web server was used to analyze the data, and acceptable metabolites were manually input using the HMDB number. The small metabolites pathway data bank (SMPDB), which contains 99 compounds based on normal human metabolic pathways, was used for enrichment and pathway analysis. MetaboAnalyst provides the list of pathways in which these metabolites are found.
13C-Metabolic flux analysis
For [1,2]-13C-glucose flux analysis, DMEM was supplemented with 2.5 mM 1,2-13C glucose and 2.5 mM unlabeled glucose for a total of 5 mM glucose with a 50% label. A volume of 100 μl of cell culture medium from 1,2-13C glucose experiment was treated with 400 μl of methanol. After centrifugation, the supernatant was transferred to a LC-MS sampling vial and dried under gentle nitrogen flow. The sample was reconstituted into 100 ml of 80% methanol for LC-MS injection. Metabolite separation was performed on an ACE PFP-C18 column (1.7 μM x 1 mm×100 mm) and analyzed on a ABSciex 6500+ with a Multiple Reaction Monitoring (MRM) mode. The glycolysis, pentose cycle, PDH, PC, PDH/PC, and PDH + PC fluxes were calculated using methods reported by Madhu et al. 5. Briefly, carbon was tracked in two ion pairs (precursor ion/product ion): one from position 1 to 4 (M0, m/z 146/102, loss of carboxyl group at the position of carbon 5) and the other from position 1 to position 5 (M0, m/z 146/128, loss of water). The two 13C atoms (M2) originate from pyruvate (after coming from glucose) that enters either by pyruvate carboxylase (PCX), which is responsible for carbons 2 and 3, or pyruvate dehydrogenase (PDH) (carbons 4 and 5). The ion pair (m/z 148/103, M1) monitors two 13C in the precursor ion at positions of 4 and 5, and the product ion of one 13C at position 4. This ion pair reflects the activity of PDH. The C1-4 glutamate fragment’s M2 (m/z 148/104) shows that pyruvate enters the oxaloacetate pool through PCX and that there is 13C present at the second and third carbons5,6,57. The presence of both 13C at the second and third carbon positions or the presence of both 13C at the fourth and fifth carbon positions is reflected in M2 of the glutamate C1-5 fragment (m/z 148/130), which adds up to the total TCA cycle flow. During glycolysis, [1,2]-13C-glucose will generate two 13C labeled pyruvate ([2,3]-13C pyruvate) (M2, m/z 91/91). With pentose cycle via pentose phosphate pathway (PPP), 13C on the first position of glucose will be lost and will subsequently generate one 13C labeled lactate (M1, m/z 90/90) after glycolysis. The fraction of glucose converted to pentose phosphate in the pentose cycle is indicated by the lactate labeling and can be defined as: Pentose cycle = (M1/M2)/ (3 + M1/M2)5,6,57.
When flux was assessed in the U13C-glutamine labeling experiment, labeled glutamine was added to be 50% of the total DMEM glutamine concentration. Methanol extraction from [1,2]-13C-glucose and U13C glutamine-labeled cell pellets were dried under gentle nitrogen flow. The dried samples were derivatized with a methyl-moximation (with 15 mg/ml methoxyamine in pyridine, 30 oC for 90 min) and MTBSTFA (at 70oC for 60 min). The samples were then analyzed with an Agilent GC-MS, with an electron impact mode and a DB-5MS column (Agilent) following our protocol5. The data were analyzed with Mass Hunter Quantitative Analysis software (Agilent). The enrichment was calculated after subtraction from the background of the non-labeled treatment samples. Fractional enrichments from G to R is sigma (stands for sigma mean), and it is equal to the weighted mean average of the metabolite’s enrichment (sigma mn = 1 xm1+2xm2+3xm3, etc.).
Mitochondrial DNA quantification
DNA was isolated from ShCtrl and ShOpa1 cells using DNA extraction lysis buffer (Qiagen). The mitochondrial DNA (mtDNA) content was determined using qPCR (Applied Biosystems A25742 PowerUP SYBR green master mix). For mitochondrial DNA (mtDNA; Nd1) 5’-GGC TCC TTC TCC CTA CAA ATA C-3’ and 5’-TGT TTC TGC AAG GGT TGA AAT G-3’. For nuclear DNA (nDNA; Cox4) 5’-ATG TTG ATC GGC GTG ACT AC-3’ and 5’-AGT GGG CCT TCT CCT TCT-3’ were used. The ratio of mtDNA to nDNA (mtDNA/nDNA) reflects the relative mtDNA content.
Generation of conditional knockout mice
All mouse studies were carried out in conformity with the Institutional Animal Care and Use Committee (IACUC) of Thomas Jefferson University’s applicable rules and regulations. Opa1fl/fl on mixed C57BL/6–129/SvEv background were described previously52. Aggrecan-CreERT2 mice were from Jackson Laboratories (stock #019148). For all experiments, skeletally mature 3-month-old female and male mice of all genotypes received an intraperitoneal injection of 100 mg/kg tamoxifen (Sigma-Aldrich, St. Louis, MO, USA) dissolved in palm oil (Sigma-Aldrich) for 3 consecutive days to activate Cre recombinase. OPA1 conditional knock-out (Opa1cKO:AcanCreERT2Opa1fl/fl) and control (Opa1CTR: Opa1fl/fl) mice were analyzed after 3 (7-month-old), 9 (12-month-old), and 17 (20-month-old) months after tamoxifen injections. Mice were housed in specific pathogen-free conditions in individually ventilated cages with a consistent light/dark cycle, food, water, and controlled temperature.
Histological analysis of intervertebral disc
Spines were dissected and immediately fixed in freshly made 4% paraformaldehyde (PFA) for 48 hours, followed by decalcification in 20% EDTA at 4 °C prior to embedding in paraffin. Lumbar and caudal motion segments from 7-month-old (6-7 mice/genotype, 25-30 lumbar and 12-14 caudal discs/genotype), 12-month-old (8-9 mice/genotype, 32-36 lumbar and 18-19 caudal discs/genotype) 20-month-old (9-11 animals/genotype, 36-44 lumbar and 27-33 caudal discs/genotype) WT and Opa1cKO mice were processed. 7 μm coronal sections were stained with 1% SafraninO, 0.05% Fast Green, and 1% Hematoxylin and imaged on an Axio Imager A2 microscope using 5x/0.15 N-Achroplan or 20x/0.5 EC Plan-Neofluar (Carl Zeiss) objectives, Axiocam 105 color camera, and Zen2™ software (Carl Zeiss AG, Germany). 4 blinded observers used a Modified Thompson grading scale for the NP and AF compartments to score histological sections11,47,61. Picrosirius red staining (Polysciences, 24901) was used to measure collagen fibril thickness, and pictures were captured using a polarizing light microscope (Eclipse LV100 POL; Nikon) and a 4x /0.25 Pol /WD 7.0 (Nikon) objective. The color thresholds for green (thin), yellow (middle), and red (thick) fibers were established using the Nikon NIS Elements Viewer program26,77. For all samples, the color threshold values remained constant. Since phenotypic outcome at each of the spinal levels is governed by the unique interactions between the environmental (mechanical), biological, and genetic factors, each disc was considered as a unique sample11,47,61.
Immunohistochemistry and confocal analysis
Mid-coronal 7 μm thick disc tissue sections were de-paraffinized and incubated in microwaved citrate buffer for 20 min, proteinase K for 10 min at room temperature, or Chondroitinase ABC for 30 min at 37 °C for antigen retrieval. Sections were blocked in 5% normal serum (Thermo Fisher Scientific, 10000 C) in PBS-T (0.4% Triton X-100 in PBS) and incubated with antibody against OPA1 (1:250, Abcam, ab157457), LC3B (1:200, Novus Biologicals, NB100-2220) GLUT-1 (1:200, Abcam, ab40084), CA3 (1:200; Abcam, 196835), KRT19 (1:3, DSHB, TROMA-III/supernatant, AB-2133570), Collagen X (1:500, Abcam, ab58632) Collagen I (1:100; Thermo Fisher, PA1-26204), COMP (1:200, Abcam, ab231977), MMP13 (1:200, Abcam, ab39012), ACAN/aggrecan (1:50; Millipore Sigma, AB1031), ARGXX. (1:200; Abcam, ab3773). When using mouse primary antibodies, Mouse on Mouse Kit (Vector laboratories, BMK2202) was used for blocking during primary antibody incubation. Tissue sections were thoroughly washed and incubated with Alexa Fluor−488 or 594 or 647 conjugated secondary antibodies (Jackson ImmunoResearch Lab, Inc.), at a dilution of 1:700 for 1 h at room temperature in the dark. The sections were washed with PBS-T (0.4% Triton X-100 in PBS) and mounted with ProLong® Gold Antifade Mountant with DAPI (Thermo Fisher Scientific, P36934). All mounted slides were visualized using a Zeiss LSM 800 Axio Inverted confocal microscope (Plan-Apochromat 5x/0.16 or 10x/0.45). ImageJ 1.52i (NIH) was used for all quantifications. Images were threshold to generate binary images, then NP and AF compartments were contoured manually using the Freehand Tool. The ROI were examined using the Area Fraction measurement.
RNA-Sequencing and bioinformatic analysis
Total DNA-free RNA (RIN > 8) extracted from NP tissues of Opa1cKO and WT mice using RNeasy mini columns (Qiagen) (n = 6 mice/genotype) was used for total RNA sequencing using standard protocols (Azenta Life Sciences, Burlington, MA, USA) (GSE270889). Significantly up- and downregulated DEGs (FDR < 0.05) were analyzed using the GTAC-CompBio Analysis Tool (PercayAI Inc., St. Louis, MO) as described previously27. The Assertion Engine module within CompBio was used to determine global level and thematic similarities between Opa1cKO and transcriptomic profiles of human degenerated NP tissue samples (GSE70362)5,29.
ScRNA-Seq data analysis
Mouse intervertebral discs samples at 8-week old (GSE186549)30 and at P30 (GSM7496680)31, 8-week-old rat intervertebral disc (GSE211407)32, and healthy young human nucleus pulposus tissue (GSM7831813)33 were determined. Rat disc single-cell RNA-seq paired-end run read one (R1) and read two (R2) FASTQ files were downloaded from European Nucleotide Archive. Using 10x Genomics Cloud Analysis platform and single cell 3’ gene expression library type, FASTQs were aligned to the Rattus Norvegicus. Single-cell data analysis outputs were generated and visualized using the Loupe Cell Browser. Mouse and human data were attained from Gene Expression Omnibus. For both mouse and human ScRNA seq, the Seurat package (version 4.3.0) was used to show aggrecan, OPA1, brachyury/T, keratin 18 and keratin 19-expressing cells.
Micro-computed tomography (μCT) analyses
μCT imaging was performed on the lumbar spine of 7, 12, and 20-month WT and Opa1cKO mice using the high-resolution μCT scanner (Skyscan 1272, Bruker, Belgium). Lumber segments L2-6 were placed in PBS and scanned with an energy of 50 kVp and current of 200 mA, resulting in 15 mm3 voxel size resolution. Trabecular parameters were assessed in the 3D reconstructed trabecular tissue using Skyscan CT analysis (CTAn) software by contouring the region of interest (ROI). The bone volume percentage (BV/TV), trabecular number (Tb. N.), trabecular thickness (Tb. Th.), and trabecular separation (Tb. Sp.) of the resulting datasets were all evaluated. Cortical bone volume (BV), cross-sectional thickness (Cs. Th.), mean cross-sectional bone area (B. Ar), and mean cross-sectional tissue area (T. Ar) were all measured in two dimensions. A standard curve was established with a mineral density calibration phantom pair (0.25 g/cm3 CaHA and 0.75 g/cm3 CaHA) to determine mineral density. Intervertebral disc height and the length of the vertebral bones were measured and averaged along the dorsal, midline, and ventral regions in the sagittal plane. Disc height index (DHI) was calculated as previously described26,77.
Mouse hindlimbs (right limb) were harvested, the surrounding muscles were removed, and limbs were fixed in formalin (10%, 48 h). MicroCT analysis of the femur, tibia, and knee joint were performed on a Bruker SkyScan 1275 scanner as we have previously described34. MicroCT reconstructions and quantitative analysis of tibial subchondral bone volume fraction (BV/TV), trabecular thickness (Tb.Th), trabecular separation (Tb.Sp), and subchondral bone plate thickness (SCBP) were performed using the SkyScan CT Analyzer (CTan) and CT Vox software on coronal slices that spanned the medial and lateral tibial plateaus.
Histological and histomorphometry analysis of OA
Hindlimbs were decalcified (EDTA, 19%, 21 days), processed, paraffin embedded, and sectioned along the coronal plane, as we have described34. Mid-coronal sections were stained with H&E, or toluidine blue, and OA severity was analyzed by Articular Cartilage Structure (ACS) score (0-12 scale), toluidine blue score (0–12 scale), and osteophyte scoring (0–3 scale) on the medial and lateral tibial plateaus (MTP, LTP) and femoral condyles (MFC, LFC). Briefly, the ACS and toluidine blue scores for OA damage are calculated using a 0-12 scale where: 0=normal/no damage; 1-3=minimal damage; 4-6= mild damage; 7-9=moderate damage; and 10-12=severe damage. Osteophytes (0–3 scale) were scored using H&E-stained sections, as we have described, were: 0=no osteophyte; 1=small osteophyte (same as cartilage thickness), 2=medium osteophyte (up to 3 times the thickness of cartilage); 3=large osteophyte (greater than 3 times the thickness of cartilage). All sections were scored by a blinded individual who has experience of the grading schemes. Synovial hyperplasia was assessed using a 0-3 scale as described earlier78. In joints that presented with synovial ossification, a maximal synovial hyperplasia score was assigned. Analysis was performed by a blinded scorer with experience in the OA scoring techniques. Detailed histomorphometric analysis of articular cartilage thickness and area, calcified cartilage thickness and area, and subchondral bone thickness and area were analyzed on the MTP and LTP using ImageJ software as we have previously described34. Joint tissue damage and OA severity can differ between lateral and medial compartments within the same joint. As such, all histological and histomorphometric analyses were conducted using separate images (10X) taken from the lateral and medial compartments. Lateral and medial scores or histological quantification is presented separately to demonstrate these differences, which is consistent with our previous work34.
Statistical analysis
Statistical analysis was performed using Prism9 (GraphPad, La Jolla, CA, USA). The quantitative data are represented as mean ± SEM or Box and whiskers plots showing all data points with median and interquartile range and maximum and minimum values. Data distribution was checked with the Shapiro-Wilk normality test, and differences between two groups were assessed by t-test or Mann-Whitney test as appropriate. One-way ANOVA, whereas non-normally distributed data were analyzed using or Kruskal–Wallis test, with an appropriate post-hoc test (Sidak’s multiple comparisons test) used for comparisons between more than two groups. Analysis of Modified Thompson Grading data distribution and fiber and fiber thickness distribution were performed using chi-square test; p < 0.05.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data generated or analyzed during this study are included in this article. Transcriptomic data analyzed in this study are available in the GEO database (GSE270889, GSE70362). Source data are provided with this paper.
References
Disease, G. B. D., Injury, I. & Prevalence, C. Global, regional, and national incidence, prevalence, and years lived with disability for 354 diseases and injuries for 195 countries and territories, 1990-2017: a systematic analysis for the Global Burden of Disease Study 2017. Lancet 392, 1789–1858 (2018).
Madhu, V., Guntur, A. R. & Risbud, M. V. Role of autophagy in intervertebral disc and cartilage function: Implications in health and disease. Matrix Biol. 100-101, 207–220 (2021).
Collins, J. A., Diekman, B. O. & Loeser, R. F. Targeting aging for disease modification in osteoarthritis. Curr. Opin. Rheumatol. 30, 101–107 (2018).
Merceron, C. et al. Loss of HIF-1alpha in the notochord results in cell death and complete disappearance of the nucleus pulposus. PLoS One 9, e110768 (2014).
Madhu, V. et al. The mitophagy receptor BNIP3 is critical for the regulation of metabolic homeostasis and mitochondrial function in the nucleus pulposus cells of the intervertebral disc. Autophagy 19, 1821–1843 (2023).
Madhu, V. et al. Hypoxic regulation of mitochondrial metabolism and mitophagy in nucleus pulposus cells is dependent on HIF-1alpha-BNIP3 Axis. J. Bone Min. Res 35, 1504–1524 (2020).
Otte, P. Basic cell metabolism of articular cartilage. Manometric studies. Z. Rheumatol. 50, 304–312 (1991).
Gavriilidis, C., Miwa, S., von Zglinicki, T., Taylor, R. W. & Young, D. A. Mitochondrial dysfunction in osteoarthritis is associated with down-regulation of superoxide dismutase 2. Arthritis Rheum. 65, 378–387 (2013).
Collins, J. A. et al. Oxidative stress promotes peroxiredoxin hyperoxidation and attenuates pro-survival signaling in aging chondrocytes. J. Biol. Chem. 291, 6641–6654 (2016).
Loeser, R. F. et al. Overexpression of peroxiredoxin 3 in cartilage reduces the severity of age-related osteoarthritis but not surgically induced osteoarthritis in mice. ACR Open Rheumatol. 4, 441–446 (2022).
Novais, E. J. et al. Comparison of inbred mouse strains shows diverse phenotypic outcomes of intervertebral disc aging. Aging Cell 19, e13148 (2020).
Pernas, L. & Scorrano, L. Mito-morphosis: Mitochondrial fusion, fission, and cristae remodeling as key mediators of cellular function. Annu Rev. Physiol. 78, 505–531 (2016).
Song, Z., Ghochani, M., McCaffery, J. M., Frey, T. G. & Chan, D. C. Mitofusins and OPA1 mediate sequential steps in mitochondrial membrane fusion. Mol. Biol. Cell 20, 3525–3532 (2009).
Olichon, A. et al. The human dynamin-related protein OPA1 is anchored to the mitochondrial inner membrane facing the inter-membrane space. FEBS Lett. 523, 171–176 (2002).
Frezza, C. et al. OPA1 controls apoptotic cristae remodeling independently from mitochondrial fusion. Cell 126, 177–189 (2006).
Gilkerson, R. W., Selker, J. M. & Capaldi, R. A. The cristal membrane of mitochondria is the principal site of oxidative phosphorylation. FEBS Lett. 546, 355–358 (2003).
Quintana-Cabrera, R. et al. The cristae modulator Optic atrophy 1 requires mitochondrial ATP synthase oligomers to safeguard mitochondrial function. Nat. Commun. 9, 3399 (2018).
Tezze, C. et al. Age-associated loss of OPA1 in muscle impacts muscle mass, metabolic homeostasis, systemic inflammation, and epithelial senescence. Cell Metab. 25, 1374–1389 e1376 (2017).
Choi, H. et al. Hypoxia promotes noncanonical autophagy in nucleus pulposus cells independent of MTOR and HIF1A signaling. Autophagy 12, 1631–1646 (2016).
Okatsu, K. et al. Phosphorylated ubiquitin chain is the genuine Parkin receptor. J. Cell Biol. 209, 111–128 (2015).
Mookerjee, S. A., Gerencser, A. A., Nicholls, D. G. & Brand, M. D. Quantifying intracellular rates of glycolytic and oxidative ATP production and consumption using extracellular flux measurements. J. Biol. Chem. 292, 7189–7207 (2017).
Mookerjee, S. A., Nicholls, D. G. & Brand, M. D. Determining maximum glycolytic capacity using extracellular flux measurements. PLoS One 11, e0152016 (2016).
Spinelli, J. B. et al. Fumarate is a terminal electron acceptor in the mammalian electron transport chain. Science 374, 1227–1237 (2021).
Henry, S. P. et al. Generation of aggrecan-CreERT2 knockin mice for inducible Cre activity in adult cartilage. Genesis 47, 805–814 (2009).
Novais, E. J., Diekman, B. O., Shapiro, I. M. & Risbud, M. V. p16(Ink4a) deletion in cells of the intervertebral disc affects their matrix homeostasis and senescence associated secretory phenotype without altering onset of senescence. Matrix Biol. 82, 54–70 (2019).
Boneski, P. K. et al. Abcc6 null mice-a model for mineralization disorder PXE shows vertebral osteopenia without enhanced intervertebral disc calcification with aging. Front Cell Dev. Biol. 10, 823249 (2022).
Winkler, E. S. et al. Human neutralizing antibodies against SARS-CoV-2 require intact Fc effector functions for optimal therapeutic protection. Cell 184, 1804–1820 e1816 (2021).
Rath, S. et al. MitoCarta3.0: an updated mitochondrial proteome now with sub-organelle localization and pathway annotations. Nucleic Acids Res 49, D1541–D1547 (2021).
Kazezian, Z. et al. Gene expression profiling identifies interferon signalling molecules and IGFBP3 in human degenerative annulus fibrosus. Sci. Rep. 5, 15662 (2015).
Tan, Z. et al. Progenitor-like cells contributing to cellular heterogeneity in the nucleus pulposus are lost in intervertebral disc degeneration. Cell Rep. 43, 114342 (2024).
Zhang, C. et al. Single cell RNA sequencing reveals emergent notochord-derived cell subpopulations in the postnatal nucleus pulposus. FASEB J. 38, e23363 (2024).
Rohanifar, M. et al. Single cell RNA-sequence analyses reveal uniquely expressed genes and heterogeneous immune cell involvement in the rat model of intervertebral disc degeneration. Appl Sci. (Basel) 12, 8244 (2022).
Chen, F. et al. Serglycin secreted by late-stage nucleus pulposus cells is a biomarker of intervertebral disc degeneration. Nat. Commun. 15, 47 (2024).
Collins, J. A. et al. Cartilage-specific Sirt6 deficiency represses IGF-1 and enhances osteoarthritis severity in mice. Ann. Rheum. Dis. 82, 1464–1473 (2023).
Delettre, C. et al. Nuclear gene OPA1, encoding a mitochondrial dynamin-related protein, is mutated in dominant optic atrophy. Nat. Genet 26, 207–210 (2000).
Yu-Wai-Man, P. et al. Multi-system neurological disease is common in patients with OPA1 mutations. Brain 133, 771–786 (2010).
Hudson, G. et al. Mutation of OPA1 causes dominant optic atrophy with external ophthalmoplegia, ataxia, deafness and multiple mitochondrial DNA deletions: a novel disorder of mtDNA maintenance. Brain 131, 329–337 (2008).
Murley, A. & Nunnari, J. The emerging network of mitochondria-organelle contacts. Mol. Cell 61, 648–653 (2016).
Birgisdottir, A. B., Lamark, T. & Johansen, T. The LIR motif - crucial for selective autophagy. J. Cell Sci. 126, 3237–3247 (2013).
Liao, C. et al. Dysregulated mitophagy and mitochondrial organization in optic atrophy due to OPA1 mutations. Neurology 88, 131–142 (2017).
Zaninello, M. et al. Inhibition of autophagy curtails visual loss in a model of autosomal dominant optic atrophy. Nat. Commun. 11, 4029 (2020).
Valverde, D. P. et al. ATG2 transports lipids to promote autophagosome biogenesis. J. Cell Biol. 218, 1787–1798 (2019).
Imai, K. et al. Atg9A trafficking through the recycling endosomes is required for autophagosome formation. J. Cell Sci. 129, 3781–3791 (2016).
Razi, M., Chan, E. Y. & Tooze, S. A. Early endosomes and endosomal coatomer are required for autophagy. J. Cell Biol. 185, 305–321 (2009).
Botte, A. et al. Ultrastructural and dynamic studies of the endosomal compartment in Down syndrome. Acta Neuropathol. Commun. 8, 89 (2020).
Gosavi, N., Lee, H. J., Lee, J. S., Patel, S. & Lee, S. J. Golgi fragmentation occurs in the cells with prefibrillar alpha-synuclein aggregates and precedes the formation of fibrillar inclusion. J. Biol. Chem. 277, 48984–48992 (2002).
Choi, H. et al. A novel mouse model of intervertebral disc degeneration shows altered cell fate and matrix homeostasis. Matrix Biol. 70, 102–122 (2018).
Krishnan, Y. & Grodzinsky, A. J. Cartilage diseases. Matrix Biol. 71-72, 51–69 (2018).
Brand, M. D. The efficiency and plasticity of mitochondrial energy transduction. Biochem Soc. Trans. 33, 897–904 (2005).
Salin, K., Auer, S. K., Rey, B., Selman, C. & Metcalfe, N. B. Variation in the link between oxygen consumption and ATP production, and its relevance for animal performance. Proc. Biol. Sci. 282, 20151028 (2015).
Sanchez-Rodriguez, R. et al. OPA1 drives macrophage metabolism and functional commitment via p65 signaling. Cell Death Differ. 30, 742–752 (2023).
Zhang, Z. et al. The dynamin-related GTPase Opa1 is required for glucose-stimulated ATP production in pancreatic beta cells. Mol. Biol. Cell 22, 2235–2245 (2011).
Amini, P. et al. Neutrophil extracellular trap formation requires OPA1-dependent glycolytic ATP production. Nat. Commun. 9, 2958 (2018).
Chao de la Barca, J. M. et al. Metabolomics hallmarks OPA1 variants correlating with their in vitro phenotype and predicting clinical severity. Hum. Mol. Genet 29, 1319–1329 (2020).
Eisenberg, T. et al. Cardioprotection and lifespan extension by the natural polyamine spermidine. Nat. Med. 22, 1428–1438 (2016).
Singh, P. et al. Taurine deficiency as a driver of aging. Science 380, eabn9257 (2023).
Izzo, L. T. et al. Acetylcarnitine shuttling links mitochondrial metabolism to histone acetylation and lipogenesis. Sci. Adv. 9, eadf0115 (2023).
Pereira, R. O. et al. OPA1 deletion in brown adipose tissue improves thermoregulation and systemic metabolism via FGF21. Elife 10, e66519 (2021).
Martinez-Reyes, I. & Chandel, N. S. Mitochondrial TCA cycle metabolites control physiology and disease. Nat. Commun. 11, 102 (2020).
Schworer, S. et al. Fibroblast pyruvate carboxylase is required for collagen production in the tumour microenvironment. Nat. Metab. 3, 1484–1499 (2021).
Silagi, E. S. et al. Lactate efflux from intervertebral disc cells is required for maintenance of spine health. J. Bone Min. Res 35, 550–570 (2020).
Sarver, J. J. & Elliott, D. M. Mechanical differences between lumbar and tail discs in the mouse. J. Orthop. Res. 23, 150–155 (2005).
McCormack, D. J., Puttock, D. & Godsiff, S. P. Medial compartment osteoarthritis of the knee: a review of surgical options. EFORT Open Rev. 6, 113–117 (2021).
Cicuttini, F. M. et al. Compartment differences in knee cartilage volume in healthy adults. J. Rheumatol. 29, 554–556 (2002).
Burtscher, J., Millet, G. P., Place, N., Kayser, B. & Zanou, N. The muscle-brain axis and neurodegenerative diseases: the key role of mitochondria in exercise-induced neuroprotection. Int J. Mol. Sci. 22, 6479 (2021).
Marques-Aleixo, I. et al. Physical exercise improves brain cortex and cerebellum mitochondrial bioenergetics and alters apoptotic, dynamic and auto(mito)phagy markers. Neuroscience 301, 480–495 (2015).
van Rossom, S. et al. Knee Joint Loading in Healthy Adults During Functional Exercises: Implications for Rehabilitation Guidelines. J. Orthop. Sports Phys. Ther. 48, 162–173 (2018).
Gan, X., Huang, S., Yu, Q., Yu, H. & Yan, S. S. Blockade of Drp1 rescues oxidative stress-induced osteoblast dysfunction. Biochem Biophys. Res Commun. 468, 719–725 (2015).
Ballard, A. et al. The tethering function of mitofusin2 controls osteoclast differentiation by modulating the Ca(2+)-NFATc1 axis. J. Biol. Chem. 295, 6629–6640 (2020).
Gruber, H. E. et al. Mitochondrial gene expression in the human annulus: in vivo data from annulus cells and selectively harvested senescent annulus cells. Spine J. 11, 782–791 (2011).
Kuhl, I. et al. Transcriptomic and proteomic landscape of mitochondrial dysfunction reveals secondary coenzyme Q deficiency in mammals. Elife 6, e30952 (2017).
Huang, M. et al. Mitochondrial dysfunction-induced H3K27 hyperacetylation perturbs enhancers in Parkinson’s disease. JCI Insight 6, e138088 (2021).
Widjaja, A. A. et al. Inhibition of IL-11 signalling extends mammalian healthspan and lifespan. Nature 632, 157–165 (2024).
Palikaras, K., Lionaki, E. & Tavernarakis, N. Coordination of mitophagy and mitochondrial biogenesis during ageing in C. elegans. Nature 521, 525–528 (2015).
Fivenson, E. M. et al. Mitophagy in neurodegeneration and aging. Neurochem Int 109, 202–209 (2017).
Risbud, M. V. et al. Nucleus pulposus cells express HIF-1 alpha under normoxic culture conditions: a metabolic adaptation to the intervertebral disc microenvironment. J. Cell Biochem 98, 152–159 (2006).
Johnston, S. N., Madhu, V., Shapiro, I. M. & Risbud, M. V. Conditional Deletion of HIF-2alpha in Mouse Nucleus Pulposus Reduces Fibrosis and Provides Mild and Transient Protection From Age-Dependent Structural Changes in Intervertebral Disc. J. Bone Min. Res 37, 2512–2530 (2022).
Rowe, M. A. et al. Reduced osteoarthritis severity in aged mice with deletion of macrophage migration inhibitory factor. Arthritis Rheumatol. 69, 352–361 (2017).
Acknowledgements
We thank Drs. Irwin Kurland and Yunping Qiu, Stable Isotope and Metabolomics Core Facility, Albert Einstein College of Medicine, for metabolic flux analyses. We thank Dr. Gyorgy Csordas and Timothy Schneider, and Thomas Jefferson University for help with TEM studies. This study was supported by NIH grants R01AR055655, R01AR074813, and R01AG073349 to MVR. JC acknowledges support from NIA R01AG078609. HS is supported by NIH grant R35GM144103 and the Ethan and Karen Leder HAP Scholar Award.
Author information
Authors and Affiliations
Contributions
Conceptualization: V.M., M.V.R.; Methodology: V.M., M.V.R.; Investigation: V.M., M.H.M., A.C., K.S., K.I., O.H., P.K.B., R.A.B., J.A.C.; Visualization: V.M.; Supervision: M.V.R.; Writing - original draft: V.M., M.V.R.; Writing - review and editing: V.M., K.S., R.A.B., M.V.R., J.A.C., H.S.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Madhu, V., Hernandaz-Meadows, M., Coleman, A. et al. The loss of OPA1 accelerates intervertebral disc degeneration and osteoarthritis in aged mice. Nat Commun 16, 5996 (2025). https://doi.org/10.1038/s41467-025-60933-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-60933-9
This article is cited by
-
Targeting the mitochondrial metabolite-dynamics-MDVs-MitoEVs axis: a new frontier in osteoarthritis management
Journal of Translational Medicine (2026)
-
Multi-layered integrated shielding: engineering ferroptosis-resistant mesenchymal stem cells for precision therapy of intervertebral disc degeneration
Apoptosis (2026)