Abstract
Radial spokes (RSs) are conserved multimolecular structures attached to the axonemal microtubule doublets and are essential for the motility control of both cilia and sperm flagella. CFAP91, an RS3 protein, is implicated in human male infertility, yet its molecular function remains poorly understood. Here, we demonstrate that Cfap91 knockout (KO) mice exhibit impaired sperm flagellum formation and male infertility. Using a transgenic rescue model expressing FLAG- and BioID2-tagged CFAP91, we reveal that CFAP91 immunoprecipitates with RS3 proteins CFAP251 and LRRC23, whose localization is disrupted in Cfap91 KO sperm flagella. In addition, proximity labeling in mature spermatozoa identifies EFCAB5 as a sperm-specific CFAP91-proximal component. We show that Efcab5 KO males exhibit reduced sperm motility and fertility. Our findings establish CFAP91 as an essential scaffolder of RS3 assembly and EFCAB5 as a sperm-specialized movement regulator, advancing understanding of axonemal specialization in mammalian spermatozoa and its relevance to male infertility.
Similar content being viewed by others
Introduction
Flagella drive the movement of haploid spermatozoa in the female reproductive tract1, defects of which often lead to male infertility2,3. The main component of sperm flagella is the axoneme, a microtubule-based structure composed of two microtubule singlets (central pair) surrounded by nine peripheral microtubule doublets (doublet microtubules)4 (Fig. 1a). Based on the microtubules, associated proteinaceous structures such as dynein arms, radial spokes (RSs), and microtubule inner proteins are arranged5. Noteworthy, the axoneme is not only present in sperm flagella but also in motile cilia. Although both flagellar and ciliary axonemes are thought to share the same origin6, recent studies have identified that the sperm axoneme consists of extra structural elements in comparison to the ciliary axoneme5,7.
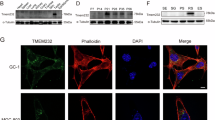
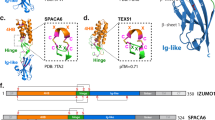
a A schematic diagram of the radial spoke in mouse sperm flagella and human respiratory cilia. CP central pair, DMT doublet microtubule, RS radial spoke. b Atomic model of RS3 components. Data was retrieved from the paper by Leung et al.35. c, d RT-PCR of Cfap91 utilizing cDNA from multiple mouse organs (c) or mouse postnatal testes (d). Actb was used as a loading control. BR brain, TH thymus, LU lung, HE heart, LI liver, SP spleen, KI kidney, TE testis, OV ovary. e A schematic drawing of the KO strategy of Cfap91. f Ratio of Cfap91+/+, Cfap91+/-, and Cfap91-/- mice obtained from the mating of Cfap91+/- mice in the B6D2 or B6D2/129 background. n = 89 for B6D2 background and n = 107 for B6D2/129 background. g Fertility tests of Cfap91+/+ and Cfap91-/- males. Data from each male mouse was individually color-coded. n = 30 plugs examined over 3 males for WT mice, n = 27 plugs examined over 3 males for Cfap91-/- mice. Data were presented as mean ± SD. h Sections of stage VII-VIII seminiferous tubules from Cfap91+/- and Cfap91-/- males. i Immunohistochemistry of stage II-III, stage VII-VIII, and stage X-XI seminiferous tubules from Cfap91+/- and Cfap91-/- males.
RSs are T-shaped structures extended from each of the nine doublet microtubules towards the central pair8. Broadly, RSs are critical for motility control in cilia and flagella9, and defects in the RS structure are linked with male infertility10,11,12. Each of the three RSs, namely RS1, RS2, and RS3, is composed of multiple proteins that form different shapes13, all of which, can be separated into head, neck, and stalk, according to the spatial localization of each compartment11,14 (Fig. 1a). Along the doublet microtubules, RS1, RS2, and RS3 are localized repeatedly in the same order in a 96 nm periodicity15,16,17. Remarkably, RS3 is evolutionarily less conserved between species and structurally diversified between cell types; RS3 is present in Chlamydomonas reinhardtii in a truncated form (RS3s)18. Moreover, the bridge structure linking RS2 to RS3 is only present in the sperm axoneme but not in the axoneme in other known cell types7 (Fig. 1a).
During the last phase of spermatogenesis, namely spermiogenesis, axonemes assemble on the centrosome and further attach to the haploid sperm nucleus19,20. Spermiogenesis can be divided into 16 steps in mice21, with axonemes forming during step 2–34. The formation of the axoneme largely depends on intraflagellar transport (IFT), which consists of large protein complexes that are responsible for bidirectional transport in cilia and flagella22,23. From step 8, the skirt-like manchette is observed in the caudal part of the sperm nucleus21, which functions in carving out the base of the sperm nucleus24. Anomalies in each of these two processes may lead to male infertility21,25.
A previous study has identified two biallelic variants of Cilia and Flagella Associated Protein 91 (CFAP91) in male infertile patients, which led to severe astheno-teratozoospermia, a condition that is accompanied by diminished sperm motility and abnormal sperm morphology26. The underlying mechanism of CFAP91 in regulating sperm flagellum biogenesis is not fully understood. According to the TreeFam database27, Cfap91 is conserved in 84 species out of 109 eukaryotes and is present in all mammals (40/40) (Supplementary Fig. 1a). Equally important, CFAP91 is also conserved in humans and major experimental animals (C. reinhardtii, Capra hircus, Drosophila melanogaster, Danio rerio, Mus musculus) (Supplementary Fig. 1b). In C. reinhardtii, FAP91 (ortholog of CFAP91) was found to form a complex with FAP251 (ortholog of CFAP251)28,29. This CFAP91-CFAP251 complex is found to be involved in the correct formation of RS3 in Tetrahymena thermophila29,30. Recently, by integrating artificial intelligence modeling and cryo-electron microscopy/tomography, CFAP91 has been shown to localize in RS3 and bind with CFAP251 in mouse ependymal cilia and human respiratory cilia31,32. Notably, according to a high-resolution study published recently, CFAP91 has been shown to exhibit a distinctive localization in bovine sperm axoneme: the N-terminal region of CFAP91 is localized from the RS2 base to the RS3 base through the base plate of the nexin-dynein regulatory complex (N-DRC) and is intertwined with the protofilament formed by CCDC39/4033,34. Meanwhile, the C-terminal region extends from the RS3 base to the RS3 neck35 (Fig. 1b). In the present work, by recruiting cell biological, proteomics, and genetic techniques, we revealed that depletion of CFAP91 led to abnormal recruitment of its binding partners and resulted in impaired sperm tail elongation. Moreover, we employed proximity labeling in mature spermatozoa and identified EFCAB5 as a sperm-specific CFAP91-proximal protein, in which deletion affected sperm motility.
Results
Generation of Cfap91 KO mice
To elucidate the organic expression profile of Cfap91, reverse transcription polymerase chain reaction (RT-PCR) was performed on multiple adult mouse tissues. Cfap91 exhibited a predominant expression in mouse testes with a weak expression in the lungs (Fig. 1c). RT-PCR using mouse testicular tissues from postnatal day 10 to day 35, demonstrated that Cfap91 started to express in mouse testes from postnatal day 14 (Fig. 1d). This period corresponds to the first presence of spermatocytes during the first wave of spermatogenesis36.
To understand the role of Cfap91 in spermatogenesis, we generated Cfap91 knockout (KO) mice. We electroporated Cas9 and two guide RNAs (gRNAs) that targeted the genomic region of Cfap91 into 64 zygotes obtained from mating (B6D2F1 x B6D2F1) (Fig. 1e). Next, we transferred 62 two-cell embryos into the oviduct of two pseudopregnant females and obtained 8 pups; 6 of which possess a large deletion in Cfap91. Subsequently, the Cfap91 mutant F0 mouse was mated with the B6D2F1 wild type (WT) mouse, which produced Cfap91+/- F1 mice; Cfap91+/- F1 mice were then caged with each other to obtain Cfap91-/- mice. Later, we confirmed there were 32,204 base pairs (bps) of deletion and 4 bps of insertion in the genomic region of Cfap91 (Supplementary Fig. 1c). This deletion was confirmed by genomic PCR (Supplementary Fig. 1d). Similar to the KO studies of some axonemal proteins1, we could not obtain Cfap91-/- mice according to the Mendelian ratio, only 5.6% of the pups were Cfap91-/- mice in the Cfap91 heterozygous mating of the B6D2 background (5/89 pups) (Fig. 1f). Even then, they all died before 8 weeks of age. As mutations in other Cfap genes were reported to cause hydrocephalus37,38, we hypothesized that Cfap91-/- mice in the B6D2 background may also carry hydrocephalus. An earlier report established that mice in the 129S6/SvEvTac (129) background have less susceptibility to hydrocephalus39. Therefore, we crossed B6D2 Cfap91+/- mice with 129 background WT mice. Mice produced from this mating are defined as B6D2/129 background mice. We caged B6D2/129 Cfap91+/- male and female in 3 individual cages and were able to obtain Cfap91-/- mice under the Mendelian ratio (28/107 pups, 26.2%) (Fig. 1f). Afterwards, we used the mice in the B6D2/129 background for further experiments.
Ablation of Cfap91 resulted in male infertility, accompanied by defective spermiogenesis
To test the fertility, we caged Cfap91-/- males with WT females and found that Cfap91-/- males were infertile in 3 trials (Fig. 1g). To discover why Cfap91-/- males were infertile, we first performed a gross inspection of their testes. Although there was no alteration in their body weight when compared with Cfap91+/- males (Supplementary Fig. 2a), the weight of Cfap91-/- testes was significantly decreased (Supplementary Fig. 2b, c). We then analyzed the cross-sections of Cfap91-/- testes. It should be noted that sections of seminiferous tubules can be categorized into 12 stages in mice according to their histology40. In stage VII-VIII, elongated spermatids are localized near the center of the seminiferous tubules, with tails aligning in the center of the lumens. In contrast, we could not find the sperm tails in the seminiferous tubules of Cfap91 KO males (Fig. 1h). To inspect sperm tail biogenesis in Cfap91-/- testes, we performed immunohistochemistry (IHC) on testis sections with an anti-acetylated tubulin antibody and found shorter sperm tails in Cfap91-/- mice (Fig. 1i). Furthermore, morphology of round spermatids was comparable between Cfap91+/- males and Cfap91-/- males, whereas elongating/elongated spermatids showed abnormal head morphology in Cfap91-/- males (Supplementary Fig. 2d). These data suggest that Cfap91 KO males undergo defective spermiogenesis.
Cfap91 KO males showed oligo-astheno-teratozoospermia
We then examined the epididymis and cauda epididymal spermatozoa of Cfap91-/- males. It showed that Cfap91-/- males had a smaller and significantly lighter epididymis (Supplementary Fig. 3a, b). When the cauda epididymal section was inspected, only a few sperm heads were found in Cfap91-/- males (Supplementary Fig. 3c). Consistently, fewer number of spermatozoa were found in the cauda epididymis of Cfap91-/- males (Supplementary Fig. 3d). Furthermore, spermatozoa from Cfap91-/- cauda epididymis showed abnormal morphology (Fig. 2a), including shorter tails (Fig. 2b) and abnormal heads (Fig. 2c). In addition, Cfap91 KO spermatozoa were completely immotile (Fig. 2d).
a Phase contrast images of cauda epididymal spermatozoa from Cfap91+/- and Cfap91-/- males. b Sperm tail length of cauda epididymal spermatozoa from Cfap91+/- and Cfap91-/- males. Data from each male mouse was individually color-coded. n = 267 spermatozoa examined over 3 males for Cfap91+/- mice, n = 300 spermatozoa examined over 3 males for Cfap91-/- mice. Data were presented as mean ± SD. An unpaired two-tailed t-test was performed for statistical analysis. P < 1.0E-15. c Ratio of the abnormal head of cauda epididymal spermatozoa from Cfap91+/- and Cfap91-/- males (n = 3 males for each genotype). Data were presented as mean ± SD. An unpaired two-tailed t-test was performed for statistical analysis. P = 1.5E-8. d Ratio of motile spermatozoa from the cauda epididymis of Cfap91+/- and Cfap91-/- males. Cfap91-/- males showed no motile spermatozoa (n = 5 males for each genotype). Data were presented as mean ± SD. e, f A schematic diagram of the EGFP-Tuba3a transgene driven by a Clgn promoter (e). The results of genomic PCR performed with the corresponding primers are shown (f). g Immunoblotting of EGFP in testes and cauda epididymal spermatozoa of WT and EGFP-Tuba3a TG males. ACTB was used as a loading control. h Imaging of manchette using EGFP-Tuba3a TG mice with Cfap91+/- and Cfap91-/- genotypes. Hoechst 33342 (H33342) was used to visualize the nuclei of spermatids. Images are arranged from left to right with the progression of spermiogenesis.
Since motile cilia also possess axonemes with radial spokes, we examined ependymal and tracheal motile cilia in Cfap91-/- mice (B6D2/129 background), using an anti-acetylated tubulin antibody. However, there were no apparent differences found between Cfap91+/- and Cfap91-/- mice (Supplementary Fig. 3e), indicating that ciliary elongation was endurable in Cfap91-/- mice in contrast to disrupted sperm flagellum formation.
As Cfap91 KO spermatozoa showed abnormal sperm head morphology, we attempted to visualize the morphological change of sperm heads together with the manchette. As the manchette mainly consists of TUBA3 (alpha-tubulin) and TUBB4 (beta-tubulin) among tubulin proteins41, we generated a transgenic (TG) mouse expressing TUBA3A fused with an N-terminal EGFP under testis-specific Clgn promoter (Fig. 2e). We confirmed that the transgene was incorporated into the genome by genomic PCR (Fig. 2f), and confirmed the expression of EGFP-TUBA3A by immunoblotting in testes but not in cauda epididymal spermatozoa lacking the manchette (Fig. 2g). We then crossed EGFP-Tuba3a TG mice with Cfap91 mutant mice to analyze the manchette-dependent morphological changes of sperm heads in Cfap91-/- males (Fig. 2h). For early elongating spermatids, no overt abnormalities were found in manchettes and sperm heads of Cfap91-/- mice. However, in later steps, Cfap91-/- males showed longer manchettes, accompanied by abnormal head morphology, indicating that disrupted sperm head morphogenesis was likely caused by abnormally elongated manchettes.
We then applied transmission electron microscopy (TEM) on testes and cauda epididymis to observe ultrastructural defects. In testis samples, Cfap91 KO spermatids showed defective microtubule structures (Fig. 3a–c). Cauda epididymal spermatozoa of Cfap91-/- males also showed disordered axoneme structures (Fig. 3d–h). In addition, abnormalities were found in the mitochondrial sheath, outer dense fiber, and fibrous sheath in Cfap91 KO spermatozoa (Fig. 3d–h). In late spermiogenesis, the electron-dense annulus structure migrates distally away from the sperm head, which marks the boundary of the midpiece and principal piece, and allows the formation of mitochondrial sheath42. However, in cauda epididymal spermatozoa from Cfap91-/- males, the annulus could not migrate far enough to the distal region (Fig. 3h). Overall, our results indicate that spermiogenesis was abnormal in Cfap91-/- males, which resulted in oligo-astheno-teratozoospermia.
a–f Cross-sections of spermatids from the testes (a–c), and spermatozoa in the cauda epididymis (d–f) of Cfap91+/- and Cfap91-/- males. g, h Longitudinal sections of spermatozoa in the cauda epididymis of Cfap91+/- and Cfap91-/- males. Genotypes of Cfap91 are indicated on the upper right of each figure. Abnormal proximal annulus found in Cfap91 KO spermatozoa is shown with black triangles.
Cfap91 transgene was able to rescue the fertility of Cfap91 -/- males
To confirm whether the infertility found in Cfap91-/- males was caused by the absence of Cfap91 but not by other factors, we generated Cfap91 TG mice. We tagged BioID243 and 3×FLAG in the C-terminus of CFAP91, where its expression was regulated by a testis-specific Clgn promoter (Fig. 4a). Hereby, mice carrying this transgene are referred to as Cfap91 TG mice. The genotype of Cfap91 TG mice was confirmed by genomic PCR (Supplementary Fig. 4a) and the protein expression of CFAP91-BioID2-3×FLAG was validated by immunoblotting (Fig. 4b). We then found that Cfap91-/- TG males showed a comparable sperm tail length, motile sperm rate, abnormal sperm head rate, and sperm count to Cfap91+/- males (Supplementary Fig. 4b–f). On top of that, Cfap91-/- TG males showed similar fertility when compared with WT males (Fig. 4c). These results indicate that the oligo-astheno-teratozoospermia of Cfap91-/- mice is indeed caused by the absence of CFAP91.
a A schematic diagram of the Cfap91-BioID2-3×FLAG transgene. Mice carrying this transgene are referred to as Cfap91 TG mice. b Immunoblotting of CFAP91-BioID2-3×FLAG with an anti-FLAG antibody using testes or cauda epididymal spermatozoa of WT and Cfap91-/- TG males. c Fertility tests on Cfap91+/+ and Cfap91-/- TG males. Data from each male mouse was individually color-coded. n = 23 plugs examined over 3 males for WT mice, n = 15 plugs examined over 3 males for Cfap91-/- TG mice. Data were presented as mean ± SD. An unpaired two-tailed t-test was performed for statistical analysis. d Immunohistochemistry on the sections of seminiferous tubules with an anti-FLAG antibody in WT and Cfap91-/- TG males. e Immunocytochemistry on spermatids obtained from WT and Cfap91-/- TG males with an anti-FLAG antibody. f Immunoblot analyses on sperm head-tail separated lysates. IZUMO1 and acetylated tubulin (Ac-TUB) served as controls of the head and tail fractions, respectively. g Immunoblot analyses on fractionated sperm lysates. BASIGIN (BSG), Ac-TUB, and AKAP4 served as controls for Triton-soluble, SDS-soluble, and SDS-resistant fractions, respectively.
CFAP91 is localized in sperm tails and fractionated with axonemal proteins
To examine the localization of CFAP91, we performed IHC using an anti-FLAG antibody to detect CFAP91-BioID2-3×FLAG in Cfap91-/- TG males. In testicular sections, CFAP91 signals were found in sperm tails (Fig. 4d). To better understand the spatiotemporal localization of CFAP91, we isolated germ cells from the testis and performed immunocytochemistry (ICC). We found that CFAP91 localizes in both sperm tail and cytoplasm during the round spermatid phase, with signals in the cytoplasm being largely diminished when entering the elongating spermatid phase (Fig. 4e). This localization pattern suggests that cytoplasmic CFAP91 is likely transported to sperm tails during spermiogenesis. CFAP91 signals were then found in whole tails of mature spermatozoa collected from Cfap91-/- TG cauda epididymis (Supplementary Fig. 4g). To confirm our findings, we performed immunoblotting on sperm head-tail separated lysates from Cfap91-/- TG mice. CFAP91 was detected in the tail lysate but not in the head lysate (Fig. 4f). We further questioned which compartment CFAP91 localizes to in sperm tails. We therefore performed sperm fractionation, in which sperm tails were fractionated into three compartments mainly containing transmembrane and cytosolic proteins, axonemal proteins, and outer dense fiber/fibrous sheath proteins (Triton-soluble, SDS-soluble, and SDS-resistant compartments, respectively)3,44. CFAP91 showed signals in the SDS-soluble compartment (Fig. 4g), suggesting that CFAP91 may be an axoneme-associated protein in spermatozoa.
CFAP91 immunoprecipitates with RS3 proteins during spermiogenesis
To explore the relationship between CFAP91 and other proteins, we performed immunoprecipitation-mass spectrometry (IP-MS) on the testis lysate of Cfap91-/- TG males using an anti-FLAG antibody (Supplementary Fig. 5a). We found that 45 proteins were significantly enriched in the CFAP91 immunoprecipitates (Fig. 5a and Supplementary Data 1). Among the immunoprecipitates, seven proteins, including CFAP91, AK7, AK9, CATIP, CFAP251, LRRC23 and MDH1B, have been shown to localize in RS3 of mammalian spermatozoa (Fig. 5a and Supplementary Data 2), consistent with CFAP91 localization in sperm RS335. We confirmed the presence of CFAP251 and LRRC23 in the CFAP91 immunoprecipitates with immunoblotting (Fig. 5b). Since the anti-LRRC23 antibody works for IP, association of CFAP91 and LRRC23 in testes was further confirmed by performing IP and subsequent immunoblotting for CFAP91-3xFLAG (Fig. 5c). During spermiogenesis, CFAP91 co-localizes with CFAP251 and LRRC23 in sperm tails (Supplementary Fig. 5b). To test if this CFAP91-CFAP251/LRRC23 complex is also present in the early stage of spermatogenesis, we performed co-IP on postnatal 15 days (P15) testes, before the appearance of haploid spermatids with tails36. Intriguingly, CFAP251 but not LRRC23 immunoprecipitated with CFAP91 in P15 testes (Fig. 5d and Supplementary Fig. 5c), suggesting that the CFAP91-CFAP251 complex is formed first during spermatogenesis.
a A volcano plot of the results from IP-MS studies using an anti-FLAG antibody. When comparing Cfap91-/- TG males to WT males, proteins with fold change >2 and P < 0.05 are considered as significantly upregulated, and dots are color-coded in magenta. An unpaired two-tailed t-test was performed for statistical analysis. b Immunoblotting was performed after IP with an anti-FLAG antibody. Signals of CFAP251, LRRC23, IFT140, and BBS2 were found in Cfap91-/- TG but not WT. IZUMO1 served as a loading control of inputs. c Immunoblotting was performed after IP with an anti-LRRC23 antibody or rat IgG, on Cfap91-/- TG testicular lysate. A band of FLAG was found in the IP product using the anti-LRRC23 antibody. d Immunoblotting was performed after IP using P15 testes from WT and Cfap91-/- TG males. A band of CFAP251 was found in Cfap91-/- TG but not WT, while bands of LRRC23 were not found in either Cfap91-/- TG or WT. Acetylated tubulin (Ac-TUB) served as a loading control for input lysates. e Immunohistochemistry of Cfap91+/- and Cfap91-/- testicular sections. LRRC23 and CFAP251 were co-localized with Ac-TUB, while this co-localization was not found in Cfap91-/- testicular sections. f Immunoblot analyses of testes and cauda epididymal spermatozoa from WT and Cfap91-/- males. Alpha-tubulin (α-TUB) served as a loading control.
We next questioned how the absence of Cfap91 impacted its immunoprecipitates and applied IHC on Cfap91-/- testis sections. We demonstrated that CFAP251 and LRRC23 could not localize in sperm tails during spermiogenesis in Cfap91-/- males (Fig. 5e). Similarly, the protein levels of CFAP251 and LRRC23 showed no alteration in Cfap91-/- testes in immunoblotting; however, they were downregulated in Cfap91 KO spermatozoa (Fig. 5f). In addition, IFT140, which is responsible for sperm tail elongation, was not present in WT epididymal spermatozoa45, but IFT140 remained in the spermatozoa of Cfap91-/- males (Fig. 5f), suggesting that sperm tail formation may halt in the absence of CFAP91. Overall, these data suggested that the localization of CFAP91-associating RS3 proteins was disrupted in sperm tails of Cfap91-/- males.
CFAP91 immunoprecipitates with BBSome proteins during spermiogenesis
In addition to RS3 proteins, 5 members of the Bardet-Biedl syndrome (BBS) protein family, BBS1, BBS2, BBS5, BBS7 and BBS9, were detected in CFAP91 immunoprecipitates (Fig. 5a, b). Previous reports indicated that BBS1/2/4/5/7/8/9/18 form a complex called BBSome and function in protein transport in cilia/flagella46. KO of individual members of BBSome has been shown to cause the shortening of sperm flagella in mice47,48,49,50,51. When comparing with our previous IP-MS results of MYCBPAP, a central pair apparatus protein essential for sperm tail elongation52, these BBS proteins were only found in CFAP91 immunoprecipitates (Supplementary Fig. 5d). Furthermore, in Cfap91+/- testicular cells, BBS2 was present in the cytoplasm and along the sperm tails. Meanwhile, in Cfap91-/- testicular cells, the signal of BBS2 was not distributed evenly and formed foci in the cytoplasm (Supplementary Fig. 5e). These results suggest that CFAP91 may exhibit its effect on flagellum formation with BBS proteins.
Proximity labeling in mature spermatozoa was able to detect EFCAB5 as a sperm-specific RS3 protein
Exploring protein bindings for sperm axonemal proteins in mature spermatozoa has been challenging, as typical lysis buffers that can solubilize axonemes may alter protein-protein interactions by changing the conformation of proteins53. To explore the interacting proteins of CFAP91, we applied proximity labeling in mature spermatozoa. For proximity labeling, we tagged BioID2 to CFAP91, which biotinylates the proteins surrounding CFAP91 in a radius of 10 nm in the presence of biotin43. Mature spermatozoa from Cfap91-/- TG cauda epididymis were cultured in a medium with biotin for 16 h to let BioID2 biotinylate its proximate proteins, then spermatozoa were lysed in a 0.4% SDS sample buffer. Subsequently, the biotinylated proteins were pulled down by streptavidin (SA) and analyzed by MS (Fig. 6a). To follow up, immunoblot analyses showed that CFAP91-BioID2-3×FLAG was pulled down by SA (Fig. 6b and Supplementary Data 3), indicating that self-biotinylation occurred. Twenty-four proteins were exclusively detected in the pulldown product of Cfap91-/- TG mice; among them, CFAP91 and EFCAB5 were identified with significantly more spectra than all proteins detected (Fig. 6c). We confirmed that EFCAB5 was pulled down by SA in mature spermatozoa of Cfap91-/- TG mice by immunoblotting (Fig. 6b). In addition, proximity labeling identified not only two RS3 proteins, CFAP251 and AKAP14, but also an RS1/2 protein, RGS22 (Fig. 6c, d), which was predicted to interact with EFCAB5 in the RS2-RS3 bridge of bovine spermatozoa35 (Fig. 1a, b).
a A schematic drawing of the application of BioID2 in mature spermatozoa. Spermatozoa from WT and Cfap91-/- TG males were incubated in a medium supplemented with biotin for 16 h. Collected spermatozoa were lysed, and biotinylated proteins were pulled down by streptavidin (SA). b Immunoblotting was performed after the pull-down of biotinylated proteins in WT and Cfap91-/- TG males. EFCAB5 was not detected in the input, likely due to low protein abundance and/or low solubilization efficiency (0.4% SDS compared to 1% SDS or 6 M urea in other experiments). IZUMO1 served as a loading control for inputs. c Molecular weight and total spectra of twenty-four proteins only identified in Cfap91 KO TG spermatozoa. CFAP91 and EFCAB5 were identified with the most peptides among all twenty-four proteins. A linear regression line with 95% confidence bands is shown. Slope = 0.06 and Y-intercept = 5.38. d Venn diagram of CFAP91 immunoprecipitates, proximity labeling-identified proteins, and mammalian sperm RS3 proteins. e RT-PCR of Efcab5 utilizing cDNA from multiple mouse organs. Actb was used as a loading control. f Immunoblot analyses of human spermatozoa. g immunoblotting using testes and cauda epididymal spermatozoa from WT and Cfap91-/- males with an anti-EFCAB5 antibody. Alpha-tubulin (α-TUB) served as a loading control. h Fractionation using WT spermatozoa. Signals of EFCAB5, LRRC23, and CFAP251 were found in the SDS-soluble fraction.
Efcab5 is predominantly expressed in mouse testes (Fig. 6e). When we compared the expressions of Efcab5, Cfap91, Cfap251 and Lrrc23 in the testis with other abundantly expressed tissues (lung and brain), we found that Efcab5 showed significantly higher expression ratio compared to other genes (Supplementary Fig. 6a). EFCAB5 is conserved among all mammals, 47 out of 58 Chordata, but it is missing in all of the Arthropoda included in TreeFam database27 (Supplementary Fig. 6b). Moreover, EFCAB5 is not conserved in C. reinhardtii or T. thermophila. We confirmed that EFCAB5 exists in human spermatozoa by immunoblotting (Fig. 6f). Furthermore, immunoblot analyses revealed that EFCAB5 was absent in Cfap91 KO spermatozoa (Fig. 6g). When sperm fractionation analyses were performed on WT mouse spermatozoa, EFCAB5, CFAP251, and LRRC23 all showed signals in the axoneme compartment like CFAP91 (Fig. 6h). These results indicate that EFCAB5 is a sperm-specific RS3 protein in mice.
EFCAB5 is vital for sperm motility
According to a previous human male infertility study, EFCAB5 expression is significantly downregulated in the testes of ejaculatory azoospermia patients54, suggesting that EFCAB5 may play a role in human spermatogenesis. To verify its role in male fertility, we generated Efcab5-/- mice. We electroporated Cas9 and two gRNAs that targeted the genomic region of Efcab5 into 64 zygotes obtained from mating (B6D2F1 x B6D2F1) (Fig. 7a). We transferred 56 two-cell embryos into the oviduct of two pseudopregnant females and obtained 13 pups, 2 of which possess a large deletion in Efcab5. By subsequent mating of F0 mice, Efcab5-/- mice were obtained, with a large deletion in the genomic region of Efcab5 confirmed (Supplementary Fig. 7a and b). It was observed that Efcab5-/- males showed a slightly decreased fertility in mating test (Fig. 7b). No apparent anomalies were confirmed in the morphology of male reproductive organs (Supplementary Fig. 7c–g) or the mature spermatozoa in Efcab5-/- males (Fig. 7c). In contrast to Cfap91-/- males, CFAP251 and LRRC23 were present in the testes and spermatozoa of Efcab5-/- males (Fig. 7d).
a A schematic drawing of the KO strategy of Efcab5. b Fertility tests of Efcab5+/+ and Efcab5-/- males. Data from each male mouse was individually color-coded. n = 46 plugs examined over 5 males for WT mice, n = 48 plugs examined over 5 males for Efcab5-/- mice. Data were presented as mean ± SD. An unpaired two-tailed t-test was performed for statistical analysis. P = 8.5E-8. c Phase contrast images of cauda epididymal spermatozoa from WT and Efcab5-/- males. d Immunoblotting of testes and cauda epididymal spermatozoa from WT and Efcab5-/- males, with anti-EFCAB5, anti-CFAP251, anti-LRRC23, and anti-RGS22 antibodies. ACTB served as a loading control. e Ratio of motile and progressive spermatozoa in WT and Efcab5-/- males after 10 min and 120 min of incubation in a capacitation medium. n = 5 males for each genotype. Data were presented as mean ± SD. An unpaired two-tailed t-test was performed for statistical analysis. P = 2.5E-04 for progressive motility after 120 min of incubation. f VCL, VSL, and VAP of cauda epididymal spermatozoa of WT and Efcab5-/- males after 10 min and 120 min of incubation in a capacitation medium. n = 5 males for each genotype. Data were presented as mean ± SD. An unpaired two-tailed t-test was performed for statistical analysis. g α-angle of cauda epididymal spermatozoa in WT and Efcab5-/- males after 10 min and 120 min of incubation in a capacitation medium. Data from each male mouse was individually color-coded. n = 45 spermatozoa examined over 3 males for each genotype and time point. Data were presented as mean ± SD. One-way ANOVA and Tukey’s multiple comparisons test with adjustment were used for statistical analysis. P = 7.0E-14 for WT mice between 10 min and 120 min of incubation and P = 7.1E-14 for 120 min of incubation between WT and Efcab5-/- mice.
When we analyzed the motility of Efcab5 KO spermatozoa (Supplementary Movie 1), we found a significant decrease in the percentages of both motile and progressive spermatozoa after 10 min and 120 min of incubation in capacitation medium (Fig. 7e). Moreover, curvilinear velocity (VCL), straight line velocity (VSL), and average path velocity (VAP) were decreased after 10 min of incubation; VSL and VAP were decreased after 120 min of incubation compared to WT spermatozoa (Fig. 7f). To test if the sperm motility pattern was also defective in Efcab5-/- males, we analyzed maximal bending angles of flagella relative to sperm heads (α-angle), which is used to quantitatively analyze hyperactivation55,56. Consistent with earlier reports, the α-angle of the control spermatozoa increased after 2 h of incubation in the capacitation medium, compared to 10 min of incubation55,56. However, this increase of α-angle was not seen in Efcab5 KO spermatozoa (Fig. 7g), suggesting that the decreased fertility of Efcab5-/- males is caused by abnormal flagellar motility patterns.
As EFCAB5 localizes in the sperm-specific RS2-RS3 bridge, we propose that EFCAB5 may enable sperm-specific regulation of flagellar motility in this structure. In bovine spermatozoa, the RS2-RS3 bridge structure is formed by EFCAB5 and RGS22. It has been suggested that binding of EFCAB5 to RGS22 in RS2 may induce an intensive conformational change of RGS2235. We then analyzed the amount and localization of RGS22 in Efcab5-/- testes, but no overt abnormalities were found (Fig. 7d and Supplementary Fig. 8a). EFCAB proteins possessed one or multiple EF-hand calcium-binding domains, and we found that EFCAB5 was able to bind with Ca2+ in this domain according to AlphaFold predictions57 (Supplementary Fig. 8b). Moreover, Ca2+ influx occurs during capacitation to regulate downstream functions for hyperactivation58,59. To test if Ca2+ concentration can alter the EFCAB5-RGS22 binding, we expressed EFCAB5 and RGS22 in HEK293T cells and performed immunoblotting following IP in the presence and absence of Ca2+. As a result, RGS22 immunoprecipitated with EFCAB5 despite the concentration of Ca2+ (Supplementary Fig. 8c), indicating that EFCAB5 can bind to RGS22 independently of Ca2+.
Discussion
Previously, human male patients lacking functional CFAP91 have been reported as infertile26. However, the role of Cfap91 in spermatogenesis has not been fully discovered. In this study, we employed the CRISPR/Cas9 system to generate Cfap91-/- mice and subsequently demonstrated that Cfap91 is also essential for murine male fertility (Fig. 1g). Cfap91-/- mice exhibited abnormal sperm flagellum elongation (Fig. 1h, i), which resulted in short and immotile tails (Fig. 2a, b, d). Ultrastructural analyses on testicular and cauda epididymal spermatozoa showed that the integrity of sperm axonemes in Cfap91-/- males was damaged (Fig. 3a–f). As Cfap91 KO spermatozoa showed abnormal flagellum elongation, we could not identify if CFAP91 plays a role in regulating sperm flagellar motility. However, inactivation of Cfap91 orthologs in C. reinhardtii, Trypanosoma brucei, and T. thermophila resulted in decreased mobility of flagella26,30,60. These earlier studies suggest that CFAP91 may also function in murine sperm motility regulation.
Imaging of manchette is usually performed by ICC or IHC using an anti-tubulin antibody61; yet, in this study, we generated EGFP-Tuba3a TG mice and demonstrated its usage in Cfap91-/- mice. As expected, EGFP-Tuba3a TG mice showed fluorescence in the manchette, and we were able to clarify that the abnormal sperm head shape was associated with abnormal manchette in Cfap91-/- males (Fig. 2h). Several KOs of sperm axonemal proteins that do not localize in the manchette exhibited defective manchette-mediated head morphogenesis12,62, which suggests that CFAP91 may not be directly involved in the manchette elongation as well. Rather, the abnormal manchette formation may be a secondary effect of impaired flagellum formation. Intriguingly, EGFP-TUBA3A is not localized in sperm flagella (Fig. 2g), even though TUBA3A is considered a component of the sperm doublet microtubules5. This may be because N-terminally fused EGFP inhibits the uptake of TUBA3A into the doublet microtubules. Overall, the EGFP-Tuba3a TG mouse is a powerful tool for understanding manchette-mediated sperm head morphogenesis.
To understand the molecular function of CFAP91, we generated Cfap91 TG mice. After performing IP-MS, we demonstrated that 45 proteins were significantly upregulated in CFAP91 immunoprecipitates (Fig. 5a). Of these 45 proteins, we especially focused on CFAP251 and LRRC23. Previously, CFAP251 orthologs were found to interact with CFAP91 orthologs in C. reinhardtii and T. thermophila28,29. This interaction was found to be conserved in mouse ependymal cilia and human respiratory cilia31,32. In these four species, CFAP251 was hypothesized to be localized at the base of RS3, suggesting that the localization of CFAP251 in the RS3 stalk is conserved from protists to mammals. CFAP251 was downregulated in the spermatozoa of human patients carrying a pathogenic variant in CFAP9126. Furthermore, human male infertile patients carrying CFAP251-deficient alleles were found to have shorter sperm tails63, indicating that CFAP251 is essential for sperm flagellum formation. Hence, depletion of CFAP91 disabled the incorporation of CFAP251 into the sperm flagella (Fig. 5e), which may be the essential reason that Cfap91-/- males produced shorter sperm tails. In contrast, LRRC23 truncation has been shown to cause the loss of the RS3 head in mouse spermatozoa without overt abnormalities in flagellum elongation11. Recently, structural analyses on bovine spermatozoa reported that LRRC23 localizes in the head of RS3 and was supported by the C-terminus of CFAP91 that localizes along the RS3 neck35 (Fig. 1b). We confirmed that LRRC23 was pulled down by the C-terminal half of CFAP91 (Supplementary Fig. 8d). However, the physical interaction between CFAP91 to LRRC23 was not found in bovine sperm RS335, suggesting that the interaction between CFAP91 and LRRC23 is indirect.
Interestingly, in C. reinhardtii, a shorter RS3 can be formed only by orthologs of CFAP61/91/25131, still they are enough for flagellum elongation. Furthermore, in mammalian RS3, only the absence of CFAP61/91/25126,62,63,64 impairs sperm tail length, but not other known RS3 members (AK965, AKAP1466, LRRC2311,67, LRRD168, PKA69,70, STYXL171; it is not clear for Ak7-/- males since the study was conducted at 42-day-old mice, and full-size sperm tails may be formed in adult mice72), suggesting that the basic unit sufficient for flagellum elongation may be CFAP61/91/251 in mouse spermatozoa. In the P15 testes before the appearance of sperm tails, we were able to immunoprecipitate CFAP251 but not LRRC23 with CFAP91 (Fig. 5d). This conveys that CFAP251-CFAP91 may be assembled as an integral part of the axoneme elongation. In contrast, LRRC23, which tunes motility, may later be assembled on the C-terminus region of CFAP91 that extends along the RS3 neck. It should be noted that we were also able to detect CFAP61 in IP-MS samples with high fold change, although it was not statistically significant (Supplementary Data 1; P = 0.13).
Based on our results that indicate CFAP91 localization to the mouse sperm RS3, we performed proximity labeling experiments using mature spermatozoa, which led us to identify EFCAB5 with high peptide numbers (Fig. 6c). When we were preparing this manuscript, an atomic model of bovine sperm RS3 was published35, which also indicates the localization of EFCAB5 in the RS3 (Fig. 1b). We also detected EFCAB5 with IP-MS using Cfap91 TG mice, but it was not statistically significant compared to control WT mice (Supplementary Data 1; P = 0.08). The reason why EFCAB5 was not significantly enriched in CFAP91 immunoprecipitates (Fig. 5a) could be multifactorial. The most likely cause may be the existence of numerous types of spermatogenic cells in testicular IP-MS analyses. The highly organized axoneme structures are formed in stages, and EFCAB5 may attach to the basic structure of the RS3 in later stages, in which the flagella are highly compacted and not soluble in the IP lysis buffer. It is possible that EFCAB5 is evolutionarily appended to the RS3 basic structure when RS3 evolved from its ancestors, consistent with EFCAB5 being a critical component for sperm-specialized movement. Our analyses using HEK293T cells confirmed that the interaction of EFCAB5 and RGS22, an RS2 protein, is independent of Ca2+ (Supplementary Fig. 8c). The constant connection of RS2 and RS3 through RGS22/EFCAB5 may be important for the regulation of sperm-specialized movement.
In summary, this study suggests that CFAP91 localizes to sperm RS3 and is involved in sperm tail biogenesis by recruiting CFAP251 into the assembling axoneme. Moreover, we report an EGFP-Tuba3a TG mouse line that can be used for the live imaging of the manchette. This study also demonstrates proximity labeling in mature spermatozoa, which has the advantage of detecting adjacent structural proteins in axonemes that are difficult to dissolve in mild lysis buffers. Our proximity labeling analysis indicates that EFCAB5 is a sperm axoneme-specific RS3 protein that is vital for sperm motility. This study widens the understanding of etiology and pathobiology of human male infertility.
Methods
Maintenance and generation of mice
All mouse experiments were approved by the Animal Care and Use Committee at the Research Institute for Microbial Diseases, Osaka University (Approval numbers: #Biken-AP-H30-01 and #Biken-AP-R03-01). BDF1 and ICR mice were purchased from Japan SLC (Shizuoka, Japan) and CLEA Japan (Tokyo, Japan). Mice in the 129×1/SvJmsSlc background were purchased from Japan SLC. All mice were housed in a specific-pathogen-free animal facility and were able to access unlimited food and water voluntarily. The animal room was maintained on a 12-h light/12-h dark cycle. Male mouse experiments were conducted when they reached 8 weeks of age.
To generate KO mice with the CRISPR/Cas9 system, pairs of gRNAs were designed to target the genomic region of Cfap91 and Efcab5. Each gRNA was incubated with Cas9 (Thermo Fisher Scientific, Waltham, MA) and trans-activating crRNA (tracrRNA) (Sigma-Aldrich, St Louis, MO) to induce the formation of Cas9-gRNA ribonucleoprotein (RNP). RNPs are subsequently electroporated into mouse zygotes, which are obtained from the natural mating of BDF1 males with superovulated BDF1 female mice. For the generation of EGFP-Tuba3a TG mice, EGFP-Tuba3a cDNA was cloned into an expression vector containing a Clgn promoter (Addgene #173686). For Cfap91 TG, Cfap91 cDNA was inserted into CAG-MCS-BioID2-3×FLAG plasmid73 (Addgene #186812). Then, the CAG promoter was replaced by a Clgn promoter74 (Addgene #173686). Both plasmids were linearized by ClaI and PacI (New England Biolabs, Essex, MA), and pronuclear injection of linearized plasmids was performed on mouse zygotes. In both cases of generating KO and TG mice, the zygotes were cultured to the two-cell stage in potassium-supplemented simplex optimized medium75 (KSOM) after electroporation or injection and then transplanted into the ampulla of pseudopregnant ICR female mice. Sanger sequencing and genomic PCR of offspring obtained from pseudopregnant females were performed to check their genotype. Genomic PCR was performed with the following conditions for 40 cycles: 94 °C for 30 s, 65 °C for 30 s, 72 °C for 30 s (except for F3–R3, as the last step was extended to 120 s). The sequences of gRNAs and primers are listed in Supplementary Data 4.
RT-PCR
Total RNA from various tissues and testicular RNA in postnatal testes were extracted by TRIzol (Thermo Fisher Scientific) and reversed transcribed by SuperScript III First-Strand Synthesis System (Thermo Fisher Scientific). Products from reverse transcription were used as templates for PCR. PCR cycles were set in the following conditions: 94 °C for 30 s, 60 °C for 30 s, and 72 °C for 60 s (For Actb, the last step was set to 30 s). The sequences of primers used for RT-PCR are listed in Supplementary Data 4.
Fertility test
For the Cfap91 KO and Cfap91 TG mouse lines, the fertility (pups/plugs) of three males was tested. For the Efcab5 KO mouse line, the fertility of five KO males was tested. Each male was housed with three WT females for 11 weeks, and the presence of vaginal plugs and the number of offspring were counted every weekday.
Histological analysis
Testis and epididymis specimens were immersed in Bouin’s Fixative (Polysciences, Inc., Warrington, PA) for 6 h at room temperature. After gradual dehydration, specimens were embedded in a paraffin block and sliced into sections with a thickness of 5 μm. Sections were dewaxed and treated with 1% periodic acid (Nacalai Tesque, Kyoto, Japan) for 10 min, Schiff’s reagent (Wako, Osaka, Japan) for 20 min, and Mayer’s Hematoxylin Solution (Wako) for 30 s, with a brief wash by tap water after each step. Sections were mounted with Entellan new (Sigma-Aldrich) and observed by an Olympus BX-53 microscope (Tokyo, Japan).
Immunohistochemistry (IHC) and immunocytochemistry (ICC)
For IHC, tissues were immersed in 4% paraformaldehyde (PFA; Thermo Fisher Scientific) for 6 h at 4 °C, and were transferred to 15% and 30% sucrose for a day at 4 °C before the samples were embedded with Optimal Cutting Temperature (OCT) Compound (Sakura Finetek, Tokyo, Japan). Specimens were then sliced into 8 μm-thick sections and briefly washed with phosphate-buffered saline (PBS). Then, the specimens were treated with 0.1% Triton X-100 (Nacalai Tesque) in PBS and were blocked with 3% bovine serum albumin (BSA; Sigma-Aldrich) in PBS. Then, the specimens were incubated overnight at 4 °C with primary antibodies that were diluted in 3% BSA in PBS (Supplementary Data 5). On the following day, the specimens were washed with PBS and incubated for 2 h with secondary antibodies that were diluted in 3% BSA in PBS. Finally, the specimens were washed with PBS, incubated with 0.05% of Hoechst 33342 (Thermo Fisher Scientific), and mounted with Epredia Immu-Mount (Thermo Fisher Scientific) prior to the observation under an Olympus BX-53 microscope or a Nikon C2 Eclipse Ti microscope (Nikon, Tokyo, Japan).
For ICC, spermatozoa and spermatids were spread onto slides and were air-dried before fixation with 4% PFA for 15 min at room temperature. The slides were washed with PBS, blocked, probed with antibodies, and mounted as described in IHC.
Sperm parameter inspection
To measure sperm tail length, the ratio of abnormal sperm heads, and sperm count, we released mature spermatozoa from the cauda epididymis into PBS using fine forceps and scissors. Spermatozoa were then spread onto a slide, and abnormal sperm heads were inspected and photographed using an Olympus BX-53 microscope. Tail lengths were analyzed using Fiji76. Spermatozoa were further diluted in water for immobilization, and the number of spermatozoa was counted using a hemocytometer.
For sperm motility measurements, cauda epididymal spermatozoa were released into TYH capacitation medium77 and incubated at 37 °C under 5% CO2 in air. For Cfap91-/- males, we measured sperm motility after only 10 min of incubation, as spermatozoa lost motility completely. For Efcab5-/- males, spermatozoa were incubated for 10 min and 120 min, which represents before and after capacitation78. We analyzed sperm motility using the CEROS II sperm analysis system (software version 1.5; Hamilton Thorne Biosciences, Beverly, MA). For the analysis of the α-angle of spermatozoa, we captured videos of sperm movement with an Olympus BX-53 microscope equipped with a high-speed camera HAS-L1 (Ditect, Tokyo, Japan). The video was recorded at 200 frames per second. The α-angle of spermatozoa was analyzed using Fiji76.
Imaging of manchette using EGFP-Tuba3a TG male mice
Seminiferous tubules of EGFP-Tuba3a TG males were released by breaking the tunica albuginea. Seminiferous tubules were gently pipetted to remove the attachment of Leydig cells and were squeezed by fine forceps to release the spermatids into PBS. Spermatids were then stained with 0.1% Hoechst 33342 and imaged with an Olympus BX-53 microscope.
Ultrastructural analysis
Ultrastructural analysis by TEM was performed as previously described79. In brief, anesthetized male mice underwent perfusion fixation using 4% PFA, after which the testes were further fixed in 4% PFA. The samples were fixed again with 1% glutaraldehyde. Further, the testes were immersed in a solution of 1% osmium tetroxide (OsO4) and 0.5% potassium ferrocyanide. Following dehydration with ethanol, the specimens were incubated in propylene oxide (PO) before being placed in a mixture of PO and epoxy resin. This mixture was subsequently replaced with pure epoxy resin. Ultrathin sections were cut and stained, and the images were captured using a JEM-1400 Plus electron microscope (JEOL, Tokyo, Japan) operating at 80 kV, equipped with a Veleta 2 K × 2 K CCD camera (Olympus).
Immunoblotting (IB)
IB was performed as previously described80. In the case that the lysis buffer was not specified, testicular proteins and sperm proteins were lysed by Triton lysis buffer [1% Triton X-100, 50 mM NaCl, 20 mM Tris-HCl, 1× protease inhibitor cocktail (Nacalai Tesque)] and urea lysis buffer (6 M urea, 2 M thiourea, 2% sodium deoxycholate), respectively. For human spermatozoa, samples were obtained from fertile donors with their consent. The experiments using human samples were approved by the Ethics Committee of the Research Institute for Microbial Diseases, Osaka University (#28-4-2). The uncropped and unprocessed scans of IB are provided in the Source Data file.
Sperm head-tail separation
Cauda epididymal spermatozoa were collected in PBS and sonicated by Branson SLPe Digital Sonifier (Branson Ultrasonics, Brookfield, CT). Sonicated samples were gently transferred to the top of 90% Percoll (GE Healthcare, Chicago, IL) in PBS and centrifuged to separate sperm heads and tails. Finally, the separated heads and tails were resuspended in a sample buffer (60 mM Tris-HCl pH 6.8, 2% SDS, 10% glycerol, 0.025% Bromophenol Blue) and incubated at 95 °C for 5 min for protein extraction. A detailed protocol can be found in the previous report3.
Sperm fractionation
Sperm fractionation was performed as previously reported52. In brief, cauda epididymal spermatozoa were treated in the order of Triton lysis buffer, 1% SDS lysis buffer (1% SDS, 75mM NaCl, 24 mM EDTA, 1× protease inhibitor cocktail), and sample buffer, at 4 °C for 2 h, room temperature for 1 h, and 95 °C for 5 min, respectively. Samples were centrifuged, and the supernatant was taken and replaced with subsequent lysis buffer after the first and second protein extraction.
Cell maintenance and transfection
HEK293T cells81 were cultured in Dulbecco’s Modified Eagle Medium (DMEM; Thermo Fisher Scientific) supplemented with 10% fetal bovine serum (FBS; Sigma-Aldrich) and 1% penicillin and streptomycin (Thermo Fisher Scientific) at 37 °C under 5% CO2 in air and was subcultured every 3 or 4 days. Transfection of cells was carried out by the calcium phosphate-DNA precipitate method when cells reached 70–80% confluency. Cells were harvested 24 h post-transfection and were subjected to subsequent experiments.
Immunoprecipitation (IP)
Magnetic beads-based IP was performed. Cell lysates were subjected to IP using the Dynabeads Co-Immunoprecipitation Kit (Thermo Fisher Scientific). IP was carried out under the recommended protocol of the manufacturer. The IP product was subjected to immunoblotting or MS analyses.
Antibodies
Antibodies used in this study are indicated in Supplementary Data 5. Antibodies against BASIGIN82, EGFP83, IZUMO184, LRRC2367 were generated in the previous studies. The antibody against EFCAB5 was generated in this study against the sequence ‘CGSRRGSGTDQGQHRGSV’. The Anti-1D485 antibody was a gift from Dr. Martin M. Matzuk.
Mass spectrometry
Proteins were analyzed by nanocapillary reversed-phase liquid chromatography-tandem mass spectrometry (LC-MS/MS). The analysis was performed using a C18 column (IonOpticks, Victoria, Australia) integrated into a nanoLC system (Bruker Daltonics, Billerica, MA), which was coupled to a timsTOF Pro mass spectrometer (Bruker Daltonics) and the CaptiveSpray nano-electrospray ion source (Bruker Daltonics). The raw data were processed using DataAnalysis software (Bruker Daltonics), and protein identification was carried out using MASCOT (Matrix Science, Tokyo, Japan) with the SwissProt database. Quantitative values and fold changes were subsequently calculated using Scaffold 5 (Proteome Software, Portland, OR).
Proximity labeling of cauda epididymal spermatozoa
Cauda epididymal spermatozoa were collected into TYH medium supplemented with 250 μM biotin (Sigma-Aldrich) and incubated at 37 °C under 5% CO2 in air for 16 h. Subsequently, spermatozoa were collected and washed with PBS twice. Protein extraction from spermatozoa was performed according to earlier reports43, with several modifications. SDS sample buffer [50 mM Tris (pH 7.4), 500 mM NaCl, 0.4% SDS, 1 mM dithiothreitol (DTT), and 1× protease inhibitor cocktail] was used as a lysis buffer. After incubating spermatozoa in the lysis buffer for 1 h at 4 °C, Triton X-100 was added to the lysis buffer till Triton X-100 reached 2% for the final concentration. Tubes were then intensely vortexed for 2 min. Then, 50 mM Tris (pH 7.4) was added to the tube until the final concentration of Triton X-100 reached 1%. Samples were then briefly vortexed and centrifuged at 15,000 × g for 20 min, and the supernatant was collected and incubated with Pierce Streptavidin Magnetic Beads (Thermo Fisher Scientific) overnight at 4 °C. After incubation, beads were washed with wash solutions three times for streptavidin pull down. The recipe for wash solutions can be found in the indicated paper86. Followed by the three washes, 2× SDS sample buffer was incubated with the beads at 95 °C for 5 min to elute the proteins attached to the beads. Eluates were then analyzed by immunoblotting and MS.
Structural analysis
Cryo-EM density maps and fitted models were visualized by UCSF ChimeraX87.
Statistical analysis and reproducibility
All experiments were performed at least three times with similar results. Statistical analysis was performed using Graphpad Prism 9 (GraphPad Software, MA, USA). Data are all indicated as mean ± SD. An unpaired two-tailed t-test was performed for statistical analyses unless otherwise specified. The level of significance was set at P < 0.05, and the P values were directly indicated in the figures.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The authors declare that the data that support the findings of this study are available from the corresponding author upon request. Proteomic data is available with accession codes JPST003879 and PXD065243. Cryo-EM density maps and fitted model were retrieved from previous studies with accession codes EMDB-35888 [https://www.ebi.ac.uk/emdb/EMD-35888]31, EMDB-50664 [https://www.ebi.ac.uk/pdbe/entry/emdb/EMD-50664] and 9FQR35. Source data are provided with this paper.
References
Miyata, H., Morohoshi, A. & Ikawa, M. Analysis of the sperm flagellar axoneme using gene-modified mice. Exp. Anim. 69, 374–381 (2020).
Yuan, S. et al. SPATA6 is required for normal assembly of the sperm connecting piece and tight head-tail conjunction. Proc. Natl. Acad. Sci. USA 112, E430–E439 (2015).
Castaneda, J. M. et al. TCTE1 is a conserved component of the dynein regulatory complex and is required for motility and metabolism in mouse spermatozoa. Proc. Natl. Acad. Sci. USA 114, E5370–E5378 (2017).
Miyata, H., Shimada, K., Kaneda, Y. & Ikawa, M. Development of functional spermatozoa in mammalian spermiogenesis. Development 151, dev202838 (2024).
Zhou, L. et al. Structures of sperm flagellar doublet microtubules expand the genetic spectrum of male infertility. Cell 186, 2897–2910.e19 (2023).
Khan, S. & Scholey, J. M. Assembly, functions and evolution of archaella, flagella and cilia. Curr. Biol. 28, R278–R292 (2018).
Leung, M. R. et al. The multi-scale architecture of mammalian sperm flagella and implications for ciliary motility. EMBO J. 40, e107410 (2021).
Curry, A. M. & Rosenbaum, J. L. Flagellar radial spoke: a model molecular genetic system for studying organelle assembly. Cell Motil. Cytoskeleton 24, 224–232 (1993).
Zhu, X., Liu, Y. & Yang, P. Radial spokes—a snapshot of the motility regulation, assembly, and evolution of cilia and flagella. Cold Spring Harb Perspect Biol. 9, a028126 (2017).
Zhang, X. et al. Differential requirements of IQUB for the assembly of radial spoke 1 and the motility of mouse cilia and flagella. Cell Rep. 41, 111683 (2022).
Hwang, J. Y. et al. LRRC23 truncation impairs radial spoke 3 head assembly and sperm motility underlying male infertility. Elife 12, RP90095 (2023).
Abbasi, F. et al. RSPH6A is required for sperm flagellum formation and male fertility in mice. J Cell Sci 131, jcs221648 (2018).
Pigino, G. & Ishikawa, T. Axonemal radial spokes: 3D structure, function and assembly. Bioarchitecture 2, 50 (2012).
Grossman-Haham, I. et al. Structure of the radial spoke head and insights into its role in mechanoregulation of ciliary beating. Nat. Struct. Mol. Biol. 2020 28:1 28, 20–28 (2020).
Dentler, W. L. & Cunningham, W. P. Structure and organization of radial spokes in cilia of Tetrahymena pyriformis. J. Morphol. 153, 143–151 (1977).
Leung, M. R. et al. Structural specializations of the sperm tail. Cell 186, 2880–2896.e17 (2023).
Goodenough, U. W. & Heuser, J. E. Substructure of inner dynein arms, radial spokes, and the central pair/projection complex of cilia and flagella. J. Cell Biol. 100, 2008–2018 (1985).
Barber, C. F., Heuser, T., Carbajal-González, B. I., Botchkarev, V. V. & Nicastro, D. Three-dimensional structure of the radial spokes reveals heterogeneity and interactions with dyneins in Chlamydomonas flagella. Mol. Biol. Cell 23, 111–120 (2012).
O’Donnell, L. Mechanisms of spermiogenesis and spermiation and how they are disturbed. Spermatogenesis 4, e979623 (2014).
Contreras, C. T., Hoyer-Fender, S., Kherraf, Z. E. & Yuan, S. The Transformation of the centrosome into the basal body: similarities and dissimilarities between somatic and male germ cells and their relevance for male fertility. Cells 10, 2266 (2021).
Yogo, K. Molecular basis of the morphogenesis of sperm head and tail in mice. Reprod. Med. Biol. 21, e12466 (2022).
Ma, Y. et al. Structural insight into the intraflagellar transport complex IFT-A and its assembly in the anterograde IFT train. Nat. Commun. 2023 14:1 14, 1–12 (2023).
Rosenbaum, J. L. & Witman, G. B. Intraflagellar transport. Nat. Rev. Mol. Cell Biol. 2002 3:11 3, 813–825 (2002).
Dunleavy, J. E. M., O’Bryan, M. K., Stanton, P. G. & O’Donnell, L. The cytoskeleton in spermatogenesis. Reproduction 157, R53–R72 (2019).
Reiter, J. F. & Leroux, M. R. Genes and molecular pathways underpinning ciliopathies. Nat. Rev. Mol. Cell Biol. 18, 533 (2017).
Martinez, G. et al. Biallelic variants in MAATS1 encoding CFAP91, a calmodulin-associated and spoke-associated complex protein, cause severe astheno-teratozoospermia and male infertility. J. Med Genet 57, 708–716 (2020).
Ruan, J. et al. TreeFam: 2008 update. Nucleic Acids Res. 36, D735 (2008).
Dymek, E. E. & Smith, E. F. A conserved CaM- and radial spoke–associated complex mediates regulation of flagellar dynein activity. J. Cell Biol. 179, 515–526 (2007).
Urbanska, P. et al. The CSC proteins FAP61 and FAP251 build the basal substructures of radial spoke 3 in cilia. Mol. Biol. Cell 26, 1463–1475 (2015).
Bicka, M. et al. Cfap91-dependent stability of the RS2 and RS3 base proteins and adjacent inner dynein arms in tetrahymena cilia. Cells 11, 4048 (2022).
Walton, T. et al. Axonemal structures reveal mechanoregulatory and disease mechanisms. Nature 618, 625 (2023).
Meng, X. et al. Multi-scale structures of the mammalian radial spoke and divergence of axonemal complexes in ependymal cilia. Nat. Commun.15, 1–16 (2024).
Ghanaeian, A. et al. Integrated modeling of the Nexin-dynein regulatory complex reveals its regulatory mechanism. Nat. Commun. 2023 14:1 14, 1–15 (2023).
Ma, M. et al. Structure of the decorated ciliary doublet microtubule. Cell 179, 909–922.e12 (2019).
Leung, M. R. et al. Structural diversity of axonemes across mammalian motile cilia. Nature. https://doi.org/10.1038/s41586-024-08337-5 (2025).
Ernst, C., Eling, N., Martinez-Jimenez, C. P., Marioni, J. C. & Odom, D. T. Staged developmental mapping and X chromosome transcriptional dynamics during mouse spermatogenesis. Nat. Commun. 2019 10:1 10, 1–20 (2019).
Morimoto, Y. et al. Nonsense mutation in CFAP43 causes normal-pressure hydrocephalus with ciliary abnormalities. Neurology 92, e2364 (2019).
McKenzie, C. W. et al. CFAP54 is required for proper ciliary motility and assembly of the central pair apparatus in mice. Mol. Biol. Cell 26, 3140–3149 (2015).
Lee, L. Riding the wave of ependymal cilia: genetic susceptibility to hydrocephalus in primary ciliary dyskinesia. J. Neurosci. Res. 91, 1117–1132 (2013).
Meistrich, M. L. & Hess, R. A. Assessment of spermatogenesis through staging of seminiferous tubules. 299–307. https://doi.org/10.1007/978-1-62703-038-0_27 (2013).
Gadadhar, S., Hirschmugl, T. & Janke, C. The tubulin code in mammalian sperm development and function. Semin. Cell Dev. Biol. 137, 26–37 (2023).
Pleuger, C., Lehti, M. S., Dunleavy, J. E. M., Fietz, D. & O’Bryan, M. K. Haploid male germ cells - The Grand Central Station of protein transport. Hum. Reprod. Update 26, 474–500 (2020).
Kim, D. I. et al. An improved smaller biotin ligase for BioID proximity labeling. Mol. Biol. Cell 27, 1188–1196 (2016).
Cao, W., Gerton, G. L. & Moss, S. B. Proteomic profiling of accessory structures from the mouse sperm flagellum. Mol. Cell. Proteom. 5, 801–810 (2006).
San Agustin, J. T., Pazour, G. J. & Witman, G. B. Intraflagellar transport is essential for mammalian spermiogenesis but is absent in mature sperm. Mol. Biol. Cell 26, 4358 (2015).
Weihbrecht, K. et al. Keeping an eye on Bardet-Biedl syndrome: a comprehensive review of the role of Bardet-Biedl syndrome genes in the eye. Med. Res. Arch. 5, https://doi.org/10.18103/mra.v5i9.1526 (2017).
Davis, R. E. et al. A knockin mouse model of the Bardet-Biedl syndrome 1 M390R mutation has cilia defects, ventriculomegaly, retinopathy, and obesity. Proc. Natl. Acad. Sci. USA 104, 19422–19427 (2007).
Nishimura, D. Y. et al. Bbs2-null mice have neurosensory deficits, a defect in social dominance, and retinopathy associated with mislocalization of rhodopsin. Proc. Natl. Acad. Sci. USA 101, 16588–16593 (2004).
Mykytyn, K. et al. Bardet-Bledl syndrome type 4 (BBS4)-null mice implicate Bbs4 in flagella formation but not global cilia assembly. Proc. Natl. Acad. Sci. USA 101, 8664–8669 (2004).
Zhang, Q. et al. BBS7 is required for BBSome formation and its absence in mice results in bardet-biedl syndrome phenotypes and selective abnormalities in membrane protein trafficking. J. Cell Sci. 126, 2372–2380 (2013).
Bentley-Ford, M. R. et al. A mouse model of BBS identifies developmental and homeostatic effects of BBS5 mutation and identifies novel pituitary abnormalities. Hum. Mol. Genet. 30, 234 (2021).
Wang, H. et al. MYCBPAP is a central apparatus protein required for centrosome–nuclear envelope docking and sperm tail biogenesis in mice. J. Cell Sci. 137, jcs261962 (2024).
Winogradoff, D., John, S. & Aksimentiev, A. Protein unfolding by SDS: the microscopic mechanisms and the properties of the SDS-protein assembly. Nanoscale 12, 5422–5434 (2020).
Spiess, A. N. et al. Cross-platform gene expression signature of human spermatogenic failure reveals inflammatory-like response. Hum. Reprod. 22, 2936–2946 (2007).
Hwang, J. Y. et al. Dual sensing of physiologic pH and calcium by EFCAB9 regulates sperm motility. Cell 177, 1480–1494.e19 (2019).
Qi, H. et al. All four CatSper ion channel proteins are required for male fertility and sperm cell hyperactivated motility. Proc. Natl Acad. Sci. USA 104, 1219–1223 (2007).
Abramson, J. et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 630, 493–500 (2024).
BALDI, E. et al. Intracellular Calcium accumulation and responsiveness to progesterone in capacitating human spermatozoa. J. Androl. 12, 323–330 (1991).
Breitbart, H. Intracellular calcium regulation in sperm capacitation and acrosomal reaction. Mol. Cell Endocrinol. 187, 139–144 (2002).
Dymek, E. E., Heuser, T., Nicastro, D. & Smith, E. F. The CSC is required for complete radial spoke assembly and wild-type ciliary motility. Mol. Biol. Cell 22, 2520–2531 (2011).
Morohoshi, A. et al. Nexin-Dynein regulatory complex component DRC7 but not FBXL13 is required for sperm flagellum formation and male fertility in mice. PLoS Genet. 16, e1008585 (2020).
Huang, T. et al. Absence of murine CFAP61 causes male infertility due to multiple morphological abnormalities of the flagella. Sci. Bull. (Beijing) 65, 854–864 (2020).
Li, W. et al. Biallelic mutations of CFAP251 cause sperm flagellar defects and human male infertility. J. Hum. Genet. 64, 49–54 (2019).
Liu, S. et al. CFAP61 is required for sperm flagellum formation and male fertility in human and mouse. Development 148, dev199805 (2021).
Sha, Y. et al. Deficiency in AK9 causes asthenozoospermia and male infertility by destabilising sperm nucleotide homeostasis. EBioMedicine 96, 104798 (2023).
Xu, Z. et al. AKAP14 is dispensable for mouse fertility. Cells Dev. 182, 204032 (2025).
Zhang, X. et al. LRRC23 is a conserved component of the radial spoke that is necessary for sperm motility and male fertility in mice. J Cell Sci. 134, jcs259381 (2021).
Suzuki, A. et al. Individual disruption of 12 testis-enriched genes via the CRISPR/Cas9 system does not affect the fertility of male mice. J. Reprod. Immunol. 163, 104252 (2024).
Nolan, M. A. et al. Sperm-specific protein kinase A catalytic subunit Cα2 orchestrates cAMP signaling for male fertility. Proc. Natl. Acad. Sci. USA 101, 13483–13488 (2004).
Skålhegg, B. S. et al. Mutation of the Cα subunit of PKA leads to growth retardation and sperm dysfunction. Mol. Endocrinol. 16, 630–639 (2002).
Chen, Y. et al. STYXL1 regulates CCT complex assembly and flagellar tubulin folding in sperm formation. Nat. Commun. 2024 15:1 15, 1–15 (2024).
Fernandez-Gonzalez, A., Kourembanas, S., Wyatt, T. A. & Mitsialis, S. A. Mutation of murine adenylate kinase 7 underlies a primary ciliary dyskinesia phenotype. Am. J. Respir. Cell Mol. Biol. 40, 305–313 (2009).
Oura, S., Ninomiya, A., Sugihara, F., Matzuk, M. M. & Ikawa, M. Proximity-dependent biotin labeling in testicular germ cells identified TESMIN-associated proteins. Sci. Rep. 12, 22198 (2022).
Ikawa, M. et al. Calmegin is required for fertilin alpha/beta heterodimerization and sperm fertility. Dev. Biol. 240, 254–261 (2001).
Erbach, G. T., Lawitts, J. A., Papaioannou, V. E. & Biggers, J. D. Differential growth of the mouse preimplantation embryo in chemically defined media. Biol. Reprod. 50, 1027–1033 (1994).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods. 9, 676–682 (2012).
Toyoda, Y. & Yokoyama, M. The early history of the TYH medium for in vitro fertilization of mouse ova. J. Mamm. Ova Res. 33, 3–10 (2016).
Miyata, H. et al. Sperm calcineurin inhibition prevents mouse fertility with implications for male contraceptive. Science 350, 442–445 (2015). 1979.
Shimada, K., Kato, H., Miyata, H. & Ikawa, M. Glycerol kinase 2 is essential for proper arrangement of crescent-like mitochondria to form the mitochondrial sheath during mouse spermatogenesis. J. Reprod. Dev. 65, 155–162 (2019).
Wang, H. et al. Golgi associated RAB2 interactor protein family contributes to murine male fertility to various extents by assuring correct morphogenesis of sperm heads. PLoS Genet. 20, e1011337 (2024).
Tiscornia, G., Singer, O. & Verma, I. M. Production and purification of lentiviral vectors. Nat. Protoc. 1, 241–245 (2006).
Kaneda, Y. et al. TEX38 localizes ZDHHC19 to the plasma membrane and regulates sperm head morphogenesis in mice. Proc. Natl. Acad. Sci. USA 122, e2417943122 (2025).
Hayashi, K. & Shirao, T. Change in the shape of dendritic spines caused by overexpression of drebrin in cultured cortical neurons. J. Neurosci. 19, 3918–3925 (1999).
Ikawa, M. et al. Calsperin is a testis-specific chaperone required for sperm fertility. J. Biol. Chem. 286, 5639–5646 (2011).
Molday, L. L. & Molday, R. S. 1D4: a versatile epitope tag for the purification and characterization of expressed membrane and soluble proteins. Methods Mol Biol 1177, 1–15 (2014).
Le Sage, V., Cinti, A. & Mouland, A. J. Proximity-dependent biotinylation for identification of interacting proteins. Curr. Protoc. Cell Biol. 73, 17.19.1–17.19.12 (2016).
Meng, E. C. et al. UCSF ChimeraX: tools for structure building and analysis. Protein Sci. 32, e4792 (2023).
Acknowledgements
We thank Keiko Murata for sequence analysis, Hiroko Omori for ultrastructural analysis, Akinori Ninomiya and Hiroko Kato for mass spectrometry analysis (Core Instrumentation Facility, Research Institute for Microbial Diseases, Osaka University). We also thank Natsuki Furuta and Kaito Yamamoto for their technical assistance, and Ferheen Abbasi and Joanna J. Luna for critical reading of this manuscript. We appreciate the experimental discussion with the members of the Department of Experimental Genome Research. This study was supported by the Japan Society for the Promotion of Science (JSPS) KAKENHI grants (JP23K05831 to K.S., JP22H03214, JP23K18328, JP25K02773 to H.M., and JP19H05750, JP21H04753, JP21H05033, JP23K20043 to M.I.); Japan Agency for Medical Research and Development (AMED) grant (JP23jf0126001 to M.I.); a Takeda Science Foundation grant to K.S., H.M., and M.I.; JST FOREST (JPMJFR211F to H.M.); the Eunice Kennedy Shriver National Institute of Child Health and Human Development (P01HD087157 and R01HD088412 to M.I.); and the Bill & Melinda Gates Foundation (Grand Challenges Explorations grant INV-001902 to M.I.). Molecular graphics and analyses performed with UCSF ChimeraX, developed by the Resource for Biocomputing, Visualization, and Informatics at the University of California, San Francisco, with support from National Institutes of Health R01-GM129325 and the Office of Cyber Infrastructure and Computational Biology, National Institute of Allergy and Infectious Diseases.
Author information
Authors and Affiliations
Contributions
Conceptualization, H.W., K.S., H.K., S.O., N.Y., M.I., and H.M.; Formal analysis, H.W., K.S., A.P., M.K., H.K., and S.O.; Investigation, H.W., K.S., A.P., Y.O., M.K., and H.M.; Supervision, N.Y., M.I., and H.M.; Funding acquisition, K.S., M.I., and H.M.; Manuscript preparation, H.W., K.S., M.I., and H.M.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, H., Shimada, K., Pham, A.H. et al. Proximity labeling of axonemal protein CFAP91 identifies EFCAB5 that regulates sperm motility. Nat Commun 16, 8238 (2025). https://doi.org/10.1038/s41467-025-63705-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-63705-7