Abstract
Gut mycobiome dysbiosis has been implicated in inflammatory bowel disease (IBD). However, it remains unknown whether specific fungal species identified by sequencing directly contribute to IBD pathogenesis. Here, based on analysis of three fecal metagenome datasets of IBD cohorts and a previously established cultivated gut fungi catalog, we identify an IBD-depleted intestinal fungus Clavispora lusitaniae strain P4013B. We show P4013B attenuates DSS-induced colitis in wild-type, antibiotics-treated, and germ-free mice through activation of aryl hydrocarbon receptor (AHR). Using an activity-guided isolation strategy, we identify the P4013B metabolite indole-3-ethanol (IEt) as the AHR agonist mediating the anti-colitis activity. We further validate the role of IEt via engineering strains that overexpress pyruvate decarboxylases producing high yields of IEt. Tea polysaccharide enhanced the anti-colitis activity of P4013B by promoting its proliferation and colonization in the colon. Together, these results suggest that C. lusitaniae P4013B may be explored as a potential probiotic for the treatment and prevention of IBD.
Similar content being viewed by others
Introduction
Gut mycobiome, as an integral part of gut microbiome, plays a vital role in intestinal ecology and human health. The dysbiosis of gut fungal community has been implicated in multiple disease states, such as digestive system neoplasms1, autoimmune diseases2,3, cardiovascular diseases4 and so on. Notably, the association between gut fungal community and specific diseases has always been established based on the deep-sequencing technologies, yet it is currently unknown whether the fungal strains with differential abundance in disease cohorts have the essential roles in disease progression. To elucidate the precise role of gut mycobiome in disease pathogenesis, we have established a catalog of cultivated gut fungi (CGFs) that comprises 760 fungal genomes and over 11,000 gut fungal strains obtained from the feces of healthy volunteers and identified significant common disease-related variations in gut mycobiome composition5. Especially, dysbiosis in the gut mycobiome was profoundly associated with inflammatory bowel disease (IBD) and we found specific fungi attenuated experimental colitis in mice via secretion of active metabolites2,5. In present study, we aimed to further characterize the gut mycobiome signatures in IBD patients and discover the specific fungi with therapeutic potential for the treatment of IBD.
IBD is the general term for Crohn’s disease (CD) and ulcerative colitis (UC), and characterized by chronic relapsing digestive disorders and inflammation, which affect millions of individuals globally6,7. Especially in newly industrialized countries like China and India with the epidemiological stage of IBD in the “Acceleration in Incidence”, the prevalence will increase tremendously in the coming decades7,8. Despite the advances made in the medical management of IBD, including monoclonal antibodies and novel oral small molecules such as IL23p19 antagonists, selective Janus kinase inhibitors and sphingosine-1-phosphate receptor modulators6,9, the effective therapy for IBD also faces the great challenges with respect to adverse reactions, cost, feasibility and recurrence. Instead, growing evidences suggest that microbiota therapeutics, mainly refer to fecal microbiota transplantation (FMT), represents a promising innovative therapeutic strategy for the treatment of IBD10,11. The goal of the microbiota therapeutics is to correct the dysbiosis of gut microbiota through whole gut microbiome replacement10,11. As a small but crucial component of the gut microbiota, the abundance of specific fungal species including but not limited to Saccharomyces boulardii12, Saccharomyces cerevisiae13, and Candida metapsilosis2 were significantly altered in IBD cohorts and the transplantation of these fungal strains exhibited improvement effects against colitis via maintaining gut microbiota homeostasis, regulating host intestinal immunity, or secreting active metabolites14. Instead, Malassezia restricta and Candida albicans are enriched in IBD patients and exacerbated colitis through eliciting innate inflammatory responses in mice15,16,17. Therefore, it is very necessary to identify the pathogens and probiotics from the gut mycobiota in individual species instead of mycobiome level. More fungal species with health benefits need to be discovered in multi-kingdoms fungal community to develop some novel approaches for preventing and/or managing IBD.
In this work, a Clavispora lusitaniae strain (P4013B), is identified through composition analysis of the gut mycobiota in IBD cohorts, following cultivation from human feces. The protective effects of C. lusitaniae P4013B are comprehensively evaluated in experimental colitis models in mice. Its therapeutic mechanisms and active metabolites are thoroughly investigated through integration of transcriptomic analysis, knockout mice, and a bioassay-guided isolation strategy. Additionally, a higher yield of bioactive metabolite from P4013B is achieved through genetic engineering and the incorporation of herbal polysaccharides, enhancing its therapeutic potential for IBD. The discovery of P4013B lays the groundwork for its future clinical application in the prevention and treatment of IBD.
Results
Clavispora lusitaniae is depleted in IBD cohorts and attenuates colitis in mice
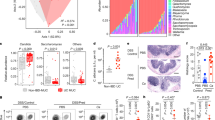
To explore the fungal characteristics in patients with IBD, we profiled the gut mycobiomes using three publicly available fecal metagenome datasets (i.e., HeQ_201718, WengY_201919, and YanQ_20245) comprising a total of 169 IBD patients and 228 healthy controls of Chinese origin. At the phylum and genus levels, our analysis revealed that the gut fungal compositions of IBD patients and healthy controls were relatively similar, with permutational multivariate analysis of variance (PERMANOVA) p-values > 0.05 for both taxonomic levels after adjusting for sex, age, body mass index, and study effect (Fig. 1a, b). However, at the species level, multivariate analysis demonstrated a significant shift in the overall mycobiome of IBD patients compared to healthy controls (Fig. 1c), with an adjusted PERMANOVA p-value of 0.001. Using random-effect meta-analysis across the three datasets, we identified 13 species with significantly differed relative abundances between patients and controls (meta-analysis q < 0.05; Fig. 1d). Among these, three species including Trametes sp. AH28-2, Pichia kudriavzevii, and Clavispora lusitaniae were more abundant in healthy controls, while 10 species such as Microascales sp. c195, Kazachstania servazzii, Candida albicans A, and Trichophyton indotineae, were enriched in IBD patients. Notably, the enrichment of K. servazzii, C. albicans, and the depletion of P. kudriavzevii in IBD patients have also been observed in previous global surveys of IBD populations5. Moreover, within-dataset case-control comparisons via Wilcoxon rank-sum tests confirmed that Trametes sp. AH28-2, P. kudriavzevii, and C. lusitaniae were consistently enriched or showed a trend of enrichment in healthy controls across each dataset (Fig. 1d), suggesting that these species may be potential markers of IBD-depleted gut fungi.
a, b Phylogenetic composition of the gut mycobiome of IBD patients and healthy controls at the phylum (a) and genus (b) levels. The bars indicate the relative abundance of each phylum/species and are colored according to their taxonomic assignments. Only species with an average relative abundance of > 2% are presented. c Principal coordinate analysis (PCoA) of the gut mycobiome in three Chinese fecal metagenomic datasets. The first two principal coordinates (PCs) and their explained variances are presented. Upper and left boxplots show the disease scores in the first two PCs. Statistics: two-sided Wilcoxon rank-sum test. Box plots show median (line), IQR (box), and 1.5 × IQR (whiskers). Outliers are shown as individual points. d Meta-analysis (left panel) and Wilcoxon rank-sum test analysis (right panel) of the gut fungal signatures of IBD. Forest plot displays the Hedges’g standardized mean differences of gut fungi between IBD patients and healthy controls. Solid red lines indicate the 95% confidence intervals. Heatmap showing the enrichment of each species in each case-control comparison. Wilcoxon rank-sum test: *, q < 0.05; **, q < 0.01. n = 169 IBD patients and n = 178 healthy controls in (a–d). e Schematic diagram of the experimental study design for the transplantation and functional evaluation of the clinically isolated Clavispora lusitaniae P4013B strain. f Disease Activity Index (DAI) of mice after 3% DSS induction. g, h Gross morphology and length of colons. i Colonoscopy and H&E-staining of colons. Scale bars, 50 μm. j Expression levels of inflammation-associated proteins in colons. k Plasma concentration of FITC-dextran after oral administration for 4 h. l Immunofluorescence analysis of colonic ZO-1. Scale bars, 50 μm. m Expression levels of tight junctions (TJs) and adherens junctions (AJs) proteins in colons. Statistics: one-way ANOVA with Tukey’s multiple comparison test (f, h, k). The data represent the means ± SEM, n = 6 biological replicates in (f, h, k). Source data are provided as a Source Data file.
To investigate the potential effects of healthy control-enriched fungi on IBD pathology, three species (Trametes sp. AH28 − 2, P. kudriavzevii, and C. lusitaniae) were orally administrated and colonized into Specific Pathogen-Free (SPF) mice with dextran sulfate sodium (DSS)-induced colitis. Firstly, three strains from each species (PG5501, F2008 and P4013B) were selected from the cultivated gut fungi (CGF) catalog we established previously5, based on their frequency and abundance during the fungal cultivation and isolation from the feces of healthy volunteers. Then, the mice received oral transplantation of the strains prior to DSS-induced colitis (Fig. 1e). Interestingly, no improvement was observed after colonization of Trametes sp. AH28 − 2 PG5501, or P. kudriavzevii F2008 (Supplementary Fig. 1a, b), although we found that only colonization of C. lusitaniae P4013B attenuated colitis-related clinical symptoms scored as disease activity index (DAI), including body weight loss, diarrhea, and rectal bleeding (Fig. 1f). Furthermore, P4013B colonization significantly alleviated DSS-induced colon shortening and pathological injures, including diffuse hemorrhages, erosions, and ulcers under colonoscopy, and extensively destroyed structural integrity of villi, crypt damage, and inflammatory infiltration by H&E-staining (Fig. 1g–i), indicating improvement of colonic inflammation and gut barrier function by P4013B colonization. Indeed, p65 phosphorylation and protein levels of TNF-α and IL6 were remarkably decreased by P4013B colonization in the colon of DSS-induced mice (Fig. 1j). Meanwhile, the mice with P4013B colonization exhibited the lower plasma concentration of FITC-dextran after oral administration, and higher protein expression of tight junctions (TJs) and adherens junctions (AJs), including ZO-1, occludin, E-cadherin and β-catenin (Fig. 1k–m). Moreover, pasteurized P4013B exerted a weaker protection against DSS-induced colitis than live P4013B (Fig. 1f–m), indicating that the colonization of live P4013B in the gastrointestinal tract facilitated its protective effect. The successful colonization of P4013B in mouse gut was also verified by fecal metagenomic sequencing (Supplementary Fig. 2a). In addition, both live and pasteurized P4013B showed no impact on the body weight of mice during colonization (Supplementary Fig. 2b), which fully suggested that P4013B is a non-pathogenic strain. These results indicated that P4013B colonization attenuated DSS-induced colitis in mice through improvement of gut barrier function and inhibition of colonic inflammation.
P4013B-derived metabolites attenuate colitis independent of inherent gut microbiota
The primary modes by which the gut microbiota interacts with the host include direct interactions with the intestinal mucosa via immunity or neuromodulation, interference with other gut microbiota, and the secretion of bioactive metabolites2,20. To investigate the interaction mechanism between C. lusitaniae P4013B and the host, the colonization effect of P4013B on colitis was also evaluated in germ-free (GF) mice and antibiotic-treated (ABX) mice, respectively, to explore its potential interactions with the native gut microbiota. GF mice were orally transplanted with P4013B for two weeks prior to DSS administration, and the severity of colitis was assessed (Fig. 2a). Fecal samples from the GF mice with P4013B colonization were cultured on agar plates, revealing numerous white colonies (Fig. 2b). Additionally, fungal colonization of P4013B in colon tissue was confirmed by fluorescence in situ hybridization (FISH) analysis (Fig. 2c), demonstrating successful monocolonization.
a After colonization without or with P4013B for two weeks, germ-free mice were administered 3% DSS in drinking water for 7 days to induce acute colitis. b P4013B colonies after incubation of stool samples from mice with or without P4013B colonization on agar plates. c P4013B colonization in colon visualized by fluorescence in situ hybridization (FISH). Scale bars, 50 μm. d The body weight change, diarrhea and rectal bleeding scores after 3% DSS induction. e Gross morphology and length of colons. f H&E-staining of colons. Scale bars, 50 μm. g Immunofluorescence analysis of colonic ZO-1. Scale bars, 50 μm. h Expression levels of AJ proteins (E-cadherin and β-catenin) and TJ protein occludin in colons. i Expression levels of inflammation-associated proteins (p65, TNF-α, and IL6) in colons. Statistics: two-tailed Student’s t test (d, e). The data represent the means ± SEM, n = 5 biological replicates in (d, e). j, k Colonic crypts were isolated and cultured for 6 days in the absence or presence of LPS (5 μg/mL) or P4013B fermentation broth (5%). j Representative images of organoid formation. Scale bars, 50 μm. k H&E-staining (Scale bars, 50 μm) and Immunofluorescence analysis of ZO-1 in colonic organoid (Scale bars, 10 μm). l, m Normal human colon epithelial cells NCM460 were treated by LPS (1 μg/mL) in the absence or presence of P4013B fermentation broth (5%). l Immunofluorescence analysis of ZO-1 and Occludin in NCM460 cells. Scale bars, 50 μm. m Expression levels of AJ proteins (E-cadherin and β-catenin) and inflammation-associated proteins (p65, TNF-α, and IL6) in NCM460 cells. Source data are provided as a Source Data file.
In GF mice, P4013B monocolonization significantly alleviated colitis, as evidenced by reduced body weight loss, diarrhea, and rectal bleeding, as well as elongated colon length, less severe histopathological damage, improved gut barrier function, and lower levels of colonic inflammation (Fig. 2d–i). Similarly, ABX-treated mice, which had depleted gut microbiota, exhibited the reduced colitis severity after P4013B colonization, with less colitis-related symptoms, lower gut permeability, and decreased inflammation (Supplementary Fig. 3). These findings fully suggested that P4013B attenuates colitis independent of the native gut microbiota.
To further assess whether P4013B metabolites directly contribute to the protective effects against colitis, colonic organoids and the human colon epithelial cell line NCM460 were used. Crypts isolated from SPF mice were cultured and exposed to LPS to induce inflammation, with or without P4013B fermentation broth (5%) for 6 days. Notably, LPS-induced inhibition of organoid germination and formation was largely reversed by the P4013B fermentation broth (Fig. 2j). H&E staining and immunofluorescence analysis revealed that treatment with P4013B fermentation broth ameliorated the disorganization and thinning of the intestinal epithelium and upregulated the expression of tight junction protein ZO-1 (Fig. 2k), indicating the significant improvement in gut barrier function.
Similarly, NCM460 cells treated with P4013B fermentation broth exhibited higher cell viability and formed confluent monolayers with increased expression of ZO-1 and occludin after LPS stimulation (Fig. 2l). Furthermore, P4013B fermentation broth also inhibited the expression of adherens junction proteins (E-cadherin and β-catenin) and inflammation-associated proteins (p65, TNF-α, and IL-6) in LPS-stimulated NCM460 cells (Fig. 2m). These results suggest that P4013B exerts the direct protective effects against colonic inflammation in vitro, independent of immune or nervous system regulation.
P4013B protects against colitis dependent on AHR activation
RNA-seq analysis was conducted to further elucidate the underlying mechanism about the protective effect of P4013B against colitis. The colons from DSS-induced SPF mice with or without P4013B colonization were used for RNA extraction and transcriptome sequencing. As expected, P4013B colonization led to a distinctive expression profile (Supplementary Fig. 4a). A total of 495 differentially expressed genes (DEGs) was identified using the cut-off of |log2 fold change | >2 and padj<0.05, of which 277 DEGs were down-regulated and 218 DEGs were up-regulated (Fig. 3a). Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis revealed that DEGs were significantly enriched at the various pathways related to wound healing21, regulation of VEGFs22,23, extracellular matrix assembly and organization24, positive regulation of GPCRs25, protein activation cascade, and regulation of body fluid levels (Supplementary Fig. 4b), which was the common response to mucosal injury and repair during colitis. Among DEGs, the top 5 DEGs with highest |padj| values were Cyp1b1, Prl2a1, Scara5, Ighv10-3, and Kng2 (Fig. 3a and Supplementary Fig. 4b), which were involved in xenobiotic metabolism26, immunoreactivity27, intestinal fibrosis28,29, intestinal adaptive immune response30, and intestinal inflammation31, respectively. The mRNA expressions of five DGEs in the colons were profoundly induced upon P4013B colonization by qPCR assay, particularly Cyp1b1 with a 7.6-fold increase (Fig. 3b), suggesting a transcriptional regulation. CYP1B1 is a member of the cytochrome P450 superfamily involved in the metabolism of numerous physiological substrates including polycyclic aromatic hydrocarbons26, which can be regulated by several nuclear receptor superfamily members (NRs) for real-time response to xenobiotics. The molecular pathways involved in the regulation of Cyp1b1 mediated by NRs, including AHR, AR, ER, and Sp132, were then extracted from the transcriptome sequencing data (Supplementary Fig. 4c). A Welch’s two-sample t test of PC1 scores revealed that AHR activation showed a trend toward significance (t = 2.68, p = 0.076), whereas no meaningful effects were observed for other pathways: AR (t = 0.48, p = 0.68), ER (t = − 0.93, p = 0.43), and SP1 (t = 0.37, p = 0.74) (Supplementary Fig. 4c). Furthermore, qPCR analysis of canonical downstream targets for AR, ER, and SP1 revealed no significant differences between DSS group and P4013B treatment group (Supplementary Fig. 4d). However, qPCR assay showed that the typical downstream target genes of AHR pathways, mainly involved in various metabolic enzymes (Cyp1a1, Cyp1a2, Cyp1b1, Aldh3a) and pro-/anti-inflammatory cytokines (Il10, Il22, Il17a, Tnf-α, Il1b), were transcriptionally regulated upon P4013B colonization (Fig. 3c). Protein-protein interaction (PPI) network was constructed using STRING11.5 and revealed that AHR was the hub target with degrees of 10 among these DEGs (Fig. 3d). Gene Set Enrichment Analysis (GSEA) also indicated that IL17 signaling pathway and PPAR signaling pathway were significantly enriched in P4013B colonization group (Supplementary Fig. 4e), both of which were reported to be regulated by or have the crosstalk with AHR pathway33,34. Furthermore, the protein expression of colonic CYP1A1 was upregulated in SPF mice with P4013B colonization (Supplementary Fig. 5a, b). Similar results were also found in germ-free mice, in which P4013B monocolonization induced the mRNA expressions of AHR target genes (Cyp1a1, Cyp1a2, Cyp1b1) as well as the protein expression of colonic CYP1A1 (Supplementary Fig. 5c, e). However, AHR expression was almost unchanged upon P4013B colonization (Supplementary Fig. 5a and d). These results indicated that the AHR signaling pathway was activated in the colon of DSS-induced mice with P4013B colonization.
a Volcano plot of all detected genes in the colons of mice with DSS-induced colitis with or without P4013B colonization. b qPCR analysis of the top five significantly upregulated genes with significant statistical differences. c qPCR analysis of AHR target genes. Colored bars indicate standardized log2 expression intensities. Red and blue represent upregulation and downregulation, respectively. d Protein-protein interaction (PPI) network of DEGs and AHR. Node size is proportional to node degree. Node color reflects differential expression levels. e Agonistic effect of P4013B fermentation broth (5%) and AHR agonist FICZ determined by luciferase reporter assays using HEK293T cells transfected with AHR and CYP1A1 promoter plasmids. f–i NCM460 cells were incubated with P4013B fermentation broth (5%) in the absence or presence of AHR antagonists GNF351 (1 μM). f Immunofluorescence analysis of nuclear translocation of AHR. Scale bars, 10 μm. g AHR levels in the cytoplasm and nucleus. h qPCR analysis of AHR target genes. i Protein expression levels of AHR target genes. Statistics: two-tailed Student’s t test (b); one-way ANOVA with Dunnett’s test(e); one-way ANOVA with Tukey’s test (h). The data represent the means ± SEM, n = 3 biological replicates in (b, e, h). j Ahr-/- mice were administered 3% DSS in drinking water for 7 days to induce acute colitis after colonization without or with P4013B for two weeks. k Body weight change and DAI after 3% DSS induction. l Gross morphology and length of colons. m Colonoscopy and H&E-staining of colons. Scale bars, 50 μm. n Immunofluorescence analysis of colonic ZO-1. o Western blot analysis of TNF-α and IL6 in colons. p Representative images of organoid formation. Colonic crypts from Ahr-/- mice were isolated and cultured for 6 days in the absence or presence of LPS (5 μg/mL) or P4013B fermentation broth (5%). Scale bars, 50 μm. Statistics: two-tailed Student’s t test (k, l). The data represent the means ± SEM, n = 6 biological replicates in (k, l). Source data are provided as a Source Data file.
To directly validate whether P4013B metabolites could act as AHR agonizts, AHR activation in vitro was evaluated using luciferase reporter assays and NCM460 cells after incubation with P4013B fermentation broth. We found that P4013B fermentation broth showed agonistic action with 6-Formylindolo[3,2-b]carbazole (FICZ) as a positive control (Fig. 3e). Incubation with P4013B fermentation broth facilitated the nuclear translocation of AHR in NCM460 cells, which was blocked by GNF351, an AHR antagonist (Fig. 3f, g). AHR activation by P4013B fermentation broth subsequently induced a wholesale upregulation of AHR target genes in both mRNA and protein levels, which were substantially suppressed by GNF351 (Fig. 3h, i). These findings clearly suggested that the metabolites in P4013B fermentation broth could significantly activate AHR pathway.
To further clarify the relationship between AHR activation and resistance to colitis, Ahr-/- mice were used to evaluate the potential effect of P4013B colonization on DSS-induced colitis (Fig. 3j). Although metagenomic sequencing verified successful colonization of P4013B (Supplementary Fig. 5f), we found that P4013B failed to attenuated DSS-induced colitis in Ahr-/- mice, including gradually deteriorating DAI, colon shortening, mucosal injury, gut barrier loss, and inflammatory infiltration (Fig. 3k–o). The expression of AHR target genes was not changed regardless of P4013B colonization in Ahr-/- strain (Supplementary Fig. 5g–i). Additionally, the formation of colonic organoids from the crypts of Ahr-/- mice was not responsive to P4013B fermentation broth (Fig. 3p). These data revealed that AHR was essential for the improvement of P4013B on colitis. Taken together, P4013B attenuated DSS-induced colitis in vitro and in vivo through AHR activation by P4013B-dervied metabolites.
P4013B-derived Indole-3-ethanol (IEt) attenuates colitis through AHR activation
To finally elucidate that the active metabolite derived by P4013B was responsible for the protective effect against colitis, an activity-guided isolation strategy was established by integrating luciferase reporter assays and chromatographic method. The fermentation broth of P4013B was separated and fractionated by semi-preparative high-performance liquid chromatography (pre-HPLC) equipped with ODS column, and the fractions were screened against AHR activation by luciferase reporter assays (Fig. 4a). Totally eight fractions were obtained and fractions 6-8 showed the strong agonistic action on AHR at 50 μg/mL (Fig. 4b). They were subsequently extracted and isolated from the fermentation broth to obtain four metabolites which were identified unambiguously as indole-3-lactic acid (ILA), indole-3-carboxaldehyde (ICA), indole-3-ethanol (IEt, also known as tryptophol) and Indole (Ind) based on a wide array of spectroscopic data, including 1H-, 13C-NMR, and HR-MS (Fig. 4b, Supplementary Figs. 6–25 and Supplementary Table 1–4). Further confirmation was achieved by comparison with authentic standards of ILA, ICA, IEt, and Ind (Supplementary Figs. 6–25 and Supplementary Table 1–4). LC-MS/MS analysis showed that IEt was the major metabolite with a yield of 25 μM in the fermentation broth of P4013B (Fig. 4c).
a Schematic diagram of the strategy for AHR agonistic activity-guided isolation. The fermentation broth of P4013B was separated and fractionated by semi-preparative high-performance liquid chromatography (pre-HPLC), the fractions of which were screened against AHR activation by luciferase reporter assays. b AHR agonizts isolated from the fermentation broth of P4013B. The red line was a representative chromatogram of the fermentation broth of P4013B by pre-HPLC. The black line was the levels of AHR activation upon fractions treatment by luciferase reporter assays using HEK293T cells transfected with AHR and CYP1A1 promoter plasmids. The fold-change represented the ratio of the metabolite-activated luminescence to control luminescence. The chemical structures and retention times of metabolites with potent agonistic activities against AHR were indicated in the chromatogram. c The concentrations of P4013B metabolites after fermentation for 24 h determined by LC-MS/MS. d Agonistic effect of P4013B metabolites determined by luciferase reporter assays. e Dose-response curves of AHR activation upon IEt treatment by luciferase reporter assays using HEK293T cells transfected with AHR and CYP1A1 promoter plasmids. f In silico molecular docking assays for the interaction between IEt and AHR. g The agonistic effects of IEt on WT and mutant AHR in HEK293T cells by luciferase reporter assays. Statistics: one-way ANOVA with Dunnett’s test (d, e). The data represent the means ± SEM, n = 3 biological replicates in (b, c, d, e, g). h–k NCM460 cells were incubated with IEt (10 μM) in the absence or presence of AHR antagonists GNF351 (1 μM). h Immunofluorescence analysis of nuclear translocation of AHR. Scale bars, 10 μm. i AHR levels in the cytoplasm and nucleus. j qPCR analysis of AHR target genes. k Western blot analysis of AHR target genes. lm. The fecal concentrations of IEt in human (30 healthy subjects and 29 IBD patients, l) and DSS-induced germ-free mice (n = 6 biological replicates, m) with or without P4013B colonization. Statistics: one-way ANOVA with Tukey’s test (j); two-tailed Student’s t test (l, m). The data represent the means ± SEM, n = 3 biological replicates in (j). Source data are provided as a Source Data file.
To directly assess the agonistic activity of P4013B metabolites against AHR, luciferase reporter assays were conducted in HEK293T cells transfected with AHR and CYP1A1 promoter plasmids. Among these metabolites, IEt showed the strongest agonistic effect with a 2.6-fold increase (Fig. 4d). And IEt activated AHR in a concentration-dependent manner with EC50 of 10.9 ± 1.2 μM (Fig. 4e). In silico molecular docking assay indicated that IEt could entry into the ligand binding pocket in the PAS-B domain of human AHR and interact with the residue Cys333 and Ser336 through H-bond interaction (Fig. 4f). The hydrogen bond formation directly contributes to stabilizing protein-ligand complexes and both Cys333 and Ser336 residues were identified as key residues influencing ligand binding via H-bond interactions35,36, which were selected as potential targets for further validation through site-directed mutagenesis. Plasmids encoding AHR with single mutation (C333A or S336A) and dual mutation (C333A/S336A) were constructed and transfected into HEK293 cells with CYP1A1 promoter for further luciferase reporter assay. Compared to wild-type (WT) AHR, mutated AHR (C333A or S336A) led to a weaker induction of CYP1A1 promoter activity upon IEt stimulation, while AHR with dual mutation completely abolished IEt-induced CYP1A1 promoter activity (Fig. 4g). These results indicated that IEt acted as an AHR agonist. Furthermore, NCM460 cells were used for the evaluation of AHR activation in vitro. We found that incubation with IEt induced the nuclear translocation of AHR in NCM460 cells, which was blocked by GNF351 (Fig. 4h, i). Meanwhile, both mRNA and protein expressions of AHR target genes, CYP1A1, CYP1A2, and CYP1B1, were induced in the presence of IEt, which were substantially suppressed by GNF351 (Fig. 4j and k), suggesting AHR activation by IEt in NCM460 cells. These findings confirmed that the novel metabolite IEt derived from P4013B was an AHR agonist.
Notably, LC-MS/MS analysis found that IEt was detectable in the stool samples from both healthy subjects and IBD patients and slightly decreased in IBD patients (Fig. 4l). In the mouse model of colitis fed with standard diet, regardless of SPF mice, germ-free mice, and Ahr-/- mice, on the contrary, P4013B colonization significantly increased the fecal concentration of IEt (Fig. 4m and Supplementary Fig. 26). Especially, the fecal level of IEt was increased 2.5-fold after P4013B transplantation in germ-free mice (Fig. 4m), suggesting that the protective effect of P4013B against colitis might be attributed to its metabolite IEt via AHR activation.
To directly address the biological activity of IEt in vivo, we investigated the susceptibility to DSS-induced colitis after orally administration of IEt in WT mice or Ahr-/- mice. IEt treatment significantly attenuated DSS-induced colitis symptoms characterized by body weight loss, diarrhea, rectal bleeding, colon shortening, mucosal injury and pathological changes, increased levels of inflammatory factors, and decreased expression of tight junction proteins, indicating improvement of colonic inflammation and gut barrier function by IEt (Fig. 5a–g). Meanwhile, mRNA expressions of AHR target genes, including Cyp1a1, Cyp1a2, and Cyp1b1, as well as the protein level of CYP1A1 in the colon, were markedly increased by IEt treatment (Fig. 5h–i), indicating activation of AHR pathway by IEt. Furthermore, colonic crypts from WT mice were isolated to evaluate the potential effect of IEt on organoid formation in the presence of LPS. Incubation of IEt enhanced the organoid-forming efficiency, ameliorated the disorganized and thinning of the intestinal epithelium, and upregulated the expression of ZO-1 and CYP1A1 (Fig. 5j, k), all of which indicating a coordinated repairment of gut barrier function and AHR activation in vitro. These results suggested that IEt attenuated colitis and activated AHR pathway in mice and in colonic organoid. On the contrary, IEt treatment fail to alleviate the colitis symptoms induced by DSS in Ahr-/- mice (Supplementary Fig. 27a–d). Elevated inflammatory level and intestinal permeability, as well as AHR target genes in the colon, were not changed by IEt treatment (Supplementary Fig. 27e–h). Similarly, organoid-forming dysfunction induced by LPS could not be rescued by IEt in the colonic crypts of Ahr-/- mice (Supplementary Fig. 27i). Additionally, incubation of IEt induced a coordinated upregulation of TJ and AJ proteins and downregulation of pro-inflammatory cytokines in LPS-induced NCM460 cells, which was abolished by GNF351 (Supplementary Fig. 27j and k). These results confirmed that IEt attenuated colitis through AHR activation.
a IEt (200 mg/kg) was administered via oral gavage every 12 h for 7 days, and were administered 3% DSS in drinking water at 7th day to induce acute colitis. b The body weight change, diarrhea and rectal bleeding scores after 3% DSS induction. c Gross morphology and length of colons. d Colonoscopy and H&E-staining of colons. Scale bars, 50 μm. e Plasma concentration of FITC-dextran after oral administration for 4 h. f Immunofluorescence analysis of colonic ZO-1 and Western blot analysis of TJs and AJs proteins. Scale bars, 50 μm. g Inflammation-associated proteins in colons. h qPCR analysis of AHR target genes in the colons. i Immunofluorescence and Western blot analysis of colonic CYP1A1. Statistics: two-tailed Student’s t test (b, c, e, h). The data represent the means ± SEM, n = 6 biological replicates in (b, c, e, h). j, k Colonic crypts were isolated and cultured for 6 days in the presence of LPS (5 μg/mL) with or without IEt (10 μM). j Representative images of organoid formation. Scale bars, 50 μm. k H&E-staining and Immunofluorescence analysis of ZO-1 and CYP1A1 in colonic organoid. Scale bars, 10 μm. n = 10 biological replicates (j). Source data are provided as a Source Data file.
Pyruvate decarboxylase-derived IEt suppresses DSS-induced colitis in mice through AHR activation
To exclude potential effects of fungal strain background and further clarify the biosynthesis of IEt, we engineered the additional Escherichia coli BL21 strains to produce IEt for assessing the in vivo impacts induced by microbial IEt. For this purpose, enzymes involved in IEt formation from tryptophan were identified and transformed into an engineered strain E. coli BL21, which were then screened for IEt production (Fig. 6a). IEt is commonly biosynthesized by the well-known Ehrlich pathway37 in Saccharomyces cerevisiae through three enzymatic steps: transamination, decarboxylation and reduction (Fig. 6a). According to this metabolic pathway, five enzymes were identified as key enzymes catalyzing IEt formation from tryptophan by P4013B through whole-genome sequencing and functional gene annotation using KEGG database (Fig. 6b and c; Supplementary Table 5). Specifically, tryptophan is firstly transaminated by aromatic aminotransferases aro8_1158 to form indole-3-pyruvic acid (IPYA); IPYA is then catabolized by two pyruvate decarboxylases pdc_5239 and/or pdc_2062 to obtain indole-3-acetaldehyde (IAD); IAD is finally catabolized by two alcohol dehydrogenases ndh1_1589 and frmA_5117 to produce IEt (Fig. 6c). Using P4013B genome as template, we obtained five PCR products with molecular weights of 1464-1733 kb (Fig. 6d), which matched with the sequencing results. Subsequently, the PCR products were ligated at NdeI (CATATG)-XhoI (CTCGAG) sites in plasmid pET30a(+) to reconstitute the pathway for IEt generation in E. coli BL21. Five strains (E. coli_1158, E. coli_1589, E. coli_2062, E. coli_5117, E. coli_5239) and control strain (E. coli_pET30a(+)) were successfully established with overexpression of the specific protein of interest using anti-His antibody by Western blot analysis (Fig. 6e). As expected, the chemical transformation from tryptophan to IEt by the engineered functional E. coli strain was significantly accelerated and further observed the elevated production of IEt using LC-MS/MS analysis (Fig. 6f). Among them, E. coli_2062 and E. coli_5239 gave the highest yield of IEt with a 71.3-fold and 71.1-fold increase, respectively (Fig. 6f), which were subsequently used for functional evaluation in mice. After oral transplantation of E. coli_2062 or E. coli_5239 and colitis induction by DSS, mice developed an improved colitis phenotype with a reduction of DAI, colon shortening, mucosal damage, and pathological changes, and an upregulation of ZO-1, indicating improvement of colonic inflammation and gut barrier function by E. coli_2062 and E. coli_5239 (Fig. 6g–k). Meanwhile, mRNA expressions of AHR target genes, including Cyp1a1, Cyp1a2, and Cyp1b1, as well as the protein level of CYP1A1 in the colon, were markedly increased by the colonization of E. coli_2062 and E. coli_5239 (Fig. 6l and m), suggesting that AHR pathway was activated in the colon with the colonization of the functional engineered strains. Furthermore, the fecal concentrations of IEt were increased by 100% and 137% in the mice colonized with the high-producing strains (E. coli_2062 and E. coli_5239, respectively) as compared to those with E. coli_ pET30a(+) colonization, respectively (Fig. 6n). These findings fully demonstrated that functional engineered strains with high-yield IEt by pyruvate decarboxylases 2062 and 5239 cloned from P4013B could attenuate DSS-induced colitis via AHR activation. In other words, the protective effect of P4013B really depends on the presence of a IEt-generating pathway.
a Schematic diagram of the cloning strategy for the construction and screening of strains with overexpression of IEt-producing enzymes and high production of IEt. b P4013B genome map with annotation of genes involved in the Ehrlich pathway responsible for the biosynthesis of IEt. c Schematic diagram of IEt biosynthetic pathways in P4013B. d PCR analysis of genes encoding IEt-producing enzymes in P4013B. e Western blot analysis of recombinant expression of IEt-producing enzymes in engineered E. Coli BL21 strains. IEt-producing genes with His tag were cloned separately into pET30a(+) plasmids, which were then transduced into E. Coli BL21. f production of IEt in the fermentation broth of engineered E. Coli BL21 strains determined by LC-MS/MS. n = 3 biological replicates (f). g Mice were colonized with E. coli_2062 or E. coli_5239 for two weeks and were administered 3% DSS in drinking water at 15th day to induce acute colitis. h DAI after 3% DSS induction. i Gross morphology and length of colons. j Colonoscopy and H&E-staining of colons. Scale bars, 50 μm. k Plasma concentration of FITC-dextran after oral administration for 4 h. l Immunofluorescence analysis of colonic ZO-1 and CYP1A1. Scale bars, 50 μm. m qPCR analysis of AHR target genes in the colons. n IEt concentrations in the feces of mice determined by LC-MS/MS. Statistics: one-way ANOVA with Dunnett’s test (h, i, k, m, n). The data represent the means ± SEM, n = 6 biological replicates in (h, i, k, m, n). Source data are provided as a Source Data file.
Tea Polysaccharide enhances the anti-colitis activity of P4013B as a prebiotic
In order to facilitate the better therapeutic effect of 4013B in treating IBD, we screened for potential candidates that can promote the growth of P4013B from a total 72 natural herbal polysaccharides. P4013B was cultured with herbal polysaccharide (5 mg/mL) over 24 h and their growth curve were monitored to assess the proliferation (Fig. 7a). Results showed multiple polysaccharides caused varying degrees of promotion on P4013B proliferation and Tea Polysaccharides (TPS, F7) exhibited the strongest capacity to facilitate the growth of P4013B (Fig. 7b). The growth curves of P4013B showed an extended logarithmic growth phase after incubation with TPS (Fig. 7c). The promotion on P4013B proliferation by TPS exhibited concentration dependent and could be observed as low as 1 mg/mL (Fig. 7d). TEM assay showed number of P4013B cells with the production of clavate ascospores were largely increased by incubation with TPS (Fig. 7e). Flow analysis indicated that TPS decreased the proportion of dead cells from 3.18% to 0.93% (Fig. 7f). These findings clearly indicated that TPS promoted the proliferation of P4013B. Subsequently, an RNA-seq transcriptomic approach was employed to identify genes potentially driving the promoted proliferation of P4013B. TPS induced a globally transcriptional regulation of P4013B genome with a total of 1208 DEGs ( | log2 fold change | >2 and padj<0.05) that consist of 466 down-regulated DEGs and 742 up-regulated DEGs (Fig. 7g). Gene Ontology (GO) pathway and KEGG enrichment analysis indicated that DEGs are predominantly involved in ribosome and mitochondrial gene expression, metabolism of amino acids and monocarboxylic/carboxylic acids, and oxidative phosphorylation and thermogenesis (Fig. 7h), confirming their functionally important roles in cell proliferation. The most significantly changed DEGs involved in the enriched pathways were mainly assigned to metabolic enzymes (CAT1, SIM1, FOB63_000225, FOB63_000488, FOB63_000162), transporters (ITR1, FOB63_000305, FOB63_001415), and reproduction-related genes (HHT2 and PIR1), all of which were regulated by TPS incubation as assessed by qPCR assays (Fig. 7i). These findings suggested that TPS induced the proliferation of P4013B through stimulation of metabolism and cell division.
a Schematic diagram of the strategy for screening of prebiotics for P4013B proliferation from herbal polysaccharides. b Effects of 72 herbal polysaccharides on P4013B proliferation. c Growth curves of P4013B cultured with or without TPS. Statistics: two-tailed Student’s t test (c). The data represent the means ± SEM, n = 3 biological replicates in (c). d Effects of TPS (0–5 mg/mL) on the growth of P4013B. P4013B was cultured with TPS (0-5 mg/mL) for 36 h, and random sampling with different dilutions (1, 10, 100, 1000 folds) were cultured on ager plate for more 24 h. e Electron microscope of P4013B cultured with or without TPS. f Flow cytometry analysis of dead cells of P4013B cultured with or without TPS. P4013B was cultured with TPS for 36 h and stained with PI for flow cytometry analysis. g Comparative heatmap analysis of differentially expressed genes in P4013B cultured with or without TPS. Colored bars indicate standardized log2 expression intensities. Red and blue represent upregulation and downregulation, respectively. h The most significantly changed pathways according to the GO pathway and KEGG enrichment analysis. Dot size represents the number of genes annotated to each term. Red and blue represent upregulation and downregulation, respectively. i qPCR analysis of differentially expressed genes involved in cell proliferation, metabolic activity, and transport. j Schematic of gut microbiota depletion, P4013B colonization, TPS treatment and colitis induction. k Relative P4013B abundance in feces by metagenomic sequencing. P values on the top were calculated by the two-sided Wilcoxon rank-sum test. The boxplot conventions are consistent with the description in Fig. 1C. l IEt concentrations in the feces of mice by LC-MS/MS. m DAI after 3% DSS induction. n Gross morphology and length of colons. o colonoscopy and H&E-staining of colons. Scale bars, 50 μm. Statistics: one-way ANOVA with Tukey’s test (i, m, n). The data represent the means ± SEM, n = 6 biological replicates in (i, k, l, m, n). Source data are provided as a Source Data file.
In vivo facilitation on P4013B proliferation and consequent biological activity against colitis were further evaluated in DSS-induced SPF mice and ABX mice after oral administration of P4013B or TPS alone or in a combination (Fig. 7j and Supplementary Fig. 28a). In SPF mice, P4013B colonization, TPS treatment, and the combination of both attenuated DSS-induced colitis with reduced clinical colitis symptoms, alleviated mucosal damage, and improvement of colonic inflammation and gut barrier function, and the protective potency of three treatment was in the following order: the combination of P4013 with TPS treatment > P4013B colonization > TPS treatment (Supplementary Fig. 28b–e), suggesting an additive effects of P4013B and TPS. Especially, fecal IEt were increased by the treatment of TPS in the mice with P4013B colonization (Supplementary Fig. 28f), further suggesting that TPS promoted the colonization of P4013B in mice and subsequently enhanced the protective effect of P4013B against colitis. To exclude the potential interaction of inherent gut microbiota, we investigated the biological effects of TPS on the colonization and protection of P4013B in ABX mice with DSS-induced colitis (Fig. 7j). Metagenomic sequencing analysis showed that the relative abundance of P4013B in the feces of mice administered TPS was significantly increased, accompanied by an increased level of fecal IEt as assessed by LC-MS/MS (Fig. 7k, l). Interestingly, TPS failed to improve DSS-induced colitis symptoms in ABX mice, but it certainly facilitated the improvement of P4013B colonization against colitis induced by DSS (Fig. 7m–o). These findings confirmed that TPS facilitated the colonization of P4013B in vivo and subsequently enhanced the protective effect of P4013B against colitis.
Discussion
Correcting the dysbiosis of gut microbiota by FMT induces remission of IBD in the clinic, suggesting its determinative role in IBD progress. In the present study, we identified a human intestinal fungal strain, C. lusitaniae P4013B, from the CGFs we established previously5. P4013B is depleted in IBD cohorts and attenuated DSS-induced colitis in mice. Further study revealed that P4013B-derived metabolite IEt facilitated its protective effect against colitis through activation of intestinal AHR. Genetically engineering strains with overexpression of IEt-producing enzyme pyruvate decarboxylases, as well as combined herbal polysaccharide TPS, enhanced the protective effect of P4013B due to a higher yield of IEt. Our findings suggested that C. lusitaniae P4013B represents a promising probiotic for microbiota therapeutics in the prevention and treatment of IBD.
Human intestinal fungus C. lusitaniae P4013B is implicated in IBD and attenuates DSS-induced colitis in mice
The association between the dysbiosis of gut fungal community and the course of IBD has been demonstrated in geographically distinct IBD cohorts5,13,14. However, a direct causal relationship between fungal dysbiosis and IBD has not been definitively established in humans due to the lack of systemic gut fungal reference genomes, the incomplete annotation of fungi in current genomic databases, and the non-availability of fungal strains captured by deep sequencing, all of which limiting the direct evaluation of fungal effect on IBD progression. Recently, it becomes feasible by the utilization of the fungal strain resources comprising 12,453 fungal isolates from human feces in our laboratory5. Intestinal fungal species with differential abundance in IBD, like Trichosporon coremiiforme c31, Candida sp. c21, Lodderomyces elongisporus c112, and Candida metapsilosis, were found for the first time to attenuate colitis in mice after intestinal colonization2,5. Along with the thorough research, we found that C. lusitaniae P4013B is enriched in the feces of healthy subjects in three IBD-related Chinese cohorts. In addition, based on gut fungal analyzes from global population cohorts, C. lusitaniae appears to be more associated with a healthy gut environment in terms of prevalence38,39,40. However, whether the differences in abundance between IBD and healthy populations are universally applicable across different populations requires further validation through deeper sequencing and larger study cohorts in future research (Supplementary Fig. 29 and Supplementary Table 7).
The validation experiments showed that P4013B colonization attenuated DSS-induced colitis independent of inherent gut microbiota in conventional, ABX, and germ-free mice, respectively (Figs. 1, 2 and Supplementary Fig. 3). C. lusitaniae is a saprophytic yeast that has usually been isolated from the environment, food, and gastrointestinal tract of animals and humans41. It has been used in the biotechnology industry to produce ethanol and xylitol and eliminate toxic metabolites in fermented food and alcoholic beverages42,43,44,45,46. To our knowledge, it is the first time that human gut-colonizing C. lusitaniae has been found to attenuate colitis, although similar results were observed from the evaluation of C. lusitaniae 30 isolated from traditional fermented dairy food in China47. Considering the widely prevalence of C. lusitaniae in the environment and organism, P4013B may provide health benefits in the control of colitis as a probiotic fungus.
P4013B-derived metabolite IEt protects against colitis in mice through activation of intestinal AHR
As an organism, gut microbiota governs the homeostasis of metabolism, immunity, and endocrine in the host gastrointestinal tract through directly interacting with target cells or secreting the active metabolites20,48,49. In our previous studies, gut fungal species attenuated colitis through activating FXR in enterocytes via secreting secondary metabolites2,5. Given the results of the ineffectiveness of Pasteurized P4013B and the promotion of organoid formation in vitro (Figs. 1 and 2), the active metabolites of P4013B were considered to contribute to the direct protective effect against colitis. To test this hypothesis, we described an integrated systems approach for activity-guided isolation and identification of active metabolites of P4013B, combining transcriptome sequencing, luciferase assay, and chromatographic and spectroscopic methods. We found that P4013B attenuated colitis dependent on AHR activation and P4013B metabolite IEt acted as an AHR agonist to facilitate protection against colitis, which was unambiguously verified by knockout mice in vivo and site-directed mutagenesis in vitro (Figs. 3, 4 and Supplementary Figs. 4–26). AHR is a ligand-activated transcription factor to sense environmental and endogenous xenobiotics50. Intestinal AHR is considered to be a potential therapeutic target for the control of intestinal homeostasis through the management of mucosal immunity and maintaining intestinal barrier integrity. Specifically, AHR activation by dietary ligands, such as tryptophan metabolites, promotes intestinal stem cell proliferation through Wnt-β-catenin signaling, upregulates epithelial tight junction proteins dependent on the Nrf2 pathway, and maintains the balance between controlled regeneration and malignant transformation of enterocytes through Yap/Tead transcriptional regulation51,52,53. In addition, AHR in the immune cells, such as CD4+ T, ILC3, γδ T, and dendritic cells, controls the production of IL10 and IL22 to preserve the intestinal immune responses and prevent the DNA damage response in epithelial cells54,55,56. In our study, improvement of colonic inflammation and gut barrier function was observed in DSS-induced mice with P4013B colonization or IEt administration upon AHR activation (Figs. 1, 5 and Supplementary Fig. 27), which confirmed the potential clinical value of P4013B for the treatment of IBD mediated by the AHR pathway. More importantly, as a gut fungus isolated from human feces, IEt can be produced using a 24 h sustained release system in the gastrointestinal tract for uninterruptedly activation of intestinal AHR. Plus, the direct colonization of P4013B in the intestinal epithelium, the effective treatment of IBD can be reached at a low concentration of IEt. Indeed, relative high dosage of IEt (200–600 mg/kg) was used to treat DSS-induced colitis in mice in our and previous studies57, suggesting the high practicality and feasibility for utilization of IEt-producing P4013B in IBD treatment. Other microbial tryptophan metabolites, such as indole derivatives (indole-3-acetic acid, indole-3-pyruvate, indole-3-aldehyde, indole-3-propionic acid, and indole-3-lactic acid), xanthurenic and kynurenic, are considered to facilitate the alleviation of colitis by targeting intestinal AHR55,57,58,59. Among various microbial tryptophan metabolites, IEt, as the major metabolite in P4013B (Fig. 4), exhibits excellent agonistic characteristics with a potential agonistic activity, a low EC50 value and a broad range of effective concentration (Fig. 4)60. Taken together, P4013B-derived IEt represents a more effective therapy for IBD.
Yield improvement of IEt is an effective strategy to enhance the protective effect of P4013B
Considering the small portion of gut microbiota, the yield improvement of IEt represents an effective strategy to enhance the protective effect of P4013B. In the current study, two approaches were tried to enhance the production of IEt. First, genetically engineering strains with overexpression of IEt-producing enzymes were established and evaluated for their anti-colitis effect in mice (Fig. 6). Gut microbiota is now considered as a metabolism organ for the multifunction, versatility, and bio-compatibility of their enzyme system. Specific enzymes of gut microbiota facilitate the production of bioactive substances and consequently exert influence on disease progression, such as the bile salt hydrolase and 3α-hydroxysteroid dehydrogenase in the metabolism of bile acids61,62,63, the tyrosine decarboxylase in the production of tyramine64, the enzymes encoding by the choline utilization (cut) gene cluster involved in trimethylamine N-oxide production65,66. We found strains with heterologous recombination of pdc of P4013B exhibited the high production of IEt and significantly attenuated colitis in mice (Fig. 6). These findings provided an alternative solution for the potential clinical applications of IEt-producing P4013B in immunocompromised patients with the risk of invasive fungal infections67,68.
The second strategy to regulate P4013B proliferation is the combined herbal polysaccharides. Natural products, especially herbal polysaccharides, have showed great potential for modulation of immune and inflammation, in which gut microbiota predominantly functions as the medium69,70. Therefore, we screened herbal polysaccharides for the promotion of P4013 proliferation, and finally identified Tea Polysaccharide (TPS) that remarkedly enhanced the growth and intestinal colonization of P4013B and consequently strengthened its protection effect against colitis in mice (Fig. 7 and Supplementary Fig. 28). It has been reported that polysaccharides are metabolized by gut microbiota to generate short-chain fatty acids, which feeds back into growth promotion of microbiota69. Besides, polysaccharides may interact with and impact the immune homeostasis in the gastrointestinal tract and hence provide an environment suitable for the generation of microbial communities71. We found TPS induced a globally transcriptional regulation of P4013B genome predominantly related to metabolism and transport, and reproduction (Fig. 7) in vitro, suggesting a direct promotion of P4013 proliferation. Notably, TPS attenuated colitis in a microbiota-dependent manner (Fig. 7 and Supplementary Fig. 28), indicating the potential effect of TPS in improvement of ecological environment and adjustment for microflora balance. These findings not only demonstrate that IEt-producing P4013B is a promising organism with health benefits for the treatment of IBD, but also provide the solid foundation for the development of the combination of TPS and P4013B as probiotic products.
In discussing the novelty of our study, we noted that the protective effects of IEt on gut barrier function in DSS-induced colitis via AhR signaling has been reported previously57. But in the present study, we have newly identified a human-gut fungus C. lusitaniae that produces IEt and holds promise for preventing and treating IBD. C. lusitaniae can stably colonize the gut and continuously convert tryptophan to IEt at physiological concentrations by fungal pyruvate decarboxylase, achieving therapeutic AHR activation. This human-gut fungus approach not only circumvents the potential toxicity of high-dose IEt but also enhances localized delivery to the intestinal mucosa. Additionally, a higher yield of bioactive metabolite from the human-gut fungus was achieved through genetic engineering of fungal pyruvate decarboxylase and the incorporation of herbal polysaccharides TPS, enhancing its therapeutic potential for IBD. Collectively, our findings bridge a key translational gap of human-gut fungus in AHR-targeted therapies, offering a sustainable, low-risk alternative to conventional drug administration.
In conclusion, Clavispora lusitaniae is significantly depleted in patients with inflammatory bowel disease (IBD), and re-colonization with C. lusitaniae strain P4013B alleviates colitis in mice. P4013B enhances gut barrier function and reduces colonic inflammation by activating the AHR pathway. The active major metabolite IEt of P4013B acts as an AHR agonist, contributing to the protective effect of P4013B. Strategies to enhance IEt production, either through genetic engineering or in combination with herbal polysaccharide TPS, further promote the protective effect of P4013B against colitis. Together, P4013B and TPS show promise as probiotic and prebiotic agents for the treatment and prevention of IBD.
Methods
Human subjects
Fresh fecal samples collected from patients with IBD and healthy volunteers were frozen and stored at − 80 °C until metabolite determination by LC-MS/MS. The study was approved by the Ethics Committee of Xinhua Hospital affiliated with Dalian University (XH2020A008). Informed consent was obtained from all volunteers.
Animals
Animal experiments were approved and performed in accordance with the guidelines of the Institutional Animal Care and Use Committee of Dalian Medical University (approval no. AEE19044). Male C57BL/6 J mice (8–10 weeks old) were obtained from the Experimental Animal Center of Dalian Medical University. Germ-free male C57BL/6 J mice and AHR knockout (Ahr-/-) mice were provided by Cyagen Biosciences (Suzhou, China). The germ-free mice were raised in gnotobiotic isolators and given sterilized food and water. All mice were fed with standard chow (SPF grade) and water ad libitum in a specific pathogen-free (SPF) grade facility under controlled conditions (22–24 °C, 40–60% humidity) and maintained on a 12 h light/dark cycle.
Cell culture
The NCM460 cell line was acquired from American Tissue Culture Collection (ATCC, Manassas, VA, USA) and cultured in DMEM supplemented with 10% fetal bovine serum at 37 oC and 5% CO2. The culture medium was changed every two to three days, and cells were passaged when confluence reached about 80–90%.
P4013B was isolated previously in our laboratory2. Inoculating from solid medium, P4013B was cultured in liquid MTB medium at 32 °C for 48 h. Pasteurization consisted of heat treatment at 70 °C for 30 min of fresh P4013B.
Processing and profiling of publicly fecal metagenome datasets
Three publicly available fecal metagenome datasets from IBD and healthy cohorts, namely HeQ_201718, WengY_201919, and YanQ_20245, were downloaded from the National Center for Biotechnology Information (NCBI) database, with accession codes PRJEB15371, PRJNA429990, and PRJEB67456, respectively. Three datasets comprise 169 patients and 178 healthy controls. Moreover, three publicly available fecal shotgun metagenomic datasets from non-Chinese IBD cohorts, namely NielsenHB_201438, SchimerM_201839, and JiangS_202240, were downloaded from the NCBI database, with accession codes PRJEB1220, PRJNA389280, and PRJNA400072, respectively. To ensure high-quality data, we processed each metagenomic sample using fastp. The raw reads underwent several quality control steps, including trimming tails and removing low-quality reads based on the following criteria: (1) reads shorter than 90 bp; (2) reads with a mean Phred quality score below 20; (3) reads where more than 30% of bases had a Phred quality score below 20; (4) reads with a mean complexity score under 30%; and (5) unpaired-end reads. The filtered reads were then mapped to the GRCh38 reference genome using Bowtie2 with default parameters to remove human-derived sequences.
To construct a comprehensive human-associated fungal genome reference, we expanded the previously established cultivated gut fungi (CGF) catalog, which contains 760 genomes, by incorporating an additional 1490 publicly available human-associated fungi (PHF) genomes from the NCBI RefSeq database (updated June 2024). These genomes were clustered into 317 non-redundant human-associated fungal species using an average nucleotide identity (ANI) threshold of 95%. Based on this reference genome collection, we generated the gut mycobiome composition of all fecal metagenomes using a recently developed method. In brief, this method involves mapping metagenomic reads to the gene sets of the reference genome collection, resulting in the fungal relative abundances of 317 species. The relative abundance profiles at the phylum and genus levels were obtained by summing species-level abundances.
Statistical analyzes were conducted using the R 4.0.1 platform. Principal coordinates analysis (PCoA) was performed with the R vegan package based on the Bray-Curtis dissimilarly at the species level. Permutational multivariate analysis of variance (PERMANOVA) was realized with the vegan package (adonis analysis), and the p-value was generated based on 1000 permutations. For meta-analysis, we (1) converted the species abundance profiles to log10-transformed proportions, (2) employed the Hedges’g standardized mean difference statistic to calculate the pooled effect size using the escalc function from the R metafor package, and then 3) assessed was the between-study heterogeneity using Cochrane’s Q test and the I2 index. The meta-analysis p-values were obtained from the random effects models. The Wilcoxon rank-sum test was implemented using the function ‘wilcox.test’. The q-value was used to evaluate the false discovery rate (FDR) for the correction of multiple comparisons and was calculated based on the R fdrtool package.
Animal study design
A mouse model of colitis was induced by administration of 3% (w/v) DSS in drinking water for 5–7 days. In all cases, mice were sacrificed when the body weight was lower than 17 g or the scores of body weight loss, diarrhea, or rectal bleeding were 4 based on animal welfare considerations. Before sacrifice, mice were anesthetized with isoflurane-mixed gas (2.5%) to reduce pain. Tissue was collected for subsequent analysis. During the course of the experiment, the body weight, stool consistency and bleeding were measured every day to assess the disease activity index (DAI). Briefly, the DAI score is defined as follows, for weight loss, 0: no loss; 1: 1–5% loss; 2: 5–10% loss; 3: 10–20% loss and 4: > 20% weight loss; For stool consistency, 0: normal; 2: loose stool; 4: diarrhea; While for stool bleeding, 0: no blood; 2: presence and 4: gross blood.
In the initial evaluation study of three human intestinal fungal strains, the SPF mice were randomly divided into 4 groups (n = 6), including one DSS group and three colonization groups. Mice in the DSS group were inoculated by gavage with 0.2 mL saline for three weeks. Mice in the colonization group were inoculated by gavage with 0.2 mL saline containing 108 CFU of Clavispora lusitaniae P4013B, Trametes sp. AH28-2 PG5501 or Pichia kudriavzevli F2008 per mouse for three weeks. At the 15th day, all mice were administered with 3% (w/v) DSS in the drinking water for 7 days, followed by 2 days of regular drinking water before sacrifice.
In the functional evaluation study, the SPF mice were randomly divided into 4 groups (n = 6), including: (1) Vehicle control group, (2) DSS group, (3) DSS + live P4013B group, and (4) DSS + pasteurized P4013B group. Mice were inoculated by gavage with 0.2 mL saline (Groups 1 and 2), live P4013B (108 CFU per mouse, Group 3), or pasteurized P4013B (108 CFU per mouse, Group 4) for three weeks, respectively. At the 15th day, the mice in Groups 2–4 were administered 3% (w/v) DSS in the drinking water for 7 days, followed by 2 days of regular drinking water before sacrifice.
To exclude the effect of gut microbiota, the effect of P4013B colonization on colitis was investigated in the antibiotics-treated (ABX) mice and germ-free (GF) mice. The mice were randomly divided into 2 groups (n = 6 for ABX mice and (n = 5 for GF mice), including: (1) DSS group, (2) DSS + P4013B group. ABX mice were received an antibiotic cocktail (ampicillin 1 mg/mL, neomycin 1 mg/mL, vancomycin 0.5 mg/mL and metronidazole 1 mg/mL) in the drinking water for 5 d prior to P4013B colonization. Then, the study design was as same as stated above.
To evaluate the effect of P4013B-derived metabolite, the SPF mice were randomly divided into 2 groups (n = 6), including: (1) DSS group, (2) DSS + IEt group. IEt (200 mg/kg) was dissolved in 50% PEG-400 and administered via oral gavage every 12 h for 14 d. Control mice received an equal vehicle. At the 8th day, mice were subject to colitis induction by DSS.
To verify the key role of the AHR pathway, Ahr-/- mice were randomly divided into 2 groups (n = 6): (1) DSS group and (2) DSS + P4013B group or DSS + IEt group. The detail was the same as described above in SPF mice.
To evaluate the key role of IEt-producing enzymes, SPF mice were randomly divided into 3 groups (n = 6), including: (1) DSS + E. coli_ pET30a group, (2) DSS + E. coli_2062 group, (3) DSS + E. coli_5239 group. Mice were orally administrated E. coli strains (109 CFU per mouse) for two weeks, respectively. At the 8th day, mice were subject to colitis induction by DSS.
To investigate the effect of the combination of Tea Polysaccharide (TPS) and P4013B, SPF mice were randomly divided into 4 groups (n = 6), including: (1) DSS group, (2) DSS + P4013B group, (3) DSS + TPS group, and (4) DSS + P4013B + TPS group. TPS (100 mg/kg) was dissolved in saline and orally administrated alone or in combination of P4013B. Other operations were the same as described above in SPF mice. Similarly, the same study design was conducted in ABX mice.
Isolation and identification of metabolites from P4013B
To obtain sufficient metabolites for the bioactivity assay, P4013B was fermented in 300 L medium at 32 °C for 6 days and extracted using EtOAc. After the EtOAc had been evaporated in a vacuum, the residue was used to investigate active metabolites. First, 1 g of residue was analyzed by HPLC and separated by preparative HPLC to obtain 8 chromatographic fractions (Fr.1-8). After solvent evaporation, residues corresponding to Fr. 1-8 were obtained, and their agonistic effect on AHR was evaluated using luciferase assays. Active fractions were used as reference samples to isolate active metabolites from the EtOAc extract. The target subfractions containing the same constituents as Fr.6-8 were further separated by preparative HPLC to produce the Indole-3-lactic acid (ILA), Indole-3-carboxaldehyde (ICA), Indole-3-ethanol (IEt) and Indole (Ind) metabolites, which were subjected to HPLC analysis as follows: mobile phase CH3OH-H2O (0.5 ‰ CF3COOH), flow rate 0.8 mL/min. The chemical structures of the purified metabolites ILA, ICA, IEt and Ind and their authentic standards were determined using 1H NMR, 13C NMR, and HR-ESI-MS spectroscopic measurements. All 1H, 13C NMR, and HR-MS data are shown in Supplementary Figs. 6–25 and Supplementary Table 1–4.
Molecular docking assay
To evaluate the interaction and binding patterns between IEt and AHR, the structural model of human AHR (PDB: 8QMO) was downloaded from the RCSB Protein Data Bank (RCSB PDB, https://www.rcsb.org). The entire molecular docking process in this study was conducted using BIOVIA Discovery Studio (Dassault Systèmes, San Diego, USA). The AHR structure was prepared using the ‘Prepare Protein’ procedure, which included inserting missing atoms in incomplete residues, modeling missing loop regions, deleting alternate conformations (disorder), removing water molecules, standardizing atom names, and protonating titratable residues. Meanwhile, the ligand IEt was prepared using the ‘Prepare Ligand’ procedure, which involved removing duplicates, enumerating isomers and tautomers, and generating 3D conformations. The CHARMM36 force field was employed to represent the protein and ligand structures. Docking simulations were performed using a standard LibDock protocol, where protein features are referred to as hotspots. After a final energy minimization step (allowing the ligand poses to be flexible), the top-scoring ligand poses were saved. The rigid poses were placed into the active site of AHR, and the hotspots were matched as triplets. For the docking process, IEt was input, and the AHR-IEt complexes with the highest LibDock scores were selected from the docking results and depicted in full text. PyMOL and LigPlot + were used for visualization72.
Design and application of the FISH probe
The FISH probe consists of an 18S rRNA sequence conjugated at its 5’-terminus with Cy5 fluorophore: 5’-CCGTAGGTGAACCTCCGG-Cy5-3’.
Freshly isolated sterile murine colonic tissues were fixed, paraffin-embedded and sectioned. Sections underwent deparaffinization followed by rehydration through graded alcohols. Following this, tissue sections underwent Proteinase K-mediated permeabilization for 10 min at room temperature. Subsequent pre-hybridization was conducted at 37 °C for 20 min, followed by probe hybridization under light-protected conditions at 37 °C for 2 h. Sections were then subjected to stepwise stringency washes, counterstained with DAPI, and mounted with antifade reagent prior to image acquisition via laser scanning confocal microscopy.
Measurement of metabolites
The concentrations of fungal metabolites in fecal specimens from humans or mice were determined by LC-MS/MS. Fecal specimens from humans collected in our previous study2 were used. Frozen samples were dried on Freeze Dryer for 12 h. The dried sample was weighed, crushed and dissolved in 1 mL methanol. After 30 min of ultrasound, the supernatant was centrifuged and then blowdried. Immediately prior to LC-MS analysis, the samples were resuspended in 50% acetonitrile. The sample analysis was performed on an AB Sciex Qtrap 5500 LC-MS/MS. Quantification of metabolites was determined by integrating the extracted ion count of the exact masses of the metabolites, which were determined using commercial standards. The MRM transitions and the related optimized declustering potential (DP), entrance potential (EP), collision energy (CE), and collision cell exit potential (CXP) for the different analytes are listed in Supplementary Table 8. Analyst 1.6.3 software (Applied Biosystems) was used to control the equipment and for data acquisition and analysis. Standard curves in which known amounts of metabolite were utilized to determine the amount of each metabolite in each sample, which was normalized to the dry weight of the fecal samples.
RNA isolation and qPCR
Total RNA was isolated from the colon tissues of mice or P4013B using AG RNAex Pro reagent (Accurate Biotechnology) and purified using lithium chloride precipitation to remove DSS. cDNA was generated from 1 μg of total RNA using a MonScript™ RTIII All-in-One Mix (Monad Biotechnology, Wuhan, China) and was amplified using SYBR Green qPCR Mix (Monad Biotechnology, Wuhan, China) with an ABI PRISM® 7500 Real-Time PCR System (Applied Biosystems, Thermo Fisher Scientific, USA). Relative target mRNA expression was normalized to the housekeeping gene β-actin using the comparative ΔΔCt method. The primer sequences are summarized in Supplementary Table 9.
Western blot and immunofluorescence analysis
Western blot proteins were extracted from NCM460 cells or colon by Cell lysis buffer and Nuclear and Cytoplasmic Protein Extraction Kit (Beyotime, Shanghai, China) supplemented with protease inhibitor and phosphatase inhibitor. Protein concentration was measured by Pierce BCA Protein Assay Kit (Beyotime, Shanghai, China) according to the manufacturer’s instructions. Extracted proteins were separated by 10% SDS-PAGE gel electrophoresis and transferred onto PVDF membranes with a pore size of 0.45 µm (Merck, Darmstadt, Germany). Western blots were probed with specific primary antibodies and incubated with the corresponding secondary antibodies. Proteins were visualized using High-sig ECL western blotting substrate (Tanon, Shanghai, China). β-Actin was used as a standard protein.
NCM460 cells were grew on 20 mm Circle Microscope Cover Glass. And cells were incubated with IEt or P4013B for 24 h and fixed with 4% paraformaldehyde for 10 min. Colonic tissues fixed in neutral 10% buffered formalin were cut into 5 μm thick sections in paraffin. Then, cells or tissues were stained with primary antibodies and secondary fluorescein isothiocyanate-conjugated antibodies. Cell nuclei were stained with DAPI (Beyotime, Shanghai, China). Fluorescence images were captured using a confocal laser scanning microscope (Leica SP8, Germany).
The primary antibodies used in Western blot and immunofluorescence analysis are summarized in Supplementary Table 10.
AHR activity assay
The HEK 293 T cells were placed in a 96-well plate at 8000 cells/well. At 50-60% confluency, the cells were transfected with 0.4 µg each of AHR and CYP1A1 promoter plasmids (Youbao Biotechnology) per well. After 6 h, the media was replaced and the cells were treated for 24 h with P4013B metabolites or DMSO of equivalent volume. After 24 h, cells were lysed with PLB, and 40 µL of lysate was added to a 96-well plate. Afterwards, 50 µL of firefly luciferase substrate was added to each well, and luminescence was measured using a Tecan-Spark Microplate Luminometer instantly. Immediately after, 50 µL of the Renilla substrate was added to each well, and luminescence was measured. Luciferase activity was determined by subtracting a blank and calculating the ratio of the glowworm luciferase signal to the Renilla luciferase signal.
Preparation of overexpressed strains
The recombinant plasmids were transfected into an engineered strain Escherichia coli BL21, and uniformly coated on an LB plate containing kanamycin. The next day, a single colony was picked from agar plates and incubated in LB media with kanamycin until the OD600 values reached 0.6. At which point, Isopropyl β-d-1-thiogalactopyranoside (IPTG) was added to induce the expression of the target proteins with a final concentration of 1 mM at 16 °C. After 16 h of induction, the cells were washed and resuscitated into TM1 buffer (glucose 30 g/L, KH2PO4 5 g/L, MgSO4 0.5 g/L, NaCl 1 g/L). At this point, with the addition of Trp, the overexpressed strains were incubated for 24 h at 37 °C for the measurement of metabolites.
Screening for bioactive polysaccharides
P4013B was cultured with virus herbal polysaccharides (5 mg/mL) for 24 h, and the OD600 nm values of the culture were determined by a microplate reader. The proliferation without polysaccharide was taken as 100%. The different colors represented different relative proliferation rates. White and red represented weak and strong proliferation potentials, respectively. 72 herbal polysaccharides used for the screening of prebiotic effects were listed in Supplementary Table 6.
Growth curves
P4013B was cultured to logarithmic growth and then passaged to low-glucose (0.1%) MTB at 1:10,000. The control group and the TPS (5 mg/mL) group were set up, and the 600 nm absorbance value was measured every 4 h until 36 h.
Flow cytometry analysis
P4013B was incubated with TPS (5 mg/mL) for 24 h and then centrifuged at 5000 × g for 3 min, washed and resuspended in phosphate-buffered saline. Dead cell viability staining was performed using propidium iodide (PI) as detailed by the manufacturer (Beyotime, Shanghai, China). A BD Accuri™ C6 Plus Flow Cytometer instrument (BD Biosciences, New York, USA) was used to determine the percentage of PI-positive cells.
Statistical analysis
Experimental subjects/preparations were randomized to groups, and group assignments, data recording, and data analysis were blinded to the operator. All experiments were independently repeated at least three times with consistent results. Specifically, hematoxylin and eosin (HE) staining, fluorescence microscopy, and western blotting (WB) analyzes were each performed with three biological replicates (n = 3), unless otherwise stated. All data were presented as means ± standard error of the mean (SEM) and analyzed using GraphPad Prism V.9.5 (GraphPad Software, USA). All measurements of sample size (n) were taken from distinct biological replicates. Statistical significance between two groups was compared using a two-sided Student’s t test, and statistical significance from more than two groups were compared using one-way ANOVA followed by post-hoc Tukey’s test.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Fecal metagenome datasets from IBD and healthy cohorts were downloaded from the National Center for Biotechnology Information (NCBI) database, with accession codes PRJEB15371, PRJNA429990, PRJEB67456, PRJEB1220, PRJNA389280, and PRJNA400072.
The RNA-seq data for mouse colons generated in this study have been deposited in the Sequence Read Archive (SRA) database under the accession codes PRJNA1266285 and PRJNA1271697. The mouse fecal metagenome data generated in this study have been deposited in the Sequence Read Archive (SRA) database under the accession codes PRJNA1297366 and PRJNA1281509.
The data supporting the findings from this study are available within the manuscript and its supplementary information.Source data are provided in this paper.
Code availability
The data processing and analysis codes, genomic and functional annotations, gut mycobiome profiles, and statistical scripts used in this study are publicly available at https://github.com/yexianingyue/Cultivated-Gut-Fungi.
References
Aykut, B. et al. The fungal mycobiome promotes pancreatic oncogenesis via activation of MBL. Nature 574, 264–267 (2019).
Huo, X. et al. Cultivated human intestinal fungus Candida metapsilosis M2006B attenuates colitis by secreting acyclic sesquiterpenoids as FXR agonists. Gut 71, 2205–2217 (2022).
Sun, H. et al. Gut commensal Parabacteroides distasonis alleviates inflammatory arthritis. Gut 72, 1664–1677 (2023).
Wang, X. et al. Candida albicans accelerates atherosclerosis by activating intestinal hypoxia-inducible factor2α signaling. Cell Host Microbe 32, 964–979 (2024).
Yan, Q. et al. A genomic compendium of cultivated human gut fungi characterizes the gut mycobiome and its relevance to common diseases. Cell 187, 2969–2989 (2024).
Ma, C., Solitano, V., Danese, S. & Jairath, V. The future of clinical trials in inflammatory bowel disease. Clin. Gastroenterol. Hepatol. 23, 480–489 (2025).
Wang, R., Li, Z., Liu, S. & Zhang, D. Global, regional and national burden of inflammatory bowel disease in 204 countries and territories from 1990 to 2019: a systematic analysis based on the Global Burden of Disease Study 2019. Bmj. Open. 13, e065186 (2023).
Kaplan, G. G. & Windsor, J. W. The four epidemiological stages in the global evolution of inflammatory bowel disease. Nat. Rev. Gastroenterol. Hepatol. 18, 56–66 (2021).
Ma, C. et al. Interpreting modern randomized controlled trials of medical therapy in inflammatory bowel disease. Nat. Rev. Gastroenterol. Hepatol. 21, 792–808 (2024).
Yadegar, A. et al. Fecal microbiota transplantation: current challenges and future landscapes. Clin. Microbiol. Rev. 37, e0006022 (2024).
Bethlehem, L. et al. Microbiota therapeutics for inflammatory bowel disease: the way forward. Lancet Gastroenterol. Hepatol. 9, 476–486 (2024).
Dalmasso, G. et al. Saccharomyces boulardii inhibits inflammatory bowel disease by trapping T cells in mesenteric lymph nodes. Gastroenterology 131, 1812–1825 (2006).
Sokol, H. et al. Fungal microbiota dysbiosis in IBD. Gut 66, 1039–1048 (2017).
Mukherjee, P. K. et al. Mycobiota in gastrointestinal diseases. Nat. Rev. Gastroenterol. Hepatol. 12, 77–87 (2015).
Limon, J. J. et al. Malassezia is associated with Crohn’s sisease and exacerbates colitis in mouse models. Cell Host Microbe 25, 377–388 (2019).
Li, X. V. et al. Immune regulation by fungal strain diversity in inflammatory bowel disease. Nature 603, 672–678 (2022).
Li, X. V., Leonardi, I. & Iliev, I. D. Gut mycobiota in immunity and inflammatory disease. Immunity 50, 1365–1379 (2019).
He, Q. et al. Two distinct metacommunities characterize the gut microbiota in Crohn’s disease patients. Gigascience 6, 1–11 (2017).
Weng, Y. J. et al. Correlation of diet, microbiota and metabolite networks in inflammatory bowel disease. J. Dig. Dis. 20, 447–459 (2019).
Lavelle, A. & Sokol, H. Gut microbiota-derived metabolites as key actors in inflammatory bowel disease. Nat. Rev. Gastroenterol. Hepatol. 17, 223–237 (2020).
Sedhom, M. A. et al. Neutralisation of the interleukin-33/ST2 pathway ameliorates experimental colitis through enhancement of mucosal healing in mice. Gut 62, 1714–1723 (2013).
D’Alessio, S. et al. VEGF-C-dependent stimulation of lymphatic function ameliorates experimental inflammatory bowel disease. J. Clin. Invest. 124, 3863–3878 (2014).
Scaldaferri, F. et al. VEGF-A links angiogenesis and inflammation in inflammatory bowel disease pathogenesis. Gastroenterology 136, 585–595 (2009).
Levi-Galibov, O. et al. Heat Shock Factor 1-dependent extracellular matrix remodeling mediates the transition from chronic intestinal inflammation to colon cancer. Nat. Commun. 11, 6245 (2020).
Trilleaud, C. et al. Agonist anti-ChemR23 mAb reduces tissue neutrophil accumulation and triggers chronic inflammation resolution. Sci. Adv. 7, eabd1453 (2021).
Li, F., Zhu, W. & Gonzalez, F. J. Potential role of CYP1B1 in the development and treatment of metabolic diseases. Pharmacol. Ther. 178, 18–30 (2017).
Stevens, F. M. & Shaw, C. Prolactin-like immunoreactivity in human small-intestinal mucosa. Br. Med. J. 284, 1014–1015 (1982).
Huang, J. et al. Genetic and epigenetic silencing of SCARA5 may contribute to human hepatocellular carcinoma by activating FAK signaling. J. Clin. Invest. 120, 223–241 (2010).
Xiong, Z. et al. Small extracellular vesicles derived from adipose mesenchymal stem cells alleviate intestinal fibrosis by inhibiting the FAK/Akt signaling pathway via MFGE8. J. Gastroenterol. 59, 1092–1106 (2024).
Zhu, Y. et al. The multilevel extensive diversity across the cynomolgus macaque captured by ultra-deep adaptive immune receptor repertoire sequencing. Sci. Adv. 10, eadj5640 (2024).
Stadnicki, A. et al. Kallikrein-kininogen system activation and bradykinin (B2) receptors in indomethacin induced enterocolitis in genetically susceptible Lewis rats. Gut 43, 365–374 (1998).
Sissung, T. M., Price, D. K., Sparreboom, A. & Figg, W. D. Pharmacogenetics and regulation of human cytochrome P450 1B1: implications in hormone-mediated tumor metabolism and a novel target for therapeutic intervention. Mol. Cancer Res. 4, 135–150 (2006).
Dou, H. et al. Aryl hydrocarbon receptor (AhR) regulates adipocyte differentiation by assembling CRL4B ubiquitin ligase to target PPARγ for proteasomal degradation. J. Biol. Chem. 294, 18504–18515 (2019).
Fu, J. et al. Shared epitope-aryl hydrocarbon receptor crosstalk underlies the mechanism of gene-environment interaction in autoimmune arthritis. Proc. Natl. Acad. Sci. USA 115, 4755–4760 (2018).
Dai, S. et al. Structural insight into the ligand binding mechanism of aryl hydrocarbon receptor. Nat. Commun. 13, 6234 (2022).
Song, Y. et al. Molecular and structural basis of interactions of vitamin D3 hydroxyderivatives with aryl hydrocarbon receptor (AhR): An integrated experimental and computational study. Int. J. Biol. Macromol. 209, 1111–1123 (2022).
Gong, X. et al. Effects of tryptophan and phenylalanine on tryptophol production in Saccharomyces cerevisiae revealed by transcriptomic and metabolomic analyses. J. Microbiol. 60, 832–842 (2022).
Nielsen, H. B. et al. Identification and assembly of genomes and genetic elements in complex metagenomic samples without using reference genomes. Nat. Biotechnol. 32, 822–828 (2014).
Schirmer, M. et al. Dynamics of metatranscription in the inflammatory bowel disease gut microbiome. Nat. Microbiol. 3, 337–346 (2018).
Jiang, S. et al. Establishing a novel inflammatory bowel disease prediction model based on gene markers identified from single nucleotide variants of the intestinal microbiota. Imeta 1, e40 (2022).
Rojas, O. C., Montoya, A. M. & Treviño-Rangel, R. J. Clavispora lusitaniae: From a saprophytic yeast to an emergent pathogen. Fungal Biol. 128, 1933–1938 (2024).
Ochoa-Chacón, A. et al. Fermentation performance of a Mexican native Clavispora lusitaniae strain for xylitol and ethanol production from xylose, glucose and cellobiose. Enzym. Microb. Technol. 160, 110094 (2022).
Zhao, Y. et al. Advancements in fermented beverage safety: Isolation and application of Clavispora iusitaniae Cl-p for ethyl carbamate degradation and enhanced flavor profile. Microorganisms 12, 882 (2024).
Díaz, M. A. et al. Protection of citrus fruits from postharvest infection with Penicillium digitatum and degradation of patulin by biocontrol yeast Clavispora lusitaniae 146. Microorganisms 8, 1477 (2020).
Estrada-Ávila, A. K., González-Hernández, J. C., Calahorra, M., Sánchez, N. S. & Peña, A. Xylose and yeasts: A story beyond xylitol production. Biochim. Biophys. Acta Gen. Subj. 1866, 130154 (2022).
Fan, G. et al. Isolation and identification of a high-yield ethyl caproate-producing yeast from Daqu and optimization of its fermentation. Front. Microbiol. 12, 663744 (2021).
Meng, Y. et al. Saccharomyces cerevisiae I4 Showed Alleviating Effects on Dextran Sulfate Sodium-Induced Colitis of Balb/c Mice. Foods 11, 1436 (2022).
Kayama, H., Okumura, R. & Takeda, K. Interaction between the microbiota, epithelia, and immune cells in the intestine. Annu. Rev. Immunol. 38, 23–48 (2020).
Gomes, A. C., Hoffmann, C. & Mota, J. F. The human gut microbiota: Metabolism and perspective in obesity. Gut Microbes 9, 308–325 (2018).
Stockinger, B., Shah, K. & Wincent, E. AHR in the intestinal microenvironment: safeguarding barrier function. Nat. Rev. Gastroenterol. Hepatol. 18, 559–570 (2021).
Metidji, A. et al. The environmental sensor AHR protects from inflammatory damage by maintaining intestinal stem cell homeostasis and barrier integrity. Immunity 49, 353–362 (2018).
Singh, R. et al. Enhancement of the gut barrier integrity by a microbial metabolite through the Nrf2 pathway. Nat. Commun. 10, 89 (2019).
Shah, K. et al. Cell-intrinsic Aryl Hydrocarbon receptor signalling is required for the resolution of injury-induced colonic stem cells. Nat. Commun. 13, 1827 (2022).
Gronke, K. et al. Interleukin-22 protects intestinal stem cells against genotoxic stress. Nature 566, 249–253 (2019).
Michaudel, C. et al. Rewiring the altered tryptophan metabolism as a novel therapeutic strategy in inflammatory bowel diseases. Gut 72, 1296–1307 (2023).
Xie, L. W. et al. Microbiota-derived I3A protects the intestine against radiation injury by activating AhR/IL-10/Wnt signaling and enhancing the abundance of probiotics. Gut Microbes 16, 2347722 (2024).
Scott, S. A., Fu, J. & Chang, P. V. Microbial tryptophan metabolites regulate gut barrier function via the aryl hydrocarbon receptor. Proc. Natl. Acad. Sci. USA 117, 19376–19387 (2020).
Lamas, B. et al. CARD9 impacts colitis by altering gut microbiota metabolism of tryptophan into aryl hydrocarbon receptor ligands. Nat. Med. 22, 598–605 (2016).
Wang, G. et al. Microbiota-derived indoles alleviate intestinal inflammation and modulate microbiome by microbial cross-feeding. Microbiome 12, 59 (2024).
Meynier, M. et al. AhR/IL-22 pathway as new target for the treatment of post-infectious irritable bowel syndrome symptoms. Gut Microbes 14, 2022997 (2022).
Collins, S. L., Stine, J. G., Bisanz, J. E., Okafor, C. D. & Patterson, A. D. Bile acids and the gut microbiota: metabolic interactions and impacts on disease. Nat. Rev. Microbiol. 21, 236–247 (2023).
Song, X. et al. Microbial bile acid metabolites modulate gut RORγ(+) regulatory T cell homeostasis. Nature 577, 410–415 (2020).
Sun, L. et al. Gut microbiota and intestinal FXR mediate the clinical benefits of metformin. Nat. Med. 24, 1919–1929 (2018).
Li, C. et al. Enterococcus-derived tyramine hijacks α(2A)-adrenergic receptor in intestinal stem cells to exacerbate colitis. Cell Host Microbe 32, 950–963 (2024).
Zhu, W. et al. Gut microbial metabolite TMAO enhances platelet hyperreactivity and thrombosis risk. Cell 165, 111–124 (2016).
Roberts, A. B. et al. Development of a gut microbe-targeted nonlethal therapeutic to inhibit thrombosis potential. Nat. Med. 24, 1407–1417 (2018).
Banday, A. Z. et al. False-positive HIV serology, Candida lusitaniae pneumonia, and a novel mutation in the CYBB gene. Immunobiology 226, 152110 (2021).
Du, H. et al. Candida auris: Epidemiology, biology, antifungal resistance, and virulence. PLoS Pathog. 16, e1008921 (2020).
Holscher, H. D. Dietary fiber and prebiotics and the gastrointestinal microbiota. Gut Microbes 8, 172–184 (2017).
Martel, J. et al. Anti-obesogenic and antidiabetic effects of plants and mushrooms. Nat. Rev. Endocrinol. 13, 149–160 (2017).
Murphy, E. J. et al. Polysaccharides-Naturally Occurring Immune Modulators. Polymers 15, 2373 (2023).
Wu, Y. Z. et al. Bysspectin A, an unusual octaketide dimer and the precursor derivatives from the endophytic fungus Byssochlamys spectabilis IMM0002 and their biological activities. Eur. J. Med. Chem. 145, 717–725 (2018).
Acknowledgements
This work was financially supported by the National Natural Science Foundation of China (No. 82225048 to X.M., 82474340 to X.H.), Liaoning Provincial Natural Science Foundation (2024-MS-147 to X.H.), Liaoning Revitalization Talents Program (XLYC2402004 to C.W.), 1 + X Research Project of the Second Hospital of Dalian Medical University (LYYH2024002 to X.M.).
Author information
Authors and Affiliations
Contributions
X.H., F.W., Y.Wang, Y.L., and J.L. contributed toward the animal studies. C.W., F.W., Y.Wang, Z.M., Y.L., J.L., and R.Y. isolated and identified the metabolites. F.W., Z.M., Y.Wu., and Z.Y. performed in vitro experiments and MD simulations. Z.X., S.L., Y.Wu., X.F., and X.T. processed and profiled fecal metagenome datasets. Z.Y., X.F., and X.T. collected fresh human stools. X.H., F. W., C. W., and X. M. wrote the manuscript. X. H., C. W., and X. M. conceived and designed the experiments, interpreted the data and revised the manuscript. All authors discussed the results and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Philip Busbee, and the other anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wu, F., Wang, Y., Mai, Z. et al. Human intestinal fungus Clavispora lusitaniae attenuates colitis through Pyruvate decarboxylase-derived Indole-3-ethanol. Nat Commun 16, 9980 (2025). https://doi.org/10.1038/s41467-025-64914-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-64914-w