Abstract
Colorectal cancer (CRC) development involves microbial and metabolic dysbiosis, with gut microbial β-glucuronidases (gmGUSs) potentially impacting carcinogenesis through de-glucuronidation of diverse important molecules. Here, we identify 550 gmGUSs from a public cohort, employing 114 alignment references, three structural domains, and seven conserved residues. Stage-specific shifts include enrichment of mini-Loop2 (a category defined by two active site-adjacent loop regions) and species-level gmGUS dysregulation (e.g., Bacteroides cellulosilyticus) in CRC. GUS biomarkers display modest efficacy in classifying CRC and adenoma patients from controls, though with limited generalizability, and in predicting CRC outcomes (AUCs > 0.8). Taxonomic and metabolic association analyses highlight microbe-gmGUS-metabolite (MGM) axis perturbations, including increased Alistipes and Fusobacterium, enriched mucin and flavonoid degraders, as well as amino acid and vitamin metabolism alterations linked to CRC progression. In vitro enzyme assays show that the identified gmGUSs possess differential substrate activities. Furthermore, RNA-seq of HCT116 cells co-cultured with BC.G3 (one of the differential gmGUSs from B. cellulosilyticus) reveals upregulation of RNA transcription, DNA replication, and protein folding, shedding preliminary light on its potential effects in CRC progression. Here, we define disturbance of MGM axis in colorectal tumorigenesis and offer potential early diagnostic biomarkers and therapeutic targets for CRC.
Similar content being viewed by others
Introduction
Host UDP-glucuronosyltransferases (UGTs)-mediated glucuronidation is one of the common metabolic pathways of numerous biologically important endogenous compounds, like hormones, neurotransmitters, and bile acids (BAs), as well as xenobiotics, like drugs and dietary carcinogens1. A recent study of urinary glucuronide profiling unraveled 685 structurally diverse glucuronides with 181 annotated2, underscoring the importance of glucuronidation in metabolic homeostasis. Gut microbial β-glucuronidases (gmGUSs, E.C. 3.2.1.31) are enzymes from the glycoside hydrolase (GH) family (GH family 2 in the majority and GH family 30 and 79 the minority) that catalyze the hydrolysis of glucuronidated conjugates of endogenous or exogenous compounds to release the aglycones3. The deconjugation reaction reverses the host UGTs-mediated glucuronidation process, facilitates the reabsorption and enterohepatic recirculation1. The liver UGTs-gmGUSs axis works coordinately to maintain the glucuronidation homeostasis and determine the circulating level and local intestinal exposure of the parent compounds and their metabolites.
Colorectal cancer (CRC) is a worldwide leading cause of cancer morbidity and mortality with an increasing incidence in young adults4. The gut microbial dysbiosis, along with enrichment of CRC-promoting species like Fusobacterium nucleatum, polyketide synthase-positive Escherichia coli, and enterotoxigenic Bacteroides fragilis, plays critical roles in CRC onset and development5. A recent study highlighted the stage-specific alterations in gut microbial and metabolic composition during CRC progression6. Elevated fecal GUS activity has been observed in CRC patients compared to the healthy cohort7, suggesting a potential gmGUS dysregulation in CRC. GUS enzymes are widely distributed in gut microbiota8. Notably, CRC-enriched species like B. fragilis, Enterococcus faecalis and E. coli are also GUS-harboring species1,9. These evidences, albeit limited, indicate gmGUS-mediated disturbance of glucuronidation homeostasis in CRC, which may also show stage-specific changes and contribute to CRC initiation and/or progression.
Despite these insights, the mechanistic link between gmGUSs and CRC development remains to be explored. On one hand, deconjugation by gmGUSs allows the release of CRC-associated toxic aglycones to undergo enterohepatic recirculation, increasing their intestinal and systemic exposure and the following harmful effects. Typical examples include dietary carcinogens like 2-amino-3-methylimidazo[4,5-f]quinoline10 and 2-amino-1-methyl-6-phenylimidazo[4,5-b]pyridine (both formed in well-cooked meat and are DNA-adducts)11. A number of endogenous compounds like neurotransmitters (serotonin and norepinephrine), estrogens (17β-estradiol) and BAs1 also undergo enterohepatic circulation and were reported to be related to CRC. Serotonin is linked to more severe colitis symptoms and promotes CRC cells toward expansion12. Norepinephrine enhances colon cancer progression via the norepinephrine-CREB1-miR-373 axis, as observed in vitro on HCT116 and RKO cells and in vivo using nude mice and clinical specimens13. Evidence suggests both anticancer and colorectal carcinogenesis activity of 17β-estradiol, depending on the signaling pathways involved14. An elevation of primary and secondary BAs in CRC was observed in many studies15. Secondary BAs promote CRC development by influencing T cell differentiation and activating the Wnt/β-catenin and NF-κB signaling pathways, as well as intrinsic apoptotic pathways and BA receptors FXR and TGR515. On the other hand, mucins, glycosaminoglycans like chondroitin, heparan and hyaluronan, and plant polysaccharides like xylan, cellulose and pectin, all contain glucuronic acid (GlcA)8, making them susceptible to modulation by gmGUSs. Mucin degradation disrupts mucosal structure and epithelial integrity, facilitating pathogen adhesion and invasion16. Breakdown of glycosaminoglycans like chondroitin and heparan significantly impacts host health by impacting gut microbiota colonization and proliferation17. GlcA-containing plant polysaccharides serve as vital dietary fibers, influencing both the composition and functionality of gut microbiota16. Additionally, the common product GlcA released from de-glucuronidation processes is a weak acid. It can not only serve as a carbon source for certain microbial populations1 but make the gut lumen a more acidic environment, reshaping the microbial structure. A higher GUS activity7 and enrichment of some GUS-harboring species in CRC patients1 imply more active de-glucuronidation events and a greater CRC risk. Taken together, the change of the microbe-gmGUS-metabolite (MGM) axis should be among the early events of CRC. Unraveling the gmGUS profile as well as the connections between microbes, gmGUSs and metabolites may not only offer significant mechanistic insights into CRC etiology, but also provide potential diagnostic biomarkers and therapeutic targets for CRC.
Recently, a gmGUS atlas in the healthy cohort from the Human Microbiome Project was identified, adopting a traditional workflow for gmGUS identification, including the alignment of the gene sets to four reference GUSs followed by screening the aligned hits for the presence of seven GUS conserved residues essential for hydrolyzing glucuronide-containing substrates. These conserved residues include the NxKG motif, catalytic E residues and N and Y motifs, which distinguish the carboxylic acid moiety unique to glucuronic acid relative to the primary alcohol in the nearly isostructural galactose8. However, this method is likely to produce false positive GUS sequences through pairwise alignment, and given the high structural diversity of gmGUSs, the four reference GUSs used for identifying putative gmGUSs are under-representative. Currently, the continuously accumulating GUS sequences in public databases allow the expansion of reference proteins for increasing the coverage. Moreover, the inclusion of Glyco_hydro_2_N (PF02837), Glyco_hydro_2 (PF00703), and Glyco_hydro_2_C (PF02836), three architectural domains of GUS enzymes18, can remove false positive hits obtained from pairwise alignment to enhance accuracy.
In this work, a comprehensive approach for gmGUS identification is proposed, which employs a combination of 114 manually checked public gmGUSs as references, refined with three architectural domains of GUS enzymes and seven conserved residues of bacterial GUSs. The approach is then applied to identify gmGUSs from a public CRC cohort containing 571 individuals6. Subsequently, taxonomic annotation is conducted to build the atlas of the identified gmGUSs. Variations in GUS loop categories, GUS-harboring species and gmGUSs along CRC stages are recognized, followed by discovering discriminative gmGUSs for CRC early diagnosis and the building of the GUSscore model to predict clinical outcome. Associations between differential microbes, gmGUSs and metabolites are characterized to construct the MGM axis, aiming to shed light on the role of the MGM axis in CRC development. Finally, in vitro enzyme assays are used for functional validation, and cell experiments followed by RNA-seq are implemented to provide preliminary insights into the potential role of gmGUSs in CRC progression.
Results
Identification of gmGUSs and construction of an atlas of human gmGUSs
To discover gmGUSs, a gene catalog containing 10,947,761 Unigenes was constructed firstly using a public metagenomic dataset6 consisting of healthy controls (normal and a few polyps, n = 247), MP (multiple polypoid adenomas with low-grade dysplasia, n = 66), S0 (polypoid adenomas with high-grade dysplasia, n = 73), SI/II (stages I and II CRC, n = 111) and SIII/IV (stages III and IV CRC, n = 74) groups.
The schematic diagram of identification and annotation of gmGUSs from Unigenes was presented in Fig. 1a. First, 114 GUS references were obtained from the National Center for Biotechnology Information database after manual checks (Supplementary Fig. 1 and Supplementary Data 1). The amino acid sequences of Unigenes were then aligned to the references, resulting in 69,503 putative gmGUSs. Subsequent removal of false positives from pairwise alignment with three architectural domains and seven conserved residues resulted in 820 putative gmGUSs. Finally, 550 gmGUSs were annotated to species-level (Supplementary Data 2). The rarefaction curves reached plateaus at around 200 samples, suggesting that the number of samplings is sufficient, with very few new gmGUSs undetected (Fig. 1b).
a Flow chart of the pipeline used for identification of gmGUSs from Unigenes (n = 10,947,761), which employed a combination of 114 reference gmGUSs, three architectural domains of GUS enzymes (PF02836, PF02837 and PF00703), and seven GUS conserved residues. b Rarefaction curves showing the number of gmGUSs in healthy controls (n = 247), CRC patients (S0, SI/II and SIII/IV groups, n = 258), and total samples (n = 571) after 100 random samplings. c Taxonomic cladogram of 550 gmGUSs. The inner to outer circles represent the hierarchical taxonomic levels of kingdom, phylum, class, order, family, genus, and species, respectively. Different colors represent different phyla, and the bar chart indicates the number of annotated gmGUSs for respective species. The asterisks represent the top two orders with the highest gmGUS number. Source data are provided as a Source Data file.
A taxonomic cladogram of the 550 gmGUSs was shown in Fig. 1c. The majority of gmGUSs originated from the phyla Bacteroidota (60%) and Bacillota (29.6%). Two orders, namely Bacteroidales and Eubacteriales, were the main contributors, accounting for 57.1 and 26.4% of the gmGUS pool, respectively. The 550 gmGUSs were assigned to 173 species, with the top three species (Bacteroides ovatus, Bacteroides cellulosilyticus, and Bacteroides faecium) contributing to 12% (Supplementary Fig. 2).
Distribution of loop structures
Two active site-adjacent loop regions, which potentially influence substrate preference, were categorized into six loop types, namely Loop 1 (L1), mini-Loop 1 (mL1), Loop 2 (L2), mini-Loop 2 (mL2), mini-Loop 1,2 (mL1,2), and No Loop (NL) according to the lengths of loops8. The gmGUSs span over a wide range of length (320-2429 AA, median 800.5 AA) with the majority (99.5%) classified into six loop categories. NL type was the richest (45.1%), followed by mL1 (25.1%), L1 (13.5%), L2 (8.6%), mL2 (4.6%), and mL1,2 (2.7%) (Fig. 2a and Supplementary Fig. 3). The top five genera of each loop category were shown in Fig. 2b. Bacteroides was the most represented in all loop categories, while its contribution varied widely, ranging from 24.3% (L1) to over 60% of the two minor groups (mL1,2 and mL2). The less abundant four genera were largely different across loop categories. For instance, Faecalibacterium, Parabacteroides, Clostridium, Alistipes, Akkermansia, and Phocaeicola were secondary to Bacteroides for NL, mL1, L1, L2, mL2, and mL1,2, respectively. Interestingly, mL2 and mL1,2 GUSs only came from five and six genera, respectively, while the top five genera account for 56, 63, 47.3, and 78.7% of NL, mL1, L1, and L2 types, respectively, indicating vast taxonomic sources of these GUS categories.
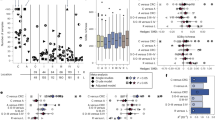
a Distribution of gmGUS length colored by loop category. No coverage means predicted proteins lack sequencing information for one or both loop regions. b Genus composition with percentages of gmGUSs per genus indicated. c Violin plot showing the abundance distribution of mini-Loop 2. d PCoA based on the gmGUS profile. No significant differences across groups are established by PERMANOVA. e Violin plot showing the total abundance (left) and number (right) of gmGUSs across groups. f Left heatmap representing mean cumulative GUS abundance (CGA) (transformed into Z scores) of nine species with significant differences in CGA between healthy controls and one of the CRC stages; middle heatmap displaying log2 fold change (log2FC) of comparisons; dot plot illustrating feature trends, with increases in red triangles and decreases in blue inverse triangles. g Violin plot illustrating differences in cumulative GUS number for three species among the nine depicted in Fig. 2f, exhibiting significant variations (P < 0.1). h Left heatmap representing the mean relative abundance (transformed into Z scores) of six species with significant differences among the nine species shown in Fig. 2f, between different groups; middle heatmap displaying log2FC; dot plot illustrating species trends using the SILVA LTP method, with decreases in blue inverse triangles and increases in red triangles. Species trends validated with MetaPhlAn2 and mOTUs4 are represented by rectangles with black borders and gray fills, respectively. Healthy (n = 247); MP (n = 66); S0 (n = 73); SI/II (n = 111); SIII/IV (n = 74). For boxplots, the middle line represents the median; the box bounds indicate the first and third quartiles (Q1 and Q3), which form the interquartile range (IQR); the upper whisker extends to the maximum value within Q3 + 1.5×IQR; the lower whisker extends to the minimum value within Q1 − 1.5×IQR. P values in Fig. c, e, g are from a two-sided Wilcoxon rank-sum test without multiple testing correction. Source data are provided as a Source Data file.
The abundance of mL2 was significantly enriched in SIII/IV compared with the healthy counterparts (P < 0.001; fold change = 1.52) (Fig. 2c). Furthermore, the GUS number of mL2 also increased in SIII/IV with a fold change of 1.20 (P = 0.005) (Supplementary Fig. 3).
Species-level disturbance of gmGUSs in CRC
The overall GUS community structure (β-diversity), as well as the total abundance and total number of gmGUSs displayed no differences between controls and different CRC stages (Fig. 2d, e and Supplementary Data 3, 4).
The cumulative GUS abundance (CGA) in nine species was significantly different between controls and different CRC stages (Fig. 2f and Supplementary Data 5). Specifically, CGA was enriched in CRC stages for six species: B. faecium in MP, Bacteroides helcogenes in S0, B. cellulosilyticus, Dorea longicatena, and Akkermansia muciniphila in SIII/IV, Bacteroides nordii in both SI/II and III/IV. Conversely, CGA decreased in SI/II for Subdoligranulum variabile and Bifidobacterium bifidum, and in SIII/IV for Lachnospira eligens. To assess the impact of GUS copy number variation (CNV) on GUS abundance of the nine species, MIDAS219 was employed. The results revealed that six species with sufficient coverage for analysis generally showed no statistical differences between groups, except for a significant decrease in GUS CNV of A. muciniphila in SIII/IV compared to healthy individuals (Supplementary Fig. 4). The changes of cumulative GUS number of three species, including B. faecium, B. cellulosilyticus, and B. nordii correspond well to their CGA changes (Fig. 2g). Additionally, the changes of bacterial relative abundance of six species showed consistency with the CGA findings (Fig. 2h).
gmGUS signatures of different CRC stages
Totally, there are 38 differential gmGUSs at different CRC stages compared to controls (Supplementary Fig. 5 and Supplementary Data 6). Apparent gmGUS shifts already occurred in early stages of CRC, and the number of differential gmGUSs generally increased with CRC progression (10, 13, 12, and 19 differential gmGUSs in MP, S0, SI/II, and SIII/IV, respectively). Interestingly, all 13 downregulated gmGUSs were stage-specific (Fig. 3a). In contrast, among the 25 upregulated gmGUSs, 56% were stage-specific, indicating large overlay in gmGUSs upregulation among different CRC stages.
a Hierarchical taxonomic tree of the 38 differential gmGUSs between different CRC stages and healthy controls. Loop categories are shown on the right side of the tree: Loop 1 (L1), mini-Loop 1 (mL1), Loop 2 (L2), mini-Loop 2 (mL2), mini-Loop 1,2 (mL1,2), No Loop (NL). On the right of the hierarchical taxonomic tree, colored heatmap showing the mean abundances (transformed into Z scores) of corresponding gmGUS per group, dot plot showing the decreased (inverse triangle colored by blue) or increased (triangle colored by red) tendency of 38 gmGUSs among different comparisons, and colored dot-line chart indicating the prevalence of 38 gmGUSs among healthy controls and different stages of CRC. b ROC curves of the two random forest models classifying CRC patients (n = 258) and adenoma patients (n = 66) from healthy controls (n = 247) based on the test set. The AUCs are displayed in the lower right-hand corner. c Mean decrease accuracy (MDA) of the top 20 selected features from the CRC classifier. d Forest plot displaying eight gmGUSs associated with progression-free survival based on 46 CRC samples. The hazard ratio (HR) and P values are obtained through two-sided univariate Cox regression analysis without multiple testing correction. e Risk factor diagram of the GUSscore model, illustrating the distribution of GUSscore, survival time, and abundance of model gmGUSs across the 37 training samples (reordered by GUSscore). The cutpoint is determined with the R package survminer. f Progression-free survival curve showing outcomes between high- (n = 26) and low-GUSscore (n = 11) groups in the training set. The P value is obtained through the log-rank test. g Six-year receiver operating characteristic curves evaluating the performance of the GUSscore model in predicting survival outcomes in the training (n = 37) and test (n = 9) sets. Source data are provided as a Source Data file.
Differences of the 38 gmGUSs within confounding factors including age, gender, BMI, Brinkman index, and alcohol consumption were evaluated (Supplementary Data 7). Par_chongii.GUS1 and Lac_eligens.GUS2 exhibited gender differences. Lac_eligens.GUS2, Ech_strongylocentroti.GUS, and Stu_stutzeri.GUS significantly correlated with age, Brinkman index, and alcohol consumption, respectively. After adjusting for associated confounders, only the difference for Lac_eligens.GUS2 between SIII/IV and healthy controls remained significant.
A hierarchical tree was constructed to clarify taxonomic, loop classification and stage patterns of the 38 significant gmGUSs (Fig. 3a). These gmGUSs span over four phyla, 13 genera, and 19 species. The contributors mainly belong to phyla Bacteroidota and Bacillota, containing 60.5 and 28.9% of the 38 gmGUSs, respectively. Among the 13 genera these gmGUSs distributed, Bacteroides, Parabacteroides, and Faecalibacterium accounted for 44.7, 13.2, and 7.9%, respectively. It was interesting to note four species-level patterns of the contributors to these significant gmGUSs. The first was the elevation in S0 and SIII/IV, with B. cellulosilyticus as the main contributor. The second pattern was characterized by gmGUSs from B. nordii, which displayed particularly high abundance in SIII/IV. Third, gmGUSs that were specifically elevated in SIII/IV were mainly attributed to species such as Ruminococcus torques and D. longicatena. Lastly, the decreases in specific CRC stages were mainly characterized by Faecalibacterium prausnitzii, B. bifidum and Oscillospira bacterium. Notably, the second and fourth patterns displayed lower prevalence compared to the other two. In terms of loop classification, these gmGUSs covered all six loop categories, with NL (36.8%) and mL1 (34.2%) accounting for the majority. Interestingly, NL and mL1 constituted MP-specific changes, while the differential gmGUSs of S0 to SIII/IV CRC originated from diverse loop types, indicating that NL and mL1 GUSs are the main players throughout CRC development and advanced CRC stage involves more complex metabolic disturbance.
gmGUS biomarkers for early detection of CRC
To evaluate the potential of gmGUSs as early diagnostic markers for CRC, two random forest classifiers were utilized to distinguish CRC and adenoma patients from controls, achieving moderate AUCs of 0.827 and 0.815 for the test set, respectively (Fig. 3b). The selected gmGUS biomarkers for the CRC classifier were primarily distributed in B. nordii, B. cellulosilyticus, and Bacteroides caccae (Fig. 3c), while those for the adenoma classifier were mainly in B. caccae and B. nordii (Supplementary Fig. 6). Subsequently, the performance of CRC classifier was evaluated on datasets from three countries, yielding suboptimal AUCs of 0.727, 0.677, and 0.707 for Australia, Germany, and France, respectively (Supplementary Fig. 7a). These results indicate limited generalizability of the current model across cohorts and suggest cautious clinical utility. Furthermore, differential analysis validated only 12, 10, and 4 out of the 38 significant gmGUSs in Australia, Germany, and France, respectively (Supplementary Fig. 7b and Supplementary Data 8), highlighting potential cross-cohort heterogeneity of GUS composition.
Additionally, the multi-class model (including healthy individuals, adenoma patients, and CRC patients) achieved an AUC of ~0.75 (Supplementary Fig. 8a, b). Furthermore, by integrating 13 gmGUSs (primarily from B. nordii and B. cellulosilyticus) with 32 bacterial biomarkers, the discriminating power of the CRC classifier was enhanced with an AUC of 0.875 (Supplementary Fig. 8c, d). The top species, weighted by MeanDecreaseAccuracy, included Fusobacterium animalis, Peptostreptococcus stomatis, Gemella morbillorum, Parvimonas micra, and Bulleidia moorei, which are critical biomarkers reported previously20.
Construction and validation of the GUSscore model
The 38 differential gmGUSs were further analyzed with the survival status of the 46 patients in a follow-up study lately published21. Eight of the differential gmGUSs were significantly associated with progression-free survival, with five from B. cellulosilyticus (Fig. 3d and Supplementary Data 9). The optimal model was established using five gmGUSs selected from 550 gmGUSs (Supplementary Fig. 9a, b). Following the coefficients, GUSscore was calculated using the formula: GUSscore = 0.2513090 × Bac_cellulosilyticus.GUS18 + 0.3141506 × Bac_cellulosilyticus.GUS5 + 0.1534125 × Osc_bacterium.GUS15 + 1.0682435 × Osc_bacterium.GUS7 + 0.2044853 × Osc_bacterium.GUS8. With a cutpoint of 0.05 determined by the R package survminer (Supplementary Fig. 9c), training samples were divided into two groups: high-GUSscore (n = 26) and low-GUSscore (n = 11). The model gmGUSs were all increased in the high-GUSscore group (Fig. 3e and Supplementary Fig. 10a). Survival analysis revealed that the high-GUSscore group exhibited significantly shorter survival times than the low-GUSscore (Fig. 3f and Supplementary Fig. 10b). Subsequently, time-dependent ROC curves demonstrated the prognostic accuracy of GUSscore for 6-year survival, with AUCs of 0.934 and 0.875 for the training and test sets, respectively (Fig. 3g and Supplementary Fig. 10c).
gmGUS-associated ecological network across CRC stages
To gain insight into the potential interplays between gmGUSs and microbiota, stage-specific co-abundance networks were constructed using differential species and gmGUSs of respective stages. Totally, 179 GUS-correlated and CRC-associated species were found (Fig. 4a and Supplementary Data 10). The number of GUS-associated species, GlcA utilizers, GUS-harboring species, and associations increased steadily with CRC progression (Fig. 4b, c and Supplementary Fig. 11).
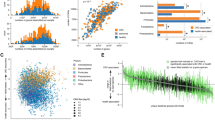
a Co-abundance network displaying correlations using differential species and gmGUSs from comparisons between different CRC stages and healthy controls. Mean correlations were recalculated between species and gmGUSs annotated to the same species. Node color represents functional group, while border color represents abundance trend, with red for increases and blue for decreases in CRC stages. Species from the same genus have a yellow-green elliptical background; those from the same functional groups are marked with corresponding background colors. Other functioners include mucin degrader, flavonoid degrader, equol producer, primary bile acid converter, and putrescine fermenter, marked by a purple elliptical background. Node shape represents feature type: yellow rectangles for gmGUSs and circles for species. Line color represents negative (blue) or positive (red) correlations, with thickness indicating correlation strength. Glucuronic acid (GlcA) utilizers and GUS-harboring species are marked with red and blue stars, respectively. b Barplot showing percentage of GUS-associated species relative to the number of differential species across different stages, with blue indicating species enriched in healthy controls and red indicating species enriched in CRC stages. c Barplot displaying the number of GlcA utilizers and GUS-harboring species across different CRC stages, with blue indicating species enriched in healthy controls and red indicating species enriched in CRC stages. d Dot plot showing the enrichment of functional groups at different CRC stages. The node size represents the species ratio, and the node color represents log2(k), where k represents the number of GUS-correlated species with a specific function in a specific stage. *P < 0.05. e Dot plot showing the enriched genera. The node size represents the species ratio. The node color represents the P values, with gray indicating P > 0.05. The genera color indicates the tendency of the corresponding GUS-associated species, with blue indicating control-enriched, red indicating stage-enriched, and black indicating genera with both control-enriched and stage-enriched species. P values in Fig. d, e are from one-sided Fisher’s exact test without multiple testing correction. Source data are provided as a Source Data file.
Next, the 179 species were manually categorized based on their functional roles to gain a deeper understanding of their potential contribution to CRC pathogenesis (Fig. 4a, d and Supplementary Data 11). Significant decreases were observed in taxa responsible for degrading plant glycans (primary degraders) like cellulose, xylan, and pectin in MP and SI/II. Pathogens, defined as species isolated from clinical specimens, specifically Schaalia odontolytica, P. micra, and Clostridium symbiosum, were statistically enriched in SIII/IV, suggesting a heightened infectious burden in advanced CRC. Noteworthily, SIII/IV showed stage-specific GUS-associated diverse functional groups, including sulfate reducers (SRBs), mucin degraders, flavonoid degraders, equol producers, primary BA converters and putrescine fermenters, which were generally increased.
Genus-level enrichment analysis of GUS-associated species revealed notable stage-specific shifts (Fig. 4e). Lachnospira spp., which are primary degraders and butyrate producers6, decreased in MP, whereas Prevotella and Parabacteroides, which have been reported as CRC-enriched flora6,22, increased in S0. In SI/II, Bifidobacterium and Ruminiclostridium were reduced, while an increase of Alistipes was observed. Bifidobacterium spp. are gut commensals, while Alistipes has been reported as potential pathogens in CRC23. Fusobacterium spp., which are recognized to play a role in colorectal carcinogenesis5, were specifically enriched in SIII/IV.
Stage-specific biological link maps of gmGUSs in CRC
Fecal metabolites and microbial KEGG orthology (KO) were analyzed to reveal the potential mechanistic links between gmGUSs, metabolites, and host processes during CRC progression. Changes of GUS-correlated metabolites were only found in SIII/IV, indicating the GUS-associated global metabolic reprogramming lay behind gut microbial disturbance. As shown in Fig. 5a and Supplementary Data 12, 13, totally, 44 GUS-correlated fecal metabolites were detected, with the majority (63.6%) involved in amino acid (AA) metabolism. The 11 enriched metabolites belong to phenylalanine metabolism (3), branched-chain amino acids (BCAA) metabolism (2), and lysine degradation (2), most (6/11) being negatively associated with GUSs of L. eligens. Among the 33 metabolites downregulated, 17 were involved in polyamine metabolism (5), methionine metabolism (5), histamine metabolism (5) and lysine degradation (2). GUSs of B. cellulosilyticus, Cellulosimicrobium cellulans, Parabacteroides goldsteinii, and R. torques were all upregulated and negatively correlated with downregulated metabolites, indicating that the four species are the main players of the downregulation. Functional pathway analysis revealed significant enrichment of beta-alanine metabolism, phenylalanine, tyrosine and tryptophan biosynthesis, and lysine degradation (Fig. 5b). Disease analysis identified enrichment of these GUS-associated metabolites in patients with gastrointestinal diseases and inflammatory arthritis, suggesting a potential role of these metabolites in the pathogenesis of these conditions (Fig. 5b).
a Co-abundance network displaying correlations using differential metabolites and gmGUSs from comparisons between SIII/IV patients and healthy controls. Mean correlations were recalculated between metabolites and gmGUSs annotated to the same species. Node color represents the functional category. The elliptical background color of nodes indicates the specific amino acid metabolism pathways. Node border color represents feature trends, with red for increases in the SIII/IV group and blue for decreases. Node shape represents feature type, with a rectangle representing gmGUSs and a circle representing metabolites. Line color represents negative (blue) or positive (red) correlations, with thickness reflecting correlation strength. b Enriched KEGG pathways (left) and human diseases (right) based on GUS-correlated metabolites. The node size represents the ratio of metabolites in each category, and node color represents the adjusted P value (FDR), with gray indicating FDR >0.05. P values are obtained using MetaboAnalyst with FDR correction. c Venn diagram showing the overlap of GUS-associated KOs in the comparative analysis between healthy controls and CRC patients at different stages. d Enriched KEGG pathways based on 684 GUS-correlated KOs. The node size represents the number of KOs in each KEGG pathway, and node color represents the FDR. Only KEGG pathways with FDR < 0.05 are displayed. P values are obtained using ClusterProfiler with FDR correction. e Dot plot displaying the annotated GUS-correlated and CRC-associated KOs at four CRC stages for each KEGG pathway. A total of 318 KOs were annotated to 56 KEGG pathways that can be categorized into ten functional categories. The corresponding number of metabolites from Fig. 5a is also displayed. The node size represents the number of features. The node color represents the tendency of features, with red indicating an increase in CRC stages, blue indicating a decrease in CRC stages, and yellow indicating the presence of both control-enriched and stage-enriched features. ABC ABC transporters, PTS phosphotransferase system. Source data are provided as a Source Data file.
The GUS-correlated KOs and their enriched functions were also analyzed. Totally, 684 GUS-correlated and CRC-associated KOs were identified, with 95, 129, 205, and 391 KOs in the MP, S0, SI/II, and SIII/IV, respectively (Fig. 5c). Out of the 684 KOs, 58.2% (n = 398) were annotated to 86 KEGG pathways (Supplementary Data 14), with seven showing statistical enrichment (Fig. 5d). The core functional pathways of the 398 KOs were manually checked to eliminate redundancy. It was found that 79.9% of them (n = 318) were categorized into 56 KEGG pathways across ten functional categories (Fig. 5e and Supplementary Data 15).
AA metabolism was the main GUS-correlated category, including 14 pathways. KOs for biosynthesis of aromatic AAs (histidine, phenylalanine, and tryptophan) were decreased, while those involved in their degradation were increased in CRC. This was alongside decreased metabolites of histidine metabolism and increased metabolites linked to phenylalanine metabolism in SIII/IV (Fig. 5a). Furthermore, KOs for polyamine metabolism were increased, accompanied by decreases of five GUS-correlated polyamine derivatives in SIII/IV. Two GUS-associated BCAAs, leucine and valine, along with lysine (all of which are essential AAs), were increased in SIII/IV, coupled with decreased biosynthesis and increased degradation in CRC. Moreover, KOs for proline and glycine metabolism were increased in CRC alongside decreased hydroxyproline in SIII/IV.
Dysregulated carbohydrate metabolism was evidenced by simultaneous increases and decreases in most related KOs. However, the metabolism of ascorbate and aldarate, butanoate, and propanoate were consistently increased in several CRC stages. Another important category, vitamin biosynthesis, showed distinct patterns with increases in vitamin K (VK) and folate, and decreases in cobalamin (VB12) in CRC. Furthermore, biosynthesis of bacterial cell envelope components, including lipopolysaccharide (LPS) and O-antigen nucleotide sugar, as well as metabolism of glycerolipids and glycerophospholipids, were increased, along with increases of ABC transporters for phospholipids, LPS, and lipoproteins, suggesting potential proliferation of harmful bacteria in CRC. Additionally, the phosphotransferase system, particularly for N-acetylgalactosamine and fructoselysine/glucoselysine, was upregulated. Moreover, energy metabolism also underwent significant changes, including increases in oxidative phosphorylation, methanogenesis, and nitrogen metabolism, along with a decrease in photosynthesis. Notably, in sulfur metabolism, aprFamily and cysFamily KOs corresponding to dissimilatory and assimilatory pathways, respectively24, were decreased, while asrFamily KOs, which use sulfite as a substrate to produce hydrogen sulfide (H2S), were increased. In the genetic information processing category, KOs responsible for DNA replication and base excision repair were increased in SIII/IV, suggesting potential alterations in genomic stability, while KOs involved in RNA degradation and aminoacyl-tRNA biosynthesis were decreased, indicating possible changes in protein synthesis in CRC.
Differential glucuronide processing in vitro
To functionally validate the gmGUSs identified, 17 gmGUSs were selected from 38 differential gmGUSs for in vitro enzyme assays (Supplementary Data 16), including two novel gmGUSs without online BLAST hits (CC.G, SS.G), four with low BLAST identity (<90%) (BH.G2, ES.G, FP.G3, OB.G7, Supplementary Data 17), four B. cellulosilyticus gmGUSs and three B. nordii gmGUSs across different loop types, and four core gmGUSs (DL.G2, FP.G7, PG.G2, and RT.G) in the MGM axis. Except for SS.G and BN.G5, most (15/17) purified gmGUSs exhibited moderate to strong deconjugation activity against the probe substrates p-nitrophenyl-β-D-glucuronide (PNPG) and/or 4-methylumbelliferyl-β-D-glucuronide (4MUG) (Fig. 6a, Supplementary Fig. 12, and Supplementary Data 18). Notably, the GUS proteins also exhibited a distinct preference towards the two small molecular substrates. For instance, ES.G, RT.G, BN.G4, and BC.G18 preferred processing PNPG over 4MUG, while FP.G3 processed 4MUG effectively but not PNPG. SS.G and BN.G5 only showed weak activity against the hydrolysis of 4MUG and PNPG. Given the structural diversity of glucuronides2 and the observed substrate preferences of gmGUSs, they may catalyze the deconjugation of other glucuronide substrates more efficiently.
a Barplot displaying the GUS activity of 17 purified candidate gmGUSs using the standard substrates 4-methylumbelliferyl-β-D-glucuronide (4MUG) and p-nitrophenyl-β-D-glucuronide (PNPG). GUS activity is calculated as nmol of PNP or 4MU liberated/mg protein/min. Each bar represents three biological replicates. All data were presented as means with standard errors. The 17 gmGUSs were divided into three groups based on tertiles (33.3% and 66.7% percentiles) of the mean values of GUS activity: low (≤23.7; labeled in blue), moderate (23.7-127.5; labeled in orange), and high (>127.5; labeled in red). b Cell viability measured at OD450. All data are presented as means with standard deviations. P value between BC.G3 (Bac_cellulosilyticus.GUS3, 10 ng/mL) cocultured HCT116 cells (n = 3) and controls (incubated with stocking buffer, n = 3) is obtained using the ANOVA test without multiple testing correction. c Volcano plot illustrating the differentially expressed genes (DEGs) between the BC.G3 cocultured group (n = 3) and controls (n = 3). P values are obtained using DESeq2 with FDR correction. d Barplot showing enriched pathways based on 384 upregulated DEGs (Up) and 158 downregulated DEGs (Down) as determined by Metascape analysis with Benjamini-Hochberg correction. Only pathways with q values <0.05 are displayed, with the count of DEGs for each enriched pathway listed to the right of the bars. e The log2FoldChange barplot (Up) is accompanied by a heatmap displaying the expression (transformed into Z scores) of the top 40 DEGs across groups. Following the heatmap, a dot plot illustrates the mean abundances (transformed into Z scores) and the percentage of expressing epithelial cells between colorectal tumors and adjacent normal tissues from a public human CRC atlas (c295) in the Single Cell Portal (https://singlecell.broadinstitute.org/single_cell/study/SCP1162/human-colon-cancer-atlas-c295). Below the dot plot, the log2FoldChange of the respective genes compares colorectal tumors to adjacent normal tissues. Among the 40 genes, only 28 genes present in the single-cell atlas are displayed. Source data are provided as a Source Data file.
Role of gmGUS on human CRC cells
Considering that NL GUSs and B. cellulosilyticus GUSs made the most significant contribution to the CRC-enriched gmGUSs and B. cellulosilyticus GUSs are also key gmGUSs in the MGM axis, Bac_cellulosilyticus.GUS3 (BC.G3), an NL GUS from B. cellulosilyticus, was chosen for co-culture with CRC cells (HCT116 and HT29). BC.G3 induced a modest increase in the viability of HCT116 cells (anova P = 0.042; fold change = 1.115) (Fig. 6b and Supplementary Data 19) but showed no effect on HT29 cells (Supplementary Fig. 13). RNA-seq analysis revealed 542 differentially expressed genes (DEGs) (384 upregulated and 158 downregulated) between HCT116 cells cocultured with BC.G3 and control HCT116 cells (Fig. 6c and Supplementary Data 20). Functional enrichment analyses indicated that the upregulated DEGs were involved in RNA metabolism, splicing and transcription, DNA metabolism and replication, cell cycle, transcriptional regulation by TP53, nucleotide excision repair, protein folding, mitochondrial-associated pathways, and regulation of apoptosis (Fig. 6d and Supplementary Data 21). Gene Set Enrichment Analysis revealed the upregulation of pathways involving cell differentiation, ATP synthesis, and rRNA processing (Supplementary Fig. 14). Interestingly, 25 out of the top 28 upregulated DEGs were also elevated in the epithelial cells of colorectal tumors compared to adjacent normal tissues, according to a public human CRC atlas (c295) in the Single Cell Portal (Fig. 6e). These findings suggest these genes may be relevant to CRC pathogenesis, although their role in gmGUS-mediated effects remains preliminary and requires further functional validation.
Discussion
This study provides a global overview for understanding the role of gmGUSs in CRC pathogenesis. By integrating multi-omics profiles, an MGM axis was constructed to link the dysregulation of key gmGUSs to the functionality of the gut microbiota and metabolic disturbances. Additionally, this study underscores potential diagnostic biomarkers and therapeutic targets for CRC, and provides preliminary insights into the roles of gmGUSs in CRC development, suggesting that gmGUS variations are potentially both indicative of and contribute to CRC progression.
Compared to the previous GUS identification method8, which relied on four references for alignment and seven conserved residues for filtration, resulting in 279 gmGUSs from 139 samples (HMGC279), the current approach includes an expanded reference database to enhance sensitivity and coverage for potential GUS identification, along with domain-based structural verification to reduce false positives. Ultimately, this study nearly doubled the number of identified gmGUSs (543 vs. 279) and assigned species (168 vs. 87) from 247 controls, suggesting that gmGUSs are more widespread and diverse than previously recognized. Despite these enhancements, this approach may not cover non-GH2 GUSs, as a minority of gmGUSs belong to GH30 and GH793, while the domains used in this study are derived from GH2 analysis18. Considering that CAZy classification for the HMGC279 with confirmed lengths reveals that all enzymes have conserved GH2 domains8, the identification may miss non-GH2 GUS hits, but should be very few. Additionally, like most alignment-based systems, this method remains limited in discovering novel gmGUSs that deviate from the references, potentially leading to false negatives in the gene catalog. Furthermore, it is constrained in differentiating structural homologs (e.g., β-galactosidases) from GUSs. With the development of this area and the accumulation of data, including experimentally validated true positive and true negative GUS sequences, a robust and effective learning-based model for GUS identification will be developed to address these limitations25. Moreover, future integration of structural alignments like AlphaFold26, will facilitate the development of a more comprehensive gmGUS atlas.
In this study, NL GUSs, though underrepresented in references (13.2%), were predominant and accounted for the majority of the differential gmGUSs, suggesting their overlooked significance and largely unknown functions. Categories L1, mL1, and L2 GUSs appear to be more efficient in de-glucuronidating small substrates than mL2, mL1,2, and NL GUSs27. mL2 GUSs, which favor large substrates, exhibited a significant increase in SIII/IV. These shifts of GUS pattern may echo the shift of substrate spectrum throughout carcinogenesis and the disruption of glucuronidation homeostasis, which potentially contribute to CRC development. Notably, the alignment-based loop classification may introduce biases caused by loop edge ambiguity28. Given the structural diversity of urinary glucuronides2 and functional varieties even within gmGUSs of the same loop type29, future functional validation is essential to clarify the roles of these categories in CRC and to promote the establishment of a meticulous system that offers higher biological relevance.
Herein, controversial trends in gmGUSs within species were noticed. For example, among the 38 significant gmGUSs, one GUS from P. goldsteinii decreased in S0, while two from the same species increased in late-stage CRC. Additionally, elevated fecal GUS activity has been observed previously in CRC patients compared to healthy cohorts7. However, not all gmGUSs were elevated in CRC in the current research, with some gmGUSs and GUS-harboring species were significantly decreased, suggesting a complex imbalance between the upregulation and downregulation of gmGUSs in CRC. Given the structural variability of glucuronides2 and the observed substrate preferences of gmGUSs, the significance of the complex regulation of gmGUSs in CRC needs to be explored with physiologically relevant substrates.
Interestingly, B. cellulosilyticus made the biggest contribution to the CRC-enriched gmGUSs. Moreover, preliminary RNA-seq data from co-culture experiments suggest that BC.G3 may elicit transcriptional responses in CRC cells that are associated with proliferation-related pathways. B. cellulosilyticus is a common species inhabiting the human gut. It ferments versatile cellulose to produce short-chain fatty acids (SCFAs)30. Administration of B. cellulosilyticus attenuated murine colitis in vivo31. Thus, this species has been widely accepted as a beneficial species. However, the elevation of B. cellulosilyticus in fecal microbiota has been reported in several diseases, including Parkinson’s disease32 and type 1 diabetes33, as well as in dextran sulfate sodium-induced colitis mice34. Besides, a previous study has shown that the growth of B. cellulosilyticus is significantly favored at pH >735. A protein-rich diet, which is believed to be a risk factor of CRC, can increase the colonic pH as a result of ammonia release from protein fermentation36. The increase of B. cellulosilyticus in disease status may lead to the degradation of mucin, which could be attributed to the presence of mucin-degrading GHs in the genomes of B. cellulosilyticus37. These findings indicate that B. cellulosilyticus may outcompete other mucin-degrading bacteria when complex carbohydrates are inadequate or absent in the host’s diet38. As a consequence, exacerbated mucin utilization by gut microbiota enriched with B. cellulosilyticus plus insufficient mucin production due to butyrate decrease39 weakens gut barrier function and disrupts immune homeostasis, potentially contributing to CRC development38.
Additionally, gmGUSs display their potential as early diagnostic biomarkers and therapeutic targets for CRC. While the CRC classifier showed modest performance (AUC = 0.827) within the study cohort, its variable and lower performance in external independent cohorts indicates that substantial generalizability issues remain, consistent with the broader challenge of translating microbial biomarkers across studies20,40. Notably, the enhanced AUC (0.875) from integrating gmGUSs with bacterial biomarkers is promising but exploratory. Caution is warranted regarding clinical translation. Additionally, the observed heterogeneity of GUS composition across cohorts, along with previous meta-studies20,40, which suggest poor transportability of microbial biomarkers from one dataset to other and employing large, diverse training sets facilitates the development of more accurate diagnostic models, highlight the necessity of using multi-cohorts to globally evaluate the potential of gmGUSs as biomarkers for CRC in the future. Recent studies indicate that gmGUSs-mediated de-glucuronidation accounts for severe side effects of CRC chemotherapy (such as the chemotherapy drugs irinotecan and regorafenib)29, reactivation of dietary carcinogens10,11, and the circulating levels of various endogenous compounds linked to colorectal carcinogenesis1. The established stage-specific changes of gmGUSs and the MGM axis in this study, support further discovery and applications of gmGUS inhibitors in CRC management and medication.
Noteworthily, enrichment of several functional species was specifically identified in late-stage CRC. Mucin degraders can damage the protective mucus layer, enhancing disease susceptibility16. Moreover, the observed elevated transport and phosphorylation of N-acetylgalactosamine, a component of mucin16, also suggest enhanced mucin degradation in CRC patients. SRBs produce H2S, which inhibits butyrate oxidation, thereby compromising the integrity of the colonocyte barrier24. Equol producers metabolize phytoestrogens daidzein and genistein to equol, a metabolite with high affinity to estrogen receptors, thus may contribute to CRC progression through estrogen homeostasis41. Flavonoid degradation reduces bioavailability and anticarcinogenic effects of beneficial flavonoids in CRC42. Furthermore, natural flavonoids can moderately to strongly inhibit E. coli GUS43, and GUSs from the other two species also exhibit varied inhibition propensities with dietary flavones29, suggesting that flavonoid degradation may suppress negative feedback on GUS activity. Collectively, these bacterial alterations are likely a response or contributor to the changing colonic microenvironment during CRC progression.
The study highlights the role of gmGUS in regulating AA homeostasis. The observed degradation and decreased biosynthesis of essential AAs, coupled with the decrease of primary degraders, suggest potentially elevated dietary protein and insufficient dietary fiber intake in CRC patients. Supporting this, a high-protein and low-carbohydrate diet, commonly associated with CRC44, has been shown to cause excessive AAs in feces compared to maintenance diets45. Additionally, elevated fecal GUS activity in individuals consuming a high meat diet compared to a nonmeat regimen46 further underscores the association between gmGUSs and AA changes on CRC progression. The decreased histamine, a histidine derivative with antitumor effects47, may lead to increased gmGUS activity by relieving GUS release48, potentially promoting CRC progression. Conversely, aromatic AAs were increased in SIII/IV. Intestinal metabolism of these AAs can yield suspected co-carcinogens phenylpropionate45, which is also GUS-correlated and SIII/IV-enriched, and gut barrier-disrupting metabolites phenol and p-cresol49. Notably, p-cresol undergoes extensive first-pass intestinal and hepatic metabolism, producing p-cresol sulfate and p-cresol glucuronide, which are thought to primarily mediate its toxic effects50, suggesting that gmGUSs may promote CRC by modulating exposure of the toxic tyrosine and phenylalanine derivatives. Moreover, excessive AA metabolism can produce harmful intestinal compounds, including ammonia, phenolic compounds, and polyamines, which can disrupt host homeostasis and alter diversity, abundance and metabolism of the gut microbiota51, potentially contributing to CRC development. Disturbance of polyamine homeostasis, marked by elevated levels of putrescine fermenter, Anaerovorax odorimutans, and polyamine biosynthesis precursors, arginine and lysine, as well as depleted acetyl-polyamines, was observed in this study. Furthermore, gases generated through AA metabolism can be utilized by hydrogenotrophic microorganisms (primarily methanogenic archaea, acetogenic bacteria and SRBs)45, exacerbating the microbial dysbiosis in CRC patients and facilitating CRC development.
Beyond AA metabolism, vitamin synthesis emerges as another critical category in GUS-mediated colon carcinogenesis. VK can promote CRC progression by activating pregnane X receptor, which stimulates the expression of NPC1L1 and CD36, both implicated in CRC tumorigenesis52,53,54. Lipophilic VK undergoes enterohepatic circulation, typically conjugated with GlcA to enhance solubility for excretion55. Thus, elevated VK synthesis, alongside increased gmGUS level to facilitate VK recirculation, may enhance VK exposure, contributing to CRC development. Folate is involved in one-carbon metabolism, and imbalances in this pathway may disrupt regulatory mechanisms in purine and pyrimidine biosynthesis, which are vital for DNA and RNA synthesis, and methylation reactions, ultimately impacting CRC development56. Excessive folate intake has been associated with elevated CRC risk in individuals with precursor lesions based on high-dose folate studies56. Moreover, folate can inhibit the glucuronidation activity of UGTs in vitro57, suggesting that it may potentially affect the UGT-gmGUS axis and disrupt glucuronidation homeostasis in CRC.
Intriguingly, associations between gmGUSs and bacterial motility were advised in this study. Bacterial motility, as indicated by GUS-correlated KOs for flagellar assembly and chemotaxis, was reduced in CRC. The downregulation of csrA, a post-transcriptional regulator that can promote motility58, supports the observed decrease of motility. Besides, this study observed increased bacterial communication, with rbsD and lsrF, crucial genes involved in processing quorum-sensing signal autoinducer-259, being GUS-correlated and CRC-enriched. Collectively, these findings underscore a complex interplay between gmGUS and bacterial behavior, warranting further investigation into the underlying mechanisms. Furthermore, LPS biosynthesis was elevated, corresponding with gmGUS changes. LPS is a large glycolipid composed of three structural domains: lipid A, the core oligosaccharide, and the O antigen60. GlcA can be added to the third heptose of the inner core oligosaccharide of LPS61. The variety of O-antigens, which function as receptors for bacteriophages and significantly influence host immune responses62, is also noteworthy. Indeed, there are 186 documented O-antigens within the E. coli typing schemes, with four reported to contain GlcA62. Hence, GlcA released through de-glucuronidation by gmGUSs may participate in LPS biosynthesis in addition to being used as an energy source, enriching harmful Gram-negative bacteria such as Alistipes, Bacteroides, Fusobacterium, and Prevotella, enhancing their resistance to antimicrobial compounds and modulating host immune responses in CRC patients60.
The current study had the following limitations: (1) As individual metadata, such as dietary factors are associated with fecal GUS activity46 and CRC development44, lack of these information in the study cohort limit the interpretation of results; (2) The abundance calculation may be biased by gene duplications, although GUS copy number variation analysis was conducted to discuss the biases, further analyses at genomic-level are needed to refine the results; (3) The CRC classifier displays limited generalizability across cohorts, suggesting cautious clinical utility and the need for further multi-cohort studies; (4) The GUSsore model was constructed based on limited sample size, leading to potential overfitting effects. Future collection of additional CRC samples with survival status is needed to enhance the robustness of the model; (5) This study concentrated on GUS-associated perturbations of fecal metabolites in CRC, potentially missing certain glucuronides due to the limited detection sensitivity of non-targeted metabolome; (6) The in vitro and in vivo effects of different gmGUSs on various substrates were not assessed in this study; (7) This study is descriptive and does not establish direct biological causality. Despite these limitations, the results of this study are still valid for three reasons. Firstly, the relationship between gmGUSs and CRC is assessed by a study cohort with big population size and stage information. Secondly, since glucuronidation is a common metabolic pathway of numerous endogenous and exogenous molecules and the MGM axis in CRC patients is largely unknown, investigation of the gmGUS signatures and the MGM axis provides valuable insights into the GUS-associated roles in CRC pathogenesis, potentially enabling improved treatment strategies and diagnostic tools. Thirdly, GUS activity was validated using in vitro enzyme assays with two standard substrates, and subsequent cell experiments, followed by RNA-seq, provided preliminary insights into the role of gmGUSs in CRC progression. Further studies incorporating multi-cohorts, targeted glucuronide profiling2, functional validation (e.g., knockout studies, pathway inhibition), and wet-lab experiments like CRC organoids and in vivo models63, should be conducted to address the limitations of this work, and to offer a more accurate portrait of the roles of gmGUSs and the microbe-gmGUS-glucuronide axis in colorectal carcinogenesis.
Methods
All research protocols and procedures in this study were approved by the Sub Panel on Biomedical Science & Engineering Research Ethics (Human Participants) of the University of Macau (Approved No. BSERE18-APPO24-ICMS).
The study cohort
Publicly available fecal metagenomic sequencing data for 576 subjects from Japan in the healthy controls (normal and a few polyps, n = 251), MP (multiple polypoid adenomas with low-grade dysplasia, n = 67), S0 (polypoid adenomas with high-grade dysplasia, n = 73), SI/II (stages I and II CRC, n = 111), and SIII/IV (stages III and IV CRC, n = 74) groups were obtained from DDBJ Sequence Read Archive (DRA) as DRA006684 and DRA0081566. Among the 576 samples, five were excluded from analysis due to sequencing data yielding less than 1 G.
Gene catalog construction and determining the abundance of unigenes
Firstly, a gene catalog was constructed based on the downloaded metagenomic sequencing data. Specifically, the assembly of clean reads was executed using SOAPdenovo (Version 2.04, parameters: -d 1 -M 3 -u -F -V 1 -K 57). Then, clean reads were mapped against the assembled contigs using SOAP2 (Version 2.21, parameters: -r 2 -m 200 -x 400). Unmapped reads were co-assembled on all samples using MEGAHIT64 (Version 1.2.9, parameters: --k-list 57 --min-contig-len 500) to discover low-abundance genes that could not be assembled from individual samples. Genes were predicted on contigs using MetaGeneMark65 (prokaryotic GeneMark.hmm Version 3.38, parameters: -a -d -f G) and a non-redundant gene catalog was constructed with CD-HIT66 (Version 4.8.1, parameters: -c 0.95 -G 0 -aS 0.9 -g 1 -n 5 -d 0). To determine gene abundance, reads were realigned to the gene catalog with BWA (Version 0.7.17-r1188, default parameters). The abundance of genes was calculated by counting the number of mapped reads per million and normalizing by gene length.
Identification of gut microbial GUSs
Reference GUS proteins were retrieved and downloaded from the NCBI database using the search keyword “beta glucuronidase” in 25/09/2023, limiting the taxonomic information to bacteria. Sequences marked by the word “candidate” or “putative”, and sequences originating from other enzymes and sequences with unknown functions were manually excluded from the searching result. Finally, 114 GUS protein sequences were maintained and used as references for the following gmGUS identification. gmGUSs were retrieved from the gene catalog using the following criteria: (i) aligning to 114 GUS proteins with >25% identify and E values below 0.05 by blastp (Version 2.12.0 + , default parameters); (ii) containing all three architectural domains of GUS enzymes (PF02836, PF02837 and PF00703)18 by hmmsearch (Version 3.3.2, default parameters) with E values below 0.05; and (iii) preserving the essential and specific active-site residues for bacterial GUSs, including the N and Y motif, NxKG motif and catalytic E residues8.
Loop classification and taxonomic annotation of gut microbial GUSs
Loop classification of gmGUSs were determined as previously described8. Specifically, by using ClustalO (Version 1.2.3, default parameters), multiple sequence alignment was conducted based on the identified gmGUS proteins and eight references with NCBI accession numbers WP_005931592.1, NP_416134.1, WP_012740861.1, WP_005639106.1, WP_004298526.1, pdb|3CMG | A, WP_035447612.1, and WP_007841259.1. Then, the loop category for each gmGUS was determined by the length of the Loop 1 region and the Loop 2 region. The abundance of loop categories was calculated by summing the abundance of gmGUSs belonging to the same loop category. Taxonomic information of gmGUSs were annotated using Kraken267 (Version 2.0.7-beta, default parameters) with an advanced database created by extracting sequences of bacteria, archaea, fungi, and virus from NCBI genebank database (accession date: 17/10/2023). The cumulative GUS abundance of bacteria were calculated by summing the abundance of gmGUSs annotated to the same bacteria.
The validation cohort
Three independent CRC datasets were included, comprising a total of 332 samples from France (FRA), Australia (AUS), and Germany (GER). These datasets were selected based on the availability of CRC stage samples and healthy controls, and their availability of raw metagenomic sequencing data. Specifically, the FRA dataset68 consisted of 53 CRC patients and 61 matched controls. The AUS dataset69 included 63 controls and 38 CRC patients. Lastly, the GER dataset20 contained 60 control samples with 57 CRC patients.
Construction of a random forest classifier
Using the profile of gmGUSs, the samples were randomly divided into a training set and a test set using the R package caret (Version 7.0-1). Random forest classifiers were trained on 80% of the data and tested on the remaining 20% of the data using the R package randomForest (Version 4.7-1.1). Specifically, features were selected from the 550 identified gmGUSs based on the training set using the Boruta algorithm (Version 8.0.0) with 500 seeds. gmGUSs present in ≥10% of the seeds were ultimately selected. Hyperparameter tuning (ntree: 500,600,700,800,900,1000; mtry:1 to the number of selected features) was implemented via nested cross-validation, followed by Random Forest model construction based on the training set. Variable importance was calculated for the random forest models. The performance of the models was measured as a receiver operating characteristic curve and AUC analysis when applied to the test set and three independent CRC datasets using the R package pROC (Version 1.16.2).
Development and construction of the GUSscore model
In the study cohort, progression-free survival information of 46 CRC patients was obtained from a preprint by Yachida’s group21. The 28 differential gmGUSs from comparisons of SI/II vs. healthy and SIII/IV vs. healthy were included for univariate Cox regression analysis based on the 46 samples using the R package survival (Version 3.7-0). For GUSscore model construction, these patients were randomly divided into a training set (80%, n = 37) and a test set (20%, n = 9) using the R package caret (Version 7.0-1). Then, gmGUSs present in ≥10% of the Boruta feature selection using 500 seeds were selected for LASSO model evaluation based on the training set using the R package glmnet (Version 4.1-8). The optimal LASSO model was established using the min lambda, and the GUSscore was calculated using the following formula:
The coefficient (\(i\)) represented the coefficient obtained from LASSO analysis, and the abundance (\(i\)) denoted the abundance of gmGUS (\(i\)). Using the cutpoint determined with the R package survminer (Version 0.4.9), the training samples were then divided into two subgroups: the high-GUSscore group (GUSscore >= cutpoint) and the low-GUSscore group (GUSscore <cutpoint). Survival probabilities were calculated using the R package survival, and the prediction efficiency was assessed with the R package timeROC (Version 0.4). Later, the test set was also split into two subgroups based on the established cutpoint, followed by survival and ROC analysis to evaluate the performance of the model.
Expression and purification of β-glucuronidases
The GUS DNA was cloned into the pET28a expression plasmid with an N-terminal 6x-Histidine tag. The resulting vector (pET28a-gmGUS) was sequenced on both strands using an ABI 3730 sequencer (Perkin-Elmer, CA, USA). Then, the recombinant plasmid was transformed into Escherichia coli BL21-DE3 competent cells. The cells were grown at 37 °C until an OD600 of 0.6 was reached, induced with isopropyl-1-thio-D-galactopyranoside (IPTG), and then harvested by centrifugation. After cell resuspension, the cells were disrupted by sonication and then centrifuged to obtain the soluble proteins from the supernatant. The cell-free extracts were applied to a Ni-NTA column to collect fractions. Fractions showing appreciable p-nitrophenyl-β-D-glucuronide (PNPG) or 4-methylumbelliferyl-β-D-glucuronide (4MUG) activity were combined and concentrated by ultrafiltration (Amicon Ultra 30 kDa, Merck Millipore, Darmstadt, Germany). Protein concentrations were determined using a Nanodrop Spectrophotometer, and purity was determined by SDS-PAGE (8% gels).
Enzymatic assays for GUS activity
PNPG and 4MUG are commercially available (purity > 98%). PNPG was dissolved directly in phosphate-buffered saline (PBS) to 3 mM. For 4MUG, a 60 mM stock solution was prepared in dimethyl sulfoxide (DMSO) and subsequently diluted with PBS to 3 mM. In vitro assays were conducted at 100 μL total volume in 96-well, clear-bottom assay plates (Costar) at 37 °C. The reactions consisted of 80 μL PBS, 10 μL enzyme, and 10 μL substrate (at a final concentration of 300 μM). PNP formation was measured via absorbance at 405 nm using a SpectraMax M5 Microplate Reader (Molecular Devices, Silicon Valley, CA, USA). The formation of 4MU was measured with fluorescence detection using excitation and emission wavelengths of 350 and 450 nm, respectively. Each GUS was tested with three biological replicates.
Cell culture
Human CRC cell lines, HCT116 (catalog no. SNL-077) and HT29 (catalog no. SNL-069), were purchased from SUNNCELL Biotechnology Co., Ltd. (Wuhan, China). All cell lines were authenticated by the supplier using STR profiling prior to delivery. Cells were cultured in McCoy’s 5 A (modified) Medium (Gibco, Grand Island, NY) supplemented with 10% fetal bovine serum and 1% penicillin/streptomycin (Life Technologies, Shanghai, China), and antibiotics (100 μg/mL streptomycin and 100 U/mL penicillin) in a 5% CO2 incubator at 37 °C.
Cell viability assay
A Cell Counting Kit-8 (CCK-8) assay (MCE, Shanghai, China) was performed to test the effect of Bac_cellulosilyticus.GUS3 on cell viability. Cells were seeded in 96-well plates (3000 cells per well) for 24 h and incubated with GUS enzyme (10 ng/69.92 kDa/mL) for 0, 24, 48, 72 h. After incubation, 10 μL CCK-8 solution was added to each well and incubated for another 2 h. The absorbance density was measured on a 96-well plate reader at a wavelength of 450 nm. Each reaction was set up in technical triplicate, and the average (absorbance) value was used for analysis. Three biological replicates (samples prepared and seeded on different days) were performed for each condition.
RNA-seq analysis
HCT116 cells were seeded in 6-well plates (30 W cells per well) for 24 h and incubated with Bac_cellulosilyticus.GUS3 (10 ng/69.92 kDa/mL) for 48 h. The controls were incubated with the stock buffer for 48 h. Three biological replicates were used for RNA-seq experiments. After incubation, the cells were harvested and suspended with RNA-later and stored at -80 °C until sequencing. RNA-seq samples were prepared from three biological replicates per experimental condition. Total RNA was isolated using the Trizol kit (Ambion/Invitrogen, USA), and RNA quality and integrity were confirmed via Agilent Bioanalyzer 2100. Next, mRNA libraries were generated and sequenced using Illumina NovaSeq X Plus PE150 at Novogene (Tianjin, China). After quality control, clean reads were mapped to the hg38 genome using HISAT2 (Version 2.2.1) with default parameters. Then, featureCounts (Version 2.0.6) was used to count the reads mapped to each gene and generate the FPKM profile. Differentially expressed genes (DEGs), which were defined as |log2(fold change)| >1.2 and adjusted P < 0.05, were calculated from DESeq2 (Version 1.42.0). Functional enrichment of DEGs was analyzed through Metascape (https://metascape.org/). Gene Set Enrichment Analysis was conducted using R package clusterProfiler (Version 4.10.1).
Statistical analysis
Rarefaction analysis was implemented by randomly sampling with replacement 100 times and estimating the total number of gmGUSs that could be identified from these samples. PCoA and PERMANOVA were performed by the ape package and vegan package in R software (Version 4.3.2) based on the profile of gmGUSs, respectively.
The Wilcoxon rank-sum test and FDR correction was used to identify statistically different gmGUSs and GUS-harboring species. Only features that existed in at least 10% samples were considered in the analyses. Features with an adjusted P value <0.05 and an absolute log2 fold change >1 were considered statistically different. The differences in the abundance of species (based on the SILVA LTP method), bacterial KOs and metabolites between healthy controls and CRC patients at different stages, as well as the species profile based on MetaPhlAn2 (Version 2.6.0), were respectively extracted from the Supplementary Tables S15-1, S15-5, S15-6, and S9 of the study cohort mentioned in the Methods section6. Species profile from mOTUs4 was generated using high-quality reads with default parameters.
MIDAS219 was employed to analyze GUS copy number variation (CNV) of species based on their publicly available genomes from the GTDB. GUSs were identified from the genes of the collected genomes based on the established procedure, and the GUS CNV for each species was calculated using MIDAS2 with default parameters. Due to the limited sample size for Bacteroides nordii with default parameters, parameters recommended by MIDAS2 for genotyping low-abundance species were applied for the analysis of B. nordii (--select_by median_marker_coverage,unique_fraction_covered --select_threshold=0,0.5). Then, the Wilcoxon rank-sum test was used to evaluate the differences in species GUS CNV between groups.
Differences of statistically significant gmGUSs within confounding factors were assessed using the Wilcoxon rank-sum test (For categorical variable, gender) and Spearman’s correlation analysis (for continuous variable, including age, BMI, Brinkman Index, and alcohol consumption). An analysis of Variance (ANOVA) model (using the aov function in the R package stats) was utilized to evaluate differences in gmGUSs between CRC stages and healthy individuals, adjusting for associated confounding factors.
Based on the supplementary tables of the study cohort, which include the species, metabolic, and KO profile, differential gmGUSs were correlated with the differential species, metabolites and KOs at each of the four CRC stages vs. healthy controls, respectively, by using Spearman’s correlation analysis with the R package psych (Version 2.3.3). Only correlations with FDR <0.05 and an absolute correlation coefficient (r) >0.2 were considered as statistically correlated. The networks were visualized by Cytoscape (Version 3.10.1). To simplify the network, the correlations between species/metabolites and gmGUSs annotated to the same species were recalculated by taking their mean correlations.
The enrichment of functional groups and genera from GUS-associated species at the four CRC stages was tested using the Fisher’s exact test, with the following formula:
Where N represents the total number of GUS-correlated species, M represents the number of GUS-correlated species with a specific function or belonging to a specific genus, n represents the total number of GUS-correlated species in a specific stage, and m represents the number of GUS-correlated species with a specific function or belonging to a specific genus in a specific stage.
The species ratio of Fisher’s exact test was calculated with the following formula:
KEGG pathways of GUS-correlated KOs were identified by searching the KEGG database. Functional enrichment of GUS-correlated metabolites and GUS-correlated KOs were performed with the R package MetaboAnalyst (Version 3.3.0) and ClusterProfiler (Version 4.6.2), respectively. Disease enrichment of GUS-correlated metabolites was performed with the R package MetaboAnalyst.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The raw RNA sequencing data generated in this study have been deposited in the Genome Sequence Archive (GSA) under accession code HRA011183. The processed gene catalog and GUS sequence data are available at Figshare [https://doi.org/10.6084/m9.figshare.29037680]. Source data are provided with this paper.
Code availability
All codes have been deposited at GitHub [https://github.com/yr2008-UM/CRC_GUS] and have been assigned the [https://doi.org/10.5281/zenodo.17082894]70.
References
Pellock, S. J. & Redinbo, M. R. Glucuronides in the gut: Sugar-driven symbioses between microbe and host. J. Biol. Chem. 292, 8569–8576 (2017).
Chen, Z. Q. et al. Chemical isotope labeling and dual-filtering strategy for comprehensive profiling of urinary glucuronide conjugates. Anal. Chem. 96, 13576–13587 (2024).
Wang, P. P., Jia, Y. F., Wu, R. R., Chen, Z. Q. & Yan, R. Human gut bacterial beta-glucuronidase inhibition: an emerging approach to manage medication therapy. Biochem. Pharmacol. 190, 114566 (2021).
Vuik, F. E. et al. Increasing incidence of colorectal cancer in young adults in Europe over the last 25 years. Gut 68, 1820–1826 (2019).
Qu, R. et al. Role of the gut microbiota and its metabolites in tumorigenesis or development of colorectal cancer. Adv. Sci. 10, e2205563 (2023).
Yachida, S. et al. Metagenomic and metabolomic analyses reveal distinct stage-specific phenotypes of the gut microbiota in colorectal cancer. Nat. Med. 25, 968–976 (2019).
Kim, D. H. & Jin, Y. H. Intestinal bacterial beta-glucuronidase activity of patients with colon cancer. Arch. Pharm. Res. 24, 564–567 (2001).
Pollet, R. M. et al. An atlas of beta-glucuronidases in the human intestinal microbiome. Structure 25, 967–96 (2017).
Cheng, Y. W., Ling, Z. X., Li, L. J. The intestinal microbiota and colorectal cancer. Front. Immunol. 11, 615056 (2020).
Humblot, C. et al. beta-Glucuronidase in human intestinal microbiota is necessary for the colonic genotoxicity of the food-borne carcinogen 2-amino-3-methylimidazo[4,5-f]quinoline in rats. Carcinogenesis 28, 2419–2425 (2007).
Zhang, J. et al. Gut microbial beta-glucuronidase and glycerol/diol dehydratase activity contribute to dietary heterocyclic amine biotransformation. BMC Microbiol. 19, 99 (2019).
Battaglin, F. et al. Neurotransmitter signaling: a new frontier in colorectal cancer biology and treatment. Oncogene 41, 4769–4778 (2022).
Han, J. et al. Norepinephrine-CREB1-miR-373 axis promotes progression of colon cancer. Mol. Oncol. 14, 1059–1073 (2020).
Wu, Z. et al. 17beta-estradiol in colorectal cancer: friend or foe?. Cell Commun. Signal 22, 367 (2024).
Cai, J., Sun, L. & Gonzalez, F. J. Gut microbiota-derived bile acids in intestinal immunity, inflammation, and tumorigenesis. Cell Host Microbe 30, 289–300 (2022).
Koropatkin, N. M., Cameron, E. A. & Martens, E. C. How glycan metabolism shapes the human gut microbiota. Nat. Rev. Microbiol. 10, 323–335 (2012).
Dong, J., Cui, Y. & Qu, X. Metabolism mechanism of glycosaminoglycans by the gut microbiota: bacteroides and lactic acid bacteria: a review. Carbohydr. Polym. 332, 121905 (2024).
Talens-Perales, D., Gorska, A., Huson, D. H., Polaina, J. & Marin-Navarro, J. Analysis of domain architecture and phylogenetics of family 2 glycoside hydrolases (GH2). PLoS ONE 11, e0168035 (2016).
Zhao, C., Dimitrov, B., Goldman, M., Nayfach, S., Pollard, K. S. MIDAS2: metagenomic intra-species diversity analysis system. Bioinformatics 39, btac713 (2023).
Wirbel, J. et al. Meta-analysis of fecal metagenomes reveals global microbial signatures that are specific for colorectal cancer. Nat. Med. 25, 679–67 (2019).
Shibata, T. et al. Microbiome spectra and prevalent colibactin-associated mutational process in Japanese colorectal cancer. Preprint at Research Square https://doi.org/10.21203/rs.3.rs-4967540/v1 (2024).
Gagnière, J. et al. Gut microbiota imbalance and colorectal cancer. World J. Gastroenterol. 22, 501–518 (2016).
Parker, B. J., Wearsch, P. A., Veloo, A. C. M. & Rodriguez-Palacios, A. The genus Alistipes: gut bacteria with emerging implications to inflammation, cancer, and mental health. Front. Immunol. 11, 906 (2020).
Kushkevych, I. et al. Recent advances in metabolic pathways of sulfate reduction in intestinal bacteria. Cells 9, 698 (2020).
Li, Y. et al. HMD-ARG: hierarchical multi-task deep learning for annotating antibiotic resistance genes. Microbiome 9, 40 (2021).
Abramson, J. et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 630, 493–500 (2024).
Candeliere, F. et al. beta-Glucuronidase pattern predicted from gut metagenomes indicates potentially diversified pharmacomicrobiomics. Front. Microbiol. 13, 826994 (2022).
Creekmore, B. C. et al. Mouse gut microbiome-encoded beta-glucuronidases identified using metagenome analysis guided by protein structure. Msystems 4, e00452-19 (2019).
Wang, P. et al. Inhibition and structure-activity relationship of dietary flavones against three Loop 1-type human gut microbial beta-glucuronidases. Int. J. Biol. Macromol. 220, 1532–1544 (2022).
He, X. et al. In vitro colonic fermentation profiles and microbial responses of cellulose derivatives with different colloidal states. J. Agric. Food Chem. 70, 9509–9519 (2022).
Neff, C. P. et al. Diverse intestinal bacteria contain putative Zwitterionic capsular polysaccharides with anti-inflammatory properties. Cell Host Microbe 20, 535–547 (2016).
Jo, S. et al. Oral and gut dysbiosis leads to functional alterations in Parkinson’s disease. Npj Parkinsons Dis 8, 87 (2022).
Zhang, L. et al. Islet autoantibody seroconversion in type-1 diabetes is associated with metagenome-assembled genomes in infant gut microbiomes. Nat. Commun. 13, 3551 (2022).
Guo, M. R. et al. Sucralose enhances the susceptibility to dextran sulfate sodium (DSS) induced colitis in mice with changes in gut microbiota. Food Funct. 12, 9380–9390 (2021).
Raba, G., Adamberg, S. & Adamberg, K. Acidic pH enhances butyrate production from pectin by faecal microbiota. FEMS Microbiol. Lett. 368, fnab042 (2021).
Russell, W. R. et al. High-protein, reduced-carbohydrate weight-loss diets promote metabolite profiles likely to be detrimental to colonic health. Am. J. Clin. Nutr. 93, 1062–1072 (2011).
Glover, J. S., Ticer, T. D. & Engevik, M. A. Characterizing the mucin-degrading capacity of the human gut microbiota. Sci. Rep. 12, 8456 (2022).
Christensen, L. et al. Microbial enterotypes beyond genus level: Bacteroides species as a predictive biomarker for weight change upon controlled intervention with arabinoxylan oligosaccharides in overweight subjects. Gut Microbes 12, 1847627 (2020).
Burger-van Paassen, N. et al. The regulation of intestinal mucin MUC2 expression by short-chain fatty acids: implications for epithelial protection. Biochem. J. 420, 211–219 (2009).
Thomas, A. M. et al. Metagenomic analysis of colorectal cancer datasets identifies cross-cohort microbial diagnostic signatures and a link with choline degradation. Nat. Med. 25, 667–678 (2019).
Mayo, B., Vázquez, L. & Flórez, A. B. Equol: a bacterial metabolite from the daidzein isoflavone and its presumed beneficial health effects. Nutrients 11, 2231 (2019).
Gupta, A. et al. Association of Flavonifractor plautii, a flavonoid-degrading bacterium, with the gut microbiome of colorectal cancer patients in India. Msystems 4, e00438–19 (2019).
Sun, C. P. et al. The study of inhibitory effect of natural flavonoids toward beta-glucuronidase and interaction of flavonoids with beta-glucuronidase. Int. J. Biol. Macromol. 143, 349–358 (2020).
Hughes, R., Magee, E. A. & Bingham, S. Protein degradation in the large intestine: relevance to colorectal cancer. Curr. Issues Intest. Microbiol. 1, 51–58 (2000).
Davila, A. M. et al. Intestinal luminal nitrogen metabolism: role of the gut microbiota and consequences for the host. Pharm. Res. 68, 95–107 (2013).
Reddy, B. S., Weisburger, J. H. & Wynder, E. L. Fecal bacterial beta-glucuronidase: control by diet. Science 183, 416–417 (1974).
Krell, T. et al. Histamine: a bacterial signal molecule. Int. J. Mol. Sci. 22, 6312 (2021).
Busse, W. W. & Sosman, J. Histamine inhibition of neutrophil lysosomal enzyme release: an H2 histamine receptor response. Science 194, 737–738 (1976).
Andriamihaja, M. et al. The deleterious metabolic and genotoxic effects of the bacterial metabolite p-cresol on colonic epithelial cells. Free Radic. Biol. Med. 85, 219–227 (2015).
Rong, Y. & Kiang, T. K. L. Characterizations of human UDP-glucuronosyltransferase enzymes in the conjugation of p-Cresol. Toxicol. Sci. 176, 285–296 (2020).
Dai, Z., Wu, Z., Hang, S., Zhu, W. & Wu, G. Amino acid metabolism in intestinal bacteria and its potential implications for mammalian reproduction. Mol. Hum. Reprod. 21, 389–409 (2015).
Drury, J. et al. Inhibition of fatty acid synthase upregulates expression of CD36 to sustain proliferation of colorectal cancer cells. Front. Oncol. 10, 1185 (2020).
He, J. et al. NPC1L1 knockout protects against colitis-associated tumorigenesis in mice. BMC Cancer 15, 189 (2015).
Yan, H. et al. The relationship among intestinal bacteria, vitamin K and response of vitamin K antagonist: a review of evidence and potential mechanism. Front. Med. 9, 829304 (2022).
Barkhan, P. & Shearer, M. J. Metabolism of vitamin K1 (phylloquinone) in man. Proc. R. Soc. Med. 70, 93–96 (1977).
Kok, D. E. et al. Bacterial folate biosynthesis and colorectal cancer risk: more than just a gut feeling. Crit. Rev. Food Sci. Nutr. 60, 244–256 (2020).
Pande, S. et al. Inhibition of uridine 5’-diphospho-glucuronosyltransferases A10 and B7 by vitamins: insights from in silico and in vitro studies. Silico Pharm. 12, 8 (2024).
Timmermans, J. & Van Melderen, L. Post-transcriptional global regulation by CsrA in bacteria. Cell Mol. Life Sci. 67, 2897–2908 (2010).
Marques, J. C. et al. LsrF, a coenzyme A-dependent thiolase, catalyzes the terminal step in processing the quorum sensing signal autoinducer-2. Proc. Natl Acad. Sci. USA 111, 14235–14240 (2014).
Bertani, B. & Ruiz, N. Function and biogenesis of lipopolysaccharides. EcoSal Plus 8, https://doi.org/10.1128/ecosalplus.ESP-0001-2018 (2018).
Klein, G. et al. Molecular and structural basis of inner core lipopolysaccharide alterations in Escherichia coli: incorporation of glucuronic acid and phosphoethanolamine in the heptose region. J. Biol. Chem. 288, 8111–8127 (2013).
Samuel, G. & Reeves, P. Biosynthesis of O-antigens: genes and pathways involved in nucleotide sugar precursor synthesis and O-antigen assembly. Carbohydr. Res. 338, 2503–2519 (2003).
Su, A. C. Y. et al. Lactococcus lactis HkyuLL 10 suppresses colorectal tumourigenesis and restores gut microbiota through its generated alpha-mannosidase. Gut 73, 1478–1488 (2024).
Li, D., Liu, C. M., Luo, R., Sadakane, K. & Lam, T. W. MEGAHIT: an ultra-fast single-node solution for large and complex metagenomics assembly via succinct de Bruijn graph. Bioinformatics 31, 1674–1676 (2015).
Zhu, W., Lomsadze, A. & Borodovsky, M. Ab initio gene identification in metagenomic sequences. Nucleic Acids Res. 38, e132 (2010).
Fu, L., Niu, B., Zhu, Z., Wu, S. & Li, W. CD-HIT: accelerated for clustering the next-generation sequencing data. Bioinformatics 28, 3150–3152 (2012).
Wood, D. E., Lu, J. & Langmead, B. Improved metagenomic analysis with Kraken 2. Genome Biol. 20, 257 (2019).
Zeller, G. et al. Potential of fecal microbiota for early-stage detection of colorectal cancer. Mol. Syst. Biol. 10, 766 (2014).
Feng, Q. et al. Gut microbiome development along the colorectal adenoma-carcinoma sequence. Nat. Commun. 6, 6528 (2015).
Chen, J. R., Li, Y., Tang, S., Jin, W. Y. & Yan, R. Gut microbial β-glucuronidases and their role in the microbiome-metabolite axis in colorectal cancer. Zenodo https://doi.org/10.5281/zenodo.17082894 (2025).
Acknowledgements
The high-performance computing resources and services used in this work were supported by the High-Performance Computing Center of Information and Communication Technology Office, University of Macau. This work was supported by the Science and Technology Development Fund of Macao SAR (0091/2021/A2 to R.Y., 0098/2019/A2 to R.Y., SKL-QRCM(UM)−2023-2025 to R.Y.), Shenzhen-Hong Kong-Macau Science and Technology Program Category C (SGDX20210823103805038 to R.Y.), and University of Macau (MYRG-GRG2023-00241-ICMS-UMDF to R.Y.).
Author information
Authors and Affiliations
Contributions
J.C. analyzed data and wrote the manuscript. R.Y. revised the manuscript and supervised this project. Y.L. and W.J. performed the expression and purification of β-glucuronidases, as well as the in vitro enzyme assays. S.T. provided the GlcA-utilizing species list and prepared samples for RNA sequencing. S.T. and Y.L. conducted the in vitro cell experiments.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Samuel Miravet-Verde, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Chen, J., Li, Y., Tang, S. et al. Gut microbial β-glucuronidases and their role in the microbiome-metabolite axis in colorectal cancer. Nat Commun 16, 10660 (2025). https://doi.org/10.1038/s41467-025-65679-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-65679-y