Abstract
The CCKBR agonists induce neocortical long-term potentiation of excitatory synaptic transmission and enhance memory formation, while its antagonists weaken the potentiation in the amygdala and alleviate depression-like behaviors. However, the mechanism that drives CCKBR dependent long-term potentiation remains elusive. There is also no signaling pathway-biased CCKBR agonist to modulate the potentiation. Here, we discover a β-arrestin biased CCKBR agonist MF-8 with IC50 = 0.9 nM. The activation of CCKBR with MF-8 fails to induce the potentiation but efficiently induces CCKBR endocytosis. Multi-Electrode Array results demonstrate that the potentiation is dependent on Gαq/11-Ca2+ and Gαs-cAMP signaling pathways. The potentiation is entirely blocked by MF-8 through β-arrestin signaling. Furthermore, MF-8 effectively inhibits the formation of cue-to-cue associative fear memory. These results reveal the signal pathway preference of the CCKBR long-term potentiation and identify a blocker of the potentiation, which provides us with broader insights into developing drugs targeting CCKBR.
Similar content being viewed by others
Introduction
G protein-coupled receptors (GPCRs) are the most prominent family of membrane proteins in eukaryotes, with more than 800 members in this family1. These proteins facilitate the transmission of diverse signals across the cell membrane, including gas molecules, polypeptides, photons, and small molecules2. Given their critical roles in mediating intracellular and extracellular signaling, these membrane proteins represent optimal targets for drug development. Current statistics indicate that GPCR-targeted drugs account for nearly 34% of all marketed drugs worldwide1,2.
Signal transduction upon GPCR activation was initially thought to be only G protein-dependent until arrestins were also found to play a significant role in GPCR signaling3. Besides, each GPCR was initially supposed to activate downstream signaling pathways through one unique G protein4. However, recent findings have shown that the same GPCR can activate different G proteins5,6, and arrestins can function either dependently or independently of G proteins7,8,9. The arrestin family contains four members, including arrestin1, arrestin2 (β-arrestin1), arrestin3 (β-arrestin2), and arrestin4. The human G protein family includes 16 Gα, 5 Gβ, and 12 Gγ4,10. There are indications that GPCR activation is biased towards different subtypes of G proteins and arrestins4,10. Therefore, even the same GPCR produces different biological effects due to the different types of G proteins and arrestins it activates. Based on this theory, drugs with high selectivity for receptor types or bias for signaling pathways may be specifically applied in treating specific diseases while avoiding numerous potential side effects during disease treatments. Hence, several signaling pathway-biased agonists and antagonists are being developed to treat specific diseases. A classic example is oliceridine, a Gαi-biased agonist of μ-opioid receptor (μ-OR) approved by the Food and Drug Administration (FDA) in 2020 for treating acute to moderate pain11. Oliceridine effectively enhances the analgesic effect while minimizing β-arrestin-mediated side effects such as tolerance, addiction, and respiratory depression11.
Cholecystokinin (CCK) is one of the most abundant neuropeptides in the nervous system and acts in the form of sulfated CCK8s in the brain12,13,14. CCKAR, CCKBR, and GPR173 were discovered as receptors of CCK, and they serve important functions in the brain. For instance, CCKAR is involved in spatial memory and emotion15,16, CCKBR in associative memory, spatial learning, motor skill, and depression17,18,19,20, and GPR173 is related to inhibitory plasticity21. Studies have reported that CCKAR can activate Gαq/11, Gαi/o, Gαs, Gα12/13, and β-arrestin1&25,6,21,22,23. while CCKBR can also activate these signaling pathways5,6,21,23. Nevertheless, the relationship between these signaling pathways and CCK receptors’ functions in the brain has not been thoroughly understood. We recently demonstrated that a CCKBR agonist, HT-267, could be used to alleviate anterograde amnesia in CCK-deficient and aged Alzheimer’s disease mice24, and a CCKBR antagonist YM022 displayed an antidepressant-like effect in a chronic social defeat stress mouse model19. CCKBR-mediated cognitive function, emotion, and movement occur through the CCKBR-dependent long-term potentiation (CCKBR-LTP) of excitatory synaptic transmission, but the signaling mechanism of this LTP is still unclear17,19,20,24,25. Hence, revealing the signaling pathway mechanism of the CCKBR-LTP is crucial for the development of drugs targeting this receptor.
It has been more than 20 years since the cloning and purification of the CCKBR receptor in 199226, and most previous studies have focused on its Gαq/11-mediated signaling pathway27,28,29. CCKBR can also increase the excitability of small-sized dorsal root ganglion (DRG) neurons through Gαo-mediated inhibition of A-type K+ channels, thereby enhancing pain susceptibility30. The development of biased agonists and antagonists is beneficial not only for studying the mechanism of action of receptors but also for treating diseases more precisely and effectively2. Although there are many signaling pathways downstream of CCKBR, we found only one report about biased antagonists of CCKBR, and there is no report about biased CCKBR agonists. GV150013x is a Gαq/11-biased CCKBR antagonist with no antagonistic effect on the β-arrestin signaling pathway31. However, no study has systematically evaluated the bias of this antagonist or other agonists and antagonists toward individual CCKBR signaling pathways. Therefore, finding biased agonists and antagonists of CCKBR will benefit the development of CCKBR drugs and the study of its mechanism.
In this work, we use a Multielectrode Array (MEA)-based electrophysiological method to reveal the mechanism of CCKBR-LTP in the auditory cortex (AC). Meanwhile, we identify a β-arrestin biased CCKBR agonist MF-8 through fluorescence resonance energy transfer (FRET)-based binding, bioluminescence resonance energy transfer (BRET), cyclic AMP (cAMP), calcium, receptor internalization, and PRESTO-Tango assays. This biased agonist can reduce other CCKBR signaling pathways, including Gαs-induced cAMP and Gαq/11-induced calcium signaling, thereby inhibiting the CCKBR-LTP. The blockade of MF-8 to the LTP is abolished by mutating the key site N3536.55. which determines the β-arrestin signaling bias induced by MF-8. The β-arrestin blocker barbadin and knockdown of β-arrestin expression also alleviate the inhibitory effect of MF-8 on theta burst stimulation (TBS)-induced CCKBR-LTP, suggesting that the inhibition is mediated by the β-arrestin signaling pathway. Furthermore, MF-8 effectively inhibits the formation of cue-to-cue associative fear memory. In conclusion, we uncover the mechanism of CCKBR-LTP in the AC and identify a β-arrestin-biased CCKBR agonist that blocks the LTP and CCKBR-dependent cue-to-cue associative fear memory via the β-arrestin signaling pathway.
Results
MF-8 binds to CCKBR with high affinity
CCK4 (Supplementary Table. 1) contains four amino acids at the carboxyl terminus of CCK8s, and it is considered the shortest CCK fragment that activates CCKBR32. In our previous screening campaign, we found that a CCK4-modified compound HT-178 (Supplementary Fig. 1D) exhibited a good activation effect on CCKBR downstream calcium signaling. However, we found a very interesting phenomenon. A slight change in the substituent on the amino group in the red box of HT-178 (Supplementary Fig. 1D) greatly affected the activation of calcium signals downstream of CCKBR. Since the downstream signaling pathway of CCKBR could be intricate5,6,21,23, we speculated that this slight change might not change the affinity for CCKBR, but only could change the bias for the downstream signaling pathway of CCKBR. Therefore, we selected compounds with different substituents at the amino position, as well as some other compounds based on CCK4 modification, for systematic biological evaluation (Supplementary Table. 1). They were first assayed based on established calcium mobilization and PRESTO-Tango assays to differentiate their Gαq/11-Ca2+ (Table. 1) and β-arrestin2 response (Table. 1). MF-7, MF-8, MF-9, MF-10, MF-11, MF-12, and MF-13 performed poorly in activating Gαq/11-Ca2+ signaling but were highly potent in inducing β-arrestin2 signaling (Table. 1). Both calcium and PRESTO-Tango assays were performed using CCK8s as a balanced agonist control. This was necessary to minimize system bias and quantitatively evaluate their bias level. According to the bias factor β33 (Table 1, Fig. 1A), we demonstrated that MF-8 is highly β-arrestin biased compared to Gαq/11-Ca2+ signaling with β > 2 (Fig. 1A–C).
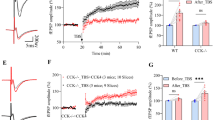
A β value was calculated based on the data from PRESTO-Tango and calcium mobilization assays. See also Table 1. CCK8s was used as the balanced agonist, n = 4. B Normalized dose-response curves of CCK8s (blue, n = 4) and MF-8 (red, n = 5) to CCKBR in calcium mobilization assay. C Normalized dose-response curves of CCK8s (blue, n = 5) and MF-8 (red, n = 4) in the PRESTO-Tango assay. D The schematic of the FRET-based binding system. E The dose-dependent response curve of MF-8 (red, n = 4) and CCK8s (blue, n = 5) at 293T cells expressing transfected HaloTag-CCKBR in the FRET assay. For (A–C) and (E), data were presented as means ± s.e.m.
To further identify these compounds, we constructed a binding system based on FRET. This system mainly consists of two components: a fluorescent probe and a modified CCKBR (Fig. 1D, E). We attached rhodamine and a HaloTag protein to the amino terminus of CCK5 and the amino terminus of CCKBR, respectively (Fig. 1D, E). After transient transfection of HaloTag-tagged CCKBR into 293 T cells, we labeled CCKBR with Fluo-Halo (Fig. 10, Supplementary Figs. 1C, 3, 4). Via this binding test, we determined that the binding affinity of MF-8 to CCKBR is close to CCK8s (IC50(MF-8) = 0.9 nM, IC50 (CCK8s) = 1.1 nM) (Fig. 1E). The above PRESTO-Tango and binding assays prove that MF-8 is a highly efficient CCKBR agonist.
The CCKBR-LTP in the AC is not dependent on β-arrestin signaling
Several critical physiological functions, such as mood, memory, and movement within the nervous system, are dependent on LTP. We have previously revealed that CCKBR regulates LTP in the AC and facilitates the formation of associative memory17. For recording CCKBR-LTP, coronal slices (300 μm) containing AC were recovered in artificial cerebrospinal fluid (aCSF) at 28 °C for 1.5 h, then positioned in a MED64 probe with continuous aCSF perfusion. Minutely field excitatory postsynaptic potentials (fEPSPs) were recorded at 30% maximal response until stable, then after 10 min of drug-treated aCSF perfusion, low-frequency electrical stimulation (LFS) was applied at 1 Hz for 60 cycles at 70% maximal response. Finally, minutely fEPSPs were re-recorded for over 60 min (Fig. 2A). Consistent with previous results, LFS along with CCK8s induced LTP in the AC (Fig. 2B, C). Since the EC50 of MF-8 equals 4.54 nM in the PRESTO-Tango assay (Fig. 1C), and its IC50 is 0.9 nM in the binding assay (Fig. 1E), therefore, 20 nM MF-8 coupled with LFS was used for LTP induction (Fig. 2B, C). However, this condition failed to induce LTP (Fig. 2B, C). We then increased MF-8’s concentration to 200 nM together with LFS to evaluate whether a higher concentration of MF-8 can induce LTP. Although a slight LTP was observed, there was no significant difference in fEPSPs before and after LFS and MF-8 treatment (Fig. 2B, C). Considering that MF-8 preferentially activates β-arrestin signaling of CCKBR, the above results demonstrate that CCKBR-LTP in AC is not dependent on β-arrestin signaling.
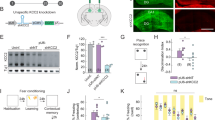
A The flow chart of the whole fEPSP recording with the representative diagram for the positions of multi-electrode arrays in the mouse AC (close to the microscope) and the drug-LFS-induced-LTP paradigm above the flow chart (Created in BioRender. Zhang, M. (2025) https:// BioRender.com/s14o899). B Normalized amplitudes of fEPSPs and (C) average change in fEPSP amplitude (%) before and after LFS with 200 nM CCK8s treatment for 10 min (light blue, N = 3 mice, 8 slices, P < 0.0001), 20 nM MF-8 (light purple, N = 2 mice, 6 slices, P = 0.7621), 200 nM MF-8 (pinkish purple, N = 2 mice, 5 slices, P = 0.5561) in C57 mice. D Normalized amplitudes of fEPSPs and (E) average change in fEPSP amplitude (%) before and after LFS with the 10-min treatment of 200 nM CCK8s in AC of C57. For (D, E), measurements were taken following a 1-h preincubation period of 100 μM barbadin (purple, N = 3 mice, 8 slices, P < 0.0001) or aCSF preincubation (bice, N = 3 mice, 5 slices, P < 0.0001). F The virus expression of AAV2/9-mCaMKIIa-MasterRNAi30e(NC)-eGFP-WPER(upper panels), AAV2/9-mCaMKIIa -MasterRNAi30e (mArrb2)-eGFP-WPER(central panels) and AAV2/9-mCaMKIIa-MasterRNAi30e (mArrb1)-eGFP-WPER (lower panels) in AC. Blue: DAPI, green: virus, red: β-arrestin; Scale bar: 50 μm. G Normalized amplitudes of fEPSPs and (H) average change in fEPSP amplitude (%) before and after LFS with the 10-min treatment of 200 nM CCK8, whose AC expressed AAV2/9-mCaMKIIa-MasterRNAi30e(NC)-eGFP-WPER (shRNA(NC), tangerine, N = 3 mice, 6 slices, P < 0.0001), AAV2/9-mCaMKIIa-MasterRNAi30e(mArrb2)-eGFP-WPER (shRNA (β-arrestin1), green, N = 3 mice, 8 slices, P < 0.0001) and AAV2/9-mCaMKIIa -MasterRNAi30e(mArrb1)-eGFP-WPER (shRNA(β-arrestin2), ocean, N = 3 mice, 7 slices, P < 0.0001). For (C, E, H), data were means ± s.e.m. values. Two-way ANOVA were used with ***P < 0.001, ****P < 0.0001; ns, not significant; Šídák's correction.
A β-arrestin signaling blocker, barbadin34, was also applied during LTP recording to further verify whether the CCKBR-LTP depends on the β-arrestin signaling pathway. We induced the LTP with CCK8s and LFS after the blocker was incubated with the brain slices for 1 h, but the LTP still maintained the same level as the control group (Fig. 2D, E). In addition, after β-arrestin1 and β-arrestin2 in the mouse AC were knocked down by adeno-associated virus (AAV)-mediated shRNA (Fig. 2F), the CCKBR-LTP could still be induced normally (Fig. 2G, H). Taken together, these pharmaceutical interventions and gene expression regulation studies reveal that the CCKBR-LTP is not dependent on β-arrestin signaling.
CCKBR-LTP in AC is Gαq/11 and Gαs dependent
Previous reports have indicated that CCKBR can activate a series of Gαi, Gαs, Gαq/11, and β-arrestin signaling pathways5,6,22,23,28, but the specific signaling pathway that determines CCKBR-dependent LTP remains unclear. To address this issue, we investigated the mechanism of the CCKBR-LTP formation by using a series of signaling pathway blockers. As TBS is an alternative method to induce the CCKBR-LTP17, we first induced LTP in AC using TBS (Fig. 3A, I), and confirmed that the LTP was CCKBR-dependent through treatment with the specific CCKBR antagonist YF476 (Fig. 3B, I). We then used Gαq/11 signaling pathway blockers U-73122, 2-APB, and YM254890 to confirm whether Gαq/11 is involved in the formation of the LTP (Fig. 3C–E, I). We found that all Gαq/11 inhibitors significantly and almost completely inhibited the formation of LTP (Fig. 3C–E, I). Also, the antagonist of the Gαs signaling pathway, H-89, partially inhibited LTP formation (Fig. 3G, I). However, PTX and barbadin, inhibitors of the Gαi and β-arrestin signaling pathway, respectively, could not inhibit the LTP (Fig. 3F, H, I). In addition, we used MF-8 to block TBS-induced CCKBR-LTP, and simultaneously applied chemogenetic elements hM3d(Gq)35 and rM3D(Gs)36 to activate Gq and Gs, respectively, under the induction of Clozapine N-oxide (CNO). Although MF-8 could block the LTP, the inhibitory effect of MF-8 on the LTP could be abolished by both hM3dq(Gq) and rM3D(Gs) under the activation of CNO (Supplementary Fig. 6A–F). These results indicate that the CCKBR-LTP in the AC depends on Gαq/11 and Gαs signaling pathways, especially the Gαq/11 signaling pathway.
A–G Normalized amplitudes of fEPSPs before and after TBS with one-hour preincubation with aCSF (Steel, N = 5 mice, 7 slices), 100 nM YF476 (Red, N = 3 mice, 7 slices), 10 μM U-73122 (Moss, N = 5 mice, 8 slices), 50 μM 2-APB (Mocha, N = 6 mice, 12 slices), 20 μM YM254890 (Dark purple, N = 3 mice, 6 slices), 100 μM Barbadin (Blue, N = 3 mice, 6 slices), 10 μM H-89 (Ocean, N = 5 mice, 10 slices) in C57 mice with representative trace of single fEPSP before (translucent color) and after (solid color) TBS. H Normalized amplitudes of fEPSPs before and after TBS, with 300 ng PTX injected into the AC of C57 24–48 h before recording (grape, N = 3 mice, 6 slices) with the representative of a single fEPSP before (translucent color) and after (solid color) TBS on above. I The average change in fEPSP amplitude (%) before (circle, translucent color) and after (square, solid color) TBS of groups. Vehicle: Steel, N = 5 mice, 7 slices, P < 0.0001; YF476: Red, N = 3 mice, 7 slices, P = 0.4533; U-73122: Moss, N = 5 mice, 8 slices, P = 0.4578; 2-APB: Mocha, N = 6 mice, 12 slices, P > 0.9999; YM254890: Dark purple, N = 3 mice, 6 slices, P = 0.901; Barbadin: Blue, N = 3 mice, 6 slices, P < 0.0001; H-89: Ocean, N = 5 mice, 10 slices, P = 0.105; PTX: grape, N = 3 mice, 6 slices, P < 0.0001. Data were presented as means ± s.e.m. values. Two-way ANOVA: ****P < 0.0001; ns, not significant; Šídák's corrected.
MF-8 is a β-arrestin-biased and CCKBR-selective agonist
To further explore the role of MF-8 in CCKBR-mediated LTP, we investigated the agonist’s cell signaling pathway bias and CCK receptor subtype selectivity through diverse cell pathway assays. Regarding the activation of the Gαi family (Gαi1, Gαi2, Gαi3, Gαoa, Gαob, and Gαz), MF-8 showed a partial activation effect on each of the Gαi subtypes (Fig. 4A–F). For the Gα12 family, MF-8 induced weak activation on Gα12 under the high concentration condition, and the potency of MF-8 on Gα13 was about 500 times weaker than CCK8s (Fig. 4G, H). MF-8 showed partial activation effects on Gαq/11 and Gαss, but had no activity on Gαsl (Fig. 4I–L). However, MF-8 had a higher potency than CCK8s in the BRET assay for the recruitment of both β-arrestin1 and β-arrestin2 (Fig. 4M, N). The β-arrestin results were consistent with a higher affinity of MF-8 than CCK8s (Fig. 1E). We then used the results of the BRET-based test of G protein (Fig. 4A–L) and β-arrestin (Fig. 4M, N) subunits to draw a radar graph of MF-8 relative to CCK8s activity (Fig. 4O). The graph indicates that MF-8 has a clear preference for β-arrestin1 and β-arrestin2. Besides, the results of cAMP assays suggest that at a high concentration of 10 µM, the activation rate of CCKBR by MF-8 was only about 17% of CCK8s’ activation (Supplementary Fig. 7A), which is consistent with the results of BRET-based Gαss and Gαsl assays (Fig. 4I, J). We also calculated the activity of MF-8 relative to CCK8s based on the results of calcium mobilization (Fig. 1B), PRESTO-Tango (Fig. 1C), and cAMP assays (Supplementary Fig. 7A), and the results suggest that MF-8 has a higher β-arrestin preference (Supplementary Fig. 7B). Overall, compared with the endogenous agonist CCK8s, MF-8 is a potent CCKBR agonist with a strong bias for β-arrestin signaling.
A–L TRUPATH profiling in 293T-CCKBR cells. A–F Concentration-response curves for CCK8s (blue) and MF-8 (red) at 293T-CCKBR cells across 6 Gαi proteins. G, H Concentration-response curves for CCK8s (blue) and MF-8 (red) at 293T-CCKBR cells across two Gα12/13 proteins. I, J Concentration-response curves for CCK8s (blue) and MF-8 (red) at 293T-CCKBR cells across two Gαs proteins. K, L Concentration-response curves for CCK8s (blue) and MF-8 (red) at 293T-CCKBR cells across two Gαq proteins. M, N Concentration-response curves for CCK8s (blue) and MF-8 (red) at 293 T cells transfected with CCKBR across β-arrestins determined by BRET-based assays. O Relative activities for MF-8 and CCK8s against CCKBR. G-protein and β-arrestin signaling profiles for MF-8 and CCK8s using the results of Fig. 4A–N. Relative activities were represented as Δlog (Emax/EC50). All data were presented as means ± s.e.m. The number of independent repeated trials n is shown on the graph.
Apart from CCKBR, GPR173 and CCKAR are CCK receptors. Hence, we also investigated MF-8’s activation activity on CCKAR and GPR173. CCK8s could activate the β-arrestin and Gαq/11 signaling pathways downstream of CCKAR (Fig. 5A, B). Although MF-8 weakly activated the β-arrestin signaling at 10 µM, it did not exhibit any agonistic effect on the Gαq/11 signaling of CCKAR (Fig. 5A, B). We previously found that CCK8s could trigger the release of calcium ions in monoclonal CHO cells overexpressing GPR17321, but MF-8 did not increase intracellular calcium levels in these cells (Fig. 5C). We also applied a BRET-based assay to test the activation of MF-8 on β-arrestin1/2 downstream of GPR173. CCK8s and PNX-14 were positive controls, as PNX is also a GPR173 ligand37. Although both PNX-14 and CCK8s could activate these two subtypes downstream of GPR173, MF-8 had no activating effect on them, even at 10 μM (Fig. 5D, E). The above results indicate that MF-8 is a highly selective β-arrestin-biased agonist for CCKBR.
A The concentration-response curve of CCK8s (red, n = 4) and MF-8 (gray, n = 3) at 293T-CCKAR cells in calcium mobilization assays. B The concentration-response curve of CCK8s (blue, n = 4) and MF-8 (gray, n = 3) at HTLA cells expressing modified CCKAR in PRESTO-Tango assays. C The concentration-response curve of CCK8s (red, n = 3) and MF-8 (gray, n = 3) at CHO-GPR173 cells in the calcium mobilization assay. D, E The concentration-response curve of CCK8s (blue, n = 4), MF-8 (gray, n = 5) and PNX-14 (red, n = 6) at 293T-GPR173 cells in the BRET-based β-arrestin assay ((D) for β-arrestin1; (E) for β-arrestin2). All data were presented as means ± s.e.m.
MF-8 is able to induce CCKBR internalization
The canonical β-arrestin signaling pathway mediates endocytosis after GPCR activation38,39. Consequently, we hypothesized that MF-8 could also induce CCKBR’s endocytosis. We designed three assay systems using β-arrestin1 and β-arrestin2 to detect fluorescence distribution changes in cells after receptor activation. 1) EYFP was fused to the N-terminus of truncated β-arrestin1 (1–382aa) (Fig. 6A). 2) Fusion expression of GFP at the C-terminus of β-arrestin2 (Fig. 6D). 3) Fusion expression of EYFP at the N-terminus of β-arrestin1 (Supplementary Fig. 10A). We co-expressed these fused fluorescent protein-expressed β-arrestins with HaloTag-tagged CCKBR in 293 T cells (Fig. 6A, D, Supplementary Fig. 10A). We labeled CCKBR with the fluorescent dye Rho-2CF3 by targeting HaloTag (Fig. 6A, D, Supplementary Fig. 10A) and then traced the changes in the distribution of β-arrestin and CCKBR under the stimulation of CCK8s or MF-8. We found that under control conditions, CCKBRs were mainly distributed on the cell membrane, while β-arrestin1 and β-arrestin2 were diffusely distributed within the cells (Fig. 6B, E, Supplementary Figs. 10B, 11). Within the CCK8s and MF-8 treatment groups, dense co-labeled signals of β-arrestins and CCKBR were evident in the cells (Fig. 6B, E, Supplementary Figs. 10B, 11). The results confirm that MF-8 and CCK8s can both induce receptor endocytosis. To compare the effects of endocytosis, we quantified the number of co-labeled signals in each group (Fig. 6C, F, Supplementary Fig. 10C). The number of co-labeling in CCK8s and MF-8 groups was significantly higher than that of the control group (Fig. 6C, F, Supplementary Fig. 10C). Meanwhile, the number of co-labeling in the MF-8-treated group was not significantly different from that of the CCK8s-treated group (Fig. 6C, F, Supplementary Fig. 10C), indicating that the level of endocytosis induced by MF-8 was equivalent to that of the endogenous agonist CCK8s. The above results demonstrate that MF-8 can effectively induce endocytosis of CCKBR.
A, D The test system schematic of (A) β-arrestin1(1-382aa) and (D) β-arrestin2-mediated CCKBR internalization assays. B, E The representative results correspond to (A) and (D). Left column: cells with Rho-2CF3 but without CCKBR agonist. Middle column: cells treated with Rho-2CF3 and 100 nM CCK8s. Right column: cells treated with Rho-2CF3 and 100 nM MF-8. Scale bar: 5 μm. C Quantification of endocytosis levels through colocalization analysis of EYFP-β-arrestin1(1-382aa) and HaloTag-tagged CCKBR. Control: blue, n = 30 cells; MF-8: red, n = 30 cells; CCK8: green, n = 34 cells. F Quantification of endocytosis levels through colocalization analysis of β-arrestin2-GFP and HaloTag-tagged CCKBR. Control: blue, n = 30 cells; MF-8: red, n = 29 cells; CCK8: green, n = 33 cells. For (C, F), all data were presented as means ± s.e.m., derived from three independent experiments; ordinary one-way ANOVA with Bonferroni’s multiple comparison was used. ****P < 0.0001; ns, not significant.
CCKBR-mediated calcium, cAMP accumulation, and LTP are blocked by MF-8 with high potency
Based on the results from our previous experiments, we hypothesized that the key signaling pathways (Gαq/11 and Gαs) mediated by CCKBR would also be inhibited by MF-8. To confirm that, we applied calcium mobilization and cAMP assays to examine whether MF-8 could reduce the intracellular calcium release and cAMP accumulation induced by CCK8s. We first tested the response of CCK8s and MF-8 using 293 T cells infected with a control virus. The results indicated that CCK8s and MF-8 had no agonistic effect on these cells (Supplementary Fig. 12). In the calcium mobilization assay, MF-8 was then incubated with 293T-CCKBR cells using a series of gradient concentrations for two hours, we then proceeded to add CCK8s at a final concentration of 15 nM to detect the increase in intracellular calcium (Fig. 7A). We found that MF-8, like YF476, inhibited the increase in calcium flux induced by CCK8s with IC50 = 22.7 nM (Fig. 7A). Simultaneously, we also examined cAMP accumulation after 2-h incubation of MF-8 with 293T-CCKBR cells (Fig. 7B). The results showed that MF-8, like YF476, inhibited the increase of cAMP induced by CCK8s with IC50 = 16 nM (Fig. 7B). Both calcium and cAMP assays showed that MF-8 inhibited the key signaling pathway of CCKBR-mediated LTP with high potency.
A Dose-dependent response of MF-8 (blue, n = 3) and YF476 (red, n = 3) blocking 15 nM CCK8s-induced calcium mobilization in 293T-CCKBR cells. B Dose-dependent response of MF-8 (blue, n = 10) and YF476 (red, n = 10) blocking 200 nM CCK8s-induced cAMP accumulation in 293T-CCKBR cells. C The TBS protocol and the schematic flow chart for MEA recording (Created in BioRender. Zhang, M. (2025) https://BioRender.com/fmc3p2t). TBS paradigm: 100 Hz (x5), 5 Hz (x4), 0.1 Hz (x10). D CCKBR-LTP in the AC was blocked by MF-8. Left: fEPSPs amplitude traces before and after TBS with Vehicle (gray, 8 brain slices from 4 mice) and MF-8 (blue, 10 brain slices from 4 mice). Right: The average change in fEPSPs amplitude (%) before (translucent color) and after (solid color) TBS of groups. Vehicle: gray, 8 brain slices from 4 mice, P = 0.0005; MF-8: blue, 10 brain slices from 4 mice, P = 0.0581. Data were presented as means ± s.e.m. Two-way ANOVA: **P < 0.01; ns, not significant; Šídák's corrected.
Based on the previously mentioned results, we hypothesized that MF-8 could suppress Gαq/11 and Gαs-mediated CCKBR-LTP in AC. We incubated 200 nM MF-8 with brain slices for 2 h and then examined the effect of MF-8 on TBS-induced CCKBR-LTP in AC (Fig. 7C). Compared with the control group, MF-8 significantly and completely inhibited the CCKBR-LTP (Fig. 7D). These results suggest that the β-arrestin-biased CCKBR agonist MF-8 can suppress LTP formation by blocking CCKBR-mediated cAMP and calcium signaling.
MF-8 blocks CCKBR-LTP in a β-arrestin dependent manner
To further verify whether MF-8 inhibits CCKBR-LTP through β-arrestin signaling, we attempted to discover the key amino acid sites in CCKBR that are associated with this pathway. Simply relying on molecular docking, such as autodock, cannot obtain accurate docking postures, especially for highly flexible polypeptide molecules40. Here, we used autodock vina to perform molecular docking of CCKBR and MF-8 (Supplementary Data 1), and then we selected the docking results based on the conformation and orientation of gastrin-17 in the Cyro-EM gastrin-17-CCKBR complex41. Molecular dynamics simulations with GROMACS were used to evaluate the binding mode of CCKBR and MF-8 (Supplementary Data 2). We monitored interactions formed between the receptor and ligand during the three simulations using multiple bioinformatics tools (Supplementary Figs. 13–15). Then point mutations in CCKBR combined with a BRET-based β-arrestin assay and PRESTO-Tango assay were applied to verify hydrogen bond interactions of MF-8 and CCKBR (Fig. 8A, Supplementary Fig. 16). As shown in Fig. 8A, CCKBR could form stable hydrogen bond interactions with MF-8 through residues N1152.65, T1112.61, H3767.39, Y3807.43, N3536.55, Y1894.60, H207ECL2, R3566.58, and H3647.27 (Fig. 8A, B, Supplementary Figs. 13–16), which was similar to the bonds formed by gastrin-17-CCKBR41. Among them, N1152.65 and H3647.27 could not form stable hydrogen bond interactions with the CCK4 part of gastrin-1741, which might explain why MF-8 could act as a potent agonist of CCKBR.
A The representative 3D structure of the ligand-receptor complex highlighting hydrogen bond interactions. The ligand backbone is depicted in light green, and the receptor and key residues are depicted in orange. Atoms Br, O, N, and H are colored magenta, red, blue, and white, respectively. Hydrogen bonds are indicated by light blue dashed lines. Residue labels are annotated with Ballesteros-Weinstein numbering. B Effects of key residue mutations in the MF-8 recognition pocket of CCKBR in response to the stimulation with MF-8 in BRET-based β-arrestin assays. The heat map of ΔpEC50 is colored according to the ΔpEC50 where ΔpEC50 = pEC50 of mutant—pEC50 of wild type. The heat map of Emax (%) is colored according to the Emax of mutations relative to that of the wild type. C Normalized amplitudes of fEPSPs and (F) Average change in fEPSPs amplitude (%) before and after TBS in AC slices of CCKBR-KO mice expressing rAAV-hsyn-mCCKBR-P2A-EGFP (pink, 9 slices from 6 mice, P < 0.0001) and rAAV-hsyn-EGFP (gray, 15 slices from 4 mice, P = 0.3543). D Normalized amplitudes of fEPSPs and (H) Average change in fEPSPs amplitude (%) before and after TBS in AC slices of CCK-BR-KO mice expressing rAAV-hsyn-hCCKBR-P2A-EGFP. Preincubation of 200 nM MF-8 blocked the CCKBR-LTP in the AC (red, N = 3 mice, n = 8 slices, P = 0.802). TBS induced CCKBR-LTP in the AC of the Vehicle group (blue, N = 3 mice, n = 6 slices, P < 0.0001). E Normalized amplitudes of fEPSPs and (J) Average change in fEPSPs amplitude (%) before and after TBS in AC slices of CCK-BR-KO mice expressing rAAV-hsyn-hCCKBR (Asn353Ala) -P2A-EGFP. Preincubation of 200 nM MF-8 did not block the CCKBR-LTP (Moss, N = 4 mice, n = 6 slices, P = 0.0001). Preincubation of the vehicle did not influence CCKBR-LTP in the vehicle group (Mocha, N = 3 mice, n = 7 slices, P = 0.0003). L Normalized amplitudes of fEPSPs and (N) Average change in fEPSPs amplitude (%) before and after TBS following 2-h preincubation of barbadin (Blue, N = 3 mice, n = 5 slices, P = 0.0064), or barbadin + 200 nM MF-8 (Red, N = 3 mice, n = 6 slices, P = 0.0059) with AC slices of C57. M Normalized amplitudes of fEPSPs and (P) Average change in fEPSPs amplitude (%) before and after TBS following 2-h incubation of 200 nM MF-8 with AC slices of C57 expressing AAV2/9-mCaMKIIa-MasterRNAi30e(NC) -eGFP-WPER (Green, N = 3 mice, n = 8 slices, P > 0.9999), AAV2/9-mCaMKIIa -MasterRNAi30e(mArrb1)-eGFP-WPER (Red, N = 3 mice, n = 5 slices, P < 0.0001), and AAV2/9-mCaMKIIa -MasterRNAi30e(mArrb2)-eGFP-WPER (Dark blue, N = 3 mice, n = 9 slices, P < 0.0001). G, I, K, O, Q Representative of single fEPSP before (translucent color) and after (solid color) TBS. Data were means ± s.e.m. For (F, H, J, N, P) Two-way ANOVA Šídák's multiple comparisons tests were used: ****P < 0.0001; ***P < 0.001; ns, not significant.
Considering the drastic effect of N3536.55 on MF-8-mediated β-arrestin signaling (Fig. 8B, Supplementary Fig. 16) and the weak effect of this site on the Gαq signaling pathway41, we speculated that mutation of this site would not affect TBS-induced CCKBR-LTP, but would eliminate the blockade of MF-8 on the TBS-induced CCKBR-LTP since the LTP mainly depends on the CCKBR-Gαq/11-mediated signaling pathway. We then adopted CCKBR knockout mice to verify the integrity of TBS-induced CCKBR-LTP in the AC (Fig. 8C). The TBS-induced LTP was completely abolished in CCKBR knockout mice (Fig. 8C, F, G). We also overexpressed CCKBR in AC via AAV injection (Supplementary Fig. 17), and TBS induced CCKBR-LTP formation regardless of whether we overexpressed the human or mouse CCKBR (Fig. 8C, F, G, D, H, I). Meanwhile, 200 nM of MF-8 completely abolished TBS-induced CCKBR-LTP in the AC of human CCKBR-overexpressing CCKBR-KO mice (Fig. 8D, H, I). However, when we overexpressed the human CCKBR with the N3536.55A mutation, we found that although TBS induced CCKBR-LTP formation, this LTP was not blocked by MF-8 (Fig. 8E, J, K). We also used barbadin to inhibit β-arrestin signaling and then determined whether MF-8 could inhibit TBS-induced LTP in the presence of barbadin. The results indicated that barbadin could indeed eliminate the inhibitory effect of MF-8 on this LTP (Fig. 8L, N, O). Meanwhile, AAV-mediated shRNA was applied to knock down β-arrestin1 and β-arrestin2 (Fig. 2F). The inhibitory effect of MF-8 on the LTP was significantly reduced after the two β-arrestins were knocked down (Fig. 8M, P, Q). These results suggest that MF-8 reduces CCKBR-LTP via a β-arrestin-dependent signaling pathway.
MF-8 treatment inhibits the CCKBR-LTP mediated cue-to-cue fear memory
Enhanced fear memories are thought to be closely linked to anxiety, depression, and post-traumatic stress disorder42,43,44. Our previous study found that the CCKBR-LTP in AC can enhance cue-to-cue fear memory, while CCKBR antagonists could inhibit the formation of fear memory17. This study found that the CCKBR-LTP in AC is mainly mediated by the Gαq/11 and Gαs signaling pathways. MF-8, as a β-arrestin biased CCKBR agonist, can inhibit the CCKBR-LTP. Therefore, MF-8 may also be able to inhibit the CCKBR-LTP mediated cue-to-cue fear memory. Here, we used the previous behavioral paradigm and found that MF-8 can indeed inhibit cue-to-cue fear memory (Fig. 9), which highlights the translational value of MF-8 in fear memory-related disorders.
A Diagram of MF-8 treatment on the day before tone-to-tone pair training, tone-to-tone pair training (day 1–3, f1 = 1k Hz, f2 = 4k Hz), baseline tests and conditioning of footshock with f2 (day 4), and freezing tests (day 5). B Bar charts show freezing percentages of the C57 mice with saline (gray, N = 12 mice) and MF-8 (red, N = 15 mice) injection. One-way ANOVA with Tukey’s multiple comparisons tests were used: ****p < 0.0001; **P < 0.01; NS, not significant. The exact P-values were shown in the source data.
Discussion
In this study, we developed a series of CCKBR agonists with β-arrestin bias using a multidisciplinary approach. Through in vitro electrophysiology, we demonstrated that CCKBR-LTP formation in AC depends on both Gαq/11 and Gαs signaling pathways, but more especially on the Gαq/11 signaling pathway. Meanwhile, MF-8, the agonist with the strongest β-arrestin bias, shows attenuating effects on both Gαq/11 and Gαs signaling pathways and can also block CCKBR-LTP. By analyzing CCKBR and ligand binding modes, we confirmed that N3536.55 is a key site affecting MF-8’s β-arrestin bias. After mutating this site, the agonist no longer has an antagonistic effect on CCKBR-LTP, which proves that MF-8 blocks CCKBR-LTP through β-arrestin signaling. In summary, we developed a CCKBR agonist with strong β-arrestin bias and revealed the signaling pathway mechanism of CCKBR-LTP. This biased agonist can mediate the blockade of CCKBR-LTP through β-arrestin signaling pathway.
The bias of signaling pathways is crucial for drug development. Few studies have focused on medications that bias CCKBR receptor signaling pathway. However, in our preliminary screening, we discovered that although MF-8 responded poorly in calcium mobilization assay, it exhibited comparable potency and efficiency to those of endogenous CCKBR ligand CCK8s in the PRESTO-Tango assay. To confirm the affinity of this agonist with CCKBR, we designed a FRET test system and discovered that the compound showed better affinity than CCK8s. These results indicate that MF-8 is a unique type of CCKBR agonist.
LTP formation is crucial to the function of the nervous system and it is important for biological functions such as memory, emotion, and movement45. Our previous research has demonstrated that CCKBR regulates associative memory, spatial learning and memory, motor learning, and emotion by mediating the formation of LTP17,18,19,20,24. Other previous research on the mechanism of LTP mainly emphasized the roles of NMDA receptors and AMPA receptors46,47. GPCRs play an important role in synaptic transmission through both presynaptic and postsynaptic mechanisms. The activation of Gαi/o-coupled GPCRs can suppress synaptic transmission by inhibiting presynaptic voltage-gated calcium channels48,49. These GPCRs can also activate postsynaptic inward rectifying potassium channels (GIRKs) to inhibit neuronal excitability50. Conversely, the inhibition of Gαi/o activity has been demonstrated to block the formation of LTP in hippocampal CA151. Group1 metabotropic glutamate receptor mediates LTP via Gαq/11 in hippocampal area CA152. Furthermore, the activation of the melanocortin-4 receptor (MC4R) can enhance LTP in hippocampal CA1 via the Gαs signaling pathway53. 5-HT2A and 5-HT7 receptors can enhance LTP in the developing prefrontal cortex in a glutamate-independent manner through Gαs signaling54. β-arrestin2 mediates opioid-induced LTP, thereby regulating opioid-mediated analgesia and allodynia55. Different GPCRs in different brain regions regulate LTP via different signaling pathways.
Evidence suggests that CCKBR can activate various G protein signaling pathways5,6,27,28,29. Moreover, we have also confirmed through multiple cell-based assays that CCKBR activates these G proteins and β-arrestin-mediated signaling pathways. MF-8 has a similar structure to CCK4 and has potent agonist activity in the PRESTO-Tango assay but weak activity in the calcium mobilization assay. In addition, endogenous CCK can induce the formation of LTP in the auditory cortex17, and β-arrestin has also been reported to mediate synaptic plasticity55. Therefore, we initially selected a concentration based on cell test results that could fully activate β-arrestin signaling pathway to test whether LTP can be induced. However, neither 20 nM nor 200 nM MF-8 successfully induced LTP, while CCK8s did. These findings led us to hypothesize whether other signaling pathways besides β-arrestin signaling pathway mediate the formation of LTP. By using antagonists of multiple G protein signaling pathways, we found that CCKBR-LTP is predominately mediated by Gαq/11 and Gαs. The poor potency of MF-8 in the calcium and cAMP assays also explains the drug’s inability to induce LTP at both 20 and 200 nM.
One may ask why does CCKBR-LTP depend on multiple G protein signaling pathways, and what is the potential mechanism behind CCKBR-LTP formation? 1) CCKBR-mediated LTP may depend on various G protein and β-arrestin signaling pathways across diverse neuronal circuits. Both excitatory CCK-positive neurons in the entorhinal cortex (EC) and medial geniculate body (MGB) can project to AC25,30. However, the EC to AC projection tends to induce heterosynaptic LTP56, whereas the MGB to AC projection potentiates homosynaptic LTP25. Different neural circuits may be regulated by different G protein signaling pathways. 2) Neuron types downstream of each circuit may be different. CCKBR receptors are expressed in diverse neuronal types in the cortex, such as pyramidal cells27, astrocytes28,29, PV+ basket cells27, and DRG neurons30. The downstream signaling pathways of the same receptor vary in different cell types57. Similarly, the signaling pathways mediated by CCKBR are potentially different in diverse cell types. For instance, CCKBR on hippocampal pyramidal cells signals through the canonical Gαq/11-mediated pathway to trigger endocannabinoid-mediated signaling events, while CCKBR on neighboring PV+ basket cells couples to the pertussis-toxin-sensitive Gαi/o pathway27. The current protocol for inducing CCKBR-LTP cannot distinguish between heterosynaptic and homosynaptic LTP. Mixed neural circuits and cell types may ultimately lead to the CCKBR-LTP showing dependence on Gαq/11 and Gαs signaling pathways. 3) CCKBR may also potentiate the LTP by increasing the expression and membrane content of AMPA receptors in the postsynaptic neurons. LTP formation largely depends on the activation of NMDA and AMPA receptors46,47. However, our previous research confirmed that CCK induced LTP even if NMDA receptor was inhibited17, suggesting that CCKBR could not induce LTP by regulating NMDA receptors. Therefore, CCKBR may influence LTP formation by regulating the expression of AMPA receptors on the cell membrane.
Biased drugs are very promising for disease treatments, because their precise targeting may reduce side effects. Our research team discovers β-arrestin-biased CCKBR agonist and this agonist has a weakening effect on Gαq/11 and Gαs signaling pathway-dependent CCKBR-LTP in AC. It has also been reported that β-arrestin-biased agonists can block GPCR-coupled downstream pathways. Carvedilol, a β-arrestin-biased agonist of the β2-adrenergic receptor (β2AR), inhibits the G protein signaling pathways of this receptor58. Activating the β-arrestin signaling pathway of the β2AR enhances skeletal muscle strength and induces hypertrophy, while the activation of its G protein signaling pathways leads to cardiac arrhythmia58. Therefore, β-arrestin-biased agonists of β2AR effectively prevent unwanted side effects and thus enhance skeletal muscle strength. Although the potential therapeutic advantages of β-arrestin-biased CCKBR agonists in treating neurological diseases and minimizing side effects remain undetermined, the discovery of such agonist provides us with a profound understanding of CCKBR functions in the nervous system and provides more perspectives and strategies for the development of drugs targeting CCKBR and other GPCRs. On the other hand, although we have proposed a binding mode of MF-8 and CCKBR through computational methods coupled with cell-based assays, this might not reveal the precise activation mechanism of the β-arrestin signaling pathway downstream of CCKBR. Decoding the structure of the MF-8 and CCKBR complex using cryo-electron microscopy may reveal the precise binding mechanisms and provide insights into the development of biased drugs that target specific CCKBR signaling pathways.
Methods
Ethics statement
All animal experimental procedures were reviewed and approved by the Animal Subjects Ethics Sub-committees of the City University of Hong Kong. The following mice were used for electrophysiology: C57BL/6 J (C57) and CCKBR-KO mice with a 129S1 background (Stock # 006369, Jackson Laboratory). All mice were randomly allocated into the experimental and control groups. Male C57 and CCKBR-KO mice, with 8–24 weeks, were used for MEA recording. Male C57 with 6–8 weeks were used for animal behavior analysis. Mice were housed at 20–24 °C with 40–60% humidity under a 12-h light/12-h dark cycle (dark from 9:00 to 21:00) and given food and water ad libitum.
Plasmid construction
Tango-CCKBR was constructed in our previous work21. To examine the effects of mutations to alanine (A) at Asn115, Gln204, His207, and Asn353 on β-arrestin signaling, Tango-CCKBR (N115A), Tango-CCKBR (Q204A), Tango-CCKBR (H207A), and Tango-CCKBR (N353A) were generated by replacing the CCKBR sequences with point mutations into Tango-CCKBR. To construct pcDNA3.1(+)-HaloTag-CCKBR, the full length of Homo sapiens CCKBR (NM_176875.4) complementary DNA was cloned into the pcDNA3.1(+) vector (Invitrogen) containing an hGH signal sequence followed by a HaloTag sequence before the receptor sequence. Full length of Homo sapiens GPR173 (NM_018969.5), CCKAR (NM_000730), and CCKBR ((NM_176875.4)) were cloned into the multiple cloning site (MCS) of pLVX-puro vector to generate pLVX-puro-GPR173, pLVX-puro-CCKAR, and pLVX-puro-CCKBR. The coding sequence of β-arrestin2 (NM_004313) with GFP conjugated to its carboxyl terminus was cloned into pcDNA3.1(+) vector to generate pcDNA3.1(+)-βarr2-GFP. To construct pcDNA3.1(+)-YFP-βarr1, the coding sequence of β-arrestin1 (NM_004041.5) with YFP conjugated to its amino terminus was cloned into pcDNA3.1(+) vector. pcDNA3.1(+)-YFP-βarr1_1-382aa was constructed by replacing nucleotides GAT encoding Asp-383 with nucleotides TAG encoding a stop codon of YFP-βarr1 in pcDNA3.1(+)-YFP-βarr1. pcDNA3.1(+)-Flag-CCKBR, pcDNA3.1(+)-Flag-CCKBR mutants, pcDNA3.1(+)-rGFP-CAAX, pcDNA3.1(+)-β-arrestin1-RlucII, and pcDNA3.1(+)-β-arrestin2 -RlucII were obtained from PPL (Public Protein/Plasmid Library, China). All final recombinant vectors were confirmed by sequencing analysis.
Generation of GPCRs overexpression-cell lines
HEK293T, 293T-puro (HEK293T cells infected with empty lentivirus), CCKAR-overexpressing HEK293T (293T-CCKAR), CCKBR-overexpressing HEK293T (293T-CCKBR), and GPR173-overexpressing HEK293T (293T-GPR173) cells were maintained in DMEM medium (Thermo Fisher Scientific), supplemented with 10% fetal bovine serum (Gibco) and 1% Penicillin-Streptomycin-Neomycin (PSN) Antibiotic Mixture (Thermo Fisher Scientific) at 37 °C in a humidified atmosphere of 5% CO2.
The procedure for generating 293T-puro, 293T-GPR173, 293T-CCKAR and 293T-CCKBR is outlined as follows. To produce the lentivirus required for GPCRs overexpression, HEK293T cells were seeded for 70%–90% confluency in 6-well tissue culture plates and incubated for 10–12 h in the mixture of calcium phosphate transfection reagent (31.25 µl 2 M CaCl2 solution, 250 µl 2 × HBS (0.05 mol/L HEPES, 0.012 mol/L D-(+)-Glucose, 0.28 mol/L NaCl, 0.023 mol/L KCl, 0.0015 mol/L Na2HPO4), 218.75 µl H2O) and 1 ml fresh medium, with 4 µg pLVX-puro-GPCR plasmids (Public Protein/Plasmid Library, Nanjing, China) or pLVX-puro, 3 µg pSPAX2 plasmid (Addgene, plasmid #12260) and 1.2 µg pMD2.G plasmid (Addgene, plasmid #12259). The cells were cultured for an additional 36 h in their normal medium before collecting the virus supernatant and infecting HEK293T cells. Meanwhile, around 1 × 106 HEK293T cells were seeded in 6-well plates, each well for 12 h, 0.5 mL virus supernatant, and 0.1% polybrene (Sigma) in 1 mL fresh normal medium were mixed and added. The second infection was performed at the interval of 24 h, and the virus supernatant mix was replaced by the fresh normal medium of HEK293T after 6–8 h of each time infection. After 2 days of the second infection, puromycin (Invitrogen) was added at the final concentration of 10 µg/mL for one week. The screened cells were going to be used for subsequent assays. Signaling pathway-based cell assays were used to validate the 293T-CCKAR (Supplementary Fig. 9) and 293T-CCKBR cell lines (Supplementary Fig. 8).
Calcium mobilization assays
HEK293T (ATCC, CRL-3216) cells stably expressing GPCRs were plated at 6 × 104 (1 × 104) cells per well in black and optical-bottom 96/384-well plates (Corning). The next day, the Ca2+ release response of cells was measured by a Fluo-8 No Wash Calcium Assay Kit (AAT-Bioquest) according to the manufacturer’s instructions. All measurements of calcium mobilization assay were performed by EnVision 2104 Multilabel Reader (Perkin Elmer) or CLARIOstar® Plus Multi-Mode Plate Reader (BMG LABTECH). The relative luminescence was exported into an Excel sheet and processed with Excel and GraphPad Prism. The cells were cultured at 37 °C, 5% CO2 in DMEM medium (ThermoFisher Scientific) containing 10% FBS (Natocor-Industria Biológica) and washed with HHBS once before incubating with Fluo-8.
The procedure for calcium mobilization assay of CHO-GPR17321 was identical to the other GPCR-overexpressing HEK293T cells apart from the culture medium. CHO-GPR173 were cultured at 37 °C and 5% CO2 in DMEM/F12 medium (Thermo Fisher Scientific) containing 10% FBS (Thermo Fisher Scientific).
PRESTO-Tango assays
PRESTO-Tango assays were performed as described in59 with some modifications. HTLA cells (A gift from the R. Axel’s Laboratory) grown in 6-well plate were transfected with 1 µg Tango-CCKAR21 or Tango-CCKBR21 per well according to the instructions of Lipo8000 (Beyotime). The transfected cells were seeded in poly-L-lysine-coated 384-well plates (JingAn Biological) at 10000 to 15000 cells per well in 50 µl of medium after 24-h culture. The medium was replaced with fresh medium containing a series concentration of compounds after 24-h culture in 384-well plate. Medium and drug solutions were removed from each well after ~18-h incubation with cells, and 20 µl per well of Bright-Glo solution (Promega) diluted 20-fold in assay buffer (1x HBSS, 2.5 mM probenecid, 20 mM HEPES, pH = 7.4) was added into each well. After ~15 min incubation at room temperature, luminescence was measured in FlexStation 3 (Molecular Devices) or CLARIOstar® Plus Multi-Mode Plate Reader (BMG LABTECH). The relative luminescence was exported into an Excel sheet and processed with Excel and GraphPad Prism.
BRET-based β-arrestin assays
The procedure of BRET-based β-arrestin assays was modified based on a previously reported paper60. HEK293T cells were seeded into 6-well culture plates and maintained in DMEM supplemented with 10%(v/v) FBS and 1%(v/v) Penicillin-Streptomycin-Neomycin Antibiotic Mixture (Gibco) at 37 °C in a humidified incubator with 5% CO₂. Plasmids encoding CCKBR (pcDNA3.1(+)-ssflag-CCKBR), an effector-RlucII fusion protein (pCDNA3.1(+)-β-arrestin1/2 -RlucII), and pCDNA3.1(+)-rGFP-CAAX were transiently transfected into cells using Lipofectamine 3000 (ThermoFisher Scientific) according to the manufacturer’s protocol. For BRET-based β-arrestin assays of GPR173, 293T-GPR173 cells were used and only plasmids containing β-arrestin1/2-RlucII and rGFP-CAAX were transfected. Twenty-four hours after transfection, cells were trypsinized, resuspended in DMEM, and seeded into white clear-bottom 96-well microplates (Greiner Bio-One) at a density of 5 × 10⁴ cells per well. The BRET substrate Coelenterazine 400a (Nanolight Technologies) was added to each well to a final concentration of 5 μM. BRET signals were acquired using a CLARIOstar microplate reader (BMG Labtech) equipped with dual emission filters (480 nm for RlucII and 530 nm for rGFP). Background luminescence was measured using untransfected cells, and background-subtracted BRET ratios were calculated as the 530 nm/480 nm emission intensity ratio.
Immunofluorescence
For brain slice immunofluorescence in Fig. 2D1, Supplementary Fig. 6A, and 17, mice were anesthetized with pentobarbital sodium and transcranial-perfused with 30 ml cold PBS and 30 ml cold 4% (w/v) PFA in PBS. Brain tissue was removed, postfixed with 4% PFA, and treated with 30% (w/v) sucrose in PBS at 4 °C for two days. Brain tissue was sectioned on a cryostat (Leica CM3500) with 30 μm thickness and preserved with antifreeze buffer (20% v/v glycerin and 30% v/v ethylene glycol diluted in PBS) at −20 °C. Brain sections in Supplementary Figs. 6A and 17 were washed three times with PBS and then incubated with DAPI for 30 min. Next, the sections were washed three times with PBS and mounted with 70% (v/v) glycerol in PBS on a slide. Brain sections in Fig. 2D1 were rinsed three times with PBS and blocked with blocking buffer (10% v/v goat serm in PBS with 0.25% v/v Triton X-100) for 1 h at room temperature. Sections were incubated with primary antibodies (1:50 dilution, β-arrestin1/2 antibody, SANTA, Catlog No: SC-74591) at 4 °C for 36 ~ 48 h. After washing thrice (15 min for each) in PBS, sections were incubated with the corresponding secondary antibodies (1:200 dilution, Goat anti-mouse 594, Jackson ImmunoResearch, Catlog No: 115-585-003) and DAPI for two hours at room temperature. We mounted all sections with 70% (v/v) glycerol in PBS on a slide after rinsing them with PBS three times. Images of the sections were acquired using Nikon Confocal Imaging System AXR and processed with Nikon NIS element and ImageJ (NIH).
FRET-based binding assays
For FRET-based binding assays using CCKBR N-terminal labeled with HaloTag, cultivation of 293 T cells was performed in DMEM supplemented with 10% fetal bovine serum (FBS) and 1% Penicillin-Streptomycin-Neomycin Antibiotic Mixture at 37 °C in a humidified 5% CO₂ atmosphere. The 293 T cells were transiently transfected with pcDNA3.1(+)-HaloTag-CCKBR using Lipofectamine™ 3000 Transfection Reagent (ThermoFisher, Catalog Number: L3000015) according to the manufacturer’s instructions. The transfected cells were seeded at 15,000-20,000 cells per well in a black transparent-bottomed 384-well microplate and allowed to adhere overnight. For fluorescence resonance energy transfer (FRET) measurements, serial dilutions of tested compounds were prepared. A dual-labeled detection system was established using 500 nM SRB-CCK5 and 1000 nM Fluo-Halo. Prior to assay initiation, growth medium was aspirated from designated wells. Aliquots of 20 μL of the prepared solution with compounds were dispensed into each well, followed by the addition of 40 μL combined probe solution to achieve final concentrations of 500 nM SRB-CCK5 and 1000 nM Fluo-Halo in a total reaction volume of 60 μL. After 5-min incubation at room temperature, FRET signals were measured simultaneously through dual excitation/emission channels (475 nm/520 nm, 475 nm/586 nm). The final concentration-dependent response curve was plotted using GraphPad Prism after subtracting the FRET ratio (F586nm/F520nm) values of the control group.
TRUPATH assays (BRET2 assays)
The TRUPATH suite (Kit#1000000163)61, which includes 14 optimized BRET Gαβγ biosensors, was obtained from Addgene. We used Lipo8000TM Transfection Reagent (Beyotime Biotechnology) for plasmid transfection according to the manufacturer’s instructions. The Gαβγ biosensor’s plasmids with the best single pathway resolution were transfected into HEK293T cells for 24 h, using a 2:1:1:1 DNA ratio of receptor: Gα-RLuc8: Gβ: Gγ-GFP2 (1 µg per construct for 6-well dishes). After 24 h transfection, cells were harvested and incubated in poly-L-lysine-coated white, clear-bottom 96-well assay plates (Greiner Bio-One) at a density of 4 × 104–6 × 104 cells per well. The next day, we inspected the cell density and plastered the white backing (PerkinElmer) to the bottom of the plate. After cautiously aspirating the growth medium, 60 μl BRET assay buffer (1× Hank’s balanced salt solution (HBSS) + 20 mM HEPES, pH 7.4) was added to each well, followed by the addition of 10 μl freshly prepared 50 μM coelenterazine 400a (Nanolight Technologies). After 5 min, cells were treated with 30ul of gradient concentration drug solution. The plate was read using a CLARIOstar plate reader (BMG LABTECH Ltd) with 395 nm (RLuc8-coelenterazine 400a) and 510 nm (GFP2) emission filters, with an integration time of 1 s per well. Plates were read serially 4 times, and the measurements from the last read were used for all analyses. BRET2 ratios were calculated as the ratio of the 510 nm emission to the 395 nm emission.
CCKBR internalization assays
HEK293T cells were transfected at a 1:1 ratio with pcDNA3.1(+)-Halotag-CCKBR and β-arrestins subunit (pcDNA3.1( + )-YFP-βarr1, pcDNA3.1( + )-YFP-βarr1_1-382aa, or pcDNA3.1(+)-βarr2-GFP respectively) by Lipo8000TM (Beyotime Biotechnology). The next day, the cells were reseeded onto poly-L-lysine-coated coverslips. After 24 h, the cells were incubated with drugs at specific concentrations and 500 nM Rho-2CF3 for 1 ~ 2 h. The cells were then fixed with 4% PFA in PBS for 15 min at room temperature. Afterward, the cells were washed thrice for five minutes in PBS. Thereafter, the cells’ attached coverslips were mounted on glass slides with Anti-Fade Fluorescence Mounting Medium (Abcam, ab104135) for imaging with Super Resolution Confocal-Leica Stellaris STED. For culture cell immunofluorescence in Fig. S11, the fixed cells were co-stained with DAPI (Chem Cruz) for 10 min at room temperature. After three washings with PBS, we mounted them on a microscope slide for imaging. For the analysis, all detection, colocalization, and quantification were performed using ComDetv.0.5.5 plugin for ImageJ (https://github.com/UU-cellbiology/ComDet). The colocalization between β-arrestin-fused fluorescent proteins and Rho-2CF3-labeled CCKBR was assessed for statistical differences. A minimum of 30 cells was utilized in each group for analysis.
GloSensor™ cAMP accumulation assays
The 293T-CCKBR cells were cultured in DMEM (Gibco) with 10% Fetal Bovine Serum (FBS) and 1% penicillin-streptomycin (P/S) in a humidified 37 °C incubator with 5% CO2. The cells were transfected by the transfection mixture, which was diluted with Opti-MEM medium with a ratio of 2 μl Lipo8000TM (Beyotime Biotechnology) per 1 μg pGloSensor™−22F. On the day after transfection, transfected cells were transferred and incubated overnight in white, clear-bottom Poly-L-Lysine-coated 96-well microplates with approximately 40000 cells per well. On the following day, the white backing (PerkinElmer) was sealed to the bottom of the plate before measurement. The medium was replaced by 50 μl D-luciferin drug buffer, which was made with the Equilibration medium (20 mM HEPES pH 7.5, 1 x Hank’s Balanced Salt Solution (HBSS, Gibco), 0.1% w/v Bovine Serum Albumin (BSA)) with 2% GloSensor™ cAMP Reagent stock solution. Cells were incubated for 2 h at room temperature with D-luciferin buffer. After that, the tested compounds were gradient diluted in HHBS buffer (20 mM HEPES, pH 7.5, 1x HBSS) at 2x final concentration. 50 μL gradient diluted drugs were added to each well of the assay plate. The Plate was read using a CLARIOstar plate reader after 30 min. All drugs or compounds used here were dissolved in dimethyl sulfoxide (DMSO) and diluted to 10 mM stocks stored at −20 °C before use.
Chemicals
YF476 (Cat. No. HY-14850) and A-71623 (Cat. No. 2411/1) were purchased from Sigma-Aldrich and Tocris Bioscience, respectively. Barbadin (Cat. No. HY-119706) and YM254890 (Cat. No. HY-111557) were purchased from MCE. CCK4, CCK8, and CCK8s were chemically synthesized by Bankpeptide Biotechnology. PNX-14 was chemically synthesized by Hefei ScierBio-Tech Co., Ltd. MF-1, MF-2, MF-3, MF-4, MF-5, MF-6, MF-7, MF-8 (Supplementary Fig. 1), MF-9, MF-10, 5r11, MF-12, and MF-13 were synthesized by Shanghai Top-peptide Bio Co.,ltd and NovoPro Bioscience Inc. Rho-2CF3 was synthesized according to our previous work62. HPLC analysis results ensured that the purity of chemicals was above 95%. Mass spectrometry further confirmed the molecular weight. We listed the chemical information in Supplementary Table. 1.
4(5)-((2-(2-((6-chlorohexyl)oxy)ethoxy)ethyl)carbamoyl)−2-(6-hydroxy-3-oxo-3H-xanthen-9-yl)benzoic acid (Fluo-Halo).
The route (Fig. 10) describes the synthetic scheme of Fluo-Halo
To a solution of 5(6)-Carboxyfluorescein (50 mg, 0.13 mmol) in dry DMF (5 ml) was added HBTU (65.62 mg, 0.17 mmol). After being stirred at room temperature for 20 min, the mixture was added HaloTag (O2) amine (59 mg, 0.27 mmol) and DIEA (46 μL, 0.27 mmol). After being stirred at room temperature for 3 h, the reaction was quenched by NH4Cl (0.5 ml). The mixture was extracted by DCM (3 × 20 ml), and the combined organic phase was washed by H2O2 (6 × 20 ml), dried over Na2SO4, filtered and concentrated in vacuo. The solvent was evaporated and the residue was purified by silica gel flash column chromatography using dichloromethane: methanol (30: 1 to 10: 1) to afford Fluo-Halo (37.2 mg, 48.1 % yield) as a yellow solid. 1H NMR (400 MHz, Chloroform-d) δ 8.40 (s, 1 H), 8.14 (d, J = 4.0 Hz, 1 H), 8.03 (s, 1 H), 7.96 (s, 0.5 H), 7.81 (s, 0.5 H), 7.81 (s, 0.5 H), 7.19 (d, J = 8.0 Hz, 0.5 H), 6.62 (d, J = 4.0 Hz, 1 H), 6.58 (s, 1 H), 6.52 (dd, J = 16.0, 12.0 Hz, 2 H), 6.44 (d, J = 8.0 Hz, 2 H), 3.67-3.60 (m, 4 H), 3.55-3.46 (m, 7 H), 3.38 (t, J = 8.0 Hz, 1 H), 1.72–1.67 (m, 2 H), 1.60-1.48 (m, 2 H), 1.38-1.25 (m, 4 H). 13C NMR (101 MHz, DMSO-d6) δ 168.24, 168.11, 164.71, 164.60, 159.68, 159.68, 151.87, 140.61, 136.19, 134.75, 129.30, 128.23, 126.52, 124.31, 123.36, 122.35, 112.77, 112.77, 109.16, 109.16, 109.11, 109.11, 102.29, 102.29, 70.24, 69.67, 69.58, 69.50, 69.37, 68.79, 68.65, 45.40, 32.05, 29.11, 26.16, 24.97. MS (ESI) (m/z): [M + H]+ calcd. for C31H33ClNO8+, 582.2; found, 582.2.
HaloTag(O2)amine, HBTU, DIEA, dry DMF, r.t., yield 48.1%.
N-(9-(4-(N-(4-((1H-indol-3-yl)methyl)−14-amino-13-benzyl-7-butyl-10-(carboxymethyl)−2,5,8,11,14-pentaoxo-3,6,9,12-tetraazatetradecyl)sulfamoyl)−2-sulfophenyl)−6-(diethylamino)−3H-xanthen-3-ylidene)-N-ethylethanaminium (SRB-CCK5).
Peptide GW(Nle)DF-NH2 (NovoPro, 15 mg, 0.025 mmol) and Lissamine rhodamine B sulfonyl chloride (17.5 mg, 0.03 mmol) were dissolved in 2 mL DMF, then the DIEA (9 μL, 0.052 mmol) was added. The mixture was stirred at room temperature for 3 h. The product SRB-CCK5 (12 mg, 41%) was purified by HPLC (Fig. S5). MS (ESI) (m/z): [M]+ calcd. for C59H70N9O13S2+, 1176.5; found, 1176.9 (Supplementary Fig. 5).
Molecular docking and molecular dynamics simulations
The Cryo-EM structure of the CCKBR in complex with gastrin-17 (PDB ID: 7F8V)41 was obtained from the RCSB Protein Data Bank. Non-protein components were removed, and the missing intracellular loop 3 (ICL3, residues 250–325) was reconstructed using MODELER (version 10.6)63, and the model with the lowest zDOPE (normalized Discrete Optimized Protein Energy) score and without traversing the intracellular gap between TM5 and TM6 was selected for subsequent molecular docking. The protonation state of the protein was assigned by adding hydrogens in UCSF ChimeraX (version 1.9)64 at pH 7.0. MF-8 was protonated at pH 7.0 using MarvinSketch (version 23.4.0, http://www.chemaxon.com/products/marvin/marvinsketch/), and no tautomers were identified. Three methods were then used to generate diverse 3D conformers. MarvinSketch provided one low-energy conformer. Open Babel (version 3.1.1)65 and RDKit (version 2024.09.2) (https://www.rdkit.org) generated 10 conformers separately, and the lowest-energy structure from each was selected as the input conformation for docking (Supplementary Data 1). The receptor was prepared using ADFR Suite (version 1.9)66, and ligands were processed with Meeko (version 0.6.1) (https://github.com/forlilab/Meeko). Molecular docking was performed using AutoDock Vina (version 1.2.5)67, generating 20 binding poses for each input conformer. Docked poses were visually inspected and those that preserved a backbone orientation consistent with gastrin-17 and formed multiple similar conformations across docking runs were considered, the one with the highest docking score was selected for molecular dynamics (MD) simulations.
For MD simulations (Supplementary Data 2), the selected receptor-ligand complex was embedded into a heterogeneous 100 × 100 Å lipid bilayer composed of POPC and cholesterol in a 55:45 ratio at pH 7, with the receptor orientation determined using PPM2.068. Protonation states of the receptor and ligand were assigned according to pH 7. The system was solvated with 22.5 Å of water on both sides of the membrane, neutralized with 0.15 M KCl, and assembled using Amber force fields (FF19SB69 for protein, Lipid2170 for lipid, GAFF271 for ligand parameters, and OPC model72 for water). The steps were carried out using CHARMM-GUI (version 3.7)73. Simulations were performed in triplicate using GROMACS (version 2023.3)74, including energy minimization, equilibration under the NPT ensemble at 303.15 K and 1 atm, and a 200 ns production run.
Binding interactions, including hydrogen bonds, salt bridges, and hydrophobic interactions, were tracked using GROMACS (version 2023.3), PLIP (version 2.4.0)75, and ProLIF (version 2.0.3)76. The key binding pose was visualized in ChimeraX.
Local virus injection in the AC
Animals were anesthetized with pentobarbital (100 mg/kg) supplemented with atropine (0.05 mg/kg, Sigma). The anesthetized mice’s skull was fixed to the stereotaxic device (RWD Life Science). The auditory cortex (AP: −2.5 to −3.0 mm posterior to the bregma, ML: 4.1 to 4.3 mm to the middle, DV: −0.600 mm from the endocranium) was labeled. Then, craniotomy was performed on the labeled site. Adeno-associated viral type 9 (AAV9) vector expressing mouse CCKBR (rAAV-CaMkIIa-mCCKBR-P2A-EGFP-WPRE-hGH, 5.0E+12vg/ml, BrainVTA) or human CCKBR (rAAV-hsyn-hCCKBR-P2A-EGFP, 5.00E + 12 μg/mL, BrainVTA) or human CCKBR with the 353 site mutation fused with an enhanced GFP (rAAV-hsyn-hCCKBR(Asn353AIa) -P2A-EGFP, 5.00E + 12 μg/mL, BrainVTA) or AAV2/9-mCaMKIIa-MasterRNAi30e(NC)-eGFP -WPER (Taitool, Shanghai, China, 2E + 12VG/ml, Cat. No. S1273-9, targeting sequence for: 5’-GCTGAGTACTTCGAAATGTCA-3’) or AAV2/9-mCaMKIIa-MasterRNAi30e(mArrb1) (Taitool, Shanghai, China, 2E + 12VG/ml, targeting sequence for mouse β-arrestin1: 5’-TCCTGGTGGATCCTGAGTATC-3’)-eGFP-WPER or AAV2/9-mCaMKIIa-MasterRNAi30e -(mArrb2)#1-eGFP-WPER (Taitool, Shanghai, China, 2E + 12VG/ml, targeting sequence for mouse β-arrestin2: 5’-GGCTTGTGGAGTAGACTTTGA-3’) or AAV2/9-mCaMKIIa -MasterRNAi30e(mArrb2)#2-eGFP-WPER (Taitool, Shanghai, China, 2E + 12VG/ml, targeting sequence for mouse β-arrestin2: 5’-GAGGAACTCTGTGCGGCTTAT-3’) or rAAV-CaMKIIa -hM3D(Gq)-EGFP-WPRE (Taitool, Shanghai, China, 1.79E + 13VG/ml, Cat. No. S0140-9-H20) or AAV2/9-mCaMKIIa-rM3D(Gs)-TS-EGFP-ER2-WPRE (Taitool, Shanghai, China, 1E + 13VG/ml, Cat. No. S0571-9) or AAV2/9-mCaMKIIa-EGFP-ER2-WPRE (Taitool, Shanghai, China, 1E + 13VG/ml, Cat. No. S0241-9) was injected into the two sides of AC of CCK-BR-KO mice. To knock down β-arrestin2, #1 and #2 viruses were mixed and injected at a 1:1 ratio. rAAV-hsyn-EGFP (5.0E+12vg/ml, BrainVTA) was injected into the AC of CCK-BR-KO mice as a negative control. The diluted AAV vectors with aCSF were injected into 4 selected locations at a speed of 10 nL/min (Nanoliter Injector, Shanghai-GAOGE).
Multielectrode array (MEA) assays
Fresh aCSF was prepared in advance from 119 mM NaCl, 2.5 mM KCl, 1.25 mM NaH2PO4, 24 mM NaHCO3, 12.5 mM glucose, 2 mM CaCl2·4H2O, and 2 mM MgSO4·7H2O, ~25 °C).
The mice were euthanized with gaseous isoflurane, after which their brain was extracted and immersed in 95% O2 and 5% CO2 bubbled cold aCSF for approximately 1 min. 300 μm coronal brain slices with auditory cortex were obtained using a vibratome (Leica VT1000S) and were incubated in a 28 °C, 95% O2 and 5% CO2 bubbled aCSF for at least 1.5 h.
The target area of the incubated slice was placed on a probe (MED-PG515A), and an anchor (Warner Instruments, Harvard) overlayed the slice to stabilize it. We adjusted the position of the slice through the camera (DP70, Olympus) connected to an inverted microscope until the region of interest could be recorded. The fresh gas bubbled aCSF was perfused into the probe, and the temperature was kept at 32-34 °C throughout the recording period. All the recordings were carried out using the MED64 system (MED, Panasonic Alpha-Med Sciences). Firstly, the I/O curve was recorded to obtain the recording voltage (the voltage value that can induce 30% of maximal response) and stimulation voltage (the voltage value that can induce 75% of maximal response). Secondly, we recorded the basal signal of synaptic response with recording voltage until more than 30 min of stable signal was obtained. Afterward, a group of specific high/low-frequency stimulation with/without drug treatments was applied before re-recording signals under recording voltage for more than an hour. Mean fEPSP amplitude/slope before and after the stimulation were analyzed using two-way ANOVA. For comparison of the LTP magnitude between different treatments, the averaged value of the last 20 min of recordings was compared statistically. The data was exported into an Excel sheet and processed with Excel and GraphPad Prism.
Cue-to-cue fear memory
This method is similar to the one we reported previously17. A brief description is given below. Two tones were selected at a frequency of 1 kHz (f1) and 4 kHz (f2) with a duration of 3 s. The tones were set at 70 dB SPL, which could evoke neuronal responses in the auditory cortex. During the first three days of training (days 1–3), f1 and f2 were presented in pairs with 0.5 s interval to C57 mice for 25 trials in each session. We conducted 4 sessions of training for each day, with 5 min interval between each session. On day 4, a baseline test was performed to calculate the percentage of freezing over a period of 10 s after the f1 and f2 tones were presented separately. The f2 tone was then conditioned for 3 trials with a footshock. A delayed conditioning was adopted, in which the f2 tone was presented firstly and the footshock and f2 tone were terminated simultaneously. On day 5, we performed a post-conditioning test to f1 and f2 tone, and the percentage of freezing was calculated. MF-8 was intraperitoneally injected at 100 μg/kg on the day before each training session, and the same volume of saline was injected into the control group mice. Tones were digitally generated using a Tucker-Davis Technologies (TDT, Alachua, FL) workstation and delivered by a coupled electrostatic speaker. The electrical stimuli of the footshock were generated using a TDT workstation and delivered by an isolator (Isoflex, AMPI, Israel). The current of the electrical stimuli of the footshock was set at 50 ~ 200 μA with a duration of 0.5 ms. The freezing percentage of freezing was analyzed with Smart 3.0 video-tracking software.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data generated and analyzed in this study are provided in this published article, the Supplementary Information, and the Source Data files. All docked models generated with different methods in this study are provided in the Supplementary Data 1. Input files, initial, final configurations, and other necessary parameters for the molecular dynamics simulations described in this study are provided in the Supplementary Data 2 and Supplementary Information Table. The MEA recording, calcium, cAMP, FRET-based binding, Presto-Tango, TRUPATH, BRET-based β-arrestin, and animal behavioral assay data generated in this study are provided in the Source Data file. Source data is available as a Source data file. Source data are provided with this paper.
References
Yang, D. et al. G protein-coupled receptors: structure- and function-based drug discovery. Signal Transduct. Target. Ther. 6, 7 (2021).
Zhang, M. et al. G protein-coupled receptors (GPCRs): advances in structures, mechanisms, and drug discovery. Signal Transduct. Target. Ther. 9, 88 (2024).
Wang, W., Qiao, Y. & Li, Z. New insights into modes of GPCR activation. Trends Pharmacol. Sci. 39, 367–386 (2018).
Calebiro, D., Koszegi, Z., Lanoiselée, Y., Miljus, T. & O’Brien, S. G protein-coupled receptor-G protein interactions: a single-molecule perspective. Physiol. Rev. 101, 857–906 (2021).
Inoue, A. et al. Illuminating G-protein-coupling selectivity of GPCRs. Cell 177, 1933–1947.e1925 (2019).
Masuho, I. et al. Rules and mechanisms governing G protein coupling selectivity of GPCRs. Cell Rep. 42, 113173 (2023).
Jean-Charles, P. Y., Kaur, S. & Shenoy, S. K. G protein-coupled receptor signaling through β-arrestin-dependent mechanisms. J. Cardiovasc. Pharmacol. 70, 142–158 (2017).
Wei, H. et al. Independent beta-arrestin 2 and G protein-mediated pathways for angiotensin II activation of extracellular signal-regulated kinases 1 and 2. Proc. Natl Acad. Sci. USA 100, 10782–10787 (2003).
Shenoy, S. K. et al. beta-arrestin-dependent, G protein-independent ERK1/2 activation by the beta2 adrenergic receptor. J. Biol. Chem. 281, 1261–1273 (2006).
Spillmann, M. et al. New insights into arrestin recruitment to GPCRs. Int. J. Mol. Sci. 21, https://doi.org/10.3390/ijms21144949 (2020).
Che, T., Dwivedi-Agnihotri, H., Shukla, A. K. & Roth, B. L. Biased ligands at opioid receptors: Current status and future directions. Sci. Signal. 14, https://doi.org/10.1126/scisignal.aav0320 (2021).
Beinfeld, M. C., Meyer, D. K., Eskay, R. L., Jensen, R. T. & Brownstein, M. J. The distribution of cholecystokinin immunoreactivity in the central nervous system of the rat as determined by radioimmunoassay. Brain Res. 212, 51–57 (1981).
Dockray, G. J., Gregory, R. A., Hutchison, J. B., Harris, J. I. & Runswick, M. J. Isolation, structure and biological activity of two cholecystokinin octapeptides from sheep brain. Nature 274, 711–713 (1978).
Agersnap, M., Zhang, M. D., Harkany, T., Hökfelt, T. & Rehfeld, J. F. Nonsulfated cholecystokinins in cerebral neurons. Neuropeptides 60, 37–44 (2016).
Matsushita, H. et al. Spatial memory impairment in OLETF rats without cholecystokinin-a receptor. Neuropeptides 37, 271–276 (2003).
Yamamoto, Y. et al. Increased anxiety behaviour in OLETF rats without cholecystokinin-a receptor. Brain Res. Bull. 53, 789–792 (2000).
Chen, X. et al. Cholecystokinin release triggered by NMDA receptors produces LTP and sound-sound associative memory. Proc. Natl Acad. Sci. USA 116, 6397–6406 (2019).
Su, J. et al. Entorhinohippocampal cholecystokinin modulates spatial learning by facilitating neuroplasticity of hippocampal CA3-CA1 synapses. Cell Rep. 42, 113467 (2023).
Zhang, X. et al. Cholecystokinin B receptor antagonists for the treatment of depression via blocking long-term potentiation in the basolateral amygdala. Mol. psychiatry 28, 3459–3474 (2023).
Li, H. et al. Cholecystokinin facilitates motor skill learning by modulating neuroplasticity in the motor cortex. eLife 13, https://doi.org/10.7554/eLife.83897 (2024).
He, L. et al. A novel CCK receptor GPR173 mediates potentiation of GABAergic inhibition. J. Neurosci.: Off. J. Soc. Neurosci. 43, 2305–2325 (2023).
Yang, F. et al. Phospho-selective mechanisms of arrestin conformations and functions revealed by unnatural amino acid incorporation and (19)F-NMR. Nat. Commun. 6, 8202 (2015).
Zeghal, M., Laroche, G., Freitas, J. D., Wang, R. & Giguère, P. M. Profiling of basal and ligand-dependent GPCR activities by means of a polyvalent cell-based high-throughput platform. Nat. Commun. 14, 3684 (2023).
Zhang, N. et al. Cholecystokinin B receptor agonists alleviates anterograde amnesia in cholecystokinin-deficient and aged Alzheimer’s disease mice. Alzheimer’s Res. Ther. 16, 109 (2024).
Li, X. et al. Cholecystokinin modulates age-dependent Thalamocortical Neuroplasticity. eLife https://doi.org/10.7554/elife.101513.1 (2024).
Wank, S. A., Pisegna, J. R. & de Weerth, A. Brain and gastrointestinal cholecystokinin receptor family: structure and functional expression. Proc. Natl Acad. Sci. USA 89, 8691–8695 (1992).
Lee, S. Y., Földy, C., Szabadics, J. & Soltesz, I. Cell-type-specific CCK2 receptor signaling underlies the cholecystokinin-mediated selective excitation of hippocampal parvalbumin-positive fast-spiking basket cells. J. Neurosci. 31, 10993–11002 (2011).
Müller, W., Heinemann, U. & Berlin, K. Cholecystokinin activates CCKB-receptor-mediated Ca-signaling in hippocampal astrocytes. J. Neurophysiol. 78, 1997–2001 (1997).
Crosby, K. M. et al. Cholecystokinin switches the plasticity of GABA synapses in the dorsomedial hypothalamus via astrocytic ATP release. J. Neurosci. 38, 8515–8525 (2018).
Yu, S. et al. Cholecystokinin type B receptor-mediated inhibition of A-type K(+) channels enhances sensory neuronal excitability through the phosphatidylinositol 3-kinase and c-Src-dependent JNK pathway. Cell Commun. Signal.: CCS 17, 68 (2019).
Magnan, R. et al. Distinct CCK-2 receptor conformations associated with β-arrestin-2 recruitment or phospholipase-C activation revealed by a biased antagonist. J. Am. Chem. Soc. 135, 2560–2573 (2013).
Steigerwalt, R. W. & Williams, J. A. Binding specificity of the mouse cerebral cortex receptor for small cholecystokinin peptides. Regul. Pept. 8, 51–59 (1984).
Rajagopal, S. et al. Quantifying ligand bias at seven-transmembrane receptors. Mol. Pharmacol. 80, 367–377 (2011).
Beautrait, A. et al. A new inhibitor of the β-arrestin/AP2 endocytic complex reveals interplay between GPCR internalization and signalling. Nat. Commun. 8, 15054 (2017).
Armbruster, B. N., Li, X., Pausch, M. H., Herlitze, S. & Roth, B. L. Evolving the lock to fit the key to create a family of G protein-coupled receptors potently activated by an inert ligand. Proc. Natl Acad. Sci. USA 104, 5163–5168 (2007).
Guettier, J. M. et al. A chemical-genetic approach to study G protein regulation of beta cell function in vivo. Proc. Natl Acad. Sci. USA 106, 19197–19202 (2009).
Treen, A. K., Luo, V. & Belsham, D. D. Phoenixin activates immortalized GnRH and kisspeptin neurons through the novel receptor GPR173. Mol. Endocrinol. (Baltim., Md.) 30, 872–888 (2016).
Ferguson, S. S. et al. Role of beta-arrestin in mediating agonist-promoted G protein-coupled receptor internalization. Science 271, 363–366 (1996).
DeWire, S. M., Ahn, S., Lefkowitz, R. J. & Shenoy, S. K. Beta-arrestins and cell signaling. Annu. Rev. Physiol. 69, 483–510 (2007).
Warren, G. L. et al. A critical assessment of docking programs and scoring functions. J. Med. Chem. 49, 5912–5931 (2006).
Zhang, X. et al. Structures of the human cholecystokinin receptors bound to agonists and antagonists. Nat. Chem. Biol. 17, 1230–1237 (2021).
Coutellier, L. & Usdin, T. B. Enhanced long-term fear memory and increased anxiety and depression-like behavior after exposure to an aversive event in mice lacking TIP39 signaling. Behav. Brain Res. 222, 265–269 (2011).
Hori, H. et al. Fear memory regulation by the cAMP signaling pathway as an index of reexperiencing symptoms in posttraumatic stress disorder. Mol. psychiatry 29, 2105–2116 (2024).
Careaga, M. B. L., Girardi, C. E. N. & Suchecki, D. Understanding posttraumatic stress disorder through fear conditioning, extinction and reconsolidation. Neurosci. Biobehav. Rev. 71, 48–57 (2016).
Nicoll, R. A. A brief history of long-term potentiation. Neuron 93, 281–290 (2017).
Diering, G. H. & Huganir, R. L. The AMPA receptor code of synaptic plasticity. Neuron 100, 314–329 (2018).
Lüscher, C. & Malenka, R. C. NMDA receptor-dependent long-term potentiation and long-term depression (LTP/LTD). Cold Spring Harb. Perspect. Biol. 4, https://doi.org/10.1101/cshperspect.a005710 (2012).
Kajikawa, Y., Saitoh, N. & Takahashi, T. GTP-binding protein beta gamma subunits mediate presynaptic calcium current inhibition by GABA(B) receptor. Proc. Natl Acad. Sci. USA 98, 8054–8058 (2001).
Takahashi, T., Forsythe, I. D., Tsujimoto, T., Barnes-Davies, M. & Onodera, K. Presynaptic calcium current modulation by a metabotropic glutamate receptor. Sci. (N. Y., N. Y.) 274, 594–597 (1996).
Lüscher, C., Jan, L. Y., Stoffel, M., Malenka, R. C. & Nicoll, R. A. G protein-coupled inwardly rectifying K+ channels (GIRKs) mediate postsynaptic but not presynaptic transmitter actions in hippocampal neurons. Neuron 19, 687–695 (1997).
Goh, J. W. & Pennefather, P. S. A pertussis toxin-sensitive G protein in hippocampal long-term potentiation. Sci. (N. Y., N. Y.) 244, 980–983 (1989).
Miura, M., Watanabe, M., Offermanns, S., Simon, M. I. & Kano, M. Group I metabotropic glutamate receptor signaling via Galpha q/Galpha 11 secures the induction of long-term potentiation in the hippocampal area CA1. J. Neurosci. 22, 8379–8390 (2002).
Shen, Y., Fu, W. Y., Cheng, E. Y., Fu, A. K. & Ip, N. Y. Melanocortin-4 receptor regulates hippocampal synaptic plasticity through a protein kinase A-dependent mechanism. J. Neurosci. 33, 464–472 (2013).
Ogelman, R. et al. Serotonin modulates excitatory synapse maturation in the developing prefrontal cortex. Nat. Commun. 15, 1368 (2024).
Chen, G. et al. β-arrestin-2 regulates NMDA receptor function in spinal lamina II neurons and duration of persistent pain. Nat. Commun. 7, 12531 (2016).
Li, X. et al. Interhemispheric cortical long-term potentiation in the auditory cortex requires heterosynaptic activation of the entorhinal projection. iScience 26, 106542 (2023).
Malhotra, D., Shin, J., Solnica-Krezel, L. & Raz, E. Spatio-temporal regulation of concurrent developmental processes by generic signaling downstream of chemokine receptors. eLife 7, https://doi.org/10.7554/eLife.33574 (2018).
Kim, J. et al. The β-arrestin-biased β-adrenergic receptor blocker carvedilol enhances skeletal muscle contractility. Proc. Natl Acad. Sci. USA 117, 12435–12443 (2020).
Kroeze, W. K. et al. PRESTO-Tango as an open-source resource for interrogation of the druggable human GPCRome. Nat. Struct. Mol. Biol. 22, 362–369 (2015).
Avet, C. et al. Effector membrane translocation biosensors reveal G protein and βarrestin coupling profiles of 100 therapeutically relevant GPCRs. eLife 11, https://doi.org/10.7554/eLife.74101 (2022).
Olsen, R. H. J. et al. TRUPATH, an open-source biosensor platform for interrogating the GPCR transducerome. Nat. Chem. Biol. 16, 841–849 (2020).
Zhang, J. et al. De novo designed self-assembling rhodamine probe for real-time, long-term and quantitative live-cell nanoscopy. ACS nano 17, 3632–3644 (2023).
Webb, B. & Sali, A. Comparative protein structure modeling using MODELLER. Current Protocols in Bioinformatics 54, 5.6.1-5.6.37, https://doi.org/10.1002/cpbi.3 (2016).
Meng, E. C. et al. UCSF ChimeraX: tools for structure building and analysis. Protein Sci.: a Publ. Protein Soc. 32, e4792 (2023).
O’Boyle, N. M. et al. Open babel: an open chemical toolbox. J. Cheminform. 3, 33 (2011).
Ravindranath, P. A., Forli, S., Goodsell, D. S., Olson, A. J. & Sanner, M. F. AutoDockFR: advances in protein-ligand docking with explicitly specified binding site flexibility. PLoS Comput. Biol. 11, e1004586 (2015).
Trott, O. & Olson, A. J. AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 31, 455–461 (2010).
Lomize, M. A., Pogozheva, I. D., Joo, H., Mosberg, H. I. & Lomize, A. L. OPM database and PPM web server: resources for positioning of proteins in membranes. Nucleic Acids Res. 40, D370–D376 (2012).
Tian, C. et al. ff19SB: amino-acid-specific protein backbone parameters trained against quantum mechanics energy surfaces in solution. J. Chem. theory Comput. 16, 528–552 (2020).
Zhang, X. et al. Structural basis for the ligand recognition and signaling of free fatty acid receptors. Sci. Adv. 10, eadj2384 (2024).
Wang, J., Wolf, R. M., Caldwell, J. W., Kollman, P. A. & Case, D. A. Development and testing of a general amber force field. J. Comput. Chem. 25, 1157–1174 (2004).
Izadi, S., Anandakrishnan, R. & Onufriev, A. V. Building water models: a different approach. J. Phys. Chem. Lett. 5, 3863–3871 (2014).
Jo, S., Kim, T., Iyer, V. G. & Im, W. CHARMM-GUI: a web-based graphical user interface for CHARMM. J. Comput. Chem. 29, 1859–1865 (2008).
Abraham, M. J. et al. GROMACS: High performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1-2, 19–25 (2015).
Schake, P., Bolz, S. N., Linnemann, K. & Schroeder, M. PLIP 2025: introducing protein-protein interactions to the protein-ligand interaction profiler. Nucleic Acids Res. 53, W463–w465 (2025).
Bouysset, C. & Fiorucci, S. ProLIF: a library to encode molecular interactions as fingerprints. J. Cheminform. 13, 72 (2021).
Acknowledgments
This work was supported by funding of JFH from the following: Hong Kong Research Grants Council, General Research Fund: CityUHK 11103922, CityUHK 11104923, CityUHK 11104524. Hong Kong Research Grants Council, Collaborative Research Fund: C1002-24W, C5053-22G. Hong Kong Research Grants Council, Senior Research Fellow Scheme: SRFS2324-1S02. Hong Kong Health Bureau, Health and Medical Research Fund: 09203656. Innovation Technology Commission of the Hong Kong SAR, China: Health@InnoHK program. We also thank the following charitable foundations for their generous support to JFH: Wong Chun Hong Endowed Chair Professorship, Charlie Lee Charitable Foundation, and Fong Shu Fook Tong Foundation. Open Access made possible with partial support from the Open Access Publishing Fund of the City University of Hong Kong.
Author information
Authors and Affiliations
Contributions
H.S., M.F.Z., and J.F.H. designed the experiments. H.S., M.F.Z., and J.F.H. wrote the draft paper. H.S., M.F.Z., J.F.H., S.W., C.H.L., Y.P.G., S.T.B., X.W., and G.C. revised the paper. H.S. and M.F.Z. designed the cell-based assays. H.S., M.F.Z., X.F.H., and G.C. conducted the cell-based assays. H.F.C. helped in the BRET-based β-arrestin tests. M.F.Z. and S.W. conducted the brain-slice experiments. H.S. and T.C. conducted the molecular docking analysis. J.Z., P.Z.W. and H.Y.S. helped in the binding assay and chemical synthesis. X.W., S.W., and H.S. performed the animal behavior test.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Niklas Doering, who co-reviewed with Gerhard Wolber, Chunyu Zeng, who co-reviewed with Yu Junyi, and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Shi, H., Zhang, M., Hu, X. et al. Discovery of a β-arrestin-biased CCKBR agonist that blocks CCKBR-dependent long-term potentiation. Nat Commun 16, 10938 (2025). https://doi.org/10.1038/s41467-025-65962-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-65962-y