Abstract
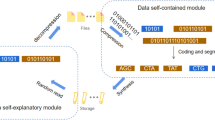
DNA-based storage is an alternative solution for archiving vast cold data. Its future development aims to meet the demands of hot data storage, which requires rapid random access and efficient data modification. Here, we present a DNA origami nanostructure-enabled linked data storage (DONLDS) system that implements a linked list architecture. This system uses distinct DNA origami shapes as nodes to store diverse data (English letters, numerals, Chinese characters) in binary locations, achieving a storage density of 222.22 Gbit/cm2. Pointers, defined by DNA strands at the nanostructure edges, establish data positions. Furthermore, detachable DNA strands serve as instructions, enabling dynamic linking of pointers for accurate storage and their reversible detachment for dynamic data retrieval. The DONLDS system eliminates the need for full-structure traversal, enables parallel data storage, and supports data insertion and removal. This highlights its adaptability and accuracy in managing complex datasets.
Similar content being viewed by others
Introduction
DNA, serving as the fundamental material for carrying and transmitting information in organisms, exhibits information storage capacity and stability1,2,3,4,5,6,7. DNA storage technology typically employs synthetically engineered DNA sequences to store data, necessitating data to be written and read in a predetermined sequence8,9,10. With the development of advanced DNA synthesis and sequencing techniques, DNA storage is encompassing terabytes of text, audio, and image information which is characterized by high storage density, minimal electromagnetic interference, and reduced risk of data loss. This characteristic renders current DNA storage technologies particularly adept at preserving cold data (data accessed infrequently, prioritizing high-density, long-term archival)11,12,13,14. However, the future trajectory of DNA storage is to preserve hot data (data requiring frequent, rapid access and modification), encompassing rapid random access and effective data modification functionalities that current DNA synthesis-based sequential storage cannot perform15,16,17,18,19,20. Hence, researchers have delved into myriad innovative methods to enable data modification without synthesis. For instance, leveraging single-stranded DNA overhang sequences that bind primers at room temperature allows users to rename, delete, or lock files21. Additionally, some methods utilize natural DNA strands as information carriers, such as DNA punch cards22 or nanopore sensor-based DNA hard drives23,24. Despite offering alternative pathways, these methods often necessitate the redistribution and reorganization of entire storage units, resulting in significant data migration and processing overhead.
Linked storage, different to sequential storage in computer memory storage paradigms, effectively manages frequent data insertions and deletions by adjusting the pointers of storage nodes, thereby avoiding extensive data migration, thereby substantially reducing the complexity of data processing. To achieve DNA-based linked storage, nodes for storage and pointers for data migration are needed. Previous studies have proved that DNA origami nanostructures25, fabricated by a long scaffold DNA with numerous short staple DNA strands, not only inherit the encoding capacity26,27 for storage but have shape identifiability28,29,30,31, unparalleled addressability32,33,34,35 and structural controllability36,37,38. For instance, the DNA PAINT technology29 was employed to read information via spot arrays and reconfigurable DNA origami structures39,40 to achieve data changes at the molecular level. As an information storage medium, DNA origami holds significant potential to meet the demands of large-scale data storage and processing, poised to play a crucial role in the future of information science. Nevertheless, information storage based on DNA nanostructures, involving the specific identification and efficient reversible recombination of storage units for data modification and deletion, has yet to be explored.
Here, we have developed a DNA origami nanostructure-enabled linked data storage (DONLDS) system to create nodes, pointers, and operations for data traversal, insertion, and deletion. These nodes are fabricated by triangular, rectangular, and cross-shaped DNA origami nanostructures (DONs) with binary locations represented for English letters, Arabic numerals, and Chinese characters, the storage density of which reach 222.22 Gbit/cm2. Pointers are featured by a series of DNA strands at the edges of DONs for the definition of data position. The precise manipulation of data can be achieved through a series of detachable DNA strands, which are not only capable of performing connection operations to tightly integrate the nodes, thereby guaranteeing the high fidelity of data storage, but also adept at executing disconnection operations to sever the inter-node bonds, thereby facilitating the dynamic storage and retrieval of data. The durability and reusability of these data connections have been validated through repeated detachment and reattachment tests. Furthermore, we have demonstrated the ability of a linked list based on DONs to store the string ‘DNA helix 1953’ and edit it with data deletions and insertions for its transformation to ‘DNA storage 1988’. The flexible, efficient, and scalable DONLDS eliminates the traversal of the entire data structure, facilitates the parallel storage of diverse data types and supports the seamless insertion and removal of data, highlighting its adaptability and accuracy in managing complex datasets.
Results
Creation of the linked list in the DONLDS system
The linked storage employs nodes as its fundamental units, interconnected through pointers to forge an adaptable data chain. Each node encompasses not only data but also pointers directed towards adjacent nodes, thereby ensuring data continuity. Benefitting from the programmability and the self-assembly properties of DNA molecules.
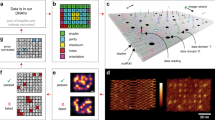
In DONLDS system, we utilized triangular, rectangular, and cross-shaped DNA origami nanostructure (TDON, RDON, CDON) as nodes, each of which had two sets of pointers. Typically, DONs with defined shape were fabricated by a long scaffold and hundreds of staple strands (Fig. 1a and Supplementary Fig. 1). To establish precise and orderly connections between nodes, we designed a series of strands with specific sequences as pointers, which were situated at the edges of DON nodes and adorned with grouping sequence coding (Supplementary Table 5). Furthermore, we introduced DNA strands as instructions to exercise exact control over the sequence for node connections. The sequences of the operation strands were meticulously crafted to be the reverse complement of the target pointer strands, ensuring a 16-nucleotide complementary base pairing between each pair of pointer and operation strands, thereby guaranteeing the specificity and stability of node connections (Fig. 1b). We assessed orthogonality among the 120 designed 16-nt pointer strands (Supplementary Table 5) by computing all pairwise Hamming distances. Visualization of the similarity matrix (Supplementary Fig. 3) showed that more than 99.27% of off-diagonal pairs scored ≤8 (<50% similarity), demonstrating low inter-sequence similarity and cross-reactivity risk across the design. Unlike traditional memory allocation, DONLDS system did not require contiguous memory nodes for data storage but dynamically allocates space at various locations in memory based on the size and type of data (Supplementary Fig. 2). This dynamic allocation and non-continuous storage approach allowed the system to flexibly accommodate fluctuations in data volume, obviating the need to reserve space for fixed-size data blocks. For example, to store 1Kbit of data, we divide the data into 10 layers, and each layer requires 100–144 types of pointer strands and operation strands (Supplementary Table 1). Atomic force microscopy (AFM) images illustrated how nodes of varying shapes achieve orderly connections under the guidance of operation strands (Fig. 1c and Supplementary Figs. 4–6). Moreover, the flexibility of memory allocation extended beyond the node linkage from the same type of DON nodes, enabling allocation across different node types. The multiple-node linkage was extended to interconnect diverse types of nodes through serial operations, enabling concurrent processing. This approach could provide the possibility of storing multiple types of information (Fig. 1d and Supplementary Fig. 7).
a In the design of DON nodes, each structure is divided into two main components: the area within the green box on the left represented the prior pointer, while the area within the red box on the right served as the next pointer. b The design scheme of pointer strands in DON nodes, as well as the synthesis process of the linked structure. c A Schematic diagram of memory unit connections based on three shapes of DON. The tetramer linked DONs are depicted in AFM images, shown images are from one experiment representative of 3 independent replicates. d The DONLDS achieves flexible expansion between nodes through serial operations, parallel processing, and supports the storage of diverse information types. The multiple-node linkage is shown as in the AFM image, shown images are from one experiment representative of 3 independent replicates. Scale bar: 50 nm.
Definition of data domain in DONLDS system
In this study, we conceptualized and crafted three DON nodes with the purpose of facilitating high-efficiency data operation. In devising our data storage scheme, we focused on optimizing the layout of the data domains. Leveraging the addressability of DON nodes in which staple strands have their own position, we were able to precisely utilize the central part of DON nodes as data domains. To denote the binary ‘1’ on the surface of the DON nodes, we implemented connections at SA linked domains (red dots), coupling streptavidin (SA) with biotin-modified data strands, only unlabeled biotin staples located at the SA linked domains are defined as code ‘0’. Because the size of SA is ~6 nm41, we adopted a minimum spacing of 20 nm between adjacent SA connection domains, thus effectively preventing neighboring effects and minimizing potential cross-interference (Fig. 2a).
a DONs are delineated by a black linear framework signifying its scaffold strands, complemented by blue filigrees that represent the staple strands, collaboratively assembling a robust DNA nanostructure. Red dots indicate potential SA conjugation sites for data encoding, where SA presence/absence defines bit states (1/0). b The synthesis and data encoding process of DON nodes involves the utilization of M13 single-stranded DNA extracted from Escherichia coli as the scaffold, the data strands along with SA are affixed to these nanostructures, thereby accomplishing the encoding process. The AFM image depicting all domains of the three DONs, each interfaced with SA for the storage of information, shown images are from one experiment representative of 3 independent replicates. Scale bar: 50 nm. c The size of each node used by DONLDS for storing information, as well as its storage capacity and density, shown images are from one experiment representative of 3 independent replicates. Scale bar: 50 nm.
The encoding process started with mixing the scaffold and staple strands together within a reaction system to induce their spontaneous self-assembly through annealing from 95 °C to 25 °C. The data writing process was achieved by covalently coupling SA-decorated data strands, thus enabling precise data placement on the DON nodes. Ultimately, AFM images confirm site-specific conjugation of SA to biotin-modified domains (Fig. 2b and Supplementary Fig. 8). To comprehensively evaluate the performance of these three distinct types of DON nodes, we conducted an in-depth comparative analysis of their size, capacity, and density. The results revealed that three types of DON nodes exhibited similar storage potential (Fig. 2c and Supplementary Tables 2–3). Specifically, each RDON node boasts an area of approximately 5400 nm2, which was capable of 12 bits of binary data. CDON and TDON nodes possess areas of around 4500 nm2 and 5072 nm2, respectively, each of which could store 8 bits of binary data. After calculating the storage density of these DONs, we define the maximum number of bits storable per cubic centimeter. The calculations indicated that the RDON boasts the highest storage density at 222.22 Gbit/cm2, followed by the CDON at 177.76 Gbit/cm2, with the TDON at 157.78 Gbit/cm2.
Establishment of English alphabet information storage by TDON nodes
To demonstrate the programmability of the DON nodes, we employed TDON for encoding English alphabets and pointers. Each TDON node was equipped with a data domain to encode in a clockwise spatial arrangement and the pointer domain to install pointer strands (Fig. 3a). Then, we developed a set of 8-bit binary codes for the 26 English letters. To ensure unambiguous spatial orientation, we used a fixed 3-bit orientation code (‘010’) and a streptavidin (SA) marker on the inner edge of side “a” to break rotational symmetry. This combination eliminates ambiguities from origami flipping and rotational misalignment, followed by a 5-bit alphabet coding set for letter differentiation. Take the English alphabet ‘A’ for example, the whole process would include four Analogous to the scaffold loop in CDON, the RDON dimer employs its blue orientation domain to break structural symmetry steps: inputting 8-bit binary sequence ‘01000010’ to the TDON node-based encoding system, reading the nodes by AFM, outputting the nodes into a binary sequence, and converting into ‘A’ (Fig. 3b). Following the rules, we input the corresponding binary data and created 26 distinct TDON nodes, which were read precisely by AFM. The experimental data indicated that SA could effectively dock with the predetermined storage domains, ensuring information storage accuracies of 91.17 ± 0.86% to 95.64 ± 2.05% for TDON nodes (Fig. 3c and Supplementary Figs. 9–11).
a A schematic diagram of the TDON data domain and pointer domain is provided. In the floor plan, the area within the gray line frame represents the attachment position of SA, which is the data domain, and the area within the blue frame is the pointer domain of TDON. In the three-dimensional view, the black short cylinders represent the orientation code domain, and the orange cylinders represent the Alphabetic code domain. The data in the data domain is read in a clockwise direction. b A flowchart for the writing and reading process of English letters is illustrated. The figure describes the English alphabet alongside its corresponding binary code table and provide a stepwise depiction of the encoding procedure for the letter ‘A’ within the TDON node. c The AFM images showcase the storage of all 26 English letters on the TDON nodes, shown images are from one experiment representative of 3 independent replicates. d The assembly process of ten different TDON nodes is depicted. The AFM images illustrate the synthesis of five distinct dimer letter combinations: ‘HE’, ‘L1L2’, ‘O1W’, ‘O2R’, and ‘L3D’. e The graph shows the frequencies of the dimer nodes to be 18.62 ± 0.31% ~ 19.87 ± 0.35%, with a node mismatch rate of 3.83 ± 0.95%. The sample size is denoted as N. Error bars represent Standard Deviation (SD). Data are from n = 3 independent experiments, presented as mean ± SD. Individual data points from each experiment are overlaid on the bars. Scale bar: 50 nm. Source data are provided as a Source Data file.
After the achievement of TDON-based storage nodes which emerged as the fundamental building block of the linked list data structure, we investigated the ability of the instructions to execute commands. The phrase ‘HELLO WORLD’ was demonstrated and decomposed into five English alphabet pairs independently, namely ‘HE’, ‘L1L2’, ‘O1W’, ‘O2R’, and ‘L3D’. After anchoring five sets of DNA strands with specially designed DNA sequences as pointers onto the TDON nodes respectively, the instruction of linkage could be executed by introducing operation strands which generated 5 dimeric TDON nodes with an accuracy of 90.73 ± 1.36% to 92.07 ± 1.78% (Fig. 3d and Supplementary Fig. 12). Additionally, to verify the specificity between operation strands and pointer strands, we conducted instructions by co-incubating all ten monomer storage nodes with their corresponding operation strands in a single reaction system. The results showed that these ten nodes were able to connect precisely according to the predetermined connection objectives, resulting in the formation of five groups of dimeric nodes. Excluding the unpaired single nodes, the frequencies of the dimer nodes within the reaction system were 19.87 ± 0.35% (HE), 19.43 ± 0.46% (L1L2), 18.62 ± 0.31% (O1W), 19.25 ± 0.32% (O2R), and 18.99 ± 0.77% (L3D), respectively (Fig. 3e and Supplementary Fig. 13).
Construction of digital information storage based on CDON nodes
At the left and right sides of each CDON node, we meticulously designed six pointer strands with unique DNA sequences to facilitate the pairing of operation strands, thereby ensuring the stability and specificity of inter-node connections. To ascertain the reversibility and stability of information interconnectivity between CDON nodes, we conducted multiple rounds of connection and disconnection experiments on the data nodes. In the initial stage (stage 0), the CDON nodes remained mutually independent. After the introduction of operation strands to execute linkage instruction, the CDON nodes combined together to form dimers in stage 1. Here, the operation strands not only connected two CDON nodes, but had the toehold part for disconnection. Hence, the operation strands to execute disconnection instruction could result in two independent CDON nodes via toehold-mediated strand displacement, which is a fundamental mechanism in DNA nanotechnology where transient binding at short single-stranded “toehold” regions initiates branch migration to displace hybridized strands (stage 2). Of course, with the re-introduction of operation strands, the CDON nodes reconstituted the dimeric structures (stage 3, Fig. 4a). The statistical analysis of the average accuracy rates from stage 0 to stage 3 were 79.87% (connection) and 95.34% (disconnection) (Supplementary Fig. 14).
a Experiment on Node connection and disconnection Based on CDON. At stage 0, which is the initial stage, the data nodes are in a separate stage. Upon transitioning to stage 1, the data nodes change from a separate stage to a combined stage. Subsequently, at stage 2, the data nodes dissociate from the combined stage back to a separate stage. Finally, at stage 3, the data nodes once again enter a combined stage. Scale bar: 200 nm. b CDON nodes store Arabic numerals information from 1–10. Scale bar: 50 nm. c Head nodes and data nodes. In the head nodes, the confirmation point is blue, the north latitude point is green, the south latitude point is gray, the west longitude point is red, and the east longitude point is orange. In the data nodes, the unit degree point is yellow, the unit minute point is light blue, the data domain ‘0’ is white, and the data domain ‘1’ is black. d A world map displaying the binary codes and locations of four cities, with color-coded pins using blue for New York, orange for Beijing, green for Brasilia, and purple for Sydney. e CDON nodes store the latitude and longitude information of each city, New York: 40°43’N, 74°00’W, Beijing: 39°56’N, 116°20’E, Brasilia: 15°47’S, 47°56’W, Sydney: 33°55’S, 150°53’E. Scale bar: 50 nm.
We also employed CDON nodes for digital information storage. The unique loop structure positioned on the right side of the cross-shaped storage nodes acted as a vital orientation reference, which could unambiguously determine the spatial orientation and configuration of the complete storage node. We selected four dedicated coding domains (positions 1-4 in Supplementary Fig. 15) within CDON nodes to implement 4-bit binary encoding for digital information storage. The AFM images vividly illustrated the ten CDON nodes for digital information storage, The data retention accuracy of CDONs ranged from 91.17 ± 1.64% to 95.94 ± 1.04% across three independent experiments (Fig. 4b and Supplementary Fig. 16).
To show the application of these CDON nodes, we constructed a linked memory to represent geographical coordinates which were capable of pinpointing precise locations on Earth and providing functionalities such as navigation, mapping services, and positioning. Because geographical coordinates usually contain latitude and longitude data, we used header nodes to show the latitude and longitude and data nodes to display the data (Fig. 4c). Within the CDON header node, the blue circle denoted the header node’s positioning point, while the gray circles were allocated for the storage of urban latitude and longitude. In addition, we established four binary positions within the header node: green for northern latitude, gray for southern latitude, red for western longitude, and orange for eastern longitude. For the data nodes, each was capable of storing 8 bits of binary information, which were converted into 2-digit decimal numerical data according to the encoding rules. In light of the extensive range and specific precision requirements for data denoted in degrees, it was customary to allocate one or two nodes for its retention. The storage demanded for such data spans from 0 to 12 bits of binary information, allowing for a nimble adaptation to the varying scales of data storage needs. By contrast, data annotated in minutes was more concise, requiring merely a single node to encapsulate, with its storage space confined to within 0 to 8 bits of binary information. This design ingeniously enhanced the efficiency of storage space while ensuring the high fidelity of data representation. Then, we assigned 7-bit binary codes and linked five or six CDONs for four cities (New York, Beijing, Brasilia, and Sydney), each representing a different global direction for storage (Fig. 4d). For example, a five-node structure was utilized to store the geographical coordinates of New York: 40°43’N, 74°00’W, in which the decimal latitude and longitude information (40°43’, 74°00’) was translated into binary data (01001100010000110111010011001100) and stored within the CDON nodes. The AFM images demonstrated the storage of the geographical coordinates for four cities (Fig. 4e and Supplementary Figs. 17–20).
Encoding Chinese characters information by RDON nodes
Chinese characters, which own a large amount of characters and abundant meanings, are usually encoded by 16 bits of binary numbers in the computer. To store Chinese characters efficiently and accurately, we established a 16-bit binary storage scheme according to GB2312, in which each Chinese character was represented by 4 Arabic numerals. We listed some of the Chinese characters with their zone-bit codes (Fig. 5c). Because of the larger storage space required for Chinese characters, two RDON nodes as a unit were utilized for one Chinese character instead of a single RDON node. Considering the symmetry of RDON nodes and the order of Chinese characters, the RDON unit was divided meticulously into three domains, including the index domain, orientation domain, and Chinese character domain. The index domain was encoded by four red marker circles to establish the reading order of the characters, ensuring the accuracy of information retrieval. The orientation domain was indicated by three blue marker circles, which could ensure the information in the correct direction. Analogous to the scaffold loop in CDON, the RDON dimer employs its blue orientation domain to break structural symmetry (Fig. 5a). The orientation domain cooperates with the scaffold loop to enforce uniform node orientation during substrate adsorption, eliminating data readout ambiguity caused by structural flipping. The Chinese character domain was responsible for storing the binary encoding information of Chinese characters with 16 gray marker circles in four rows, which were read from left to right and from up to down (Fig. 5a). Taking the Chinese character ‘HOU’ for example, it was firstly searched in GB2312 to obtain its zone-bit code 2681. Then, the number was encoded into a 16-bit binary RDON unit via a coding mapping procedure. Finally, the generated RDON unit could be read and decoded into ‘HOU’ (Fig. 5b). According to these encoding standards, eight Chinese characters, which mean noble morality, strong mind, seek the truth and practice earnestly, were encoded into eight independent RDON nodes and validated by AFM images and their height analysis (Fig. 5c and Supplementary Figs. 21–28).
a Diagram illustrating the division of index, orientation, and Chinese character domains within a dimer RDON nodes. b The process for storing and retrieving Chinese characters. Initially, the character ‘HOU’ is identified, followed by the retrieval of its zone-bit code, which is transcoded into binary and inscribed within the RDON node. After the information is read using an AFM, the binary code is reconverted back to the zone-bit code, ultimately leading to the retrieval of the character ‘HOU’. c The Chinese character zone-bit code and RDON nodes store eight Chinese characters arranged in the following reading order: ‘HOU DE’; ‘HONG YI’; ‘QIU SHI’; ‘DU XING’. These characters represent the meanings of ‘profound virtues’, ‘magnanimous and firm’, ‘seeking truth’, and ‘persevering practice’. d Tetramer RDONs with expanded storage capacity have a storage capacity of 48 bits and a storage density of 222.22 Gbit/cm2. Storing the Chinese phrase in the tetrameric RDONs: ‘DAO KE DAO, FEI CHANG DAO’. Analysis of AFM images show that the information stored in the two DONs is successfully decoded and transformed into a phrase from the Chinese book Tao Te Ching: ‘DAO KE DAO, FEI CHANG DAO’ (shown images are from one experiment representative of 3 independent replicates). Scale bar: 50 nm.
Beyond monomeric RDON nodes, we leveraged RDON’s modular expandability to augment storage capacity specifically within the Chinese character domain (excluding the index and orientation domains). We demonstrated storage of the traditional Chinese maxim ‘DAO KE DAO FEI CHANG DAO’ from the Tao Te Ching using an RDON tetramer, achieving 48-bit capacity at 222.22 Gbit/cm2 density. This resolves the long-text storage bottleneck, and the modular architecture underpins future chain or grid-based large-scale DONLDS system (Fig. 5d, Supplementary Figs. 29–30).
The linked data storage of different types of information
The distinguished feature of DONLDS system lies in its capacity as a storage medium to facilitate precise control over the connection and disconnection between DON nodes, which greatly facilitates efficient insertion and removal of data elements. In this process, there is no need for extensive rearrangement or reconfiguration of the entire structure, thereby maintaining the consistency and integrity of the data structure. The workflow executed by the DONLDS system illustrated the processes of data insertion and deletion. (Fig. 6a–b). Then we turned to delineate the intricate process of data insertion and deletion by RDON nodes. Prior to data manipulation, a linked list was constructed by the linkage of RDON nodes 1, 2 and 4 with linking strands. When executing the insertion operation between nodes 2 and 4, the detachment strands were introduced and nucleated at the toehold domains to initiate the strand-displacement reactions, which generated the disconnection of nodes 2 (P, denoting ‘previous’ node) and 4. Then, two new linking strands were added to form the linkage of the pointer of node 3 (S, denoting ‘subsequent’ node) with that of nodes 2 (P) and 4, thereby facilitating the completion of the data insertion process. The deletion operation aimed to remove node 2 (P) from a list consisting of four nodes. This was achieved by simply disconnecting node 2 (P) from its adjacent nodes and establishing a new linkage between nodes 1 and 3. Here, the operation of pointer replacement was achieved through a sophisticated DNA strand displacement, which ensured the high precision, reliability, and stability of the data list editing process at the molecular scale. This strand displacement mechanism leverages the properties and complementarity of DNA molecules, accomplishing precise adjustment of pointers between nodes through a process of carefully designed sequence matching and strand exchange (Fig. 6c and Supplementary Fig. 31).
a A flowchart for the insertion of DON nodes. b A flowchart for the deletion of DON nodes. c Establish a linked list and carry out the insertion and deletion of nodes. d Created linked list 1 and performed deletion and insertion operations to establish linked list 2. T, R, and C denote distinct DNA origami nanostructure nodes: triangular (TDON), rectangular (RDON), and cross-shaped (CDON). e The TDON nodes store ‘DNA’ English letter information, the RDON nodes store ‘HELIX’ Chinese character information, and the CDON nodes store ‘1953’ digital information. Establish the information connection among each storage node and achieve the storage of ‘DNA-HELIX-1953’ information. f Remove the messages ‘helix’ and ‘1953’, then re-encode and input new storage nodes, ‘Storage’ and ‘1988’. g A new linked list is constructed to realize the storage of ‘DNA-STORAGE-1988’ information. Scale bars: 50 nm.
To further underscore the scalability of DON nodes in data storage, we crafted an interconnection mechanism among variously shaped DONs. These strategic linkages not only augmented the synergy between nodes but also markedly enhanced flexibility and capacity in the overall storage system. As depicted in Fig. 6a, our storage nodes consist of three TDON, four RDON, and two CDON nodes, each boasting a unique base-pairing scheme and a complex folding architecture. Initially, we constructed a bidirectional linked list comprising nine nodes, and through the execution of deletion and insertion operations, the initial list 1 was transformed into list 2 (Fig. 6d).
The paramount advantage of the DONLDS technology lies in its dynamic memory space allocation capabilities. Following data encoding and storage, we sequentially linked each structure with detachable operation strands. To ensure the stability and reliability of data transmission, we designed six to eight sets of distinct pointer strands between different-shaped DONs, each of which was a unique sequence for high-specificity recognition with the operation strands. This allowed each DON to interlock and complement one another like precision gears, forming a highly modular and expandable linked list. We encoded the English letters ‘D’, ‘N’, and ‘A’ by TDON nodes, Chinese characters ‘LUO XUAN’ for ‘HELIX’ by RDON nodes, and the CDON nodes for the historically significant digits ‘1953’, signifying the year of the discovery of the DNA double helix (Fig. 6e and Supplementary 32–34). In constructing our DONs linked list, we specifically designated the triangular DON node as the head node. To distinctly mark this node, A SA connection domain was embedded within the edge “a” of TDON as an identifying feature of the head node. Subsequently, we connected the RDON and CDON nodes to the head node by the predetermined connection logic. AFM images validated the interconnection of the nine distinct storage nodes. By analyzing the height variations presented in the images and converting them into corresponding binary data, we stored the complete multi-type textual information ‘DNA-HELIX-1953’ (Fig. 6e and Supplementary Fig. 37). This experimental outcome robustly demonstrates the exceptional capabilities of the DONLDS technology in the splicing of multi-type storage nodes, proving its ability to seamlessly integrate data between nodes of varying shapes and types, while maintaining data storage accuracy and stability with high efficiency.
Subsequently, we undertook the transition from linked list 1 to linked list 2. In the preliminary stage, we disconnected the data linkage between the TDON node T3 and the RDON node R1. This was achieved by introducing inter-node unlocked strands, deftly substituting the operation strands within the nodes, thereby facilitating the operation of data deletion. Thereafter, within the newly generated RDON nodes, we encoded and stored the Chinese character ‘storage’, while the CDON nodes recorded the number ‘1988’, signifying the year the concept of DNA storage was first proposed (Fig. 6f and Supplementary Figs. 35–36). Ultimately, by introducing the operation strands of nodes T3 and R5, as well as R8 and C3, and performing base-complementary pairing with their pointer strands, the insertion operation of the ‘STORAGE-1988’ linked list was executed. The AFM images depicted the storage of the ‘DNA-STORAGE-1988’ sequence, with the detailed imagery showcasing the integrity of the data (Fig. 6g and Supplementary Fig. 38).
To minimize error accumulation, we employed a composite reading strategy involving cross-verification of multiple nine-node units. Decoding thresholds were established as follows: a signal connection (SA) exceeding 80% was interpreted as ‘1’, a connection below 20% as ‘0’, and connections between 20% and 80% were considered invalid bits. Analysis of 24 randomly selected samples (totaling 1728 bits) revealed a 100% recovery rate when compared bitwise with the pre-defined ‘DNA-STORAGE-1988’ sequence (Supplementary Figs. 38–39). This experimental result further validated the potential application value of DNA nanotechnology-based data processing systems in the realm of information storage and manipulation. To accurately predict the storage success rate for larger-scale data, we use a refined computational model. This model is described in detail in the supplementary methods section, which includes parameterized mathematical formulas that systematically calculate the data storage success rate under corresponding conditions by taking the raw bit error rate as an input variable (Supplementary Table 4 and Fig. 40).
Discussion
The DONLDS leveraged a design of DON nodes and a linking mechanism based on DNA strands, demonstrating ingenuity in the field of information storage. The core components of DONLDS consisted of DONs of various shapes, including triangular, rectangular, and cross-shaped nodes. Each node encompassed a data domain dedicated to the storage of binary information and a pointer domain which was responsible for interconnecting the nodes.
In terms of data storage capability, we have exemplified the application of three distinct DON nodes (triangular, rectangular, and cross-shaped) in the storage of textual information. We demonstrated the storage of 26 English letters, 10 Arabic numerals, and 16 representative Chinese characters using GB2312 code, confirming the DONLDS’s efficiency and accuracy. The crowning feature of the DONLDS lay in its dynamic memory space allocation. Following data encoding and storage, the system sequentially connected structures through detachable operation strands and designed multiple sets of specific sequence pointer strands to ensure stable and reliable data transmission. Utilizing ten distinct TDON nodes, the English letters that comprise ‘HELLOWORLD’ were shown with the executed data connections according to a predefined program. Experimental results revealed a connection accuracy rate of 96.17%, substantiating the specificity and precision of the operation and pointer strand connections. Furthermore, through multiple rounds of connection and disconnection experiments with CDON nodes, we confirmed the reversibility and stability of information interconnection between DON nodes. By adjusting the operation strands between DONs, we performed insertion and deletion operations on storage nodes. We also demonstrated the process of node insertion and deletion, efficiently executing parallel operations by merely altering the direction of the pointers without the need to relocate other nodes.
Moreover, the flexibility of memory allocation was not confined to nodes of the same type but extended across different types, optimizing resource utilization and enhancing the adaptability and efficiency of the DONLDS. By interconnecting DONs of varying shapes, we showed the simultaneous storage of a variety of information types, including English characters, Chinese characters, and digits. AFM images validated the interconnection of different DON nodes for ‘DNA-HELIX-1953’, showcasing DONLDS’s exceptional prowess in the splicing of multiple node types and data integration. To verify data deletion and insertion operations, we constructed a 9-node bidirectional linked structure. Node deletion was achieved through thermodynamic annealing and strand displacement reactions, while insertion operations integrated new data directly via base pairing principles. In our experiments, we severed and reconnected specific nodes, encoding and storing the ‘DNA-STORAGE-1988’ information, with AFM images clearly depicting the integrity of the data. These results further substantiated the potential application value of DNA nanotechnology-based data processing systems in the realm of information storage and manipulation. While DONLDS currently utilizes TDONs, RDONs, and CDONs for ease of fabrication, achieving optimal linked-data storage necessitates balancing multiple factors: pointer domain complexity, data capacity/density, structural stability, addressability, and synthesis feasibility. For instance, hexagonal DONs could offer greater pointer complexity, whereas circular DONs maximize data storage area but present challenges for placement precision and structural rigidity. Future designs will systematically explore these shape-dependent trade-offs.
DNA storage and DNA computing currently face ‘insufficient rates’ primarily due to the complexity and lengthy nature of the synthesis, incubation, and read/write processes. However, researchers are striving for breakthroughs in synthesis with emerging technologies such as enzymatic DNA synthesis10,42,43 and high-density microarray chips44,45,46. Under mild conditions, enzymatic DNA synthesis enables faster base coupling and can be carried out in parallel on high-throughput array chips to increase writing speed greatly. High-density chips, on the other hand, employ nanoscale fabrication and localized heating to overcome challenges posed by optical diffraction limits and electrode crosstalk. In addition, the coupling of DNA and semiconductor technologies holds considerable promise: using electrode arrays for parallel DNA synthesis47, combined with cross-disciplinary innovations in microfluidics48,49, optics50, and automated DNA synthesis equipment51, offers vast possibilities for DNA computing, DNA motors, and many other DNA-based nanomaterials. Although the costs for large-scale synthesis and post-processing are still relatively high, ongoing advancements in batch production, semiconductor processes, and enzyme engineering will continue to drive down reagent and synthesis expenses. This will, in turn, accelerate the broader application of DNA nanotechnology and open new frontiers for DNA storage and DNA computing.
Although a single DON achieves high local density, system-level density is reduced by essential replicas. This redundancy counters stochastic DNA self-assembly errors, ensuring molecular data fidelity, which is a crucial trade-off for reliability. Thus, DONLDS delivers node-level density, while system-level gains depend on fabrication optimization and algorithmic advances. With the continuous optimization and advancement of DNA nanotechnology, it is anticipated that the storage capacity of DONLDS will experience significant enhancement. These advancements include technologies for the large-scale synthesis of DNA origami, which aim at reducing synthesis costs and expanding storage capacity52,53. Continued advancements in AFM techniques, such as the application of AI-AFM fusion to RNA dynamics, alongside techniques enabled by High-Speed Atomic Force Microscopy (HS-AFM) like label-free spectroscopy at microsecond resolution, millisecond-scale imaging of single-molecule complexes, and visualization of transporter diffusion kinetics, hold the potential to improve DON data read speeds by orders of magnitude54,55,56,57. Although current limitations in read/write speed, cost, and per-node capacity pose challenges for the immediate commercialization of DONLDS, experimental results demonstrate that single-node data accuracy exceeds 91%. Although the raw bit error rate is relatively high, ranging from 1.11% to 1.44%, any desired level of reliability can theoretically be achieved through redundancy and error-correcting codes (ECC). However, the required redundancy increases substantially with data volume. For example, using a 10-fold redundant encoding scheme with majority voting, we improved the system-level correctness for reading 1 kB of data from nearly zero to over 99.9%, demonstrating that practical data reliability can be attained in DONLDS through appropriate system-level design. For larger data volumes, such as 1 MB, more sophisticated ECC schemes (e.g., Reed–Solomon) and higher redundancy ratios would be required. Therefore, before DONLDS-based storage systems can be practically deployed, they must incorporate such robust ECC and redundancy schemes. At this stage, DONLDS should be regarded as a proof-of-concept platform for exploring dynamic data operations (e.g., insertion and deletion) in molecular storage systems. Its scalability ultimately depends on synergistic breakthroughs in coding theory, materials science, and readout technologies.
Methods
Materials
All chemicals were obtained from Sigma (USA). All short strands were purchased from Sangon (ULTRAPAGE purification, Shanghai, China). Biotin-modified oligonucleotides were supplied by Sangon and purified by high-performance liquid chromatography (HPLC). M13mp18 DNA was purchased from Bioruler (cat. no. B3003, Jiangsu, China), The sample was stored in 500 μL of water at a concentration of 100 nM. The streptavidin powder was obtained from Sigma (cat. no. S4762, USA). All solutions were prepared with Milli-Q water (resistivity = 18.2 MΩ cm).
Synthesis of DON nodes
The single-stranded DNA of the M13mp18 virus was selected as the scaffold strand. The scaffold strands (100 nM) and staple strands (100 μM) were mixed at a 1:10 molar ratio in 1× TAE/Mg2+ buffer solution in the experiment. The components of the buffer include 40 mM Tris (to maintain pH, Sigma, cat. no. 252859), 20 mM acetic acid (Sigma, cat. no. 338826), 12.5 mM magnesium acetate (Sigma, cat. no. 228648), and 2 mM EDTA (Sigma, cat. no. 798681). The mixed DNA solution was then transferred to a PCR instrument (Thermo Fisher Veriti Fast, cat. no. 4375305) for precise temperature control. Initially, the solution was heated to 95 °C for 5 min to denature the single-stranded DNA and ensure homogeneity of the mixture. The temperature was then gradually decreased to 10 °C at a rate of 1 °C per minute, facilitating a slow annealing process that promotes the formation of stable and precise hybrid structures between the staple and scaffold strands.
Purification of DON nodes
Excess DNA staple strands were purified using ultra centrifugal filters 100 K ultrafiltration tubes (Merck millipore, cat. no. UFC5100BK). The sample solution was combined with 400 μL of 1× TAE/Mg2+ solution and transferred to the ultrafiltration tube. It was then centrifuged at 25 °C and 3000 g for 10 min to eliminate residual liquid from the tube’s base. Subsequently, 400 μL of 1 × TAE/Mg2+ solution was added to the tube for centrifugation, and this process was repeated three times. Following the repetitions, the ultrafiltration tube was inverted, placed in a new ultrafiltration tube holder, and centrifuged at 1000 g for 10 min to achieve purified DON. Finally, UV quantification was performed on the purified monomeric DON to determine the concentration of each DON.
Synthesis of data strands
Streptavidin powder was dissolved in 1 × PBS/Mg2+ buffer to prepare a 1 mg/mL stock solution. This stock was then diluted in the same buffer to a final working concentration of 5 μM for the experiments. The components of the buffer include 10 mM sodium dihydrogen phosphate (Sigma, cat. no. 71500), 2 mM disodium hydrogen phosphate (Sigma, cat. no. 71643), 137 mM sodium chloride (Sigma, cat. no. S7653), 2.7 mM potassium chloride (Sigma, cat. no. P3911), and 12.5 mM magnesium acetate (Sigma, cat. no. 228648). The 3’ end of staple strands was modified with biotin to enable its connection to streptavidin (SA). The biotinylated staple strands (100 μM), following modification, was then mixed with SA (5 μM) in a precise 1:1 molar ratio and thoroughly agitated to ensure uniformity of the mixture. After homogenization, the solution was placed on a shaker and incubated for 2 h at a constant temperature of 37 °C to facilitate effective binding between the biotinylated staple strands and SA. Excess SA were purified using ultra centrifugal filters 50 K ultrafiltration tubes (Merck Millipore, cat. no. UFC5050BK).
The data writing process of DON nodes
The purified DON nodes (2.5 nM) were mixed with data strands (100 μM) in a 1:10 molar ratio, and the mixed DNA solution was transferred to a thermal cycler (Thermo Fisher Veriti Fast, cat. no. 4375305) for an accurate annealing procedure. Firstly, the solution was heated to 50 °C, and then gradually decreased to 25 °C at a rate of 1 °C per minute. This completed the data writing process.
The serial operation of DON nodes
The serial operation of DON nodes is achieved through base pairing between the operation strands and their edge pointer strands. Mix DON nodes at a molar ratio of 1:1 and then add equal concentrations of operation strands to the solution system. Finally, place in a thermal cycler (Thermo Fisher Veriti Fast, cat. no. 4375305) for the annealing process as follows: heat to 45 °C at a rate of 1 °C per second, then cool from 45 °C to 25 °C in 2 h (decrease by 0.33 °C per minute).
Reading the information on DON nodes
The DON nodes were analyzed using a Microscope MultiMode VIII atomic force microscope (Bruker, USA). Initially, 5–10 μL of the DON was deposited drop by drop onto the mica surface for 3 min. Afterward, the mica was rinsed with 400 μL ultra-pure water, and completely dried using a washing ear ball. The mica sheet was then positioned on the AFM stage for imaging in Tapping mode in air, with parameter adjustments optimized for data capture. The results were analyzed by the Nanoscope analysis (Bruker).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The main data supporting the results in this study are available within the paper and its Supplementary Information. There are no data from third-party or publicly available datasets. Other source data that support the findings of this study are available from the corresponding authors upon request. Source data are provided with this paper. The source data generated in this study have been deposited in the zenodo database under accession code 10.5281/zenodo.17414132. Source data are provided with this paper.
References
Church, G. M., Gao, Y. & Kosuri, S. Next-generation digital information storage in DNA. Science 337, 1628–1628 (2012).
Goldman, N. et al. Towards practical, high-capacity, low-maintenance information storage in synthesized DNA. Nature 494, 77–80 (2013).
Grass, R. N., Heckel, R., Puddu, M., Paunescu, D. & Stark, W. J. Robust chemical preservation of digital information on DNA in silica with error-correcting codes. Angew. Chem. Int. Ed. 54, 2552–2555 (2015).
Koch, J. et al. A DNA-of-things storage architecture to create materials with embedded memory. Nat. Biotechnol. 38, 39–43 (2020).
Matange, K., Tuck, J. M. & Keung, A. J. DNA stability: a central design consideration for DNA data storage systems. Nat. Commun. 12, 1358 (2021).
Matange, K., Tuck, J. M. & Keung, A. J. CRISPR-Cas encoding of a digital movie into the genomes of a population of living bacteria. Nature 547, 345–349 (2017).
Song, L. et al. Robust data storage in DNA by de Bruijn graph-based de novo strand assembly. Nat. Commun. 13, 5361 (2022).
Antkowiak, P. L. et al. Low cost DNA data storage using photolithographic synthesis and advanced information reconstruction and error correction. Nat. Commun. 11, 5345 (2020).
Newman, S. et al. High density DNA data storage library via dehydration with digital microfluidic retrieval. Nat. Commun. 10, 1706 (2019).
Palluk, S. et al. De novo DNA synthesis using polymerase-nucleotide conjugates. Nat. Biotechnol. 36, 645–650 (2018).
Bancroft, C., Bowler, T., Bloom, B. & Clelland, C. T. Long-term storage of information in DNA. Science 293, 1763–1765 (2001).
Shendure, J. et al. DNA sequencing at 40: past, present and future. Nature 550, 345–353 (2017).
van der Valk, T. et al. Million-year-old DNA sheds light on the genomic history of mammoths. Nature 591, 265–269 (2021).
Yang, S. et al. DNA as a universal chemical substrate for computing and data storage. Nat. Rev. Chem. 8, 179–194 (2024).
Banal, J. L. et al. Random access DNA memory using Boolean search in an archival file storage system. Nat. Mater. 20, 1272–1280 (2021).
Bögels, B. W. A. et al. DNA storage in thermoresponsive microcapsules for repeated random multiplexed data access. Nat. Nanotechnol. 18, 912–921 (2023).
Organick, L. et al. Random access in large-scale DNA data storage. Nat. Biotechnol. 36, 242–248 (2018).
Zhang, J., Hou, C. & Liu, C. CRISPR-powered quantitative keyword search engine in DNA data storage. Nat. Commun. 15, 2376 (2024).
Meiser, L. C. et al. Reading and writing digital data in DNA. Nat. Protoc. 15, 86–101 (2020).
Sadremomtaz, A. et al. Digital data storage on DNA tape using CRISPR base editors. Nat. Commun. 14, 6472 (2023).
Lin, K. N., Volkel, K., Tuck, J. M. & Keung, A. J. Dynamic and scalable DNA-based information storage. Nat. Commun. 11, 2981 (2020).
Tabatabaei, S. K. et al. DNA punch cards for storing data on native DNA sequences via enzymatic nicking. Nat. Commun. 11, 1742 (2020).
Chen, K. et al. Digital data storage using DNA nanostructures and solid-state nanopores. Nano Lett. 19, 1210–1215 (2019).
Chen, K., Zhu, J., Bošković, F. & Keyser, U. F. Nanopore-based DNA hard drives for rewritable and secure data storage. Nano Lett. 20, 3754–3760 (2020).
Rothemund, P. W. K. Folding DNA to create nanoscale shapes and patterns. Nature 440, 297–302 (2006).
Dickinson, G. D. et al. An alternative approach to nucleic acid memory. Nat. Commun. 12, 2371 (2021).
Zhang, Y. et al. DNA origami cryptography for secure communication. Nat. Commun. 10, 5469 (2019).
Aghebat Rafat, A., Sagredo, S., Thalhammer, M. & Simmel, F. C. Barcoded DNA origami structures for multiplexed optimization and enrichment of DNA-based protein-binding cavities. Nat. Chem. 12, 852–859 (2020).
Banerjee, A. et al. Single-molecule analysis of DNA base-stacking energetics using patterned DNA nanostructures. Nat. Nanotechnol. 18, 1474–1482 (2023).
Douglas, S. M. et al. Self-assembly of DNA into nanoscale three-dimensional shapes. Nature 459, 414–418 (2009).
Tikhomirov, G., Petersen, P. & Qian, L. Fractal assembly of micrometre-scale DNA origami arrays with arbitrary patterns. Nature 552, 67–71 (2017).
Chao, J. et al. Solving mazes with single-molecule DNA navigators. Nat. Mater. 18, 273–279 (2019).
Chatterjee, G., Dalchau, N., Muscat, R. A., Phillips, A. & Seelig, G. A spatially localized architecture for fast and modular DNA computing. Nat. Nanotechnol. 12, 920–927 (2017).
Tikhomirov, G., Petersen, P. & Qian, L. Programmable disorder in random DNA tilings. Nat. Nanotechnol. 12, 251–259 (2017).
Voigt, N. V. et al. Single-molecule chemical reactions on DNA origami. Nat. Nanotechnol. 5, 200–203 (2010).
Ji, W. et al. Encoding signal propagation on topology-programmed DNA origami. Nat. Chem. 16, 1408–1417 (2024).
Kim, M. et al. Harnessing a paper-folding mechanism for reconfigurable DNA origami. Nature 619, 78–86 (2023).
Woo, S. & Rothemund, P. W. K. Programmable molecular recognition based on the geometry of DNA nanostructures. Nat. Chem. 3, 620–627 (2011).
Fan, S. et al. Proximity-induced pattern operations in reconfigurable DNA origami domino array. J. Am. Chem. Soc. 142, 14566–14573 (2020).
Fan, S. et al. Information coding in a reconfigurable DNA origami domino array. Angew. Chem. Int. Ed. 59, 12991–12997 (2020).
Fan, X. et al. Single particle cryo-EM reconstruction of 52 kDa streptavidin at 3.2 Angstrom resolution. Nat. Commun. 10, 2386 (2019).
Kawabe, H. et al. Enzymatic synthesis and nanopore sequencing of 12-letter supernumerary DNA. Nat. Commun. 14, 6820 (2023).
Verardo, D. et al. Multiplex enzymatic synthesis of DNA with single-base resolution. Sci. Adv. 9, eadi0263 (2023).
Atwater, J. et al. Combinatorial synthesis of macromolecular arrays by microchannel cantilever spotting (µCS). Adv. Mater. 30, 1801632 (2018).
Rozkiewicz, D. I., Brugman, W., Kerkhoven, R. M., Ravoo, B. J. & Reinhoudt, D. N. Dendrimer-mediated transfer printing of DNA and RNA microarrays. J. Am. Chem. Soc. 129, 11593–11599 (2007).
Costa, J. A., Dentinger, P. M., McGall, G. H., Crnogorac, F. & Zhou, W. Fabrication of inverted high-density DNA microarrays in a hydrogel. ACS Appl. Mater. Interfaces 11, 30534–30541 (2019).
Feng, D., Xu, C., Ma, B., Zhao, C. & Liu, H. Gel-based electrochemical DNA synthesis for quasi-solid-state data storage. Chem. Eng. J. 487, 150485 (2024).
Piao, Y. et al. Bead-based DNA synthesis and sequencing for integrated data storage using digital microfluidics. Angew. Chem. Int. Ed. 64, e202416004 (2025).
Li, K. et al. DNA-DISK: automated end-to-end data storage via enzymatic single-nucleotide DNA synthesis and sequencing on digital microfluidics. Proc. Natl. Acad. Sci. USA 121, e2410164121 (2024).
Lee, H. et al. Photon-directed multiplexed enzymatic DNA synthesis for molecular digital data storage. Nat. Commun. 11, 5246 (2020).
Wang, C. et al. Cost-effective DNA storage system with DNA movable type. Adv. Sci. 12, 2411354 (2025).
Zhang, C. et al. Parallel molecular data storage by printing epigenetic bits on DNA. Nature 634, 824–832 (2024).
Weng, Z. et al. Massively parallel homogeneous amplification of chip-scale DNA for DNA information storage (MPHAC-DIS). Nat. Commun. 16, 667 (2025).
Degenhardt, M. F. S. et al. Determining structures of RNA conformers using AFM and deep neural networks. Nature 637, 1234–1243 (2025).
Heath, G. R. & Scheuring, S. High-speed AFM height spectroscopy reveals µs-dynamics of unlabeled biomolecules. Nat. Commun. 9, 4983 (2018).
Kanaoka, Y. et al. AFM observation of protein translocation mediated by one unit of SecYEG-SecA complex. Nat. Commun. 16, 225 (2025).
Jiang, Y. et al. HS-AFM single-molecule structural biology uncovers basis of transporter wanderlust kinetics. Nat. Struct. Mol. Biol. 31, 1286–1295 (2024).
Acknowledgements
This work was supported by the National Key Research and Development Program of China (2024YFA1209402), the NSFC (22274081 (J.C.), 62288102 (L.W.), T2188102 (C.F.)), the Natural Science Foundation of Jiangsu Province, Major Project (BK20212012 (L.W.)), and the New Cornerstone Science Foundation.
Author information
Authors and Affiliations
Contributions
J.C. conceived the project. J.C., L.W., C.F., and C.Z. designed the experiments. C.Z. carried out the experiments. C.Z., M.X., and J.C. collected and analyzed the data. C.Z., J.C., C.F., and L.W. wrote the manuscript. All authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhang, C., Xie, M., Wang, L. et al. Linked data storage using DNA origami nanostructures. Nat Commun 16, 11422 (2025). https://doi.org/10.1038/s41467-025-66274-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-66274-x