Abstract
Extracellular vesicles (EVs) carry complex glycoproteins critical mediators of disease progression and hold great potential for liquid biopsy applications. However, their low abundance presents significant analytical challenges. This study develop a versatile platform for EV N-glycoproteomic analysis (EVGpro), which enables tandem EV capture, EV protein digestion, and enrichment of N-glycopeptides. Functionalized magnetic nanochains with cell-penetrating peptides (octa-arginine, R8) enable efficient EV isolation, followed by on-chain hydrolysis to expose PEG2000 for ultrasensitive glycopeptide enrichment. The streamlined EVGpro platform identified up to 17 times more EV glycoproteins than ultracentrifugation across seven biofluids, profiling 398 EV glycoproteins from just 1 μL tear. Applied to seminal plasma, EVGpro uncover disease-specific glycoprotein signatures and identified previously unreported subtypes of asthenozoospermia. This robust and scalable approach advances the sensitive detection of disease biomarkers and molecular subtypes from diverse biofluids to enable precision medicine.
Similar content being viewed by others
Introduction
Extracellular vesicles (EVs) are membrane-bound particles secreted by nearly all cells into the extracellular space, providing valuable insights into organ function, cellular activities, and disease states1,2. The phospholipid bilayer of EVs protects their cargo, including proteins, from enzymatic degradation, thereby rendering them ideal for early disease detection and monitoring via liquid biopsy1,3,4,5. EV proteins have emerged as promising biomarkers for the diagnosis and prognosis of various diseases1,6,7, such as prostate cancer8, breast cancer9,10, and male infertility4,11,12. Protein glycosylation, a prevalent post-translational modification occurring in 50–70% of proteins, is crucial in numerous biological processes3,13,14. Furthermore, the majority of clinically approved protein biomarkers are glycosylated proteins15,16. Notably, EVs are particularly rich in glycoproteins, which are essential for intercellular communication, immunomodulation, and disease pathogenesis2,3,9. The unique glycosylation patterns of these N-glycoproteins can provide insights into the cellular origins and pathological states of EVs, enhancing their potential as specific biomarkers for targeted therapies17,18. Despite the promising role of EV proteins in various diseases, the characterization of EV-associated N-glycoproteins remains underdeveloped, revealing a substantial gap in our understanding of their functional roles and potential applications.
Due to the low abundance of glycopeptides, enrichment before analysis is important for effective high-throughput identification and quantification. Traditional methods for EV N-glycoproteins enrichment involve two distinct steps: isolation of EVs and enrichment of N-glycoproteins from enzymatic digests of EV protein lysates. Ultracentrifugation (UC) and size-exclusion chromatography (SEC) are usually used to isolate EVs4,19,20,21. Recently, Su et al. developed a charge-based fractionation method using Capto Core 700 and Q-Sepharose resins for EV enrichment22, and Hou et al. integrated size-based filtration and electrophoretic separation to enrich EVs23. The hydrophilic interaction chromatography (HILIC) is widely used for N-glycoprotein enrichment24,25,26. However, the challenges inherent in analyzing EV N-glycoproteins arise from the trace number of EVs in biofluids and the low abundance of N-glycopeptides within EVs25,27. For example, Hillary et al. achieved the enrichment of 1,222 EV N-glycopeptides and 578 N-glycoproteins from 1 mL of plasma using UC and hydrazide chemistry3. Whereas Bai et al. employed an exosome isolation kit to extract EVs from 200 µL of serum, then subsequently enriched N-glycopeptides from EV enzymatic digests using poly(NIPAM co-hydrazide) materials, and finally identified only 161 N-glycoproteins27. Notably, hydrazide chemistry-based enrichment methods compromised the structural integrity of glycans. Alternatively, magnetic beads-based methods offer a promising alternative for EV protein enrichment, combining rapid magnetic response, stable chemical properties, and ease of surface functionalization18,19,24,25. Simplified approaches using magnetic beads can enhance the sensitivity of EV N-glycoprotein analysis and provide better solutions for analyzing precious biofluid samples.
In this study, we developed a sensitive magnetic nanochains-based strategy to tandemly enrich EV and N-glycoproteins (EVGpro) from the ultratrace of biofluids. The EVGpro employs octa-arginine (R8) for membrane penetration and PEG2000 for enhanced hydrophilicity, allowing for capture and lysis of EV, reduction, alkylation, and digestion of N-glycoproteins, as well as N-glycopeptide enrichment in a single tube. This method outperforms the traditional UC method and applies to various biofluids, including plasma, urine, saliva, pleural effusion, cerebrospinal fluid, ascites, tears, and seminal plasma. Approximately 17.5% of adults globally experience infertility, with male factors contributing to about 50% of these cases28. Asthenozoospermia (AZS), one of the most prevalent forms of male infertility, is marked by reduced sperm motility and accounts for 20-40% of male infertility29,30,31. Although EVs from AZS seminal plasma could affect sperm motility, investigations into changes in EV glycoproteins and their molecular subtypes are still lacking12. Our EVGpro analysis of AZS revealed different EV N-glycoprotein signatures, and two distinct molecular subtypes of AZS in both discovery and validation cohorts, which might enhance the understanding of the pathogenesis and precision therapy of AZS.
Results
Preparation and characterization of magnetic nanochains
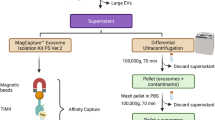
The EVGpro nanochains were Fe3O4 magnetic nanochains (Fe3O4-MN) decorated with PEG2000-R8 functional molecules. The EVs in biofluids were specifically captured using cell-penetrating peptide (octa-arginine, R8) of EVGpro. Following EVs lysis on the nanochains, trypsin was employed to simultaneously digest the EV proteins and cleave R8 from EVGpro, thereby exposing the PEG2000. Subsequently, N-glycopeptides were enriched by highly hydrophilic PEG2000 of EVGpro in the same tube based on the hydrophilic interaction liquid chromatography (HILIC) mechanism (Fig. 1A). This streamlined method using the same nanochain material in one tube avoids sample loss due to tube transfer and sequential enrichment using different materials and is suitable for N-glycoproteomic analysis of EVs of diverse biofluids.
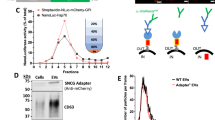
A EVs from diverse biofluids are initially captured using the cell-penetrating peptide R8 of EVGpro. After on-nanofibril protein lysis, trypsin is used to digest the proteins and R8 of EVGpro to expose PEG2000. Subsequently, EV N-glycopeptides are enriched by highly hydrophilic PEG2000 of EVGpro in the same tube based on the hydrophilic interaction liquid chromatography (HILIC) mechanism. B The representative conformation of the dominant cluster from the clustering analysis of round 1 during the convergence phase of simulation (150-200 ns), with hydrogen bonds in yellow. Blue, green, and pink represent C atoms, red represents O atoms, white indicates H atoms, dark blue indicates N atoms, and the pink loop indicates R8. C, D The SEM (C) and TEM (D) images of the nanochains. Scale bar, 200 nm for SEM and 100 nm for TEM. The experiment was repeated 3 times with similar results. E The zeta potential of Fe3O4 (coral red), Fe3O4-MN (cerulean blue), Fe3O4-MN@SH (green), and EVGpro nanochains (lavender pink) (means ± standard deviation (SD), n = 3 independent experiments). F The magnetic hysteresis curves of Fe3O4 (sky blue), Fe3O4-MN (crimson), Fe3O4-MN@SH (lilac), and EVGpro nanochains (rosy pink). G Transmittance of EVGpro (blue) and Fe3O4 (pink) during magnetic adsorption as a function of time. The zeta potential and transmittance are presented as means ± SD (n = 3 independent experiments). CSF, cerebrospinal fluid, EVs, extracellular vesicles, HILIC, hydrophilic interaction chromatography, R8, octa-arginine, Fe3O4-MN, Fe3O4 magnetic nanochains, EVGpro, EV N-glycoproteomic analysis. The “Biofluid source” section was created in BioRender. (2025) https://BioRender.com/5xrzn5h, and the “Identification” section was created in BioRender. (2025) https://BioRender.com/lq01r4m. Source data are provided as a Source Data file.
The pod-like Fe3O4-MN was synthesized through the self-assembly of nanoparticles induced by an external magnetic field, followed by the in situ polymerization of adsorbed acrylic acid precursor to form hydrogel shell. Then the SH-group was modified to Fe3O4-MN by reacting carboxyl groups with NH2-PEG1000-SH amides. At last, the nanochains were functionalized with R8 through a click chemistry reaction between SH and MAL-PEG2000-R8. The sequential amide and click chemistry processes ensured the high-efficiency and high-purity functionalization of R8 by preventing side reactions of R8 itself.
Cell-penetrating peptides (CPPs) are short peptides with exceptional membrane permeability, capable of delivering various bioactive substances into cells. Arginine- and lysine-rich cationic CPPs exhibit strong affinity for the cytoplasmic membrane under physiological conditions. One of the most well-studied CPPs, R8, binds to negatively charged cell membranes primarily through electrostatic interactions32. The hydrophilic phosphate head of phosphatidylcholine served as a model for the phospholipid bilayer of EVs. Through molecular docking studies (Supplementary Fig. 1A), R8 formed putative binding interactions with EVs with a docking energy of -2.6 kcal/mol, suggesting that R8 of EVGpro nanochains can stably enrich EVs in biofluids. Furthermore, R8 established nine hydrogen bonds and three salt bridges with sphingolipids. Notably, all salt bridging and 67% (6/9) of the hydrogen bonds were bound to the phosphate group of sphingolipids, indicating strong non-covalent binding interactions.
All-atom molecular dynamics simulations were extended to 200 ns to validate long-term binding stability, with the 150–200 ns interval showing convergent dynamics. The root mean square deviation (RMSD) of the R8-sphingolipid complex exhibited fluctuations of less than 0.5 Å across three independent trajectories, indicating structural equilibration after 150 ns (Supplementary Fig. 1B). To further investigate the stable binding conformations observed during the converged phase of the simulation (150–200 ns), clustering analysis was performed on this time window for each round of the three simulations. The most dominant cluster from each round was extracted and analyzed. The structural alignment of the dominant cluster conformations showed good overlap among the three rounds, indicating consistent and reproducible binding pose across the three simulation rounds (Supplementary Fig. 1C).
Representative dominant cluster conformations showed stable hydrogen bonds between ARG-6 and the sphingolipid phosphate headgroup (Fig. 1B, Supplementary Fig. 1D, E), with the other arginine residues (ARG-3 and ARG-7) engaging in hydrophobic interactions with lipid tails. Root mean square fluctuation (RMSF) analysis of R8 Cα atoms during 150–200 ns of simulations showed backbone fluctuations <0.6 Å, indicating structural equilibration (Supplementary Fig. 1F). The complex maintained 2–3 hydrogen bonds during 150–200 ns of simulations (Supplementary Fig. 1G). Molecular Mechanics/Generalized Born Surface Area (MM/GBSA) calculations yielded a mean binding free energy of -17.5 kcal/mol (Supplementary Table 2). The convergent stable binding results with hydrogen-bond interaction in the molecule dynamics simulation provided theoretical basis for R8 to enrich EVs in biofluids, consistent with the well-known membrane-penetrating capability of R8 as a classic CPP33,34.
The EVGpro nanochains were individual polymer-wrapped flexible nanochains with an average length of 36 μm, typically exhibiting a curved morphology as observed through scanning electron microscopy (SEM) and transmission electron microscopy (TEM) (Fig. 1C, D). The modification process was monitored using zeta potential measurement, Fourier transform infrared spectra (FTIR), and magnetic hysteresis curves. As shown in Fig. 1E, Fe3O4 had a zeta potential of −10.03 mV, which became highly negative (-36.57 mV) after introducing carboxyl groups. Subsequent immobilization of SH groups increased the zeta potential to -16.17 mV, and functionalization with R8 elevated it to +31.97 mV. Additionally, the FTIR showed a strong absorption peak at 1674 cm−1 and an additional peak at 1,728 cm−1, associated with the stretching vibration of C = O in the keto and carboxyl group respectively, indicating successful functionalization of the EVGpro (Supplementary Fig. 2A). The magnetic hysteresis curves in Fig. 1F revealed the superparamagnetic properties of EVGpro, with a saturation magnetization of 24.53 emu·g−1, which was sufficient for the rapid enrichment of EVs. As shown in Fig. 1G, the EVGpro achieved nearly complete magnetic separation within 5 min, while the Fe3O4 nanoparticles required 12 min. The transmittance of the EVGpro solution just rose from 20.32% to 73.57% after 48 h, also demonstrating good suspensibility (Supplementary Fig. 2B).
Optimization of nanochains-based EVGpro method for EV enrichment and proteomic analysis in biofluid
The EV capture efficiency of EVGpro nanochains depends mainly on the amount of EVGpro used and the incubation duration. Using an initial volume of 50 μL of seminal plasma, performance was assessed with increasing amounts of EVGpro. The number of unique proteins and peptides from EVs rose from 1162 to 1474 and 8175 to 10,608, respectively. The enrichment reached a plateau when using 0.5 mg to 1 mg EVGpro (Fig. 2A, Supplementary Data 1), indicating near-complete capture of seminal plasma EVs. Regardless of the amount of EVGpro used, Gene Ontology analysis of cell components (GO-CC) consistently showed exosomes as the most enriched term. For instance, with 0.5 mg of EVGpro, the -log10(P-value) associated with exosomes was 1.84 times higher than the second term and 4.84 times higher than the third term (305 vs. 166, 305 vs. 63, Supplementary Fig. 3A).
A The number of identified EV proteins using different amounts of EVGpro nanochains (means ± SD, n = 3 independent experiments). B TEM image of EVs eluted by EVGpro nanochains. The experiment was repeated 3 times with similar results. C Nanoparticle tracking analysis (NTA) of the particle size distribution of EVs enriched by EVGpro (blue) and UC (pink), respectively. D, E Comparison of purity (D) and recovery rate (E) of seminal plasma EVs captured by UC (pink) and EVGpro nanochains (blue) (means ± SD, n = 3 independent experiments). The purity is defined as the ratio of the number of EV particles measured by NTA to the mass of EV proteins measured by the bicinchoninic acid assay. F Proteomic analysis of seminal plasma EVs enriched by UC (pink) and EVGpro (blue) (means ± SD, n = 3 independent experiments). G The Venn diagram shows the overlap among seminal plasma proteins identified by UC (pink) and EVGpro (blue), and those in the Vesiclepedia database (green). UC, ultracentrifugation. Source data are provided as a Source Data file.
Moreover, the effects of incubation time were assessed. The results indicated that increasing the incubation duration from 30 min to 120 min did not alter the quantity of captured EV proteins, with identified proteins ranging from 1285 to 1343 (Supplementary Fig. 3B, Supplementary Data 1). This indicated complete capture of EVs in the initial 30 min of incubation. Therefore, the 0.5 mg EVGpro and 30 min culture time were used in subsequent analyses.
Nanochains-based EVGpro method outperforms the UC approach in EV enrichment and subsequent proteomic and N-glycoproteomic analysis
The UC method serves as the gold standard for EV enrichment35. To assess the performance of EVGpro nanochains relative to UC, the features of enriched EVs from seminal plasma were evaluated using a series of experiments, including Western blot (WB), nanoparticle tracking analysis (NTA), and proteomic analysis7,19. We found greater advantages of EVGpro over UC. TEM further validated the conventional cup-shaped morphology of seminal plasma EVs captured using EVGpro, with sizes consistent within the expected range for EVs (Fig. 2B). WB analysis of EV markers showed higher levels of TSG101, CD9, and CD81 in EVGpro-enriched EVs compared with those using UC. We also observed higher levels of APOE, an EV corona protein36, in EVGpro-enriched EVs. Meanwhile, we analyzed ACPP, an abundant soluble seminal plasma protein, which was lower in EVGpro-enriched EVs, indicating reduced contamination of soluble seminal plasma proteins37 (Supplementary Fig. 3C). NTA analysis showed comparable size distributions between EVs isolated by EVGpro and UC, yet EVGpro enriched 1.45 times more EV particles per equal sample volume and achieved 2.91 times greater purity (18.97 × 1012 particles/mg) than that of UC (6.52 × 1012 particles/mg) (Fig. 2C, D). The purity of EVs isolated by UC was comparable to the published studies38,39,40. Furthermore, EVGpro demonstrated higher recovery efficiency, retrieving 85.42% of particles compared to 34.38% using UC’s (Fig. 2E). Additionally, EVGpro utilized a reduced EV enrichment time of approximately 0.5 h compared with 5 h for UC.
Moreover, proteomics analysis of the EVs enriched using EVGpro and UC was performed. The results revealed that the EVGpro method identified 41.3% more peptides and 41.1% more proteins compared to UC (9778 vs. 6921 EV peptides; 1376 vs. 975 EV proteins; Fig. 2F, G, Supplementary Data 1). Nanochains-based EVGpro method achieved the identification of 1,376 EV proteins from one-twentieth of the EV proteins from 50 μL of seminal plasma, surpassing those reported by Poliakov41, who identified 438 EV proteins using UC and size exclusion chromatography (SEC) from 100 μg of exosome-like vesicles. Zhang et al. identified 1517 EV proteins using UC and sucrose density gradient centrifugation from 20 μg EV proteins from seminal plasma42. EVGpro’s efficacy in EV protein identification is further supported by lower P-values of exosome term enrichment (Supplementary Fig. 3D). In conclusion, EVGpro has emerged as a superior method for efficient, high-yield, and high-purity enrichment of EVs, offering substantial advancements for EV research and clinical applications43.
Optimization of nanochains-based EVGpro method for enrichment of N-glycopeptides
N-glycopeptides comprised only 3-5% of the peptides in the enzymatic hydrolysate, with their detection substantially hampered by abundant non-N-glycopeptides in mass spectrometry analysis. To optimize the N-glycoproteomic analysis of EV proteins, different loading buffer conditions for N-glycopeptide enrichment were evaluated. The concentration of organic solvents in the loading buffer can impact their retention on HILIC and thus influence N-glycopeptide enrichment. Trifluoroacetic acid (TFA), an ion-pairing reagent, mitigates hydrophilic and non-specific electrostatic interactions of non-N-glycopeptides on adsorbents without affecting those of hydrophilic N-glycopeptides.
Various concentrations of acetonitrile (ACN) in the loading buffer were tested, ranging from 60% to 90% (Fig. 3A, Supplementary Data 2). Results indicated that 85% ACN yielded optimal conditions, identifying 1,871 N-glycopeptides corresponding to 414 N-glycoproteins. The percentage was 36.41% for N-glycopeptides and 75.90% for N-glycoproteins. However, ACN concentrations above 85% adversely affected the analysis due to non-N-glycopeptide interference. Using a higher TFA concentration (1%) maximized N-glycoprotein identification, possibly by promoting more ion pairs in the loading buffer (2,158 N-glycopeptides from 483 N-glycoproteins, Fig. 3B, Supplementary Data 2). In conclusion, enriching EV N-glycopeptides with a loading buffer containing 85% ACN and 1% TFA yielded superior N-glycoproteomic analysis results using EVGpro nanochains.
A, B The performances of different ACN (A) and TFA (B) concentrations in the loading buffer of the EVGpro method for EV N-glycopeptides and N-glycoproteins identification. (means ± SD, n = 3 independent experiments). C–E The number of EV N-glycopeptides and N-glycoproteins (C) (means ± SD, n = 3 independent experiments), the overlap (D), and intensity ranking (E) of EV N-glycoproteins identified by EVGpro (blue) and UC (pink) method. F Comparison of the N-glycoproteins identified using the EVGpro method (pink) in our study with Yang et al.’s data (LTQ Orbitrap Velos) (blue)60. ACN, acetonitrile, TFA, trifluoroacetic acid. Source data are provided as a Source Data file.
The performance of the EVGpro method in identifying EV N-glycopeptides was evaluated by comparing it to the classical UC method. Our findings revealed that the EVGpro identified 254.93% more EV N-glycopeptides and 169.83% more EV N-glycoproteins than UC (2158 vs. 608 N-glycopeptides; 483 vs. 179 N-glycoproteins; Fig. 3C, Supplementary Data 2). Moreover, EVGpro encompassed 77.03% of the EV N-glycoproteins identified by UC (Fig. 3D). Importantly, identifying low-abundance proteins is crucial for comprehensively understanding biological sample compositions. EVGpro enrichment identified more low-abundance EV N-glycoproteins (Fig. 3E). Additionally, 341 proteins were identified as N-glycoproteins in human seminal plasma compared to those reported in published literature (Fig. 3F).
Nanochains-based EVGpro method performs well on microliters of diverse biofluids
Diverse biofluids, such as plasma, urine, saliva, PE (pleural effusion), CSF (cerebrospinal fluid), ascites, and tears, contain EVs that are amenable to enrichment using UC. The EVGpro nanochains method was systematically compared with UC across various biofluids, and the performances in EV enrichment were evaluated through NTA, proteomic, and N-glycoproteomic analysis. Plasma, known for its biomarker potential, often requires substantial volumes for effective EV enrichment due to its abundance of high-abundance proteins3. Our findings demonstrated that from 10 μL of plasma, the EVGpro method enriches EV particles 2.86-fold more than UC (14 × 1010 particles vs. 4.9 × 1010 particles for EVGpro vs. UC, Fig. 4A) with similar size distributions (50–200 nm, Supplementary Fig. 4A). Enzymatic digestion of EVs from 10 μL of plasma enabled sequentially proteomic and N-glycoproteomic analyses, identifying 543 EV proteins and 1480 EV N-glycopeptides (224 N-glycoproteins), respectively, surpassing UC by 1.57- and 36.10- (10.67-) fold (543 vs. 346, 1,480 vs. 41, 224 vs. 21, Fig. 4B, C, Supplementary Data 3).
A NTA of EV particles from plasma, urine, saliva, PE, CSF, ascites, and tears enriched by EVGpro (blue) and UC (pink), respectively (means ± SD, n = 3 biological replicates). B, C The number of EV proteins (B) and EV N-glycoproteins (C) identified by UC (pink and green) and EVGpro (blue and lavender) from the same amounts of biofluids (10 μL of plasma, 200 μL of urine, 50 μL of saliva, 50 μL of PE, 200 μL of CSF, 50 μL of ascites, and 5 μL of tear). 5% of the EV protein digests were used for EV proteomic analysis and the remaining was utilized for N-glycopeptides enrichment. (means ± SD, n = 2 independent experiments). D–F The overlap among identified EV proteins from plasma (D), CSF (E), and tear (F) by EVGpro (pink) and those in the literature (blue and green)44,45,94,95. PE, pleural effusion. Source data are provided as a Source Data file.
Plasma-derived EV proteins and N-glycopeptides have also been characterized by various methods in previous studies. For example, Qu et al. identified 208 EV proteins from 0.9 mL plasma using SEC coupled with a minute™ hi-efficiency exosome isolation kit44. This study reported fewer proteins compared to our findings from just 10 μL of plasma using the EVGpro method (Fig. 4D). The -log10(P-value) for exosomes identified by EVGpro was 4.95 times higher than those identified by SEC coupled with exosome isolation kit, respectively44. Furthermore, our EVGpro method also identified higher number of N-glycopeptides, which surpassed other published methods, including the exosome isolation kit coupled with poly-(NIPAM-co-hydrazide) material (283 N-glycopeptides, 161 N-glycoproteins from 200 μL)27 and UC with hydrazide magnetic beads (1336 N-glycopeptides, 525 N-glycoproteins from 5.5 mL)9 (Supplementary Fig. 5A). These findings underscored the efficiency of the EVGpro method in enriching EVs and identifying proteins and N-glycopeptides from low volumes of complex biofluids, highlighted its remarkable advantages and potential for advancing clinical applications.
Biofluids like CSF are characterized by limited volume and lower concentrations of EV particles7,19. Using 200 μL of CSF, the EVGpro nanochains enriched EV particles 1.94 times more than UC (6 × 1010 particles vs. 3.1 × 1010 particles, Fig. 4A, Supplementary Fig. 4B). EVGpro also identified 2154 proteins, 2330 N-glycopeptides, and 604 N-glycoproteins, which represent 2.35-, 17.13-, and 10.60-fold increases over UC, respectively (2154 vs. 916, 2330 vs. 136, 604 vs. 57, Fig. 4B, C, Supplementary Data 4). In CSF, the EVGpro method provided 3.85–16.38 times increase in the number of identified EV proteins compared to conventional methods (Fig. 4E). Meanwhile, using 5 μL of tear, EVGpro identified 2113 proteins, 1952 N-glycopeptides, and 493 N-glycoproteins, which represent 2.16-, 38.27-, and 17-fold increases over UC, respectively (2113 vs. 976, 1952 vs. 51, 493 vs. 29, Fig. 4B, C, F, Supplementary Data 5). EVGpro also yielded 1.06 times more proteins than that enriched by the EXODUS device (1992 proteins from 30 μg EV proteins)45. Furthermore, EVGpro demonstrated superior capability in EV enrichment and N-glycopeptide identification across other biofluids such as urine, saliva, PE, and ascites compared to the UC method. NTA results indicated that EVGpro yielded 1.42-1.96 times more EV particles across various biofluids (urine: 1.96 times, saliva: 1.42 times, PE: 1.79 times, ascites: 1.63 times) (Fig. 4A, Supplementary Fig. 4D–G). Using as little as 200 μL of urine, EVGpro identified higher numbers of proteins, N-glycopeptides, and N-glycoproteins compared to UC (proteins: 1920 vs. 772, N-glycopeptides: 2202 vs. 120, N-glycoproteins: 500 vs. 63, Supplementary Data 6). Additionally, EVGpro outperformed UC with as little as 50 μL each of saliva, PE, and ascites (Fig. 4B, C, Supplementary Data 7–9). These results underscored the versatility and robustness of EVGpro across diverse biological fluids, positioning it as a pivotal tool for advancing research and clinical applications in EV biology.
In urine, our results surpassed other methods such as UC (1,917 proteins) and SEC method (1159 proteins)46 (Supplementary Fig. 5B). In saliva, the EVGpro method provided a 171%–673% increase in the number of identified EV proteins compared to conventional methods47,48 (Supplementary Fig. 5C). Similarly, EVGpro yielded 14.76 times more N-glycopeptides and 8.2 times more N-glycoproteins than the Fe3O4@TiO2@Cys material from urine (134 N-glycopeptides, 64 N-glycoproteins)25 (Supplementary Fig. 5D). These findings underscored the effectiveness of the developed EVGpro enrichment method for EV N-glycopeptides across common clinical biofluids, showcasing improvements over conventional methods and existing literature reports.
EVGpro nanochains enable proteomic and N-glycoproteomic of EVs from ultratrace of biofluids
Integrating data-independent acquisition (DIA) into proteomic and N-glycoproteome analysis enhances protein coverage, quantitative accuracy, and reproducibility compared to data-dependent acquisition (DDA)-based approaches49. Despite these advantages, DIA N-glycopeptide methods have faced considerable challenges, and there have been no advancements in the study of EV N-glycopeptides using the DIA approach49. To quantify proteins and N-glycopeptides using DIA, spectral libraries were generated that included 26,534 peptides from 3487 proteins and 6037 N-glycopeptides belonging to 1281 N-glycoproteins. A comparative analysis of library-DIA, direct-DIA, and DDA methods for EV proteomics and N-glycoproteomics was conducted.
The results demonstrated the superiority of the library-DIA method over others (1869 vs. 1612 and 1088 for proteins, 4339 vs. 2627 and 1963 for N-glycopeptides, 621 vs. 508 and 391 for N-glycoproteins) (Fig. 5A, Supplementary Fig. 6A, Supplementary Data 10). The library-DIA method identified 89.35% of proteins and 92.86% of N-glycoproteins identified by DDA (Fig. 5B, Supplementary Fig. 6B). This enhancement in peptide and N-glycopeptide identification was attributed to DIA’s utilization of a broader range of precursor ions in MS mode, resulting in increased dynamic range and higher quantitative sensitivity50. Importantly, this study represented the application of the DIA approach in EV research for investigating glycosylation.
A, B The number of identified N-glycoproteins (A) (means ± SD, n = 3 independent experiments) and their overlap (B) in seminal plasma EV using DDA (pink), direct-DIA (blue), and library-DIA (green). C, D The number of proteins (C) and N-glycopeptides (D) identified by EVGpro from 1 μL seminal plasma using library-DIA method based on magnetic stirrer (blue) and 3D-shaking rotator (pink), respectively (means ± SD, n = 3 independent experiments). E, F The number of proteins (E) and N-glycopeptides (F) identified by EVGpro from 1 μL tear using the DIA method (means ± SD, n = 3 independent experiments). DDA, data-dependent acquisition; DIA, data-independent acquisition. Source data are provided as a Source Data file.
For precious samples from long-term cohorts, the performance of the EVGpro nanochains was evaluated on the ultratrace of biofluid by enriching EVs from 1 μL of seminal plasma and subsequently identifying EV proteins or N-glycopeptides. Under an alternating magnetic field, the magnetic nanochains rotate to generate eddy currents or laminar currents to homogenize ultratrace samples, and the hydrogel shell of EVGpro nanochains gave better dispersion and suspension to counteract potential sedimentation51. We used the magnetic stirring-based homogenization method, and found that the magnetic stirring-based EVGpro method identified 1.23, 1.33, and 1.16 times more proteins, N-glycopeptides, and N-glycoproteins, than 3D-shaking-based homogenization method, respectively (1620 vs. 1320, 2207 vs. 1658, 526 vs. 453) (Fig. 5C, D, Supplementary Data 11). What’s more, the EVGpro method also identified 1965 proteins and 1502 N-glycopeptides (398 N-glycoproteins) from 1 μL of tear, respectively (Fig. 5E, F, Supplementary Data 11). It is possible that exogenous magnetic stirring can enhance the efficiency of EV capture using the EVGpro method. However, reducing the biofluid volume to 1 μL posed the primary challenge in achieving effective homogenization and dispersion of magnetic nanochains, as standard shaking methods were inadequate due to poor dispersion. The EVGpro protocol is expected to be suitable for ultra-low amounts of biofluids, providing substantial depth in identifying EV proteins and N-glycopeptides.
Nanochains-based EVGpro method identifies N-glycoprotein signatures of AZS
Over the past 45 years, total sperm count has decreased by 62.3%, with the rate of decline accelerating52. AZS represents the most prevalent type of male infertility, characterized by multifaceted etiologies that necessitate further investigation28,31,53,54. Previous studies have reported abnormalities in EV proteins in seminal plasma from AZS patients, yet the alterations in EV N-glycoproteins remain uncharacterized12. This study enriched EV proteins and N-glycopeptides from seminal plasma samples of 20 AZS and 20 normal sperm (NS) individuals (discovery cohort) according to the World Health Organization 2010 guidelines using the EVGpro method55,56,57,58. Subsequently, quantitative proteomics and N-glycoproteomics analyses were performed using DIA (Fig. 6A). The basic information of our research cohort, including age, progressive motility, and total motility, was shown in Supplementary Data 12.
A Overview of the strategy for analyzing the seminal plasma EV proteome and N-glycoproteome of AZS patients and controls with NS using EVGpro nanochains. B PCA of seminal plasma EV N-glycopeptides in the discovery cohort (NS = 20 (rose red), AZS = 20 (lake blue)). C Volcano plot displays differential expression of seminal plasma EV N-glycopeptides (NS = 20, AZS = 20, FC > 1.5 or FC < 1/1.5 and P-value < 0.05, two-sided independent Student’s t-test). X-axis: log2 FC (AZS/NS ratio), y-axis: -log10 P-value. Up (scarlet), down (lime green), notsig (sky blue). D Heatmap displayed differential expression of seminal plasma EV N-glycopeptides (NS = 20, AZS = 20, FC > 1.5 or FC < 1/1.5 and P-value < 0.05, two-sided independent Student’s t-test). The color range, from red ( + 4) to blue (-4), indicates the row z-score of normalized N-glycopeptides expression levels. E Dot plot of the log2 (FC) of differentially expressed EV N-glycoproteins and that of corresponding proteins. Line: navy blue, up-up: red, down-up: green, down-down: cyan, up-down: purple, center: coral. F, G GO (F) and KEGG pathway (G) terms enriched in downregulated N-glycoproteins in AZS relative to NS. H The SHAP values for the two different features driving the predictions of the machine learning classifier in the validation cohort (NS = 34, AZS = 34). I The optimal number of protein features to distinguish AZS and NS. J ROC curves to distinguish AZS from NS in the verification cohort. K, L Abundance distributions of the two differentially expressed N-glycoproteins, PRELP (K) and DPP4 (L), in the verification cohort (NS = 34 (sky blue), AZS = 34 (scarlet)), with error bars representing the median and interquartile range. The P-value was calculated using two-sided independent Student’s t-test. AZS asthenozoospermia, NS normal sperm, PRM parallel reaction monitoring, PC1 principal component 1, FC fold change, BP biological process, CC cellular component, MF molecular function, SHAP SHapley Additive exPlanations, AUC area under the curve. Created in BioRender. (2025) https://BioRender.com/lq01r4m. Source data are provided as a Source Data file.
The results revealed the quantification of 2475 EV proteins, which distinguished AZS from NS samples based on principal component analysis (PCA) visualization (Supplementary Fig. 7A, Supplementary Data 13). Among these proteins, 287 exhibited differential expression, with 137 down-regulated (fold change (FC) < 1/1.5, P-value < 0.05) and 150 up-regulated (FC > 1.5, P-value < 0.05) in AZS (Supplementary Fig. 7B, C). For EV N-glycopeptides in the discovery cohort, a total of 5807 EV N-glycopeptides were identified, corresponding to 765 distinct N-glycoproteins (Supplementary Data 13). Most of the N-glycoproteins (183/196, 93.37%) were reported by both Xin et al.’s and Yang et al.’s studies. A total of 369 N-glycoproteins were identified, which had not been reported to be glycosylated in human seminal plasma59,60 (Supplementary Fig. 7D). Among these N-glycopeptides, 472 showed differential expression, with 196 upregulated (FC > 1.5, P-value < 0.05) and 276 downregulated (FC < 1/1.5, P-value < 0.05) (Fig. 6B–D).
To characterize the features of EV protein N-glycosylation in the seminal plasma of AZS, changes in the EV protein expression and N-glycosylation levels were compared. It was found that only 20 proteins were differentially expressed at both the protein and N-glycosylation levels between AZS and NS groups (Supplementary Fig. 7E, Supplementary Data 13). Most of the proteins differentially regulated at N-glycosylation levels were unchanged at their protein level (70/103, 67.96%) (Fig. 6E), indicating that N-glycosylation analysis offers a distinct dimension for resolving AZS. Further Gene Ontology (GO) analysis showed enrichment in terms of cell adhesion, membrane protein intracellular domain proteolysis, and polysaccharide biosynthetic process. Additionally, N-glycoproteins exhibited diverse enzymatic activities, including maltose alpha-glucosidase, aminopeptidase, and metallocarboxypeptidase activities (Fig. 6F). According to Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway annotation, EV proteins with down-regulated glycosylation levels in AZS were primarily associated with efferocytosis, ECM-receptor interaction, and renin-angiotensin system (Fig. 6G).
To identify N-glycosylation signatures related to AZS, we analyzed receiver operating characteristic (ROC) curves of EV N-glycopeptides, revealing 452 N-glycopeptides with an area under the curve (AUC) > 0.7 and high discriminatory power (Supplementary Data 13). Additionally, tissue expression distribution analysis using the Human Protein Atlas showed that all proteins differentially regulated at the glycosylation level in AZS were expressed in the male reproductive system. Notably, six were testis-specific (SLIT1, PLAUR, IZUMO4, ACRBP, ZPBP, and SPACA1), three epididymis-specific (SCGB2A1, SPINT4, and ELSPBP1), one seminal vesicle-specific (CRISP3), and two prostate-specific (TGM4 and ANO7). Among these, testis-specific N-glycoproteins ACRBP achieved an AUC of 0.99, while epididymis-specific N-glycoproteins ELSPBP1 achieved an AUC of 0.95, effectively distinguishing AZS from NS in the discovery cohort (Supplementary Data 13).
EV N-glycoprotein signatures were validated using targeted N-glycoproteomics via parallel reaction monitoring and parallel accumulation serial fragmentation (PRM-PASEF) on an independent validation cohort of 34 AZS and 34 NS19. Notably, five N-glycoproteins (PRELP, DPP4, MUC6, BGN, and MSLN) demonstrated an AUC exceeding 0.81 in distinguishing AZS from NS in the validation cohort. SHapley Additive exPlanations (SHAP) analysis was performed to rank the importance of these differentially expressed N-glycoproteins identified by PRM. The SHAP values, which reflect the impact of each N-glycoprotein on the model’s output, ranked in order of importance as PRELP and DPP4 (Fig. 6H). The performance of various N-glycoprotein combinations was assessed by calculating the mean AUC for different feature numbers (Fig. 6I). The analysis revealed that the mean AUC reached its maximum with two features, specifically PRELP and DPP4. Notably, this optimal combination achieved a AUC of 1.0 in the validation cohort Fig. 6J–L).
Nanochains-based EVGpro method identifies subtypes of AZS according to N-glycoprotein signatures
AZS is prevalent among male infertility cases, with its etiologies remaining poorly understood61. Although EVs from AZS seminal plasma could affect sperm motility, investigations into changes in their molecular subtypes are still lacking12. Our above N-glycoproteomic analysis of EV in seminal plasma showed distinct signatures associated with AZS, prompting further investigation into potential AZS subtypes at the EV protein glycosylation level. Consensus clustering analysis of the discovery cohort identified two molecular subtypes: AZS-C1 and AZS-C2, characterized by differential N-glycoprotein levels (Fig. 7A). This two-subtype classification was further validated by PCA, clearly segregating AZS-C1 and AZS-C2 samples into two clusters (Fig. 7B). Analysis of EV proteins differentially regulated at the glycosylation level between AZS-C1 and AZS-C2 revealed 12 N-glycoproteins enriched in AZS-C1 and 17 in AZS-C2 (FC > 1.2 or FC < 1/1.2, P-value < 0.05, Supplementary Data 13, Fig. 7C). Among these, all (29/29) of N-glycoproteins had AUC values exceeding 0.7 in the discovery cohort. These findings demonstrated effective AZS classification into two molecular subtypes based on these EV N-glycoproteins. SHAP analysis was conducted on the above 29 N-glycoproteins identified in the discovery cohort, yielding importance rankings for distinguishing AZS subtypes. The top-ranking ten N-glycoproteins were CLIC1, RAB4B, PCDH7, GRN, ANPEP, TSPAN6, MME, GDF11, NPC2, and PRELP. Then, the optimal combination of ten N-glycoproteins achieved a perfect AUC value of 1.0 (Fig. 7D–G).
A Consensus clustering analysis categorized AZS samples in the discovery cohort into two subtypes: AZS-C1 (n = 9) and AZS-C2 (n = 11). B Distribution of samples from AZS subtypes, AZS-C1 (n = 9) (red) and AZS-C2 (n = 11) (blue), in the PCA plot of the discovery cohort. C Heatmap displays differential expression of EV N-glycoproteins between AZS subtypes AZS-C1 and AZS-C2 (P-value < 0.05, two-sided independent Student’s t-test). The color range, from red ( + 2) to blue (-2), indicates the row z-score of normalized N-glycoproteins expression levels. D SHAP values for the ten important features (N-glycoproteins) distinguishing AZS-C1 (n = 9) and AZS-C2 (n = 11) subtypes in the discovery cohort. E The ROC curves of the ten important N-glycoproteins used to distinguish AZS subtypes, AZS-C1 (n = 9) and AZS-C2 (n = 11), in the discovery cohort. F, G Abundance distributions of the N-glycoproteins RAB4B (F) and CXADR (G) in the discovery cohort. Error bar, median with interquartile range. AZS-C1 (n = 9) (sky blue), AZS-C2 (n = 11) (scarlet). The P-value was calculated using two-sided independent Student’s t-test. H AZS samples in the verification cohort were also categorized into two subtypes AZS-C1 and AZS-C2 by K-means clustering analysis. (I) The ROC curves to distinguish AZS-C1 (red) and AZS-C2 (cyan) in the verification cohort. (J) Abundance distributions of N-glycoproteins PLA1A in the verification cohort. AZS-C1 (n = 12) (sky blue), AZS-C2 (n = 22) (scarlet). The P-value was calculated using two-sided independent Student’s t-test. Error bar, median with interquartile range. Source data are provided as a Source Data file.
To confirm the existence of subtypes of AZS distinguished by N-glycopeptides, an additional analysis was performed using a validation cohort of 68 samples with PRM-PASEF-based targeted N-glycoproteomics. Clustering analysis confirmed the presence of two distinct AZS subtypes, AZS-C1 and AZS-C2 (Fig. 7H). Combining DPP4, PLA1A, and PRELP proteins achieved an AUC of 1.0, completely differentiating AZS-C1 from AZS-C2 (Fig. 7I, J). These findings, combined with results from the discovery cohort, underscore the effectiveness of EV N-glycoproteins in classifying AZS into distinct subtypes and provide a robust foundation for future research into the role of protein glycosylation in AZS pathogenesis. This also highlights the potential of clinical proteomics in promoting early diagnosis and tailoring treatment strategies for patients.
Discussion
EV N-glycoproteins hold substantial potential for disease diagnosis and prognosis2,9,11,17,18. However, their study has been constrained by the challenges associated with the tandem enrichment of EVs and N-glycopeptides from biofluids. Additionally, their profiles in conditions such as AZS remain underexplored. In this study, we developed the EVGpro method—a highly sensitive and advanced approach designed to sequentially enrich EVs and N-glycopeptides within a single tube. This method facilitated multi-omic analysis of EV protein expression and N-glycosylation from microliters of biofluids. The application of EVGpro to the N-glycoproteomic analysis of seminal plasma EVs identified two molecular subtypes of AZS in male fertility.
Using nanochains coated with PEG2000-cell penetrating peptide (octa-arginine, R8) for the enrichment of EVs and their N-glycopeptides, EVGpro nanochains enable sequential proteomic and N-glycoproteomic analysis from ultratrace of biofluids. Magnetic beads, commonly used for EV enrichment and proteomics in plasma, urine, CSF, saliva, etc19,20,21, often suffer from slower separation due to weak magnetic forces from small particles and can be lost during washing and tube transfer. In contrast, our nanochains achieved faster magnetic separation and higher recovery during enrichment. This enhanced efficiency allows for effective analysis of EVs and N-glycopeptides from the ultratrace of samples. The EVGpro method was expected to be suitable for automation in large-scale proteomic or N-glycoproteomic studies.
Our EVGpro approach has proven robustness across various biofluids, including seminal plasma, plasma, urine, saliva, PE, CSF, ascites, and tears. It outperforms the classical UC method and sets a benchmark compared to the existing literature3,9,27,43,44. 1965 EV proteins, 1502 EV N-glycopeptides, and 398 EV N-glycoproteins were identified from just 1 μL tear, respectively. The EVGpro method enables in-depth EV proteomic and N-glycoproteomic analysis on ultratrace and precious clinical samples. Previous methods were generally tested on single biofluid, and N-glycoproteomics analysis of EV proteins has been seldom explored7,25,27. Our comprehensive testing across diverse biofluids demonstrated that EVGpro is a universal method for proteomic and N-glycoproteomics analysis and is anticipated to facilitate identify potential biomarkers in various pathological conditions. This universality provides a more convenient and effective approach to clinical proteomics. Furthermore, intact N-glycopeptides can be flourished from the EVGpro enrichment, enabling the analysis of EV N-glycopeptides with intact glycans. Glycans are known important biomarker sources62,63. In the future, the EVGpro platform can be adapted to explore site-specific glycans to decipher N-glycoprotein microheterogeneity and discover more disease-specific biomarkers.
AZS is a prevalent type of male infertility, yet its characterization remains limited due to multifaceted etiologies4,29. Seminal plasma EVs have been reported to be related to sperm viability, maturation, and function4,12,64, and might provide valuable insights into the pathophysiology. Using EVGpro, a total of 5807 EV N-glycopeptides corresponding to 765 distinct N-glycoproteins were quantified in seminal plasma in the discovery cohort. Validation in an independent cohort of 34 AZS and 34 control samples revealed five N-glycoproteins with AUCs exceeding 0.81, including PRELP, DPP4, MUC6, BGN, and MSLN. MUC6 is a glycosylated secretory mucin mainly detected in male seminal vesicles65. BGN modulates FGFR3c signaling, which is involved in spermatogonia regulation66. MSLN can regulate the G2-M transition of the cell cycle and ensure proper meiotic progression during spermatogenesis67.
We identified two AZS molecular subtypes, AZS-C1 and AZS-C2, based on distinct EV N-glycopeptide profiles validated through PRM-PASEF and consensus clustering. N-glycopeptides such as DPP4, PRELP, and PLA1A showed high AUC values, supporting this classification. Although AZS presents with diverse underlying causes and treatment, subtype classification has not previously been reported. Our identification of these subtypes in AZS may help elucidate the molecular mechanisms underlying AZS and aid in developing personalized treatment strategies. Among the N-glycopeptides differentiating subtypes of AZS, DPP4 was a prostasome-bound protein transferred to spermatozoa through fusion and present in seminal plasma20,41,42,43,68,69. The elevated expression of DPP4 correlates positively with reduced sperm velocity in the AZS group70. Furthermore, DPP4 was highly expressed in the AZS-C1 subtype, which was confirmed in the validation cohort. The EV N-glycopeptide signatures in seminal plasma can help uncover two molecular subtypes of AZS and may contribute to subtype analysis across various diseases.
In summary, we developed a sensitive EVGpro-based method that enables the tandem enrichment of EV proteins and EV N-glycoproteins. This method is universally applicable to various tissue fluids and effective with ultratrace volumes of samples, outperforming the classical UC method and existing literature reports. Beyond identifying characteristic EV N-glycoproteins associated with AZS, the EVGpro-based method classified AZS—a prevalent cause of male infertility—into two distinct subtypes. It may improve biomarker identification, elucidate disease subtypes, aid in characterizing molecular mechanisms, and contribute to developing personalized medicine strategies.
Methods
Ethical statement
Approval for the study was obtained from the ethical committees of Nanjing Medical University (Permission Number: 2023-541) and the Sir Run Run Hospital of Nanjing Medical University (Permission Number: 2023-SR-023). Written informed consent was secured from all participants, between 18 and 40 years of age.
For the seminal plasma, we focused solely on male patients because seminal plasma occurs only in males. The other biofluids were a mix of male and female donors, and did not include a comparison of sexes. Seminal plasma samples were collected from healthy donors categorized as having normal sperm (NS) and individuals with asthenozoospermia (AZS). NS seminal plasma samples were obtained from healthy semen donors with sperm concentration of at least 15 × 106/mL, total motility of at least 40%, and progressive motility of at least 32%, while the AZS seminal plasma samples were from donors having sperm count more than 15 × 106/mL but less than 32% forward-moving sperm in the semen analysis according to the 2010 guidelines of the World Health Organization55,56,57,58,71. All individuals in the NS group had proven fertility as having fathered at least one child, while the AZS patients had failed to achieve pregnancy after 12 months of regular unprotected intercourse. The discovery cohort included 20 NS and 20 AZS samples, which resulted in 91.53% of differentially expressed N-glycopeptides exhibiting Cohen’s d ≥ 1.0 and 99.98% power to detect group differences, exceeding the standard exploratory research threshold of 80% power72. Additionally, 34 NS and 34 AZS samples were further analyzed using targeted quantification via parallel reaction monitoring (PRM).
The remaining seven biofluids were all collected from ten donors: plasma, urine, saliva, and tear samples from healthy donors, while pleural effusion (PE) and ascites were from patients with cirrhosis and fluid accumulation. Cerebrospinal fluid (CSF) samples were from individuals requiring diagnostic evaluation of CSF (30–70 years of age). All samples were centrifuged at 3000 g for 15 min. Aliquots of each seminal plasma sample from the discovery cohort were pooled, and samples of different biofluids were pooled by biofluid type to reduce individual variability during the testing and development of the EVGpro method. Clinical characteristics are summarized in Supplementary Data 12.
Chemicals and reagents
Polyvinylpyrrolidone (PVP K-30) was purchased from Beijing Boao Tuoda Technology Co., Ltd (Beijing, China). Tannins, sodium acetate (NaAC), acrylic acid (AA), ethylene dimethacrylate (EGDMA), 4-(2-Hydroxyethyl) piperazine-1-ethanesulfonic acid (HEPES), 2-(N-Morpholino) ethanesulfonic acid (MES), Tris (2-carboxyethyl) phosphine (TCEP), 2-chloroacetamide (CAA), triethylammonium bicarbonate (TEAB), Tris-HCl, trifluoroacetic acid (TFA), acetonitrile (ACN), ammonium bicarbonate, triethylamine, ammonium formate, urea, sodium chloride (NaCl), Tris(hydroxymethyl)aminomethane (Tris), β-glycerophosphate, sodium fluoride (NaF), and formic acid (FA) were all provided by Sigma-Aldrich (Shanghai, China). PAGE gel rapid preparation kit was purchased from Shanghai Epizyme Biotech Co., Ltd. EDTA-free protease inhibitor was obtained from Thermo Fisher Scientific. Iron(III) chloride hexahydrate (FeCl3·6H2O), ethylene glycol (EG), 1-ethyl-3-(3′-dimethylaminopropyl) carbodiimide (EDC), N-hydroxysuccinimide (NHS), 4-nonylphenyl-polyethylene glycol (NP-40), and n-dodecyl-β-D-maltopyranoside (DDM) were bought from Shanghai Aladdin Biochemical Technology Co., Ltd (Shanghai, China). T-octylphenoxypolyethoxyethanol (tritonX-100) was supplied by Shanghai (China) Biotechnology Engineering Co., Ltd. PBS (Gibco), 2-hydroxy-2-methylpropiophenone (HMPP) obtained from Shanghai Bide Pharmaceutical Technology Co., Ltd (Shanghai, China). NH2-PEG1000-SH was bought from Chongqing Yusi Pharmaceutical Technology Co., Ltd (Chongqing, China). Maleimide-PEG2000-R8 (MAL-PEG2000-R8) was synthesized by Ruixi Co., Ltd (Shanxi, China).
Molecular docking
The AutoDock Vina (version 1.1.2) was utilized to construct the binding mode between R8 and sphingolipid molecules73. Before docking, the charge state and the lowest energy conformation of the compound were calculated via the LigPrep module in Maestro 13.1 software. A docking box (20 Å × 20 Å × 20 Å) was set around the headgroup of the sphingolipid. The above molecules were converted to PDBQT format via ADFRsuite 1.03 for AutoDock Vina docking. During docking, the global search exhaustiveness was set at 32 with other parameters default. The highest-scoring pose (-2.6 kcal/mol) was selected for molecular dynamics validation. Finally, PyMol 2.5.4 was employed for the visual analysis of the docking results.
Molecular dynamics simulation
Three parallel all-atom molecular dynamics simulations were conducted using the AMBER 22 software package, with initial structures derived from the molecular docking pose of R8 and sphingolipid generated by AutoDock Vina (version 1.1.2) where the highest-scoring pose (-2.6 kcal/mol) was selected74. The sphingolipid was parameterized using the GAFF2 force field, and R8 was described by the ff14SB protein force field75,76. Protonation states were assigned, and hydrogen atoms were added via the LEaP module77. The system was solvated in a 10-Å truncated octahedral box containing TIP3P water molecules, with Na+/Cl− ions added to neutralize the charge (8 Cl− ions for charge balance) (Supplementary Table 1).
Energy minimization was performed in two stages: 2500 steps of the steepest-descent method followed by 2500 steps of the conjugate-gradient method, reducing residual forces to below 0.05 kcal/mol/Å. The system was then heated from 0 K to 298.15 K over 200 ps under constant volume (NVT ensemble) to equilibrate solvent molecules, followed by a 500-ps NVT simulation to ensure uniform solvation. A subsequent 500-ps NPT (constant-pressure, constant-temperature) simulation at 1 atm and 298.15 K was performed to equilibrate the system density, after which a 200-ns production run under NPT conditions was conducted. Bond lengths involving hydrogen atoms were constrained using the SHAKE algorithm, and long-range electrostatic interactions were computed via the particle-mesh Ewald (PME) method with a 10-Å nonbonded cutoff. The Langevin thermostat (collision frequency = 2 ps−1) and Parrinello-Rahman barostat were used to maintain temperature and pressure, respectively, with an integration time step of 2 fs78,79,80. Trajectories were saved every 10 ps, and data from the equilibrated phase (150–200 ns) were analyzed for structural and energetic properties. To assess the structural fluctuations of R8 throughout the simulation, root mean square fluctuations (RMSF) analysis was performed using the Cα atoms of the R8 molecule, with data extracted from the equilibrated phase of 150–200 ns timeframe of simulation. The simulation input/output files are available at https://zenodo.org/uploads/15898317. The binding free energy between the sphingolipid and R8 in all systems was calculated using the Molecular Mechanics/Generalized Born Surface Area (MM/GBSA) method during the 150–200 ns simulation period81,82,83. The binding free energy was given by
ΔGbind was binding free energy, ΔEvdW was van der Waals energy, ΔGelec-solv was electrostatic contribution to solvation, which was computed via the generalized Born model (igb = 2)84. ΔGnonelec-solv was non-polar contribution to solvation, which was calculated as the product of the surface tension (γ) and the solvent-accessible surface area (SASA) via the formula ΔGnonelec-solv = 0.0072 × ΔSASA85. Entropic contributions were calculated based on the Hou et al. study82.
Preparation of EVGpro magnetic nanochains
-
(a)
Fe3O4 synthesis: Fe3O4 was synthesized by combining 8 g of PVP K-30, 0.072 g of tannins, and 30 mL of EG in a three-necked flask and stirred at 80 °C for 20 min. After cooling to room temperature, 0.702 g of FeCl3·6H2O was added and stirred for 30 min. Subsequently, 2.8 g of NaAC was introduced and stirred for 30 min. The resulting mixture was then heated at 200 °C for 12 h in a Teflon-lined stainless steel autoclave. After cooling, the Fe3O4 product was collected using a magnet and washed with ethanol three times.
-
(b)
Fe3O4 magnetic nanochains (Fe3O4-MN) preparation: Fe3O4 along with 0.6 g of AA, 50 mg of EGMDA, and 70 mg of HMPP, were thoroughly mixed. Subsequently, 50 μL of the above mixture, 450 μL of EG, and 450 μL of water were placed in a petri dish and exposed to a magnetic field for 2 min. Following this, ultraviolet light was turned on for 4 min. The resulting Fe3O4-MN was collected with a magnet and cleaned with ethanol and water successively.
-
(c)
Surface functionalization: the carboxyl-derivatized Fe3O4-MN was activated with EDC (20 mg) and NHS (10 mg) for 30 min. Subsequently, 5 mg of NH2-PEG1000-SH was added, and the mixture was rotated for 6 h at room temperature. The resulting Fe3O4-MN@SH was washed three times with water, and was dispersed in 100 mL of 10 mM HEPES buffer (pH = 7.4) containing MAL-PEG2000-R8, and rotated for 30 min. Finally, the EVGpro nanochains were separated using a magnet and washed three times with water.
Functional characterization of magnetic nanochains
The magnetic nanochains were examined using a scanning electron microscope (SEM, Zeiss Ultra Plus, Germany) and a transmission electron microscope (TEM, JEM-2100, Japan)20. The zeta (ζ) potentials of nanochains were recorded by Zetasizer (Malvern). The saturation magnetization curve was measured on a vibrating sample magnetometer (Lakeshore-7404, USA). The Fourier-transformed infrared spectroscopy (FT-IR) characterization has been performed using the Thermo In 10 Fourier transform infrared spectrophotometer.
EVs isolation by ultracentrifugation
To isolate extracellular vesicles (EVs) by ultracentrifugation, seminal plasma from each individual was subjected to rapid thawing at 37 °C, followed by centrifugation at 3000 g for 30 min at 4 °C, and subsequent centrifugation at 20,000 g for 60 min at 4 °C. Supernatants were further centrifuged at 199,700 g for 90 min at 4 °C in an Optima XE Ultracentrifuge (Beckman Coulter). Pellets were resuspended in PBS, followed by another centrifugation at 199,700 g for 90 min at 4 °C11. Pellets were stored at -80 °C for preservation. For other biofluids, EVs were isolated by centrifugation at 100,000 g, following the same procedures. The size of the EVs was measured using ZetaView (Particle Metrix, Meerbusch, Germany)20.
EV isolation and N-glycopeptides enrichment by EVGpro
For seminal plasma EVs isolation using EVGpro, 50 μL of seminal plasma from each individual was mixed with 950 μL of PBS and 100 μL of 0.1% NP40/TritonX-100 in a 1.5-mL tube. EVGpro nanochains were added to the mixture and placed on a rotator at 20 rpm at room temperature for 0.5 h, magnetically separated to discard the supernatant, and wash once with 1 mL 0.01% NP40/Triton X-100 and twice with 1 mL PBS by magnetic separation followed by the supernatant removal. The EVs on EVGpro or EVs from UC were lysed at 95 °C for 10 min in the lysis buffer of 50 mM Tris·HCl (pH = 8.5), 40 mM CAA, 10 mM TCEP, and 0.01% DDM. After cooling to room temperature, the protein solution was diluted with 5-fold TEAB (50 mM) and digested in Lys-C at a 1/100 (w/w) enzyme/protein ratio at 37 °C for 3 h, and in trypsin at 1/50 (w/w) overnight at 37 °C. The enzymatic digestion was terminated by adding 10% TFA to a final concentration of 1%. After magnetic separation, 5% of the digests were used for proteomic analysis, and the remaining was utilized for N-glycopeptides enrichment. For proteomic analysis of the seminal plasma EVs, samples from six individuals were pooled to generate one replicate of the three biological replicates.
In the EVGpro method, N-glycopeptides were enriched in the same tube in hydrophilic interaction liquid chromatography (HILIC) mode. Briefly, the peptide solution was mixed with 100% ACN containing 1% TFA to achieve a final concentration of 85% ACN (1% TFA), incubated for 20 min at 37 °C, washed three times with 300 µL of loading buffer (ACN/H2O/TFA, 85/14/1, v/v/v), eluted with 100 µL of elution buffer (ACN/H2O/TFA, 20/79.7/0.3) for 10 min. Following magnetic separation, the supernatant was collected into a centrifuge tube and lyophilized in a vacuum centrifuge (Labconco CentriVap). The lyophilized N-glycopeptides were dissolved in 200 µL of ammonium bicarbonate buffer (25 mM) containing 0.3 µL of PNGase F (New England BioLabs), incubated at 37 °C for 12 h to remove the N-glycans, and desalted using a C18 cartridge before LC-MS/MS analysis. For other biofluids, the EV proteins and glycopeptide enrichment procedure was also performed as outlined above.
Western blotting
The EVs were lysed in protein lysis buffer (8 M urea, 75 mM NaCl, 50 mM Tris (pH 8.2), 1% EDTA-free protease inhibitor, 1 mM NaF, and 1 mM β-glycerophosphate) for 30 min, followed by centrifugation at 12,000 g for 30 min at 4 °C. The supernatant of the EV lysate was heated at 95 °C for 10 min. After cooling to room temperature, 5 μg EV proteins were separated using SDS/PAGE and transferred onto a polyvinylidene fluoride membrane. The HEK293T cells (ATCC, CRL-3216) were used as a negative control. The primary antibodies used are mouse anti-CD9 (dilution: 1:2000, catalog number: 60232-1-IG, Proteintech), mouse anti-CD81 (dilution: 1:2000, 66866-1-IG, Proteintech), rabbit anti-TSG101 (dilution: 1:2000, 28283-1-AP, Proteintech), rabbit anti-Calnexin (dilution: 1:2000, catalog number: 10427-2-AP, Proteintech), mouse anti-APOE (dilution: 1:5000, catalog number: 83728-4-RR, Proteintech), and mouse anti-ACPP (dilution: 1:3000, catalog number: 60176-1-Ig, Proteintech). The secondary antibodies used are goat anti-mouse IgG (dilution: 1:5000, catalog number: 31430, Thermo Fisher Scientific) and goat anti-rabbit IgG (dilution: 1:5000, catalog number: 31460, Thermo Fisher Scientific). The EVs in UC were from the particles obtained by centrifugation at 20,000 × g.
Nanoparticle tracking analysis (NTA)
The EVGpro with captured EVs underwent three consecutive steps of elution with 200 μL 200 mM triethylamine solution, and was subjected to magnetic separation to collect EVs in the resultant supernatant. The size of EVs from EVGpro or UC was measured using ZetaView (Particle Metrix, Meerbusch, Germany)20. Before use, the ZetaView was calibrated with 100 nm polystyrene particles diluted by 250,000 fold with pure water.
EV recovery rate analysis
To calculate the recovery rate, EVs isolated by UC were measured by NTA, and divided into two equal aliquots. Each aliquot was subjected to EVs capture by UC or EVGpro nanochains. After the capture process, NTA was performed once again to measure the particle number of the recaptured EVs. The recovery rate was calculated as the recaptured EV particles divided by the original EV particles.
Fractionation with high-pH reverse-phase chromatography
To construct spectra library of seminal plasma EVs N-glycoproteomic, two schemes were employed. On one hand, the N-glycopeptides enriched from EVGpro nanochains underwent fractionation using a high-pH reverse-phase column (XBridge BEH130 C18, 300 μm × 150 mm, 1.7 μm, Waters) coupled with the M-class HPLC system (Waters). The mobile phase comprised 20 mM ammonium formate (pH = 10.0) as mobile phase A and 100% ACN with 20 mM ammonium formate (pH = 10.0) as mobile phase B. The separation flow rate was set at 4 μL/min. The elution gradient was programmed as follows: 0-14 min, 3% B; 14–15 min, 3–8% B; 15–39 min, 8–29% B; 39–43 min, 29–41% B; 43–44 min, 41–100% B; 44–52 min, 100% B; 52–53 min, 100–3% B; 53–73 min, 3% B. Fractions were collected using the non-adjacent pooling scheme with 1 min per tube. For example, samples collected at 1 min, 21 min, 41 min, and 61 min were combined to form fraction 1. In total, twenty fractions were obtained and subsequently dried.
On the other hand, desalted EV peptides were fractionated using an Agilent HPLC system coupled with a C18 column (Waters BEH C18, 2.1×150 mm, 1.7 μm). The separation flow rate was 0.4 mL/min. The gradient elution program was as follows: 0% B for 2 min, 0–5% B in 4 min, 5–10% B. Ten fractions were collected using the non-adjacent pooling scheme at a rate of 30 s/tube and dried using vacuum centrifugation. These fractions were then subjected to N-glycopeptide enrichment by adding 85% ACN containing 1% TFA and EVGpro nanochains. A total of thirty fractions generated from the above two schemes were ultimately used for MS analysis to construct the EV N-glycopeptide spectra library. To construct the EV protein spectra library for EV proteomic analysis, ten fractions were collected using the non-adjacent pooling scheme at a rate of 1 min/tube from the pooled EVGpro-enriched and UC-enriched EV peptides using the M-class HPLC system.
3D-shaking and magnetic stirring incubation
For 3D-shaking incubation: 1 μL seminal plasma or tear was combined with 300 μL PBS and 0.1 mg EVGpro in 0.5-mL tube. The mixture was incubated for 0.5 h using a rotator at 20 rpm at room temperature, and separated by a magnetic stand with supernatant discarded. 10 μL lysis buffer was added. Subsequently, lysis, enzymatic digestion, and N-glycopeptide enrichment steps were the same as those for the conventional-volume samples. In brief, the sample was washed three times with 100 μL 85% ACN 1% TFA, followed by 50 μL elution buffer (ACN/H2O/TFA, 20/79.7/0.3, v/v/v) to elute the N-glycopeptides, followed by lyophilization and removal of the N-glycans.
For magnetic stirring incubation: 1 μL seminal plasma or tear was mixed with 30 μL PBS and 0.1 mg EVGpro on a glass slide. The mixture was incubated on magnetic stirrers at 50 rpm at room temperature for 30 min, and magnetic separated using a magnet (900 GS). The 10 μL lysis buffer was added. Subsequently, the mixture was transferred to a 0.5-mL centrifuge tube. The processes of lysis, enzymatic digestion, N-glycopeptide enrichment, washing, elution, and N-glycans removal were performed using the same method as that of the 3D-shaking incubation.
LC-MS/MS analysis
The peptides digested from tear protein samples were analyzed using an Orbitrap Fusion Lumos mass spectrometry system (Thermo Fisher) with the Easy-nLC 1200 (ThermoFisher). Mobile phase A: 0.1% FA, mobile phase B: 80% ACN (0.1%FA). The analytical column (75 μm x 150 mm, 1.7 μm, CoAnn Technologies) was used for peptide separation. The elution gradient was as follows: 0–5 min, 3–5% B; 5–24 min, 5–15% B; 24–45 min, 15–28% B; 45–52.3 min, 28–38% B; 52.3–52.35 min, 38-100% B; 52.35–60 min, 100% B.
For the tear sample, the MS parameters were provided in detail in Supplementary Data 586. For the full scan, a resolution of 120k was used and the normalized automated gain control (AGC) target was 250%. The scan covered a range of 350–1500 m/z. The maximal ion injection time mode was also set to custom, with a maximal ion injection time of 50 ms. Regarding the MS/MS scan, it was carried out in higher-energy collisional dissociation mode at a normalized collision energy (NCE) of 30%. The fragment spectra were detected by an Orbitrap analyzer with a resolution of 30 K and the maximum injection time was set to dynamic.
For the samples other than tear, the EV peptides and N-glycopeptides were subjected to timsTOF Pro mass spectrometer (Bruker) analysis. Homemade packed column (25 cm × 75 µm, 1.9 µm, 100 Å) was used for peptide separation. Mobile phase A: 0.1% FA, mobile phase B: 100% ACN (0.1%FA). Three elution gradient were as follows: (i) 0 –45 min, 5 –22% B; 45 –50 min, 22–40% B; 50–55 min, 40–80% B; 55–60 min, 80% B; (ii) 0–70 min, 5–22% B; 70–80 min, 22–40% B; 80–85 min, 40–80% B; 85–90 min, 80% B; (iii) 0–40 min, 2–22% B; 40–50 min, 22–37% B; 50–55 min, 37–80% B; 55–60 min, 80% B. The samples were all separated at 300 nL/min. For data–dependent acquisition (DDA), MS spectra were acquired in positive ion mode with an m/z range from 100 to 1700 Da, and a resolution set at a ‘target value’ of 40,00019. Collision energy was 20 eV for 0.6 1/K0 and 59 eV for 1.6 1/K0.
To construct EV N-glycopeptide spectra library and EV protein spectra library for DIA analysis using the timsTOF Pro, the N-glycopeptide DDA data obtained from 30 fractions were searched against the UniProt database using Spectronaut (version 18.0) to generate EV N-glycopeptide spectra library. And the peptide DDA data from 10 fractions were used to generate EV protein spectra library. Deamidation (Asn) was selected as a dynamic modification, specifically for N-glycopeptides, while other parameters were maintained as default settings. The obtained results were then imported into the py-diAID software (version 0.0.19) for the optimization of DIA-parallel accumulation serial fragmentation (PASEF) parameters87.
For proteomic analysis of the seminal plasma EV, the m/z range was from 100 to 1700 Da and the cycle time was 3.82 s. Collision energy was 20 eV for 0.6 1/K0 and 59 eV for 1.6 1/K0. The m/z precursor range windows were detailed in Supplementary Data 10. For N-glycoproteomic analysis of the seminal plasma EV, the m/z range was from 100 to 1700 Da and the cycle time was 3.82 s. Collision energy was 20 eV for 0.6 1/K0 and 59 eV for 1.6 1/K0. The m/z precursor range windows were detailed in Supplementary Data 10.
PRM quantification analysis
For targeted N-glycoproteomic quantification, PRM mode was employed19. The liquid-phase PRM method mirrors the 60-min DDA protocol, with the modification of employing PRM-PASEF for mass spectrometry analysis. Differential abundance N-glycoproteins were constructed using Skyline (24.1) for library construction and N-glycopeptide screening. The selected modifications for the PRM included deamidation (N), carboxymethylation, oxidation, and acetylation (N-term). The retention times of the selected glycopeptides varied by no more than 5 min across samples. PRM-PASEF method was employed to identify selected N-glycoproteins, each corresponding to a distinct N-glycopeptide. A resolution of 60 P was selected for ion mobility filtering.
Database searching
The PEAKS Studio X+ workstation (Bioinformatics Solutions Inc.) was utilized to analyze the DDA raw data by directly searching against the Swiss-Prot reviewed human sequence database (downloaded in Dec. 2020)20. A mass accuracy of 15 ppm was applied to precursor ions and 0.05 Da to fragment ions. The database searches included fixed modifications such as carbamidomethyl (Cys) and dynamic modifications including oxidation (Met), acetylation (protein N-terminus), and deamidation (Asn, specifically for N-glycopeptides). Peptides with a false discovery rate (FDR) of 1%, proteins with 5%, and a minimum of one unique peptide were considered. N-glycopeptides were strictly defined as having deamination modification at Asn and containing the sequence N-X-S/T (X ≠ P).
The Spectronaut (Biognosys, v18, Switzerland) was employed to analyze DIA raw data files88. For both direct-DIA and library-DIA, a maximum of two missed cleavages was permitted. The variable and fixed modifications were consistent with those in the DDA analysis. FDR cutoff of 1% was established at the peptide-spectrum match, peptide, and protein levels. The precursor Q-value cutoff was 0.01, the precursor PEP cutoff was 0.2, and the protein PEP cutoff was 0.75. The calibration and main search were dynamic.
For the direct-DIA, the DIA data file was processed by Spectronaut software in directDIATM search model with the length of peptide ranging from 7 to 52, and other parameters in default. For the library-DIA, the DIA data of EV proteins were analyzed in Spectronaut by leveraging the above EV protein spectra library. While the DIA data of EV N-glycopeptides were processed in the same software as the above EV N-glycopeptide spectra library. N-glycopeptides were strictly characterized as having deamination modification at Asn and containing the sequence N-X-S/T (X ≠ P).
Statistical analyses
Statistical analyses were conducted using GraphPad (version 8.0.2), Python (v3.12.4), and R software (v4.3.1). For quantitative analysis of N-glycopeptides across AZS and NS samples, quantified N-glycopeptides with at least 60% valid values in one group in the discovery cohort were selected. Differentially expressed proteins and N-glycopeptides were defined as having fold change >1.5 or fold change <1/1.5 and P-value < 0.05 (two-sided independent Student’s t-test). The -log10 (P-value) for enriched terms was from the Gene Ontology (GO) enrichment analysis using the DAVID. The Area under the curve (AUC) values of N-glycopeptides for differentiating between AZS and NS in the discovery cohort were obtained through [https://cloud.oebiotech.cn/task/]. Tissue expression data related to the differential expression of glycoproteins were obtained from the Human Protein Atlas (HPA) [https://www.proteinatlas.org/]. The receiver-operating characteristic (ROC) curves and AUC values for individual N-glycopeptide and panel were calculated using scikit-learn (v1.4.2) via 5-fold cross-validation with a random forest classifier89. The optimal number of N-glycoprotein features was determined by recursive feature elimination with cross-validation in sklearn.feature_selection with AUC as the scoring metric. The SHapley Additive exPlanations (SHAP) values were calculated using the Shap library (v0.46.0) to assess N-glycoprotein contributions90. The AZS subtypes analysis was performed on the discovery cohort using the ConsensusClusterPlus package (v1.70.0) and on the validation cohort using the k-means package in R (v4.3.1)91.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The proteomics and N-glycoproteomics data generated in this study have been deposited in ProteomeXchange Consortium via the iProX partner repository92,93 (identifier: PXD057512 [https://www.iprox.cn//page/project.html?id=IPX0010057000] and PXD057515 [https://www.iprox.cn//page/project.html?id=IPX0010090000]). The initial and final configurations of the molecular dynamics simulation generated in this study have been deposited at [https://zenodo.org/uploads/15898317]. Unless otherwise stated, all data supporting the results of this study can be found in the article, supplementary, and source data files. Source data are provided with this paper.
Code availability
The molecular dynamics simulation can be accessed at [https://zenodo.org/records/15898317], and the AUC code is available at [https://github.com/XuejiangGuo/EVGpro]. The code is also available on Zenodo [https://doi.org/10.5281/zenodo.15877846], all with unrestricted and free access. Code used for generating figures is available from corresponding authors upon reasonable request.
References
Tkach, M. & Théry, C. Communication by extracellular vesicles: where we are and where we need to go. Cell 164, 1226–1232 (2016).
Zhang, Y., Wu, X. & Tao, W. A. Characterization and applications of extracellular vesicle proteome with post-translational modifications. Trac-Trends Anal. Chem. 107, 21–30 (2018).
Andaluz Aguilar, H., Iliuk, A. B., Chen, I. H. & Tao, W. A. Sequential phosphoproteomics and N-glycoproteomics of plasma-derived extracellular vesicles. Nat. Protoc. 15, 161–180 (2020).
Simon, C. et al. Extracellular vesicles in human reproduction in health and disease. Endocr. Rev. 39, 292–332 (2018).
Yokoi, A. et al. Identifying high-grade serous ovarian carcinoma-specific extracellular vesicles by polyketone-coated nanowires. Sci. Adv. 9, eade6958 (2023).
Kumar, M. A. et al. Extracellular vesicles as tools and targets in therapy for diseases. Signal. Transduct. Target. Ther. 9, 27 (2024).
Welsh, J. A. et al. Minimal information for studies of extracellular vesicles (MISEV2023): from basic to advanced approaches. J. Extracell. Vesicles 13, e12404 (2024).
Dhondt, B. et al. Unravelling the proteomic landscape of extracellular vesicles in prostate cancer by density-based fractionation of urine. J. Extracell. Vesicles 9, 1736935 (2020).
Chen, I. H. et al. Analytical pipeline for discovery and verification of glycoproteins from plasma-derived extracellular vesicles as breast cancer biomarkers. Anal. Chem. 90, 6307–6313 (2018).
Vinik, Y. et al. Proteomic analysis of circulating extracellular vesicles identifies potential markers of breast cancer progression, recurrence, and response. Sci. Adv. 6, eaba5714 (2020).
Yao, L. et al. Quantitative proteomic biomarkers from extracellular vesicles of human seminal plasma in the differential diagnosis of azoospermia. Clin. Transl. Med. 11, e423 (2021).
Murdica, V. et al. Proteomic analysis reveals the negative modulator of sperm function glycodelin as over-represented in semen exosomes isolated from asthenozoospermic patients. Hum. Reprod. 34, 1416–1427 (2019).
Fang, Z. et al. Glyco-decipher enables glycan database-independent peptide matching and in-depth characterization of site-specific N-glycosylation. Nat. Commun. 13, 1900 (2022).
Cui, M. et al. Cyst fluid glycoproteins accurately distinguishing malignancies of pancreatic cystic neoplasm. Signal Transduct. Target. Ther. 8, 406 (2023).
Schjoldager, K. T., Narimatsu, Y., Joshi, H. J. & Clausen, H. Global view of human protein glycosylation pathways and functions. Nat. Rev. Mol. Cell Biol. 21, 729–749 (2020).
Pinho, S. S. & Reis, C. A. Glycosylation in cancer: mechanisms and clinical implications. Nat. Rev. Cancer 15, 540–555 (2015).
Freitas, D. et al. Different isolation approaches lead to diverse glycosylated extracellular vesicle populations. J. Extracellular Vesicles 8 (2019).
Wang, Z. et al. Dual-selective magnetic analysis of extracellular vesicle glycans. Matter 2, 150–166 (2020).
Sun, J. et al. Profiling phosphoproteome landscape in circulating extracellular vesicles from microliters of biofluids through functionally tunable paramagnetic separation. Angew. Chem.-Int. Ed. 62, e202305668 (2023).
Sun, J. et al. Synergistically bifunctional paramagnetic separation enables efficient isolation of urine extracellular vesicles and downstream phosphoproteomic analysis. ACS Appl. Mater. Interfaces 13, 3622–3630 (2021).
Wei, D. et al. Targeted phosphoproteomics of human saliva extracellular vesicles via multiple reaction monitoring cubed (MRM3). Anal. Chem. 96, 1223–1231 (2024).
Su, X. et al. Enhanced proteomic profiling of human plasma-derived extracellular vesicles through charge-based fractionation to advance biomarker discovery potential. J. Extracell. Vesicles 13, e70024 (2024).
Hou, G. et al. Electric field assisted tangential flow filtration device for highly effective isolation of bioactive small extracellular vesicles from cell culture medium. Anal. Chem. 96, 13345–13351 (2024).
Xiong, Y. T., Li, M. M., Liu, Y. H., Liang, X. M. & Qing, G. Y. Enrichment driven glycoproteomics: New materials, new methods, and beyond. Trac-Trends in Analytical Chemistry 168 (2023).
Chen, Y. J. et al. Specific enrichment of urinary exosomes and exosomal glycopeptides by coefficient affinity of integrated L-cysteine and titania. Chinese Chem. Lett. 34 (2023).
Qing, G. Y., Yan, J. Y., He, X. N., Li, X. L. & Liang, X. M. Recent advances in hydrophilic interaction liquid interaction chromatography materials for glycopeptide enrichment and glycan separation. Trac-Trends in Analytical Chemistry 124 (2020).
Bai, H. et al. Development a hydrazide-functionalized thermosensitive polymer based homogeneous system for highly efficient N-glycoprotein/glycopeptide enrichment from human plasma exosome. Talanta 186, 513–520 (2018).
Zhao, J. Q. et al. Dietary fat and fatty acid consumptions and the odds of asthenozoospermia: a case-control study in China. Hum. Reprod. Open 2023, hoad030 (2023).
Guo, Y. et al. Proteomics analysis of asthenozoospermia and identification of glucose-6-phosphate isomerase as an important enzyme for sperm motility. J. Proteom. 208, 103478 (2019).
Cavarocchi, E., Whitfield, M., Saez, F. & Touré, A. Sperm ion transporters and channels in human asthenozoospermia: genetic etiology, lessons from animal models, and clinical perspectives. Int. J. Mol. Sci. 23, 3926 (2022).
Zhang, B. et al. A DNAH17 missense variant causes flagella destabilization and asthenozoospermia. J. Exp. Med. 217 (2020).
Xu, J. et al. Cell-penetrating peptide: a means of breaking through the physiological barriers of different tissues and organs. J. Controlled Release 309, 106–124 (2019).
Milletti, F. Cell-penetrating peptides: classes, origin, and current landscape. Drug Discov. Today 17, 850–860 (2012).
Kawaguchi, Y. et al. Dipicolylamine/metal complexes that promote direct cell-membrane penetration of octaarginine. Bioconjugate Chem. 30, 454–460 (2019).
Chen, Y. et al. Exosome detection via the ultrafast-isolation system: EXODUS. Nat. Methods 18, 212–218 (2021).
Tóth, E.Á. et al. Formation of a protein corona on the surface of extracellular vesicles in blood plasma. J. Extracell. Vesicles 10 (2021).
Agarwal, A., Selvam, M., Baskaran, S., Finelli, R. & Leisegang, K. Unraveling the footsteps of proteomics in male reproductive research: A scientometric approach. Antioxid. Redox Signal. 39, 760–775 (2021).
Takov, K., Yellon, D. M. & Davidson, S. M. Comparison of small extracellular vesicles isolated from plasma by ultracentrifugation or size-exclusion chromatography: yield, purity and functional potential. Journal of Extracellular Vesicles 8 (2019).
Jeppesen, D. K. et al. Comparative analysis of discrete exosome fractions obtained by differential centrifugation. J. Extracell. vesicles 3, 25011–25011 (2014).
Webber, J. & Clayton, A. How pure are your vesicles? Journal of extracellular vesicles 2 (2013).
Poliakov, A., Spilman, M., Dokland, T., Amling, C. L. & Mobley, J. A. Structural heterogeneity and protein composition of exosome-Like vesicles (prostasomes) in human semen. Prostate 69, 159–167 (2009).
Zhang, X., Vos, H. R., Tao, W. & Stoorvogel, W. Proteomic profiling of two distinct populations of extracellular vesicles isolated from human seminal plasma. Int. J. Mol. Sci. 21, 7957 (2020).
Yang, C. et al. Comprehensive proteomics analysis of exosomes derived from human seminal plasma. Andrology 5, 1007–1015 (2017).
Qu, X. et al. Proteomic analysis of circulating extracellular vesicles identifies potential biomarkers for lymph node metastasis in oral tongue squamous cell carcinoma. Cells 10, 2179 (2021).
Hu, L. et al. Interaction network of extracellular vesicles building universal analysis via eye tears: iNEBULA. Sci. Adv. 9 (2023).
Zheng, H. et al. Deconstruction of heterogeneity of size-dependent exosome subpopulations from human urine by profiling N-glycoproteomics and phosphoproteomics simultaneously. Anal. Chem. 92, 9239–9246 (2020).
Sun, Y. et al. Comparative proteomic analysis of exosomes and microvesicles in human saliva for lung cancer. J. Proteome Res. 17, 1101–1107 (2018).
Sun, Y. et al. Systematic comparison of exosomal proteomes from human saliva and serum for the detection of lung cancer. Analytica Chim. Acta 982, 84–95 (2017).
Dong, M. et al. Urinary glycoproteins associated with aggressive prostate cancer. Theranostics 10, 11892–11907 (2020).
Lou, R. et al. Benchmarking commonly used software suites and analysis workflows for DIA proteomics and phosphoproteomics. Nat. Commun. 14, 94 (2023).
Chong, W. H. et al. Stirring in suspension: Nanometer-sized magnetic stir bars. Angew. Chem.-Int. Ed. 52, 8570–8573 (2013).
Levine, H. et al. Temporal trends in sperm count: a systematic review and meta-regression analysis of samples collected globally in the 20th and 21st centuries. Hum. Reprod. Update 29, 157–176 (2023).
Lu, X., Huang, Y., Zhang, H. & Zhao, J. Effect of diabetes mellitus on the quality and cytokine content of human semen. J. Reprod. Immunol. 123, 1–2 (2017).
Miao, Y. et al. Organochlorine pesticide exposures, metabolic enzyme genetic polymorphisms and semen quality parameters among men attending an infertility clinic. Chemosphere 303, 135010 (2022).
Zhang, X. et al. CEP112 coordinates translational regulation of essential fertility genes during spermiogenesis through phase separation in humans and mice. Nat. Commun. 15, 8465 (2024).
Kumar, U. et al. TEX13B is essential for metabolic reprogramming during germ cell differentiation. Hum. Reprod. 39, 1390–1403 (2024).
Pham, T. D. et al. Intracytoplasmic sperm injection versus conventional in vitro fertilization in infertile couples with normal total sperm count and motility: does sperm morphology matter? Human Reprod. 40 (2024).
Wang, L., Li, H. & Zhou, W. Prehypertension in male affects both semen quality and pregnancy outcomes in their first single blastocyst frozen-thawed embryo transfer cycles. Fertil. Steril. 123, 406–414 (2025).
Xin, M. et al. Precision structural Interpretation of site-specific N-glycans in seminal plasma. J. Proteome Res. 21, 1664–1674 (2022).
Yang, X. et al. Proteomic analysis of N-glycosylation of human seminal plasma. Proteomics 15, 1255–1258 (2015).
Saraswat, M. et al. Human spermatozoa quantitative proteomic signature classifies normo- and asthenozoospermia. Mol. Cell. Proteom. 16, 57–72 (2017).
Zhang, Q., Ma, C., Chin, L.-S., Pan, S. & Li, L. Human brain glycoform coregulation network and glycan modification alterations in Alzheimer’s disease. Sci. Adv. 10, eadk6911 (2024).
Li, Y. et al. Urinary extracellular vesicle N-glycomics identifies diagnostic glycosignatures for bladder cancer. Nat. Commun. 16, 2292 (2025).
Ma, Y., Ma, Q. W., Sun, Y. & Chen, X. F. The emerging role of extracellular vesicles in the testis. Hum. Reprod. 38, 334–351 (2023).
Russo, C. L. et al. Mucin gene expression in human male urogenital tract epithelia. Hum. Reprod. 21, 2783–2793 (2006).
Winge, S. B. et al. Biglycan is a novel binding partner of fibroblast growth factor receptor 3c (FGFR3c) in the human testis. Mol. Cell. Endocrinol. 399, 235–243 (2015).
Liu, D., Liao, C. & Wolgemuth, D. J. A role for cyclin A1 in the activation of MPF and G2-M transition during meiosis of male germ cells in mice. Developmental Biol. 224, 388–400 (2000).
Wang, J. et al. Proteomic analysis of seminal plasma from asthenozoospermia patients reveals proteins that affect oxidative stress responses and semen quality. Asian J. Androl. 11, 484–491 (2009).
Wu, Y. et al. Quantitative proteomic analysis of human seminal plasma from normozoospermic and asthenozoospermic individuals. Biomed. Res. Int. 2019, 1–10 (2019).
Turunen, T., Magris, M., Malinen, M. & Kekäläinen, J. Seminal-plasma-mediated effects on sperm performance in humans. Cells 11, 2147 (2022).
Zhou, L. et al. Structures of sperm flagellar doublet microtubules expand the genetic spectrum of male infertility. Cell 186, 2897–2910.e19 (2023).
Skates, S. J. et al. Statistical design for biospecimen cohort size in proteomics-based biomarker discovery and verification studies. J. Proteome Res. 12, 5383–5394 (2013).
Trott, O. & Olson, A. J. Software news and update autodock vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Computational Chem. 31, 455–461 (2010).
Salomon-Ferrer, R., Case, D. A. & Walker, R. C. An overview of the amber biomolecular simulation package. Wiley Interdiscip. Rev.-Computational Mol. Sci. 3, 198–210 (2013).
Wang, J., Wolf, R. M., Caldwell, J. W., Kollman, P. A. & Case, D. A. Development and testing of a general amber force field. J. Computational Chem. 25, 1157–1174 (2004).
Maier, J. A. et al. ff14SB: Improving the accuracy of protein side chain and backbone parameters from ff99SB. J. Chem. Theory Comput. 11, 3696–3713 (2015).
Mark, P. & Nilsson, L. Structure and dynamics of the TIP3P, SPC, and SPC/E water models at 298 K. J. Phys. Chem. B 105, 9954–9960 (2001).
Kräutler, V., Van Gunsteren, W. F. & Hünenberger, P. H. A fast SHAKE:: Algorithm to solve distance constraint equations for small molecules in molecular dynamics simulations. J. Computational Chem. 22, 501–508 (2001).
Larini, L., Mannella, R. & Leporini, D. Langevin stabilization of molecular-dynamics simulations of polymers by means of quasisymplectic algorithms. J. Chem. Phys. 126, 104101 (2007).
Sagui, C. & Darden, T. A. Molecular dynamics simulations of biomolecules: Long-range electrostatic effects. Annu. Rev. Biophysics Biomolecular Struct. 28, 155–179 (1999).
Chen, Y., Zheng, Y., Fong, P., Mao, S. & Wang, Q. The application of the MM/GBSA method in the binding pose prediction of FGFR inhibitors. Phys. Chem. Chem. Phys. 22, 9656–9663 (2020).
Hou, T., Wang, J., Li, Y. & Wang, W. Assessing the performance of the MM/PBSA and MM/GBSA methods. 1. The accuracy of binding free energy calculations based on molecular dynamics simulations. J. Chem. Inf. Modeling 51, 69–82 (2011).
Genheden, S. & Ryde, U. The MM/PBSA and MM/GBSA methods to estimate ligand-binding affinities. Expert Opin. Drug Discov. 10, 449–461 (2015).
Nguyen, H., Roe, D. R. & Simmerling, C. Improved generalized born solvent model parameters for protein simulations. J. Chem. Theory Comput. 9, 2020–2034 (2013).
Weiser, J., Shenkin, P. S. & Still, W. C. Approximate atomic surfaces from linear combinations of pairwise overlaps (LCPO). J. Computational Chem. 20, 217–230 (1999).
Guo, Y. et al. Single-cell multi-omics analysis of in vitro post-ovulatory-aged oocytes revealed aging-dependent protein degradation. Mol. Cell. Proteom. 24, 100882 (2025).
Skowronek, P. et al. Rapid and in-depth coverage of the (phospho-) proteome with deep libraries and optimal window design for dia-PASEF. Mol. Cell. Proteom. 21, 100279 (2022).
Bruderer, R. et al. Optimization of experimental parameters in data-independent mass spectrometry significantly increases depth and reproducibility of results. Mol. Cell. Proteom. 16, 2296–2309 (2017).
Zhang, Y., Yang, H., Yu, Y. & Zhang, Y. Application of nanomaterials in proteomics-driven precision medicine. Theranostics 12, 2674–2686 (2022).
Cai, Z. et al. Synthetic augmentation of cancer cell line multi-omic datasets using unsupervised deep learning. Nat. Commun. 15, 10390 (2024).
Contreras-Sanz, A. et al. Proteomic profiling identifies muscle-invasive bladder cancers with distinct biology and responses to platinum-based chemotherapy. Nat. Commun. 16, 1240 (2025).
Chen, T. et al. iProX in 2021: connecting proteomics data sharing with big data. Nucleic Acids Res. 50, D1522–D1527 (2022).
Ma, J. et al. iProX: an integrated proteome resource. Nucleic Acids Res. 47, D1211–D1217 (2019).
Li, M. et al. Isolation of exosome nanoparticles from human cerebrospinal fluid for proteomic analysis. Acs Appl. Nano Mater. 4, 3351–3359 (2021).
Chatterjee, M. et al. C1q is increased in cerebrospinal fluid-derived extracellular vesicles in Alzheimer’s disease: A multi-cohort proteomics and immuno-assay validation study. Alzheimers Dement. 19, 4828–4840 (2023).
Acknowledgements
The authors wish to thank the National Natural Science Foundation (82221005 (X.G.), 82227808 (Z.X.), 82371606 (X.G.), 82371623 (Y.L.)), the National Key Research and Development Program of China (2021YFC2700200, X.G.), the Fundamental Research Funds for the Central Universities (2242023K5005, Z.X.), the Jiangsu Funding Program for Excellent Postdoctoral Talent (No. 2023ZB240, J.S.), the China Postdoctoral Science Foundation (No. 2024M750406, J.S.), and the Open Research Fund of Southeast University and Jiangsu Province Hospital (2024-K02, Z.X.).
Author information
Authors and Affiliations
Contributions
M.L., X.S., H.T., Y.D., J.S., D.W., X.Y., Y.W., and X.Z. investigation. M.L., B.W., Y.L.W., B.L., Y.L., Z.X., and X.G. data curation. M.L., Z.G., Y.G., Y.L., Z.X., and X.G. designed the study. M.L., B.L., X.L., X.S., J.N., and Y.Z. prepared samples for proteomic and glycoproteomic analysis. M.L., Y.D., J.S., and Y.C. carried out mass-spectrometry experiments. H.T., Y.L., and X.G. collected clinical samples. J.G. conducted the molecular docking and molecular dynamics simulation. M.L., Y.L., Z.X., and X.G. writing-review & editing.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Alessandro Arcovito who co-reviewed with Giorgia Canini; Albert Salas-Huetos, Xu Yang, and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Li, M., Song, X., Tu, H. et al. A functionally tunable magnetic nanochains platform for N-glycoproteomic analysis of extracellular vesicles from ultratrace biofluids. Nat Commun 16, 7646 (2025). https://doi.org/10.1038/s41467-025-63075-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-63075-0