Abstract
Cellular senescence contributes to age-related neurodegeneration, yet its manifestation varies across brain cell types and senescence-inducing stressors. Here, we investigated senescence hallmarks in five human brain cell lines – astrocytes, endothelial cells, microglia, oligodendrocytes, and dopaminergic-like neurons – using chronic 5-Bromodeoxyuridine treatment and validated our findings in primary cells and alternative toxin-induced models. Principal component analysis and transcriptional network inference identified both common and cell-type-specific senescence-associated transcriptional regulators (SATRs). Functional studies of TFAP4, a key SATR, revealed its role in modulating senescence phenotypes in a cell-type-dependent manner, with decreased TFAP4 expression observed in Parkinson’s Disease patient tissue and in vivo models. These results delineate distinct senescence profiles across brain cell types and highlight transcriptional regulators that may underlie senescence heterogeneity, offering insights into targeted therapeutic strategies for neurodegenerative diseases.
Similar content being viewed by others
Introduction
Parkinson’s Disease (PD) is the second most common age-related neurodegenerative disorder and is characterized by the progressive loss of dopaminergic (DA) neurons in the substantia nigra pars compacta (SNpc)1. Although age is the biggest risk factor for PD, the exact causes of DA neuron loss are unclear and no current treatment options exist to prevent or slow the degeneration. There is evidence of local inflammation in the midbrain of PD patients prior to symptom onset, as well as findings showing activated glial cells in the midbrain of PD patients2; however, an understanding of the underlying causes of this inflammaging phenotype are unknown. An emerging view, however, is that cellular senescence pathways are key mechanisms contributing to inflammation and DA neuron loss in aging and PD2,3.
Cellular senescence is a state of irreversible cell-cycle arrest in mitotic cells resulting from long-term exposure to a wide variety of stressors, such as DNA damage, oxidative stress, oncogene activation, or mitochondrial dysfunction4, and has more recently been described in post-mitotic neurons in the context of aging and neurodegeneration5. Senescence is characterized by a suite of key hallmarks, including proliferation arrest, increased senescence-associated β-galactosidase (SA β-gal) activity, elevated levels of cyclin-dependent kinase (CDK) inhibitors p21/p16/p19, expression of the senescence-associated secretory phenotype (SASP), senescence-associated heterochromatin foci (SAHF), the nuclear release of alarmin HMGB1, mitochondrial and/or lysosomal dysfunction, and morphological changes including increased nuclear size due to loss of nuclear envelope protein lamin B12,6,7,8,9. Importantly, not all of these phenotypes are present in every senescent cell or model of senescence, and there are widely reported differences in the expression of these features based on the method of senescence induction, the cell type being studied, and the timepoint at which senescence is assayed9.
Recent advancements have been made to provide “minimum criteria” to define a senescent cell in vivo based on observed phenotypes9; however, senescent cells in vivo are very low in number and can be transient in their presence as they can be cleared within days by immune cells10,11. With age, however, due to reduced immune system efficiency, these senescent cells commonly accumulate2. In contrast, in vitro models of senescence are advantageous, where an unlimited number of cells can be generated and molecular investigations into the nuanced differences in senescent phenotypes across different cell types can be made more accessible. Additionally, recent investigations have determined that senescence occurs in post-mitotic cells of the brain, such as cortical neurons in Alzheimer’s Disease (AD) and DA neurons in human and mouse models of PD, and can spread to other brain cell types in both normal aging and in the context of neurodegenerative diseases12,13,14. The characterization of cell-type-specific senescence profiles would allow for more precise, effective, and safe therapeutic interventions, particularly in complex diseases like PD, where multiple cell types contribute to pathology15. These characterizations would also help to unravel the dual roles of senescence in health and disease, ensuring that therapies target the harmful nature of senescent cells while preserving their beneficial functions16.
We systematically characterized a DNA-damage-mediated method of senescence induction in human cell lines to delineate a cell-type-specific profile of senescence utilizing chronic 5-Bromodeoxyuridine (BrdU) treatment, an established inducer of classical replicative senescence in various cell types17,18. This treatment was performed in five human cell lines, including astrocytes, endothelial cells, microglia, oligodendrocytes, and DA-like neurons. Importantly, our findings highlight cell-type-specific profiles of key hallmarks displayed in cellular senescence. After establishing this as a robust in vitro model of senescence, we confirmed this model in primary cell culture along with comparisons to other established senescence triggers such as UV irradiation, hydrogen peroxide (H2O2) treatment, paraquat (PQ) treatment, and inflammation-induced senescence19,20,21,22. Critically, we used the transcriptional profiles of the different senescent human brain cell lines to successfully isolate a unifying senescence profile independent of cell identity. We utilized regulatory network inference23 to reveal common upstream senescence-associated transcriptional regulators (SATRs), which we show to modulate specific hallmarks of senescence dependent on the cell type. We focused on characterizing the SATR TFAP424 and we interestingly observed a decrease in TFAP4 in human PD patient midbrain tissue and multiple models of aging and senescence12. Lastly, we recapitulated many of these findings in a previously characterized PQ-treated in vivo mouse model of PD19, demonstrating inflammaging and senescence in both midbrain astrocytes and DA neurons, and most interestingly, decreased levels of TFAP4. Through the investigations presented here, we have provided a detailed characterization of a senescence profiles in various brain models, which elucidated the role of cell-type-specific SATRs.
Results
Profiling canonical senescence hallmarks in human cell lines
To identify a senescence profile in each of the human cell lines, we used a common DNA damage-inducing stressor-based approach. We treated with 100 µM BrdU25 or a DMSO control for 7 days in SVG-A (astrocytes), HBEC-5i (endothelial cells), HMC3 (microglia), HOG (oligodendrocytes), and SK-N-MC (neurons with some DA-like features, such as moderate dopamine-beta-hydroxylase activity (ATCC)) lines and analyzed various well-established hallmarks of a senescence phenotype (Figs. 1A and S1A). First, we observed a significant increase in the percentage of SA β-gal positive cells following 7-day BrdU treatment across all five cell types (Figs. 1B and S1B). Additionally, when we maintained the BrdU-treated astrocyte cells for an additional 7 days (14 days total), we observed a significant increase in SA β-gal positive cells, indicating a stability of the senescence phenotype (Fig. S1C). Next, we characterized the proliferation arrest of BrdU-treated cells, a defining hallmark of senescence. We measured the growth rates across treatment groups for 48 h and observed a halt in proliferation following BrdU treatment in all cell types (Figs. 1C and S1D). We measured γH2AX (pS139) foci (SAHF) as a molecular senescence-associated marker of DNA damage and observed a significant increase in the number of γH2AX foci per cell in astrocytes, endothelial cells, and neurons (Fig. S2A, B). Next, we characterized nuclear size using staining for lamin B1 and observed a significant increase in nuclear area across all cell types except microglia (Figs. 1D, E and S3C, D). Interestingly, staining for CDK inhibitor and canonical early-stage senescence marker p217 in BrdU-treated cell lines showed a specific increase in astrocytes, endothelial cells, and oligodendrocytes, but no significant p21 upregulation was observed in microglia or in neurons (Fig. S2A, C).
A 7-day treatment timeline of human cell lines with DMSO control or 100 µM BrdU (Created in BioRender. Riessland, M. (2025) https://BioRender.com/h27e828). Features of senescence analyzed 8 days after cell plating. B Quantification of percentage of SA β-gal positive astrocytes (n = 8, 5), endothelial cells (n = 6, 6), microglia (n = 3, 4), oligodendrocytes (n = 8, 8), and neurons (n = 8, 8), where each pair represents replicates of 7-day treatment with DMSO control (grey) or 100 µM BrdU (red), respectively. C Growth rates (divisions/hour) of DMSO (grey) and 100 µM BrdU (red) treated human cell lines (n = 3 replicates). D Airyscan confocal images of human cell lines following DMSO and BrdU treatment with staining for lamin B1 (green) and Mitotracker Red CMXRos (red). Scale bars are 5 µm. E Quantification of Mitotracker Red CMXRos staining including mitochondrial mass (top), mitochondrial membrane potential (based on Mitotracker staining intensity) (middle), and mitochondrial eccentricity (bottom) and nuclear area (square µm) based on immunofluorescence staining of lamin B1 in astrocytes (n = 9, 11), endothelial cells (n = 9, 12), microglia (n = 10, 12), oligodendrocytes (n = 9, 12), and neurons (n = 9, 12), from replicates of 7-day DMSO control (grey) and 100 µM BrdU-treated (red) cells, respectively. F Z-score heatmap showing canonical senescence marker expression (including senescence-associated secretory phenotype (SASP) factors) in all cell types (astrocytes—green, endothelial cells—orange, microglia—purple, oligodendrocytes—yellow, neurons—brown) based on bulk RNAseq of DMSO (blue) and BrdU (pink) treated cell lines. G TPM expression values of highlighted senescence markers LMNB1, CDKN1A, CDKN2D, and CDKN2A and SASP factors IL6ST, IGFBP2, MMP9, and IL18 in DMSO (grey) and BrdU (red) treated cell lines. (n = 3 replicates). H Kegg Pathway Analysis from Enrichr34,35,36 of significantly dysregulated genes highlighting biologically relevant enriched pathways. Data was analyzed by two-way ANOVA with Šídák’s multiple comparisons test (B, C, E & G) and a one-sided Fisher’s exact test with Benjamini–Hochberg correction (H). All graphs show mean with error bars depicting standard deviation (ns, p > 0.05, * p < 0.05, ** p < 0.01, *** p < 0.001). Source data and exact p-values are provided as a Source Data file.
To further characterize our BrdU-induced model of senescence, we performed bulk RNA-seq of DMSO and BrdU-treated samples from all five cell lines. Extracting canonical senescence and SASP differentially expressed genes (DEGs), we revealed both common and unique transcriptional alterations following senescence induction (Fig. 1F). Specifically, nuclear envelope marker LMNB1 was decreased to significant levels in astrocytes, endothelial cells, and microglia with BrdU treatment (Fig. 1G). Focusing on genes encoding CDK inhibitors, we observed that CDKN1A (p21) was significantly increased in all cell types except microglia with BrdU treatment (Fig. 1G). We validated p21 protein levels via Western blotting and detected significant increases in BrdU-treated astrocytes, endothelial cells, and oligodendrocytes but not in microglia, consistent with observed transcriptional alterations (Fig. S2D). Alternatively, microglia, along with all other cell types, demonstrated a significant increase in the senescence driver CDKN2D (p19)26 (Fig. 1G). Surprisingly, CDKN2A (p16) was significantly downregulated in astrocytes, endothelial cells, and microglia, but was unchanged in neurons and was not expressed in oligodendrocytes following BrdU treatment (Fig. 1G). These transcriptional alterations indicate the induction of an early-stage senescence phenotype characterized by p21 and/or p19 expression rather than an upregulation of p16, contributing to longer-term maintenance of senescence7. We saw a wide variety of transcriptional alterations in SASP markers, with cell-type-specific differential upregulation of established SASP factors (IL6ST, IGFBP2, MMP9, and IL18) (Fig. 1G). Interestingly, KEGG pathway analysis revealed significantly enriched senescence-associated pathways across cell types, but in a cell-type-dependent manner (Fig. 1H).
We then investigated key senescence hallmarks through the characterization of mitochondrial and lysosomal dysfunction. We performed functional assays to unravel the mitochondrial changes in each senescent cell type using measurements of the oxygen consumption rate (OCR) based on a Seahorse XF Analyzer and Mito Stress Test in DMSO and BrdU-treated cells (Fig. S3A, B). Using MitoTracker Red CMXRos staining (Figs. 1D and S3C), we observed a significant increase in mitochondrial mass (mitochondria/cell) in senescent astrocytes, endothelial cells, and oligodendrocytes (Fig. 1E). Quantification of the mean intensity of MitoTracker Red CMXRos revealed a significant decrease in senescent astrocytes, oligodendrocytes, and neurons, highlighting a decrease in mitochondrial membrane potential (Fig. 1E). However, senescent microglia showed a significant increase in mitochondrial membrane potential based on MitoTracker Red CMXRos staining intensity (Fig. 1E). The mitochondria in the senescent cells also displayed a significantly reduced eccentricity based on their punctate appearance, as compared to the tubular, elongated mitochondria seen in the control cells (Fig. 1E). Next, we used LysoTracker Deep Red staining to quantify the number of lysosomes per cell and saw a significant increase in senescent endothelial cells and oligodendrocytes, indicating a lack of lysosomal turnover (Fig. S3D, E), which is in line with a characteristic senescence-associated increase in lysosomal mass27. Additionally, we used a cathepsin D activity assay to monitor lysosomal function revealed a significant increase in cathepsin D activity in senescent astrocytes, endothelial cells, and oligodendrocytes, with a trending increase in microglia and neurons (Fig. S3F). Taken together, these results demonstrate that BrdU-induced DNA damage was sufficient to induce hallmarks of senescence in all five human cell lines, but in a cell-type-specific manner. Where one cell type may display many senescence hallmarks, no one cell type perfectly recapitulated all established canonical senescence hallmarks.
Cell-type-specific responses to senolytic treatment
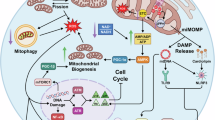
As a final investigation of comparisons amongst the cell types in the context of DNA damage-induced senescence, we characterized the responsiveness of each cell line, following DMSO or BrdU treatment, to well-established senolytic drugs: dasatinib (D) + quercetin (Q) and navitoclax (ABT-263) (Fig. 2A)28,29. First, we investigated our RNA-seq dataset for the expression of anti-apoptotic factors, which serve as targets of these senolytics. Interestingly, we observed a cell-type-specific upregulation of MCL1, BCL2, ABL1, and BCL2L1 (Fig. 2B). We tested the viability of all five cell types by exposing senescent cells to either 50 nM or 100 nM D + 10 µM or 20 µM Q and 1 µM, 10 µM, or 20 µM navitoclax for 48 h. Senescent oligodendrocytes were the most sensitive to D + Q (Fig. 2F), whereas senescent astrocytes and microglia required a higher D + Q concentration to be effectively targeted (Fig. 2C, E). Interestingly, senescent neurons and endothelial cells were relatively resistant to the same concentrations of D + Q (Fig. 2D, G). Senescent astrocytes and oligodendrocytes were the most sensitive to the tested concentrations of navitoclax (Fig. 2H, K), followed by microglia (Fig. 2J). Navitoclax treatment in senescent endothelial cells and neurons was generally less effective (Fig. 2I, L), where loss of non-senescent cells began at the same drug concentrations. Comparing the cell-type-specific responses to the senolytics to the expression of the anti-apoptotic factors (Fig. 2B), we observed a correlation in the degree of upregulation across the various factors and the sensitivity to each senolytic. For example, we saw a significant upregulation in MCL1, BCL2, ABL1, and BCL2L1 in astrocytes which concurrently displayed the greatest decrease in viability following senolytic treatment (Fig. 2B, C, H). Overall, these senolytic treatments emphasize the cell type specificity in targeting senescent cells with senolytics across the five different cell types.
A Schematic depicting mechanism of action of senolytics Dasatinib + Quercetin and Navitoclax on anti-apoptotic pathways active in senescent cells (Created in BioRender. Plessis-Belair, J. (2025) https://BioRender.com/lgfnmsj). B Z-score heatmap showing anti-apoptotic marker expression (MCL1, BCL2, ABL1, BCL2L1) in all cell lines (astrocytes—green, endothelial cells—orange, microglia—purple, oligodendrocytes—yellow, neurons—brown) based on bulk RNAseq following 7-day DMSO (blue) or BrdU (pink) treatment. C–G Normalized percent cell survival of astrocytes (C), endothelial cells (D), microglia (E), oligodendrocytes (F), and neurons (G) following 7-day DMSO (grey) and 100 µM BrdU (red) treatment with 48-h dasatinib (D) + quercetin (Q) treatment at concentrations of 50 nM D + 10 µM Q and 100 nM D + 20 nM Q (n = 3 replicates). H–L Normalized percent cell survival of astrocytes (H), endothelial cells (I), microglia (J), oligodendrocytes (K), and neurons (L) following 7-day DMSO (grey) and 100 uM BrdU (red) treatment with 48-h navitoclax treatment at concentrations of 1 µM, 10 µM, and 20 µM (n = 3 replicates). Data in (C–L) analyzed by two-way ANOVA with Šídák’s multiple comparisons test. All graphs show mean with standard deviation (ns, p > 0.05, * p < 0.05, ** p < 0.01, *** p < 0.001). Source data and exact p-values are provided as a Source Data file.
Profiling canonical senescence hallmarks in primary cell models
To establish whether BrdU treatment recapitulated the observed senescence hallmarks in primary cell culture, we treated primary human astrocytes for 7 days with 100 µM BrdU and detected a significant increase in the percentage of SA β-gal positive cells as compared to DMSO control (Fig. 3A). Next, we investigated alterations in nuclear morphology and mitochondrial dynamics and observed a significant increase in nuclear area and a significant decrease in lamin B1 signal intensity (Fig. 3B, C). Additionally, we observed an increase in mitochondrial mass (mitochondria/cell) and a decrease in mitochondrial membrane potential (Fig. 3B, C). We further determined a significant increase in the number and intensity of γH2AX foci (Fig. 3D, E), correlated with an increase in p21 intensity and an accumulation of lysosomes (Fig. 3D, E). Lastly, we performed qPCRs for senescence markers and observed a significant upregulation of CDKN1A, CDKN2A, and CDKN2D, as well as significant downregulation of LMNB1 transcripts (Fig. 3F). The concept that BrdU specifically targets cells undergoing active DNA replication was leveraged to confirm these findings in an additional primary astrocyte model in a brain co-culture, as primary post-mitotic neuronal cells should be resistant to BrdU incorporation. We co-cultured primary cortical neurons and astrocytes from P0 murine pups and subsequently treated them with DMSO and 100 µM BrdU for 7 days (Fig. 3G), leading to a significant increase in the percentage of SA β-gal positive cells (Fig. 3H). Additionally, DNA-damage as shown by elevated γH2AX intensity was sufficiently induced by BrdU treatment in the primary astrocytes, but not in the primary neurons (Fig. 3I), which was consistent with observations of an astrocyte-specific decrease in lamin B1 (Fig. 3J, K). Taken together, we were able to recapitulate our BrdU-induced model of senescence in primary astrocyte cell culture and co-culture.
A Representative images of SA β-gal staining in primary human astrocytes following 7-day DMSO or 100 µM BrdU treatment. Quantification of the percentage of SA β-gal positive cells following DMSO (grey) or BrdU (red) treatment (n = 3 replicates). B Confocal images of lamin B1 (green) and Mitotracker Red CMXRos (red) (n = 9 replicates). Scale bars are 33 µm. C Quantification of immunofluorescence staining of nuclear area (square µm), lamin B1 intensity, mitochondrial mass, and mitochondrial membrane potential (intensity) (based on (B), n = 51–115 cells). D Confocal images of γH2AX (green), p21 (red), and Lysotracker Deep Red (white) (n = 9 replicates). Scale bars are 33 µm. E Quantification of immunofluorescence staining of p21 intensity, γH2AX intensity, number of γH2AX foci per cell, and number of lysosomes per cell based on (D) (n = 68–112 cells). F Quantitative PCR of DMSO (grey) and 100 µM BrdU (red) treated primary human astrocytes for CDKN1A, CDKN2A, CDKN2D, and LMNB1 (n = 3 replicates). G Schematic depicting isolation of primary mouse cortical co-cultures with 7-day DMSO or BrdU treatment (Created in BioRender. Plessis-Belair, J. (2025) https://BioRender.com/8ttgr0z). H Representative images of SA β-gal in primary mouse co-cultures with 7-day DMSO control or 100 µM BrdU treatment. Quantification of the percentage of SA β-gal positive cells with DMSO (grey) or BrdU (red) treatment (n = 3 replicates). I Representative confocal images of γH2AX (green), Gfap (red), and Map2 (white) staining. Quantification of γH2AX intensity in Map2+ neurons (top, n = 122 DMSO,199 BrdU cells) and Gfap+ astrocytes (bottom, n = 253 DMSO, 362 BrdU cells). Scale bars are 10 µm. J Representative confocal images of lamin B1 (green) and Gfap (red) staining. Quantification of lamin B1 intensity in Gfap+ astrocytes (n = 314 DMSO, 368 BrdU cells). K Representative confocal images of lamin B1 (green) and Map2 (white) staining. Quantification of lamin B1 intensity in Map2+ neurons (n = 173 DMSO, 184 BrdU cells). Scale bars are 10 µm. Data analyzed by unpaired two-tailed t-test. All graphs show mean with standard deviation (ns, p > 0.05, * p < 0.05, ** p < 0.01, *** p < 0.001). Source data and exact p-values are provided as a Source Data file.
Toxin-dependent senescence hallmarks in a human astrocyte cell line
Just as there is no single biomarker that completely encompasses the definition of a senescence cell, there is wide heterogeneity in senescence expression based on the cell type and the trigger(s) to which those cell types are exposed30. Therefore, we sought to compare the expression of various senescence hallmarks in a toxin-dependent context in a human astrocyte cell line. To do so, we utilized previously established triggers of senescence in cell culture, including hydrogen peroxide (H2O2), paraquat (PQ), UV irradiation (UV), and inflammation through senescent astrocyte conditioned media (S/N)19,20,21,22. We then characterized the toxin-dependent senescence profile after 7-day treatments in an astrocyte cell line (Fig. 4A). We observed a significant increase in the percentage of SA β-gal positive cells in PQ, UV, and S/N-treated astrocytes but not in H2O2-treated cells (Figs. 4B and S4A). qPCR for senescence markers revealed multiple toxin-dependent changes. CDKN1A expression was significantly upregulated only in PQ treated astrocytes, with non-significantly trending upregulations in H2O2 and UV-treated cells (Fig. 4C). LMNB1 expression was significantly downregulated in PQ-treated astrocytes, like BrdU-treated cells (Fig. 1G), but was significantly upregulated with UV treatment (Fig. 4C). CDKN2A expression was also significantly downregulated in PQ-treated astrocytes, like BrdU-treated cells (Fig. 1G), but was interestingly significantly upregulated in with UV treatment (Fig. 4C). Lastly, CDKN2D was significantly upregulated in UV and S/N-treated astrocytes, consistent with BrdU-treated cells (Fig. 1G), whereas it was significantly downregulated with PQ treatment (Fig. 4C). Additional investigations into senescence phenotypes using immunostaining revealed toxin-dependent responses (Fig. S4B–E). Lamin B1 intensity was significantly decreased only with UV treatment (Fig. 4D, E). Staining with Mitotracker CMXRos revealed an accumulation of mitochondria in PQ-treated astrocytes in line with BrdU treatment (Fig. 1E), but Mitotracker intensity increased in H2O2, PQ, and UV-treated astrocytes (Fig. 4D, E). Increased γH2AX intensity and foci were found in PQ and UV-treated astrocytes (Fig. 4F, G). Interestingly, only PQ-treated astrocytes displayed increased nuclear area and elevated p21 intensity (Fig. 4F, G). Lastly, we observed a significant accumulation of lysosomes in UV and S/N-treated astrocytes (Fig. 4F, G). Altogether, PQ and UV-treated astrocytes demonstrated the widest range of expression across the tested senescence hallmarks, whereas S/N treatment showed a milder response. 7-day H2O2 treatment in astrocytes displayed many trending senescence phenotypes but comparatively failed to significantly display senescence induction. These results further highlight the robustness of our BrdU-induced in vitro senescence model through the characterization of toxin-dependent senescence expression within a single cell type.
A Schematic depicting various toxins used as triggers to induce senescence in human astrocyte cell line using 7-day treatments (Created in BioRender. Riessland, M. (2025) https://BioRender.com/te8tv71). B Quantification of the percentage of SA β-gal positive astrocytes following 20 µM hydrogen peroxide (H2O2) treatment (green), 100 µM paraquat (PQ) treatment (purple), UV irradiation (blue), or BrdU-treated astrocyte supernatant (S/N) (orange) as compared to respective controls (grey) (n = 3 replicates). C Quantitative PCR of control (grey) and toxin-treated (H2O2 green, PQ purple, UV blue, S/N orange) astrocyte cell line for senescence markers CDKN1A, CDKN2A, CDKN2D, and LMNB1 (n = 3 replicates). D Representative confocal images (merge) showing immunofluorescence staining of lamin B1 (green) and Mitotracker Red CMXRos (red) in control and toxin-treated astrocytes (top-bottom: H2O2, PQ, UV, S/N). Scale bars are 33 µm. E Quantification of immunofluorescence staining of lamin B1 (top), the number of mitochondria per cell (middle), and Mitotracker Red CMXRos intensity (bottom) in control and toxin-treated astrocytes (n = 9 replicates). F Representative confocal images (merge) showing immunofluorescence staining of γH2AX (green), p21 (red), and Lysotracker Deep Red (white) in control and toxin-treated astrocytes (top-bottom: H2O2, PQ, UV, S/N). Scale bars are 33 µm. G Quantification of immunofluorescence staining of the number of γH2AX foci per cell and overall γH2AX intensity, nuclear area (square µm), and p21 intensity based on images shown in (D) and (F) (n = 9 replicates). Data analyzed by two-way ANOVA with Šídák’s multiple comparisons test. All graphs show mean with standard deviation (ns, p > 0.05, * p < 0.05, ** p < 0.01, *** p < 0.001). Source data and exact p-values are provided as a Source Data file.
PCA reveals a cell-type-specific senescence profile
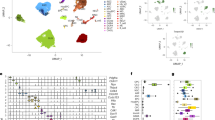
We next determined whether we could isolate a cell-type-dependent profile of senescence in an unsupervised manner. We performed a principal component analysis (PCA) as a form of dimensionality reduction on the RNA-seq data from our five human brain cell lines (astrocytes, endothelial cells, microglia, oligodendrocytes, and neurons) and treatment groups (DMSO and 100 µM BrdU). The first three principal components in a PCA dataset have been previously described to contain biologically relevant information, whereas higher-order components are typically correlated with irrelevant information or noise31,32. PCA was able to capture 94% of the explained cumulative variance within the first five principal components, and 75.1% within the first three (Fig. 5A). Comparing the correlation coefficients across different cell types and treatment groups showed a strong correlation within cell type groups, suggesting that cells will predominantly maintain their cell identity even following chronic BrdU treatment (Fig. 5B). Based on our phenotypical clustering of both cell type and treatment group, we concluded that PC1 clusters conditions by cell-type-independent of the treatment group, whereas PC2 continues to maintain cell type identity yet is influenced by treatment (Fig. 5C). Strikingly, PC3 is capable of unanimously isolating a senescence signature and separating BrdU samples from DMSO controls, independent of cell types (Fig. 5C). When observing the genetic contributions to the PCA loadings from PC3, one can identify that the most enriched positive and negative loadings associated with PC3 are inversely contributing to PC1, or cell type (Fig. 5D). Notably, genes such as SLC7A1, HSP90AB1, PARP4, POMP1, and ENO1, are all observed to be contributing to the senescence phenotype as described by PC3 (Fig. 5D). Combining contributions from PC1 and PC3 allows us to isolate a cell-type-specific signature of BrdU-induced senescence (Fig. 5E). To further characterize these cell-type-specific contributions, we directly compared treatment groups through PCA and show that treatment contributes to the greatest amount of variance within cell type comparisons through PC1 as shown by distinct clustering of treatment groups (Fig. 5F). Next, we compared the contribution of those gene loadings across cell types and interestingly, the top 10 most highly enriched genes contributing to PC1 (senescence) in each cell type were specific to that cell type yet were never inversely observed in other cell types (except for MYH9 in microglia) (Fig. 5G). Altogether, each human brain cell type had its own senescence identity following DNA-damage induction with BrdU treatment. Despite this, we were able to isolate a senescence signature independent of cell type based on the dimensionality reduction of the transcriptomic data.
A Percentage of explained variance of each dimension in the principal component analysis (PCA) of 7-day DMSO control and 100 µM BrdU-treated human cell lines (astrocytes, endothelial cells, microglia, oligodendrocytes, and neurons). B Correlation matrix of DMSO control and BrdU-treated human cell lines. C PCA loadings from PC1, PC2, and PC3 of DMSO control (black) and BrdU (red) treated human cell lines. D PCA loadings of genetic contributions towards PC1, PC2, and PC3 from (C). (Red) and (blue) data points constitute positive and negative loadings from PC3, respectively. E Two-dimensional graphical visualizations of PC1 and PC2, and PC1 and PC3 from (C) with the top genetic contribution vectors overlayed from PC3. F PC1 loadings from a PCA directly comparing 7-day treated DMSO control (black) and 100 µM BrdU (red) within cell types. G PCA loadings of the top 10 genetic contributors to PC1 for each cell type plotted across all cell types (50 genes per cell type). Data from (D) and (G) were analyzed using exploratory multivariate analysis (PCA) to identify features contributing to major axes of variance. Data derived from RNA-seq analysis from Fig. 1 (n = 3 replicates). Source data are provided as a Source Data file.
Identifying senescence-associated transcriptional regulators
The identification of a common senescence gene expression signature prompted us to investigate common regulatory gene networks. To determine specific transcriptional perturbations, we inferred and reconstructed a transcriptional network utilizing the ARACNe-AP algorithm and a previously established set of human transcription factors (TF)23,33. Based on this reconstructed transcriptional network, we utilized one- and two-tailed gene set enrichment analyses comparing each treatment group across each cell type to isolate the regulon activity of each TF (Fig. 6A). We then further normalized the differential enrichment scores (NES) for gene set size and unbiasedly took the top enrichment scores for activated (positive) and repressed (negative) regulators upon senescence induction and found significantly enriched regulators with both common and cell-type-specific enrichment (Fig. 6B). We further compared the differential expression of these regulators across cell types and ranked them based on their average expression (Fig. 6B). We then focused on a subset of transcriptional regulators meeting two requirements: the transcriptional regulator was (a) both significantly enriched and differentially expressed across cell types and (b) had a known biologically relevant function in cell fate determination (e.g., senescence, apoptosis, cell cycle etc.) (Fig. S5A). The resulting SATRs were further distinguished by observing the downstream transcriptional targets. We focused on SOX12, CHCHD3, PA2G4, and TFAP4, and as a comparison to commonly enriched regulators, we also included ERF as a regulator that is significantly enriched in most but not all cell types and is also differentially expressed (Fig. 6B). We then isolated a list of predicted transcriptional targets and observed that SATR TFAP4 had the highest mutual information (MI) scores with targets, suggesting greater regulatory influence compared to other SATRs (Fig. 6C). Enrichr34,35,36 reaction pathway analysis of these downstream targets highlighted various cell cycle dysregulation pathways for PA2G4, TFAP4 and CHCHD3, along with defects in vitamin metabolism and cholesterol biosynthesis pathways for SOX12 (Fig. 6D). Other notable significantly enriched pathways involved DNA repair, translation, and SASP (Fig. 6D). There were no significant pathways observed for ERF (Fig. 6D). Lastly, investigations into the expression of these SATRs revealed a general age-dependent decline in the human SNpc (Fig. S8; RNA-sequencing data from 179 human SNpc samples obtained from the Genotype-Tissue Expression (GTEx) Project37). Overall, we were able to isolate SATRs which were predicted to contribute to the senescence signature following BrdU-induced DNA damage across all five cell types.
A Schematic depicting analytic approach using ARACNe-AP algorithm and a previously established set of human transcription factors62 along with two-tailed gene set enrichment analyses (GSEA2) comparing each treatment group across each cell type to isolate the activity of each regulon (Created in BioRender. Plessis-Belair, J. (2025) https://BioRender.com/k7y2xkh). Positive and negative regulon targets were separated by Pearson’s correlation. Statistical significance was assessed using permutation testing (n = 1000), with p-values adjusted by the Benjamini–Hochberg method. B Gene expression z-score from the top 39 activated and repressed transcriptional regulators of DMSO control (blue) vs. 100 µM BrdU treatment (pink) from all 5 cell types (astrocytes—green, endothelial cells—orange, microglia—purple, oligodendrocytes—yellow, neurons—brown) and normalized enrichment score (NES) of 100 µM BrdU-treated (pink) samples. C Top 5 mutual information (MI) scores from TFAP4, PA2G4, CHCHD3, ERF, and SOX12 transcriptional networks. D Regulator Reaction Pathways from Enrichr34,35,36 analysis of inferred PA2G4, SOX12, CHCHD3, ERF, and TFAP4 transcriptional networks (padj < 0.05). No significant pathways isolated from ERF transcriptional network. Data from (D) was analyzed by a one-sided Fisher’s exact test with Benjamini–Hochberg correction. Source data are provided as a Source Data file.
To functionally characterize the selected SATRs, we knocked down (KD) ERF, CHCHD3, SOX12, PA2G4, and TFAP4 in all five human cell lines (Figs. S5A–S7). In addition to their predicted biological pathways (Fig. 6D), each of these selected regulators have been previously described in the context of cell cycle dysregulation, apoptosis, and cellular senescence (Fig. S5A)24,38,39,40,41,42. We sought to determine which senescence hallmarks could be recapitulated solely by the repression of the various SATRs as compared to our BrdU-induced senescence model. First, we examined the percentage of SA β-gal positive cells following treatment of all five cell types with scrambled control (siSCR) and regulator KD (siRegulator). KD of ERF, SOX12, and PA2G4 resulted in a significant increase in the percentage of SA β-gal positive astrocytes and KD of ERF led to a significant increase in SA β-gal positive microglia (Fig. S5B). Next, we performed qPCRs for senescence markers. With KD of ERF, we observed a significant senescence-like upregulation of CDKN1A in neurons and downregulation of LMNB1 in astrocytes and oligodendrocytes (Fig. S5C). siRNA KD of CHCHD3 showed significant upregulation of CDKN1A in astrocytes and upregulation of CDKN2A in neurons with significant downregulation of LMNB1 in microglia (Fig. S5D). SOX12 KD resulted in the significant downregulation of LMNB1 in astrocytes and oligodendrocytes (Fig. S5E). KD of PA2G4 showed a significant increase in CDKN1A in microglia and oligodendrocytes as well as a significant upregulation of CDKN2A in microglia, whereas no significant changes in LMNB1 levels were seen in any cell type (Fig. S5F). Lastly, KD of TFAP4 resulted in a significant upregulation of CDKN1A in astrocytes, oligodendrocytes, and neurons and an interestingly significant increase of CDKN2D levels in microglia (Fig. S5G), which mirrored what we saw with BrdU treatment (Fig. 1G). Additionally, TFAP4 KD astrocytes, microglia, oligodendrocytes, and neurons all displayed significantly diminished LMNB1 levels (Fig. S5G). Based on the functional characterization of our selected SATRs, we can conclude that each regulator contributes to a cell-type-specific signature of subsets of cellular senescence phenotypes. Importantly, no single regulator was able to fully recapitulate all the tested hallmarks in all cell types, suggesting a pathway-specific regulation of senescence hallmarks.
Cell-type-specific loss of TFAP4 predicts senescence phenotypes
TFAP4 has been well-established as a regulator of p21/p16 expression via MIR22HG43, which is a miRNA that we have previously shown to regulate DA neuron senescence via PD risk gene GBA44. First, PCA of TFAP4 transcriptional targets identified through gene set enrichment analysis revealed a senescence-associated signature in all cell types (Fig. 7A, B). Based on our results showing that KD of TFAP4 led to gene expression changes characteristic of a senescence phenotype (Fig. S5) and previous findings that loss of TFAP4 and associated increases in MIR22HG resulted in senescence phenotypes24 (Fig. 7C), we chose to focus on TFAP4 to further characterize senescence hallmarks associated with siRNA KD (Fig. 7D, E). We looked at MIR22HG levels following TFAP4 KD in all five cell types and observed a significant increase in microglia, oligodendrocytes, and neurons with TFAP4 KD (Fig. 7F). We immunostained siSCR and siTFAP4-treated cells for lamin B1 and saw a significant decrease in lamin B1 intensity with TFAP4 KD in microglia and oligodendrocytes (Figs. 7G and S9A, B). Interestingly, we saw a significant increase in nuclear area in TFAP4 KD astrocytes (Fig. 7H). Based on our earlier observations of mitochondrial and lysosomal dysfunction across cell types following BrdU treatment (Figs. 1 and S3), we used Mitotracker Red CMXRos and Lysotracker Deep Red staining in TFAP4 KD cells to investigate whether a loss of TFAP4 led to a disruption or lack of turnover of mitochondria and/or lysosomes in any cell type and observed a significant increase in the number of mitochondria per cell (Figs. 7J and S9A) and in the number of lysosomes per cell (Figs. 7K and S9B) in astrocytes. We further characterized siTFAP4 cells for γH2AX foci as a molecular marker of DNA damage and showed that astrocytes displayed an increased number of γH2AX foci/cell following TFAP4 KD (Figs. 7I and S9A). Taken together, these results further demonstrate that targeting a SATR such as TFAP4 can recapitulate senescence hallmarks in a cell-type-dependent manner, particularly in astrocytes.
A Two-tailed Gene Set Enrichment Analysis of TFAP4 from inferred transcriptional network in BrdU-treated cells. Left—Activated (red) or inhibited (blue) target elements positions in ranked gene list based on log2FoldChange. Center—Normalized Enrichment Scores (NES) and associated p-values. Right—TFAP4 Transcripts per Million (TPM) values in DMSO (grey) and BrdU (red) treated cells. Positive and negative regulon targets separated by Pearson’s correlation. Statistical significance assessed using permutation testing (n = 1000) and p-values adjusted by Benjamini–Hochberg method. Data derived from RNA-seq analysis from Fig. 1 (n = 3 replicates). B Two-dimensional graphical visualizations based on PCA of TFAP4 transcriptional targets representing PC1 and PC2; top genetic contribution vectors overlayed from PC1. C Schematic depicting TFAP4 regulation of MIR22HG24, inducing senescence (Created in BioRender. Riessland, M. (2025) https://BioRender.com/xzt9mn0). D Normalized TFAP4 protein expression from Western blots (Fig. S6) in siSCR (grey) and TFAP4 KD (red) cells (n = 3 replicates). E Fold change TFAP4 RNA levels from quantitative PCRs (Fig. S7) in siSCR (grey) and TFAP4 KD (red) cells (n = 3 replicates). F Quantitative PCR of siSCR (grey) and siTFAP4 (red) treated cells of MIR22HG levels (n = 3 replicates). G Quantification of lamin B1 immunofluorescence staining in siSCR (grey) and siTFAP4 (red) treated cells (n = 9 replicates). H Quantification of nuclear area (µm2) based on lamin B1 staining in siSCR (grey) and siTFAP4 (red) treated cells (n = 9 replicates). I Quantification of γH2AX staining in siSCR (grey) and siTFAP4 (red) treated human cell lines (n = 9 replicates). J Quantification of Mitotracker Red CMXRos in siSCR (grey) and siTFAP4 (red) treated cells (n = 9 replicates). K Quantification of Lysotracker Deep Red in TFAP4 KD (red) and controls (grey) (n = 9 replicates). L Confocal images of age-matched control and PD patient human midbrain sections: TFAP4 (red), TH (white), and GFAP (green). Scale bars are 5 µm. Data analyzed by two-way ANOVA with Šídák’s multiple comparisons test. All graphs show mean with standard deviation (ns, p > 0.05, * p < 0.05, ** p < 0.01, *** p < 0.001). Source data and exact p-values are provided as a Source Data file.
Investigations into TFAP4 expression in both in vitro and in vivo brain models of PD and senescence further revealed a loss of TFAP4 in a cell-type-specific manner. Immunostaining of aged-matched control and PD patient tissue revealed a loss of TFAP4 in DA neurons and astrocytes in the SNpc (Fig. 7L). Further investigations into our well-established SATB1-KO DA neuronal model of PD and senescence12 demonstrated a significant downregulation of TFAP4 (Fig. S10A). Concurrently, BrdU-treated primary human astrocytes and primary mouse astrocytes in a co-culture (Fig. 3) both showed a significant downregulation of TFAP4 (Fig. S10B, C, E). Altogether, the loss of TFAP4 across multiple models of PD and senescence confirmed its potential as a crucial SATR.
Paraquat induces DA neuron loss and midbrain inflammation
It has previously been shown that the environmental toxin PQ is capable of inducing DA neuron loss and induction of astrocyte senescence both in vitro and in vivo19. To further establish this in vivo model in the context of PD-relevant pathology and midbrain inflammation, we wanted to recapitulate the findings previously described19 at a later timepoint (2 weeks). We observed a significant loss of DA neurons in the midbrain of PQ-treated mice, with no significant changes in the numbers of astrocytes or microglia (Fig. 8A, B). This observed loss was confirmed to be exclusive to DA neurons, as the total number of NeuN positive neurons in the midbrain was unchanged following PQ treatment (Fig. S11A). Morphological analysis of astrocytes revealed significant changes in astrocytic branching, with PQ-treated astrocytes showing a decreased number and length of branches consistent with a decreased astrocytic domain area45 (Fig. 8C). Further investigations into microglial reactivity showed amoeboid-like morphology with decreased ramification as described by increased cell body volume, decreased branching and increased branch length46,47 (Fig. 8D). This glial atrophy is indicative of a glial transition into a disease-like state with PQ treatment, consistent with an induction of midbrain inflammation.
A Confocal slide scanner images of Gfap (green), Iba1 (red), and TH (white) staining in the midbrain of saline and PQ treated mice. Scale bars are 100 µm. Substantia nigra (SN) indicated with dashed line. B Quantification of the % of Gfap+ astrocytes, % of Iba1+ microglia, and % of TH+ neurons per total number of cells (DAPI) based on images shown in (A) in saline (grey) and PQ treated (purple) mouse midbrain sections (n = 4 replicates). C Morphological analysis of Gfap+ (green) astrocytes in saline and PQ treated mouse midbrains with skeletonization and quantification of domain area (µm2), # of branches, # of junctions, and average branch length (µm) in saline (grey) and PQ treated (purple) midbrain astrocytes (n = 411 Saline, 540 PQ cells). D Morphological analysis of Iba1+ (red) microglia in saline and PQ treated mouse midbrains with skeletonization and quantification of cell body volume (µm3), # of branches, # of junctions, and average branch length (µm) in saline (grey) and PQ treated (purple) midbrain astrocytes (n = 685 Saline, 1400 PQ cells). Scale bars are 5 µm. Data analyzed by unpaired two-tailed t-test. All graphs show mean with standard deviation (ns, p > 0.05, * p < 0.05, ** p < 0.01, *** p < 0.001). Source data and exact p-values are provided as a Source Data file.
Paraquat induces senescence hallmarks and alters TFAP4 levels
We sought to utilize this model of PD-relevant midbrain inflammation to characterize senescence hallmarks in a cell-type-specific manner in vivo. Previously, Chinta et al. described a senescence-like phenotype in midbrain astrocytes 1-week following PQ treatment through immunostaining of lamin B1 and Hmgb119. We were able to recapitulate a significant decrease in lamin B1 and Hmgb1 signal intensity in astrocytes 2-weeks following PQ treatment (Fig. 9A, B, D). Additionally, we observed a significant decrease in Hmgb1 in DA neurons, but no significant changes in lamin B1 intensity (Fig. 9A, B, D). Interestingly, we observed similar changes in Hmgb1 expression in BrdU-treated primary mouse astrocytes in a co-culture (Fig. S10D, F). Investigating the expression of SATR Tfap4 in the midbrain of PQ-treated mice, we saw a significant decrease in both astrocytes and DA neurons (Fig. 9C, D). Taken together, the observed DA neuron loss, midbrain inflammation, expression of senescence hallmarks, and loss of Tfap4 in PQ-treated mice suggests senescence pathways and our identified SATRs as PD-relevant pathological drivers underlying aging and neurodegenerative phenotypes.
A–C Confocal Images of Lamin B1, Hmgb1, and Tfap4 (red) in dopamine (DA) neurons (TH, white) and astrocytes (Gfap, green) in the midbrain of saline and PQ treated mice. Scale bars are 33 µm. D Quantification of intensity from (A–C) in DA neurons and astrocytes of Lamin B1 (n = 260 Saline, 563 PQ neurons, and n = 699 Saline, 846 PQ astrocytes), Hmgb1 (n = 716 Saline, 1087 PQ neurons, and n = 1965 Saline, 2205 PQ astrocytes), and Tfap4 (n = 117 Saline, 244 PQ neurons, and n = 1361 Saline, 1436 PQ astrocytes) intensity. Data analyzed by unpaired two-tailed t-test. All graphs show mean with standard deviation (ns, p > 0.05, * p < 0.05, ** p < 0.01, *** p < 0.001). Source data and exact p-values are provided as a Source Data file.
Discussion
The work presented here highlights cell type specificity in the expression of many established senescence hallmarks, while also shedding light on the alternative context of trigger-specific expression of diverse senescence profiles. Chronic treatment with BrdU in astrocyte, endothelial, microglial, oligodendrocyte, and neuronal cell lines was able to induce the expression of several established hallmarks of cellular senescence across cell types, including elevated SA β-gal (Fig. 1B) and proliferation arrest (Fig. 1C). Importantly, when we cultured these senescent BrdU-treated astrocytes for an additional 7-days, they maintained the increase in SA β-gal (Fig. S1C). All cell types other than microglia displayed both BrdU-dependent increases in the nuclear area (Fig. 1E) and in p21 expression levels (Figs. 1G and S2A, C, D). Interestingly, senescent microglia displayed elevated CDKN2D (p19) levels, which is in line with reports of p19-positive microglia from P301S tau mice in the context of AD48. All other cell types also showed a significant upregulation of p19 (CDKN2D) expression following BrdU treatment, emphasizing its potential as a more representative CDK inhibitor driving senescence in this context. When we applied the same BrdU treatment to primary astrocyte cell culture models, we observed a similar robust expression of most assayed senescence hallmarks, and saw an additional significant upregulation of CDKN2A (p16) following BrdU treatment (Fig. 3F). As we demonstrate in the context of BrdU treatment in the human cell lines, neither p16 nor p21 is a universal marker of senescence across cell types, at least 7 days following BrdU-induced DNA damage. Thus, p21/p16 alone is not sufficient to define the senescence phenotype across the five cell types, further highlighting the importance of assaying various senescence hallmarks. In comparison, markers such as SA β-gal, proliferation arrest, and increased nuclear size and/or decreased LMNB1 were more universally observed across cell types when exposed to the same method of senescence induction. We also investigated cell-type-specific responsiveness to well-established senolytic compounds D + Q and navitoclax28,29 and observed high sensitivity of senescent oligodendrocytes and astrocytes to both senolytics (Fig. 2), with BrdU-treated microglia also showing navitoclax sensitivity. The differences we observed in the ability of the two well-characterized senolytic cocktails to eliminate BrdU-induced senescent cell lines offer an interesting perspective for the future development of senolytic drugs, which should further consider varying responsiveness based on the senescence profile of different cell types. Overall, we highlight that although the same senescence-inducing stressor was used, each cell type expressed its own senescence profile based on variability in the hallmarks observed as well as the degree of their expression.
We sought to address the notion that a single cell type may be exposed to a variety of different stressors that may contribute to the development of a senescence phenotype. Utilizing our in vitro model of senescence, we compared treatments with various established triggers of senescence (hydrogen peroxide (H2O2), paraquat (PQ), UV irradiation (UV), and inflammation through BrdU astrocyte conditioned media (S/N)19,20,21,22) within a human astrocyte cell line and demonstrated toxin-dependent senescence-associated responses. PQ and UV treatments induced an increase in the percentage of SA β-gal positive cells (Fig. 4B), significant and trending increases in CDKN1A (p21) levels (Fig. 4C), decreases in lamin B1 levels (Fig. 4C–E), and elevated DNA-damage marker γH2AX expression (Fig. 4F, G). Interestingly, H2O2 and S/N treatment induced milder hallmark expression as compared to the other toxins used (Fig. 4). Taken together, our data emphasize the importance of assaying multiple phenotypes when defining a senescent cell, considering not only the cell type under investigation but also the context in which senescence is being induced.
Following our central theme of cell-type-dependent expression of senescence phenotypes, our unbiased isolation of senescence profiles of human cell types through PCA emphasized our ability to isolate both unique and common senescence signatures from different cell types (Fig. 5). Therefore, a common and cell-type-specific senescence profile predicts upstream SATRs, which can functionally regulate observed senescence responses both within and across these different cell types (Fig. 6)49. Our investigations into SATRs such as ERF, CHCHD3, SOX12, PA2G4, and TFAP4 highlight that each regulator can recapitulate some hallmarks of senescence, but not all, and does so in a cell-type-dependent manner (Figs. 6 and S5). The observed age-dependent decline of the expression of these SATRs further points toward their regulatory role in senescence and aging (Fig. S8).
We chose to focus on the SATR TFAP4 based partially on its previous association with senescence in the context of directly downregulating p16 and p21 in fibroblasts43 and induction based on varied c-Myc levels50. Most interestingly, TFAP4 has been shown to suppress DNA damage and senescence via repression of MIR22HG (miR-22-3p)24 (Fig. 7C), which we have recently shown to be connected to PD risk factors SATB1 and GBA in the context of lipid-induced DA neuron senescence44,51. Targeted TFAP4 KD with siRNA in the five human cell lines revealed varying expression of senescence hallmarks in a cell-type-dependent manner (Fig. 7). Our experiments using BrdU treatment did not lead to the same pattern of hallmark expression in any of the phenotypes we assessed (Figs. 1 and S1–3) which suggests that a cell-type-dependent senescence phenotype can also be dependent on the method of senescence induction (consistent with our results in Fig. 4 showing toxin-dependent senescence phenotype expression) and may be regulated by the concerted action of a subset of SATRs. In addition to our investigations into specific senescence phenotypes that may be modulated by these upstream SATRs, it is also possible to manipulate these inferred regulators in the context of senolytic drug development to specifically target certain senescent cell types in vitro and in vivo. While the use of immortalized cell lines allowed us a highly controlled experimental approach to identify cell-type-specific SATRs, we acknowledge that they do not fully resemble differentiated in vivo cells. This limitation, however, was overcome by additional testing of SATR function in primary cells, stem-cell derived DA neurons, human midbrain slices, as well as a well-characterized mouse model of PD-associated senescence and neurodegeneration.
Further investigations into TFAP4 in vitro and in vivo highlighted its altered expression in various models of senescence and PD. Our well-established SATB1-KO DA neuron model of senescence showed a significant decrease in TFAP4 (Fig. S10B) and BrdU treatment in primary human astrocytes recapitulated this decrease (Fig. S10C). Interestingly, immunostaining of age-matched control and PD patient human midbrain slices demonstrated a loss of TFAP4 in both DA neurons and astrocytes in vivo (Fig. 7L). Lastly, we utilized a PD mouse model of PQ treatment and observed a significant decrease in Tfap4 expression levels in midbrain astrocytes and DA neurons (Fig. 9C, D). The mouse model also recapitulated additional senescence hallmarks (loss of Hmgb1 and Lmnb1) (Fig. 9A–D). We also observed alterations in glial morphology indicative of midbrain inflammation in response to PQ treatment (Fig. 8), which points towards the question of the relationship between the co-occurring midbrain senescence and inflammatory phenotypes in vivo. Ultimately, the robust findings of downregulated TFAP4 across various models of senescence, aging, and PD, unify the SATR TFAP4 as a pathological driver of neurodegenerative phenotypes.
Overall, our characterization of a cell-type-specific senescence profile in various human brain cell types defines senescence in the context of neurodegeneration and specifically PD, where DA neurons of the SNpc have been shown to become senescent12. Importantly, the cell types investigated have all been shown to be capable of becoming senescent in the context of aging and neurodegeneration13,19,48,52,53. Taken together, this leads to the interesting notion that these five different cell types may have the potential to become senescent and influence brain inflammation or contribute to neurodegenerative phenotypes via the SASP. Thus, it is crucial to have defined cell-type-specific profiles of senescence to better understand the mechanisms by which senescence may spread between distinct cell types both in vitro and in vivo. Based on our results, we propose that SATRs may function as upstream regulators of these senescence phenotypes in a cell-type-dependent manner. Specifically, TFAP4, whose expression was robustly shown to be significantly decreased in various models of PD and senescence, could potentially be used as a cell-type-specific drug target for either developing senolytics, senescence biomarkers, or senescence inducers in aging and neurodegeneration.
Methods
BrdU treatment in cell culture
Cells were cultured as described in Supplementary Methods. Treatment for 7 days with 100 µM 5-bromo-2′-deoxyuridine (BrdU) (Sigma, Catalog No. B5002) dissolved in DMSO (Sigma, Catalog No. D2650) was used to induce senescence. A 0.1% DMSO solution in the appropriate cell line growth media was used as a control. Previous reports have shown that BrdU treatment activates a DNA damage response and can induce a senescence phenotype in mammalian cells, regardless of cell type17,18,25,54,55,56. Viability curves for BrdU treatment were generated for each human cell line showing optimal treatment with 100 µM BrdU for 48 h (Fig. S1A using a CCK8 viability assay (described below in “CCK8 Viability Assay”).
Senescence-associated β-galactosidase (SA β-gal) assay
To test whether cells displayed a lysosomal dysfunction phenotype consistent with entering a state of cellular senescence, a senescence-associated β-galactosidase (SA β-gal) assay (Cell Signaling Technology, Catalog No. 9860) was performed according to the manufacturer’s protocol. After the respective 7-day treatment, cells were fixed and incubated over night at 37 °C with X-gal as substrate for the SA β-galactosidase. The following day, the substrate solution was removed, 70% glycerol (in PBS) was added to the fixed cells, and wells were imaged with a microscope (Accu-Scope, EXI-410, Skye Color Camera). Analysis and quantification of images was performed using Fiji and statistical analyses were performed using Prism (GraphPad).
Live imaging and growth curve analysis
Following 7-day treatment with DMSO or 100 µM BrdU, human cell lines were passaged and seeded on 24-well plates (Sarstedt, Catalog No. NC0984607) at equal densities amongst conditions. Using the Agilent LionheartFX Live Imaging System, three regions of interest per well were imaged at 20× every 15 min for 48 h. Cell growth was standardized by starting cell number.
Immunofluorescent staining in cell culture
12 mm round glass coverslips (VWR, Catalog No. 73605-514) were washed in 70% ethanol, rinsed with PBS, and allowed to air dry for 60 min under UV light to sterilize. Coverslips were then placed into each well of a 24-well plate, 500 µL of 0.1% gelatin in H2O was added to each well, and the plate was incubated at 37 °C for at least 30 min. The gelatin solution was then removed, and cells were plated onto coverslips. Following treatment, cells were chemically fixed with 4% PFA in PBS for 20 min at room temperature. Cells were then permeabilized with 0.2% TritonX-100 in PBS for 10 min at room temperature, followed by three 5-min PBS washes and then blocking in 5% NDS/PBS/0.1% Tween20 at room temperature for 30 min. The cells were then incubated with the respective primary antibody (1:500) in 5% NDS/PBS/0.1% Tween20 overnight at 4 °C. Primary antibodies used include γH2AX (pS139) (Abcam, Catalog No. ab303656, Lot 1099105-4, Clone N1-431), γH2AX (pS139) (Cell Signaling, Catalog No. 9718, Lot 14), lamin B1 (Abcam, Catalog No. ab16048, Lot 1085092-1), p21 (Abcam, Catalog No. ab109520, Lot 1092672-16), MAP2 (AVES, Catalog No. NBP2-25156SS, Lot MAP88837977), HMGB1 (Abcam, Catalog No. ab18256, Lot 1066643-16), TFAP4 (Invitrogen, Catalog No. PA5-116031, Lot 79445811), and GFAP (Millipore, Catalog No. MAB360, Lot 4266488, Clone GA5). The following day, cells were washed 4 times for 5 min each in 0.1% Tween20-PBS and then incubated with the respective secondary antibodies (1:1000) in 5% NDS/PBS/0.1% Tween20 for 2 h in the dark at room temperature. Secondary antibodies used include Alexa Fluor 488 d@ms IgG (Invitrogen, Catalog No. A21202, Lot 3006753), Alexa Fluor 488 d@Rb (Invitrogen, Catalog No. A21206, Lot 2668665), Alexa Fluor 546 d@Rb IgG (Invitrogen, Catalog No. A10040, Lot 2892414), Alexa Fluor 647 d@Rb (Invitrogen, Catalog No. A31573, Lot 2752586), Alexa Fluor 647 d@Chk (Invitrogen, Catalog No. A78952, Lot 2789900), Alexa Fluor 488 g@rb (Invitrogen, Catalog No. A11008, Lot 1812166), Alexa Fluor 594d@ms (Invitrogen, Catalog No. A21203, Lot 2474953), and Alexa Fluor 647g@chk (Invitrogen, Catalog No. A32933, Lot X1355385). The cells were then rinsed again 4 times for 5 min each in 0.1% Tween20-PBS and then coverslips with cells were mounted onto slides (top down) with 20 μL of DAPI Fluoromount-G (Southern Biotech, Catalog No. 0100-20). Slides were then cured in the dark at room temperature overnight. Confocal images were taken with an Olympus FV3000 Laser Scanning Confocal Microscope. Laser settings (laser strength, gain, and offset) and magnification were maintained across treatment groups. Airyscan imaging was performed using Zeiss LSM 800. Images were taken at 3× zoom with 0.2 AU pinhole size and high-resolution Airyscan post-processing. All other settings were optimized by Airyscan digital processing recommendations.
Fluorescent dyes for imaging mitochondria and lysosomes
For imaging of mitochondria and lysosomes, cells were plated on sterile 12 mm round glass coverslips (as described in “Immunofluorescent staining in cell culture”). Following treatment, cells were washed 1× with Dulbecco’s PBS without calcium and magnesium (Thermo Scientific, Catalog No. 14190094) and then incubated for 15 min at 37 °C in either 500 nM Mitotracker Red CMXRos (Invitrogen, Catalog No. M7512) in PBS or 1 µM Lysotracker Deep Red (Invitrogen, Catalog No. L12492) in PBS. Following treatment, cells were chemically fixed, permeabilized, washed, blocked, and stained as described in “Immunofluorescent staining in cell culture” for additional senescence markers. Confocal images were taken with an Olympus FV3000 Laser Scanning Confocal Microscope. Laser settings (laser strength, gain, and offset) and magnification were maintained across treatment groups. Post-processing of images was performed by ImageJ and Cell Profiler as described in “Image analyses”.
RNA isolation of cell culture and RNAseq of DMSO- and BrdU-treated human cell lines
RNA was isolated from human cell lines and primary human astrocytes using the RNeasy Plus Mini Kit (QIAGEN, Catalog No. 74134) and QIAshredder (QIAGEN, Catalog No. 79656). For DMSO and BrdU-treated human cell lines, 500 ng of RNA for each sample was sent to Azenta for bulk RNAseq. Sample quality control and determination of concentration was performed using TapeStation Analysis by Azenta, followed by library preparation and sequencing. Computational analysis included in their standard data analysis package was used for data interpretation. Target genes were pulled from known lists of genes associated with senescence, senescence-associated secretory phenotype (SASP)57, mitochondrial and lysosomal function58,59. Samples used in heatmaps were normalized by z-score.
Senolytic treatment in human cell lines
Human cell lines were plated on 96-well plates and treated for 7-days with DMSO or 100 µM BrdU as described in “BrdU treatment in cell culture” Viability of the cells was then measured (see “CCK8 viability assay”) before senolytic drugs were added. Dasatinib (VWR, Catalog No. 102541-694) was dissolved in sterile DMSO and was used in final concentrations of 50 nM or 100 nM in the appropriate culture media based on the cell type being tested. Quercetin (Med Chem Express, Catalog No. HY-18085) was dissolved in sterile DMSO and was used in final concentrations of 10 µM or 20 µM in the appropriate culture media based on the cell type being tested. Navitoclax (ABT-263, Med Chem Express, Catalog No. HY-10087) was dissolved in sterile DMSO and was used in final concentrations of 1, 10, or 20 µM in the appropriate culture media based on the cell type being tested. For all senolytic drugs used, an appropriate DMSO control (with the highest % DMSO used for each drug) was applied. All senolytics were applied for 48 h and cells were incubated at 37 °C with 5% CO2.
CCK8 viability assay
To assess cellular viability, a CCK8 cell viability kit (Dojindo, Catalog No. CK04-05) was used according to the manufacturer’s protocol. In brief, cell medium was replaced by fresh medium containing 10% of CCK8 solution in the appropriate serum-containing media based on the cell type being tested. Using a SpectraMax iD3 (Molecular Devices) plate reader, an initial background absorbance reading at 450 nm was taken. Cells were then incubated for 2 h at 37 °C with 5% CO2 before absorbance was read again. For senolytic treatment in human cell lines, the CCK8 assay for assessing viability was first used after the 7-day DMSO and BrdU treatment. Cells were then washed with PBS and senolytic drugs were added (as described above) and cells were incubated for 48 h at 37 °C with 5% CO2. The CCK8 assay was then performed again to assess viability based on treatment with senolytics and results were normalized to the initial CCK8 assay performed on Day 7 following DMSO or 100 µM BrdU treatment.
Generation of primary mouse cortical co-cultures
Brains from WT C57/B6J mice were quickly dissected from P0/P1 pups in Hibernate-A/B27(0.5 mM GlutaMAX (Thermo Scientific, Catalog No. 35050061), 1% Pen/Strep, 1% B27 (Thermo Scientific, Catalog No. 17504044)). Cortices were minced and digested in Papain Digestion Medium (100 units Papain in 2 mL Hibernate-A (Thermo Scientific, Catalog No. A1247501)) for 30 min at 30 °C. An equal volume of Hibernate A/B27 was added, and the tissue was triturated 10 times with a siliconized 9-inch Pasteur pipette with a tip fire polished to an opening of 0.7–0.9 µm diameter. Supernatant was then transferred to a new tube, and the cells were pelleted by centrifugation at 80 × g for 5 min. Cells were counted by hemocytometer, and the remaining cell pellet was resuspended in an appropriate volume of Neurobasal-A/B27 (Thermo Scientific, Catalog No. 10888022) (0.5 mM GlutaMAX, 1% Pen/Strep, 1% B27) and the cells were plated at 80% of plating volume. Neurobasal media was changed after 45 min of plating and half media changes were made every two days.
UV treatment of astrocyte cell line
UV irradiation-induced senescence was performed by exposing the human astrocyte cell line to UV treatment (Sankyo Denki germicidal UV-C lamp, G30T8). Cells were exposed to 5.34 mJ/cm2 of UV-C light the day after plating, with a media change 24 h later and a second exposure to 5.34 mJ/cm2 of UV-C light 48 h after the initial exposure. Cells were then maintained in culture for a total of 7 days before markers of senescence were analyzed. Control cells were not exposed to UV treatment and were cultured for 7 days.
PQ treatment of astrocyte cell line
Treatment for 7 days with 100 µM paraquat (PQ, methyl viologen hydrate 98%) (Thermo Scientific, Catalog No. 227320050) dissolved in UltraPure Distilled Water (Invitrogen, Catalog No. 10977-015) and filtered (0.22 µm PES, Sigma, Catalog No. SCGP00525) for sterility was used to induce senescence in the human astrocyte cell line. A 0.1% water solution in the appropriate cell line growth media was used as a control.
H2O2 treatment of astrocyte cell line
Treatment for 7 days with 20 µM hydrogen peroxide (H2O2) (Fisher Chemicals, Catalog No. H325-500), diluted in UltraPure Distilled Water (Invitrogen, Catalog No. 10977-015) and filtered (0.22 µm PES, Sigma, Catalog No. SCGP00525) for sterility was used to induce senescence in the human astrocyte cell line.
Collection of supernatant (S/N) and treatment in astrocyte cell line
To generate conditioned media (supernatant, S/N) from DMSO and BrdU-treated astrocytes, we treated the human cell line for 7 days with DMSO and BrdU (as described in “BrdU treatment in cell culture”). Cells were then given fresh media and were cultured for 24 h to condition the media without the treatment being present. This S/N was then collected, filtered, snap frozen, and stored at −80 °C. For treatments, cells were given a 2:1 treatment of S/N:normal media with a DMSO S/N and BrdU S/N group. Treatment was incubated on the cells for 7-days prior to assessment of senescence phenotypes.
Principal component analysis
Principal component analysis (PCA) was performed using the FactoMineR package in R60. RNA-seq expression data was normalized by z-score for each gene list. A correlation matrix was formed for each sample condition using the built in cor() function from the normalized data set. The function princomp() was used to isolate the principal component analysis loadings from the correlation matrix. fviz_eig, fviz_pca_var, and fviz_cos2 were used to visualize the data. The graphs were constructed in GraphPad Prism.
Transcriptional regulator analysis
The transcriptional regulatory networks were inferred using the Algorithm for the Reconstruction of Accurate Cellular Networks with Adaptive Partitioning (ARACNe-AP) using IntelliJ repository33 and Bioconductor RTN package in R23,61. A list of known human transcription factors was obtained62. In short, the reconstruction of a transcriptional regulatory network (TRN) was inferred from RNA-seq data collected from all 5 cell types (astrocytes, endothelial cells, microglia, oligodendrocytes, and neurons) and from both treatment conditions (7-day DMSO or 100 M BrdU). Interactions between regulators (TFs) and potential targets were computed based on mutual information (MI), removing unstable interactions by bootstrapping (n = 1000), and then a DPI-filter was applied to remove the weakest interaction between two TFs and a common target gene. From this TRN, we used a two-tailed gene set enrichment analysis with 1000 permutations to assess the regulon-target gene interaction, separating them into a positive or negative targets using Pearson’s correlation and assessing the distribution of the targets across a log2FoldChange ranked gene list. An Enrichment Score (ES) was calculated by walking down the ranked list of target genes and increasing the running sum statistic when encountering a gene in the set and decreasing the statistic when encountering a gene not in the set. The differential Enrichment score (dES = posES-negES) was normalized to the mean enrichment score of random samples of the same gene set size (NES). P-values were calculated using log-rank statistics. The top 10 positively and negatively enriched regulons were isolated for each cell type and then further sorted by baseline expression.
RT-qPCR
RNA (200 ng) was reverse transcribed using the Applied Biosystems High-Capacity cDNA Reverse Transcription Kit (Thermo Scientific, Catalog No. 4368814) and the output volume of 20 μL was diluted in nuclease-free water to 40 μL for a cDNA working concentration of 5 ng/μL. Real-time PCR was performed using the Applied Biosystems Power SYBR Green PCR Master Mix (Thermo Scientific, Catalog No. 4367659) on a Quant Studio 3 (Applied Biosystems) with reaction specificity confirmed by melt curve analysis. All comparisons (ex. siSCR vs. siRegulator) for each qPCR reaction were run on the same qPCR plate and run in a triplicate. Human qPCR primer sequences used (5′ to 3′) are listed in Supplementary Table 1.
Staining human brain slices
Healthy and Parkinson’s disease human substantia nigra tissue was obtained from the Harvard Brain Tissue Resource Center and processed by the Banner Sun Health Research Institute (Harvard Brain Tissue Resource Center, PHS Grant R24 MH068855). For immunostaining of the human brain tissue sections, slides were removed from storage at −80 °C and allowed to dry at room temperature for 20 min. Once dry, a hydrophobic barrier was made using a Dako pen. Slides were washed twice in PBS for 10 min each while shaking at 50 rpm and then permeabilized in 0.2% Triton X-100 in PBS for 15 min. Sections were blocked for 1 h at room temperature in 2% NDS/0.1% fish gelatin (Sigma, Catalog No. 7765) in 0.1% Triton X-100 in PBS and then incubated overnight with the respective primary antibody (1:250) at 4 °C in the dark in 2% NDS/0.1% fish gelatin in 0.1% Triton X-100 in PBS. Primary antibodies used include TH (Millipore, Catalog No. ab9702, Lot 4227767), TFAP4 (Invitrogen, Catalog No. PA5-116031, Lot 79445811), and GFAP (Millipore, Catalog No. MAB360, Lot 4266488, Clone GA5). The following day, slides were washed with PBS three times for 10 min each while shaking at 50 rpm and then incubated in respective secondary antibody (1:400) at room temperature in the dark for 2 h in 2% NDS/0.1% fish gelatin in 0.1% Triton X-100 in PBS. Secondary antibodies used include Alexa Fluor 488 d@ms IgG (Invitrogen, Catalog No. A21202, Lot 3006753), Alexa Fluor 546 d@Rb IgG (Invitrogen, Catalog No. A10040, Lot 2892414), and Alexa Fluor 647 d@Chk (Invitrogen, Catalog No. A78952, Lot 2789900). Following incubation with secondary antibody, tissue sections were washed once for 5 min with PBS at room temperature and once for 5 min with 70% ethanol at room temperature. Sections were then immersed with autofluorescence eliminator reagent (Sigma, Catalog No. 2160) for 5 min to quench autofluorescence. Sections were then washed 3 times for 1 min each with 70% ethanol and then once for 5 min with PBS before mounting with DAPI Fluoromount-G (Southern Biotech, Catalog No. 0100-20). Slides were then cured in the dark at room temperature overnight. Confocal images were taken with an Olympus FV3000 Laser Scanning Confocal Microscope. Laser settings (laser strength, gain, and offset) and magnification were maintained across treatment groups.
Mice
All mouse procedures were reviewed and approved by the Institutional Animal Care and Use Committee at Stony Brook University and were carried out according to the NIH Guide for Care and Use of Laboratory Animals. Mice were housed 2–3 per cage in an environmentally controlled room (20–23 °C, 30–70% humidity, 12-h light/dark cycles) with food and water available ad libitum and cages were cleaned once per week. 6-month-old male and female C57BL/6J (Jackson laboratory) mice were injected intraperitoneally (i.p.) with saline (n = 1 male, n = 3 females) or 7 mg/kg PQ in saline (n = 1 male, n = 3 females) at 2-day intervals for a total of 6 doses19. No sex differences were observed. Animals were sacrificed 14 days after the last dose of PQ to observe long-term effects of PQ treatment.
Perfusion and cryosectioning of mouse brains
Mice were euthanized by carbon dioxide inhalation and immediately exsanguinated by cardiac puncture using a 1 mL syringe with a 25-gauge needle. The tissue was chemically fixed using a 2-min flush of PBS 1× and a 5-min perfusion of 4% PFA. Following the perfusion, the skull cap was removed, and the brain was gently teased from the cranial cavity in a rostro-caudal direction. Micro-dissection forceps were used to gently tease apart the cranial nerves. The brain was then rinsed in PBS 1× and placed in a 15 ml conical tube with 4% PFA at 4 °C with agitation overnight. The following day, the brain was transferred to 30% sucrose (w/v) for an additional 24 h. The brain was then placed in optimal cutting temperature (OCT) and stored at −80 °C until cryosectioning. Brains in OCT were placed in −20 °C for one hour prior to cryosectioning to allow for the temperature to equilibrate. Brains were then sectioned coronally in the cranial to caudal direction at 30 μm slices and placed free-floating in a 12 well microplate filled with PBS 1×. Sections were allowed to equilibrate in PBS 1× for 1 h. Sections were then rinsed with PBS 1× to remove residual OCT and were then redistributed to a new 12 well plate in PBS for immunoblotting.
Immunofluorescent staining of mouse midbrain tissue
Immunofluorescent staining was performed as described in “Immunofluorescent staining in cell culture” with slight adjustments based on previously reported protocols19. The brain slices were rinsed three times in PBS to remove residual OCT and then were permeabilized in 0.2% TritonX-100 in PBS for 1 h at room temperature. Sections were then blocked in 5% NDS/PBS/0.1% TritonX-100 for 1 h at room temperature and incubated with the respective primary antibody (1:500) in 5% NDS/PBS/0.2% TritonX-100 overnight at 4 °C. Primary antibodies used include TH (Millipore, Catalog No. ab9702, Lot 4227767), TFAP4 (Invitrogen, Catalog No. PA5-116031, Lot 79445811), GFAP (Millipore, Catalog No. MAB360, Lot 4266488, Clone GA5), Iba1 (Wako, Catalog No. 019-19741, Lot CAQ2805), NeuN (Abcam, Catalog No. ab177487, Lot GR249899-39), lamin B1 (Abcam, Catalog No. ab16048, Lot 1085092-1), and HMGB1 (Abcam, Catalog No. ab18256, Lot 1066643-16). The following day, brain slices were washed six times for 5 min each in 0.1% Tween 20-PBS and then incubated with the respective secondary antibodies (1:1000) in 5% NDS/PBS/0.2% TritonX-100 for 2 h in the dark at room temperature. Secondary antibodies used include Alexa Fluor 488 d@ms IgG (Invitrogen, Catalog No. A21202, Lot 3006753), Alexa Fluor 546 d@Rb IgG (Invitrogen, Catalog No. A10040, Lot 2892414), and Alexa Fluor 647 d@Chk (Invitrogen, Catalog No. A78952, Lot 2789900). The brain slices were then rinsed again six times for 5 min each in 0.1% Tween20-PBS before being mounted onto Superfrost Plus microscope slides (VWR, Catalog No. CA48311-703). When dry, slides were rinsed once in PBS and then incubated with 1× True Black (Biotium, Catalog No. 23007) in 70% ethanol for 30 s. Slides were then rinsed twice quickly in 70% ethanol and then washed three times for 1 min each in PBS before coverslips were added with DAPI Fluoromount-G (Southern Biotech, Catalog No. 0100-20). Slides were then cured in the dark at room temperature overnight. Confocal images were taken with an Olympus FV3000 Laser Scanning Confocal Microscope and OlympusVS-120 Slide Scanner (40×). Laser settings (laser strength, gain, and offset) and magnification were maintained across treatment groups.
Astrocyte and microglial morphological analysis
30 µm murine cortical slices were immunostained for astrocyte (GFAP), microglia (IBA1), and dopamine neuron (TH) labeling as described in “Immunofluorescent staining of mouse midbrain tissue”. TH staining was utilized to identify the Substantia Nigra pars compacta (SNpc) at ~3.08 mm Bregma. Imaging of astrocytes and microglia was performed using the Fluoview FV3000 (Olympus). Z-stacks were collected with 0.4 µm for the thickness of the tissue (1024 × 1024 pixel resolution). Before analysis, images were converted from Olympus (OIR) to Imaris (IMS) file types using the Imaris Image Converter software. Images were then rendered in 3D, and the morphological tracing was performed utilizing the Imaris FilamentTracer Autopath (no loops) pipeline. For microglia, starting points (soma) were estimated and analyzed utilizing the automated soma calculator. For astrocytes, starting points (soma) were predicted and estimated manually. Seed points were predicted and trained prior to automated detection. Filament segments were detected and manually pruned for training prior to automated detection. The number of branches, intersections, end point terminals, average branch length, were calculated based on filament traces.
Image analyses
All Olympus FV3000 images were collected at 60× magnification, 1024 × 1024 resolution, and with optimal Z-stacks. Images were analyzed in bulk through Cell profiler. Z-projections were taken from each image by maximum intensity and then separated by fluorophore. Nuclei were identified using DAPI staining and cell types were identified through specific cell markers. Mitochondria and lysosomal counts were analyzed by applying a size and intensity threshold in Cell Profiler, utilizing control samples. Counts per image were then standardized by the number of cells to give an average number of objects per cell. γH2AX foci were analyzed by applying a size and intensity threshold in Cell Profiler, utilizing control samples. These foci were then associated with each nucleus. Foci per nuclei were then averaged by the number of cells per image. p21, lamin B1, Tfap4, and Hmgb1 intensity were extracted from DAPI labeled nuclei.
Statistics
Data was analyzed using Fiji (v2.9.0), R (v4.4.3), Cell Profiler (v4.2.8), and GraphPad Prism (v10.3.1). Identification of senescence-associated transcriptional regulators was performed utilizing previously established RTN (v2.31.0) package in R. Statistical tests were performed using GraphPad’s Prism software. A threshold of p < 0.05 was considered significant. Significance was determined using the test indicated in figure legends. Results are presented as means ± SD, with individual points representing biological replicates when n < 30. All sample sizes (n) are listed as either replicates, indicating individual cell culture wells (apart from Fig. 8, which refers to brain tissue from individual mice), or the total number of cells. For images in Fig. 7L, one control and one PD patient were analyzed for TFAP4 expression. Because of an n = 1, no statistics were performed. All measurements were taken from distinct samples.
Figures S1–S3 are associated with Fig. 1. Representative SA β-gal images in Fig. S1B are associated with Fig. 1B. Representative growth curves in Fig. S1D are associated with Fig. 1C. Representative images in Fig. S2A are associated with Fig. 1E. Representative images in Fig. S3C are associated with Fig. 1E. Figure S4 is associated with Fig. 4. Representative SA β-gal images in Fig. S4A are associated with Fig. 4B. Representative images in Fig. S4B–E are associated with Fig. 4D–G. Figs. S5–S8 are associated with Fig. 6. Representative western blots of TFAP4 knockdowns in each cell type in Fig. S6 and normalized TFAP4 levels based on qPCRs in each cell type in Fig. S7 are associated with Fig. 7D and E, respectively. Figs. S9 and S10 are associated with Fig. 7. Representative images in Fig. S9 are associated with Fig. 7G–K. Figure S11 is associated with Fig. 8.
Ethics statement
All mouse procedures were reviewed and approved by the Institutional Animal Care and Use Committee at Stony Brook University (IACUC2021-00101, IACUC1556599) and were carried out according to the NIH Guide for Care and Use of Laboratory Animals.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data generated in this study for Figs. 1–9 and S1–11 are provided in the Supplementary Information/Source Data file. All exact p-values for applicable panels of Figs. 1–9 and S1–11 are included in the Source Data file. The RNA sequencing data generated in this study have been deposited in the Array Express database under accession code E-MTAB-15005. Source data are provided with this paper.
Code availability
All code used in the manuscript is described in the “Methods” section under “Transcriptional regulator analysis”, is available in the Source Data within the paper, was previously published23,33,61, and is available on GitHub (https://github.com/califano-lab/ARACNe-AP).
References
Dauer, W. & Przedborski, S. Parkinson’s disease: mechanisms and models. Neuron 39, 889–909 (2003).
Russo, T. & Riessland, M. Age-related midbrain inflammation and senescence in Parkinson’s disease. Front. Aging Neurosci. 14, 917797 (2022).
Hong, B., Ohtake, Y., Itokazu, T. & Yamashita, T. Glial senescence enhances α-synuclein pathology owing to its insufficient clearance caused by autophagy dysfunction. Cell Death Discov. 10, 50 (2024).
Gorgoulis, V. et al. Cellular senescence: defining a path forward. Cell 179, 813–827 (2019).
Hudson, H. R., Riessland, M. & Orr, M. E. Defining and characterizing neuronal senescence, ‘neurescence’, as G(X) arrested cells. Trends Neurosci. 47, 971–984 (2024).
Hayflick, L. & Moorhead, P. S. The serial cultivation of human diploid cell strains. Exp. Cell Res. 25, 585–621 (1961).
Stein, G. H., Drullinger, L. F., Soulard, A. & Dulić, V. Differential roles for cyclin-dependent kinase inhibitors p21 and p16 in the mechanisms of senescence and differentiation in human fibroblasts. Mol. Cell Biol. 19, 2109–2117 (1999).
Ogrodnik, M. Cellular aging beyond cellular senescence: markers of senescence prior to cell cycle arrest in vitro and in vivo. Aging Cell 20, e13338 (2021).
Ogrodnik, M. et al. Guidelines for minimal information on cellular senescence experimentation in vivo. Cell 187, 4150–4175 (2024).
Chaib, S., Tchkonia, T. & Kirkland, J. L. Cellular senescence and senolytics: the path to the clinic. Nat. Med. 28, 1556–1568 (2022).
Prata, L., Ovsyannikova, I. G., Tchkonia, T. & Kirkland, J. L. Senescent cell clearance by the immune system: emerging therapeutic opportunities. Semin. Immunol. 40, 101275 (2018).
Riessland, M. et al. Loss of SATB1 induces p21-dependent cellular senescence in post-mitotic dopaminergic neurons. Cell Stem Cell 25, 514–530 e518 (2019).
Musi, N. et al. Tau protein aggregation is associated with cellular senescence in the brain. Aging Cell 17, e12840 (2018).
Riessland, M., Ximerakis, M., Jarjour, A. A., Zhang, B. & Orr, M. E. Therapeutic targeting of senescent cells in the CNS. Nat. Rev. Drug Discov. 23, 817–837 (2024).
Martirosyan, A. et al. Unravelling cell type-specific responses to Parkinson’s disease at single cell resolution. Mol. Neurodegener. 19, 7 (2024).
Huang, W., Hickson, L. J., Eirin, A., Kirkland, J. L. & Lerman, L. O. Cellular senescence: the good, the bad and the unknown. Nat. Rev. Nephrol. 18, 611–627 (2022).
Michishita, E. et al. 5-Bromodeoxyuridine induces senescence-like phenomena in mammalian cells regardless of cell type or species. J. Biochem. 126, 1052–1059 (1999).
Suzuki, T. et al. Induction of senescence-associated genes by 5-bromodeoxyuridine in HeLa cells. Exp. Gerontol. 36, 465–474 (2001).
Chinta, S. J. et al. Cellular senescence is induced by the environmental neurotoxin paraquat and contributes to neuropathology linked to Parkinson’s disease. Cell Rep. 22, 930–940 (2018).
Hernandez-Segura, A., Brandenburg, S. & Demaria, M. Induction and validation of cellular senescence in primary human cells. J. Vis. Exp. 136, 57782 (2018).
Grandy, C. et al. Remodeling of the focal adhesion complex by hydrogen-peroxide-induced senescence. Sci. Rep. 13, 9735 (2023).
Hohn, A. et al. Happily (n)ever after: aging in the context of oxidative stress, proteostasis loss and cellular senescence. Redox Biol. 11, 482–501 (2017).
Fletcher, M. N. C. et al. Master regulators of FGFR2 signalling and breast cancer risk. Nat. Commun. 4, 2464 (2013).
Chou, J., Kaller, M., Jaeckel, S., Rokavec, M. & Hermeking, H. AP4 suppresses DNA damage, chromosomal instability and senescence via inducing MDC1/Mediator of DNA damage Checkpoint 1 and repressing MIR22HG/miR-22-3p. Mol. Cancer 21, 120 (2022).
Ozsvari, B., Nuttall, J. R., Sotgia, F. & Lisanti, M. P. Azithromycin and Roxithromycin define a new family of “senolytic” drugs that target senescent human fibroblasts. Aging 10, 3294–3307 (2018).
Hudson, H. R., Orr, T. C. & Orr, M. E. Investigating neuronal senescence marker, p19INK4d, in mouse and human brains. Alzheimer’s Dement. 19, e082962 (2023).
Rovira, M. et al. The lysosomal proteome of senescent cells contributes to the senescence secretome. Aging Cell 21, e13707 (2022).
Zhu, Y. et al. The Achilles’ heel of senescent cells: from transcriptome to senolytic drugs. Aging Cell 14, 644–658 (2015).
Zhu, Y. et al. Identification of a novel senolytic agent, navitoclax, targeting the Bcl-2 family of anti-apoptotic factors. Aging Cell 15, 428–435 (2016).
Sanborn, M. A., Wang, X., Gao, S., Dai, Y. & Rehman, J. Unveiling the cell-type-specific landscape of cellular senescence through single-cell transcriptomics using SenePy. Nat. Commun. 16, 1884 (2025).
Schneckener, S., Arden, N. S. & Schuppert, A. Quantifying stability in gene list ranking across microarray derived clinical biomarkers. BMC Med. Genomics 4, 1–11 (2011).
Butte, A. The use and analysis of microarray data. Nat. Rev. Drug Discov. 1, 951–960 (2002).
Lachmann, A., Giorgi, F. M., Lopez, G. & Califano, A. ARACNe-AP: gene network reverse engineering through adaptive partitioning inference of mutual information. Bioinformatics 32, 2233–2235 (2016).
Chen, E. Y. et al. Enrichr: interactive and collaborative HTML5 gene list enrichment analysis tool. BMC Bioinforma. 14, 128 (2013).
Kuleshov, M. V. et al. Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res. 44, W90–W97 (2016).
Xie, Z. et al. Gene set knowledge discovery with enrichr. Curr. Protoc. 1, e90 (2021).
The GTEx Consortium. The GTEx Consortium atlas of genetic regulatory effects across human tissues. Science 369, 1318–1330 (2020).
Darshi, M. et al. ChChd3, an inner mitochondrial membrane protein, is essential for maintaining crista integrity and mitochondrial function. J. Biol. Chem. 286, 2918–2932 (2011).
Dabiri, A., Sharifi, M. & Sarmadi, A. Knockdown of SOX12 expression induced apoptotic factors is associated with TWIST1 and CTNNB1 expression in human acute myeloid leukemia cells. Int. J. Mol. Cell Med. 10, 249–258 (2021).
Bao, Y. et al. Ebp1 p48 promotes oncogenic properties in hepatocellular carcinoma through p38 MAPK/HIF1α activation and p53 downregulation. Mol. Carcinog. 60, 252–264 (2021).
Ohtani, N. et al. Opposing effects of Ets and Id proteins on p16INK4a expression during cellular senescence. Nature 409, 1067–1070 (2001).
Chung, K. et al. The ERF transcription factor EPI1 is a negative regulator of dark-induced and jasmonate-stimulated senescence in Arabidopsis. Plant Biotechnol. (Tokyo) 33, 235–243 (2016).
Jackstadt, R., Jung, P. & Hermeking, H. AP4 directly downregulates p16 and p21 to suppress senescence and mediate transformation. Cell Death Dis. 4, e775 (2013).
Russo, T. et al. The SATB1-MIR22-GBA axis mediates glucocerebroside accumulation inducing a cellular senescence-like phenotype in dopaminergic neurons. Aging Cell 23, e14077 (2024).
Popov, A. et al. Mitochondrial malfunction and atrophy of astrocytes in the aged human cerebral cortex. Nat. Commun. 14, 8380 (2023).
Vidal-Itriago, A. et al. Microglia morphophysiological diversity and its implications for the CNS. Front. Immunol. 13, 2022 (2022).
Streit, W. J., Graeber, M. B. & Kreutzberg, G. W. Functional plasticity of microglia: a review. Glia 1, 301–307 (1988).
Brelstaff, J. H. et al. Microglia become hypofunctional and release metalloproteases and tau seeds when phagocytosing live neurons with P301S tau aggregates. Sci. Adv. 7, eabg4980 (2021).
Bitto, A. et al. Stress-induced senescence in human and rodent astrocytes. Exp. Cell Res. 316, 2961–2968 (2010).
Wang, Y. et al. Differential cellular responses by oncogenic levels of c-Myc expression in long-term confluent retinal pigment epithelial cells. Mol. Cell Biochem 443, 193–204 (2018).
Russo, T. & Riessland, M. Lipid accumulation drives cellular senescence in dopaminergic neurons. Aging (Albany NY) 16, 11128–11133 (2024).
Yamazaki, Y. et al. Vascular cell senescence contributes to blood–brain barrier breakdown. Stroke 47, 1068–1077 (2016).
Al-Mashhadi, S. et al. Oxidative glial cell damage associated with white matter lesions in the aging human brain. Brain Pathol. 25, 565–574 (2015).
Jochems, F. et al. The cancer SENESCopedia: a delineation of cancer cell senescence. Cell Rep. 36, 109441 (2021).
Masterson, J. C. & O’Dea, S. 5-Bromo-2-deoxyuridine activates DNA damage signalling responses and induces a senescence-like phenotype in p16-null lung cancer cells. Anticancer Drugs 18, 1053–1068 (2007).
Ross, H. H. et al. Bromodeoxyuridine induces senescence in neural stem and progenitor cells. Stem Cells 26, 3218–3227 (2008).
Saul, D. et al. A new gene set identifies senescent cells and predicts senescence-associated pathways across tissues. Nat. Commun. 13, 4827 (2022).
Calvo, S. E., Clauser, K. R. & Mootha, V. K. MitoCarta2.0: an updated inventory of mammalian mitochondrial proteins. Nucleic Acids Res. 44, D1251–D1257 (2016).
Smith, C. L. & Eppig, J. T. The mammalian phenotype ontology: enabling robust annotation and comparative analysis. Wiley Interdiscip. Rev. Syst. Biol. Med. 1, 390–399 (2009).
Lê, S., Josse, J. & Husson, F. FactoMineR: an R package for multivariate analysis. J. Stat. Softw. 25, 1–18 (2008).
Castro, M. A. A. et al. Regulators of genetic risk of breast cancer identified by integrative network analysis. Nat. Genet. 48, 12–21 (2016).
Lambert, S. A. et al. The human transcription factors. Cell 172, 650–665 (2018).
Acknowledgements
This work was supported in part through grants 1R01NS124735 (MR), R01AG079898 (RBS), the Hartman Center (MR), the Center for Healthy Aging (MR), startup funds from Stony Brook University (RBS), the Water Wheel Foundation (RBS), and the National Institute of General Medical Sciences of the National Institutes of Health K12GM102778 (JPB). Mitochondrial respiration measurements were performed using the Agilent Seahorse XFe96 analyzer with generous support from the Stony Brook University Biological Mass Spectrometry/Metabolomics core. Opinions, interpretations, conclusions, and recommendations are those of the author and are not necessarily endorsed by the sponsors.
Author information
Authors and Affiliations
Contributions
Conceptualization: T.R., M.R.; Methodology: T.R., M.R., J.P.B.; Investigation: T.R., M.R., J.P.B.; Visualization: T.R., J.P.B.; Funding acquisition: M.R., R.S.; Project administration: M.R., R.S.; Supervision: M.R., R.S.; Writing–original draft: T.R., J.P.B.; Writing–review & editing: T.R., M.R., J.P.B., R.S.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Jalees Rehman, who co-reviewed with Mark Sanborn; and the other, anonymous, reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Russo, T., Plessis-Belair, J., Sher, R. et al. Systematic profiling reveals distinct senescence signatures and regulators across human brain cell types. Nat Commun 16, 11059 (2025). https://doi.org/10.1038/s41467-025-66012-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-66012-3