Abstract
Lung adenocarcinoma (LUAD) is a molecularly diverse form of lung cancer characterized by distinct oncogenic driver mutations that influence both tumour biology and clinical outcomes. Understanding the interplay between these oncogenic drivers and the tumour microenvironment (TME) is crucial for improving therapeutic strategies and patient management. Here, we investigate the impact of driver mutations on the composition and spatial architecture of the TME in LUAD. Using imaging mass cytometry (IMC), we analyse tumour samples from 157 LUAD patients, integrating genomic and clinical data to link specific mutations with tumour characteristics. Unique patterns are associated with mutated KRAS and EGFR tumours with TP53 co-mutations, suggesting these co-mutations reshape the TME and promote resistance to tyrosine kinase inhibitors (TKIs). Overall, our findings highlight the complex interplay between oncogenic driver mutations and the TME in LUAD, underscoring the importance of integrating genomic and cellular data to understand the underlying tumour behaviour and prognosis.
Similar content being viewed by others
Introduction
Lung adenocarcinoma (LUAD) is the most prevalent subtype of lung cancer, distinguished by its molecular diversity and the presence of various oncogenic driver mutations1. The discovery of driver mutations in genes such as EGFR and KRAS has revolutionized treatment by enabling the development of targeted therapies2. These tyrosine kinase inhibitors (TKIs), specifically designed to inhibit recurrent mutations/rearrangements, have significantly improved patient outcomes by providing more personalized and effective treatment options. However, tumour cells eventually develop resistance, and cancer ultimately relapses3,4. The tumour microenvironment (TME) in lung cancer consists of a complex network of immune cells, stromal cells, and extracellular matrix components that interact with cancer cells. Therapeutic response is strongly modulated by the TME, which can either facilitate or impede treatment effectiveness5,6. Understanding TME dynamics is essential for developing strategies to enhance therapeutic efficacy and overcome resistance mechanisms in lung cancer treatment. Advancements in spatial profiling have revealed the intricate architecture of the tumour microenvironment, highlighting that clinical trajectories are shaped by both intrinsic cancer cell mutations and intercellular interactions among tumour-associated cell populations7. High-dimensional techniques, such as imaging mass cytometry (IMC), allow for detailed characterization of the cellular composition and spatial organization of the TME, providing insights into how specific mutations shape the tumour ecosystem.
Here, we investigate the impact of oncogenic driver mutations on the clinical characteristics and cellular composition of LUAD. By analysing samples from 157 LUAD patients using IMC, we integrate genomic and clinical data to explore correlations between specific mutations and tumour characteristics. This approach enables us to characterize how these mutations shape the TME, including intercellular interactions and spatial architecture, thereby influencing clinical outcomes. Our study underscores the importance of dissecting the crosstalk between oncogenic driver mutations and the TME. By interrogating TME differences associated with the two most prevalent LUAD drivers, KRAS & EGFR, and considering their frequent co-mutation with TP53, we gain mechanistic insight into tumour biology and the emergence of resistance to targeted therapies. This comprehensive understanding is critical for informing personalized therapeutic strategies and improving clinical outcomes in patients with LUAD.
Results
Oncogenic driver mutations are linked to distinct clinical characteristics
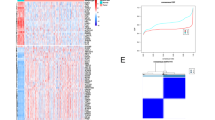
To elucidate how specific oncogenic alterations may influence the tumour and immunological landscape of LUAD, we interrogated tumours from 157 patients with pathologically confirmed LUAD, each with known genomic driver status (Supplementary Data 1). Driver alteration status was determined using the OncomineTM Precision Assay GX Gene Panel, which allows for the concurrent identification of biomarkers across 50 genes from both solid tissue and liquid biopsy samples (Fig. 1A). In total, we detected known oncogenic driver alterations in 117 patient tumours and stratified patients into six groups based on driver mutation status: KRAS (n = 71), EGFR (n = 26), MET (n = 9), PIK3CA (n = 8), BRAF (n = 3) or unknown oncogenic driver (n = 40) (Fig. 1B). TP53 mutations were detected in approximately one-third of all patients (47 patients) (Fig. 1B). Kaplan-Meier analysis revealed that patients with EGFR mutations had a significantly better probability of survival compared to patients with KRAS, PIK3CA or unknown driver mutations (Fig. 1C). Consistent with these findings, the majority of patients with KRAS and PIK3CA mutations had an expected positive history of tobacco use8,9 while EGFR mutations were enriched in female and non-smoker patients (Supplementary Fig. 1A). EGFR mutations were further linked to lower-grade lepidic and acinar histological patterns (Supplementary Fig. 1A), which are generally associated with better survival10,11. Our cohort primarily consisted of patients under the age of 75, with notable exceptions in the MET and PIK3CA groups, which were proportionally older than the other groups (Supplementary Fig. 1A). To enable spatial immune profiling, we applied highly multiplexed IMC to these 157 LUAD patient samples to generate a LUAD tumour immune atlas. Using a 35-plex antibody panel (Supplementary Table 1), we detected cancer cells, stromal cells, and various functional substates within innate and adaptive immune lineages (Fig. 1A, B). A supervised lineage assignment approach was used to classify over 600,000 single cells into 14 distinct immune cell populations, along with tumour cells and endothelial cells, using canonical lineage markers (Fig. 1D, Supplementary Fig. 1B, C). Integrating genomic and clinical data with this immune atlas provided the necessary resource to investigate how specific oncogenic alterations may influence the tumour and immunological landscape of LUAD.
A Schematic illustrating the Oncomine NGS process applied to FFPE or snap-frozen samples from 157 LUAD patients, combined with IMC acquisition for multiplexed imaging and single-cell phenotyping. B Oncoplot depicting the clinical characteristics, tumour cellular composition, and mutation status of the 157 patients in the study cohort. C Kaplan-Meier survival probability across different oncogenic driver groups, Unknown driver (n = 40), KRAS (n = 71), EGFR (n = 26), MET (n = 9), PIK3CA (n = 8), BRAF (n = 3) mutations. Statistical analysis was performed using two-sided Logrank Mantel-Cox and Gehan-Breslow-Wilcoxon tests. d Representative images of antibody staining for different oncogenic driver groups performed on tumour cores of patient with Unknown driver (n = 40), KRAS (n = 71), EGFR (n = 26), MET (n = 9), PIK3CA (n = 8), BRAF (n = 3) mutations. Scale bars, 100 µm. Source data are provided as a source data file.
Oncogenic driver mutations influence cellular frequencies in the TME of LUAD patients
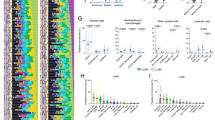
Having established this comprehensive atlas, we proceeded to analyse cellular frequencies across our cohort, examining variations within our six oncogenic driver groups (Fig. 2A, Supplementary Fig. 2A). This comprehensive analysis aimed to identify potential associations between oncogenic drivers and tumour cellular composition, thereby elucidating the relationship between the TME and the molecular characteristics of neoplastic cells. Across all groups, cancer cells, CD163- macrophages, and CD4+ helper T cells were consistently the most prevalent cell types (Fig. 2A, Supplementary Fig. 2A, B). Tumours with EGFR, PIK3CA or unknown driver alterations exhibited, on average, a higher proportion of lymphoid cells compared to myeloid cells, whereas KRAS, BRAF, and MET-driven tumours showed the opposite trend (Fig. 2B). A lower abundance of cancer cells was detected in patients with a PIK3CA mutations (Fig. 2C), suggesting higher microenvironmental involvement. The frequency of mast cells was specifically enriched in the EGFR group (Fig. 2C). Additionally, we found a lower proportion of unclassified cells in the unknown driver group, which are cells that could not be identified through our lineage antibody panel (Fig. 2C). To better understand the composition of these unclassified cells, we assessed CD45 expression and found that over 70% of these cells are CD45- and potentially include fibroblasts and other mesenchymal cells (Fig. 2D).
A Prevalence of 17 cell types, across 157 patients with LUAD as a proportion of total cells. B Prevalence of lymphoid and myeloid cells as a proportion of total cells across 6 oncogenic driver patient groups: Unknown driver (n = 40), KRAS (n = 71), EGFR (n = 26), MET (n = 9), PIK3CA (n = 8), BRAF (n = 3). Comparisons within individual oncogenic driver groups were performed using two-sided Wilcoxon tests, comparisons between oncogenic drivers were performed using two-sided Mann-Whitney tests. C Prevalence of indicated cell types as a proportion of total cells across 6 oncogenic driver patient groups, Unknown driver (n = 40), KRAS (n = 71), EGFR (n = 26), MET (n = 9), PIK3CA (n = 8), BRAF (n = 3). Two-sided Mann-Whitney tests were used for statistical analysis. D Pie chart depicting the proportion of unclassified cells that are CD45+ across 157 LUAD patients. E Prevalence of lymphoid, myeloid, cancer and endothelial cells as a proportion of total cells across 6 oncogenic driver patient groups stratified by sex: Unknown driver male (n = 23), Unknown driver female (n = 17), KRAS male (n = 20), KRAS female (n = 51), EGFR male (n = 8), EGFR female (n = 18), MET male (n = 7), MET female (n = 2), PIK3CA male (n = 4), PIK3CA female (n = 4), BRAF male (n = 2), BRAF female (n = 1). Comparisons between sex within individual oncogenic driver groups and between driver groups by sex were performed using a 2-way ANOVA. F Prevalence of indicated cell types as a proportion of total cells across 4 patient groups with different EGFR point mutations, p.E746_A750del (n = 7), p.G719A (n = 4), p.L858R (n = 9), Other (n = 6). Two-sided Mann-Whitney tests were used for statistical analysis. Source data are provided as a source data file.
To explore factors influencing the tumour microenvironment beyond driver mutations, we further stratified patients by sex and age. Among patients with PIK3CA mutations, females and those under 75 years of age exhibited higher levels of lymphocyte infiltration compared to their male counterparts and to females with unknown driver mutations (Fig. 2E, Supplementary Fig. 2C). Notably, female KRAS-mutant patients showed a higher frequency of myeloid cells than male KRAS patients and, along with female PIK3CA patients, had a lower abundance of cancer cells than females with unknown driver mutations (Fig. 2E). A similar pattern was observed in PIK3CA-mutant patients under 75 (Supplementary Fig. 2C). Interestingly, within the unknown driver mutation group, female patients had a lower frequency of endothelial cells in their tumour cores compared to male patients, suggesting reduced vascularization (Fig. 2E).
To gain a deeper understanding of how specific genetic alterations in the same driver gene can differentially impact the tumour immune microenvironment, we focused our analysis on KRAS and EGFR, given their high frequency of alteration within these genes. Mutations in KRAS and EGFR typically lead to a more active form of these proteins which can drive cancer progression12,13. The frequency of KRAS mutations in non-small cell lung cancer (NSCLC) varies but is generally reported to be around 30%1. These mutations are particularly common in adenocarcinoma, where the frequency can be as high as 40%14. The most common KRAS mutations in NSCLC occur at codon 12, with G12C, G12D, and G12V mutations being the most frequently observed alterations. KRAS G12C has specific inhibitors available due to the unique cysteine residue that allows for covalent binding15,16. In contrast, G12V and G12D lack such inhibitors, making them more challenging targets for drug development. Beyond the efficacy of the targeted inhibitor, we investigated whether these mutations could lead to changes in the TME in LUAD. In KRAS-driven tumours, we did not observe any significant changes in the frequency of immune cell types with specific point mutations in G12C (n = 27), G12D (n = 9), G12V (n = 19), or other mutations (n = 16) (Supplementary Fig. 2D). In contrast, EGFR-driven tumours exhibited several significant shifts in cellular frequencies depending on specific genetic alterations, such as p.E746_A750del (n = 7), p.G719A (n = 4), p.L858R (n = 9), and others (n = 6) (Fig. 2F, Supplementary Fig. 2D, E). EGFR mutations are present in approximately 10-30% of all NSCLC patients, with frequency variations influenced by ethnicity. The highest prevalence is observed in Asian populations, followed by North and South American, and European populations17. Three generations of TKIs targeting EGFR have been successfully used in the clinic, but acquired drug resistance ultimately overcomes efficacy3,18,19. Tumours with p.E746_A750del and p.G719A alterations showed a significant decrease in the frequency of cancer cells, while classical monocytes were increased (Fig. 2F). This could indicate a heightened immune response or an altered TME that restricts cancer cell proliferation with these mutations. Furthermore, regulatory T cells were significantly increased in tumours with p.G719A alterations, suggesting an immunosuppressive microenvironment that could impact the effectiveness of immune therapies (Fig. 2F). CD163+ macrophages were elevated in p.E746_A750del tumours, which might contribute to a pro-tumourigenic environment and influence tumour progression and metastasis (Fig. 2F). Additionally, T cells and CD163- macrophages were more prevalent in p.G719A mutant tumours, potentially indicating an active immune surveillance mechanism (Supplementary Fig. 2E). Endothelial cells were most frequently detected in tumours with p.E746_A750del and p.G719A mutations (Supplementary Fig. 2E), which could reflect enhanced angiogenesis in these specific genetic contexts. These observations underscore the distinct differences in the cellular composition of the TME based on the oncogenic EGFR driver mutation status.
TP53 co-mutation associated with immunosuppressive cell-cell interactions and avoidance patterns
Recently, osimertinib was found to be significantly less effective in patients with EGFR mutant NSCLC associated with TP53 co-mutations20. These findings carry significant clinical implications, as identifying patients with both EGFR and TP53 mutations allows for the development of more tailored and targeted treatment strategies. This could involve more intensive follow-up, early radiotherapy or ablation to address resistant tumours, or the early use of TKIs in combination with other drugs, including immunotherapy; yet how the underlying tumour and immunological landscape of NSCLC is affected by TP53 co-mutations remains poorly understood. To address this, we first compared the frequency of cells within EGFR vs EGFR + TP53 co-mutated tumours. Surprisingly, we found no significant difference in the frequency of any major cell populations (Supplementary Fig. 3); however, we hypothesized there may be functional differences based on cellular interactions amongst these cell types. To address this, we employed permutation testing to determine the likelihood of pairwise interaction or avoidance behaviours based on the spatial distribution of each population. In EGFR tumours with TP53 co-mutations compared to those without, we observed an increase in the tendency towards interactions between CD8+ and CD4+ T cells with NK cells (Fig. 3A, box i), and those NK cells were also more likely to interact with the endothelium (Fig. 3A, box ii). Physical proximity between CD8+ T cells and NK cells can amplify antitumour activity through reciprocal activation loops (e.g., IL2, IFNγ, and granzyme/perforin exchange). Cooperative CD8+ T cell–NK cell circuitry maximises target cell killing and has been linked to favourable outcomes in LUAD21. Frequent NK cell–endothelial cell contacts may reflect endothelial-mediated immunoregulation. Tumour conditioned endothelial cells can present adhesion molecules and chemokines that gate NK cell entry but also upregulate PD-L1 or release nitric oxide to blunt NK cytotoxicity which can shape vascular remodelling and immune infiltration22,23. In contrast, we found a significant decrease in the likelihood of interaction between B cells and neutrophils (Fig. 3A, box iii). Tumour-infiltrating B cells often reside in tertiary lymphoid structures (TLS) that correlate with prolonged survival and better immunotherapy response7, whereas neutrophils drive immunosuppression, metastasis and poor prognosis24. Spatial segregation of B cells from neutrophils could preserve TLS integrity and limit neutrophil derived suppression25. We also observed changes in macrophage interactions; CD163- and CD163+ macrophages had an increased tendency to interact with T cell subsets (Fig. 3A, box iv), while CD163- macrophages significantly avoided mast cells (Fig. 3A, box v), which are highly abundant in never-smokers26 and enriched for EGFR-amplified patients (Fig. 2C). Co-localisation of both macrophage subsets with T cells may underlie mixed immune signatures characteristic of TP53-mutant EGFR tumours. Overall, these context-specific interactions suggest that TP53 co-mutation reshapes EGFR-driven LUAD toward a microenvironment combining heightened cytotoxic potential with nuanced myeloid and vascular regulation. Functionally exploring such spatial crosstalk may provide clues for combinatorial immunotherapies targeting NK cell–T cell synergy, macrophage polarisation, and vascular immunomodulation.
A Heatmap depicting significant pairwise cell–cell interaction (red) or avoidance (blue), grey boxes represent interaction or avoidance patterns with less than random chance by permutation-based proximity analysis, across samples from patients with an EGFR driver mutation (n = 19) compared to EGFR driver mutation with a TP53 co-mutation (n = 7). 50,000 permutations per image. Black boxes indicate associations referenced in the text. B Prevalence of cells as a proportion of total cells compared between patients with KRAS (n = 54) and KRAS with a TP53 co-mutation (n = 17). Two sided Mann-Whitney tests were used for statistical analysis. c Heatmap depicting significant pairwise cell–cell interaction (red) or avoidance (blue), grey boxes represent interaction or avoidance patterns with less than random chance, by permutation-based proximity analysis, across samples from patients with an KRAS driver mutation (n = 54) compared to KRAS driver mutation with a TP53 co-mutation (n = 17). 50,000 permutations per image. Black boxes depict associations referenced in the text. Source data are provided as a source data file.
Next, we were interested in whether the TME changes associated with TP53 co-mutations were specific to EGFR tumours, or if they applied more generally in tumours with other genetic drivers. Given the emerging interest in targeted treatment options for KRAS-mutated NSCLC patients, we focused on these tumours to assess the putative impact of TP53 more broadly. Although we did not detect significant differences in immune cell frequencies between specific KRAS alterations (Supplementary Fig. 2D), the presence of TP53 co-mutation was associated with notable changes, including increases in classical and intermediate monocytes, as well as elevated endothelial cell populations (Fig. 3B, Supplementary Fig. 3). However, interaction analysis revealed even stronger effects, with the most prominent differences observed within the monocyte compartment. Specifically, in KRAS tumours harbouring a TP53 co-mutation compared to those without, we detected a significant increase in pairwise interactions involving intermediate monocytes with cancer cells (Fig. 3C, box i), endothelial cells (Fig. 3C, box ii), CD4⁺ helper T cells (Fig. 3C, box iii), and various monocyte and macrophage subsets (Fig. 3C, box iv). Because these intermediate monocytes lacked expression of markers associated with more differentiated cells (e.g., macrophages, dendritic cells), our findings suggest they engage in cellular interactions within the TME prior to full differentiation27. Moreover, the diversity of these interactions in KRAS TP53 co-mutated tumours specifically suggest broader functional heterogeneity.
Oncogenic driver mutations dictate spatial landscape with further differences based on TP53 co-mutation status in LUAD patients
To look beyond pairwise interactions within the TME, we aimed to delineate spatial organization differences among oncogenic driver groups, which could enhance prognostic insights7,28. We therefore calculated cellular neighbourhoods based on cellular composition of the 10 closest spatial neighbours of each cell, followed by reclassifying cells according to their spatially defined cellular neighbourhoods (CN). Through this process, we identified nine distinct neighbourhoods (CN1-9) within our cohort, which we named: CD163- macrophage enriched (CN1), tumour compartment (CN2), tumour boundary (CN3), unclassified (CN4), vascular niche (CN5), neutrophil enriched (CN6), B cell enriched (CN7), helper T cell enriched (CN8) and pan immune (CN9) (Fig. 4A, Supplementary Fig. 4A, B). The cellular neighbourhood enriched in neutrophils (CN6) was found almost exclusively in KRAS-driven tumours (Fig. 4B). PIK3CA-driven tumours were enriched in CN5 (vascular niche), indicating these tumours may be more highly vascularized (Fig. 4B). In tumours with no known driver alteration, CN4, which is enriched in unclassified cells, was almost nonexistent (Fig. 4B). This suggests that specific genomic alterations of malignant cells impact the spatial organization of the TME.
A Heatmap of 9 CNs discovered in 157 patients with LUAD calculated on 10 closest neighbouring cells. Log-rank test for survival. B Average distribution of CNs across patient groups: Unknown driver (n = 40), KRAS (71), EGFR (n = 26), MET (n = 9), PIK3CA (n = 8), BRAF (n = 3) discovered in (A). C Frequency of CN7 and CN8 as a percentage of all CNs in patient gropus: Unknown driver (n = 40), KRAS (71), EGFR (n = 26), MET (n = 9), PIK3CA (n = 8), BRAF (n = 3) discovered in (A). Two-sided Mann-Whitney tests were used for statistical analysis. D Heatmap of 9 BCNs discovered in 157 patients with LUAD calculated on 20 closest neighbouring cells. Log-rank test for survival. E Frequency of BCN8 as a percentage of all BCNs in patient groups: Unknown driver (n = 40), KRAS (71), EGFR (n = 26), MET (n = 9), PIK3CA (n = 8), BRAF (n = 3) discovered in (D). F Number of mast cells in BCNs discovered in (D). G Heatmap of 9 ECNs discovered in 26 patients with EGFR driver mutation. H Representative images of 9 cellular neighbourhoods in patients with EGFR (n = 19) or EGFR + TP53 (n = 7) co-mutations tumours discovered in (G) using Voronoi diagrams. I Average distribution of ECNs across patients with EGFR (n = 19) and EGFR TP53 (n = 7) discovered in (G). J Cell distribution within ECN5 discovered in (G). K Frequency of ECN5 in patients with EGFR (n = 19) and EGFR TP53 (n = 7) as a percentage of all ECNs discovered in (G). Two-sided Mann-Whitney test was used for statistical analysis. Source data are provided as a source data file.
Although KRAS and EGFR mutant tumours now have targeted therapeutic options, within this historical cohort we wanted to assess whether the enrichment of specific CNs was associated with overall survival outcomes. We conducted a Kaplan-Meier analysis, categorizing the frequency of CNs for each patient as either high (CNhigh; z-score ≥ 0) or low (CNlow; z-score <0). Consistent with our previous findings7, we observed that CN7 (B cell enriched) was significantly correlated with increased overall survival across all KRAS and EGFR patients (Fig. 4A), despite no significant enrichment of this CN in either of these groups (Fig. 4C). To expand on our analysis of CNs associated with favorable survival outcomes, we performed a more stringent Kaplan-Meier analysis by stratifying patients into quartiles based on CN frequency and comparing the top (z-score ≥ 75th percentile) and bottom (z-score ≤ 25th percentile) quartiles. By restricting comparisons to the top and bottom quartiles, we maximize biological contrast and statistical clarity, minimizing mid-range variability. This approach enhances detection of robust survival associations, while filtering out weaker signals. This analysis revealed CN8, characterized by an enrichment of helper T cells, to be significantly associated with improved survival, suggesting a dose-dependent effect, attributed to high helper T cell densities. The previously observed association with B cell enriched CN7 did not reach significance in the more stringent quartile comparison, indicating its effect is distributed across a wider continuum of enrichment levels among patients rather than concentrated at the extremes of enrichment (Fig. 4A). Further analysis revealed that CN8 was particularly enriched in patients with EGFR driver mutations (Fig. 4C), suggesting a possible mechanism underlying the improved survival observed in these patients (Fig. 1C). EGFR signalling can upregulate chemokines such as CXCL10/11 that attract CD4⁺ and CD8⁺ T cells, potentially explaining why helper T cell rich CN8 is common in EGFR tumours and coincides with their modest survival advantage.
To further investigate the relationship between CNs and patient survival, we explored broader spatial neighbourhoods by expanding our analysis to include the 20 nearest neighbours of each cell. These larger neighbourhoods, referred to as BCNs (Broader Cellular Neighbourhoods), were generally less defined by a single dominant cell type compared to the more distinct CNs identified using only the 10 closest neighbours (Fig. 4D, Supplementary Fig. 4C). In addition to the previously identified B cell enriched BCN4, we found two other neighbourhoods—BCN7 (Tumour Boundary 2) and BCN8 (Pan-Immune Hotspot 2)—that were positively associated with overall survival (Fig. 4D). To validate these associations, we again stratified patients into quartiles based on BCN frequency and comparing the top (z-score ≥ 75th percentile) and bottom (z-score ≤ 25th percentile) quartiles. Under this refined analysis, the survival association was lost for all BCNs except BCN8, which showed an even stronger correlation with improved survival, underscoring the robustness of this finding. Interestingly, BCN8 was evenly distributed across patients regardless of their driver mutation status, suggesting that its association with survival is independent of genetic alterations (Fig. 4E). This neighbourhood was characterized by a high abundance of immune cells, particularly mast cells, which were present in greater numbers than in any other BCN (Fig. 4F, Supplementary Fig. 4C). While some studies have shown that mast cells are associated with better survival outcomes in LUAD patients29,30, further studies are needed to clarify the role of mast cells and other immune cells in mediating this survival advantage. Together, these layers suggest that driver mutations steer, but do not fully dictate, TME architecture. Tumour intrinsic cues set the stage for certain micro niches, yet emergent, higher order immune structures can arise (or fail to arise) regardless of the underlying genotype. These findings do not negate the role of genomics; rather, they suggest that genomic alterations are necessary but may not be sufficient on their own. Driver mutations shape specific favourable cellular neighbourhoods, yet broader immune architectures such as BCN8 can enhance survival independently of tumour genotype. Thus, optimal therapeutic strategies and prognostic assessments should integrate tumour genomic data with spatially resolved immune metrics, rather than relying solely on genetics.
Given the enrichment of the helper T cell rich CN8 in EGFR-mutant tumours (Fig, 4A–C), we redefined cellular neighbourhoods specifically within this genotype (termed ECN) to evaluate the impact of TP53 co-mutations (Fig. 4G, H, Supplementary Fig. 4D). We identified a helper T cell-dominated neighbourhood (ECN5) that was significantly reduced in TP53-mutant cases, despite unchanged overall helper T cell frequency. These findings align with our pairwise interaction analysis, indicating that TP53 mutations primarily disrupt broader T cell niches rather than altering individual cell frequencies. Therefore, impaired T cell spatial organization, alongside genomic instability, may represent an under-recognized mechanism contributing to osimertinib resistance in EGFR-driven LUAD and warrants further study.
Discussion
In this study, we used highly multiplexed IMC combined with Oncomine NGS profiling of 157 LUAD tumours to explore the influence of common oncogenic driver mutations on the TME and clinical outcomes. We resolved over 600,000 cells, identifying multiple immune and stromal cell subsets, and obtained detailed spatial context. Our analysis revealed that specific oncogenic drivers distinctly influence immune and stromal cell populations, underscoring the pivotal role genetics play in shaping the TME. Notably, EGFR mutations significantly impacted the composition and frequency of cancer cells, immune cells, and endothelial cells, while KRAS mutations had minimal effects on overall cell frequencies. Importantly, our data highlight that genetic factors alone are not sufficient to fully define TME complexity or clinical outcomes. Although no significant differences in overall cell frequencies were found between EGFR-mutated tumours with or without TP53 co-mutations, spatial analysis uncovered meaningful functional differences through altered cellular interactions and organization. Specifically, TP53 co-mutations were associated with decreased abundance of a helper T cell-enriched neighbourhood, indicative of disrupted spatial niches rather than altered cell numbers. Similarly, KRAS-mutant tumours with TP53 co-mutations exhibited enhanced interactions involving intermediate monocytes, further supporting the notion that co-mutations profoundly modulate immune cell interactions rather than cell frequencies alone. These findings emphasize a critical refinement to the prevailing genomic paradigm: oncogenic driver mutations and associated co-mutations clearly influence TME composition, yet spatial organization and cellular interactions within these genetic contexts play crucial roles in mediating clinical outcomes. The differential efficacy of therapies such as osimertinib in EGFR-mutant patients harboring TP53 co-mutations underscores the necessity of integrating spatial immune metrics with genomic profiling for optimal therapeutic strategies. This study provides a large-scale spatially resolved TME analysis in the context of specific oncogenic alterations, while acknowledging limitations including modest sample sizes for rare mutations and potential effects of intratumoural heterogeneity due to restricted sampling. Our spatial analysis was performed on a single 1 mm² IMC field per tumour, and therefore, both spatial and cellular heterogeneity may not be fully represented. Previous IMC studies have shown that a single representative region can reliably reflect broad immune–stromal architectures; however, fine-scale heterogeneity may be undersampled. Future work incorporating multiregional sampling or whole slide imaging, alongside higher-dimensional spatial modalities, will be essential to confirm the generalizability of our cellular neighbourhood signatures and to refine patient classification frameworks. Nonetheless, our results clearly illustrate the clinical relevance of considering both genetic alterations and their spatial immune context for future clinical decision making, encouraging further evaluation of these patterns as a possible aid to patient management. Further investigations into spatially defined immune interactions and their functional implications for therapy resistance, especially in tumours harboring TP53 co-mutations, are warranted to enhance tailored treatment approaches and improve patient outcomes.
Methods
All research was conducted in compliance with ethical guidelines and regulations, with study approvals described in the relevant sections.
Clinical cohort
A cohort of 157 LUAD patients who underwent a lung surgical resection from March 2006 to February 2021, was collected at the Institut universitaire de cardiologie et de pneumologie de Québec – Université Laval (IUCPQ-UL). The inclusion criteria were as follows: (1) diagnosis of LUAD; (2) complete surgical resection with negative margins, and (3) availability of tissue for molecular characterization. All samples were primary treatment-naive LUADs, diagnosed by a board-certified pathologist after surgical resection or biopsy. Tissue microarrays were created by selecting one 1-mm² core from the surgical tumour specimen. Written informed consent was obtained from all patients before collecting samples and clinical data. The biobanking protocols for human samples were approved by the IUCPQ-UL ethics committee, under protocol number IRB #2022-3474, 22090.
Sample staining and IMC acquisition
The methods for sample staining, preparation, and imaging mass cytometry (IMC) have been previously described7. In short, Formalin-fixed paraffin-embedded (FFPE) slides were deparaffinized at 70 °C in EZ Prep solution (Roche Diagnostics) and underwent antigen retrieval at 95 °C in standard cell conditioning 1 solution (Roche Diagnostics) using the Ventana Discovery Ultra auto stainer platform. After a PBS wash, slides were blocked with Dako serum-free protein block (Agilent) for 45 min, then incubated overnight at 4 °C with a cocktail of metal-tagged antibodies (see Supplementary Table 1). Antibodies were conjugated by the Single Cell and Imaging Mass Cytometry Platform at the Goodman Cancer Institute using Maxpar kits (Fluidigm). Following washes in 0.2% Triton X and PBS, a secondary metal-conjugated anti-biotin antibody in Dako diluent was applied for 1 h and again washed with 0.2% Triton X and PBS. Slides were then counterstained with Cell-ID Intercalator-Ir (1:400, Fluidigm) for 30 min at room temperature, rinsed with distilled water, and air-dried. IMC images were acquired at ~1 μm resolution via 200 Hz laser ablation on the Hyperion Imaging System (Fluidigm). Raw data were processed using the Fluidigm commercial acquisition software.
Antibody optimization
All antibodies used for staining were optimized in previous studies7. Control tissues—including spleen, tonsil, appendix, placenta, thymus, normal lung, and lung adenocarcinoma (LUAD)—served as positive and negative controls for staining with the antibody panel, exception for alpha-cleaved H3. Four markers (GM-CSFR, PD-1, PD-L1, and B7-H3) did not consistently stain the control tissues and were excluded from biological interpretations.
Data transformation and normalization
The data presented in this study were not transformed, and raw IMC measurements were used directly for analysis.
Cell segmentation and lineage assignment
Cells in the IMC images were segmented using a combination of classical and modern machine learning-based computer vision algorithms31. The detailed code and description of the cell segmentation and lineage assignment pipeline are available (https://github.com/walsh-quail-labs).
Cell–cell pairwise interaction
To evaluate the significance of pairwise interactions among different cell types within individual IMC image cores, we developed a permutation-based proximity analysis. For each image, 50,000 permutations were performed in which cell type labels were randomly reassigned while spatial coordinates were preserved. Counts were generated to represent the number of times the observed interaction was more frequent (indicating interaction) or less frequent (indicating avoidance) than expected under the null distribution. Interactions were considered significantly increased if the observed frequency exceeded the 25,000th permutation (the median of the null distribution), and significantly avoidant if it fell below this threshold.
Neighbourhood identification
The code and methodology for cellular neighbourhood (CN) identification have been previously described7. Each cell was assigned to a CN based on its defining window. The prevalence of each neighbourhood within each core was normalized to sum to 100%. For oncogenic driver groups, values were z-scored, and cores with z-scores above or equal to 0 were compared with those below 0 for survival outcomes.
Statistical analysis and workflow
Image analysis was conducted using MATLAB (version 2019b) and Python (version 3.7.12), whereas statistical analyses were performed with RStudio (version 4.2.2) and GraphPad Prism (version 10.1.1). Data are expressed as mean ± s.e.m. or mean ± s.d., with a significance level set at P < 0.05 unless specified otherwise. Survival data were evaluated using Cox regression and log-rank (Mantel–Cox) test. Detailed descriptions of all statistical tests can be found in the figure legends.
NGS sequencing using Oncomine™ precision assay GX gene panel
Comprehensive molecular analysis was conducted to determine mutations/rearrangements in 50 clinically relevant genes. First, DNA/RNA was extracted from snap-frozen or formalin-fixed paraffin-embedded (FFPE) samples. It was followed by a targeted approach using the Oncomine™ Precision Assay GX Gene Panel (Thermo Fisher Scientific). Mutation analysis was performed using targeted next-generation sequencing (NGS) on the Ion Torrent Genexus platform (Thermo Fisher Scientific) following the manufacturers recommendation. The molecular analyses were reviewed by two pathologists.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Data supporting the findings in this study, including high-dimensional TIFF images, single-cell segmentation, single-cell cell types, and patient data are available at https://doi.org/10.5281/zenodo.7383627. The remaining data are available within the Article, Supplementary Information or Source Data file. Source data are provided with this paper.
Code availability
The original code used to produce the results of this study is available at https://github.com/walsh-quail-labs/IMC-Lung.
References
Collisson, E. A. et al. Comprehensive molecular profiling of lung adenocarcinoma. Nature 511, 543–550 (2014).
Imyanitov, E. N., Iyevleva, A. G. & Levchenko, E. V. Molecular testing and targeted therapy for non-small cell lung cancer: Current status and perspectives. Crit. Rev. Oncol. /Hematol. 157, 103194 (2021).
Del Re, M. et al. Concise review: resistance to Tyrosine Kinase inhibitors in non-small cell lung cancer: the role of cancer stem cells. Stem Cells 36, 633–640 (2018).
Lim, S. M., Syn, N. L., Cho, B. C. & Soo, R. A. Acquired resistance to EGFR targeted therapy in non-small cell lung cancer: Mechanisms and therapeutic strategies. Cancer Treat. Rev. 65, 1–10 (2018).
Wang, L., Jia, Q., Chu, Q. & Zhu, B. Targeting tumor microenvironment for non-small cell lung cancer immunotherapy. Chin. Med. J. Pulm. Crit. Care Med. 1, 18–29 (2023).
Wu, T. & Dai, Y. Tumor microenvironment and therapeutic response. Cancer Lett. 387, 61–68 (2017).
Sorin, M. et al. Single-cell spatial landscapes of the lung tumour immune microenvironment. Nature 614, 548–554 (2023).
Wang, X. et al. Association between smoking history and tumor mutation burden in advanced non-small cell lung cancer. Cancer Res 81, 2566–2573 (2021).
Wang, Y., Wang, Y., Li, J., Li, J. & Che, G. Clinical significance of PIK3CA gene in non-small-cell lung cancer: a systematic review and meta-analysis. Biomed. Res Int 2020, 3608241 (2020).
Wolf, J. L. et al. The value of prognostic and predictive parameters in early-stage lung adenocarcinomas: A comparison between biopsies and resections. Lung Cancer 176, 112–120 (2023).
Yasukawa, M. et al. Histological grade: analysis of prognosis of non-small cell lung cancer after complete resection. Vivo 32, 1505–1512 (2018).
Lu, S., Jang, H., Nussinov, R. & Zhang, J. The structural basis of oncogenic mutations G12, G13 and Q61 in small GTPase K-Ras4. B. Sci. Rep. 6, 21949 (2016).
Lemmon, M. A., Schlessinger, J. & Ferguson, K. M. The EGFR family: not so prototypical receptor tyrosine kinases. Cold Spring Harb. Perspect. Biol. 6, a020768 (2014).
Sorin, M. et al. Next-generation sequencing of non-small cell lung cancer at a Quebec health care cancer centre. Cancer Treat. Res Commun. 35, 100696 (2023).
Skoulidis, F. et al. Sotorasib for lung cancers with KRAS p.G12C mutation. N. Engl. J. Med 384, 2371–2381 (2021).
Jänne, P. A. et al. Adagrasib in non-small-cell lung cancer harboring a KRASG12C mutation. N. Engl. J. Med 387, 120–131 (2022).
Zhang, Y.-L. et al. The prevalence of EGFR mutation in patients with non-small cell lung cancer: a systematic review and meta-analysis. Oncotarget 7, 78985–78993 (2016).
Wang, Z.-F., Ren, S.-X., Li, W. & Gao, G.-H. Frequency of the acquired resistant mutation T790 M in non-small cell lung cancer patients with active exon 19Del and exon 21 L858R: a systematic review and meta-analysis. BMC Cancer 18, 148 (2018).
Wu, S.-G. et al. The mechanism of acquired resistance to irreversible EGFR tyrosine kinase inhibitor-afatinib in lung adenocarcinoma patients. Oncotarget 7, 12404–12413 (2016).
Hobor, S. et al. Mixed responses to targeted therapy driven by chromosomal instability through p53 dysfunction and genome doubling. Nat. Commun. 15, 4871 (2024).
Uzhachenko, R. V. & Shanker, A. CD8+ T Lymphocyte and NK cell network: Circuitry in the Cytotoxic domain of immunity. Front. Immunol. 10, (2019).
Fang, J. et al. Exploring the crosstalk between endothelial cells, immune cells, and immune checkpoints in the tumor microenvironment: new insights and therapeutic implications. Cell Death Dis. 14, 1–15 (2023).
Leone, P. et al. Endothelial cells in tumor microenvironment: insights and perspectives. Front. Immunol. 15, 1367875 (2024).
Aloe, C. et al. Emerging and multifaceted role of neutrophils in lung cancer. Transl. Lung Cancer Res. 10, 2806–2818 (2021).
Costa, S., Bevilacqua, D., Cassatella, M. A. & Scapini, P. Recent advances on the crosstalk between neutrophils and B or T lymphocytes. Immunology 156, 23–32 (2019).
Qiu, Z., Pang, G., Xu, X., Lin, J. & Wang, P. Characteristics of mast cell infiltration in lung adenocarcinoma and its impact on prognosis. Discov. Oncol. 15, 208 (2024).
Guilliams, M., Mildner, A. & Yona, S. Developmental and functional heterogeneity of monocytes. Immunity 49, 595–613 (2018).
Karimi, E. et al. Single-cell spatial immune landscapes of primary and metastatic brain tumours. Nature 1–9 https://doi.org/10.1038/s41586-022-05680-3 (2023).
Fan, F. et al. Elevated mast cell abundance is associated with enrichment of CCR2+ Cytotoxic T cells and favorable prognosis in lung Adenocarcinoma. Cancer Res 83, 2690–2703 (2023).
Bao, X., Shi, R., Zhao, T. & Wang, Y. Mast cell-based molecular subtypes and signature associated with clinical outcome in early-stage lung adenocarcinoma. Mol. Oncol. 14, 917–932 (2020).
Karimi, E. et al. Machine Learning Meets Classical Computer Vision for Accurate Cell Identification. http://biorxiv.org/lookup/doi/10.1101/2022.02.27.482183 (2022).
Acknowledgements
We thank the IUCPQ-UL Biobank of the Quebec Respiratory Health Research Network for providing access to tissue and clinical data. The molecular characterization of the tumours was supported by the SynergiQC project (CQDM) and the IUCPQ Foundation owing to a generous donation from Mr. Normand Lord. The authors are grateful for support from the Single Cell and Imaging Mass Cytometry Platform (SCIMAP, Yuhong Wei) and the Histology core facility (Nicole Robinson and Plinio Cruz) at the Rosalind and Morris Goodman Cancer Institute and the Quebec Cancer Consortium, as well as financial support from the Ministère de l’Économie et de l’Innovation du Québec through the Fonds d’accélération des collaborations en santé. The authors acknowledge support from the McGill Interdisciplinary Initiative in Infection and Immunity (Mi4). S.H. is grateful for funding from the Dr. Viktor KS Lui Studentship and holds a FRQS doctoral award. L.A.W. acknowledges funding from the Bachynski Family Foundation, the Canadian Institutes of Health Research (CIHR; PJT-162137, PJT-195919, PJT-191784), the Canadian Foundation for Innovation, and holds a Rosalind and Morris Goodman Chair in Lung Cancer Research. L.D. is supported by a CIHR Doctoral Award. Philippe Joubert holds a Junior 2 Clinical Research Scholar award from the Fond de Recherche Québec - Santé (FRQS) and holds a Canada Research Chair in Identification of predictive and diagnostic biomarkers to improve lung cancer management. Yohan Bossé holds a Canada Research Chair in Genomics of Heart and Lung Diseases.
Author information
Authors and Affiliations
Contributions
S.H., H.A., D.F.Q., P.J., and L.A.W. designed the study, reviewed the data, and wrote the manuscript. S.H., H.A., M.R., B.L., E.K., and M.S. oversaw and performed all experiments and data analysis. M.R., B.L., and E.K. wrote all the code. D.F.Q., Y.W., designed, optimized, and performed IMC protocols. S.D., L.D., D.R. provided additional data analysis and generated schematics. Clinical pathologists W.E., A.G., M.O., and P.J. curated the human tissue samples and built the patient TMAs, and collected and curated all clinical data. Y.B. performed genomic sequencing and analysis. All authors reviewed and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Xuanwen Bao and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Hartner, S., Abolfathi, H., Rezanejad, M. et al. Oncogenic driver mutations underlie the spatial tumour immune landscape of non-small cell lung cancer. Nat Commun 16, 8402 (2025). https://doi.org/10.1038/s41467-025-63465-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-63465-4