Abstract
Gut microbiota editing represents a promising therapeutic strategy for dysbiosis-associated diseases. Bacteriophages (phages), with their host specificity, enable precise microbial manipulation but face challenges such as environmental vulnerability and low bioavailability, which limit their in vivo efficacy. Here, we develop double-responsive hydrogel microspheres (HMs) via electrohydrodynamic spraying to enhance oral phage delivery. Composed of sodium alginate, hyaluronic acid, and Eudragit S100, these HMs achieve 90% encapsulation efficiency for a Salmonella-targeting phage cocktail. Such formulation significantly protects phages from gastric conditions, prolongs their intestinal retention, and enables responsive payload release in the colon. In a murine model of Salmonella Typhimurium (STm)-induced colitis, HMs-encapsulated phages (HMs-Phages) reduce intestinal STm burden by nearly 2000-fold and lower levels of proinflammatory cytokines (TNF-α, IL-6, IL-1β) to 60% of those in infected group. Notably, HMs-Phages achieve potent antibacterial efficacy comparable to ciprofloxacin while selectively targeting STm. This targeted strategy circumvents antibiotics-associated microbiota dysbiosis and diarrhea, thereby effectively restoring gut homeostasis and improving host physical health. By integrating targeted pathogen eradication with microbiota conservation, this work provides a precise toolkit for gut microbiota editing and phage therapy, offering substantial advantages over antibiotics for managing dysbiosis-related diseases.
Similar content being viewed by others
Introduction
Intestinal bacteria are the predominant microbial inhabitants of the gut and play a pivotal role in maintaining physical health. The disruption of gut homeostasis is closely associated with the occurrence and development of various diseases1,2,3,4. Intestinal pathogen infections can induce inflammatory responses and severely disrupt the balance of intestinal microbiota. Traditional antibiotic therapy indiscriminately eliminates both pathogenic and commensal/probiotic bacteria, further exacerbating the gut microbiota dysbiosis and influencing metabolic transformations5. Moreover, the rapid emergence and global dissemination of antibiotic-resistant bacteria pose a severe threat to public health6. These challenges underscore the urgent need for innovative therapeutic strategies, such as gut microbiota editing, to precisely modulate levels of specific bacterial species in the gut microbial community. However, the inherent complexity, diversity, and dynamic nature of the intestinal ecosystem pose substantial barriers to elucidating the etiology of dysbiosis and developing targeted intervention strategies. Current intervention approaches, such as probiotics, fecal microbiota transplants, and dietary changes, are limited by transient effects, safety concerns, and unstable outcomes7,8,9. These limitations highlight the necessity of advancing emerging, precise, and reliable approaches for the therapeutic modulation of the gut microbiota.
Bacteriophages (phages), which are bacterial viruses characterized by high host specificity, have a unique ability to precisely target and manipulate specific microbes within the microbiota. Beyond their host specificity, phages exhibit several desirable attributes, including rapid proliferation, abundant resources, and ease of physicochemical modification10. These features make phages promising tools for the precise modulation of the gut microbiome11. Phages have been extensively studied for treating multidrug-resistant infections, including those of the lungs, urinary tract, bones, and joints, and severe systemic conditions such as sepsis12. Despite their potential, phage-based interventions targeting the gastrointestinal tract and gut microbiota remain largely empirical and often lead to variable outcomes13. Case studies on phage therapy underscore that therapeutic efficacy depends on the bactericidal activity, specific biological properties of the phages employed, and their effective concentration at the infection site14,15. Although bicarbonate (e.g., sodium bicarbonate) has been used to facilitate the stomach passage of acid-sensitive phages, the multistep administration process, variable dosage, and potential side effects pose significant limitations to its practical application in phage therapy16,17. More importantly, successful gut microbiota editing requires additional considerations, including oral safety, intestinal retention time, and the intricate interactions between phages, target bacterial species, and other microbiota members. These factors are critical to achieve precise modulation and stable ecological outcomes.
Recently, various delivery carriers, including emulsion, liposomes, polymer particles and hydrogel microspheres, have been developed to enhance the biological function of phages during oral administration18. Among these carriers, hydrogel microspheres have garnered widespread attention owing to their prominent merits, such as high encapsulation efficiency, strong adhesion ability, excellent biocompatibility, and favorable mechanical properties. Although several studies have successfully enhanced the acid stability of encapsulated phages using hydrogel microspheres in vitro, translational research on microsphere-based carriers for phage therapy and gut microbiota editing remains limited, particularly in vivo studies19,20,21. To date, only one study reported an engineered lysogenic phage λ expressing a programmable dCas9 that precisely repressed a targeted E. coli gene in vivo using an oral microsphere carrier22. Electrohydrodynamic spraying (electrospray) offers a simple and innovative method to prepare hydrogel microspheres ranging from the nano- to micrometer scale through the cross-linking and droplet-curing of atomized precursor solution23. Notably, electrospray does not involve organic solvents or thermal processes, making it well suited for encapsulating fragile entities (e.g., probiotics and phages). Additionally, the microsphere size can be tailored by adjusting the process parameters such as the solution concentration and applied voltage.
In this study, to control phage dosage and extend intestinal retention time, we introduce a simple and green preparation method based on edible materials for encapsulating and delivering phages (Fig. 1). First, we evaluate the antibacterial properties of several Salmonella phages and identify an optimal phage combination for therapeutic application. We then use sodium alginate (SA), hyaluronic acid (HA), and Eudragit S100 (ES), all FDA-approved and generally recognized as safe (GRAS), to fabricate SA/HA/ES hydrogel microspheres for targeted phage delivery. Employing the electrospray strategy allows for flexible control over microsphere size (ranging from 100 to 900 μm), enabling customization to meet the diverse requirements of animal models in preclinical trials. SA/HA/ES hydrogel microspheres (HMs) effectively enhance the biological stability of phages and enable their responsive release into the intestine, ensuring stable and targeted phage delivery. Using a Salmonella Typhimurium (STm)-induced dysbiosis mouse model, we demonstrate that HMs-encapsulated Salmonella phages (HMs-Phages) significantly reduce intestinal STm burden and improve the physical condition of treated mice compared to free phages administered empirically. To highlight the therapeutic value of targeted phage delivery, ciprofloxacin (CIP)—a clinically used broad-spectrum antibiotic—is evaluated in parallel as a representative of indiscriminate antibacterial approaches. Importantly, our targeted approach circumvents the severe gut dysbiosis and diarrhea commonly associated with antibiotic treatment, achieving in situ gut microbiota editing. By concurrently enabling targeted pathogen eradication and commensal microbiota conservation, this study paves the way for further investigation into the etiology and therapeutics of microbiota-associated disorders.
SA/HA/ES hydrogel microspheres were fabricated to load Salmonella phages via electrohydrodynamic spraying and calcium ion crosslinking. The microspheres shielded phages from the harsh gastric environment and enabled pH-responsive release in the intestine, leading to efficient pathogen clearance, reduced proinflammatory cytokines, restoration of gut microbiota homeostasis, and alleviation of colitis symptoms. SA sodium alginate. HA hyaluronic acid. ES Eudragit S100.
Results
Screening of efficient phage combinations
Three Salmonella phages, LPST83, LPST94, and LPST153, were selected for this study based on our previous studies, which demonstrated well-defined genetic backgrounds, broad lytic ranges, and strong antibacterial abilities against Salmonella strains24,25,26. The antimicrobial activity of the phages against the model Salmonella strain was assessed using inhibition curves (Fig. 2a). Transmission electron microscopy (TEM) revealed that all three phages had typical icosahedral heads and tails (Fig. 2b), classifying them within the Caudoviricetes class27. Phylogenetic analysis identified LPST83, LPST94, and LPST153 as members of the genera Segzyvirus, Kuttervirus, and Berlinvirus, respectively24,25,26. These phages displayed differences in tail structure (long and non-contractile tail, contractile tail, and short tail) and plaque morphology (Fig. 2b and Supplementary Fig. 1). The antibacterial results indicated that when used individually, none of the three phages effectively suppressed bacterial growth, regardless of the multiplicity of infection (MOI) (Fig. 2c-e). In contrast, the phage combinations exhibited significantly improved antibacterial activity. Specifically, the combination of LPST94 and LPST153 completely inhibited bacterial growth for 24 h under high MOI conditions (MOI ≥ 10) (Fig. 2f). Similarly, the combination of all three phages effectively controlled pathogen growth for 24 h at an MOI ≥ 100 (Supplementary Fig. 2).
a Schematic diagram of the method for evaluating the antibacterial activity of phages against STm. b Transmission electron microscopy images of LPST83, LPST94, and LPST153 phages (n = 3 independent samples per phage). Scale bar, 50 nm. c–f Antimicrobial activity of single phage (LPST83, LPST94, LPST153) or phage combination (LPST94 and LPST153) against STm at different MOI. Data are presented as mean ± standard deviation (n = 3 biological replicates). g Essential conditions for phage application in pathogen control. MOI, multiplicity of infection. Source data are provided as a Source Data file.
The above findings are consistent with those of previous literatures, highlighting that the antibacterial efficacy and therapeutic outcomes of phages are closely dependent on the MOI at the infection site28. Moreover, the combination of multiple phages demonstrated superior efficacy in inhibiting pathogens compared with the use of a single phage. This enhanced performance was possibly due to each phage targeting different bacterial receptors, thus minimizing phage resistance25,26. Using fewer phage participants simplifies the operational and preparative procedures, thereby reducing application complexity. Based on these considerations, the LPST94 and LPST153 combination was selected for subsequent experiments. To further evaluate their efficacy, in vitro phage-resistance of STm was examined. The results showed that LPST94-resistant STm remained sensitive to LPST153 (Supplementary Fig. 3). These in vitro results highlighted three critical conditions for phage application: selecting effective phage participants, achieving a high phage titer, and employing a phage combination (Fig. 2g).
Fabrication and characterization of HMs-Phages
Oral administration is an effective approach to deliver phages to the intestine. However, many acid-sensitive phages are rapidly inactivated in the stomach because of the harsh acidic environment22,29. To address this issue, SA/HA/ES hydrogel microspheres were fabricated using a portable electrospraying platform to protect phages (Fig. 3a). Microscopic images and the microsphere size of the hydrogel microspheres with phages (HMs-Phages) are shown in Fig. 3b and Supplementary Fig. 4. The results showed that microspheres sizes ranged from 133 ± 19 to 890 ± 25 μm (SA/HA/ES-1 to SA/HA/ES-6) depending on the precursor solution concentration (1% to 6%), demonstrating a facile and feasible strategy for tailoring microsphere size. The increase in microsphere size was likely attributable to the higher viscosity of the polymer solution at elevated polymer concentrations, which enhanced polymer chain entanglement and led to the formation of larger atomized microdroplets, ultimately producing larger microspheres (Supplementary Fig. 5). As a comparison, non-electrosprayed SA/HA/ES microspheres were also fabricated using a conventional extrusion-dripping method, exhibiting larger particle sizes and narrow size range (1,744 ± 76 μm to 2,216 ± 25 μm) under corresponding concentrations (Supplementary Fig. 6 and Supplementary Fig. 7). This large physical size significantly limits their suitability for precision gastrointestinal delivery applications in preclinical animal models.
a Schematic diagram of the preparation platform for phage encapsulation in hydrogel microspheres. b Microscopy of different HMs-Phages (n = 3 independent samples with similar results). Scale bar, 500 μm. c SEM images of different HMs-Phages (n = 3 independent samples with similar results). Upper scale bar, 100 μm; lower scale bar, 10 μm. d Elemental mapping of HMs-Phages (SA/HA/ES-3) (n = 3 independent samples with similar results). Scale bar, 100 μm. e Fourier transform infrared spectroscopy of HMs-Phages (SA/HA/ES-3). f Encapsulation efficiency of different hydrogel microspheres for phages. Data are presented as mean ± standard deviation (n = 3 biological replicates). Phages LPST94 and LPST153 were combined for microsphere preparation and subsequent evaluations. SA sodium alginate. HA hyaluronic acid. ES Eudragit S100. HMs hydrogel microspheres. Source data are provided as a Source Data file.
Scanning electron microscopy (SEM) analysis was performed to further examine the surface morphology of the electrosprayed microspheres (Fig. 3c). A more compact and denser surface was observed with increasing precursor solution concentrations, possibly providing better protection for the phages against the harsh environments. A similar phenomenon was also observed in the SEM imaging of non-electrosprayed microspheres (Supplementary Fig. 8). To evaluate the compositional homogeneity within microspheres, cross-sectional transmission electron microscopy was performed. SA/HA/ES-3 was selected as the representative sample due to its regular morphology and uniform size distribution compared to the other groups (Fig. 3b and Supplementary Fig. 4). The TEM micrographs confirmed homogeneous dispersion of the constituent polymers (SA, HA, and ES) throughout the hydrogel network (Supplementary Fig. 9). Furthermore, fluorescence imaging revealed that Cy3-labeled HA (red fluorescence) and AF 488-conjugated ES (green fluorescence) were uniformly distributed within the microsphere matrix (Supplementary Fig. 10).
To demonstrate the successful encapsulation of phages within the microsphere, elemental mapping analysis was conducted. The results showed that the phages were uniformly dispersed in the microspheres, as evidenced by the presence of phosphorus, sulfur, and nitrogen elements (Fig. 3d). In addition, Fourier transform infrared (FTIR) spectroscopy further confirmed phage loading, as indicated by the appearance of an amide II peak at 1545 cm-1, a characteristic marker for phage capsid proteins (Fig. 3e). Consistent with these findings, all SA/HA/ES hydrogel microspheres exhibited high encapsulation efficiencies of approximately 90% (Fig. 3f), indicating a gentle and efficient method for phage loading. Moreover, the number of phages encapsulated in each microsphere rose with increasing microsphere size (Supplementary Fig. 11). Specifically, the phage content for SA/HA/ES-1, SA/HA/ES-2, SA/HA/ES-3, SA/HA/ES-4, SA/HA/ES-5, and SA/HA/ES-6 was 2.8×104, 1.0×105, 4.8×105, 7.9×105, 1.0×106, and 1.3×106 PFU per microsphere, respectively. This controllability in phage load is crucial for determining the appropriate dosage for practical applications.
Protection and release performance of HMs-Phages
Although there are no clear standards specifying the exact phage dosages for practical applications, it is generally accepted that delivering phages in sufficient quantities increases the probability of phages capturing host bacteria, thereby improving therapeutic outcomes30. Thus, it is essential to protect the phage activity from adverse conditions and ensure targeted delivery (Fig. 4a). Free phages (unencapsulated) were completely inactivated after 5 min of incubation in simulated gastric fluid (SGF) at pH 2.5 (Supplementary Fig. 12a). The microsphere composition significantly affects their protective efficacy for encapsulated cargos. To investigate this, phage-loaded microspheres with varying compositions were fabricated using a standardized precursor concentration (1% w/v), and their protective ability against internal phages in simulated gastric fluid was evaluated. Quantitative analysis revealed that after 2 h of incubation, encapsulated phages in pure SA-1 microspheres were completely inactivated (Supplementary Fig. 13). In contrast, SA/HA-1 microspheres retained approximately 2 × 103 PFU/g of viable phages, and SA/ES-1 microspheres preserved 4 × 104 PFU/g. Notably, SA/HA/ES-1 microspheres exhibited the highest phage survival, retaining around 1 × 106 PFU/g, indicating the most effective protective effect on phages. The preserved high phage viability directly correlated with antibacterial efficacy at target sites. Specifically, encapsulated phages in SA/HA/ES-1 microspheres demonstrated superior antimicrobial performance in simulated intestinal fluid (SIF) following SGF pretreatment (Supplementary Fig. 14). Based on these findings, the SA/HA/ES was identified as optimal formulation for further experiments.
a Schematic diagram of the biological function of HMs-Phages in the gastrointestinal tract. b Survival of encapsulated phages (LPST94 and LST153) in microspheres after exposure to SGF (pH 2.5) supplemented with pepsin (0.32%) at 37 °C. Data are presented as mean ± standard deviation (n = 3 biological replicates). c Cumulative release of phages from microspheres in SIF (pH 7.2) at 37 °C after pre-incubation in SGF for 2 h. Data are presented as mean ± standard deviation (n = 3 biological replicates). Phages were quantified by plating 100 µL of diluted solution with host bacteria on double-layer agar plates and incubating at 37 °C for 12 h. d Fluorescence intensity of simulated digestive fluid after incubation with SA/HA/ES-3 microspheres loaded with rhodamine B-labeled phages. Data are presented as mean ± standard deviation (n = 3 biological replicates). e Microscopy of SA/HA/ES-3 hydrogel microspheres loaded with rhodamine B-labeled phages after treatment with SGF and SIF (n = 3 independent samples with similar results). Scale bar, 500 μm. SA sodium alginate. HA hyaluronic acid. ES Eudragit S100. SGF simulated gastric fluid. SIF simulated intestinal fluid. PFU plaque-forming units of phage. Source data are provided as a Source Data file.
To further evaluate the impact of precursor solution concentration on microsphere performance under gastric conditions, SA/HA/ES microspheres prepared at varying solution concentrations were systematically evaluated. As shown in Fig. 4b, after 2 h of incubation in SGF, SA/HA/ES-3, SA/HA/ES-4, SA/HA/ES-5, and SA/HA/ES-6 microspheres exhibited higher phage survival rates (negligible loss) than SA/HA/ES-1 (2-log loss) and SA/HA/ES-2 (1-log loss), demonstrating their good protective capability against the harsh gastric environment. In SGF with a lower pH of 1.2, the survival rate of encapsulated phages decreased over time (Supplementary Fig. 15). Microspheres formed from precursor solutions with higher concentrations exhibited enhanced protective effects, which is consistent with the trends observed at pH 2.5. In addition, the acid resistance of non-electrosprayed SA/HA/ES microspheres was also performed. A similar concentration-dependent protective effect was observed (Supplementary Fig. 16), consistent with that of the electrosprayed microspheres. This enhanced protection may be related to the denser gel network formed by the solution with increased polymer concentration, which provides the interior phages with an enhanced physical barrier, preventing or reducing their exposure to acidic media.
To evaluate phage release from the hydrogel microspheres, the viability of free phages in SIF containing trypsin and bile salt was assessed. Free phages remained stable in SIF for up to 12 h without significant changes in the titer (Supplementary Fig. 12b). As illustrated in Fig. 4c, 60, 44, 38, 33, 27, and 24% of the phages were released from microspheres in different groups (SA/HA/ES-1 to SA/HA/ES-6) after 1 h of incubation in SIF, followed by sustained release over 6–10 h, contingent on the microsphere formulation. Most phages were released within the first 6 h, and a faster release behavior was observed in microspheres formulated with a low solution concentration. This behavior was likely due to the difference in the internal gel-network structure of the hydrogel microspheres.
To visualize the release profile and swelling behavior of microspheres in simulated gastrointestinal environments, rhodamine B-labeled phages were encapsulated in SA/HA/ES-3 microspheres with uniform size distribution and tracked using fluorescence imaging. Fluorescence images showed that rhodamine B-labeled phages were effectively released from microspheres in SIF, confirming successful intestine-targeted delivery (Fig. 4d). Also, the physical integrity of the SA/HA/ES-3 microspheres remained intact in stomach conditions (Fig. 4e). Upon transfer to SIF, the microspheres absorbed water and swelled rapidly within the first 4 h, and then gradually disintegrated, suggesting a pH-responsive release behavior. Phage release from the hydrogel network was primarily governed by a swelling-dissolution process. Under neutral or alkaline SIF conditions, the gradual dissolution of ES and HA, along with Ca2+ dissociation from alginate, led to the swelling and eventual disintegration of the microspheres (Supplementary Fig. 17). This highlights the flexibility of controlling the phage release by adjusting the microsphere size and precursor solution concentration.
Biosafety evaluation of HMs-Phages
Many traditional antimicrobial materials exhibit significant cytotoxicity or systemic toxicity, which seriously hinders their practical application31,32. To evaluate the clinical potential of HMs-Phages, in vitro biocompatibility experiments (MTT assay and SYTO9/PI staining) were conducted. The SA/HA/ES-1 hydrogel microspheres were selected based on their optimized size distribution (133 ± 19 μm), enabling compatibility with in vivo delivery protocols. Two types of intestinal epithelial cells (Caco-2 and HT-29) were incubated with varying concentrations of HMs-Phages (Fig. 5a). The MTT assay revealed no significant toxicity of HMs-Phages toward either Caco-2 or HT-29 cells, with over 90% cell viability maintained across all tested concentrations (Fig. 5b). Consistently, the cell staining results demonstrated no significant reduction in cell viability at any microsphere concentration compared with those of the control group, further confirming excellent cytocompatibility (Fig. 5c). The low cytotoxicity of HMs-Phages can be primarily attributed to the safety features of SA, HA, and ES, which serve as microsphere components.
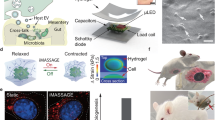
a Schematic diagram of the co-culture system of microspheres and epithelial cells for biosafety assessment. b Cytotoxicity of HMs-Phages (SA/HA/ES-1) in Caco-2 and HT-29 cells after 24 h of co-incubation. Data are presented as mean ± standard deviation (n = 5 biological replicates). c Live/Dead staining of Caco-2 and HT-29 cells after co-culture with different concentrations of HMs-Phages (SA/HA/ES-1) for 24 h (n = 3 biological replicates with similar results). Live cells were stained green, and dead cells were stained red. Scale bar, 100 μm. d Representative histological images of major organs (heart, liver, spleen, lung, and kidney) stained with hematoxylin-eosin (H&E) in the Health and HMs-Phages groups (n = 3 biological replicates with similar results). Scale bar, 100 μm. e Retention of rhodamine B-labeled free phages and HMs-Phages (SA/HA/ES-1) in the murine gastrointestinal tract at different time points (n = 3 biological replicates per time point with similar results). HMs-Phages, hydrogel microspheres with phages. Source data are provided as a Source Data file.
To further assess the biosafety of SA/HA/ES-1 hydrogel microspheres, in vivo toxicity assay was conducted. As shown in Supplementary Fig. 18, no significant differences in body weight were observed between the healthy control and oral microsphere-treated groups over the 7-day administration period. In addition, histological analysis revealed no evident signs of inflammation in the examined tissues after 7 days of microsphere administration compared with that in the healthy control group (Fig. 5d). These results demonstrated that HMs-Phages possessed excellent biocompatibility and metabolic activity, highlighting their potential as promising hydrogel-based carriers for oral phage delivery.
In vivo intestinal retention of HMs-Phages
The gastrointestinal retention of HMs-Phages was assessed in mice using a fluorescence imaging system. After the oral administration of SA/HA/ES-1 hydrogel microspheres containing rhodamine B-labeled phages, fluorescence signals were observed to track intestinal transport and retention time. As shown in Fig. 5e, mice administered with free phages displayed high fluorescence intensity in the jejunum at 1 h and in the ileum at 4 h, with fluorescence completely vanishing at 8 h because of metabolism and intestinal excretion. In contrast, HMs-Phages showed similar fluorescence signals in the stomach and small intestine at 1 h and 4 h, respectively. However, fluorescence signals in the small intestine and colon persisted for longer durations, remaining detectable at 8 and 12 h before disappearing by 24 h, indicating prolonged retention of the hydrogel microspheres in the gut. These results indicate that the SA/HA/ES hydrogel microspheres possess strong intestinal retention ability, making them ideal carriers for targeted intestinal delivery.
HMs-Phages efficiently treat Salmonella-induced colitis
A murine model of STm-induced colitis was established to investigate the therapeutic efficacy of HMs-Phages (Fig. 6a). Intestinal colonization by STm typically causes severe local or systemic infections and inflammatory responses, leading to a constant decline in body weight and food intake in mice (Fig. 6b, c). The free phage treatment group (Phages) failed to alleviate weight loss or improve food intake. In contrast, treatment with SA/HA/ES-1 microspheres with phages (HMs-Phages group) exhibited excellent therapeutic efficacy in increasing body weight and food intake, comparable to ciprofloxacin administration (CIP group). Diarrhea is a hallmark of bacterial infection and enteric inflammation. The feces of the uninfected healthy group remained firm, well-formed, and dark-brown throughout the experiment, with a fecal score of 5 points (Fig. 6d, e). In contrast, after STm infection (PBS group), the feces changed from normal to watery and the color shifted from dark-brown to yellowish-brown on day 5 (fecal score of 1 point), indicating severe diarrhea. Fecal samples from the Phages group showed marginal improvements in hardness, shape, and color on day 3 compared to the PBS group. After day 5, however, these improvements deteriorated progressively, ultimately reaching parity with the PBS group. Both the HMs-Phages and CIP interventions effectively alleviated diarrhea, with higher fecal scores. Notably, mild diarrhea reoccurred in the CIP group starting from day 5, which is presumed to be associated with antibiotic-induced gut dysbiosis. In addition, dynamic changes in bacterial and phage numbers in the mouse intestine were monitored. The Phages group exhibited minor variations in fecal bacteria numbers during treatment, with phage counts ranging from 104 to 105 PFU/g of feces (Fig. 6f, g). Furthermore, the individual LPST94 number in the murine gut of the Phages group was higher than that of LPST153, possibly due to its stronger acid tolerance than that of LPST153 (Supplementary Fig. 19). The HMs-Phages group maintained consistently high phage titers (Fig. 6g), and exhibited an approximately 2000-fold reduction in bacterial counts compared to the PBS group, leading to 100% survival protection in mice (Fig. 6f and Supplementary Fig. 20). Moreover, the individual numbers of LPST94 and LPST153 in the HMs-Phages group remained at equivalent levels, primarily due to the stable intestinal phage delivery provided by the microsphere carriers (Supplementary Fig. 19). Based on the observed findings, this sustained reduction in fecal STm numbers was principally attributed to sufficient gastric protection of the phages by the microspheres, prolonged phage retention, and targeted phage release, thereby ensuring high phage concentration and long antibacterial time at the site of action.
a Establishment of the bacterial colitis model and therapeutic interventions in mice, including streptomycin pretreatment and Salmonella infection, followed by different treatments [PBS (100 μL/day, 0 PFU/mL), HMs (100 mg/day, 0 PFU/g), Phages (100 μL/day, 109 PFU/mL), HMs-Phages (100 mg/day, 109 PFU/g), and CIP (100 μL/day, 2 mg/mL)]. b Body weight and (c) cumulative food intake in the different groups. Data are presented as mean ± standard deviation (n = 6 mice per group). d Representative fecal images and (e) stool scores of mice in the different groups on days 1, 3, 5, 7 and 9. Data are presented as mean ± standard deviation (n = 6 mice per group). f Dynamic changes in bacterial number and (g) phage titers per gram of feces in the different treatment groups (n = 6 mice per group). The central line of box plots represents the median, box bounds indicate the 25th and 75th percentiles, and whiskers extend to the minimum and maximum values. HMs hydrogel microspheres. HMs-Phages, hydrogel microspheres with phages. CIP, ciprofloxacin. Source data are provided as a Source Data file.
Colon shortening is a distinct feature of STm-induced colitis. Both the HMs-Phages and CIP groups showed a positive effect in the recovery of colon length (Fig. 7a, b). Splenomegaly is another visible characteristic of STm infection due to the activation, proliferation, and infiltration of immune cells in the spleen (Fig. 7c, d). Compared with other groups, the CIP group exhibited the strongest suppression of splenomegaly, followed by the HMs-Phages group, with no statistically significant difference between these two treatments (p > 0.05). Notably, intestinally colonized STm can invade other parenteral organs such as the spleen and liver. As shown in Fig. 7e–g, STm burdens in the spleen, liver, and colon were significantly decreased by 3-log units in the CIP group compared with the PBS group, and the HMs-Phages group exhibited comparable antibacterial efficacy. Regarding in vivo phage resistance, STm in both the Phages and HMs-Phages groups remained sensitive to LPST94 and LPST153 throughout the treatment period, indicating no phage resistance development (Supplementary Fig. 21).
Colon tissues were imaged (a) and measured for length (b), and spleens were imaged (c) and recorded for weight (d) (n = 6 mice per group). Scale bar, 10 mm. Bacterial burden was determined in spleen (e), liver (f), and colon (g) (n = 6 mice per group). h–j Proinflammatory cytokine levels in colon tissues were analyzed (n = 6 mice per group). k Representative histological images of colon tissue from different groups (n = 3 biological replicates with similar results). Scale bar, 200 μm. Statistical significance was determined by one-way ANOVA with Tukey’s post hoc test (panel b, d, h–j) or unpaired two-tailed Student’s t test (panel e-g) (*p < 0.05; **p < 0.01; ***p < 0.001; ns, no significance). Data are presented as mean ± standard deviation. Exact p-values: panel (b): Health vs. PBS, p < 0.0001; PBS vs. Phages, p = 0.1076; PBS vs. HMs-Phages, p < 0.0001; Phages vs. HMs-Phages, p = 0.0070; HMs-Phages vs. CIP, p = 0.9713; panel (d): Health vs. PBS, p < 0.0001; PBS vs. Phages, p = 0.8776; PBS vs. HMs-Phages, p = 0.0015; Phages vs. HMs-Phages, p = 0.0258; HMs-Phages vs. CIP, p = 0.8499; panel (e): PBS vs. Phages, p = 0.4972; Phages vs. HMs-Phages, p = 0.0085; HMs-Phages vs. CIP, p = 0.0464; panel (f): PBS vs. Phages, p = 0.6731; Phages vs. HMs-Phages, p = 0.0144; HMs-Phages vs. CIP, p = 0.0473; panel (g): PBS vs. Phages, p = 0.1759; Phages vs. HMs-Phages, p = 0.0246; HMs-Phages vs. CIP, p = 0.1613; panel (h): Health vs. PBS, p < 0.0001; PBS vs. HMs-Phages, p < 0.0001; HMs-Phages vs. CIP, p = 0.1026; panel (i): Health vs. PBS, p < 0.0001; PBS vs. HMs-Phages, p < 0.0001; HMs-Phages vs. CIP, p = 0.0522; panel (j): Health vs. PBS, p < 0.0001; PBS vs. HMs-Phages, p < 0.0001; HMs-Phages vs. CIP, p = 0.0179. HMs, hydrogel microspheres. HMs-Phages, hydrogel microspheres with phages. CIP, ciprofloxacin. Source data are provided as a Source Data file.
Proinflammatory cytokine levels serve as key indicators of inflammatory response severity. As presented in Fig. 7h–j, STm infection significantly elevated the levels of inflammatory cytokines (TNF-α, IL-6, and IL-1β) compared to the healthy group. Oral CIP administration significantly reduced proinflammatory cytokine levels relative to the PBS group, decreasing TNF-α by 71.7%, IL-6 by 74.1%, and IL-1β by 68.9%. Remarkably, the HMs-Phages group exhibited a comparable anti-inflammatory effect, with reductions of 60.8% for TNF-α, 62.8% for IL-6, and 54.6% for IL-1β. The alleviation of the inflammatory response by HMs-Phages treatment was mainly attributed to targeted phage delivery and effective pathogen clearance. Colonic histological analysis revealed severe tissue injury in the PBS group, which was characterized by crypt damage, reduced goblet cell numbers, and mucosal lesions (Fig. 7k). In contrast, these structural damages were significantly improved in both the HMs-Phages and CIP groups. The histopathology scores of the HMs-Phages treatment group were substantially lower than Phages groups, showing an 81.1% reduction compared with that of the PBS group (Supplementary Fig. 22).
HMs-Phages precisely edit the gut microbiota
Maintaining a balanced gut microbiota is crucial for nutritional metabolism, intestinal barrier function, and immune function. Prior to experimental interventions, the baseline gut microbiota of mice in different groups was analyzed. Taxonomic profiling revealed a similar microbial composition and abundance across all groups (Supplementary Fig. 23). β-diversity analysis further indicated no significant intergroup variation, with no apparent clustering in PCoA space, suggesting a similar baseline gut microbiota structure among experimental cohorts (Supplementary Fig. 24).
Following STm infection and different treatments, significant alterations in microbial communities were observed. Compared to the healthy group, STm infection (PBS group) drastically decreased the observed operational taxonomic units (OTUs), Shannon index, and Simpson index, suggesting a marked reduction in microbial richness and diversity (Fig. 8a–c). Interestingly, the CIP group also exhibited reduced microbial diversity, likely due to its broad-spectrum antibacterial effect. Notably, oral administration of HMs-Phages significantly restored the richness and diversity of the gut microbiota to levels approaching healthy controls. In addition, pathogen colonization significantly disrupted the gut microbiota structure, causing severe dysbiosis (Fig. 8d, e). At the family level, STm colonization significantly reduced the relative abundance of beneficial taxa such as Bacteroidaceae, Lachnospiraceae, and Prevotellaceae relative to healthy controls (Supplementary Fig. 25). These taxa are known to play key roles in dietary fiber degradation and short-chain fatty acid production33. Following treatment, HMs-Phages substantially restored the relative abundance of these beneficial microbes toward levels observed in healthy controls. At the genus level, differential abundance analysis revealed pronounced alterations in the bacterial community of the PBS group, with 23.11% of genera significantly upregulated and 17.47% significantly downregulated relative to healthy group (Supplementary Fig. 26). Compared to the PBS group, the Phages group exhibited a largely similar microbial community structure, with 99.21% of genera showing no significant abundance changes (Fig. 8f). Conversely, the HMs-Phages group exhibited a markedly distinct microbial community compared to that of PBS group, with an upregulation of 17.03% and a downregulation of 23.22% (Fig. 8g). The CIP group exhibited fewer significantly altered bacterial taxa, indicating a microbial composition more similar to that of the PBS group (Fig. 8h). Regarding STm abundance, HMs-Phages treatment led to a significant reduction compared to the other groups, achieving an efficacy comparable to that of CIP (Fig. 9a).
a Observed operational taxonomic units (OTUs), (b) Shannon diversity index, and (c) Simpson diversity index of the gut microbiota at the genus level. Data are presented as mean ± standard deviation (n = 6 mice per group). Statistical significance was determined by one-way ANOVA followed by Tukey’s post hoc test (*p < 0.05; ***p < 0.001; ns, no significance). Exact p-values: panel (a): Health vs. HMs-Phages, p = 0.9713; PBS vs. Phages, p > 0.9999; Phages vs. HMs-Phages, p < 0.0001; HMs-Phages vs. CIP, p < 0.0001; panel (b): Health vs. HMs-Phages, p = 0.9901; PBS vs. Phages, p = 0.5805; Phages vs. HMs-Phages, p < 0.0001; HMs-Phages vs. CIP, p < 0.0001; panel (c): Health vs. HMs-Phages, p = 0.9977; PBS vs. Phages, p = 0.9986; Phages vs. HMs-Phages, p < 0.0001; HMs-Phages vs. CIP, p = 0.0118. (d) Taxonomic profiling of gut microbiota at the genus level across treatment groups (n = 6 mice per group). e Principal coordinates analysis (PCoA) of microbial β-diversity based on Bray-Curtis distance (n = 6 mice per group). Volcano plot of differential bacterial abundance at the genus level between (f) PBS and Phages groups, (g) PBS and HMs-Phages groups, and (h) PBS and CIP groups (n = 6 mice per group). Dashed lines indicate thresholds (|log2(FC)| ≥ 2 and −log10(FDR) ≥ 1.301). Up, upregulated taxa (red points). Down, downregulated taxa (green points). NS non-significant taxa (gray points). FDR false discovery rate. FC fold change. HMs hydrogel microspheres. HMs-Phages, hydrogel microspheres with phages. CIP ciprofloxacin. Source data are provided as a Source Data file.
The relative abundance of (a) Salmonella, (b) Faecalibacterium, and (c) Ruminococcus. Data are presented as mean ± standard deviation (n = 6 mice per group). Statistical significance was determined by one-way ANOVA followed by Tukey’s post hoc test (**p < 0.01, ***p < 0.001. ns, no significance). Exact p-values: panel (a): PBS vs. Phages, p = 0.0017; PBS vs. HMs-Phages, p < 0.0001; Phages vs. HMs-Phages, p < 0.0001; HMs-Phages vs. CIP, p > 0.9999; panel (b): Health vs. HMs-Phages, p = 0.8688; PBS vs. Phages, p > 0.9999; Phages vs. HMs-Phages, p < 0.0001; HMs-Phages vs. CIP, p < 0.0001; panel (c): Health vs. HMs-Phages, p = 0.1961; PBS vs. Phages, p > 0.9999; Phages vs. HMs-Phages, p < 0.0001; HMs-Phages vs. CIP, p = 0.0033. d Schematic illustration of HMs-Phages regulating gut flora. Salmonella phages were delivered to the intestine via hydrogel microspheres to precisely lyse pathogens. The prebiotic ingredients (SA and HA) of the hydrogel microspheres served as nutrient substances, effectively increasing the relative abundance of beneficial bacteria and related metabolites, thereby synergistically restoring gut flora balance. HMs hydrogel microspheres. HMs-Phages, hydrogel microspheres with phages. CIP ciprofloxacin. Source data are provided as a Source Data file.
As key beneficial bacteria in the gut, Faecalibacterium and Ruminococcus play crucial roles in maintaining microbiome homeostasis and gut health by producing beneficial metabolites (e.g., butyrate) and reinforcing the intestinal mucosal barrier34. Compared to the healthy control, the abundance of Faecalibacterium and Ruminococcus was significantly reduced in the PBS group, whereas treatment with HMs-Phages restored both to levels similar to the healthy control (Fig. 9b, c). Notably, the relative abundance of these beneficial bacteria was significantly reduced following CIP treatment, likely due to its non-specific antibacterial effect. Overall, the HMs-Phages treatment precisely lyses targeted pathogenic bacteria while promoting the recovery of beneficial microbes, thereby regulating the microbiota composition toward a healthy state (Fig. 9d). This innovative delivery system enables fine-tuned manipulation of the gut microbiota in situ and enhances the therapeutic efficacy for bacterial colitis.
Discussion
Editing the gut microbiota aims to regulate the composition of the microbial community by selectively targeting specific microbial entities within the gut. In the gut microbiota, low-abundance organisms have a high possibility of extinction35. Effective gut microbiota editing requires an effective reduction in the abundance of target entities to the point of extinction. Utilizing the specific interaction between phages and bacteria, oral phage administration enables the precise suppression or targeted modification of microbial entities in situ11. Key determinants of efficiency include the working titer and duration of phages at target sites, along with their biological characteristics14,15.
In this study, we developed a phage-based strategy for the precise modulation of a single gut bacterium by combining phages with biomaterial carriers. First, we screened for a phage combination (cocktail) that exhibited potent antibacterial activity in vitro (Fig. 2f). Next, we demonstrated that the designed SA/HA/ES hydrogel microspheres effectively protected the encapsulated phages in the stomach and facilitated their responsive release into the intestine (Fig. 4b–e). The combination of these microspheres with phages enabled a targeted and sustained approach for manipulating specific bacteria in the murine gut (Fig. 8). This strategy successfully restricted specific entities to a low level for a sufficiently long period, creating a critical spatiotemporal window for the gut flora to re-establish balance (Fig. 9d). This microbiota editing strategy is a powerful tool for exploring the causal relationships between gut bacteria and host health.
Although phage-resistant STm emerged after 48 h in vitro (Fig. 2f), the pathogens in the mice remained sensitive to delivered phages throughout the experimental period (Supplementary Fig. 21). This phenomenon may be attributed to the heterogeneity of the intestinal environment. The uneven distribution of phages creates bacterial niches, which reduce the overall efficacy of phage therapy and contribute to the long-term coexistence of phages with phage-sensitive bacteria36. Nonetheless, further optimization and screening of phage combinations with prolonged antibacterial activity and a broader lytic range against Salmonella strains are required. In addition, further exploration and modeling of phage-bacteria interactions, coupled with targeted experimental validation, are pivotal for successful gut microbiota editing and enhanced therapeutic outcomes.
Our work showed that targeted phage delivery via microspheres significantly reduced Salmonella abundance in the mammalian gut and alleviated inflammation compared to treatment with free phages (Figs. 6f, 9a). Although antibiotics (CIP) exhibited potent pathogen clearance, its non-selective bactericidal effect significantly depleted beneficial commensal bacteria (e.g., Faecalibacterium, Ruminococcus), inducing pronounced gut microbiota dysbiosis. Our targeted phage delivery strategy circumvented the gut dysbiosis and diarrhea typically associated with antibiotic therapy, enabling precise in situ manipulation of specific bacterial species. Furthermore, beneficial metabolites (e.g., short-chain fatty acids) yielded from microbial metabolism of the microsphere components (SA and HA) served as nutritional substrates for commensal bacteria, thereby promoting their growth and gut homeostasis restoration37,38. Interestingly, only a slight increase in the abundance of Faecalibacterium and Ruminococcus was observed in the HMs group without phage intervention, which is likely attributable to the dominant effect of STm infection.
It is important to note that streptomycin was used to facilitate exogenous strain (STm) colonization and establish the murine colitis model by inducing transient disruption of the gut microbiota. Healthy control mice received neither streptomycin nor STm infection, providing a baseline reference with an undisturbed gut microbial community. Despite initial streptomycin pretreatment and STm challenge, phage-mediated targeted intervention of the HMs-Phages group created a crucial spatiotemporal advantage for the intestinal microbiota to re-establish balance. In future translational studies applying this phage-targeted strategy to other endogenous gut bacteria, antibiotic pretreatment can be omitted, while the screening and delivery of matched phages remain essential.
In summary, our study outlines a generalizable approach to enhance the biological stability of semi-living entities (phages) and ensure their stable intestinal delivery. This approach enables the specific modification of intestinal bacteria through noninvasive and convenient oral delivery, providing a promising platform for exploring the causal relationships between gut microbiota and disease. Furthermore, our electrospray strategy allows for the flexible control of both microsphere size and phage release rates, making it adaptable to the diverse needs of animal models in preclinical trials. These advancements hold significant potential for advancing phage therapy and deepening our understanding of gut-microbiome interactions.
Methods
Animals and ethics statement
Female C57BL/6 J mice (6-7 weeks old) were purchased from Henan Skobes Biotechnology Co., Ltd. All mice were housed in individually ventilated cages under specific pathogen-free conditions. The animal facility was maintained at 23 ± 2 °C and 50-60% relative humidity, with a 12 h light/12 h dark cycle. Mice were fed with a standard sterile diet and water ad libitum. After a one-week acclimation period, the mice were randomly assigned to the designated experimental groups. All animal procedures were conducted in accordance with the guidelines and were approved by the Animal Management and Ethics Committee of Huazhong Agricultural University (permission numbers HZAUMO-2023-0267 and HZAUMO-2025-0167). The SA/HA/ES-1 hydrogel microspheres were prioritized for oral gavage administration due to their optimal size (133 ± 19 μm), enabling smooth passage through a fine feeding needle in murine models while complying with animal welfare standards.
General materials
Sodium alginate (SA, 98%), sodium hyaluronate (HA, Mw = 10 kDa), and ciprofloxacin hydrochloride (CIP) were purchased from Shanghai Yuanye Bio-Technology Co., Ltd. (Shanghai, China). Poly (methyl methacrylate-co-methacrylic acid) (Eudragit® S100, ES) was supplied by Evonik Specialty Chemicals (Shanghai) Co., Ltd. (Shanghai, China). Streptomycin was obtained from Shanghai Aladdin Biochemical Technology Co., Ltd. Calcium chloride, sodium chloride, hydrochloric acid, sodium hydroxide, sodium citrate, sodium bicarbonate, and magnesium sulfate were bought from Sinopharm Chemical Reagent Co., Ltd. 2-Amino-2-(hydroxymethyl)-1,3-propanediol (Tris) was purchased from BioFroxx GmbH. Pepsin (1:3000) and trypsin (1:250) were obtained from Shanghai Macklin Biochemical Co., Ltd. (Shanghai, China). Fetal bovine serum, MEM medium, McCoy’s 5 A medium, penicillin-streptomycin, and phosphate buffer saline (0.01 M, pH 7.4) were bought from Thermo Fisher Scientific Co. (Beijing, China). The colonic epithelial cell lines Caco-2 (ATCC HTB-37, passage 10) and HT-29 (ATCC HTB-38, passage 14) were used for cytotoxicity assays and cultured in MEM and McCoy’s 5 A media, respectively. Gelatin (from porcine skin), thiazolyl blue tetrazolium bromide (MTT), and dimethyl sulfoxide (DMSO) were purchased from Sigma-Aldrich Co. (Missouri, USA).
Bacteria and culture conditions
Salmonella enterica serovar Typhimurium ATCC 13311, UK-1, and SL1344 were used in this study. All strains were obtained from our laboratory collection. ATCC 13311 and UK-1 were used to propagate their corresponding phages. The naturally streptomycin-resistant S. Typhimurium SL1344 (STm) was selected as the model strain for subsequent in vitro antibacterial and in vivo therapeutic experiments because of its ability to yield reliable results in animal models. ATCC 13311 and UK-1 were cultured in Luria-Bertani (LB) broth for 12 h at 37 °C with shaking at 150 rpm, while SL1344 was cultured in LB supplemented with 50 μg/mL streptomycin.
Preparation of purified phage solution
The Salmonella phages LPST83, LPST94, and LPST153 used in this study were previously stored in our laboratory24,25,26. The host bacterium for both Salmonella phage LPST83 and LPST94 was S. Typhimurium UK-1, while the host for LPST153 was S. Typhimurium ATCC13311. Phages were propagated separately by co-culturing them with their respective host bacteria for 12 h at 37 °C. The bacterial debris was removed by centrifugation (8000 × g, 10 min) and filtration (0.22 µm filters). Then, phages were further purified by transferring them from culture media into SM buffer by ultracentrifugation (154,000 × g, 1 h) and ultrafiltration (10 kDa). Final phage solutions were filtered with 0.22 μm filters and stored at 4 °C for further use.
Screening of efficient phage combination
The antimicrobial activity of phages, either individually or in combinations, against Salmonella was evaluated by inhibition curves as described in published study39. Briefly, 100 μL of STm strains (105 CFU/mL) was mixed with an equal volume of phage suspension (103-108 PFU/mL) in a 96-well microtiter plate. Each phage in the combination solution had an equal titer. The control group was treated with isochoric Luria-Bertani broth media instead of a phage suspension. The microtiter plate was incubated at 37 °C and 150 rpm, and the absorbance (600 nm) was measured at defined intervals using a BioTek Cytation5 microplate reader with Cytation5 v3.11 software (BioTek, USA). A phage combination with strong antimicrobial activity was selected for later use. The phage resistance of STm was assessed using the spot test24,36. Briefly, 3 µL of phage solution was separately spotted onto bacterial lawns formed by the phage resistance bacterial strains on Luria-Bertani agar plates. After 12 h of incubation at 37 °C, bacterial lawns with clear plaques were considered as phage sensitive.
Preparation of hydrogel microspheres with phages (HMs-Phages)
The HMs-Phages were prepared by electrohydrodynamic spraying and the calcium ion crosslinking method. The viscosity of SA/HA/ES precursor solutions was measured using a Discovery HR-2 rotational rheometer with TRIOS v5.0 software (TA Instruments, USA), and the conductivity was measured using a DDS-11A digital conductivity meter (INESA Scientific Instrument, China). In brief, hydrogel precursor polymers (SA, HA, and ES in a 1:1:1 mass ratio) were added to sterile deionized water at total polymer concentrations of 1, 2, 3, 4, 5, and 6% (w/v), followed by gentle stirring overnight at room temperature (25 °C). Afterwards, the pH of the polymer solution was adjusted to 7.4 with 0.1 M sodium hydroxide to ensure complete dissolution of ES, followed by sterilization through 0.22 μm filter. Then, purified LPST94 and LPST153 were mixed at a volume ratio of 1:1, with a total titer of 109 or 1010 PFU/mL, and subsequently added into the precursor solutions at a volume ratio of 1:9. A portable electrospinning apparatus/electrostatic sprayer (JUNADA HHE-1, China) was used to atomize and disperse precursor solution with phages into micro-droplets that were vertically deposited into a calcium chloride solution (0.2 M) and crosslinked for 1 h to obtain HMs-Phages, which were denoted as SA/HA/ES-1, SA/HA/ES-2, SA/HA/ES-3, SA/HA/ES-4, SA/HA/ES-5, and SA/HA/ES-6, respectively. The supplied voltage was 10 kV and the tip-collector distance was 10 cm. After ionic gelation, the prepared microspheres were washed three times with SM buffer containing 0.2 M calcium chloride to remove unloaded phages. Then, the final microspheres were resuspended in the same buffer and stored at 4 °C for further use. Similarly, other microsphere formulations (comprising SA, SA/HA, and SA/ES) were prepared at polymer concentrations of 1% or 3% (w/v) following the aforementioned procedure. In addition, the corresponding non-electrosprayed SA/HA/ES microspheres were prepared using a conventional extrusion-dripping method. Unlike the aforementioned electrospraying method, the hydrogel precursor solution was directly extruded and dropped into a calcium chloride solution (0.2 M) for crosslinking and solidification.
Physicochemical characterization of HMs-Phages
The macroscopic morphology of HMs-Phages was evaluated by an optical microscope (Nexcope, China), and the microsphere sizes were measured by ImageJ v1.52 software (National Institutes of Health, USA) from the digital images. Meanwhile, the surface micromorphology of HMs-Phages was observed by using a scanning electron microscope (SEM, Hitachi SU8000, Japan) with an accelerating voltage of 10 kV after lyophilization and gold spraying treatment. The element distribution of hydrogel microspheres (SA/HA/ES-3 as a representative sample) was analyzed using SEM coupled with energy dispersive spectroscopy. Fourier transform infrared spectrum of polymer powder and lyophilized HMs-Phages (SA/HA/ES-3) were recorded using a Nicolet iS50 spectrometer with OMNIC Spectra v2.1 software (Thermo Fisher Scientific, USA) with a resolution of 4 cm−1 over the wavenumber range of 400 − 4000 cm−1. The cross-sectional structure of HMs-Phages (with SA/HA/ES-3 as a representative) was examined using transmission electron microscopy (TEM, Hitachi H-7650, Japan) to confirm the uniform dispersion of polymers within the hydrogel network. The distribution of polymer components within the microspheres was further verified by fluorescence labeling. HA and ES were fluorescently labeled via amide coupling reactions with Cy3-NHS (red fluorescence) and Alexa Fluor 488-NHS (green fluorescence), respectively. Briefly, 200 mg of either HA or ES was dissolved in 5 mL of deionized water, followed by the addition of 3 mL of 1-ethyl-3-(3-dimethylaminopropyl) carbodiimide solution (100 mg/mL). After stirring for 15 min, 100 μL of fluorescent dye (1 mg/mL) was added, and the reaction was carried out at room temperature for 12 h with constant stirring. The resulting mixture was then dialyzed against 1000 mL of deionized water for 48 h using a dialysis membrane with a molecular weight cut-off of 7000 Da. Finally, the dialyzed solution was lyophilized to obtain the fluorescently labeled polymers. The electrosprayed microspheres labeled with fluorescent dyes were prepared at a 3% (w/v) polymer concentration using the aforementioned electrospray method. Fluorescence microscopy imaging was then performed using a Leica DMi8 fluorescence microscope with LAS X v3.0 software (Leica Microsystems, Germany) across different channels.
Encapsulation efficiencies of HMs-Phages
The encapsulation efficiencies were evaluated by dissolving microspheres into a microsphere-broken solution (pH 7.5) consisting of 50 mM sodium citrate, 0.2 M sodium bicarbonate and 50 mM Tris, and then measuring phage amounts based on the double-layer agar plate method24. S. Typhimurium ATCC13311 was used as the host bacterium for counting phage titer due to its high efficiency of plating against both phage LPST94 and LPST153 demonstrated in our previous study40. Briefly, 100 μL of host bacteria was mixed with 100 µL of diluted phage suspension, followed by the addition of 3 mL of molten top agar (Luria-Bertani medium with 0.5% agar). Then, the mixture was immediately poured onto pre-solidified Luria-Bertani agar plates to harden. The plates were incubated overnight at 37 °C, and the phage numbers was calculated based on the plaque formed. Encapsulation efficiency of phages in microspheres was determined as the ratio of phage released from the microspheres to the initial phage used for microsphere preparation, expressed as a percentage.
Acid resistance and release properties of HMs-Phages
The acid stability of encapsulated phages under gastric conditions was assessed by coincubation with SGF (pH 2.5 or 1.2), which was composed of 34.22 mM sodium chloride, 3.2 mg/mL pepsin and hydrochloric acid. Specifically, 100 mg of HMs-Phages was added to 1 mL of SGF and incubated at 37 °C. At certain intervals, microspheres were dissolved in a microsphere-broken solution, and the phage numbers were determined by the double-layer agar plate method mentioned above. In addition, the in vitro antibacterial efficacy of phage-loaded microspheres with varied formulations (SA, SA/HA, SA/ES, SA/HA/ES) was evaluated. Briefly, 100 mg of microspheres (prepared from 1% w/v polymer solutions) were pre-incubated in SGF (pH 2.5) for 2 h. Afterwards, the microspheres were mixed with 1 mL of STm suspension in SIF (10⁵ CFU/mL). SIF was comprised of 6.8 mg/mL KH2PO4, 10 mg/mL trypsin, 5 mg/mL bile salt, and sodium hydroxide. Following incubation, bacterial counts were quantified at specified timepoints (0, 2, 4, 8, and 12 h) to assess antimicrobial activity.
The responsive release of encapsulated phages from the microspheres was quantified using double-layer agar plate counting. Cumulative release was determined as the percentage of phages released at each time point relative to the total phage encapsulated in the microspheres. In addition, phage release was also assessed by fluorescent labeling and measurement of the fluorescence intensity in the supernatant. The rhodamine B-labeled phages were prepared by incubating phage solution with dye (40 μM) for 12 h, followed by ultrafiltration (10 kDa) to remove the excess dye41. SA/HA/ES-3 microspheres were selected as a representative due to their regular morphology and uniform size distribution. Specifically, 100 mg of HMs-Phages was added to 1 mL of SGF or SIF and incubated at 37 °C with shaking at 150 rpm. At certain intervals, the fluorescence intensity of the supernatant was measured using a microplate reader (BioTek Cytation5, USA). The excitation wavelength was set to 525 nm, while the emission was examined at 580 nm. Meanwhile, the morphology of microspheres was recorded by a Leica DMi8 fluorescence microscope with LAS X v3.0 software (Leica Microsystems, Germany). The swelling degree of microspheres in simulated digestive fluid was determined as the ratio of the swollen mass to the initial mass, expressed as a percentage.
Biosafety evaluation of HMs-Phages
The biocompatibility of HMs-Phages was evaluated by cytotoxicity experiments and in vivo toxicity studies. For the cytotoxicity test, SYTO9/PI cell staining and MTT assay were used to assess the biocompatibility of HMs-Phages, following protocols from previous reports42,43. Caco-2 and HT-29 cells were seeded into a 96-well plate (1 × 104 cells/well) and cultured for 24 h. Then, varying masses of microspheres were added into the wells and co-incubated with the cells for another 24 h. After removing the culture medium and microspheres, the cells were incubated with SYTO9/PI staining buffer for 15 min and then imaged by a fluorescence microscope (Nikon, Japan). As for the MTT assay, cells were also incubated with fresh cell medium containing MTT (500 μg/mL). After 4 h of culture, the supernatant was replaced with DMSO to dissolve the formazan crystal. The ratio of absorbance values at 570 nm between the HMs-Phages group and the control group was expressed as relative cell viability. For the in vivo toxicity study, C57BL/6 J mice (6-7 weeks old) were orally administered with 100 mg HMs-Phages (SA/HA/ES-1) daily for 7 days, and their body weights (n = 4 mice) were monitored as an indicator of overall health. Mice treated with PBS were used as the control group (n = 4). The major organs (heart, liver, spleen, lung, and kidney) were collected and stained with hematoxylin and eosin (H&E) for pathological analysis.
Intestinal retention property of HMs-Phages
C57BL/6 J mice (aged 6-7 weeks) were randomly allocated into two groups (15 mice per group) and orally administered with either 100 μL of rhodamine B-labeled phage solution (108 PFU) or 100 mg of SA/HA/ES-1 microspheres loaded with rhodamine B-labeled phages (108 PFU) using 21-gauge feeding needles. After 1, 4, 8, 12, and 24 h, the mice were euthanized using isoflurane anesthesia followed by cervical dislocation. The gastrointestinal tracts were excised for fluorescence imaging on the IVIS Spectrum using Living Image v4.4 software (PerkinElmer, USA) to assess the in vivo retention of HMs-Phages.
Animal models of Salmonella-induced colitis
C57BL/6 J mice (6-7 weeks old) were used to establish a model of Salmonella-induced colitis as previously described44. The healthy control, HMs-Phages, and CIP groups comprised 10 mice each, while the PBS, HMs, and Phages groups contained 20 mice per group. Briefly, mice were orally administered with 100 μL of streptomycin solution (200 mg/mL) after fasting for water and food for 4 h. Subsequently, mice were provided with water and food ad libitum. After streptomycin treatment for 20 h, the mice were fasted for 4 h and then orally infected with 100 μL of STm suspension (108 CFU). The healthy group received sterile water in place of streptomycin solution, and phosphate buffer instead of bacteria suspension. One day later, the infected mice received PBS buffer (PBS, 100 μL, 0 PFU), pure hydrogel microspheres (HMs, 100 mg, 0 PFU), free phages (Phages, 100 μL, 108 PFU), SA/HA/ES-1 hydrogel microspheres with phages (HMs-Phages, 100 mg, 108 PFU), and ciprofloxacin (CIP, 100 μL, 2 mg/mL) once every day by oral gavage using 21-gauge feeding needles for 8 days. Meanwhile, the body weights and food intakes of mice were recorded daily. Animals meeting predefined humane endpoints (> 25% weight loss from baseline, persistent severe diarrhea, or inability to self-feed) were humanely euthanized via terminal isoflurane anesthesia followed by cervical dislocation.
Assessment of bacterial diarrhea
Mouse feces were collected on 1, 3, 5, 7 and 9 days after different treatments. Fecal scores were then assessed by adopting a blinded method. The fecal samples were independently graded as follows: healthy feces (dark brown, spindle-shape, and hard texture, 5 points), sub-healthy feces (light brown, spindle-shape, and slight hard texture, 4 points), mild diarrheal feces (light brown, irregular-shape, and soft texture, 3 points), diarrheal feces (light brown, irregular-shape, and semi-solid texture, 2 points), severe diarrheal feces (light brown, droplet shape, and liquid state, 1 point).
Histology analysis
On day 9 after STm infection, the mice with different treatments were sacrificed by isoflurane anesthetics and cervical dislocation. The spleen weight and colon length were recorded. In addition, the colon tissues were fixed in 4% paraformaldehyde for 24 h, followed by paraffin embedding and H&E staining for pathological analysis. Histological damage was blindly scored using the previously described criteria45, as shown in Supplementary Table 1. The histological score was calculated as the sum of the inflammation, depth of lesion, and crypt damage scores, multiplied by the lesion area score.
Bacterial burden in tissues
The spleen, liver, and colon of mice in different groups were harvested for bacterial burden analysis. Specifically, portions of the small intestine, colon, and liver were dissected and weighed, while the entire spleen were weighed. These tissues were then homogenized and serially diluted in 1 mL of sterile PBS buffer. Proper dilutions were plated on the selective Luria-Bertani agar plates supplemented with streptomycin (50 μg/mL) to calculate STm number (CFU/g). In addition, the phage resistance of intestinal STm against LPST94 and LPST153 was evaluated using the spot test method. Briefly, 100 μL of fecal STm was mixed with 3 mL of molten top agar (Luria-Bertani medium containing 0.5% agar). The mixture was then dispensed onto solid Luria-Bertani agar plates to solidify. Next, 3 μL of phage solution was spotted onto the bacterial lawns. After incubating the plates at 37 °C for 12 h, bacterial lawns displaying clear plaques were considered phage-sensitive.
Inflammatory cytokine analysis
The colon tissues from mice in each group were dissected and weighed, then homogenized in 1 mL of PBS buffer and centrifuged at 4 °C (12,000 × g, 10 min) to obtain supernatants. Then, the proinflammatory cytokines (TNF-α, IL-1β, and IL-6) in the supernatants were detected by commercial ELISA kits (Jianglai biology, China).
Gut microbiome analysis
Fecal samples were collected from mice on day -1 (pre-intervention baseline) and day 9 (treatment endpoint), immediately stored in liquid nitrogen. Then, they were shipped on dry ice to Beijing Genomics Institute (BGI) for total DNA extraction and metagenomic sequencing. Metagenomic libraries were prepared using the MGIEasy Universal DNA Library Prep Kit according to the manufacturer’s instructions, and subsequently sequenced on the DNBSEQ platform following standard institutional workflows. Specifically, after sample quality control, 500 ng of meta-DNA was fragmented using an ultrasonicator (Covaris, UK), and fragments ranging from 300 bp to 700 bp were selected using magnetic beads. The selected DNA fragments were repaired, then ligated with an indexed adapter. The ligation product was amplified by PCR and circularized to get a single-stranded circular (ssCir) library. The ssCir library was subsequently amplified through rolling circle amplification to obtain a DNA nanoball (DNB). The DNB was loaded onto the DNBSEQ platform for sequencing. The microbial diversity analysis was conducted as described46. Briefly, FastQC and fastp were employed for quality control and filtering low-quality reads47. Host-derived DNA sequences from the mouse genome were removed by aligning reads to the Mus musculus reference genome (GRCm39) using Bowtie2 prior to taxonomic classification48. The remaining reads were assigned using Kraken2 in paired-end pattern, with the parameters --gzip-compressed and --use-names49. Subsequently, Bracken was employed to re-estimate read abundances from the Kraken2 output using default parameters50. Alpha diversity (observed microbial richness, Shannon, and Simpson indices) and beta diversity (Bray–Curtis dissimilarity) were calculated with the vegan package in R v4.5.0 based on observed operational taxonomic units. Differential microbial abundance analysis was performed using the DESeq2 package in MicroWorldOmics v1.3 software.
Statistical analysis and reproducibility
All the statistical analyses were performed by GraphPad Prism v8.0.2. One-way ANOVA followed by Tukey’s post hoc test and two-tailed Student’s t test were applied as appropriate. Quantitative data were presented as mean ± standard deviation. Survival data were evaluated using the Log-rank (Mantel-Cox) test. Differences between experimental groups were considered statistically significant at *p < 0.05, **p < 0.01, ***p < 0.001. For differential abundance analysis, false discovery rate (FDR) values were adjusted using the Benjamini–Hochberg correction, with significance defined as |log2(FC)| ≥ 2 and FDR < 0.05. No statistical method was used to predetermine sample size. Sample sizes or biological replicates were presented in the figure legends, and no data were excluded from the analyses. For the result shown by representative images, reproducibility was confirmed by at least three independent experiments. Animals were randomly assigned to experimental groups, and the investigators were blinded to allocation during outcome assessment.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All metagenomic sequencing data generated in this study have been deposited in the Genome Sequence Archive repository (China National Center for Bioinformation) under accession code CRA030008. All other data supporting the findings of this study are available in the article and its Supplementary files. Source data are provided in this paper.
References
Ni, J., Wu, G. D., Albenberg, L. & Tomov, V. T. Gut microbiota and IBD: causation or correlation? Nat. Rev. Gastroenterol. Hepatol. 14, 573–584 (2017).
Bishehsari, F., Voigt, R. M. & Keshavarzian, A. Circadian rhythms and the gut microbiota: from the metabolic syndrome to cancer. Nat. Rev. Endocrinol. 16, 731–739 (2020).
Bajaj, J. S. Alcohol, liver disease and the gut microbiota. Nat. Rev. Gastroenterol. Hepatol. 16, 235–246 (2019).
Park, E. M. et al. Targeting the gut and tumor microbiota in cancer. Nat. Med. 28, 690–703 (2022).
Fishbein, S. R. S., Mahmud, B. & Dantas, G. Antibiotic perturbations to the gut microbiome. Nat. Rev. Microbiol. 21, 772–788 (2023).
Zhang, Y. et al. Discovery of antimicrobial lysins from the “dark matter” of uncharacterized phages using artificial intelligence. Adv. Sci. 20, e2404049 (2024).
Caruso, R., Lo, B. C. & Núñez, G. Host-microbiota interactions in inflammatory bowel disease. Nat. Rev. Immunol. 20, 411–426 (2020).
He, R. et al. The interplay of gut microbiota between donors and recipients determines the efficacy of fecal microbiota transplantation. Gut Microbes 14, 2100197 (2022).
Llewellyn, S. R. et al. Interactions between diet and the intestinal microbiota alter intestinal permeability and colitis severity in mice. Gastroenterology 154, 1037–1046 (2018).
Chevallereau, A., Pons, B. J., van Houte, S. & Westra, E. R. Interactions between bacterial and phage communities in natural environments. Nat. Rev. Microbiol. 20, 49–62 (2022).
Rasmussen, T. S. et al. Bacteriophage-mediated manipulation of the gut microbiome-promises and presents limitations. FEMS Microbiol. Rev. 44, 507–521 (2020).
Melo, L. D. R., Oliveira, H., Pires, D. P., Dabrowska, K. & Azeredo, J. Phage therapy efficacy: a review of the last 10 years of preclinical studies. Crit. Rev. Microbiol. 46, 78–99 (2020).
Duan, Y., Young, R. & Schnabl, B. Bacteriophages and their potential for treatment of gastrointestinal diseases. Nat. Rev. Gastroenterol. Hepatol. 19, 135–144 (2022).
Pirnay, J. P., Ferry, T. & Resch, G. Recent progress toward the implementation of phage therapy in Western medicine. FEMS Microbiol. Rev. 46, fuab040 (2022).
Fang, Q. et al. Safety and efficacy of phage application in bacterial decolonisation: a systematic review. Lancet Microbe 5, e489–e499 (2024).
Wu, J. et al. Bacteriophage defends murine gut from Escherichia coli invasion via mucosal adherence. Nat. Commun. 15, 4764 (2024).
Jamalludeen, N., Johnson, R. P., Shewen, P. E. & Gyles, C. L. Evaluation of bacteriophages for prevention and treatment of diarrhea due to experimental enterotoxigenic Escherichia coli O149 infection of pigs. Vet. Microbiol. 136, 135–141 (2009).
Yang, Y. F. et al. Encapsulation and delivery of phage as a novel method for gut flora manipulation in situ: a review. J. Control. Release 353, 634–649 (2023).
Moghtader, F., Eğri, S. & Piskin, E. Phages in modified alginate beads. Artif. Cells Nanomed. Biotechnol. 45, 357–363 (2017).
Zhou, Y. et al. Encapsulation of Salmonella phage SL01 in alginate/carrageenan microcapsules as a delivery system and its application in vitro. Front. Microbiol. 13, 906103 (2022).
Malik, D. J. et al. Formulation, stabilisation and encapsulation of bacteriophage for phage therapy. Adv. Colloid Interface Sci. 249, 100–133 (2017).
Hsu, B. B. et al. In situ reprogramming of gut bacteria by oral delivery. Nat. Commun. 11, 5030 (2020).
Moreira, A. et al. Protein encapsulation by electrospinning and electrospraying. J. Control. Release 329, 1172–1197 (2021).
Guo, Y. et al. Application of a novel phage vB_SalS-LPSTLL for the biological control of Salmonella in foods. Food Res. Int. 147, 110492 (2021).
Islam, M. S. et al. Application of a broad range lytic phage LPST94 for biological control of Salmonella in foods. Microorganisms 8, 247 (2020).
Islam, M. S. et al. Characterization of Salmonella phage LPST153 that effectively targets most prevalent Salmonella serovars. Microorganisms 8, 1089 (2020).
Shkoporov, A. N. et al. Viral biogeography of the mammalian gut and parenchymal organs. Nat. Microbiol. 7, 1301–1311 (2022).
Strathdee, S. A., Hatfull, G. F., Mutalik, V. K. & Schooley, R. T. Phage therapy: from biological mechanisms to future directions. Cell 186, 17–31 (2023).
Jończyk-Matysiak, E. et al. Factors determining phage stability/activity: challenges in practical phage application. Expert Rev. Anti. Infect. Ther. 17, 583–606 (2019).
Pires, D. P., Costa, A. R., Pinto, G., Meneses, L. & Azeredo, J. Current challenges and future opportunities of phage therapy. FEMS Microbiol. Rev. 44, 684–700 (2020).
Makvandi, P. et al. Metal-based nanomaterials in biomedical applications: antimicrobial activity and cytotoxicity aspects. Adv. Funct. Mater. 30, 1910021 (2020).
Kim, H. S. et al. Advanced drug delivery systems and artificial skin grafts for skin wound healing. Adv. Drug Deliv. Rev. 146, 209–239 (2019).
Xu, Y. Q., Zhu, Y., Li, X. T. & Sun, B. G. Dynamic balancing of intestinal short-chain fatty acids: the crucial role of bacterial metabolism. Trends Food Sci. Tech. 100, 118–130 (2020).
Ren, Z. et al. Gut microbiome analysis as a tool towards targeted non-invasive biomarkers for early hepatocellular carcinoma. Gut 68, 1014–1023 (2019).
Costello, E. K., Stagaman, K., Dethlefsen, L., Bohannan, B. J. & Relman, D. A. The application of ecological theory toward an understanding of the human microbiome. Science 336, 1255–1262 (2012).
Lourenço, M. et al. The spatial heterogeneity of the gut limits predation and fosters coexistence of bacteria and bacteriophages. Cell Host Microbe 28, 390–401 (2020).
Qiang, T., Wang, J., Jiang, L. & Xiong, K. Modulation of hyperglycemia by sodium alginate is associated with changes of serum metabolite and gut microbiota in mice. Carbohydr. Polym. 291, 119359 (2022).
Zhang, C. et al. Hyaluronic acid modulates gut microbiota and metabolites relieving inflammation: a molecular weight-dependent study. Sci. Bull. 69, 2683–2687 (2024).
Zhang, Y. et al. Combine thermal processing with polyvalent phage LPEK22 to prevent the Escherichia coli and Salmonella enterica contamination in food. Food Res. Int. 165, 112454 (2023).
Islam, M. S. et al. Application of a phage cocktail for control of Salmonella in foods and reducing biofilms. Viruses 11, 841 (2019).
Meng, L. et al. Nanocapping-enabled charge reversal generates cell-enterable endosomal-escapable bacteriophages for intracellular pathogen inhibition. Sci. Adv. 8, eabq2005 (2022).
Tan, P. et al. Designing self-assembling chimeric peptide nanoparticles with high stability for combating piglet bacterial infections. Adv. Sci. 9, e2105955 (2022).
Wu, H. et al. Hyaluronic-acid-coated chitosan nanoparticles for insulin oral delivery: fabrication, characterization, and hypoglycemic ability. Macromol. Biosci. 22, e2100493 (2022).
Cao, Z., Wang, X., Pang, Y., Cheng, S. & Liu, J. Biointerfacial self-assembly generates lipid membrane coated bacteria for enhanced oral delivery and treatment. Nat. Commun. 10, 5783 (2019).
Peng, P. et al. Gastrointestinal microenvironment responsive nanoencapsulation of probiotics and drugs for synergistic therapy of intestinal diseases. ACS Nano 17, 14718–14730 (2023).
Yang, S. et al. The gut microbiome and antibiotic resistome of chronic diarrhea rhesus macaques (Macaca mulatta) and its similarity to the human gut microbiome. Microbiome 10, 29 (2022).
Chen, S., Zhou, Y., Chen, Y. & Gu, J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890 (2018).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Wood, D. E., Lu, J. & Langmead, B. Improved metagenomic analysis with Kraken 2. Genome Biol. 20, 257 (2019).
Lu, J., Breitwieser, F. P., Thielen, P. & Salzberg, S. L. Bracken: estimating species abundance in metagenomics data. PeerJ Comput. Sci. 3, e104 (2017).
Acknowledgements
This work was supported by the National Key Research and Development Program of China (2023YFD1801000, J.L.; 2025YFC3408400, J.L.), the National Natural Science Foundation of China (32322082, J.L.; 32402264, G.Z.; 22577034, Z.S.), the Fundamental Research Funds for the Central Universities (2662024JC008, J.L.; 2662025SPPY004, J.L.; 2662025SYPY005, R.Y.), the Young Top-notch Talent Cultivation Program of Hubei Province, the Natural Science Foundation of Hubei Province (2023AFA111, J.L.), and funding from State Key Laboratory for Conservation and Utilization of Subtropical Agro-bioresources (SKLCUSA-b202301, Z.S.). We thank the National Key Laboratory of Agricultural Microbiology Core Facility for assistance with the IVIS Spectrum instrument, and we are grateful to Zhe Hu for his help with instrument operation. We also thank Fangmei Zhang for SEM support and Limin He for TEM support. In addition, we thank Dr. Pengfei Zhang (Shenzhen Institute of Advanced Technology, Chinese Academy of Sciences) for assistance with the electrospray apparatus.
Author information
Authors and Affiliations
Contributions
Y.Y. conceived and designed the study, performed the experiments, analyzed the data, prepared the figures, and drafted the manuscript. R.L. contributed to metagenomic data analysis. Q.Z. assisted with animal experiments. Y.G., R.W., H.C., R.Z., and R.Y. participated in manuscript discussions and provided valuable suggestions. K.D., T.K.P., and G.P.S. provided critical feedback and assisted in manuscript revision. G.Z., Y.Z., J.L., and Z.S. conceptualized and supervised the project, provided financial support, and guided experiment design and manuscript preparation.
Corresponding authors
Ethics declarations
Competing interests
A patent application related to the phage-encapsulating hydrogel microsphere platform described in this work has been filed with the China National Intellectual Property Administration by Huazhong Agricultural University, with Y.Y., R.W., R.Y., G.Z., Y.Z., J.L., and Z.S. as inventors (application number CN202410757523.4). The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Dennis Nielsen, who co-reviewed with Torben Rasmussen; Martha Clokie and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Yang, Y., Li, R., Zhong, Q. et al. In situ gut microbiota editing: enhancing therapeutic efficacy for bacterial colitis by compatible oral hydrogel microspheres with phages. Nat Commun 16, 9785 (2025). https://doi.org/10.1038/s41467-025-65498-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-65498-1