Abstract
Lipid droplet (LD) biogenesis occurs in the endoplasmic reticulum (ER), the mechanisms of which is not completely known. Seipin (Fld1 in yeast) is a crucial ER membrane protein that defines LD biogenesis sites. Here, we show that truncated seipin, Fld1-∆LR in yeast, and the human equivalent hSeipin-∆LR, mutants lacking the conserved luminal domain region (LR), functionally complement the LD biogenesis defect of fld1∆ mutants. Fld1-∆LR foci colocalize with factors: Nem1, Ldb16, Pex30 and Yft2, which are important for LD biogenesis and these sites become enriched in diacylglycerol upon stimulation of LD formation. Fld1-∆LR forms a homo-oligomeric complex facilitated by protein-protein interactions. We show that mutating the 31st proline abrogates the functioning of Fld1-∆LR. We demonstrate the critical regulatory role of LR of seipin in partitioning triacylglycerol into LDs. We conclude that LR of seipin is dispensable for establishing functional ER sites to recruit proteins for LD biogenesis.
Similar content being viewed by others
Introduction
Lipid droplets (LDs) store metabolic energy in the form of neutral lipids (NLs; triglycerides (TAG) and sterol esters (SE)), and undergo regression upon environmental cues to release fatty acids for energy production or membrane biosynthesis. Defects in LD homoeostasis manifests in several human pathologies, including lipodystrophy, obesity, type-2 diabetes, fatty liver disease, insulin resistance and cancer (reviewed in refs. 1,2,3).
Recent work have shed light into the mechanisms of LD formation at specialized ER subdomains marked by ER membrane protein seipin (Sei1/Fld1 in yeast) and its associated factors, such as lipid droplet assembly factor 1 (LDAF1) in humans, or Ldb16, and Ldo proteins in yeast4,5,6,7,8,9. However, the mechanisms of how these domains are defined for efficient LD formation is not well understood. Apart from LD biogenesis factors, biophysical properties of the ER, ER membrane curvature, ER membrane tension, locally generated lipids, and lipid-protein interactions modulate droplet biogenesis10,11,12. In the droplet assembly cascade colocalization between seipin and lipin-complex activates discrete sites of LD formation, where seipin plays a crucial role in nucleation of NLs into nascent LDs, and its further growth and maturation by establishing functional ER-LD contact sites4,9. Relocalizing seipin or LDAF1 recruits LD biogenesis machinery to these ER sites6,13. Cells depleted of seipin results in abnormal droplet morphology, including several tiny LDs and/or a few supersized LDs, and accumulation of NLs within the ER membrane9,14,15,16,17. Mutations in BSCL2 encoded human seipin (hereafter called hSeipin) manifests in severe form of lipodystrophy, along with upper, lower, and peripheral motor neuronal disorders and encephalopathy, collectively known as seipinopathies18,19. Hence, how seipin functions is key to determine the mechanisms of LD formation and to understand the aetiology of its associated diseases.
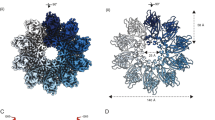
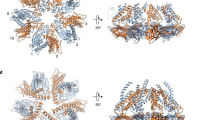
Recently resolved cryo-EM structures of human, fly, and yeast seipin revealed that it forms a large ring-shaped oligomeric structure in the ER comprising of 11, 12, and 10 subunits respectively7,20,21. Each subunit forms a toroidal structure of ~15 nm in diameter and comprises of a central luminal domain region (LR), flanked by two transmembrane domains (TMDs), and short N- and C- cytosolic regions5,22. The LR of mammalian and fly seipin contains hydrophobic helices (HHs) in the centre of the ring that associates with the luminal leaflet of the ER and binds to TAG in vivo21. Consistent with this, Molecular Dynamic Simulations (MDS), and mutation analyses have shown that the HHs can sequester nanoscale TAG8,23 and SE24 molecules within the seipin ring complex. The occurrence of NL nucleation remained low in the absence of seipin, thus suggesting direct interaction of various NL with seipin24. Hence, presence of seipin induces TAGs being oiled out into lenses at a lower concentration than TAG alone. As a result, LDs preferentially form at seipin demarcated sites, and ER TAG levels increases in the absence of seipin9,13,25. However, yeast seipin lacks HH, but its function is provided by Ldb16, a yeast-specific seipin partner protein7. MD simulation showed consistent but mild accumulation of TAG in the proximity of TMD segments, implicating the role of membrane region in LD formation7. Moreover, the TM region regulates the kinetics of TAG accumulation within the seipin ring and acts as gates for the entry of TAG molecules23. The structure also resolved two separate conformations for the TMDs that would provide TMD the flexibility to open up to accommodate additional TAG molecules within the central ring during LD biogenesis5. Consistent with this, a conserved short luminal helix near TMDs, known as locking helix (LH) or switch domain plays a crucial role in TMD orientation and flexibility5,7. A recent study using MDS has shown that hSeipin TMD segments apart from HH play critical role in TAG nucleation and assist in converting flat TAG-lenses into growing nascent LDs26.
In this study, we investigated the role of seipin TMDs in LD biogenesis in vivo. We report truncated seipin, Fld1-∆LR in yeast, and the human equivalent hSeipin-∆LR, mutants devoid of the LR, however, containing the two conserved TMDs, a small LH or switch domain adjoining TMDs, and short N- and C- terminal cytosolic regions., Fld1-∆LR localizes to ER subdomains independently of LDs like wild-type (WT) Fld1. We provide evidence that Fld1-∆LR represents functional LD biogenesis sites as it can recruit the LD biogenesis machinery, including accessory factors, Nem1, Ldb16, Pex30, and Yft2 and these sites become enriched in diacylglycerol (DAG) upon initiation of LD formation. We find that in contrast to Fld1, Fld1-∆LR restored the LD morphology defect and terbinafine growth sensitivity of fld1∆ ldb16∆ mutants like hSeipin. Similarly, expression of hSeipin-∆LR in yeast exhibits punctate localization and complemented the aberrant LD morphology and terbinafine sensitivity of fld1∆ and fld1∆ ldb16∆ cells. Intriguingly, mutating the 31st amino acid proline in Fld1-∆LR to glycine perturbs the localization of Fld1-∆LR and renders it non-functional in reversing the defects of fld1∆, and fld1∆ ldb16∆ mutants. We demonstrate that Fld1-∆LR can assemble into oligomeric complex independently of the LR, or the presence of Ldb16, a yeast specific Fld1 partner. We found that lack of LR in Fld1-∆LR/hSeipin-∆LR resulted in increased steady state TAG levels, suggesting crucial regulatory role of the LR in regulating TAG partitioning into LDs. However, Fld1-ΔLRP31G resulted in loss of TAG accumulation activity of Fld1-ΔLR. In addition, abrogation of LH or switch region mislocalizes hSeipin-∆LR, and renders it non-functional in complementing fld1∆ mutants. Together these findings shed new light into the mechanism of LD assembly. Collectively, we propose a model in which seipin lacking LR represents the minimal region of seipin that is necessary and sufficient in determining functional sites of LD biogenesis.
Results
Fld1-∆LR localizes to ER subdomains independent of LDs
To investigate the role of seipin TMDs in LD nucleation, we generated truncated version of yeast seipin (amino acids 1−285) lacking the LR, comprising transmembrane segment 1 (TM-1, residues 1−50), and transmembrane segment 2 (TM-2, residues 235−285) and expressed as mCherry fusion on a plasmid under ADH1 promoter (Fig. 1A). The fusion protein called Fld1-∆LR was expressed in yeast fld1∆ mutants co-expressing Sec63-GFP, an ER marker. The Fld1-∆LR revealed punctate localization in the ER similar to Fld1 (Fig. 1B). Intriguingly, appending mCherry with TM segment of Sec63 (residues 1-250) and expressed as a Sec63-TMD-mCherry fusion resulted in uniform ER localization (Fig. S1A, B), indicating that punctate localization is specific for Fld1-∆LR. The Fld1-∆LR-mCherry appeared as ~35 kDa protein as revealed by Western blot analysis (Fig. 1C). Previously we have reported that localization of Fld1 is independent of NL synthesis and the presence of LDs9, we wondered if localization of Fld1-∆LR requires the presence of LDs. Therefore, we expressed Fld1-∆LR in yeast mutants lacking the capacity to produce NLs. Synthesis of NLs in yeast is controlled by four acyltransferases, Lro1 and Dga1 produce TAG, whereas Are1 and Are2 synthesize SE. Yeast mutants devoid of all four enzymes are1∆ are2∆ dga1∆ lro1∆ (called 4∆KO) are devoid of NL synthesis and hence contain no detectable LDs27. Hence, we expressed Fld1 and Fld1-∆LR in 4∆KO yeast mutants lacking Fld1. Intriguingly, Fld1-∆LR localized into distinct foci like Fld1 in 4∆KO fld1∆ cells lacking any detectable LDs as evidenced by staining with BODIPY, a NL staining dye (Fig. 1D)9. This indicates that the localization of Fld1-∆LR to discrete ER sites is independent of NL synthesis, or the presence of LDs. Next, we wondered if these foci are stable over time. Previously it has been shown that seipin puncta stably associates with LD precursors thereby defining and/or stabilizing ER subdomains where droplets form9,13,16. Therefore, we visualized the mobility of Fld1-∆LR in 4∆KO fld1∆ cells by time-lapse microscopy. Most of the Fld1-∆LR spots were stable over time, with a few showing slower mobility, like WT Fld1 (Fig. S1C, D, Supplementary Movie S1 and S2), in agreement with previous observations9,16,17. Overall, these results suggest that Fld1-∆LR localizes to discrete ER subdomains independently of LDs and probably mark potential sites of LD biogenesis (Fig. 1E).
A Cartoon depicting generation of Fld1-∆LR lacking luminal domain region. B Fld1-∆LR displays ER localization like WT Fld1. Yeast fld1∆ cells harbouring mCherry fused to Fld1 or Fld1-∆LR and co-expressing Sec63-GFP were grown in SC media and visualized by fluorescence microscopy. Representative image shown from three independent experiments. Scale bar: 5 µm. C Western blot showing expression of Fld1-mCherry or Fld1-∆LR-mCherry. PGK1 serves as a loading control. D Localization of Fld1-∆LR is independent of LDs. Cells of the indicated genotypes expressing Fld1-mCherry or Fld1-∆LR-mCherry were grown in SC media, stained with BODIPY and visualized by fluorescence microscopy. Representative image shown from three independent experiments. Scale bar: 5 µm. E Illustration depicting domains contained within the Fld1-∆LR, two transmembrane segments and a short-conserved locking helix (LH) connecting the two TM-regions. TM-1, transmembrane-1; TM-2, transmembrane-2; LH- locking helix.
Fld1-∆LR functionally complements lack of seipin function
Given that Fld1-∆LR displays localization like Fld1, we wondered if these puncta are functional. Therefore, we visualized LD morphology phenotype of fld1∆ cells expressing either Fld1 or Fld1-∆LR. Fld1 mutants display aberrant LD phenotype with either many tiny droplets or a few supersized LDs, as previously reported28, a phenotype that we confirmed upon BODIPY staining (Fig. 2A). However, overexpression of Fld1-∆LR completely reversed the LD morphology defects of fld1∆ cells like Fld1 (Fig. 2B–E). Quantitative analysis revealed ~80% colocalization of LDs with Fld1-∆LR that was indistinguishable in comparison to WT Fld1 (Fig. 2F). Number and size distribution of LDs in fld1∆ cells were alike upon overexpression of either Fld1 or Fld1-∆LR (Fig. 2D, E). To investigate the LD morphology phenotype at the ultrastructural level, we performed electron microscopy of fld1∆ mutants expressing either Fld1 or Fld1-∆LR. Fld1 mutants contained electron translucent LDs either in clusters mostly in the range of 100-150 nm, or supersized LD ( > 500 nm) in comparison to WT cells (Fig. 2G, H). However, fld1∆ mutants containing Fld1-∆LR rescued the LD morphology defects like Fld1 (Fig. 2G). Quantitative analysis revealed most of the LDs having diameter within 200–500 nm for both Fld1 or Fld1-∆LR (Fig. 2H). Moreover, cells did not show clustering or supersized LD phenotype (Fig. 2G). Next, we assessed the functionality of Fld1-∆LR. Absence of Fld1 renders yeast cells sensitive to terbinafine, a sterol biosynthesis inhibitor and has been used for growth sensitivity assays previously5,29, a phenotype we confirm (Fig. 2I). We found that Fld1-∆LR fully restored the growth defect of fld1∆ mutants to the same extent of WT Fld1 in the presence of terbinafine (Fig. 2I), suggesting that Fld1-∆LR is functional. Taken together, these findings indicate that Fld1-∆LR at ER subdomains can functionally complement the absence of seipin.
A Aberrant LDs accumulate in the absence of Fld1. WT and fld1∆ cells were grown in YPD media, stained with BODIPY and subjected to fluorescence imaging. Representative image shown from three independent experiments. Scale bar: 5 µm. B Fld1-∆LR restores LD morphology defect of fld1∆ mutants. Cells expressing Fld1 or Fld1-∆LR as mCherry fusion in fld1∆ cells were cultivated in SC media, stained with BODIPY, and imaged using fluorescence microscope. White arrowheads denote colocalization between LDs and either Fld1 or Fld1-∆LR. Representative image shown from three independent experiment. Scale bar: 5 µm. C Line scans of signal intensity along the yellow lines shown in (B). D, E Quantification of LD number (D) and size distribution (E) in cells of the indicated genotypes. Data represent mean ± s.d. and were analysed with one-way ANOVA and Tukey’s comparisons. n = 100 cells. F Quantification of colocalization between LDs and either Fld1 or Fld1-∆LR. Data represent mean ± s.d and was analyzed using two-tailed unpaired t-test. n = 75 cells. G Ultrastructural analysis of LDs. Cells of the indicated genotypes were fixed, and processed for EM imaging as described in the Methods section. N nucleus, LD lipid droplet, V vacuole, ER endoplasmic reticulum, M mitochondria. Scale bar: 500 nm. H Quantification of LD size distribution of the cells shown in (G). n = 20 cells. I Terbinafine growth assay. Indicated yeast strains were grown in either YPD or SC media, serially diluted, and spotted on YPD plates containing either DMSO or terbinafine at a final concentration of 100 μg/mL. Representative image shown from three independent experiments. Source data are provided as a Source Data file.
Both TM-1 and TM-2 segments are required together to constitute a functional Fld1-∆LR
We next asked which region of Fld1-∆LR is responsible for the punctate localization in the ER. Therefore, we expressed TM-1 and TM-2 in isolation as a mCherry fusion in 4∆KO fld1∆ cells (Fig. S2A, B). TM-1-mCherry fusion showed punctate ER localization, whereas TM-2-mCherry failed to localize into puncta (Fig. S2B). To determine if TM-1 or TM-2 constructs would rescue aberrant LD phenotype of fld1∆ mutants, we expressed TM-1 and TM-2 fusions in fld1∆ background (Fig. S2C). However, both TM-1 and TM-2 fusion constructs failed to rescue LD morphology defect of fld1∆ cells (Fig. S2C). Next, we performed growth assay to assess the functionality of TM-1-mCherry and TM-2-mCherry fusions. Both TM-1 and TM-2-mCherry fusion proteins failed to rescue the growth defect of fld1∆ mutants in the presence of terbinafine (Fig. S2D). These results indicate that the information pertaining to punctate localization of Fld1 resides within the first 50 residues contained within the TM-1 region, however both TM-1 and TM-2 are essentially required together to constitute a fully functional Fld1-∆LR.
Fld1-∆LR adopts membrane topology similar to WT protein
Further, we were interested in determining the membrane topology of Fld1-∆LR. Fld1 is an integral ER membrane protein, having two TMDs, and short N- and C-terminus exposed towards the cytosol and a large LR oriented towards the ER lumen29,30. To determine whether Fld1-∆LR is correctly inserted in the ER membrane we expressed functional N- and C-terminal tagged Fld1-∆LR, GFP-Fld1-∆LR and Fld1-∆LR-mCherry in yeast fld1∆ mutants (Fig. S3A, B). Both the N- and C-terminal fusions localized into puncta and reversed the aberrant LD phenotype of fld1∆ mutants (Fig. S3B). Intriguingly, both the GFP-Fld1-∆LR, and Fld1-∆LR-mCherry constructs complemented the terbinafine sensitivity of fld1∆ mutants (Fig. S3C). This indicates that both N- and C-terminal fusions are functional in vivo. Therefore, we sought to determine the topology of these constructs. Subcellular fractionation revealed that Fld1-∆LR is enriched in a 13 K (P13) microsomal fraction, and hence the protein is membrane associated (Fig. S3D). Extraction of microsomal fraction using salt did not render the protein soluble (Fig. S3E). However, using 1% Triton, resulted in partial solubilization of Fld1-∆LR-mCherry whereas GFP-Fld1-∆LR was completely solubilized. On the other hand using 1% SDS completely solubilized both Fld1-∆LR-mCherry and GFP-Fld1-∆LR (Fig. S3E), indicating that Fld1-∆LR is an integral membrane protein like Dpm1 used as a control (Fig. S3E). Protease protection assay in the presence and absence of detergents revealed that both the N- and C-terminus of Fld1-∆LR is protease accessible, suggesting that Fld1-∆LR has even number of TMDs and N- and C-termini are exposed towards the cytoplasm (Fig. S3F). Kar2, an ER luminal chaperone served as a control and remained protected from Proteinase K digestion in the absence of detergents (Fig. S3F). Based on these results, we propose a model for the topology of Fld1-∆LR having both N- and C-terminus oriented towards cytoplasm like WT Fld1 (Fig. S3G).
Fld1-∆LR under endogenous expression levels restores fld1∆ mutant phenotype
We next asked if expression of Fld1-∆LR driven by native promoter would rescue fld1∆ mutants. Hence, GFP-Fld1 and GFP-Fld1-∆LR were expressed under endogenous promoter on a plasmid in fld1∆ background. Although the intensity of GFP foci were weak, Fld1-∆LR puncta colocalized with MDH stained LDs like WT Fld1 (Fig. 3A, Fig. S4A). Number and size distribution of LDs in fld1∆ cells were indistinguishable in the presence of either Fld1 or Fld1-∆LR (Fig. 3B, C), suggesting Fld1-∆LR complements the LD morphology defect of fld1∆ cells like Fld1. Alternately, an untagged plasmid borne Fld1-∆LR under endogenous promoter was expressed in fld1∆ mutant and reversal of LD morphology defect was observed by BODIPY staining. Cells did not show tiny or supersized LD phenotype, typical of seipin mutants (Fig. S4B). LD number and size distribution in fld1∆ cells were indistinguishable in the presence of either Fld1 or Fld1-∆LR (Fig. S4C, D). Moreover, endogenously expressed Fld1-∆LR rescued the terbinafine growth defect of fld1∆ mutants albeit to a slightly lesser degree than WT Fld1 by day 3, however, upon prolonged incubation until day 4, the growth rescue was comparable to that of WT Fld1 (Fig. 3D, Fig. S4E). These data indicate that Fld1-∆LR under native expression levels can complement the lack of function of seipin similar to WT Fld1.
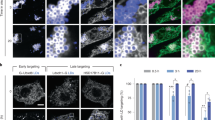
A Fld1-∆LR under endogenous expression levels. Fld1 mutants expressing either GFP-Fld1 or GFP-Fld1-∆LR under native promoter were grown in selective media, stained with LD dye MDH, and visualized by fluorescence microscopy. White arrowheads denote colocalization between LDs and either Fld1 or Fld1-∆LR. Representative image shown from three independent experiments. Scale bars: 5 µm. B, C Quantification of LD number (B) and size distribution (C) in cells of the indicated genotypes. Data represent median and were analysed with two-way ANOVA and Tukey’s multiple comparisons. n = 100 cells. D Terbinafine growth assay. Indicated yeast strains were grown in either YPD or SC media, serially diluted, and spotted on YPD plates containing either DMSO or terbinafine at a final concentration of 100 μg/mL. Plates were imaged at intervals of day 3 and day 4. Representative image shown from four independent experiments. E, F Induction of LD biogenesis in conditional yeast mutant. Inducible yeast mutant expressing either WT Fld1 or Fld1-∆LR were grown in raffinose containing media and transferred to galactose containing media for the indicated time, stained with BODIPY and subjected to fluorescence imaging. White arrowheads denote colocalization between LDs and either Fld1 (E) or Fld1-∆LR (F). Representative image shown from three independent experiments. Scale bars: 5 µm. G Quantification of LDs formed in inducible yeast mutants in the presence of either Fld1 or Fld1-∆LR. Data represent mean ± s.d. and were analysed with two-way ANOVA and Holm-Sidak’s comparisons. n = 100 cells. H, I LD marker protein colocalizes with Fld1-∆LR. Inducible yeast mutants of the indicated genotypes expressing endogenously Erg6-GFP, an LD marker protein was cultivated in raffinose containing media and shifted to galactose for the indicated time. White arrowheads depict colocalization of Erg6 with either Fld1 (H) or Fld1-∆LR (I). Representative image shown from three independent experiments. Scale bars: 5 µm. J Quantification of colocalization between Erg6 and either Fld1 or Fld1-∆LR. Data represent mean ± s.d. and were analysed with two-way ANOVA and Holm-Sidak’s comparisons. n = 150 cells. Source data are provided as a Source Data file.
Fld1-∆LR demarcates sites of de novo LD biogenesis in the ER
Given that Fld1-∆LR localizes to ER subdomains and rescues fld1∆ mutant phenotype, we wondered if these puncta are bonafide sites where de novo LDs begin to form. Therefore, we employed an inducible yeast mutant are1∆ are2∆ dga1∆ GAL-LRO1, which initiates production of de novo LDs upon galactose addition9,31. We deleted Fld1 in this strain (called 3∆ fld1∆ GAL-LRO1). The strain then expressed Fld1 or Fld1-∆LR on a plasmid (Fig. 3E, F) and chromosomally encoded Erg6-GFP (Fig. 3H, I), a protein that resides onto LDs, but is on the ER in cells devoid of LDs31,32. Before galactose addition cells lack any detectable LDs as revealed by BODIPY staining, though Fld1 and Fld1-∆LR showed punctate localization (Fig. 3E, F). Upon 2−5 h of galactose addition, BODIPY started to mark puncta that colocalized with Fld1- or Fld1-∆LR (Fig. 3E, F). These LDs remained associated with Fld1 or Fld1-∆LR upon 16 h of galactose induction (Fig. 3E, F), suggesting that mature LDs do not dissociate from Fld1-∆LR puncta and remained associated similar to Fld19. Quantitative analysis revealed comparable number of LDs formed in a time dependent manner when cells contained either Fld1 or Fld1-∆LR (Fig. 3G). This indicates that Fld1-∆LR demarcates LD biogenesis sites like Fld1. Consistent with this, Erg6-GFP becomes enriched at LD biogenesis sites 2−5 h after galactose induction (Fig. 3H, I) and displays >80% colocalization with Fld1-∆LR (Fig. 3J). Taken together, these finding suggest that Fld1-∆LR marks ER subdomains like WT Fld1 at which de novo LDs are produced.
Fld1-∆LR collaborates with LD assembly factors
If Fld1-∆LR primes ER subdomains permissive for LD biogenesis, we hypothesized that these subdomains might become enriched in LD biogenesis factors, as previously demonstrated for WT Fld19. Therefore, fld1∆ cells expressing plasmid borne copy of either Fld1-mCherry or Fld1-∆LR-mCherry and co-expressing GFP-tagged LD assembly factors; Nem1, a regulator for DAG production10, Ldb16, an Fld1 partner protein33, and Pex30, an ER membrane shaping protein34 were visualized by fluorescence microscopy and Pearson’s correlation coefficient of colocalization was determined (Fig. 4A–F). Fld1-∆LR puncta displayed association with LD biogenesis factors like WT Fld1 as it colocalized with Nem1 (Fig. 4A, B), Ldb16 (Fig. 4C, D), and Pex30 (Fig. 4E, F). This suggests that Fld1-∆LR marked ER subdomains indeed recruit proteins crucial for droplet assembly. We next asked if TAG-synthase also localizes at these Fld1-∆LR marked ER sites. Hence, we expressed Fld1-mCherry or Fld1-∆LR-mCherry on a plasmid in 4∆KO fld1∆ cells. These cells co-expressed TAG-synthase sLro1 fused to GFP on a plasmid under GAL promoter as previously described9. Intriguingly, galactose induction of sLro1 for 2 h resulted in colocalization of most of the sLro1 puncta with Fld1-∆LR like WT Fld1 (Fig. 4G, H). This indicates that Fld1-∆LR defined ER subdomains can recruit TAG-synthesizing enzyme for localized production of TAG and LD formation.
A−F Colocalization of LD assembly factors with Fld1-∆LR. Yeast fld1∆ mutants expressing either Fld1-mCherry or Fld1-∆LR-mCherry on a plasmid and co-expressing either Nem1-GFP (A), Ldb16-GFP (C), or Pex30-GFP (E) were grown in SC media and subjected to fluorescence microscopy. White arrowheads denote colocalization between LD assembly factors, Nem1, Ldb16, and Pex30 with either Fld1 or Fld1-∆LR. Representative image shown from three independent experiments. Scale bars: 5 µm. Quantitative analysis of the colocalization of either Nem1-GFP (B), Ldb16-GFP (D), or Pex30-GFP (F) with either Fld1 or Fld1-∆LR by Pearson’s correlation coefficient. Data represent mean ± s.d. and were analysed with two-tailed unpaired t-test. n = 25 cells. G TAG-synthase sLro1 colocalizes with Fld1-∆LR. 4Δ KO yeast mutant lacking Fld1 expressing mCherry fused to either Fld1 or Fld1-∆LR and co-expressing galactose inducible sLro1-GFP were shifted from raffinose to galactose containing media for the indicated time. White arrowheads depict colocalization of sLro1 with Fld1 or Fld1-∆LR. Representative image shown from three independent experiments. Scale bar: 5 µm. H Quantitative analysis of the colocalization of sLro1 with either Fld1 or Fld1-∆LR by Pearson’s correlation coefficient. Data represent mean ± s.d. and were analysed with two-tailed unpaired t-test. n = 25 cells. I, J Fld1-∆LR marked ER sites recruit yeast FITM2 protein Yft2. Yeast fld1∆ mutants containing either Fld1-mCherry or Fld1-∆LR-mCherry and co-expressing Yft2-sf-GFP were cultivated in the presence of oleic acid (OA) to stimulate LD formation for the indicated time. White arrowheads indicate colocalization of Yft2 with Fld1 or Fld1-∆LR. Representative image shown from three independent experiments. Scale bars: 5 µm. K Quantitative analysis of the colocalization of Yft2 with either Fld1 or Fld1-∆LR by Pearson’s correlation coefficient. Data represent mean ± s.d. and were analysed with two-tailed unpaired t-test. n = 25 cells. L, M Fld1-∆LR marked ER site become enriched in DAG. Cells of the indicated genotypes co-expressing ER-DAG sensor were grown in the presence of OA for the indicated time. White arrowheads denote colocalization of ER-DAG sensor puncta with Fld1 or Fld1-∆LR. Representative image shown from three independent experiments. Scale bars: 5 µm. N Quantitative analysis of the colocalization of ER-DAG sensor with either Fld1 or Fld1-∆LR by Pearson’s correlation coefficient. Data represent mean ± s.d. and were analysed with two-tailed unpaired t-test. n = 25 cells. Source data are provided as a Source Data file.
LD emergence protein Yft2 colocalizes with Fld1-∆LR upon stimulation of LD formation
Thereafter, we investigated if yeast FITM2 protein, Yft2 becomes enriched at Fld1-∆LR sites. Previously we have shown that yeast FITM2 proteins get recruited at Fld1 defined LD biogenesis sites9,35. These proteins facilitate proper emergence of LDs from the ER membrane towards the cytosol. Thus, fld1∆ cells expressing plasmid borne either Fld1-mCherry or Fld1-∆LR-mCherry and co-expressing Yft2-superfold-GFP were stimulated to produce LDs in the presence of oleic acid (OA). In the absence of OA, Yft2 shows uniform ER localization (Fig. 4I, J; 0 h), however, upon OA induction, Yft2 gets enriched into puncta that colocalized with either Fld1 or Fld1-∆LR (Fig. 4I, J; 2.5 h). We found Pearson’s correlation coefficient of Yft2 colocalizing with Fld1-∆LR to the same extent as that of WT Fld1 (Fig. 4K). This suggests that de novo droplet assembly drives recruitment of Yft2 at Fld1-∆LR marked ER sites.
DAG levels are altered at Fld1-∆LR marked LD biogenesis sites
Since LD biogenesis factors show enrichment at Fld1-∆LR sites, we wondered if DAG levels at these ER subdomains gets altered during LD biogenesis. Using a GFP-tagged ER DAG sensor previously we had demonstrated that DAG accumulates at sites of LD biogenesis9,35. Hence, we expressed ER-DAG sensor-GFP on a plasmid in fld1∆ mutants that co-expressed either Fld1-mCherry or Fld1-∆LR-mCherry. Upon OA addition of 2.5 h, the ER-DAG sensor-GFP formed puncta that colocalized with Fld1 or Fld1-∆LR (Fig. 4L, M). Pearson’s coefficient value of colocalization of Fld1-∆LR with ER-DAG sensor-GFP was similar to WT Fld1 (Fig. 4N). This suggests that levels of DAG, a substrate for TAG-synthase gets altered at Fld1-∆LR sites during de novo LD formation.
Fld1-∆LR complements the LD morphology defect of fld1∆ ldb16∆ double mutants similar to hSeipin
Next, we wanted to determine how lack of Fld1, and its partner Ldb16 from LD biogenesis sites would perturb the functioning of Fld1-∆LR. Previously it has been reported that lack of either Fld1 or Ldb16 or a fld1∆ ldb16∆ double mutant results in an indistinguishable LD morphology defect33, with few large LDs and clustered tiny LDs, a phenotype that we confirm (Fig. 5A). To investigate the role of Fld1-∆LR, we expressed plasmid borne copy of either Fld1 or Fld1-∆LR in fld1∆ldb16∆ mutants and visualized LD morphology upon BODIPY staining by fluorescence microscopy. We found that both Fld1 and Fld1-∆LR displayed punctate foci in the ER that colocalized with LDs, suggesting their localization is not dependent on the presence of Ldb16 (Fig. 5B). LD number quantification revealed ~1 LD per cell in fld1∆ldb16∆ mutants compared to ~3-4 LDs in WT cells, a phenotype that was partially rescued by expression of WT Fld1 (Fig. 5C), consistent with previous observations7,29,33. However, unlike Fld1, Fld1-∆LR reversed the LD number in fld1∆ldb16∆ mutants to that of WT levels (Fig. 5C). Similarly, quantification of LD size revealed majority population of supersized LDs having diameter >0.5 um2 in fld1∆ldb16∆ mutants (Fig. 5D). Expression of WT Fld1 resulted in an intermediate phenotype with most cells still having supersized LD distribution, however, Fld1-∆LR construct rescued the LD size distribution similar to that of WT cells (Fig. 5D). It has been reported previously that hSeipin can functionally complement the LD morphology defects of fld1∆ ldb16∆ mutants, unlike Fld129,33. Therefore, we expressed GFP tagged hSeipin in fld1∆ and fld1∆ ldb16∆ yeast mutants (Fig. 5E). GFP-hSeipin displayed punctate distribution that colocalized with MDH stained LDs (Fig. 5E). Quantitative analysis in the presence of hSeipin revealed rescue of LD number and size distribution of fld1∆ ldb16∆ mutants to that of WT cells (Fig. 5C–E). This rescue of LD number and size distribution in fld1∆ ldb16∆ mutants by hSeipin were comparable to that of Fld1-∆LR (Fig. 5C, D). Cells devoid of Fld1 or its partner Ldb16 sensitizes yeast cells to terbinafine29, a phenotype we confirm (Fig. 5F). However, unlike Fld1, expression of Fld1-∆LR in fld1∆ ldb16∆ mutants complemented the terbinafine growth sensitivity, albeit to a lower extent than WT hSeipin (Fig. 5F). Taken together, these results suggest that Fld1-∆LR can functionally restore the LD biogenesis defect of fld1∆ ldb16∆ double mutants like hSeipin.
A Lack of Fld1 or Ldb16 or both Fld1 and Ldb16 results in an indistinguishable LD morphology defect. Cells of the indicated genotypes were grown in YPD media, stained with BODIPY and imaged in a fluorescent microscope. Representative image shown from three independent experiments. Scale bars: 5 µm. B Fld1-∆LR rescues LD phenotype of fld1∆ ldb16∆ double mutant. Yeast fld1∆ ldb16∆ double mutants expressing either Fld1 alone or Fld1-∆LR as an mCherry fusion were grown in SC media, stained with LD dye BODIPY and subjected to fluorescence microscopy. White arrowheads denote colocalization between LDs with either Fld1 or Fld1-∆LR. Representative image shown from three independent experiments. Scale bars: 5 µm. C, D Quantification of LD number (C) and size distribution (D) in cells of the indicated genotypes. Data represent mean ± s.d. and were analysed with two-way ANOVA and Tukey’s multiple comparisons. n = 100 cells. E Human Seipin rescues the lack of Fld1 and Ldb16 in yeast. Yeast fld1∆ and fld1∆ ldb16∆ cells expressing GFP-hSeipin on a plasmid were grown in selective media, stained with LD dye, MDH and visualized by fluorescence microscopy. White arrowheads denote colocalization between hSeipin marked puncta and LDs. Representative image shown from three independent experiments. Scale bar: 5 µm. F Fld1-∆LR compliments terbinafine growth sensitivity of fld1∆ ldb16∆ cells like hSeipin. Indicated yeast strains were cultivated in either YPD or SC media, serially diluted, and spotted on YPD plates containing either DMSO or terbinafine at a final concentration of 100 μg/mL. Representative image shown from three independent experiments. G Western blot analysis. Western blot showing expression of Fld1-mCherry or Fld1-∆LR-mCherry or Fld1-∆LR-mCherryP31G. PGK1 serves as a loading control. H Fld1-∆LR-mCherryP31G gets mislocalized. Yeast fld1∆ and fld1∆ lbd16∆ cells expressing Fld1-∆LR-mCherryP31G were grown in selective media, stained with BODIPY and imaged. Representative image shown from three independent experiments. Scale bar: 5 µm. I, J Quantification of LD number (I) and size distribution (J) in cells of the indicated genotypes. Data represent mean ± s.d. and were analysed with two-way ANOVA and Tukey’s multiple comparisons. n = 100 cells. K Terbinafine growth assay. Yeast strains of the indicated genotypes were grown in either YPD or SC media, serially diluted, and spotted on YPD plates containing either DMSO or terbinafine at a final concentration of 100 μg/mL. Representative image from three independent experiments is shown. Source data are provided as a Source Data file.
Fld1-∆LR is mislocalized in the presence of endogenous Fld1
The fact that Fld1-∆LR is functional in fld1∆ ldb16∆ mutants unlike Fld1, suggests some gain of function of Fld1-∆LR compared to WT Fld1. To investigate this gain of function, we expressed Fld1-∆LR in ldb16∆ mutants that contains endogenous Fld1. We found that presence of Fld1-∆LR did not rescue the terbinafine sensitivity of ldb16∆ mutants (Fig. S5A) to the extent of rescue that we found in fld1∆ ldb16∆ double mutants (Fig. 5F). A plausible explanation for this could perhaps be that when endogenous Fld1 is present, i.e. in ldb16∆ mutants, it might interfere with the localization of plasmid borne copy of Fld1-∆LR at LD biogenesis sites. However, when native Fld1 is absent, i.e. in fld1∆ ldb16∆ cells, Fld1-∆LR can properly localize at LD biogenesis sites. To test this hypothesis Fld1-∆LR-mCherry was expressed in fld1∆, ldb16∆, and fld1∆ ldb16∆ mutants and visualized by fluorescence microscopy (Fig. S5B). Yeast fld1∆, and fld1∆ ldb16∆ mutants where endogenous Fld1 is missing showed punctate distribution of Fld1-∆LR, however, ldb16∆ mutants where endogenous Fld1 is still present, displayed mostly ER staining of Fld1-∆LR with few isolated foci (Fig. S5B). This suggests that in the presence of endogenous Fld1, Fld1-∆LR is mislocalized, a likely reason for it being non-functional in ldb16∆ mutants (Fig. S5A). If this is correct, we asked what would happen if we brought back endogenous Fld1 in fld1∆ ldb16∆ cells expressing Fld1-∆LR-mCherry. Hence GFP tagged Fld1 under its own promoter was co-expressed in fld1∆ ldb16∆ cells together with Fld1-∆LR-mCherry. Re-introduction of endogenous Fld1-GFP resulted in mislocalization of Fld1-∆LR-mCherry in fld1∆ ldb16∆ mutants (Fig. S5C). This suggests that presence of endogenous seipin simultaneously with Fld1-∆LR perturbs the localization and functioning of Fld1-∆LR. These interesting observations should be further investigated in future studies.
Fld1-∆LRP31G point mutant abolishes its activity
We next investigated if generating point mutation within the Fld1-∆LR would abolish its activity. Consistent with previous report of importance of tyrosine residue at position 37 (Y37) that interacts with methionine 240 (M240) for aromatic-methionine interaction, disruption of which affects ldb16 levels and results in LD morphology defects7, we mutated Y37 within TM-1, and Y248 within TM-2 to alanine (Y37A, Y248A), and expressed mutated Fld1-∆LR as mCherry fusion in fld1∆ cells. Both the mutants, Fld1-∆LRY37A, Fld1-∆LRY248A rescued the terbinafine resistance of fld1∆ cells and hence failed to impair the functioning of the Fld1-∆LR (Fig. S5D). Since our data suggests that the information for punctate localization resides within the first 50 residues containing TM-1 of Fld1, we focused on mutating key residue within this region. Hence, we mutated proline at residue 31 to glycine (P31G) within the TM-1 region of Fld1-∆LR and expressed as mCherry fusion in fld1∆ cells. Western blot analysis revealed Fld1-∆LRP31G-mCherry having similar size to Fld1-∆LR-mCherry, albeit expressed at a lower level (Fig. 5G). Fld1-∆LRP31G mutant when expressed in fld1∆ or fld1∆ ldb16∆ cells failed to localize into puncta and showed diffused ER localization (Fig. 5H). Consistent with this mislocalization, Fld1-∆LRP31G failed to rescue the LD morphology defect and terbinafine sensitivity of fld1∆ or fld1∆ ldb16∆ double mutants (Fig. 5H-K).
Further we tested if mislocalization of Fld1-∆LRP31G affected the distribution of LD biogenesis factors. Therefore, we performed colocalization of Fld1-ΔLRP31G with Nem1, Pex30, Ldb16 and Yft2. The Fld1-ΔLRP31G-mCherry failed to localize into puncta and mostly showed uniform ER distribution, together with vacuolar staining (Fig. S6A–D). However, mutant Fld1-ΔLRP31G did not affect the localization of Nem1-GFP, and Pex30-GFP, although most of the Nem1-GFP, and Pex30-GFP puncta failed to colocalize with Fld1-ΔLRP31G, albeit a few that colocalized with Fld1-ΔLRP31G (Fig. S6A, B). In the presence of mutant Fld1-ΔLRP31G majority of cells displayed uniform ER staining of Ldb16-GFP with a few cells showing isolated puncta together with ER staining (Fig. S6C) in contrast to WT Fld1-ΔLR (Fig.4C). Similarly, Yft2 failed to localize into puncta upon stimulation of LD formation in the presence of OA when cells expressed mutant form of Fld1-ΔLRP31G (Fig. S6D) in contrast to WT Fld1-ΔLR (Fig.4J). This suggests that when WT Fld1-ΔLR is present it properly localizes into discrete puncta and creates functional LD biogenesis sites and recruits all the necessary LD biogenesis factors at these ER subdomains (Fig. 4), whereas mutant Fld1-ΔLRP31G failed to localize to discrete ER subdomains and instead display uniform ER distribution a likely reason for the defective assembly of LD biogenesis sites.
Fld1-∆LRP31G results in mislocalization of nascent LD probe LiveDrop
We next determined if defective formation of LD biogenesis sites in the presence of Fld1-∆LRP31G would abrogate localization of nascent LD marker LiveDrop, an established probe derived from GPAT4 protein in flies to visualize active LD biogenesis sites16. LiveDrop probe has been used in yeast and shows dual localization both in the ER and on the LDs36. Hence we expressed GFP-tagged LiveDrop and co-expressed either Fld1-mCherry or Fld1-ΔLR-mCherry or Fld1-ΔLRP31G-mCherry on a plasmid in fld1∆ cells and stained LDs using MDH, a blue staining LD dye and visualized by fluorescence microscopy and determined Pearson’s correlation coefficient of colocalization (Fig. S7A-D). Triple colour imaging revealed WT Fld1 and Fld1-ΔLR marked -mCherry puncta colocalizing with GFP-LiveDrop and MDH stained LDs (Fig. S7A,B). Quantitative analysis revealed ~75% colocalization between LiveDrop and mCherry/MDH stained puncta (Fig. S7C, D). This suggests that Fld1-ΔLR marked LD biogenesis sites in the ER is efficiently recognized by GFP-LiveDrop. However, Fld1-ΔLRP31G mutant failed to localize into puncta and displayed mostly diffused vacuolar staining and did not rescue LD morphology defect of fld1∆ cells. Intriguingly, in these cells GFP-LiveDrop displayed mostly uniform ER localization and failed to mark sites of LD biogenesis in the ER or to localize onto LDs (Fig. S7A, B). This suggests that in the presence of mutant Fld1-ΔLRP31G LD biogenesis sites are not effectively established a likely reason for LiveDrop probe to not localize at these sites.
Fld1-∆LR assembles into oligomers
Next, we determined whether the Fld1-∆LR retains its ability to oligomerize, a crucial property of seipin5,7,20,21,37. Therefore, we generated HA-epitope tagged version of Fld1 and Fld1-∆LR on a GAL-inducible plasmid. First, we tested if Fld1-HA and Fld1-∆LR-HA fusions were functional. Hence, we expressed either Fld1-HA or Fld1-∆LR-HA in fld1∆ or fld1∆ ldb16∆ double mutants and visualized LD morphology by BODIPY staining. At 0 hour fld1∆ or fld1∆ ldb16∆ cells displayed irregular LD morphology defect, typical of seipin deletion (Fig. 6A, B). However, galactose induction for 2.5 h resulted in complete reversal of LD morphology defect of fld1∆ cells (Fig. 6A), suggesting that both the fusion constructs, Fld1-HA and Fld1-∆LR-HA are functional in vivo. However, we found that compared to Fld1-∆LR-HA, Fld1-HA showed partial complementation of LD phenotype of fld1∆ ldb16∆ double mutants (Fig. 6B), in agreement with previous results (Fig. 5B-D). Western blot analyses revealed expected size of ~17 kDa for Fld1-∆LR-HA compared to ~35 kDa for WT Fld1-HA, with highest expression at 5 h after galactose addition (Fig. 6C, D). Since Fld1-∆LR-HA is functional in fld1∆ ldb16∆ double mutants, we hypothesized that Fld1-∆LR-HA likely retains its oligomerizing property.
A, B Fld1-∆LR rescues LD phenotype of fld1∆, and fld1∆ ldb16∆ double mutants. Yeast fld1∆ (A) and fld1∆ ldb16∆ (B) mutants expressing either Fld1-HA or Fld1-∆LR-HA under galactose promoter on a plasmid were cultured in raffinose and shifted to galactose containing media for the indicated time, stained with BODIPY and imaged using fluorescence microscopy. Representative image shown from three independent experiments. Scale bars: 5 µm. C, D Western blot analysis. Yeast fld1∆ mutants expressing either Fld1-HA or Fld1-∆LR-HA driven by galactose promoter on a plasmid were cultured in raffinose and shifted to galactose containing media for the indicated time and subjected to Western blot analyses. PGK1 serves as a loading control. E, F Fld1-∆LR migrate as discrete oligomer. Immunoblots of fractions collected upon size exclusion chromatography of detergent (1% DDM, 0.1% CHS) solubilized membrane extracts of fld1∆ ldb16∆ cells expressing Fld1-HA (E) or Fld1-∆LR-HA (F) from a plasmid under GAL promoter. An aliquot of sample in collected fractions from a Superose 6 column were subjected to SDS-PAGE and immunoblotted for anti HA. This was repeated twice with similar results. F I and II; Fld1-∆LR-HA Western blots from two independent experiments. Source data are provided as a Source Data file.
Therefore, to determine if Fld1-∆LR also oligomerizes like WT Fld1, we purified Fld1-∆LR-HA, and Fld1-HA by affinity and size-exclusion chromatography from fld1∆ ldb16∆ cells as previously described7. Isolated membrane fractions were solubilized in the presence of detergents DDM (n-dodecyl-β-D-maltoside) and CHEMS/CHS (Cholesteryl hemisuccinate), and subjected to immuno-purification using anti-HA agarose beads, followed by size-exclusion analyses as described previously7. Fractionation by size-exclusion chromatography revealed that Fld1-HA eluted mostly in fractions (15.6 mL) corresponding to ~440 kDa (Fig. 6E), migrating close to the ferritin standard (16.16 mL) (Fig. S8A). This suggests that even in the absence of ldb16, WT Fld1 assembles into oligomeric complex as previously reported5,7,37. On the other hand Fld1-∆LR-HA eluted in fractions (17.60 mL) close to ~158 kDa (Fig. 6F), that corresponds to the soluble marker aldolase (158 kDa) fraction (17.47 mL) (Fig. S8B), and also in fractions corresponding to ~440 kDa (Fig. 6F), migrating close to the ferritin standard (440 kDa) (Fig. S8A). This suggests that a fraction of Fld1-∆LR protein assembles into oligomeric complex of small molecular weight than WT Fld1 protein. We cannot rule out whether these complexes are homo-oligomers or in complex with some other proteins.
To investigate if Fld1-∆LR forms homo-oligomers, we performed co-immunoprecipitation with two differently tagged versions of Fld1-∆LR. Therefore, we co-expressed FLAG and HA epitope tagged versions of Fld1-∆LR (Fld1-∆LR-FLAG, Fld1-∆LR-HA) at C-terminus in fld1∆ ldb16∆ double mutant cells. WT Fld1 tagged at C-terminus with FLAG and HA (Fld1-FLAG and Fld1-HA) were generated as control and co-expressed in fld1∆ ldb16∆ cells. Upon immunoprecipitation of Fld1-FLAG resulted in co-precipitation of Fld1-HA (Fig. 7A). Similarly, performing immunoprecipitation of Fld1-∆LR-FLAG resulted in co-precipitation of Fld1-∆LR-HA (Fig. 7B). This demonstrates that FLAG and HA tagged Fld1-∆LR molecules interact with each other that facilitates oligomeric complex formation. To further confirm the close proximity of individual protomers of Fld1-∆LR in an oligomeric complex in vivo, we employed a bimolecular fluorescence complementation (BiFC) or split Venus readout approach38,39. The N-terminal portion of Venus (VN) and C-terminal portion of Venus (VC) were fused to C-terminus of Fld1-∆LR (Fld1-∆LR-VN and Fld1-∆LR-VC) and were co-expressed in fld1∆ ldb16∆ double mutant cells. Similarly, VN and VC fusions with WT Fld1 (Fld1-VN and Fld1-VC) were generated as control. Close proximity of ~1-10 nm between the fusion proteins appended with the two halves of Venus, VN and VC, results in a BiFC signal due to reconstitution of full Venus (Fig. 7C-F). In both cases, WT Fld1 (Fig. 7C, D) and Fld1-∆LR (Fig. 7E, F), Venus fluorescence was reconstituted generating a detectable fluorescent signal upon excitation and displayed punctate distribution. Most of these puncta associated with MDH stained LDs. This indicates that protomers of either WT Fld1 or Fld1-∆LR establish protein-protein interactions to form oligomeric complex. Taken together, Fld1-∆LR can assemble into oligomeric complex independently of the LR or the presence of Ldb16. It remains to be determined whether oligomers of Fld1-∆LR would arrange in a ring complex formation like WT Fld1. Further cryogenic electron microscopy (Cryo-EM) studies would reveal the nature of oligomeric assembly of Fld1-∆LR.
A, B Co-immunoprecipitation analyses. Yeast fld1∆ ldb16∆ double mutants co-expressing pFld1-HA and pFld1-FLAG (A) or pFld1-∆LR-HA and pFld1-∆LR-FLAG (B) under GAL1 promoter were fractionated and anti-FLAG immunoprecipitates were enriched. Proteins were analysed by SDS-PAGE and immunoblotted against FLAG (IP), and HA (Co-IP). Pgk1 served as a loading control. Experiment was repeated twice with similar results. C−F Bimolecular fluorescence complementation (BiFC) analyses. Illustration depicting bimolecular fluorescence complementation (BiFC) in which the N-terminal fragment (VN) and the C terminal fragment (VC) of Venus both fused to C-terminus of either WT Fld1 (C) or Fld1-∆LR (E), combines to yield a full Venus fluorescent protein. Yeast fld1∆ ldb16∆ cells co-expressing either Fld1-VN and Fld1-VC (D) or Fld1-∆LR-VN and Fld1-∆LR-VC (F) were gown to mid-logarithmic growth phase, stained with MDH dye and visualized by confocal microscopy. Reconstitution of full length Venus was visualized as fluorescent puncta. Line scans of signal intensity along the red lines shown in merged panel of (D and F) is shown on right. (D) and (F) n = 3, biological replicates, Scale bars: 5 µm.
Human Seipin lacking the LR can functionally complement the absence of both Fld1 and Ldb16
We further investigated if hSeipin lacking the LR would act like yeast Fld1-∆LR. Thus, we created truncated version of hSeipin (amino acids 1-398), lacking LR, however, containing TM segment 1 (TM-1, residues 1–50), and TM segment 2, (TM-2, residues 222-398), and expressed as a N-terminal GFP fusion on an ADH1 driven plasmid (Fig. 8A, B). The construct called hSeipin-∆LR was expressed in fld1∆ and fld1∆ ldb16∆ double mutants. hSeipin-∆LR displayed punctate localization in ER like WT hSeipin (Fig. 8C). This suggests that LR of hSeipin is not required for punctate localization like Fld1-∆LR. MDH staining revealed hSeipin-∆LR marked puncta colocalized with LDs (Fig. 8C). Upon quantitative analysis we found that hSeipin-∆LR rescued the LD number and size distribution in fld1∆ and fld1∆ ldb16∆ cells (Fig. 8D, E), albeit to a lesser extent than WT hSeipin (Fig. 5C, D). However, hSeipin-∆LR restored the terbinafine growth sensitivity defect of fld1∆, and fld1∆ ldb16∆, albeit to a lesser extent than WT hSeipin (Fig. 8F). This indicates that hSeipin-∆LR can functionally complement the role of hSeipin in vivo like Fld1-∆LR.
A, B Cartoon depicting generation of hSeipin-∆LR. Illustration showing the hSeipin-∆LR lacking the luminal domain region and fused to GFP at N-terminus (A). Position of the two TM segments and a short, conserved LH/switch region connecting the two TM-regions (B) (C) hSeipin-∆LR localizes into puncta that associates with LDs. Yeast fld1∆ and fld1∆ ldb16∆ cells expressing GFP-hSeipin-∆LR on a plasmid were grown in selective media, stained with LD dye, MDH and visualized by fluorescence microscopy. White arrowheads depict puncta of hSeipin-∆LR that colocalizes with LD. Representative image shown from three independent experiments. Scale bars: 5 µm. D, E Quantification of LD number (D) and size distribution (E) in cells of the indicated genotypes. Data represent mean ± s.d. and were analysed with one-way ANOVA and Tukey’s multiple comparisons. n = 100 cells. F hSeipin-∆LR can functionally restore growth sensitivity of yeast mutants in the presence of terbinafine. Yeast fld1∆ and fld1∆ ldb16∆ double mutants expressing either GFP-hSeipin-∆LR or GFP-hSeipin were grown in SC media, diluted and spotted onto YPD plates containing either DMSO or terbinafine. Representative image shown from three independent experiments. G Generation of hSeipin-∆LR∆switch. Cartoon depicting hSeipin-∆LR devoid of LH/switch region. H hSeipin-∆LR∆switch fails to rescue fld1∆ mutant phenotype. Yeast fld1∆ cells expressing hSeipin-∆LR∆switch was grown in SC media, stained with BODIPY, and imaged by fluorescence microscopy (H). Representative image shown from three independent experiments. Scale bar: 5 µm. I, J Quantification of LD number (I) and size distribution (J) in cells of the indicated genotypes. Data represent mean ± s.d. and were analysed with two-way ANOVA and Tukey’s multiple comparisons. n = 100 cells. K Absence of switch region renders hSeipin-∆LR nonfunctional. Yeast fld1∆ mutants expressing either hSeipin or hSeipin-∆LR∆switch was cultivated in SC media, serially diluted and spotted onto plates with or without terbinafine. Representative image shown from three independent experiments. Source data are provided as a Source Data file.
Conserved locking helix is crucial for the functioning of truncated seipin
Previously it has been demonstrated that in between the two Fld1 TM segments and luminal ß-sandwich fold, there is a small helix (α3), known as locking helix (LH) or switch region that plays a crucial role in positioning and dynamics of Fld1 TM segments5,7. Replacing LH with a flexible linker destabilizes the Fld1 partner, Ldb16 and results in LD morphology defects7. This LH region contained a highly conserved G235, L236, and R237 sequence motif (Fig. S9A)5. To gain further insight into the role of LH, we hypothesized that absence of conserved elements within the LH might impair functioning of Fld1-∆LR. Hence, we generated G235A, L236A and R237A point mutants within the LH region of Fld1-∆LR and expressed in fld1∆ cells (Fig. S9B, C). Both G235A, L236A displayed punctate localization and these foci colocalized with LDs, however, R237A mutant showed few puncta that colocalized with LDs together with vaculoar staining (Fig. S9C). We found that none of the single mutant, G235A/L236A/R237A showed LD morphology defect typical of fld1∆ cells (Fig. S9C). Further, generating double mutant G235A L236A did not affect the localization of the Fld1-∆LR and these foci colocalized with LDs (Fig. S9B, C). A triple mutant G235A L236A R237A to remove the highly conserved motif altogether, resulted in ER localization of the Fld1-∆LR rather than punctate distribution, however, the triple mutant did not show LD morphology defect typical of fld1∆ cells (Fig. S9B, C), likely due to overexpression driven from ADH1 promoter. Quantitative analysis showed none of the mutant having LD number and size distribution similar to fld1∆ cells (Fig. S9D, E). Western blot analysis revealed single, double and triple mutants were expressed at a similar size like Fld1-∆LR, albeit to a lesser level (Fig. S9F). However, all the mutants rescued the terbinafine sensitivity of fld1∆ cells (Fig. S9G). These results suggest that mutating conserved GLR residues altogether within LH does not completely perturb the functioning of Fld1-∆LR.
We next investigated if removal of the switch region (residues 222-234) that contains conserved GLR sequence motif in hSeipin-∆LR would impair its functioning. The construct lacking switch region called hSeipin-∆LR∆switch was expressed in fld1∆ mutants (Fig. 8G, H). Although hSeipin-∆LR∆switch did not display punctate distribution, free mCherry was visualized inside vacuolar compartment (Fig. 8H). We found that hSeipin-∆LR∆switch failed to complement the LD morphology defect both in terms of LD number and size distribution and terbinafine growth sensitivity of fld1∆ mutants (Fig. 8H-K). Taken together, our results demonstrate that hSeipin-∆LR can functionally complement the absence of both Fld1 and Ldb16 in yeast cells like WT hSeipin and yeast Fld1-∆LR. Moreover, presence of switch region is crucial for the functioning of hSeipin-∆LR. It is likely that lack of functioning of the hSeipin-∆LR∆switch construct might be due to conformational change produced by the absence of the entire helix.
Fld1P31G R237G renders full length protein nonfunctional
Since P31G mutation in Fld1-∆LR renders it non-functional. We next investigated whether P31G mutation would also result in loss of function of WT Fld1. Therefore, we generated Fld1P31G-mCherry on a plasmid under native promoter and expressed it in fld1∆ cells and visualized LDs upon BODIPY staining. We found no distinguishable difference in the LD morphology when cells expressed either WT Fld1 or Fld1P31G, suggesting that Fld1P31G mutation in full length Fld1 does not render it non-functional (Fig. S10A). Fld1P31G also rescued the terbinafine growth defect similar to WT Fld1 (Fig. S10B). Consistent with this, a R178A point mutant, when introduced in the LR in isolation affects oligomerization, however, this mutation in the full length protein only modestly affected LD morphology and fully rescued terbinafine sensitivity as reported previously5. It is likely that since LR can oligomerize independently of the TM domain as previously reported5, single point mutation P31G within the first TM segment is not sufficient enough to impair the activity of the full length Fld1 protein.
Therefore we generated a double mutant, Fld1P31G R237G to mutate a conserved LH residue R237 together with P31 in WT Fld1 and expressed it on a plasmid as N-terminal mCherry fusion under native promoter in fld1∆ cells and visualized LDs upon BODIPY staining. Fld1P31G R237G displayed diffused ER localization rather than punctate distribution, and also showed vacuolar staining (Fig. S10C). Fld1P31G R237G failed to rescue the LD morphology defect of fld1∆ cells both in terms of LD number and size distribution (Fig. S10C-E) and showed a subtle growth rescue in presence of terbinafine only until the first serial dilution (Fig. S10F). Taken together R235 residue within the LH region when mutated together with P31 in WT Fld1 results in a synergistic effect and renders WT Fld1 non-functional.
Absence of Seipin LR results in increased TAG levels
Given the fact that Fld1-∆LR and hSeipin-∆LR, both devoid of the LR are functional, we wondered how presence or absence of this domain affected NL levels, since hydrophobic helices within the LR of hSeipin induces nanoscale clustering of TAG as described previously7,8,23,26. Therefore, we used yeast fld1∆ mutants that lack the capacity to form STE and can only synthesize TAG, are1∆ are2∆ fld1∆. We expressed plasmid borne copy of WT Fld1 or Fld1-∆LR under constitutive ADH1 promoter. NL analysis by thin layer chromatography (TLC) revealed similar TAG levels in the absence or presence of WT Fld1 (Fig. 9A-C). However, to our surprise, we found significantly elevated levels of TAG in the presence of Fld1-∆LR compared to WT Fld1 (Fig. 9B, C). This suggests that LR probably plays an important role in regulating TAG levels. However, increased TAG levels did not result in an increased number of LDs per se, and the LD number and size were indistinguishable compared with WT Fld1 (Fig. 2B, D, E). This indicates a crucial regulatory role of the LR of seipin in partitioning of TAG into LDs. Since WT Fld1 lacks HH, TAG binding property is provided by its partner, Ldb167. An elevated TAG level detected in the presence of Fld1-∆LR might perhaps indicate an increased TAG sequestration or gathering locally within the TM segments facilitated by Fld1-∆LR. In doing so the Fld1-∆LR complements the functions of both Fld1 and Ldb16. In agreement with this notion, the Fld1-∆LR restored the terbinafine sensitivity of fld1∆ ldb16∆ mutants, unlike Fld1 (Fig. 5F). To further validate our hypothesis, hSeipin and hSeipin-∆LR were expressed in are1∆ are2∆ fld1∆ mutants and TAG levels were determined by TLC. Intriguingly, presence of hSeipin-∆LR resulted in significant increase in TAG levels when compared to hSeipin (Fig. 9C, D). This suggests that Fld1-∆LR/hSeipin-∆LR can efficiently sequester TAG independent of the presence of LR. Next, we tested if Fld1-ΔLRP31G would result in loss of potential TAG accumulation activity of Fld1-ΔLR. Hence, Fld1-ΔLRP31G was expressed in are1∆ are2∆ fld1∆ cells and TAG levels were estimated by TLC. We found that Fld1-ΔLRP31G resulted in significantly reduced TAG levels compared to Fld1-ΔLR (Fig. 9E, F). This indicates that P31G mutation results in abolishing TAG concentration activity of Fld1-ΔLR. Collectively, our results indicate that both luminal and TMD region of seipin contribute independently to LD formation, and LR is important in regulating TAG levels.
A−D Increased TAG levels in the presence of truncated seipin lacking luminal domain region. Lipid extracts from yeast are1∆ are2∆ fld1∆ mutants (A), expressing either WT Fld1 or Fld1-∆LR (B), or expressing either hSeipin or hSeipin-∆LR (D) were resolved by thin layer chromatography (TLC). Neutral lipid separation profile from two independent experiments is shown (I & II). TAG, triacylglycerol; FFA, free fatty acids; DAG, diacylglycerol; PL, phospholipids. C Quantification of steady state TAG levels from three independent experiments is shown. Data represent mean ± s.d. and was analysed using two-tailed unpaired t-test. E, F Fld1-ΔLRP31G results in loss of TAG accumulation activity of Fld1-ΔLR. Lipid extracts from yeast are1∆ are2∆ fld1∆ mutants expressing either WT Fld1, or Fld1-∆LR, or Fld1-ΔLRP31G were resolved by thin layer chromatography (TLC). Neutral lipid separation profile from two independent experiments is shown (I & II) (F). TAG, triacylglycerol; FFA, free fatty acids; DAG, diacylglycerol; PL, phospholipids. G Quantification of steady state TAG levels from three independent experiments is shown. Data represent mean ± s.d. and was analysed using two-tailed unpaired t-test. G Stepwise model of LD biogenesis at ER subdomains established by Fld1-∆LR. Step I: Fld1-∆LR localizes to ER subdomains independently of LDs. Step II: Fld1-∆LR recruits LD biogenesis machinery including factors to prime these ER sites. Step III: Localized TAG synthesis and droplet assembly at Fld1-∆LR demarcated ER sites. Growing LD acquires LD-resident proteins and mature while remaining associated with Fld1-∆LR puncta. Source data are provided as a Source Data file.
Discussion
Seipin has emerged as a crucial LD biogenesis factor, however, the molecular mechanisms of its action is not completely understood. Accumulating evidence based on cryo-EM deduced structure and in-silico modelling data indicate that seipin’s luminal and TM regions play critical roles in LD formation with LR participating in TAG sequestration due to presence of HH within this region7,8,23 and TM contributing to TAG clustering within the ER membrane26. However, yeast seipin lacks HH, the function of which is provided by its partner Ldb16, thereby increasing local TAG concentration within the proximity of seipin complex7. Despite these advances, little is known about how seipin TM segments contribute to LD biogenesis. Here, we identified the key determinants within seipin that is essential for its function in creating functional LD biogenesis sites in the ER. We demonstrate that seipin mutants, Fld1-∆LR in yeast, and the human equivalent hSeipin-∆LR, devoid of LR represents the minimal region in seipin that is necessary and sufficient to determine functional sites of LD biogenesis.
Based on our data, we propose a model in which Fld1-∆LR comprising of both TM-1 and TM-2 segments together with the conserved LH region, and short cytosolic N- and C-terminal regions can localize to ER subdomains independently of the presence of LDs (Fig. 9G, step-I). These artificial puncta are stable and define sites that recruits LD biogenesis machinery, including factors, Nem1, Ldb16, Pex30, Yft2 and become enriched in DAG upon induction of LD formation. (Fig. 9G, step-II). At these ER subdomains, TAG-synthase localize and catalyze TAG production locally resulting in formation of NL lenses (Fig. 9G, step-II). Fld1-∆LR together with LD biogenesis factors facilitate TAG gathering locally at these ER subdomains and thereby preventing outward diffusion of NL from these sites and hence driving the formation of nascent LDs that grows and emerges towards the cytoplasm (Fig. 9G, step-III).
Fld1-∆LR can functionally complement the LD morphology defects and terbinafine growth sensitivity of fld1∆ and fld1∆ ldb16 double mutants (Figs. 2I, 5F). We do not yet know what determines punctate localization of Fld1, however, our results demonstrate that the first 50 residues of Fld1 comprising TM-1 segment is sufficient for punctate localization of Fld1-∆LR, though when TM-1 is expressed in isolation these foci are non-functional in restoring LD morphology defects typical of fld1∆ cells (Fig. S2A–D). Consistent with this, mutating 31st proline to glycine (P31G) within the TM-1 region perturbs the punctate distribution of Fld1-∆LR and abolishes the functioning of Fld1-∆LR in restoring LD morphology defect and terbinafine sensitivity (Fig. 5G–K).
Our data support the notion that LR per se is not required for punctate localization. Moreover, the LR appears to be dispensable in organizing ER subdomains permissive for droplet assembly. Fld1-∆LR devoid of LR resulted in elevated TAG levels, indicating crucial regulatory role of this domain in regulating local TAG sequestration and partitioning. The LR of yeast seipin and of other organisms form homo-oligomeric ring7,20,21, we demonstrate that Fld1-∆LR can oligomerize independently of the LR and partner protein Ldb16. Future studies should focus on deciphering the structure of Fld1-∆LR oligomeric complex by cryo-EM. Previously it has been reported that interactions between Fld1 TMs is important for the stability and functioning of Ldb167. However, the fact that Fld1-∆LR can restore the LD morphology defect and terbinafine sensitivity of fld1∆ ldb16∆ double mutants (Fig. 5B–D, F), unlike Fld1 alone suggests that Fld1-∆LR can functionally take over the roles of both proteins, Fld1 and Ldb16. We do not yet know the mechanisms for this, which should be pursued in future studies. An interesting hypothesis to investigate in the future would be to determine how presence of LR and/or its oligomerization affects the activity of Fld1 species. We found that in the presence of endogenous Fld1, Fld1-∆LR is mislocalized, a likely reason for it being non-functional in ldb16∆ mutants (Fig. S5). However, when endogenous Fld1 is absent, i.e. in the case of fld1∆ and fld1∆ ldb16∆ double mutants, Fld1-∆LR localizes to LD biogenesis sites and functionally complements the absence of seipin. This indicates that perhaps platforms/domains for LD biogenesis are pre-defined, and presence or absence of endogenous seipin can interfere with the localization of Fld1-∆LR at these domains.
Given the fact that Fld1-∆LR can form oligomeric complex, the likely gain of function of Fld1-∆LR could be attributed to increased TAG levels in the presence of Fld1-∆LR oligomeric complex (Fig. 9B, C), hence suggestive of enhanced TAG gathering/interactions locally within the TMD region in the absence of LR. In such a scenario, Fld1-∆LR could effectively take over the TAG sequestration functions of both Fld1 and Ldb16. Since the number of monomers of seipin in a complex vary between different species, it suggests that the oligomeric complex of seipin is flexible to accommodate different number of protomers. It is also likely that in the absence of LR, certain LD biogenesis factors might associate more tightly with the oligomeric assembly of Fld1-∆LR thereby enhancing TAG gathering, and droplet assembly. Consistent with this, mutating tyrosine residue at position 171 to alanine in hSeipin, resulted in compromised oligomerization in vitro, however, the mutant hSeipinY171A still formed foci and rescued seipin deficiency phenotype in cells. This indicates that other region of the protein, likely the TMD region might contribute to oligomerization and functionality in cells21. In agreement with this, MDS studies have shown that Fld1 TMs, unlike its LR are able to interact with TAG7. Consistent with this, hSeipin-∆LR devoid of LR rescued LD morphology defect and terbinafine sensitivity of fld1∆ ldb16∆ mutants like hSeipin (Fig. 8C–F) and displayed significant increase in TAG levels in comparison to hSeipin (Fig. 9C, D). However, P31G mutation abolished the TAG concentration activity of Fld1-ΔLR (Fig. 9E, F). Taken together, this indicates that Fld1-∆LR/hSeipin-∆LR can effectively trap/sequester TAG within the TM region independently of the LR in the ER membrane and thereby facilitate LD biogenesis.
Protease protection assay suggested that Fld1-∆LR adopts a similar topology to WT Fld1 (Fig. S3). Fld1-∆LR marked ER sites become enriched in LD biogenesis factors, and at these sites, DAG accumulates upon stimulation of LD formation (Fig. 4). These foci become stably docked with TAG-synthesizing enzyme, sLro1, when engaged in LD formation, hence suggesting that these platforms mimic native Fld1 demarcated ER subdomains (Fig. 4). Our data indicate that the conserved LH or switch region is crucial for the stability of the hSeipin-∆LR and probably facilitates the proper orientation of the two TMs inside the ER bilayer. Abrogation of the conserved LH region, hSeipin-∆LR∆switch failed to localize into puncta in the ER and was unable to complement fld1∆ phenotype (Fig. 8H–K). Hence, LH is important for proper functioning of the hSeipin-∆LR.
In conclusion, our findings support the notion that the LR of seipin is dispensable in defining functional LD biogenesis sites in the ER. Our data provide mechanistic insights into the role of TMs in priming ER subdomains and orchestrating the recruitment of LD assembly machinery at these sites for efficient droplet formation.
Methods
Yeast strains and plasmids
Yeast strains and plasmids used in this study are listed in Supplementary Information. Yeast single, double, and triple-deletion mutants were created by PCR-based targeted homologous recombination by replacing the ORF of the gene of interest with a gene knock-out cassette followed by a marker rescue strategy32,40. Chromosomally C-terminal tagged Erg6-GFP fusion protein was constructed using PCR-based homologous recombination41. Plasmids constructs used in the study were based on pRS415-LEU, pRS416-URA, pBS34-KanMX, pGREG575-LEU, pFA6a-GFP(S65T)-His3MX6 and listed in supplementary information. To construct pADH1-Fld1-TMD1-mCherry (amino acids 1−50), or pADH1-Fld1-TMD2-mCherry (amino acids 235−285), corresponding regions of yeast Sei1/Fld1 were amplified by PCR from S. cerevisiae genomic DNA and inserted into either pRS415/416 thereby expressing mCherry fused to the TMD1 /TMD2 region of Fld1 under the ADH1 promoter. For plasmid pADH1-Fld1-∆LR-mCherry, sequence corresponding to amino acids 1−50 and 235−285 of Fld1 were fused to mCherry and inserted into either pRS415/416 to express the Fld1-∆LR under ADH1 promoter. pADH1-Fld1-mCherry was generated by PCR amplification of full length Fld1 sequence from yeast genomic DNA, and fused to mCherry sequence amplified from pBS34, and inserted into pRS415/416 vectors having LEU/URA selection markers. PCR mediated site directed mutagenesis was performed to generate Fld1-∆LRG235A, Fld1-∆LRL236A, Fld1-∆LRR237A, Fld1-∆LRG235A L236A, Fld1-∆LRG235A L236A R237A, Fld1-∆LR P31G and cloned into pRS416 under ADH1 promoter having C-terminal mCherry fusion. Full length Fld1, and Fld1-∆LR were expressed from seipin endogenous promoter (459 bp upstream of start codon of FLD1) cloned into plasmid pGREG575-GFP/mCherry and pGREG503 by replacing GAL1 promoter using Sac1 and Spe1. Fld1P31G, and Fld1P31G R237G were generated using PCR mediated site directed mutagenesis where Fld1P31G was cloned under seipin endogenous promoter having C-terminus mCherry fusion into pGREG503, and Fld1P31G R237G was cloned under seipin endogenous promoter having N-terminus mCherry fusion into pGREG575. For oligomerization studies, full length Fld1 and Fld1-∆LR were C-terminally tagged with HA epitope tag (amplified using pFA6a-3HA-HIS3MX6) and expressed under GAL1 promoter in plasmid pGREG505. For Co-IP studies, full length Fld1 and Fld1-∆LR were C-terminally tagged with FLAG epitope tag (amplified using pFA6a-5FLAG-natMX6) and expressed under GAL1 promoter in plasmid pGREG506. Plasmids for Split Venus system (BiFC) were constructed using plasmid pKT90 (pFA6a-link-yEVenus-SpHIS5) as template. For Venus-N (VN) amino acids 1−172 were used and for Venus-C (VC) amino acids 155−238 were used as C-terminal fusion proteins with Fld1 and Fld1-∆LR, expressed under ADH1 promoter in plasmids pRS416 and pRS415 respectively. pADH1-GFP-LiveDrop was constructed using plasmid pKeima-LiveDrop42, a kind gift from Tobias Walther (Addgene plasmid #199695; RRID:Addgene_199695). LiveDrop was PCR amplified with BamHI and XhoI restriction sites and ligated in plasmid pGREG575 (with replaced ADH1 promoter) to construct pADH1-GFP-LiveDrop.
The plasmids encoding Yft2-sf-GFP, and the ER-DAG sensor (pADH1-ER-DAG sensor), and pGAL1-GFP-FFAT-sLro1 have been described previously9,35,43. We used plasmids expressing GFP fused to Sec63 to label the ER. Plasmid expressing Sec63-TMDs (1-250)-mCherry-LEU was constructed by amplifying 1-250 base pairs from Sec63 coding sequence using yeast genomic DNA fused with mCherry and cloned in pRS415-LEU. Plasmids pADH1-GFP-LEU was constructed by replacing GAL1 promoter from pGREG575 by performing digestion using Sac1 and Not1 restriction sites and inserting sequence corresponding to ADH1 promoter amplified from pRS416 plasmid by using gap repair. Plasmids encoding LD biogenesis factors Ldb16, Pex30, and Nem1 tagged with C-terminal GFP were generated by amplifying respective genes using yeast genomic DNA and fused to GFP sequence amplified from pFA6a-GFP(S65T)-His3MX6 and inserted into either pRS415-LEU or pRS416-URA. For cloning hSeipin, mRNA was isolated from HEK293T cells using TRIzol reagent as per manufacturer’s information (T9424; Sigma-Aldrich). cDNA was synthesised using RevertAid First Strand cDNA Synthesis Kit (K1621; Thermo Fisher Scientific). 50−100 ng of cDNA was used as a template to amplify hSeipin having BamH1 and Xho1 restriction sites in the forward and reverse primers respectively. hSeipin was cloned into pADH1-GFP-LEU using BamH1 and Xho1 restriction sites having N-terminal GFP tag. Truncated hSeipin (hSeipin-∆LR) was constructed by amplifying N-terminal region comprising TMD1 (amino acids 1−50) and C-terminal region comprising TMD2 (amino acids 222-398) using the full length hSeipin construct (described above), using an overlap extension PCR both TMD1 and TMD2 of hSeipin were stitched together and was further cloned into pADH1-GFP-LEU having GFP tag at N-terminus using BamH1 and Xho1 restriction sites. hSeipin -∆LRΔswitch devoid of LH/switch region comprised of TMD1 (amino acids 1–50) and TMD2 (amino acids 235-398) were fused to mCherry and inserted into pRS416 to express the hSeipin -∆LRΔswitch under ADH1 promoter. All constructs were verified by sequencing.
Media and growth conditions
WT and mutant yeast cells were grown at 30 °C in rich YPD medium (1% Bacto yeast extract, 2% Bacto Peptone, 2% glucose) unless otherwise indicated. Cells bearing plasmids were grown in synthetic complete (SC) media containing 6.7 g/l yeast nitrogen base without amino acids (United States Biologicals), an amino acid mix (Sigma Aldrich), containing either 2% glucose, galactose or raffinose. Yeast cells were cultured overnight at 30 °C, 200 rpm. Cells were diluted to an OD600 = 0.2 and log phase cells were collected at OD600 = 0.6-0.8. To grow cells in presence of fatty acids for induction of LD formation, SC media were supplemented with 0.1% oleic acid (O1383; Sigma Aldrich) in the presence of 1% Brij-58 (P5884; Sigma Aldrich).
Fluorescence microscopy
Live cell imaging was performed using inverted fluorescence microscope, Zeiss Axio Observer 7 (Carl Zeiss, Germany) equipped with a sCMOS pco.edge 4.2 camera and Nikon Eclipse Ti-2E inverted widefield fluorescence microscope equipped with Prime BSI Scientific sCMOS camera (Teledyne Photometrics). Images were captured using a plan apochromatic 100X, 1.42 NA oil immersion objective and a DAPI, FITC or TXRED filter set, taken as single slice at approximately mid-plain (unless indicted in the figure legends) and deconvolved using Zen software (version 3.8) at regularised inverse filter settings, or Nikon NIS elements (version 5.41.00) using Richardson Lucy deconvolution settings. For Z-stacks, five Z-sections were taken with a step size of 0.25 µm, 3D deconvolved using Richardson Lucy deconvolution settings (Nikon NIS elements version 5.41.00) and maximum intensity projected using Fiji (NIH). Single plane images are displayed. Images being directly compared were obtained using identical microscope settings. Time-lapse images were acquired every 1 s for 10 s with an exposure of 5000 ms and deconvolved. Live cell imaging for Bimolecular fluorescence complementation (BiFC) was performed using Leica TCS SP8 confocal laser scanning microscope (Leica Microsystems). Images were acquired using a plan apochromatic 100X/1.42 N.A. oil immersion objective lens with UV diode laser (405 nm) and argon laser at YFP excitation-emission (510−550 nm) wavelength. Images were captured as z-stacks taken with a step size of 0.25 µm, processed using LasX software (Leica microsystems) and Fiji where, representative images are maximum intensity projections. Brightness and contrast were adjusted using Adobe Photoshop 2022 (Adobe Systems). Colocalization between GFP and mCherry marked punctae in acquired images was scored manually. Values were recorded in Microsoft Excel and analysed in Prism (Version 10.2.1) (GraphPad Software) (GraphPad, La Jolla, CA).
Colocalization analysis
Acquired images were analyzed for colocalization using Pearson’s correlation coefficient using the BIOP JACoP plugin in Image J/Fiji. Briefly, merged images were fed into fiji and BIOP JACoP plugin was used at Costes Auto-Threshold for channels A, B and C. Pearson’s coefficient was recorded for ≥ 25 cells, where obtained values indicate the association between fluorescence signals (0 = no correlation, 1 = max correlation).
Staining to visualize lipid droplets in yeast
To visualize LDs in yeast, cells expressing fluorescently tagged LD marked protein, were grown in YPD/SC media to mid log phase and stained with either 0.5 µg/ml BODIPY 493/503 (D3922; Invitrogen) or 0.1 mM MDH AutoDOT (SM1000a; Abcepta) and incubated for 5 min at RT. Cells were collected at 2000 rpm for 2 min and washed twice with 1 X PBS. Cells were resuspended in 50 µL of 1X PBS and visualised using FITC (for BODIPY) and DAPI (for MDH) excitation filters of Nikon Eclipse Ti-2E inverted widefield fluorescence microscope.
Protein expression and purification
Expression and purification of Fld1 and Fld1-ΔLR was adapted and modified from previously described methods7. Yeast fld1Δ ldb16Δ mutants was transformed with either pGAL-Fld1-HA or pGAL-Fld1-∆LR-HA plasmid. Cells were cultivated with 2% raffinose, harvested and induced in the presence of 2% galactose for 5 h in 1-L culture flasks. Cells were harvested by centrifugation at 4000 g for 5 min. Cells were once washed with milliQ water and ~10 g of cell pellet was resuspended in 10 mL Buffer A (50 mM) Tris.HCl (pH 7.5), 200 mM NaCl, 1 mM EDTA (pH 8.0) with 1 mM PMSF (Roche) and 1.5 µM of Pepstatin A (Thermo Fisher), and lysed in the presence of 0.5 mm glass beads (Sigma) using Precellys Evolution Homogenizer (Bertin Technologies, Germany) with cycle 2 X 40 s at 8000 rpm with incubation on ice for 10 min between subsequent cycles. Cell homogenate/lysate was harvested by centrifugation at 18,000 g for 20 sec, and was pre-cleared at 5000 g for 10 min at 4 °C. Membranes were isolated using high speed centrifugation at 185,000 g using SW 41 Ti rotor at 4 °C for 1 h. Pelleted membranes were solubilised and incubated in 25 mL buffer A containing 1% (w/v) DDM (n-dodecyl-β-D-maltoside) (Avanti polar lipids) and 0.1% (w/v) CHEMS/CHS (Cholesteryl hemisuccinate) (Avanti polar lipids) for 1 h at 4 °C. Next, the solubilised membranes were centrifuged again at 185,000 g for 1 h at 4 °C and the supernatant was incubated with 1 mL of equilibrated anti-HA agarose (Pierce, Thermo Scientific) overnight (using rotator) at 4 °C. After incubation, the material was transferred to gravity flow columns and washed with 5 column volume of buffer A containing 0.015% (w/v) DDM and 0.0015% (w/v) CHS (Wash buffer). Proteins were eluted using HA peptide (Pierce, Thermo Scientific) first using 3 X 1 mL fractions containing 0.5 µg/mL of HA peptide, second using 2 X 1 mL of 0.2 mg/mL and lastly 1 X 1 mL fraction of 1 mg/mL of HA peptide. Eluted proteins were concentrated using 30 kDa cut off centrifugal protein concentrators (Pierce, Thermo Scientific), until the volume reached below 1 mL. The concentrated protein was run with an Akta Pure (SEC) (Cytiva) over a 24 mL Superose 6 Increase 10/300 GL size exclusion column in wash buffer at 0.3 ml/min flow rate. Fractions of 500 µL were collected and analysed by SDS-PAGE followed by Western blotting. Proteins were detected using anti-HA primary antibody at 1:1000 dilution (Cell signalling), and an anti-rabbit HRP conjugated secondary antibody (Bio-rad) at 1:3000 dilution.
Co-IP of Fld1 and Fld1-∆LR
Co-IP of Fld1 and Fld1-∆LR was performed using Pierce crosslink magnetic IP/Co-IP kit (Thermo Scientific) and using protocol adapted and modified from previously described method44. Briefly, yeast fld1Δ ldb16Δ mutant was co-transformed with pGAL-Fld1-HA and pGAL-Fld1-FLAG (for full length) and pGAL-Fld1-∆LR-HA and pGAL-Fld1-∆LR-FLAG (for Fld1-∆LR). Cells were cultivated with 2% raffinose, harvested and induced in the presence of 2% galactose. 350 O.D. units were harvested after 2.5 h of galactose induction. Cells were washed once with MilliQ and resuspended in 1 mL of lysis buffer (25 mM Tris, 150 mM NaCl, 1 mM EDTA, 0.25% NP40, 5% glycerol with 1 mM PMSF and 1.5 µM of Pepstatin A). Cells were homogenized as described previously, and lysate was collected. Lysate was incubated overnight at 4 °C on a rotator with DSS crosslinked FLAG antibody-protein A/G magnetic beads. After incubation, 3 washings were performed using lysis buffer to remove non-specific background binding, one wash was given using milliQ and proteins were eluted using low pH elution buffer and neutralised with neutralisation buffer supplied with the kit. Proteins were analysed by SDS-PAGE followed by western blotting, detected using anti-HA primary antibody at 1:1000 dilution, anti-FLAG primary antibody at 1:1000 dilution (#14793 Cell signalling), anti PGK1 primary antibody at 1:5000 dilution (Invitrogen), anti-rabbit HRP conjugated secondary antibody at 1:3000 dilution and an anti-mouse HRP conjugated secondary antibody at 1:10,000 dilution.
Yeast LD number and area quantification
LD number quantification was scored manually using Fiji by counting the number of LDs in 50 cells in duplicates. LD size was analysed using Zen blue edition (Version 3.8) and Nikon NIS elements version (5.41.00) by manually tracing the perimeter of LDs. Data was exported to Excel (Microsoft) and sorted on high to low LD area (in µm2) and analysed in GraphPad Prism (Version 10.2.1) (GraphPad, La Jolla, CA).
Neutral lipid analysis by thin layer chromatography
Cells were grown in SC media with shaking at 30 °C for 16 h until early stationary growth phase. 50 OD600 units of cells were harvested by centrifugation, washed in MQ water, and stored at −80 °C until further processing. Cell pellet was resuspended in 600 µl MQ water and lysed in the presence of glass beads using a Precellys Evolution Homogenizer (Bertin Technologies, Germany) as previously described9 The homogenate was transferred to a fresh tube, and total protein was estimated using the BCA method. Volume equivalent to 1 mg of protein was subjected to lipid extraction using modified version of Folch extraction method45. Briefly, 600 µl of cell lysate was added to 600 µl of chloroform: methanol (2:1) and mixed by vortexing for 5 min. Then, 300 µl of 0.9% NaCl was added to induce phase separation, vortexed, and centrifuged at 8500 rpm for 5 min at 4 °C. The lower organic phase was transferred to a new tube. The aqueous phase was re-extracted with 600 µl of chloroform: methanol (2:1), organic phases were pooled followed by washing with 2 M KCl. Lipid extracts were dried under the stream of N2 gas and dissolved in chloroform: methanol (1:1). The lipids were spotted onto silica gel 60 thin layer chromatography (TLC) plates (Merck, Darmstadt, Germany), along with standards and developed with hexane-diethylether-acetic acid (70:30:2)46. The TLC plate was air dried and sprayed with 10% Cupric acetate (wt/vol) in 8% phosphoric acid, dried, and charred in an oven at 145 °C for 1 h to visualize the lipid spots36. Migration front of lipids was compared to that of standard, and spots were quantified using FIJI (NIH). Data were analysed in GraphPad Prism (Version 10.2.1) (GraphPad, La Jolla, CA).
Differential fractionation, membrane association and topology
Differential fractionation, determination of membrane association, and topology of WT and chimeric seipin were performed as previously described31,47. Briefly, cells were grown in SC media at 30 °C for 16 h until early stationary growth phase. 300 OD600 units of cells were harvested by centrifugation, resuspended in 15 ml of 100 mM Tris-Cl pH 9.4 containing 10 mM DTT, and incubated at 30 °C for 20 min. Then cells were centrifuged, and pellet was resuspended in 7.5 ml of spheroplasting buffer (0.6 M sorbitol,10 mM KPi pH 7.5 containing 225 µl 1X zymolase), incubated at 30 °C for 40 min and again centrifuged. The pellet was resuspended in 12 ml lysis buffer (0.2 M sorbitol, 50 mM KOAc, 20 mM HEPES pH 6.8, 2 mM EDTA, protease inhibitor cocktail) and Dounce homogenised at least 20 times on ice. The lysate was cleared (soup was taken) and again Dounce homogenization was repeated with the pellet and supernatant fractions were pooled. This homogenate (H) fraction was subjected to differential centrifugation after keeping an aliquot of 1 ml for Western Blot analysis. First it was centrifuged at 13,000 g for 45 min at 4 °C. The pellet (P13 fraction) was resuspended in 1 ml ice cold lysis buffer and store at −20 °C until use. The supernatant was centrifuged at 21,000 g for 1 h. The pellet (P21 fraction) was dissolved in 250 µl ice cold lysis buffer and the supernatant (S21 fraction) were stored at −20 °C until use. Protein concentrations were determined using BCA method and BSA as standard. P13 fraction was subjected to membrane association, and topology assessment analyses. Proteins were precipitated with 10% trichloroacetic acid (TCA), resuspended in sample buffer and separated by SDS-polyacrylamide gel electrophoresis (SDS-PAGE) and analysed by Western blot.
Analysis of fusion proteins by Western blot
Cells were grown in YPD or SC media at 30 °C until early stationary growth phase. 3 OD600 units of cells were harvested by centrifugation. Total proteins were extracted using NaOH, precipitated using TCA, and resuspended in sample buffer as previously described48. Total proteins from cells and eluted fraction from size exclusion chromatography were separated using SDS-PAGE and analysed by Western blotting. The blots were probed with antisera against GFP (mouse, 1:2000 dilution, HRP, Invitrogen), mCherry (rabbit, 1:1000, # E5D8F, Cell Signalling Technology), Dpm1 (mouse, 1:1000, # A-6429, Invitrogen), Pgk1 (mouse, 1:5000, #459250, Invitrogen), Kar2 (mouse, 1:2000, MA5-27686, Invitrogen), anti-HA (rabbit, 1:1000, # 3724 Cell signalling), and using goat anti rabbit IgG HRP (1:3000 dilution, # 1706515; Bio-Rad) and goat anti mouse IgG HRP (1:5000 dilution, # 31430, Invitrogen). The blots were detected using PierceTM ECL Western Blotting Substrate (Thermo Fisher Scientific) and imaged in FluorChem E (Protein Simple).
Drug sensitivity assay
Cells were grown in either YPD, or SC media at 30 °C until early stationary growth phase. Six-fold serial dilutions were made using a 96 well microtiter plate and stamped onto YPD plates with and without Terbinafine (100 µg/ml) (Sigma-Aldrich), using a replica plater5. The plates were incubated at 30 °C for 3−5 days and imaged using FluorChem E (Protein Simple). Brightness and contrast were adjusted using Adobe Photoshop 2022 (Adobe Systems).
Electron microscopy of yeast
Sample preparation for EM was done as described previously43. Yeast cells were grown to early stationary growth phase, and 10 OD600 units of cells were harvested and fixed in 1 ml fixative media (1% glutaraldehyde, 0.2% paraformaldehyde, and 40 mM potassium phosphate, pH 7.0) for 10 min at RT. Cells were harvested and again resuspended in 1 ml fresh fixative media and incubated on ice for 50 min. Cells were centrifuged and washed twice with 0.9% NaCl and once with water. Cells were incubated with 2% solution of KMnO4 for 5 min at RT, centrifuged, and again resuspended with fresh solution of 2% KMmO4 for 45 min at RT for en bloc staining. The cells were then dehydrated using graded series of ethanol (30%, 50%, 70%, 80%, 90%, 95%, and 100%) for 10 min each, with two more incubations in 100% ethanol from a freshly opened bottle. The samples were subsequently embedded stepwise using Spurr’s low-viscosity resin (EMS). Samples were infiltrated for 2 h each with a 3:1, 1:1, 1:3 ethanol / embedding media mixture. Cells were incubated overnight with 100% fresh resin. The next day, cells were again resuspended in fresh 100% resin for 2–3 h, transferred into BEEM capsules (EMS), and polymerized at 70 °C for 4 days. Semi- and ultrathin sections were produced with a diamond knife (Diatome) on an ultramicrotome (Ultracut UCT; Leica Microsystems), collected on 200 mesh copper grids (EMS), poststained with uranyl acetate and lead citrate, and visualized with a Talos S200 transmission electron microscope (TEM; Thermo Fisher Scientific), operating at 200 kV. Pictures were recorded on a below-mounted 4k × 4k BM-Ceta (CMOS) camera.
Statistics and reproducibility
Unless otherwise stated, data were obtained from at least three independent experiments. The images presented are representative of the results obtained. Quantitative results are expressed as mean ± standard deviation. Statistical analysis was performed with unpaired, two-tailed Students’ t-tests using GraphPad Prism (Version 10.2.1) (GraphPad, La Jolla, CA). Ordinary one-way ANOVA and two-way ANOVA were performed with Holm-Sidak’s multiple comparisons test applied (where indicated). For both t tests and ANOVA, p-values < 0.05 were considered significant.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The authors declare that the data supporting the findings of this study are contained with the paper and its supplementary information files. Source data are provided with this paper.
References
Zadoorian, A., Du, X. & Yang, H. Lipid droplet biogenesis and functions in health and disease. Nat. Rev. Endocrinol. 19, 443−459 (2023).
Henne, W. M., Reese, M. L. & Goodman, J. M. The assembly of lipid droplets and their roles in challenged cells. EMBO J. 37, e98947 (2019).
Greenberg, A. S. et al. The role of lipid droplets in metabolic disease in rodents and humans. J. Clin. Invest 121, 2102–2110 (2011).
Kumari, R. M., Khatri, A., Chaudhary, R. & Choudhary, V. Concept of lipid droplet biogenesis. Eur. J. Cell Biol. 102, 151362 (2023).
Arlt, H. et al. Seipin forms a flexible cage at lipid droplet formation sites. Nat. Struct. Mol. Biol. 29, 194–202 (2022).
Chung, J. et al. LDAF1 and Seipin form a lipid droplet assembly complex. Dev. Cell 51, 551–563.e557 (2019).
Klug, Y. A. et al. Mechanism of lipid droplet formation by the yeast Sei1/Ldb16 Seipin complex. Nat. Commun. 12, 5892 (2021).
Prasanna, X. et al. Seipin traps triacylglycerols to facilitate their nanoscale clustering in the endoplasmic reticulum membrane. PLoS Biol. 19, e3000998 (2021).
Choudhary, V., El Atab, O., Mizzon, G., Prinz, W. A. & Schneiter, R. Seipin and Nem1 establish discrete ER subdomains to initiate yeast lipid droplet biogenesis. J. Cell Biol. 219, e201910177 (2020).
Schneiter, R. & Choudhary, V. Seipin collaborates with the ER membrane to control the sites of lipid droplet formation. Curr. Opin. Cell Biol. 75, 102070 (2022).
Thiam, A. R. & Ikonen, E. Lipid droplet nucleation. Trends Cell Biol. 31, 108–118 (2021).
Renne, M. F., Klug, Y. A. & Carvalho, P. Lipid droplet biogenesis: a mystery “unmixing”?. Semin Cell Dev. Biol. 108, 14–23 (2020).
Salo, V. T. et al. Seipin facilitates triglyceride flow to lipid droplet and counteracts droplet ripening via endoplasmic reticulum contact. Dev. Cell 50, 478−493.e9 (2019).
Fei, W. et al. Fld1p, a functional homologue of human seipin, regulates the size of lipid droplets in yeast. J. Cell Biol. 180, 473–482 (2008).
Szymanski, K. M. et al. The lipodystrophy protein seipin is found at endoplasmic reticulum lipid droplet junctions and is important for droplet morphology. Proc. Natl. Acad. Sci. USA 104, 20890–20895 (2007).
Wang, H. et al. Seipin is required for converting nascent to mature lipid droplets. Elife 5, e16582 (2016).
Salo, V. T. et al. Seipin regulates ER-lipid droplet contacts and cargo delivery. EMBO J. 35, 2699–2716 (2016).
Magre, J. et al. Identification of the gene altered in Berardinelli-Seip congenital lipodystrophy on chromosome 11q13. Nat. Genet 28, 365–370 (2001).
Rao, M. J. & Goodman, J. M. Seipin: harvesting fat and keeping adipocytes healthy. Trends Cell Biol. 31, 912–923 (2021).
Yan, R. et al. Human SEIPIN binds anionic phospholipids. Dev. Cell 47, 248–256.e244 (2018).
Sui, X. et al. Cryo-electron microscopy structure of the lipid droplet-formation protein seipin. J. Cell Biol. 217, 4080−4091 (2018).
Lundin, C. et al. Membrane topology of the human seipin protein. FEBS Lett. 580, 2281–2284 (2006).
Zoni, V. et al. Seipin accumulates and traps diacylglycerols and triglycerides in its ring-like structure. Proc. Natl. Acad. Sci. USA 118, e2017205118 (2021).
Renne, M. F., Corey, R. A., Ferreira, J. V., Stansfeld, P. J. & Carvalho, P. Seipin concentrates distinct neutral lipids via interactions with their acyl chain carboxyl esters. J. Cell Biol. 221, e202112068 (2022).
Gao, Q. et al. Pet10p is a yeast perilipin that stabilizes lipid droplets and promotes their assembly. J. Cell Biol. 216, 3199–3217 (2017).
Kim, S. et al. Seipin transmembrane segments critically function in triglyceride nucleation and lipid droplet budding from the membrane. Elife 11, e75808 (2022).
Sandager, L. et al. Storage lipid synthesis is non-essential in yeast. J. Biol. Chem. 277, 6478–6482 (2002).
Cartwright, B. R. et al. Seipin performs dissectible functions in promoting lipid droplet biogenesis and regulating droplet morphology. Mol. Biol. Cell 26, 726–739 (2015).
Wang, C. W., Miao, Y. H. & Chang, Y. S. Control of lipid droplet size in budding yeast requires the collaboration between Fld1 and Ldb16. J. Cell Sci. 127, 1214–1228 (2014).
Chaudhary, R. & Choudhary, V. Protocol to determine the topology of integral endoplasmic reticulum membrane protein in Saccharomyces cerevisiae. STAR Protoc. 6, 103727 (2025).
Jacquier, N. et al. Lipid droplets are functionally connected to the endoplasmic reticulum in Saccharomyces cerevisiae. J. Cell Sci. 124, 2424–2437 (2011).
Khatri, A. & Choudhary, V. Protocol to induce de novo lipid droplets in Saccharomyces cerevisiae. STAR Protoc. 4, 102732 (2023).
Grippa, A. et al. The seipin complex Fld1/Ldb16 stabilizes ER-lipid droplet contact sites. J. Cell Biol. 211, 829–844 (2015).
Joshi, A. S. et al. Lipid droplet and peroxisome biogenesis occur at the same ER subdomains. Nat. Commun. 9, 2940 (2018).
Choudhary, V. et al. Architecture of lipid droplets in endoplasmic reticulum is determined by phospholipid intrinsic curvature. Curr. Biol. 28, 915–926.e919 (2018).
Rogers, S. et al. Triglyceride lipolysis triggers liquid crystalline phases in lipid droplets and alters the LD proteome. J. Cell Biol. 221, e202205053 (2022).
Binns, D., Lee, S., Hilton, C. L., Jiang, Q. X. & Goodman, J. M. Seipin is a discrete homooligomer. Biochemistry 49, 10747–10755 (2010).
Sung, M. K. & Huh, W. K. Bimolecular fluorescence complementation analysis system for in vivo detection of protein-protein interaction in Saccharomyces cerevisiae. Yeast 24, 767–775 (2007).
Magliery, T. J. et al. Detecting protein-protein interactions with a green fluorescent protein fragment reassembly trap: scope and mechanism. J. Am. Chem. Soc. 127, 146–157 (2005).
Gueldener, U., Heinisch, J., Koehler, G. J., Voss, D. & Hegemann, J. H. A second set of loxP marker cassettes for Cre-mediated multiple gene knockouts in budding yeast. Nucleic Acids Res. 30, e23 (2002).
Longtine, M. S. et al. Additional modules for versatile and economical PCR-based gene deletion and modification in Saccharomyces cerevisiae. Yeast 14, 953–961 (1998).
Chung, J. et al. The Troyer syndrome protein spartin mediates selective autophagy of lipid droplets. Nat. Cell Biol. 25, 1101–1110 (2023).
Choudhary, V., Ojha, N., Golden, A. & Prinz, W. A. A conserved family of proteins facilitates nascent lipid droplet budding from the ER. J. Cell Biol. 211, 261–271 (2015).
Gerace, E. & Moazed, D. Coimmunoprecipitation of proteins from yeast. Methods Enzymol. 541, 13–26 (2014).
Folch, J., Lees, M. & Sloane Stanley, G. H. A simple method for the isolation and purification of total lipides from animal tissues. J. Biol. Chem. 226, 497–509 (1957).
Schneiter, R. & Daum, G. Analysis of yeast lipids. Methods Mol. Biol. 313, 75–84 (2006).
Choudhary, V., Jacquier, N. & Schneiter, R. The topology of the triacylglycerol synthesizing enzyme Lro1 indicates that neutral lipids can be produced within the luminal compartment of the endoplasmatic reticulum: implications for the biogenesis of lipid droplets. Commun. Integr. Biol. 4, 781–784 (2011).
Horvath, A. & Riezman, H. Rapid protein extraction from Saccharomyces cerevisiae. Yeast 10, 1305–1310 (1994).
Acknowledgements
We thank members of Choudhary’s laboratory for helpful comments on the manuscript. We thank Shikha Chaudhary and the Electron Microscopy Core Facility of All India Institute of Medical Sciences (AIIMS) for technical assistance in EM imaging. We thank Sophisticated Analytical Instrument Facility (SAIF), AIIMS, New Delhi for assistance in confocal microscopy. We thank Dr Kalpana Luthra, Dept. of Biochemistry, AIIMS, New Delhi and Dr Swarandeep Singh for technical assistance with gel permeation chromatography. We also thank the Central Instrumentation Facility (CIF), University of Delhi South Campus for assistance with Ultracentrifugation. We thank Dr Kausik Chakraborty, CSIR-IGIB, New Delhi, for providing ldb16Δ::KanMX strain. This work was supported by an Indo Swiss Joint Research Programme of Department of Biotechnology (IC-12044(11)/6/2021-ICD- DBT awarded to V.C.), an Early Career Intramural Project of the All India Institute of Medical Sciences (AIIMS), New Delhi (A-1012/2023/RS awarded to V.C.), and DBT/Wellcome Trust India Alliance Fellowship (Grant IA/I/20/2/505191 awarded to V.C.). RMK was supported by fellowship from DBT (DBT-RA/2022/July/N/2565).
Author information
Authors and Affiliations
Contributions
A.K. and R.C. established the methods. A.K. and R.C. performed the experiments. R.M.K. contributed in generation of mutant strains and preliminary experiments. A.K. performed EM experiments with assistance from EM core facility at AIIMS. A.K., R.C. and V.C. analysed and interpreted the results. A.K. prepared the figures. V.C. conceived, supervised, and acquired funding for the project. V.C. wrote and edited the final version of the manuscript with inputs from all contributing authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Khatri, A., Chaudhary, R., Kumari, R.M. et al. The luminal domain region of Seipin/Fld1 is dispensable for establishing functional ER sites for lipid droplet biogenesis. Nat Commun 16, 10642 (2025). https://doi.org/10.1038/s41467-025-65645-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-65645-8