Abstract
Intellectual disability (ID) is characterized by deficits in cognition and adaptive behavior, with few treatment options. Tumor Suppressor Candidate 3 (TUSC3) has been genetically linked to autosomal recessive ID, but its molecular mechanism and therapeutic potential remain unclear. Here we show that TUSC3 is essential for endoplasmic reticulum (ER) Mg²⁺ homeostasis and neuronal function. Using a TUSC3 knockout (KO) mouse model, we find ID-like phenotypes including impairments in learning, memory, stress adaptation, and social behavior. Mechanistically, TUSC3 forms an ER-localized Mg²⁺ transport complex with ERMA and its loss leads to ER Mg²⁺ depletion, PERK–eIF2α pathway activation, synaptic dysfunction, and neuronal vulnerability. Fibroblasts from TUSC3 mutant patients similarly exhibit ER Mg²⁺ deficiency and heightened ER stress. Magnesium supplementation restores ER Mg²⁺ levels, reduces ER stress, and rescues cognitive deficits. Our findings establish ER Mg²⁺ dysregulation as a key driver of neurodevelopmental dysfunction and a promising therapeutic target.
Similar content being viewed by others
Introduction
Intellectual disability (ID) affects approximately 1–2% of the global population, posing substantial challenges to affected individuals, families, and healthcare systems1. Characterized by deficits in intellectual functioning and adaptive behaviors, including social skills and stress adaptation, ID results from a broad range of genetic mutations, many of which have been identified through advances in genomic technologies2. Notably, autosomal recessive ID (ARID), predominantly reported in Middle Eastern populations3, accounts for about 10% of ID cases in outbred populations4. Despite this, it remains less studied than autosomal dominant or X-linked forms due to its relative rarity in diverse outbred populations and genetic heterogeneity, which complicates sample collection and gene identification.
Tumor Suppressor Candidate 3 (TUSC3) is one of the genes implicated in ARID, with pathogenic variants identified in affected individuals5,6,7,8,9. TUSC3 is an endoplasmic reticulum (ER)-localized transmembrane protein, traditionally associated with N-glycosylation8. Furthermore, Zhou and Clapham identified that TUSC3 is required for cellular magnesium (Mg²⁺) uptake in mammalian cells and vertebrate embryonic development10. While this study identifies TUSC3 as essential for maintaining cellular Mg²⁺ levels10, its specific role in ER Mg²⁺ uptake and its contribution to neuronal dysfunction and ARID via ER stress remain poorly defined.
Mg²⁺ is essential for numerous enzymatic reactions, and its intracellular homeostasis is critical for neuronal function11,12. ER Mg²⁺ is crucial for ATP stabilization, chaperone activity, and protein folding. While direct evidence linking ER Mg²⁺ levels to neuronal dysfunction is lacking, studies have shown that culturing cells in low Mg²⁺ conditions triggers ER stress and activates the unfolded protein response13. Since chronic ER stress is implicated in various neurodevelopmental and neurodegenerative disorders14, a potential link between ER Mg²⁺ homeostasis and cognitive dysfunction remains to be elucidated. However, how TUSC3 regulates ER Mg²⁺ and its role in ARID remain unexplored.
To address this gap, we generated a TUSC3 knockout (KO) mouse model to investigate its role in ARID. We demonstrate that TUSC3 regulates ER Mg²⁺ homeostasis via ERMA, an ER magnesium transporter. TUSC3 deficiency disrupts ER Mg²⁺ uptake, triggers ER stress, and impairs global protein translation, leading to neuronal dysfunction and cell death. These deficits result in ID-like behaviors, including impairments in cognitive function, adaptive behaviors. Notably, magnesium supplementation restored ER Mg²⁺ balance, alleviated ER stress, and rescued cognitive and behavioral deficits of already developed ARID in TUSC3 KO mice, identifying magnesium supplementation as a potential therapy for TUSC3-related ID.
Results
TUSC3 is required for ER Mg²⁺ uptake in multiple cell types, including ARID patient-derived fibroblasts
To elucidate the molecular mechanism by which TUSC3 loss contributes to ARID, we first examined its known role as a non-catalytic subunit of the oligosaccharyltransferase complex involved in N-glycosylation15. We analyzed CNNM3, a Mg²⁺ uptake transporter known to undergo N-glycosylation16,17, in SH-SY5Y TUSC3 knockdown (KD) cells. While the N73A mutant of CNNM3 showed a clear band shift, CNNM3 WT exhibited no such shift in TUSC3 KD cells, indicating that its glycosylation was not affected (Supplementary Fig. 1a). We further analyzed glycosylation status of endogenous proteins, including NCAM1, N-cadherin, and GluA2, in primary hippocampal neurons and whole-brain lysates from our TUSC3 KO mice (characterized later in this study). Following PNGase F treatment, all proteins analyzed showed the expected band shift on the western blot, with no differences between WT and TUSC3 KO mice (Supplementary Fig. 1b, c). These results suggest that a predicted role of TUSC3 in N-glycosylation might not be linked to ARID, consistent with a previous report showing no significant changes in N-glycosylated proteins in blood samples from ARID patients8.
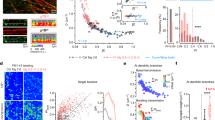
Next, given that TUSC3 regulates Mg²⁺ uptake in mammalian cells10, we examined this possibility by using an ER-targeting Mg²⁺ sensor, Mag-FRETER18,19, to monitor ER Mg²⁺ dynamics. Mag-FRETER contains an ER retention sequence (KDEL) and forms a compact structure upon binding Mg²⁺, generating a detectable FRET signal based on YFP (Fig. 1a)19,20. We confirmed ER localization of Mag-FRETER in SH-SY5Y cells (Supplementary Fig. 2a). Consistent with the previous findings19, L-lactate treatment triggered a rapid release of Mg²⁺ from the ER into the cytosol, followed by a reuptake phase where ER Mg²⁺ levels were restored (Fig. 1b). Real-time monitoring of ER Mg²⁺ dynamics revealed a sharp decrease at 21 s in both SH-SY5Y control and TUSC3 KD cells upon L-lactate treatment (Fig. 1b), accompanied by a concurrent rise in cytosolic Mg²⁺ levels (Supplementary Fig. 2b–d). The rate of Mg²⁺ release from the ER to the cytosol was comparable between SH-SY5Y control and TUSC3 KD cells (Fig. 1c). However, during the subsequent reuptake phase, SH-SY5Y TUSC3 KD cells exhibited significantly impaired Mg²⁺ uptake compared with control cells (Fig. 1d). Notably, reconstitution of SH-SY5Y TUSC3 KD cells with TUSC3-mRFP restored ER Mg²⁺ uptake to control levels (Fig. 1d, e). These findings suggest that while TUSC3 is dispensable for Mg²⁺ release from the ER, it is crucial for efficient Mg²⁺ uptake.
a Schematic of the ER-targeted KDEL-Mag-FRETER system to assess ER Mg²⁺ dynamics. Created with BioRender.com. b–d SH-SY5Y shControl and shTUSC3#3-1 cells were transfected with Mag-FRETER for 48 h and analyzed for ER Mg²⁺ uptake following L-lactate treatment (10 mM). Circle 1 indicates ER Mg²⁺ release, and Circle 2 represents ER Mg²⁺ refilling (b, left). Representative confocal images at 21 s (b, right). Quantification of normalized ER Mg²⁺ release (Circle 1) (c). Quantification of normalized ER Mg²⁺ uptake (left) and uptake rate (right) (Circle 2) (n = 6, 7, 7 cells per group) (d). e Representative confocal images of SH-SY5Y shTUSC3#3-1 cells co-expressing Mag-FRETER and TUSC3-mRFP. Scale bar: 10 μm. f–i Non-ID and ID fibroblasts were transfected with Mag-FRETER alone (f) or co-transfected with TUSC3-mRFP (g) for 48 h and analyzed for ER Mg²⁺ uptake following L-lactate treatment (10 mM). Quantification of normalized ER Mg²⁺ release (Circle 1) (h), ER Mg²⁺ uptake (i, left), and uptake rate (i, right) (Circle 2) (n = 7 cells per group). Scale bar: 10 μm. j, k SH-SY5Y shControl and shTUSC3#3-1 cells were transfected with Mag-FRETER for 48 h, and supplemented with acute MgCl₂ treatment (10 mM, 200 s) to assess ER Mg²⁺ uptake (j), quantification of ER Mg²⁺ levels at 0 s and 200 s (n = 12, 14 cells per group) (k). l, m WT and TUSC3 KO primary cortical neurons were transfected with Mag-FRETER for 48 h and analyzed for ER Mg²⁺ uptake following acute MgCl₂ treatment (10 mM, 200 s) (l), quantification of ER Mg²⁺ levels at 0 s and 200 s (n = 10, 9 cells per group) (m). n, o Non-ID and ID fibroblasts were transfected with Mag-FRETER for 48 h and supplemented with acute MgCl₂ treatment (10 mM, 200 s) to assess ER Mg²⁺ uptake (n), quantification of ER Mg²⁺ levels at 0 s and 200 s (n = 10, 7, 10 cells per group) (o). One-way ANOVA followed by Tukey’s post hoc multiple comparison test. All data are presented as mean ± S.E.M. Source data are provided as a Source data file.
To assess the relevance of our findings to human disease, we analyzed fibroblasts derived from two patients with ARID harboring biallelic mutations in TUSC3 (ID-fibroblasts)7. Using the Mag-FRETER, we found that Mg²⁺ release from the ER to the cytosol in response to L-lactate was comparable between ID- and Non-ID fibroblasts (Fig. 1f–h). Similar to SH-SY5Y cells, however, Mg²⁺ reuptake into the ER was significantly impaired in both patient fibroblasts compared with non-ID fibroblasts (Fig. 1g, i). As expected, reintroducing TUSC3-mRFP into ID-fibroblasts rescued ER Mg²⁺ uptake deficits (Fig. 1g, i). This analysis was extended to other cell types, including mouse embryonic fibroblasts (MEFs) and the colon cancer cell line HCT116, where TUSC3 KO similarly resulted in reduced ER Mg²⁺ uptake (Supplementary Fig. 2e–j). These results underscore the essential and conserved role of TUSC3 in maintaining ER Mg²⁺ homeostasis across multiple cell types, including cells derived from ARID patients.
Magnesium supplementation rescues ER Mg²⁺ levels in TUSC3-deficient cells
When we measured basal ER Mg²⁺ levels, they were significantly reduced in TUSC3-deficient cells compared with WT cells (Fig. 1j, l, n). To evaluate the immediate capacity of ER Mg²⁺ uptake, we applied an acute MgCl₂ pulse (10 mM for ~200 s) to cells transfected with Mag-FRETER. Intriguingly, we found that ER Mg²⁺ levels rapidly increased in SH-SY5Y control cells, and by 200 s, TUSC3 KD cells also showed near-complete recovery to basal control levels, albeit with lower and slower Mg²⁺ uptake (Fig. 1j, k). A similar pattern was observed in TUSC3 KO primary cortical neurons, which exhibited reduced basal ER Mg²⁺ levels relative to WT neurons, but showed partial recovery upon MgCl₂ treatment (Fig. 1l, m). Likewise, ID-fibroblasts exhibited lower ER Mg²⁺ levels than non-ID fibroblasts, and their ER Mg²⁺ was restored by MgCl₂ treatment, albeit with delayed uptake kinetics (Fig. 1n, o). These findings suggest that, while TUSC3-deficient cells have reduced basal ER Mg²⁺ levels, supplementation with high concentrations of MgCl₂ can effectively restore ER Mg²⁺ homeostasis.
TUSC3 interacts with ER magnesium transporter ERMA to facilitate Mg²⁺ uptake
TUSC3 lacks a canonical Mg²⁺ transport domain and GxN motifs21,22, suggesting that it may function by modulating another transporter rather than directly mediating Mg²⁺ transport. ERMA (ER magnesium transporter A; formerly TMEM94) is a P-type ATPase recently identified as a key mediator of ER Mg²⁺ uptake, and P-type ATPases frequently require accessory subunits for optimal activity23. We therefore hypothesized that TUSC3 functions as a regulatory component of ERMA. To test this, ER Mg²⁺ release and uptake were examined in SH-SY5Y TUSC3 KD cells overexpressing ERMA-DsRed (Fig. 2a–d). ERMA-DsRed overexpression enhanced Mg²⁺ uptake in SH-SY5Y control cells, but this effect was attenuated in TUSC3 KD cells, failing to restore ER Mg²⁺ uptake to control levels (Fig. 2b–d).
a–d SH-SY5Y shControl and shTUSC3#3-1 cells were co-transfected with Mag-FRETER and ERMA-DsRed for 48 h and analyzed using confocal microscopy (a). Scale bar: 10 μm. Representative traces showing ER Mg²⁺ uptake following L-lactate (10 mM) treatment (b). Quantification of normalized ER Mg²⁺ release (Circle 1) (c). Quantification of normalized ER Mg²⁺ uptake (left) and uptake rate (right) (Circle 2) (n = 9 cells per group) (d). e, f Co-immunoprecipitation from whole brain lysates of WT mice showing endogenous interaction between TUSC3 and ERMA. Immunoprecipitation with anti-ERMA antibody pulled down TUSC3 (e), and reciprocal IP with anti-TUSC3 antibody confirmed the presence of ERMA (f). L.C.: light chain of immunoglobulin. g Schematic representation of TUSC3-mRFP deletion mutants. h Immunoprecipitation assays between ERMA-FLAG and various TUSC3-mRFP deletion mutants. i–k SH-SY5Y shControl and shTUSC3#3-1 cells were transfected with Mag-FRETER and indicated TUSC3-mRFP deletion mutants for 48 h and analyzed for ER Mg²⁺ release and uptake following L-lactate (10 mM) treatment (i). Quantification of ER Mg²⁺ release (Circle 1) (j) and ER Mg²⁺ uptake (left) and uptake rate (right) (Circle 2) (k) (n = 10, 9, 9, 6, 9, 4, 5 cells per group). One-way ANOVA followed by Tukey’s or Dunnett’s (k) post-hoc multiple comparison test. All data are presented as mean ± S.E.M. Source data are provided as a Source data file.
To determine whether TUSC3 interacts with ERMA, we performed co-immunoprecipitation (co-IP) assays. Co-IP assays clearly showed their interaction in HEK293T cells overexpressing TUSC3-mRFP and ERMA-FLAG (Supplementary Fig. 2k, l). Consistent with this, co-IP assays using whole brain lysates revealed the interaction between endogenous TUSC3 and ERMA (Fig. 2e, f). To map the TUSC3 domain critical for this interaction, deletion (Δ) mutants were generated based on TUSC3 topology (Fig. 2g). Co-IP assays revealed that Δ197–242 (TM1 and TM2) significantly reduced TUSC3-ERMA binding, whereas Δ222–297 (TM2 and TM3) had minimal effect (Fig. 2h), indicating that the TM1 domain is essential for the interaction. To evaluate the functional relevance of this interaction, SH-SY5Y TUSC3 KD cells were reconstituted with either TUSC3 WT or deletion mutants. Unlike WT and other deletion mutants, TUSC3 Δ197–242 failed to rescue ER Mg²⁺ levels (Fig. 2i–k). Thus, TUSC3 promotes ER Mg²⁺ uptake by interacting with ERMA, and this interaction critically depends on the TM1 domain.
TUSC3 deficiency evokes ER stress, translation impairment, and cell death via Mg² dysregulation
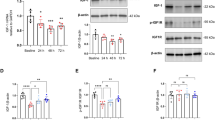
To determine how ER Mg²⁺ depletion in TUSC3-deficient cells contributes to ID phenotypes, we examined ER stress responses. Since Mg²⁺ is essential for protein folding and chaperone activity in the ER, its depletion can lead to accumulation of misfolded protein and activation of the unfolded protein response13. In the hippocampus and striatum of TUSC3 KO mice, GRP78/BiP was significantly upregulated, and activation of the PERK–eIF2α axis, including p-eIF2α and CHOP, was markedly increased (Fig. 3a–r). In contrast, the IRE1–XBP1s and ATF6 branches were less affected (Fig. 3f, i, o, r), suggesting that the PERK–eIF2α axis is the primary response of TUSC3 deficiency. CREB activity, a key regulator of learning and memory, and PI3K/AKT signaling24, its upstream modulator, were both reduced (Fig. 3g, h, p, q), linking ER stress to cognitive deficits in TUSC3 KO mice.
a Representative western blot of ER stress proteins in the hippocampus of 4-month-old female WT and TUSC3 KO mice. b–i Quantification of GRP78 (b), p-PERK (T982) (c), p-eIF2α (S51) (d), CHOP (e), XBP1s (f), p-AKT (S473) (g), p-CREB (S133) (h), and ATF6 (i) normalized to β-actin (n = 3 mice per group). j Representative western blot analysis of ER stress proteins in the striatum of 4-month-old female WT and TUSC3 KO mice (n = 3 per group). k–r Quantification of GRP78 (k), p-PERK (T982) (l), p-eIF2α (S51) (m), CHOP (n), XBP1s (o), p-AKT (S473) (p), p-CREB (S133) (q), and ATF6 (r) normalized to β-actin (n = 3 mice per group). s Non-ID and ID1/ID2 fibroblasts were chronically treated with MgT (400 μM) for 48 h and analyzed by western blotting (left). Quantification of p-eIF2α (S51) levels normalized to eIF2α (right) (n = 3 independent biological replicates). t Non-ID and ID-derived fibroblasts were pretreated with or without MgT (400 μM) for 24 h, followed by puromycin (5 μg/ml) treatment for 2 h. Representative western blot using an anti-puromycin antibody (left), and puromycin incorporation was quantified and normalized to β-actin (right) (n = 3 independent biological replicates). u SH-SY5Y shControl and shTUSC3#3-1 cells were pretreated with MgT (400 μM) for 12 h, then treated with either DMSO, thapsigargin (2 μM), tunicamycin (2 μg/ml), A23187 (2 μM), or etoposide (25 μM) for 24 h. Cells were stained with propidium iodide and Calcein-AM, and double-positive cells were quantified and normalized to Calcein-AM-positive cells. Data represent three independent biological experiments (n = 3). The exact number of cells analyzed per condition is provided in the Source Data file. v Representative western blot of primary cortical neurons (DIV7) treated with vehicle or 4-PBA (10 mM) for 24 h (left). Quantification of indicated protein levels normalized to TUBA (right) (n = 3 independent biological replicates). Two-tailed unpaired t-test (b–i, and k–r); one-way ANOVA followed by Tukey’s post hoc multiple comparison test (s, t, v); two-way ANOVA followed by Tukey’s multiple comparison test (u). Data are presented as mean ± S.E.M. or S.D. (t, u). Source data are provided as a Source data file.
ER stress inhibits protein translation via the PERK–eIF2α axis, a mechanism implicated in neurodevelopmental disorders such as Down syndrome25,26. Using the Surface Sensing of Translation (SUnSET) assay27, we found that global protein translation was significantly reduced in TUSC3 KO MEFs, coinciding with the increase of GRP78, p-PERK, p-eIF2α, and XBP1s levels (Supplementary Fig. 3a–c). Similar findings were observed in ID-fibroblasts (Supplementary Fig. 3d, e). Notably, chronic magnesium supplementation with magnesium-L-threonate (MgT, 400 μM, 48 h), which has ability to cross the blood-brain barrier12, restored protein translation and reduced p-eIF2α levels in ID-fibroblasts (Fig. 3s, t), suggesting that Mg²⁺ supplementation mitigates both ER stress and translation impairment.
Since CHOP is a pro-apoptotic factor in ER stress response28, we tested to confirm whether TUSC3 deficiency increases cell death susceptibility. Compared with control cells, SH-SY5Y TUSC3 KD cells were more vulnerable to ER stressors (thapsigargin, tunicamycin, A23187), but not to the DNA-damaging agent etoposide (Fig. 3u, Supplementary Fig. 3f). Notably, magnesium supplementation effectively mitigated ER stress-induced cell death (Fig. 3u). To further assess the role of ER stress in TUSC3-related neuronal dysfunction, TUSC3 KO primary cortical neurons were treated with 4-phenylbutyrate (4-PBA), a chemical chaperone that alleviates ER stress29. Intriguingly, 4-PBA treatment markedly restored PSD-93, PSD-95, and GluA1 levels in TUSC3 KO neurons (Fig. 3v). These findings highlight that ER Mg²⁺- stress regulates synaptic activity in TUSC3-deficient cells, linking ER Mg²⁺ homeostasis to cognitive function. Targeting ER Mg²⁺-stress may serve as a therapeutic strategy for TUSC3-related ID.
TUSC3 KO mice display abnormal embryonic development
To assess the in vivo relevance of our cellular findings, we next generated a TUSC3 KO mouse model using CRISPR-Cas9-mediated targeting of Exon 2 in the Tusc3 gene, leading to a frameshift mutation (see Methods and Supplementary Fig. 4a for detailed methodology). Genotyping and western blot analysis confirmed the successful generation of TUSC3 KO mice (Supplementary Fig. 4b). Heterozygous Tusc3 mice were bred to generate homozygous KO offspring. However, only 48 of 568 offspring (8.6%) were homozygous KO, significantly deviating from expected Mendelian ratios (Supplementary Fig. 4c). Further analysis revealed embryonic lethality, with resorption observed as early as embryonic day (E)11.5 (Supplementary Fig. 4d). Next, we assessed growth phenotypes in surviving TUSC3 KO embryos and mice, which exhibited significantly lower body weights compared with WT littermates (Supplementary Fig. 4e, f). These results indicate incomplete embryonic lethality in TUSC3 KO mice, with surviving offspring showing persistent growth deficits compared with WT mice.
TUSC3 KO mice display intellectual disability phenotypes
To investigate whether TUSC3 KO mice represent ID phenotypes, we conducted a series of behavioral tests that assess learning, memory, stress-coping ability, and social interactions, which are hallmark features of ID2. At 4 weeks of age, TUSC3 KO mice showed no significant differences compared with WT mice in the Y-maze, novel object recognition (NOR), and social interaction tests, but exhibited a significant reduction in immobility time in the tail suspension test (TST) (Supplementary Fig. 5a–e), indicating early stress-coping deficits. At 3 months of age, however, cognitive, social, and stress-related impairments emerged in TUSC3 KO mice. Three-month-old TUSC3 KO mice exhibited significantly reduced spontaneous alternation rates in the Y-maze (Fig. 4a, b). In the NOR test, TUSC3 KO mice displayed impaired novel object discrimination (Fig. 4c–e). In the passive avoidance test, TUSC3 KO mice demonstrated significantly shorter latency to re-enter the dark chamber (Supplementary Fig. 6a). Collectively, these results indicate that TUSC3 deficiency leads to significant impairments in spatial working memory, recognition memory, and associative memory, underscoring a broad disruption in cognitive function.
a, b Schematic of the Y-maze test (a) and spatial memory performance in WT and TUSC3 KO mice (b). Created with BioRender.com. Schematic of the novel object recognition (NOR) test (c) and representative tracking plots in the open field in the NOR test (d). Created with BioRender.com. e The NOR test (discrimination index) for WT and TUSC3 KO mice. f, g Schematic of the forced swim test (FST), consisting of a 120 s adaptation phase followed by 240 s of immobility measurement (f) and immobility time in the FST, assessing stress-coping ability in WT and TUSC3 KO mice (g). Created with BioRender.com. h, i Schematic of the tail suspension test (TST), with immobility recorded over 360 s (h) and immobility time in the TST for WT and TUSC3 KO mice (i). Created with BioRender.com. j, k Schematic of three-chambered social interaction test (j) and representative tracking plots from the social interaction test, showing times spent in each chamber (k). Created with BioRender.com. l, m Social interaction preference in TUSC3 KO mice, assessed by time spent with the novel mouse (l) and sniffing time (m). n–q Schematic of three-chambered social preference test (n) and representative tracking plots of WT and TUSC3 KO mice in social preference test (o). Social preference for the novel mouse of WT and TUSC3 KO mice in the social preference test (p). Sniffing time data confirms this lack of preference (q). Created with BioRender.com. In all behavioral experiments, 4-month-old mice were used: WT: 9 (5 F, 4 M); KO: 10 (6 F, 4 M). Two-tailed unpaired t-test (b, e, g, i), Two-way ANOVA followed by Tukey’s post-hoc multiple comparison test (l, m, p, q). All data are presented as mean ± S.E.M. Source data are provided as a Source data file.
Since adaptive skills, including stress response, are frequently impaired in ID30, we next assessed stress-coping ability in TUSC3 KO mice using the forced swim test (FST) and TST. FST, traditionally used as a behavioral assay for depression31, is now widely recognized as a measure of stress adaptability32,33. In both FST and TST, TUSC3 KO mice exhibited markedly reduced immobility time, indicating deficits in stress-coping ability (Fig. 4f–i). Given that social behavior is another core domain affected in ID34, we next examined social interaction and recognition in TUSC3 KO mice. In the three-chamber social interaction test, TUSC3 KO mice spent significantly less time interacting with a novel mouse (Fig. 4j–m). Similarly, in the social novelty preference test, TUSC3 KO mice failed to show a preference for a novel mouse over a familiar one, highlighting deficits in social recognition and novelty response (Fig. 4n–q). Additionally, TUSC3 KO mice exhibited hyperactivity in the open field test, with increased locomotor activity but no significant changes in anxiety-like behavior (Supplementary Fig. 6b–d). They also displayed excessive self-grooming, suggestive of repetitive behaviors (Supplementary Fig. 6e, f). Together, TUSC3 KO mice exhibit progressive behavioral impairments characteristic of ID, with stress-coping deficits emerging by 1 month of age, followed by the onset of cognitive and social deficits at 3 months.
Synaptic structure and plasticity are impaired in TUSC3 KO mice
Abnormal synaptic signaling, impaired plasticity, and changes in spine morphology are key features of various psychiatric and neurological disorders35. Synaptic dysfunction is commonly observed in individuals with ID36. We thus sought to examine levels of scaffold proteins (PSD-93, PSD-95) and AMPA receptor subunit GluA1, which are critical components of synapse structure and function37. Western blot analysis revealed reduced levels of PSD-93, PSD-95, and GluA1 in TUSC3 KO mice compared with WT mice (Fig. 5a). Additionally, neuronal activity, measured by CREB phosphorylation38, was markedly reduced in TUSC3 KO mice. These molecular deficits were accompanied by a significant reduction in brain weight of TUSC3 KO mice compared with WT mice (Supplementary Fig. 7a). Consistently, levels of synaptic markers as well as phosphorylated CREB were also decreased by TUSC3 deficiency in primary hippocampal neurons and human neuroblastoma SH-SY5Y cells (Supplementary Fig. 7b–e). These results indicate that these reductions may contribute to the synaptic dysfunction observed in TUSC3 KO mice.
a Representative western blot of PSD-93, PSD-95, GluA1, and phosphorylated CREB (S133) in whole-brain lysates of 4-month-old WT and TUSC3 KO female mice (left). Quantification of indicated protein levels (right) (n = 3). b, c Representative confocal images of immunohistochemistry for TUSC3, synaptophysin (SYP), and MAP2 in the CA1 and CA3 hippocampal regions and the striatum of 4-month-old WT and TUSC3 KO female mice (b), and quantification of SYP intensity normalized to MAP2 (c) (n = 3). Scale bar: 50 μm. d, e Representative confocal images of immunohistochemistry for TUSC3, GluA1, and MAP2 in the CA1 and CA3 hippocampal regions and the striatum of 4-month-old WT and TUSC3 KO female mice (d), and quantification of GluA1 intensity normalized to MAP2 (e) (n = 3). Scale bar: 50 μm. f, g Representative dendritic images of primary hippocampal neurons transfected with EGFP (f). Scale bar: 50 μm. Sholl analysis of dendritic branching in WT and TUSC3 KO neurons (n = 20 cells per group) (g). h, i Representative confocal images of dendritic spines of primary hippocampal neurons (DIV 12) transfected with EGFP for 48 h (h). Violin plot quantifying dendritic spine density (spines per 10 μm) in WT and KO neurons (n = 20 cells per group) (i). Scale bar: 10 μm. j–l Representative traces of sEPSCs recorded from hippocampal slices of 2-month-old WT and TUSC3 KO mice (j), quantification of sEPSC amplitude (k), and frequency (l) in WT (n = 12 cells from 5 mice; 4 F, 1 M) and TUSC3 KO (n = 8 cells from 5 mice; 2 F, 3 M). m Representative traces (left) and quantification of the eEPSC amplitude (right) in WT (n = 8 cells from 5 mice; 4 F, 1 M), TUSC3 KO (n = 8 cells from 5 mice; 2 F, 3 M). n Representative traces (left) and EPSC-PPR (right) in WT (n = 8 cells from 5 mice; 3 F, 2 M), TUSC3 KO (n = 9 cells from 6 mice; 4 F, 2 M). Two-tailed unpaired t-test. Data are presented as mean ± S.E.M. (a, c, e, and k–n) or S.D. (g, i). Source data are provided as a Source data file.
To determine which brain regions are most vulnerable to TUSC3 deficiency, we examined the hippocampus and striatum—regions critically involved in cognitive function and adaptive behaviors such as social skills and stress coping39,40. Immunohistochemistry revealed a significant reduction in synaptophysin, a presynaptic marker41, in both the hippocampus CA1 region and the striatum of TUSC3 KO mice, whereas no change was detected in the CA3 region (Fig. 5b, c). GluA1 expression was similarly reduced in these regions (Fig. 5d, e). Next, we investigated abnormalities in synaptic structure in TUSC3 KO mice. Sholl analysis of primary hippocampal neurons showed a significant reduction in dendritic branching, as evidenced by fewer crossings in TUSC3 KO neurons compared with WT neurons (Fig. 5f, g). Moreover, dendritic spine density was markedly reduced in TUSC3 KO neurons (Fig. 5h, i).
We further investigated whether the structural abnormalities of the synapse in TUSC3 KO mice are associated with altered synaptic transmission using whole-cell voltage clamp recording. The spontaneous EPSC (sEPSC) amplitude in hippocampal CA1 neurons was significantly increased in TUSC3 KO mice, whereas sEPSC frequency remained unchanged (Fig. 5j–l). Evoked EPSC (eEPSC) amplitude was reduced, while the pair-pulse ratio (PPR) was unaltered (Fig. 5m, n). Although the unchanged PPR suggests intact presynaptic release probability under brief stimulation, FM1-43 dye unloading assays revealed delayed synaptic vesicle release in TUSC3 KO hippocampal neurons following sustained depolarization (Supplementary Fig. 8a–d), indicating that TUSC3 deficiency impairs synaptic vesicle release during prolonged stimulation and represents a form of presynaptic dysfunction not detected by standard PPR measurements.
We also performed similar recordings in CA3 pyramidal neurons. Unlike in CA1, sEPSC amplitude was unchanged in CA3; however, sEPSC frequency was significantly reduced in TUSC3 KO neurons compared with WT neurons (Supplementary Fig. 8e–g). This contrast shows region-dependent vulnerability of hippocampal circuits to TUSC3 loss, reflecting postsynaptic changes such as altered receptor expression or synapse strength in CA1 and presynaptic deficits of spontaneous neurotransmitter release in CA3. When we further analyzed miniature end-plate potentials (mEPPs) at the neuromuscular junction, no significant differences in amplitude or frequency were observed between WT and TUSC3 KO mice (Supplementary Fig. 8h–j), suggesting that synaptic transmission is largely preserved in peripheral synapses. Together, these results indicate that TUSC3 deficiency leads to structural synaptic abnormalities and region-specific disruptions in synaptic transmission, with both pre- and postsynaptic components being affected in a context-dependent manner. The lack of functional impairment at the neuromuscular junction further supports a selective vulnerability of central circuits to TUSC3 loss.
Magnesium supplementation ameliorates intellectual disability symptoms caused by TUSC3 deficiency
Given the role of ER Mg²⁺-stress and neuronal dysfunction in TUSC3-deficient cells, we investigated whether magnesium supplementation could rescue or prevent ID-like phenotypes in TUSC3 KO neurons and mice. In TUSC3 KO primary cortical neurons, we observed that chronic treatment of MgT (200 μM, 48 h) restored synaptic protein levels (PSD-93, PSD-95, GluA1) and CREB activity to WT levels (Supplementary Fig. 9a, b). To clarify whether MgT is able to restore ER Mg²⁺ levels, as previously demonstrated by MgCl₂ in vitro, we measured ER Mg²⁺ accumulation kinetics in SH-SY5Y TUSC3 KD cells using the Mag-FRETER assay. Whereas MgCl₂ treatment rapidly elevated ER Mg²⁺ levels within ~200 s, MgT exhibited a slower uptake profile during the 600 s imaging window. Intriguingly, ER Mg²⁺ levels reached comparable levels to MgCl₂ treatment approximately 8 h after MgT treatment (Supplementary Fig. 9c, d). These results indicate that both MgCl₂ and MgT can replenish ER Mg²⁺ levels, albeit with distinct kinetics.
We then assessed whether chronic magnesium supplementation via drinking water (910 mg/kg/day for 4 weeks) could reverse existing ID-like phenotypes in TUSC3 KO mice. Following treatment, a battery of behavioral tests was performed (Supplementary Fig. 10a). Hyperactivity in 5-month-old TUSC3 KO mice was significantly reduced to WT levels (Supplementary Fig. 10b). Cognitive performance in the Y-maze and NOR tests was markedly improved, reaching levels comparable to WT controls (Supplementary Fig. 10c, d). In the FST, TUSC3 KO mice showed impaired stress adaptation, as indicated by early removal and reduced immobility time. MgT treatment partially improved both survival and passive coping. Stress resilience in the TST was also significantly enhanced (Supplementary Fig. 10e, f). Social interaction and social preference behaviors were significantly improved following MgT supplementation (Supplementary Fig. 10g, h). At the molecular level, western blot analysis of the brain tissues further supported these findings. Synaptic protein levels in the striatum and hippocampus of TUSC3 KO mice were significantly restored by MgT treatment, whereas minimal effects were observed in the prefrontal cortex (Supplementary Fig. 10i–k). Altogether, these results suggest that MgT supplementation effectively alleviates ID phenotypes in TUSC3 KO mice after symptom onset.
To determine whether MgT could also prevent the onset of ID phenotypes, we chronically administered MgT to 4-week-old TUSC3 KO mice for six weeks and assessed their cognitive and behavioral performance (Fig. 6a). In the Y-maze and NOR tests, MgT-treated TUSC3 KO mice exhibited cognitive performance comparable to WT levels (Fig. 6b, c and Supplementary Fig. 11a). In the passive avoidance test, MgT supplementation restored long-term memory retention in TUSC3 KO mice to WT levels (Supplementary Fig. 11b). Stress-coping abilities in the TST were maintained at WT levels and social behaviors were also preserved in MgT-treated TUSC3 KO mice (Fig. 6d–f). In contrast, total distance traveled in the open field test was not significantly improved by MgT treatment (Supplementary Fig. 11c).
a Experimental timeline. Four-week-old mice received chronic magnesium supplementation via drinking water containing MgT (910 mg/kg/day) for 42 days, followed by behavioral analysis. Created with BioRender.com. b, c Cognitive performance of MgT-treated WT and TUSC3 KO mice assessed in the Y-maze (b) and NOR test (c). d Stress-coping ability evaluated using the TST. e, f Social behavior assessed in the social interaction (e) and social novelty preference (f) tests (left). Time spent in each chamber (middle) and sniffing time (right) were measured. g, h Representative western blot of hippocampal tissues from WT and TUSC3 KO mice treated with vehicle or MgT (g). Quantification of indicated protein levels (h) (n = 3). i, j Representative confocal images of GluA1 immunohistochemistry in the CA1 (i) and CA3 (j) hippocampal regions from WT and TUSC3 KO mice treated with vehicle or MgT (n = 3). Scale bar: 50 μm. In these experiments, 10-week-old mice were used: WT: 8 (4 F, 4 M); WT + MgT: 8 (4 F, 4 M); KO: 6 (3 F, 3 M); KO + MgT: 8 (5 F, 3 M). Two-way ANOVA with Tukey’s post hoc multiple comparison test. All data are represented as mean ± S.E.M. Source data are provided as a Source data file.
At the molecular level, levels of PSD-93, PSD-95, GluA1, and NMDAR1, as well as CREB activity, were restored to WT-like levels in the hippocampus of MgT-treated TUSC3 KO mice (Fig. 6g, h). Elevated GRP78 levels observed in TUSC3 KO mice were also normalized following MgT supplementation. Consistent with these hippocampal findings, MgT treatment similarly restored CREB activity, synaptic protein levels, and GRP78 levels in the striatum (Supplementary Fig. 11d, e). In contrast, synaptic protein levels and GRP78 expression in the prefrontal cortex were not significantly altered by MgT treatment (Supplementary Fig. 11f, g). These molecular changes were further corroborated by immunohistochemistry. In the hippocampus, GluA1 level was reduced in both CA1 and CA3 regions of TUSC3 KO mice but was restored following MgT treatment (Fig. 6i, j). In the striatum, synaptophysin and GluA1 levels, which were reduced in TUSC3 KO mice, were largely preserved following MgT supplementation (Supplementary Fig. 11h, i). Additionally, AKT activity was significantly upregulated upon MgT treatment (Supplementary Fig. 11j).
These results demonstrate that MgT supplementation effectively reverses and prevents ID-like phenotypes in TUSC3 KO mice by mitigating ER stress-related deficits and restoring synaptic function. Remarkably, MgT selectively restores synaptic protein levels in the striatum and hippocampus, underscoring its potential as a therapeutic strategy for TUSC3-related ID.
Discussion
Our study demonstrates that TUSC3 plays a pivotal role in ER Mg²⁺ uptake and neuronal function, establishing a mechanistic link between ER Mg²⁺ deficiency, ER stress, and cognitive impairment. In TUSC3 KO mice, we observed key features of ID, including cognitive and adaptive deficits. We identified ERMA as a binding partner of TUSC3, forming an ER-localized Mg²⁺ transport module essential for maintaining ER homeostasis. Disruption of this module triggered ER Mg²⁺ stress, leading to synaptic protein loss and dendritic abnormalities. Magnesium supplementation restored ER homeostasis, synaptic protein levels, and behavioral performance in TUSC3 KO mice, highlighting the therapeutic relevance of targeting ER Mg²⁺ in TUSC3-associated ID (Fig. 7). While a previous study has implicated TUSC3 in cytoplasmic Mg²⁺ regulation10, our findings directly link ER Mg²⁺ imbalance to synaptic dysfunction, filling a previously unaddressed mechanistic gap.
TUSC3 deficiency impairs ER magnesium uptake by disrupting its positive modulation of ERMA, an ER Mg²⁺ transporter. This leads to reduced ER [Mg²⁺], activation of ER stress via the PERK–eIF2α axis, and suppression of global translation, including synaptic proteins (PSD-93, PSD-95, GluA1). In parallel, CREB activity is reduced through impaired PI3K/AKT signaling. Together, these changes result in synaptic dysfunction, increased neuronal vulnerability (via CHOP), and ultimately cognitive and adaptive behavioral deficits. Notably, magnesium supplementation restores ER [Mg²⁺] levels, alleviates ER stress, and rescues synaptic and cognitive phenotypes in TUSC3-deficient mice, suggesting a potential therapeutic strategy for TUSC3-associated ID.
We demonstrate that ER Mg²⁺ imbalance leads to chronic ER stress, primarily via the PERK–eIF2α axis. Given that eIF2s regulate the translation of mRNA subsets critical for neuronal plasticity42, alteration in synaptic protein levels, including PSD-93, PSD-95, and GluA1, may contribute to the abnormal behaviors in TUSC3 KO mice. Electrophysiology revealed increased sEPSC amplitude and decreased eEPSC amplitude, indicating postsynaptic dysfunction, while reduced synaptophysin and delayed FM1-43 unloading support additional presynaptic impairments in TUSC3 KO mice. Notably, the PERK–eIF2α activation occurred with minimal engagement of IRE1–XBP1 or ATF6, showing a chronic, subthreshold ER stress state that fails to elicit robust chaperone or ERAD responses43,44,45. A similar PERK-dominant UPR profile has been reported in Down syndrome46, implicating this pattern as a shared feature in neurodevelopmental disorders. Although we did not modulate the PERK–eIF2α axis directly, the previous studies have shown that pharmacological inhibition (e.g., using ISRIB) can restore protein synthesis and cognition function47,48, supporting our proposal and highlighting its therapeutic potential in TUSC3 deficiency.
It is interesting to note that supplementation with MgT, which has superior blood-brain barrier permeability compared with other Mg²⁺ formulations12, increases ER Mg²⁺ levels in TUSC3-deficient cells, leading to mitigation of ER stress and rescue of synaptic proteins. We believe that ER Mg²⁺ levels, rather than cytosol Mg²⁺ are crucial for synaptic function for the following reasons. First, MgT supplementation in TUSC3-deficient cells rescues ER Mg²⁺ levels, but not cytosolic Mg²⁺ levels, as high as untreated WT cells. Second, MgT treatment mitigates ER stress and recovers synaptic proteins specifically in TUSC3-deficient cells. Notably, ER Mg²⁺ levels were slowly restored upon Mg²⁺ exposure in TUSC3 KO cells. Although MagT1 is a highly specific magnesium channel, its minimal expression in the brain49 suggests it is unlikely to compensate for TUSC3 in neuronal Mg²⁺ regulation. Alternative or compensatory pathways, such as TRPM7, CNNM3, SLC41A1, and SLC41A2, may contribute to residual Mg²⁺ transport within the ER and other organelles17,50,51. However, their role in ER Mg²⁺ uptake remains unverified since they primarily mediate cellular Mg²⁺ influx from distinct subcellular compartments52.
Overall, our study provides important insights into the role of TUSC3 in ER Mg²⁺ regulation and its impact on neuronal function. Identifying TUSC3 as a key component of ER Mg²⁺ transport expands our understanding of how Mg²⁺ homeostasis influences cognitive function and synaptic plasticity. Interestingly, several neurodevelopmental disorders, including autism spectrum disorder (ASD) and attention-deficit hyperactivity disorder (ADHD), are associated with altered serum Mg²⁺ levels53,54,55. In these disorders, magnesium supplementation has shown beneficial effects in alleviating symptoms53,54,56. Similarly, magnesium has been widely utilized to increase neuronal activity and to alleviate pathologic symptoms in other neurodevelopmental disorder models12,57. While these findings are largely peripheral and correlative, our study provides a mechanistic basis by which ER Mg²⁺ dysregulation could impair synaptic protein synthesis and neuronal signaling. This raises the possibility that similar ER Mg²⁺-dependent stress pathways may contribute to other neurodevelopmental phenotypes beyond TUSC3 deficiency.
Our findings extend prior research on magnesium transporters such as TRPM7, CNNM3, and SLC41A1/2, which have been implicated in neuronal excitability and Mg²⁺ homeostasis58,59,60, though their roles in ER-specific uptake remain undefined. By identifying the TUSC3–ERMA module as an ER-localized Mg²⁺ transport system, our study introduces an additional layer of compartment-specific Mg²⁺ regulation that connects intracellular ion balance with synaptic integrity. Given that ER stress is a common pathological feature in neurodevelopmental and neurodegenerative disorders, including bipolar disorder, fragile X syndrome, ASD, and ALS, our identification of an ER Mg²⁺-linked stress response offers insight into a potential mechanism of neuronal vulnerability. This raises the possibility that restoring ER Mg²⁺ homeostasis could serve as a broader therapeutic strategy61,62,63,64. Although our findings highlight ER Mg²⁺ dysregulation as a key mechanism in TUSC3-related cognitive dysfunction, additional molecular pathways are likely to contribute to the broader spectrum of ID and warrant further investigation.
Translating our findings into clinical applications requires assessing the feasibility of magnesium supplementation for TUSC3-associated ID. While MgT supplementation alleviated cognitive and synaptic deficits in TUSC3 KO mice, its application in humans demands careful consideration of dosage, long-term safety11,65,66,67. Although the effective dose in mice (910 mg/kg/day) appears high, allometric scaling yields a human equivalent of ~4.4 g/day of MgT, which corresponds to ~350 mg/day of elemental Mg²⁺—a level within the recommended daily intake range (200–500 mg/day) for adults68. This suggests potential clinical applicability, particularly when combined with optimized delivery methods such as controlled-release formulations or CNS-targeted prodrugs. However, chronic supplementation may induce metabolic adaptations—such as altered intestinal absorption, renal excretion, or transporter expression—and may pose risks including calcium imbalance or osmotic stress69,70. Thus, comprehensive long-term safety studies will be essential. Despite these considerations, our findings underscore ER Mg²⁺ homeostasis as a promising therapeutic target for TUSC3-associated ID and potentially broader neurodevelopmental disorders.
Methods
Antibodies
The following primary antibodies were used in this study: ACTB (Santa Cruz Biotechnology; sc-47778; lot # J0421; WB 1:5000), TUBA (Santa Cruz Biotechnology; sc-23948; lot # G2921; WB 1:5000), TUSC3 (clone D-9; Santa Cruz Biotechnology; sc-390566; lot # J1218; IF 1:500), GRP78 (Santa Cruz Biotechnology; sc-376768; lot # C0316; WB 1:3000), p-CREB-1-S133 (clone 10E9; Santa Cruz Biotechnology; sc-81486; WB 1:3000), FLAG (Sigma-Aldrich; F1804; lot # 0000375542; WB 1:3000), RFP (MBL; PM005; lot # 048; WB 1:3000), TUSC3 (Proteintech; 16039-1-AP; lot # 00007251; WB 1:500), AKT (Cell Signaling Technology; 9272; lot # 6; WB 1:3000), P-AKT (S473) (Cell Signaling Technology; 4060; lot # 27; WB 1:3000; IF 1:250), CREB (Cell Signaling Technology; 9197; WB 1:1000), CHOP (Cell Signaling Technology; 2895; WB 1:1000), ATF6 (Cell Signaling Technology; 65880; lot # 5; WB 1:3000), MAP2 (Cell Signaling Technology; 4542; IF 1:500), NCAM1 (Cell Signaling Technology; 3576; WB 1:3000), PERK (ABclonal; A18196; lot # 3560572205; WB 1:3000), p-PERK (T982) (ABclonal; AP0886; lot # 5500040024; WB 1:3000), XBP1s (ABclonal; A17007; lot # 1153240401; WB 1:3000), eIF2α (ABclonal; A21221; lot # 360002151; WB 1:3000), p-eIF2α (ABclonal; AP0745; lot # 360003834; WB 1:3000), Puromycin (ABclonal; A23931; lot # 3523033102; WB 1:3000), PSD-93 (ABclonal; A19669; WB 1:3000), PSD-95 (ABclonal; A7889; WB 1:3000), GluA1 (ABclonal; A1826; lot # 3561869003; WB 1:3000; IF 1:250), NMDAR1 (ABclonal; A7677; lot # 5500016744; WB 1:1000), Synaptophysin (ABclonal; A6344; lot # 5500008363; WB 1:1000; IF 1:250), and N-Cadherin (BD Biosciences; 610920; WB 1:3000). Secondary antibodies included HRP-conjugated goat anti-mouse (Thermo Fisher Scientific; 31430; WB 1:10000), HRP-conjugated goat anti-rabbit (Thermo Fisher Scientific; 31460; WB 1:10000), HRP-conjugated rabbit anti-goat (Thermo Fisher Scientific; 31402; WB 1:5000), Alexa Fluor 488-conjugated goat anti-mouse IgG (H + L) (Thermo Fisher Scientific; A-11001; IF 1:500), Alexa Fluor 594-conjugated goat anti-mouse IgG (H + L) (Thermo Fisher Scientific; A-11005; IF 1:500), Alexa Fluor 488-conjugated goat anti-rabbit IgG (H + L) (Thermo Fisher Scientific; A-11008; IF 1:500), and Alexa Fluor 594-conjugated goat anti-rabbit IgG (H + L) (Thermo Fisher Scientific; A-11012; IF 1:500). The anti-human ERMA polyclonal rabbit antibody was kindly provided by Dr. Madesh and used at WB 1:500. Protein G Sepharose 4 Fast Flow (GE Healthcare; 17-0618-05) was used for immunoprecipitation experiments.
Chemicals and reagents
Anti-FLAG M2 Affinity Gel (Cat # A2220), Dimethyl sulfoxide (Cat # D2650), Tunicamycin (Cat # T7765), Paraformaldehyde (Cat # P6148), Fluoromount aqueous mounting medium (Cat # F4680), Propidium iodide (Cat # P4170), Puromycin (Cat # P7255), Etoposide (Cat # E1383), Thapsigargin (Cat # T9033), A23187 (Cat # C7522), 4-Phenylbutylic acid (Cat # P21005), L-lactate (Cat # L1750), Magnesium chloride (Cat # 208337), and Polybrene (Cat # H9268) were from Sigma-Aldrich; DMEM (Cat # LM 001-05) and IMDM (Cat # LM004-01) were from Welgene, while RPMI-1640 (Cat # 16777-145) was from Hyclone; CHANG Medium D (Cat # T105) was from Irvine Scientific. FBS (Cat # 12483020), Neurobasal Medium (Cat # 21103049), B-27 supplement (Cat # 17504044), Gentamycin (Cat # 15710060), Penicillin-Streptomycin (Cat # 15140122), and GlutaMAX (Cat # 35050061) were from Gibco; Lipofectamine 2000 (Cat # 11668019) was from Thermo Fisher Scientific; Lipofector-pMAX (Cat # AB-LF-M100) was from AptaBio; Protease inhibitor cocktail (Cat # QTPPI1012) was from Quartett; Albumin (Cat # A0100-010) and Proteinase K (Cat # P2170-001) were from GenDEPOT; The Bio-Rad protein assay dye reagent concentrate (Cat # 500-0006) was from BIO-RAD; Ethidium bromide (Cat # 1239–45–8) was from USB; Magnesium-L-threonate (Cat # QA-8149) was from Combi-Blocks Inc.; DirectPCR (Multi) (Cat # 102) was from FIAT; The Lentiviral Concentrator (Cat # 631231) was from Takara; Polyvinylidene fluoride (Cat # AE-6667-P) was from Atto. PNGase F (Cat # P0704S) was from NEB. FM1-43 (Cat # T3163) was from Invitrogen.
Cell culture and DNA transfection
Mouse primary cortical and hippocampal neurons were isolated from embryonic days 14–17 (E14–E17) of Tusc3 +/+ or Tusc3 -/- C57BL/6 J mice as previously described71. Cells were gently triturated by trypsin-EDTA and were cultured in Neurobasal Medium (Gibco) supplemented with 2% (v:v) B-27 supplement (Gibco), 10 μg/ml gentamycin (Gibco), 100 units/ml penicillin-streptomycin (Gibco), and 1% (v:v) GlutaMAX (Gibco). SH-SY5Y and HEK293T cells (ATCC) were maintained in Dulbecco’s modified Eagle medium (DMEM; Welgene Inc.) containing 10% (v:v) fetal bovine serum (FBS; Gibco), 100 units/ml Penicillin-Streptomycin. Primary neurons were transfected using Lipofectamine 2000 (Thermo Fisher Scientific), and SH-SY5Y and HEK293T cells were transfected using Lipofector-pMax (AptaBio) following the manufacturer’s instructions.
Isolation of mouse embryonic fibroblasts
Tusc3+/- pregnant mice embryos (E13.5) were used for MEFs isolation as previously described with some modifications72. After the heads, tails, limbs, and most internal organs were detached, the embryos were minced and trypsinized for 30 min (shaking per 10 min, 37 °C) and then seeded into 10 cm dishes with DMEM media. MEFs were passaged 2–3 times to obtain a morphologically homogenous culture and then expanded for further studies (split ratio 1:2). The genotyping was performed from embryos tails, and described below. For studies using MagFRETER, cells were transfected using Lipofectamine 2000 Transfection Reagent (Thermo Fisher Scientific) for 48 h. For co-expression, MEFs were transfected with TUSC3-mRFP WT.
Patient-derived fibroblasts
Patient-derived fibroblasts harboring TUSC3 mutations were obtained from a previously published study7 and used in accordance with the French National Ethics Committee guidelines. Written informed consent was obtained from all patients or their legal guardians, and the cell lines were anonymized for research use only. As a control, CCD-986sk human dermal fibroblasts were purchased from the Korean Cell Line Bank (KCLB) and cultured following the supplier’s protocol. Specifically, patient-derived fibroblasts were maintained in CHANG Medium D (T105, Irvine Scientific), while control fibroblasts were grown in Iscove’s Modified Dulbecco’s Medium (IMDM; WELGENE) supplemented with 10% FBS, 1% penicillin–streptomycin, and 1% L-glutamine. Both cell lines were incubated at 37 °C in a humidified atmosphere containing 5% CO₂.
For experiments requiring TUSC3 reconstitution, fibroblasts were transfected with a TUSC3–mRFP expression vector using Lipofectamine™ 2000 (Thermo Fisher Scientific), according to the manufacturer’s protocol. Cells were harvested and analyzed 48 h post-transfection for Mg²⁺ uptake and ER stress markers. Mg²⁺ uptake was assessed by transfecting fibroblasts with the genetically encoded Mg²⁺ sensor Mag-FRETER and examining the cells via confocal microscopy, as previously described18. ER stress markers, including GRP78 and CHOP, were evaluated by western blot, and global protein translation changes were measured using the Surface Sensing of Translation (SUnSET) assay27.
Generation of TUSC3 knockdown cells
To generate TUSC3 KD cells, HEK293T cells were co-transfected with pGIPZ control or shTUSC3 plasmids (each carrying VSV-G and PSPAX2 packaging plasmids) or using Lipofector-pMax (AptaBio), according to the manufacturer’s protocol. After 12 h, the medium was replaced with fresh DMEM supplemented with 10% FBS. Forty-eight hours post-transfection, the virus-containing supernatant was collected, passed through a 0.45 μm filter, and concentrated with a lentiviral concentrator (Takara). For transduction, SH-SY5Y cells or HCT116 cells were incubated with the concentrated lentiviral particles in the presence of 8 μg/mL polybrene for 48 h. The medium was then replaced with fresh growth medium containing 1–3 μg/mL puromycin to select successfully transduced cells. After 24–48 h of selection, single-cell clones were isolated by seeding one cell per well in a 96-well plate. Clonal expansion ensued, and KD efficiency was validated via western blot analysis. pGIPZ constructs were from Dharmacon (Horizon Discovery, United Kingdom). The target sequence of pGIPZ_sh_TUSC3#84562: 5′-TTC AGT ACC AGA ATA ATG T, pGIPZ_sh_TUSC3#84563: 5′-TTT ATC ACC ATT CAT TCG G, and pGIPZ_sh_TUSC3#351871: 5′-TCA GCT AAA AGA TTC TCC T.
Plasmid construction
The cDNAs encoding human TUSC3 and ERMA were amplified by PCR and inserted into the following vectors, pcDNA3.1, mRFP1-N1, and DsRed-N1. Deletion mutants of TUSC3 were produced by site-directed mutagenesis. All plasmid constructs were verified by DNA sequencing analysis. The sgTUSC3 targeting sequences were inserted into Lentivector V2. The target sequence of sgTUSC3 #1 is 5′-CTT GCC TAC GGC GTG AAG GAG; the target sequence of sgTUSC3#2 is 5′-GCT TGC CTA CGG CGT GAA GGA; sgTUSC3 #3 is 5′-CGC TTG CCT ACG GCG TGA AGG.
Western blot analysis
Western blotting was performed following previously established protocols71. Cell lysates and tissue samples from the hippocampus, striatum, and prefrontal cortex were homogenized in ice-cold RIPA buffer containing 50 mM Tris-Cl (pH 7.5), 150 mM NaCl, 1 mM EDTA, 1.1% (v/v) Nonidet P40 substitute, 0.25% (w/v) sodium deoxycholate, and 1 mM PMSF. The samples were then briefly sonicated and centrifuged at 12,000 × g for 10 min at 4 °C to remove debris. The supernatant was subjected to SDS-PAGE and transferred to PVDF (Atto) membrane using the Semi-Dry transfer system WSE-4025 HorizeBLOT 2 M-R (Atto). The membranes were blocked with 5% (w/v) BSA prepared in TBS-T buffer [10 mM Tris-Cl, pH 8.0, 150 mM NaCl, and 0.5% (v/v) Tween-20] for 1 h at room temperature to prevent nonspecific binding. After incubation with primary antibodies overnight at 4 °C, the membranes were washed with TBS-T, followed by incubation with HRP-conjugated secondary antibodies. Immunoreactive bands were detected using chemiluminescent detection methods, and images were captured.
Co-immunoprecipitation
Brain lysates from WT mice were prepared using NP-40 lysis buffer (50 mM Tris-HCl, pH 7.4, 150 mM NaCl, 1% NP-40, 1 mM EDTA) supplemented with a protease inhibitor cocktail (Quartett). The lysates were then briefly sonicated and centrifuged at 12,000 × g for 20 min at 4 °C, and the resulting supernatants were collected and quantified using the Bio-Rad Protein Assay (Bio-Rad). For each immunoprecipitation, 1000 μg of total protein was incubated overnight at 4 °C with 5 μL of antibodies against TUSC3 (Proteintech), ERMA (gift from Dr. Madeshi), or control rabbit IgG (Cell Signaling), under rotation. Protein–antibody complexes were captured by incubation with 40 μL of Protein G Sepharose bead slurry (GE Healthcare) for 4 h at 4 °C. The beads were then washed three times with cold PBS and eluted in 5 × Laemmli sample buffer (100 mM Tris-HCl pH 6.8, 25% glycerol, 5% SDS, 200 mM DTT, 0.02% bromophenol blue), followed by boiling at 95 °C for 5 min. Eluates were analyzed by immunoblotting.
Immunoprecipitation
For overexpression immunoprecipitation, HEK293T cells were transfected with plasmids at a 1:4 ratio using Lipofector-pMax (AptaBio). After 48 h, cells were lysed with lysis buffer (50 mM Tris-Cl, pH 7.4, 150 mM NaCl, 1 mM EDTA, 1% Triton X-100) supplemented with a protease inhibitor cocktail (Quartett) and 1 mM PMSF (Sigma-Aldrich), and cleared by centrifugation (12,000 × g, 10 min, at 4 °C). The resulting supernatant was transferred to a new tube and incubated with the primary antibody in lysis buffer overnight at 4 °C. Protein G Sepharose bead slurry (GE Healthcare) or FLAG M2 bead slurry (Sigma-Aldrich) was added and incubated at 4 °C for 4 h. The beads were washed three times with PBS, eluted with 5 × sample buffer (100 mM Tris-Cl pH 6.8, 25% glycerol, 5% SDS, 200 mM DTT, 0.02% bromophenol blue) followed by 5 min boiling at 95 °C, and analyzed by immunoblotting.
PNGase F treatment
Enzymatic deglycosylation was performed using PNGase F (NEB) under denaturing conditions following the manufacturer instructions. Briefly, total glycoprotein (20 μg) was mixed with 1 μL of 10 × Glycoprotein Denaturing Buffer and nuclease-free water to a final volume of 10 μL. The mixture was denatured by incubation at 100 °C for 10 min, followed by immediate chilling on ice. Samples were briefly centrifuged at 10,000 × g for 10 s and brought to a final reaction 20 μL by adding 2 μL of 10 × GlycoBuffer 2, 2 μL of 10% NP-40, and 6 μL of nuclease-free water. After the addition of PNGase F (500,000 units/ml, 1 μL), the reaction mixture was incubated at 37 °C for 2 h and evaluated by observing protein mobility shifts using immunoblotting.
Immunohistochemistry
Mice were anesthetized and transcardially perfused with PBS containing 10 U/mL heparin, followed by fixation with 4% (w/v) paraformaldehyde (PFA). Brains were post-fixed overnight at 4 °C and subsequently transferred to 30% (w/v) sucrose solution for cryoprotection. The tissues were embedded in OCT compound and sectioned coronally into 40-μm-thick cryosections using a Cryostat microtome (Leica). For floating immunohistochemistry, sections were washed three times in PBS and incubated in blocking solution (5% BSA and 0.1% Triton X-100 in PBS) for 2 h at room temperature. The sections were then incubated overnight at 4 °C with primary antibodies diluted in blocking solution. After washing with PBST, sections were incubated with fluorescently conjugated secondary antibodies for 1 h at room temperature. Following additional PBST washes, sections were mounted onto slides using FluoromountTM Aqueous Mounting Medium (Sigma-Aldrich). Images were captured using LSM980 confocal microscopy (Zeiss), and fluorescence intensity analysis was performed using ImageJ.
Magnesium measurement
Cells were transfected with MagFRETCyto or MagFRETER for 48 h and analyzed using confocal microscopy as described in a previously published study18. To assess intracellular magnesium levels, cells were seeded onto 35 mm confocal dishes (PLL-coating dishes for primary cortical neurons) and transfected with MagFRETCyto or MagFRETER for 48 h. Live-cell imaging was performed using an LSM980 confocal microscope (Zeiss) equipped with FRET detection modules to monitor Mg²⁺ dynamics in real-time. For FRET analysis, cells were excited at 435 nm using a single excitation laser, and fluorescence emissions were detected at 460–490 nm (Cerulean channel) and 510–550 nm (Citrine channel) to quantify energy transfer. The Citrine-to-Cerulean ratio was used to determine intracellular Mg²⁺ levels. All images were acquired at 37 °C using a 63× (NA 1.4) oil-immersion objective, and data were analyzed using built-in FRET SE analysis modules in Zen lite 2.3 software (Zeiss). To compare ER Mg²⁺ uptake across experimental groups, FRET efficiency (Citrine/Cerulean ratio) was normalized to baseline values of control for signal variability. To assess whether the Mg²⁺ deficiency in TUSC3 KD cells could be acutely rescued, cells were treated with 10 mM MgCl₂ for 200 s, and real-time changes in the Citrine/Cerulean ratio were measured.
Spine density measurement and Sholl analysis
Dendritic spine density was assessed as previously described71. In brief, hippocampal neurons (DIV 12) were transfected with pEGFP-N1 using Lipofectamine 2000 (Thermo Fisher Scientific). At DIV 14, neurons were fixed with 4% (w/v) PFA for 10 min at room temperature. Confocal microscope (LSM980, Zeiss) was used to capture images of at least 10 randomly selected EGFP-expressing neurons. Dendritic spines were manually counted along 10 μm dendritic segments using ImageJ (NIH), with classification based on morphological characteristics.
To quantify dendritic complexity, Sholl analysis was performed using ImageJ with the Sholl Analysis plugin73. Concentric circles were drawn around the soma at 10 μm intervals, and the number of dendritic intersections at each radius was counted. The Sholl regression coefficient (Rc), which quantifies dendritic branching patterns, was calculated by fitting a logarithmic regression curve to the intersection data. Statistical analysis was performed using two-way ANOVA, comparing WT and TUSC3 KO neurons across different radii to determine differences in dendritic complexity. All experiments were conducted in at least three independent cultures, with a minimum of 15 neurons per condition analyzed.
FM1-43 dye labeling and unloading assay
FM1-43 dye labeling and unloading assays were performed as previously described, with minor modifications74. To assess synaptic vesicle recycling, primary hippocampal neurons (DIV18) were first pre-washed with pre-warmed HEPES-buffered saline (HBS; 135 mM NaCl, 5 mM KCl, 2 mM CaCl₂, 1 mM MgCl₂, 10 mM glucose, and 10 mM HEPES, pH 7.4) and then incubated for 5 min in high K⁺ HBS (90 mM KCl) containing 10 μM FM1-43 dye (Invitrogen). After additional exposure to FM1-43 for 60 s, neurons were washed three times for 30 s with HBS and further rinsed 3 times with dye-free normal saline and then allowed to rest for 10 min. For dye release, cells were re-stimulated with high K⁺ HBS for 2 min while live fluorescence imaging was performed every 10 s using a confocal microscope (LSM980) equipped with a 488 nm laser and a 20× objective. The fluorescence intensities in individual synaptic boutons were quantified using ImageJ (Fiji) as previously described75. All values were background-subtracted and normalized to baseline fluorescence (F0), and unloading kinetics were assessed by analyzing the slope and total change in fluorescence (ΔF = F200 – F0).
Cell death assay
Cell Live:Dead ratios were determined as described previously71. SH-SY5Y cells were pretreated with Magnesium-L-Threonate (MgT; 400 μM) for 12 h and, cells were then cotreated with MgT (400 μM) and ER stress inducers (thapsigargin, 2 μM; tunicamycin, 2 μg/ml; A23187, 2 μM), and etoposide (25 μM) for 24 h. Viable cells were stained with calcein-AM, originally non-fluorescent but converted to fluorescent form under hydrolysis by intracellular esterases in live cells. Dead cells were stained with propidium iodide (PI), observed under a fluorescence microscope (Olympus), and the Live:Dead ratios were calculated by dividing the number of dead cells (PI-positive cells) by the number of total cells (calcein-AM-positive cells plus PI-positive cells).
Surface sensing of translation (SUnSET) assay
The SunSET assay was performed to assess global protein synthesis following previously established protocols with minor modifications27. MEFs and human fibroblasts were cultured under standard conditions before treatment. Cells were incubated with 5 μg/mL puromycin (Sigma-Aldrich) for 2 h at 37 °C to allow incorporation of puromycin into nascent polypeptide chains. Following incubation, cells were washed three times with ice-cold PBS to remove excess puromycin and then lysed using RIPA buffer. For western blot analysis, total protein extracts were separated by SDS-PAGE, transferred to PVDF membranes, and blocked with 5% (w/v) BSA in TBS-T. Membranes were then probed with anti-puromycin antibody (ABclonal) overnight at 4 °C to detect puromycin-labeled polypeptides, followed by incubation with HRP-conjugated secondary antibodies. Immunoreactive bands were visualized using chemiluminescent detection.
Animals experimentations
All experiments involving animals were performed according to the guidelines of this period according ARRIVE guidelines and the protocols were approved by the Institutional Animal Care and Use Committee guidelines of Seoul National University (240102-6-1). WT or TUSC3 KO mice on a C57BL/6 J background were generated using CRISPR-Cas9 gene-editing technology. Both male and female mice were used across all experiments. Mice were randomly assigned to experimental groups regardless of sex, as the study was not specifically designed or powered to investigate sex-dependent differences. Detailed information on the age and sex of animals used is indicated in the figure legends for each experiment using the abbreviations F (female) and M (male). All animals were housed in groups of 3–5 per cage in a temperature-controlled (22 ± 1 °C) and humidity-controlled (50–60%) facility with a 12 h light/dark cycle. Standard rodent chow and water were provided ad libitum.
Generation of TUSC3 KO mice
TUSC3 KO mice were generated by ToolGen Inc. and Macrogen Inc. (Seoul, Korea) using CRISPR-Cas9 gene-editing technology. Specifically, a single guide RNA (sgRNA) targeting a highly conserved region within Exon 2 of the mouse Tusc3 gene was designed (sequence: 5′-ATC ATG GAG TAG TTC CGA GGT GG-3′). The sgRNA was synthesized and validated in vitro for cleavage efficiency. Founder mice were screened for targeted mutations by PCR amplification of genomic DNA isolated from tail biopsies, followed by Sanger sequencing. The resulting targeted mutation in the Tusc3 locus involved a 363 bp deletion spanning Exon 2 and part of the intron between Exon 2 and Exon 3, leading to a frameshift mutation and the introduction of a premature stop codon. Successful generation of TUSC3 KO mice was further validated by western blot analysis and genotyping.
Genotyping of TUSC3 KO mice
Genomic DNA was extracted from mouse tail biopsies using DirectPCR lysis buffer (Biofiat) supplemented with Proteinase K (GenDEPOT). Specifically, 49 µL of DirectPCR lysis buffer was mixed with 1 µL of Proteinase K for each sample. After overnight digestion at 55 °C, the lysates were heat-inactivated (85–95 °C for 45 min) and used as templates for PCR-based genotyping. PCR was performed using TUSC3-specific primers designed to distinguish WT and KO alleles. The 20 µL reaction consisted of genomic DNA, primers (forward primer: 5′-GTG CAA GTA AGG TCA ATG AGG A-3′, reverse primer: 5′-CCA CCC AGA CAA TCA CTG AAT-3′), and PCR master mix. A touchdown PCR protocol was employed for higher specificity: an initial denaturation at 95 °C for 5 min was followed by 10 cycles of 95 °C for 30 s, 57 → 50 °C (decreasing 1 °C per cycle) for 30 s, and 72 °C for 30 s. This was followed by 25 cycles of 95 °C for 30 s, 50 °C for 30 s, and 72 °C for 30 s. The reaction concluded with a final extension at 72 °C for 5 min. Amplified DNA fragments were separated on a 2% agarose gel, stained with ethidium bromide, and visualized under UV illumination. The expected band sizes for the WT and KO alleles were approximately 800 bp and 400 bp, respectively.
Magnesium-L-threonate treatment
Magnesium-L-threonate (MgT) treatment was administered following previously reported dosing protocols11,65,67. WT and TUSC3 KO mice were provided with MgT-supplemented drinking water at a concentration of 910 mg/kg/day (mpk), ensuring consistent intake throughout the experimental period. Fresh MgT solution was prepared and replenished every 48 h to maintain stability and efficacy. Control mice received standard drinking water without supplementation.
Memory tests
Y-maze test
Mice were laid to the end of one arm and then explored freely in the apparatus (32.5 cm long × 15 cm high) for 7 min. The arm entry was valid when the whole body was entered entirely into each arm. The spontaneous alteration was assessed as the ratio of the number of effective alterations, an event that the mouse enters three different arms consecutively, to the number of total arm entries.
Novel object recognition
NOR test was performed with two days of habituation, one-day training, and two days of the test session. Mice were allowed to freely roam in the chamber (22 cm wide × 27 cm long × 30 cm high) for 7 min. At training, two identical objects were placed on the diagonal of the chamber. One object was replaced with a novel object once a day during the test phases. The discrimination ratio was calculated by dividing the time spent on a novel object by the sum spent on a novel object and a familiar object.
Passive avoidance
Passive avoidance was composed of one-day habituation, one-day training, and one-day test session. The apparatus (40 cm wide × 20 cm long × 20 cm high) is divided into a bright compartment with an overhead 8 W lamp and a dark compartment, of which the floor is made of electrical grids. Mice were allowed to explore freely in a bright compartment for 3 min with the door closed for conditioning. At the training session, mice were initially placed into the bright compartment and shocked by the foot grid (0.25 mA, 2 s) when both hind limbs entered the dark compartment. The latency to enter the dark compartment at the test session was observed with a 7 min cutoff point.
Forced swim test
The forced swim test was conducted to evaluate stress adaptation capability. Each mouse was placed in a container (20 cm high × 14 cm diameter) filled with water at 25 °C for 6 min. Water level was set to 16 cm. All experiments were recorded using a video camera and later analyzed to measure the immobility time. Immobility time, defined as the duration during which the mouse remained floating with minimal movement to keep its head above water, was measured. This was calculated by subtracting the total active movement time from the total test duration. Mice that exhibited signs of drowning or extreme distress were immediately removed from the water, and the time at which they were taken out was recorded. These mice were excluded from further analysis. After the test, all mice were carefully dried, placed in a pre-warmed recovery cage, and monitored before being returned to their home cages.
Tail suspension test
Each mouse was suspended by the tail using adhesive tape affixed approximately 1 cm from the tail tip and secured to a horizontal bar. The test was conducted in a soundproof behavioral testing room under consistent lighting conditions to minimize external stressors. Mice were suspended at a height ensuring that their bodies remained completely airborne and unable to contact with any surfaces. The total duration of the test was 6 min, during which the immobility time—defined as the absence of active escape-oriented movements—was recorded.
Three-chamber social interaction and social novelty test
The three-chamber social test was conducted as previously described76, with minor modifications. The testing apparatus consisted of a transparent acrylic box divided into three chambers, with small openings allowing movement between them. The wire cages used to contain stranger mice were cylindrical (11 cm height, 10.5 cm bottom diameter, 1 cm bar spacing).
For the sociability test, the test mouse was first placed in the middle chamber for 5 min for habituation. Then, a novel mouse was placed inside a wire cage in one of the side chambers, while an empty wire cage was placed in the opposite chamber. The test mouse was allowed to explore freely for 10 min and time spent in each chamber and the proximity to the cages were recorded.
For the social novelty test, after an additional 5 min of exposure to the familiar mouse, a second novel mouse was placed in the previously empty cage. The test mouse was again allowed to explore for 10 min, and social interaction was analyzed. Behavioral data, including time spent in each chamber and proximity to the cages, were quantified. The placement of Stranger 1 was counterbalanced across trials, but remained consistent within each test session.
Self-grooming test
Mice were scored for spontaneous grooming behaviors when placed individually in a clean cage. The cage was lined with a thin layer of bedding (~1 cm) in order to reduce neophobia, but prevent digging, a potentially competing behavior. Prior to the testing period, animals were allowed to habituate to the novel environment for 10 min. Each mouse was rated for 10 min on cumulative time spent grooming.
Ex vivo electrophysiology
Slice preparation
Using WT and TUSC3 KO mice, transverse hippocampal slices (300 μm) were prepared as described previously77. The mice were anesthetized using isoflurane and decapitated. Their brains were rapidly removed and placed in ice-cold, oxygenated (95% O2 and 5% CO2) dissection buffer with low Ca2+ and high Mg2+ concentrations. The dissection buffer consisted of 5 mM KCl, 1.23 mM NaH2PO4, 26 mM NaHCO3, 10 mM dextrose, 0.5 mM CaCl2, 10 mM MgCl2, and 212.7 mM sucrose. Hippocampal slices were cut and transferred to a holding chamber in an incubator filled with oxygenated artificial cerebrospinal fluid (ACSF). The ACSF contained 124 mM NaCl, 5 mM KCl, 1.23 mM NaH2PO4, 2.5 mM CaCl2, 1.5 mM MgCl2, 26 mM NaHCO3, and 10 mM dextrose. The slices were maintained at 28–30 °C for at least 30 min before recording.
Whole-cell recordings
The slices were then moved to a recording chamber and continuously perfused (2 ml/min) with oxygenated ACSF (95% O2 and 5% CO2). Recordings from CA1 and CA3 pyramidal neurons were performed, and patch pipettes (4–8 MΩ resistance) were filled with Cs-based internal solution. The solution comprised 130 mM Cs-MeSO4, 0.5 mM EGTA, 5 mM TEA-Cl, 8 mM NaCl, 10 mM HEPES, 1 mM QX-314, 4 mM ATP-Mg, 0.4 mM GTP-Na, and 10 mM Na2-phosphocreatine at a pH of 7.4 and 290–300 mOsm. The external recording solution was composed of ACSF, to which 100 μM picrotoxin was added. Neurons were voltage-clamped at −60 mV to record sEPSC, eEPSC, and paired pulse ratio (EPSC-PPR). Evoked synaptic responses were elicited by stimulating the Schaffer collateral (SC) pathway (0.2 ms current pulses) using a concentric bipolar electrode placed adjacent to the CA1 region. The maximal amplitudes of eEPSC were measured, and the half-maximal amplitude was used to record PPR. To measure EPSC-PPR, two synaptic responses were evoked by paired stimuli given at 20 ms intervals. Recordings were obtained using a Multiclamp 700 A amplifier (Molecular Devices) under visual control with differential interference contrast illumination in an upright microscope (Olympus, BX50WI). Only cells with an access resistance <20 MΩ were recorded; those with an access resistance change >20% were excluded from the analysis. Data were acquired and analyzed using pClamp 10.7 (Molecular Devices). Signals were filtered at 3 kHz and digitized at 10 kHz with Digidata 1550B (Molecular Devices).
Neuromuscular junction (NMJ) recordings
The extensor digitorum longus (EDL) muscle was dissected from TUSC3 WT and TUSC3 KO mice and then pinned to a Sylgard-coated recording chamber as described previously78. Intracellular recording was conducted in oxygenated Ringer’s solution (138.8 mM NaCl, 4 mM KCl, 12 mM NaHCO₃, 1 mM KH₂PO₄, 1 mM MgCl₂, and 2 mM CaCl₂ with a pH of 7.4). Miniature endplate potentials (mEPPs) were recorded from a junction. The data were obtained and analyzed with Axoclamp 900 A and Clampfit version 10.7 software.
Statistical analysis
All experiments were conducted independently at least three times, and data are presented as mean ± S.E.M. unless otherwise indicated. The statistical tests used and sample sizes (n) are detailed in the figure legends. Statistical analyses were performed using GraphPad Prism 10.1.0. (GraphPad). Normality was performed using the Shapiro–Wilk or Kolmogorov–Smirnov (KS) test, depending on the sample size, when the dataset was sufficiently large. If normality was not met, non-parametric tests were applied. Otherwise, an unpaired two-tailed t-test was used for comparisons between two groups, one-way ANOVA for comparisons involving three or more groups with a single independent variable, and two-way ANOVA for analyses involving two independent variables, followed by Tukey’s post-hoc test for multiple comparisons. Statistical significance was defined as P < 0.05. Graphs and figures were generated using GraphPad Prism version 10.1.0.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data generated in this study are provided in the Source Data file. Source data are provided with this paper.
References
Maulik, P. K., Mascarenhas, M. N., Mathers, C. D., Dua, T. & Saxena, S. Prevalence of intellectual disability: a meta-analysis of population-based studies. Res. Dev. Disabil. 32, 419–436 (2011).
Basel-Vanagaite, L. Genetics of autosomal recessive non-syndromic mental retardation: recent advances. Clin. Genet. 72, 167–174 (2007).
Vissers, L. E., Gilissen, C. & Veltman, J. A. Genetic studies in intellectual disability and related disorders. Nat. Rev. Genet. 17, 9–18 (2016).
Jamra, R. Genetics of autosomal recessive intellectual disability. Med. Genet. 30, 323–327 (2018).
Albobali, Y., Shahwan, M. Y., Madi, M. Y. & Al-Nuaimi, S. Two Sisters from Qatar with TUSC3 genetic mutation: psychiatric considerations. Cureus 13, e17616 (2021).
Gumus, E. A homozygous 1.16 megabases microdeletion at 8p22 including the whole TUSC3 in a three years old girl with intellectual disability and speech delay. Cell J. 22, 128–132 (2020).
El Chehadeh, S. et al. Homozygous truncating intragenic duplication in TUSC3 responsible for rare autosomal recessive nonsyndromic intellectual disability with no clinical or biochemical metabolic markers. JIMD Rep. 20, 45–55 (2015).
Molinari, F. et al. Oligosaccharyltransferase-subunit mutations in nonsyndromic mental retardation. Am. J. Hum. Genet. 82, 1150–1157 (2008).
Ozmansur, E. N., Peduk, Y., Gumus, H., Caglayan, A. O. & Per, H. A novel homozygous frameshift mutation in the TUSC3 gene identified in siblings with intellectual disability. Clin. Dysmorphol. 31, 36–38 (2022).
Zhou, H. & Clapham, D. E. Mammalian MagT1 and TUSC3 are required for cellular magnesium uptake and vertebrate embryonic development. Proc. Natl. Acad. Sci. USA 106, 15750–15755 (2009).
Li, W. et al. Elevation of brain magnesium prevents synaptic loss and reverses cognitive deficits in Alzheimer’s disease mouse model. Mol. Brain 7, 65 (2014).
Slutsky, I. et al. Enhancement of learning and memory by elevating brain magnesium. Neuron 65, 165–177 (2010).
Wang, C. et al. ATF5 attenuates low-magnesium-induced apoptosis by inhibiting endoplasmic reticulum stress via the regulation of mitochondrial reactive oxygen species. Neuroscience 535, 13–22 (2023).
Hetz, C. & Saxena, S. ER stress and the unfolded protein response in neurodegeneration. Nat. Rev. Neurol. 13, 477–491 (2017).
Mohorko, E. et al. Structural basis of substrate specificity of human oligosaccharyl transferase subunit N33/Tusc3 and its role in regulating protein N-glycosylation. Structure 22, 590–601 (2014).
Zolotarov, Y. et al. ARL15 modulates magnesium homeostasis through N-glycosylation of CNNMs. Cell Mol. Life Sci. 78, 5427–5445 (2021).
Kostantin, E. et al. Inhibition of PRL-2.CNNM3 protein complex formation decreases breast cancer proliferation and tumor growth. J. Biol. Chem. 291, 10716–10725 (2016).
Vishnu, N. et al. ERMA (TMEM94) is a P-type ATPase transporter for Mg(2+) uptake in the endoplasmic reticulum. Mol. Cell 84, 1321–1337.e1311 (2024).
Daw, C. C. et al. Lactate elicits ER-mitochondrial Mg(2+) dynamics to integrate cellular metabolism. Cell 183, 474–489.e417 (2020).
Lindenburg, L. H., Vinkenborg, J. L., Oortwijn, J., Aper, S. J. & Merkx, M. MagFRET: the first genetically encoded fluorescent Mg2+ sensor. PLoS ONE 8, e82009 (2013).
Maguire, M. E. MgtA and MgtB: prokaryotic P-type ATPases that mediate Mg2+ influx. J. Bioenerg. Biomembr. 24, 319–328 (1992).
Knoop, V., Groth-Malonek, M., Gebert, M., Eifler, K. & Weyand, K. Transport of magnesium and other divalent cations: evolution of the 2-TM-GxN proteins in the MIT superfamily. Mol. Genet. Genom. 274, 205–216 (2005).
Kuhlbrandt, W. Biology, structure and mechanism of P-type ATPases. Nat. Rev. Mol. Cell Biol. 5, 282–295 (2004).
Du, K. & Montminy, M. CREB is a regulatory target for the protein kinase Akt/PKB. J. Biol. Chem. 273, 32377–32379 (1998).
Zhu, P. J. et al. Activation of the ISR mediates the behavioral and neurophysiological abnormalities in Down syndrome. Science 366, 843–849 (2019).
Harding, H. P., Zhang, Y., Bertolotti, A., Zeng, H. & Ron, D. Perk is essential for translational regulation and cell survival during the unfolded protein response. Mol. Cell 5, 897–904 (2000).
Schmidt, E. K., Clavarino, G., Ceppi, M. & Pierre, P. SUnSET, a nonradioactive method to monitor protein synthesis. Nat. Methods 6, 275–277 (2009).
Zinszner, H. et al. CHOP is implicated in programmed cell death in response to impaired function of the endoplasmic reticulum. Genes Dev. 12, 982–995 (1998).
Yam, G. H., Gaplovska-Kysela, K., Zuber, C. & Roth, J. Sodium 4-phenylbutyrate acts as a chemical chaperone on misfolded myocilin to rescue cells from endoplasmic reticulum stress and apoptosis. Investig. Ophthalmol. Vis. Sci. 48, 1683–1690 (2007).
Tasse, M.J. & Kim, M. Examining the relationship between adaptive behavior and intelligence. Behav. Sci. 13, 252 (2023).
Slattery, D. A. & Cryan, J. F. Using the rat forced swim test to assess antidepressant-like activity in rodents. Nat. Protoc. 7, 1009–1014 (2012).
Commons, K. G., Cholanians, A. B., Babb, J. A. & Ehlinger, D. G. The rodent forced swim test measures stress-coping strategy, not depression-like behavior. ACS Chem. Neurosci. 8, 955–960 (2017).
de Kloet, E. R. & Molendijk, M. L. Coping with the forced swim stressor: towards understanding an adaptive mechanism. Neural Plast. 2016, 6503162 (2016).
Picci, G. & Scherf, K. S. A two-hit model of autism: adolescence as the second hit. Clin. Psychol. Sci. 3, 349–371 (2015).
Blanpied, T. A. & Ehlers, M. D. Microanatomy of dendritic spines: emerging principles of synaptic pathology in psychiatric and neurological disease. Biol. Psychiatry 55, 1121–1127 (2004).
Verpelli, C. & Sala, C. Molecular and synaptic defects in intellectual disability syndromes. Curr. Opin. Neurobiol. 22, 530–536 (2012).
Craven, S. E. & Bredt, D. S. PDZ proteins organize synaptic signaling pathways. Cell 93, 495–498 (1998).
Moore, A. N., Waxham, M. N. & Dash, P. K. Neuronal activity increases the phosphorylation of the transcription factor cAMP response element-binding protein (CREB) in rat hippocampus and cortex. J. Biol. Chem. 271, 14214–14220 (1996).
Baez-Mendoza, R. & Schultz, W. The role of the striatum in social behavior. Front. Neurosci. 7, 233 (2013).
Eichenbaum, H. Hippocampus: cognitive processes and neural representations that underlie declarative memory. Neuron 44, 109–120 (2004).
Wiedenmann, B. & Franke, W. W. Identification and localization of synaptophysin, an integral membrane glycoprotein of Mr 38,000 characteristic of presynaptic vesicles. Cell 41, 1017–1028 (1985).
Chesnokova, V., Pechnick, R. N. & Wawrowsky, K. Chronic peripheral inflammation, hippocampal neurogenesis, and behavior. Brain Behav. Immun. 58, 1–8 (2016).
Trinh, M. A. & Klann, E. Translational control by eIF2alpha kinases in long-lasting synaptic plasticity and long-term memory. Neurobiol. Learn Mem. 105, 93–99 (2013).
Park, S.M., Kang, T.I. & So, J.S. Roles of XBP1s in transcriptional regulation of target genes. Biomedicines 9, 791 (2021).
Yoshida, H. et al. ATF6 activated by proteolysis binds in the presence of NF-Y (CBF) directly to the cis-acting element responsible for the mammalian unfolded protein response. Mol. Cell Biol. 20, 6755–6767 (2000).
Lanzillotta, C. et al. Chronic PERK induction promotes Alzheimer-like neuropathology in Down syndrome: Insights for therapeutic intervention. Prog. Neurobiol. 196, 101892 (2021).
Coulson, R. L. et al. Translational modulator ISRIB alleviates synaptic and behavioral phenotypes in Fragile X syndrome. iScience 27, 109259 (2024).
Luhong, L. et al. PERK inhibitor (ISRIB) improves depression-like behavior by inhibitions of HPA-axis over-activation in mice exposed to chronic restraint stress. Behav. Brain Res. 471, 115122 (2024).
Matsuda-Lennikov, M. et al. Magnesium transporter 1 (MAGT1) deficiency causes selective defects in N-linked glycosylation and expression of immune-response genes. J. Biol. Chem. 294, 13638–13656 (2019).
Kolisek, M., Nestler, A., Vormann, J. & Schweigel-Rontgen, M. Human gene SLC41A1 encodes for the Na+/Mg(2)+ exchanger. Am. J. Physiol. Cell Physiol. 302, C318–C326 (2012).
Sahni, J., Nelson, B. & Scharenberg, A. M. SLC41A2 encodes a plasma-membrane Mg2+ transporter. Biochem J. 401, 505–513 (2007).
Gao, X. et al. Palmitoylation regulates cellular distribution of and transmembrane Ca flux through TrpM7. Cell Calcium 106, 102639 (2022).
Mousain-Bosc, M. et al. Improvement of neurobehavioral disorders in children supplemented with magnesium-vitamin B6. I. Attention Deficit Hyperactivity Disorders. Magnes. Res. 19, 46–52 (2006).
Starobrat-Hermelin, B. & Kozielec, T. The effects of magnesium physiological supplementation on hyperactivity in children with attention deficit hyperactivity disorder (ADHD). Positive response to magnesium oral loading test. Magnes. Res. 10, 149–156 (1997).
Almalki, A. H. et al. Exploring the association between serum magnesium level and autism spectrum disorder using validated spectrofluorimetric method. Anal. Biochem. 699, 115755 (2025).
Hemamy, M. et al. The effect of vitamin D and magnesium supplementation on the mental health status of attention-deficit hyperactive children: a randomized controlled trial. BMC Pediatr. 21, 178 (2021).
Yadav, V. K., Amrita, A., Yadav, S., Kumar, R. & Yadav, K. K. Role of magnesium supplementation in children with west syndrome: a randomized controlled clinical trial. Iran. J. Child Neurol. 16, 65–75 (2022).
Luo, Z. et al. TRPM7 in neurodevelopment and therapeutic prospects for neurodegenerative disease. Cell Calcium 120, 102886 (2024).
Sturgeon, M., Wu, P. & Cornell, R. SLC41A1 and TRPM7 in magnesium homeostasis and genetic risk for Parkinson’s disease. J. Neurol. Neuromed. 1, 23–28 (2016).
Jiang, Z.J. et al. TRPM7 is critical for short-term synaptic depression by regulating synaptic vesicle endocytosis. Elife 10, e66709 (2021).
Pfaffenseller, B. et al. Impaired endoplasmic reticulum stress response in bipolar disorder: cellular evidence of illness progression. Int. J. Neuropsychopharmacol. 17, 1453–1463 (2014).
Kawada, K. & Mimori, S. Implication of endoplasmic reticulum stress in autism spectrum disorder. Neurochem. Res. 43, 147–152 (2018).
Liu, D. C., Lee, K. Y., Lizarazo, S., Cook, J. K. & Tsai, N. P. ER stress-induced modulation of neural activity and seizure susceptibility is impaired in a fragile X syndrome mouse model. Neurobiol. Dis. 158, 105450 (2021).
Kanekura, K., Suzuki, H., Aiso, S. & Matsuoka, M. ER stress and unfolded protein response in amyotrophic lateral sclerosis. Mol. Neurobiol. 39, 81–89 (2009).
Li, W. et al. Elevation of brain magnesium prevents and reverses cognitive deficits and synaptic loss in Alzheimer’s disease mouse model. J. Neurosci. 33, 8423–8441 (2013).
Schwartz, R., Spencer, H. & Welsh, J. J. Magnesium absorption in human subjects from leafy vegetables, intrinsically labeled with stable 26Mg. Am. J. Clin. Nutr. 39, 571–576 (1984).
Xiong, Y. et al. Magnesium-L-threonate exhibited a neuroprotective effect against oxidative stress damage in HT22 cells and Alzheimer’s disease mouse model. World J. Psychiatry 12, 410–424 (2022).
Nair, A. B. & Jacob, S. A simple practice guide for dose conversion between animals and human. J. Basic Clin. Pharm. 7, 27–31 (2016).
Ajib, F.A. & Childress, J.M. Magnesium Toxicity. In StatPearls (Treasure Island, 2025).
Rosanoff, A., Weaver, C. M. & Rude, R. K. Suboptimal magnesium status in the United States: are the health consequences underestimated? Nutr. Rev. 70, 153–164 (2012).
Kim, S. H. et al. Endolysosomal impairment by binding of amyloid beta or MAPT/Tau to V-ATPase and rescue via the HYAL-CD44 axis in Alzheimer disease. Autophagy 19, 2318–2337 (2023).
Qiu, L.Q., Lai, W.S., Stumpo, D.J. & Blackshear, P.J. Mouse embryonic fibroblast cell culture and stimulation. Bio. Protoc. 6, e1859 (2016).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Gaffield, M. A. & Betz, W. J. Imaging synaptic vesicle exocytosis and endocytosis with FM dyes. Nat. Protoc. 1, 2916–2921 (2006).
Cousin, M. A., Gordon, S. L. & Smillie, K. J. Using FM dyes to monitor clathrin-mediated endocytosis in primary neuronal culture. Methods Mol. Biol. 1847, 239–249 (2018).
Yang, M., Silverman, J. L. & Crawley, J. N. Automated three-chambered social approach task for mice. Curr. Protoc. Neurosci. 8, 26 (2011).
Choi, T. Y. et al. Cereblon maintains synaptic and cognitive function by regulating BK channel. J. Neurosci. 38, 3571–3583 (2018).
Hann, S.H. et al. Depletion of SMN protein in mesenchymal progenitors impairs the development of bone and neuromuscular junction in spinal muscular atrophy. Elife 12, RP92731 (2024).
Acknowledgements
We would like to thank Dr. Muniswamy Madesh (University of Texas Health San Antonio, USA) for anti-ERMA antibody, MagFRETER, and MagFRETCyto and Dr. Michel L. Tremblay (McGill University, Canada) for FLAG-CNNM3 WT and N73A constructs. We also thank Dr. Faivre Laurence (CHU Dijon, France) for providing intellectual disability patient-derived fibroblasts. We would also like to appreciate Hyunmoon Shin (VISYOUR STUDIO, Korea) for providing technical assistance in drawing the working model. This work was supported by a CRI grant (RS-2025-00519823) and the Bio-medical Technology Development Program grant (RS 2024-00439842) funded by the National Research Foundation of Korea (NRF), Ministry of Science and ICT, and by the Korea Dementia Research Project (RS-2023-KH134817) through the Korea Dementia Research Center (KDRC), funded by the Ministry of Health & Welfare and Ministry of Science and ICT, Korea to Y.K.J.
Author information
Authors and Affiliations
Contributions
G.-R.P. and Y.-K.J. contributed to designing the study. G.-R.P. contributed to most experiments; G.-R.P. and N.-H.K. contributed to primary neuronal cultures and immunohistochemistry; C.-M.L. contributed to sgTUSC3 HCT116 cells; S.-Y.K. and S.-Y.C. contributed to electrophysiological experiments; H.-J.L. contributed to TUSC3 KO mouse; G.-R.P., J.-H.S., and Y.-K.J. contributed to analyzing data; G.-R.P. and Y.-K.J. contributed to writing the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Kotaro Oka and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Park, G., Kim, N., Kim, SY. et al. TUSC3 regulates ERMA-mediated Mg2+ uptake for synaptic function and neurodevelopment. Nat Commun 16, 9752 (2025). https://doi.org/10.1038/s41467-025-65668-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-65668-1