Abstract
Metastasis significantly impacts the mortality rate of non-small cell lung cancer (NSCLC) patients. Numerous microtubule-targeting agents (MTAs) reveal anti-metastatic efficacy, but the mechanism remains unclear. In this research, we employ KY216, a microtubule inhibitor, to generate a crystal in complex with αβ-tubulin, and illustrate that the KY216-tubulin combination binds vasohibin-2 (VASH2) to restrain NSCLC metastasis. Through crystal structure analysis, specific interaction sites between KY216 and curved tubulin are identified. KY216 decreases VASH2 levels, hindering the epithelial-mesenchymal transition (EMT) process in NSCLC. Moreover, the MTA enhances the binding of VASH2 to α-tubulin, prevents the activation of zinc finger E-box binding homolog 1 (ZEB1) by VASH2, promotes detyrosination of α-tubulin, and ultimately suppresses EMT. Additionally, KY216 elevates the levels of miR-429 to target the 3’-untranslated region (3’ UTR) of VASH2 and ZEB1 transcripts and inhibits EMT, at least partially, via the miR-429/VASH2/ZEB1 axis to block NSCLC metastasis. Overall, our investigation offers valuable insights into the roles of MTAs and VASH2 in NSCLC metastasis.
Similar content being viewed by others
Introduction
Non-small cell lung cancer (NSCLC) is the most prevalent form of lung cancer and a major contributor to global cancer-related mortality1. Despite advancements in prevention, detection, and medical treatment, metastasis remains the primary cause of death among NSCLC patients2. Distant metastasis, characterized by the spread of cancer cells to organs like the lungs, liver, brain, and bones through the blood or lymphatic system, presents a significant challenge in treatment and substantially reduces patient survival rates3,4. Therefore, a comprehensive investigation into the metastatic pathways and key signal transduction mechanisms of NSCLC is essential to develop drugs, improve therapeutic outcomes, and enhance the quality of life and prognosis for patients.
Microtubules are essential components in eukaryotic cells, playing key roles in various cellular activities like maintaining structure, substance transport, chromosome segregation, cell division, and motility5,6,7. There are two types of tubulins, α-tubulin and β-tubulin, that form heterodimers. Cancer cell growth relies on the dynamic polymerization and depolymerization of tubulin dimers8, which undergo post-translational modifications like detyrosination, acetylation, polyglutamylation, and polyadenylation. These modifications are present in tumor tissues and are linked to the epithelial-mesenchymal transition (EMT) process in cancer cells9. Targeting these dynamic processes has proven effective in developing anti-cancer drugs5,10. Microtubule-targeting agents (MTAs) are categorized as microtubule stabilizers (MSAs) like paclitaxel or microtubule destabilizers (MDAs) like colchicine and combretastatin A-4 (CA-4), based on their action11,12,13. These drugs disrupt cancer cell mitosis by targeting specific binding sites, leading to significant inhibition of cancer cell growth14,15,16. Besides, they also exhibit potent anti-metastasis activity15. However, the mechanisms regarding the impact on cancer cell migration remain uncertain and require further investigation.
Vasohibin-2 (VASH2), found within the infiltrating mononuclear cells at the angiogenic sprout front, plays a role in fostering angiogenesis17. VASH2 has been demonstrated to strongly induce EMT and angiogenesis in various cancers, such as colorectal, hepatocellular, pancreatic, and ovarian cancers17,18,19,20,21. In hepatocellular carcinoma (HCC), VASH2 promotes metastasis by upregulating zinc finger E-box-binding homeobox protein 1 (ZEB1) expression through activation of the ZEB1 promoter21. Recent studies have shown that high expression of VASH2 is significantly associated with poor prognosis in lung squamous cell carcinoma patients and promotes lymph node metastasis in lung cancer in vivo22,23. As the first de-tyrosinase discovered, VASH2 also controls the detyrosination of α-tubulin24,25. VASH2 has a transglutaminase-like function and acts as a cysteine protease in the catalytic reaction with a non-canonical Cys-His-Ser catalytic triad structure24. VASH2 comprises an N-terminal structural domain (NTD) and a C-terminal structural domain (CTD), the Cys-His-Ser catalytic triad pocket is located in these two structural domains and is strongly positively charged26,27. This positively charged property recognizes negatively charged α-tubulin tails and promotes detyrosination of α-tubulin in the catalytic pocket27. Considering the pivotal role of VASH2 in angiogenesis, cancer migration, and post-translational modification of tubulin, further investigation into its impact on lung cancer is of great biological significance.
Mammalian microRNAs (miRNAs) play a crucial role in regulating gene expression by binding to the 3’ untranslated region (UTR) of target messenger RNA (mRNA), leading to mRNA destabilization and inhibition of translation28. Specifically, the miR-200 family targets ZEB1/ZEB2 mRNAs to inhibit EMT in various cancer cells29,30,31. In lung cancer, the miR-200 family exhibits strong anti-metastatic and anti-angiogenic effects32,33. Studies have shown that miR-429 has reduced serum levels in NSCLC patients34 and is significantly suppressed in metastatic lung adenocarcinoma in mice35. Low expression of miR-429 is closely linked to shorter overall survival (OS) and disease-free survival (DFS) in NSCLC patients34,36. Furthermore, miR-429 has been demonstrated to inhibit the migration of A549 cells37.
In this study, we resolve the high-resolution X-ray crystal structure of the MTA KY21638 bound to the colchicine binding site of tubulin. Furthermore, we confirm the efficacy of KY216 in inhibiting the growth and metastasis of NSCLC both in vitro and in vivo. Mechanistic investigations reveal that KY216 suppresses the VASH2/ZEB1-induced EMT process by enhancing the VASH2/α-tubulin interaction and increasing miR-429 levels, ultimately leading to the restriction of NSCLC metastasis.
Results
High-resolution X-ray crystal structure of KY216 in complex with αβ-tubulin
In our previous studies, an N-heterocyclic-fused deoxypodophyllotoxin (DPT) analog, KY216, was designed and synthesized to act as a potent inhibitor of microtubule polymerization by targeting the colchicine-binding site and exhibited promising anti-tumor activity (Fig. 1a)38. To further investigate the mechanism of KY216, we elucidated the binding mode of KY216 with the T2R-TTL complex (consisting of two α/β-tubulin heterodimers, stathmin-like protein RB3, and tubulin tyrosine ligase), and obtained a clear and high-resolution X-ray crystal structure (PDB code 8YRK).
a Chemical structure of KY216. b Overall structure of the complex crystallized (PDB code 8YRK). RB3-SLD is orange; α-tubulin is blue; β-tubulin is pink; TTL is wheat; KY216 is marked with a black box and shown as a yellow sphere. GTP is shown as a green sphere, and GDP is shown as a purple sphere. c The electron cloud density of KY216 is presented at 2mFo-DFC with a contour level of 1.0σ. d The diagram details the molecular interactions between the KY216 (represented by yellow sticks) and the tubulin dimer, with α-tubulin illustrated as a blue cartoon and β-tubulin as a pink cartoon. It highlights one classical hydrogen bond, marked with a cyan dashed line, and three non-classical hydrogen bonds, marked with light green dashed lines. Additionally, one halogen bond is present, marked with a gray dashed line. The corresponding amino acids are represented using their single-letter codes. e Comparison of the binding modes between Podophyllotoxin and KY216. Structures of Tubulin-KY216 (pink cartoon) and Tubulin-podophyllotoxin (PDB code 1SA1, gray cartoon) are superimposed. KY216 (yellow) and podophyllotoxin (green) are shown as rods. f Superimposition of “straight” tubulin (PDB code 7TQY, gray cartoon) and “curved” Tubulin-KY216 (pink cartoon). Black arrows point out the conflict between “straight” tubulin and KY216 (yellow spheres).
We resolved the crystal to a resolution of 2.7 Å, and the X-ray data are summarized in Table 1. The high-resolution crystal structure elucidates the detailed interactions between KY216 and tubulin (Fig. 1b). KY216 exhibits a complete and uniform electron cloud density throughout the crystal (Fig. 1c). KY216 occupies the colchicine binding site that is formed by residues of strands S8, S9 and S10, loop T7 and helixes H7 and H8 of β-tubulin and loop T5 of α-tubulin (Fig. 1d). The trisubstituted phenyl chlorine atom of the E-ring is tightly embedded in a hydrophobic pocket consisting of βLeu253, βIle368 and βAla248, located in the region of the colchicine binding zone 239. More importantly, the chlorine atom is tightly bound to the main chain carbonyl of βVal236 through the formation of a halogen bond (gray dotted line), which enhances the binding stability between the trisubstituted phenyl and β-tubulin. A classical hydrogen bond is formed between the carbonyl of the D ring and the active hydrogen of the βAsp249 amide (cyan dashed line). Three non-classical hydrogen bonds (marked by green dashed lines) were formed between the A-ring dioxane hydrocarbon-based hydrogen and the βVal313 main chain carbonyl, the E-ring substituent hydrocarbon-based hydrogen and the βCys239 sulfur atom, and the D-ring carbonyl and the βAla248 hydrocarbon-based hydrogen (Fig. 1d). The formation of halogen and hydrogen bonds not only shapes the unique mode in which the compound binds to tubulin, but also substantially enhances its stability.
The tubulin-KY216 complex was compared to the tubulin-podophyllotoxin structure (PDB code 1SA1) by superimposition. It was found that KY216 and podophyllotoxin have a high degree of overlap, with a root mean square deviation (RMSD) of 0.59 (733 to 733 atoms), confirming the accuracy of KY216’s binding mode to tubulin (Fig. 1e). Overlap (RMSD = 2.319, 807 to 807 atoms) with an unlinked straight tubulin dimer (PDB code 7TQY) shows that KY216 prevents the T7 loop-associated helix H8 from stacking onto the β-sheet of the intermediate domain, thereby preventing the transition of tubulin from a curved conformation to a straight conformation. This change takes place as the T7 loop, and the helices H7 and H8 adapt to the binding of KY216 (Fig. 1f). We revealed the specific binding site of compound KY216 targeting tubulin, providing a strong structural basis for the next pharmacological mechanism studies.
KY216 inhibits NSCLC progression in vitro and in vivo
KY216 is a potent inhibitor of tubulin polymerization, demonstrating efficacy in inhibiting breast cancer cell proliferation and angiogenesis, while causing G2/M phase arrest and overcoming paclitaxel resistance38. However, its role in NSCLC remains unexplored. The half-maximal inhibitory concentration (IC50) on lung cancer cell lines was approximately 20 nM, which represents a significantly stronger inhibitory effect compared with that observed in breast cancer, colorectal cancer, and osteosarcoma cell lines (Supplementary Fig. 1a–d). Additionally, KY216 exhibited enhanced inhibitory effects on highly invasive cancer cells, including lung cancer cells (95-D and NCI-H1299), breast cancer cells (MDA-MB-231), colorectal cancer cells (HCT116), and osteosarcoma cells (143B) (Supplementary Fig. 1a–d). Additionally, the toxicity of KY216 in normal cell lines (BEAS-2B, MCF 10A, THLE-2) was lower than that of the positive drugs paclitaxel and CA-4 (Supplementary Fig. 1e).
Immunofluorescence analyses revealed that untreated NCI-H460 cells displayed a well-aligned microtubule network, which became disrupted and disordered with increasing concentrations of KY216 (Fig. 2a). In comparison to the positive drugs, KY216 significantly reduced the proliferative capacity of lung cancer cells at a lower concentration, as evidenced by clone formation and EDU assays (Fig. 2b, c and Supplementary Fig. 2a). Furthermore, low concentrations of KY216 induced a G2/M phase blockage in lung cancer cells, leading to decreased levels of proteins associated with the G2/M phase. Compared with the concentrations required for positive control drugs such as paclitaxel and CA-4 to achieve similar effects, KY216 demonstrates superior efficacy (Fig. 2d, e and Supplementary Fig. 2b). Furthermore, the study found that the conditioned medium of KY216-treated NCI-H460 and 95-D cells had a notable impact on lumen formation of human umbilical vein endothelial cells (HUVECs) and decreased vascular endothelial growth factor (VEGF) secretion by NSCLC cells (Fig. 2f, g).
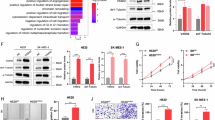
a Effects of KY216 on cellular microtubule networks (independent biological replicates n = 3). NCI-H460 cells were exposed to varying concentrations of KY216 for 48 h. Confocal microscopy was used to capture images of microtubules (labeled green with anti-α-tubulin antibody) and nuclei (stained with DAPI and labeled blue). Scale bar: 20 μm. b The formation of NCI-H460, A549, and 95-D cell colonies was significantly affected by different concentrations of KY216 (independant biological replicates n = 3). c Fluorescent expression of EDU+ cells was observed in NCI-H460 and 95-D cells. The EDU+ cells were labeled with green fluorescence, while the cell nuclei were stained with DAPI to show blue fluorescence (independent biological replicates n = 3). Scale bar: 100 μm. The images were acquired using the ImageXpress Micro® system. d, e Flow cytometry reveals that KY216 induces G2/M cycle arrest in NCI-H460 and 95-D cells (d), with significance indicated by comparison of the G2/M phase proportion in each group. Corresponding Western blot analysis is shown in (e) (independent biological replicates n = 3). f Conditioned medium from NCI-H460 and 95-D cells treated with 0, 10, 20, and 30 nM KY216 inhibited the formation of capillary-like tubular networks in HUVECs. Representative images are shown (independent biological replicates n = 3). Scale bar: 50 μm. g Determination of VEGF expression levels in supernatants of NCI-H460 and 95-D cells treated with 0, 10, 20, and 30 nM KY216 (independent biological replicates n = 3). All quantitative data are shown as means ± SEM. Values shown are P values and were calculated using the one-way ANOVA with Tukey’s comparisons test in GraphPad Prism 9.5.0 (b–g). Source data are provided as a Source Data file.
The anti-tumor effect of KY216 was evaluated in a human lung cancer NCI-H460 xenograft model in nude mice by subcutaneously implanting tumors and administering doses of 5, 10, or 20 mg/kg every other day for 14 days. A positive group received 10 mg/kg paclitaxel. Results demonstrated that KY216 significantly suppressed tumor growth without impacting mouse body weight (Fig. 3a–c and Supplementary Fig. 3a). Immunohistochemistry analysis showed that the proportion of cells staining positive for Ki67, an indicator of cell proliferation, and CD31, an indicator of angiogenesis, was markedly reduced after treatment with KY216 (Fig. 3d). Additionally, histopathological examination (H&E) of the heart, liver, spleen, lungs, and kidneys of mice showed no drug-induced abnormalities, highlighting the safety profile of KY216 (Supplementary Fig. 3b). These findings underscore the efficacy of KY216 in restraining lung cancer progression and disrupting tumor vasculature.
a Schematic of the drug administration regimen and representative images of resected tumors from the NCI-H460 xenograft model treated with KY216 (5, 10, or 20 mg/kg) or paclitaxel (10 mg/kg) (n = 6 mice per group). b The tumor volume in nude mice treated with KY216 and paclitaxel is shown, with significance indicated by comparison of tumor volume at day 14 vs day 0 (n = 6 mice per group). c The body weight of xenografted nude mice following treatment with KY216 and paclitaxel, with significance determined by comparing the body weights on Day 14 and Day 0 (n = 6 mice per group). d Immunohistochemical and haematoxylin–eosin (H&E) analysis were shown (n = 6 mice per group). Scale bar: 50 μm. e, f Invasion and migration (e) of NCI-H460 and 95-D cells (independant biological replicates n = 3) and the corresponding protein blot analysis (f) of mesenchymal markers (independant biological replicates n = 3). Scale bar: 50 μm. g, h KY216 decreased TGF-β-induced migration and invasion (g) of NCI-H460 cells (independant biological replicates n = 3). Protein blot analysis (h) was conducted to evaluate the corresponding mesenchymal markers, ZEB1 and p-Smad2 (independant biological replicates n = 3). Scale bar: 50 μm. i–k Treatment with KY216 reduced the migration and invasion abilities (i) of TGF-β-induced NCI-H460 and 95-D cells (independant biological replicates n = 3) and the protein blotting (independant biological replicates n = 3) of epithelial and mesenchymal markers, ZEB1 and p-smad2, in 95-D (j) and TGF-β-induced NCI-H460 cells (k). Scale bar: 50 μm. l Confocal microscopy images showing E-Cadherin (red, white arrows) and nuclei (DAPI, blue). Scale bar: 20 μm. All data are shown as means ± SEM. Values shown are P values and were calculated using a two-tailed Student’s t-test (b, c, e, f, h) and one-way ANOVA with Tukey’s comparisons test (d, g) in GraphPad Prism 9.5.0. Source data are provided as a Source Data file.
KY216 suppresses invasion and metastasis in highly metastatic lung cancer cells
To investigate the effect of KY216 on NSCLC metastasis, we evaluated the metastatic ability of two NSCLC cell lines, NCI-H460 and 95-D. Based on the number of cells that passed through the porous filter in the trans-well assay, we observed that 95-D cells exhibited higher migratory and invasive properties40, leading to their selection for subsequent investigations (Fig. 3e, f). TGF-β promotes the process of EMT in tumors by increasing the expression of mesenchymal markers such as Vimentin and N-cadherin and reducing the levels of epithelial markers such as E-cadherin41. Following treatment with TGF-β (10 ng/mL), there was an observed enhancement in the migration and invasion capabilities of NCI-H460 cells, along with an increase in mesenchymal marker proteins and p-Smad2 levels. Notably, KY216 was able to effectively reverse these changes by not only inhibiting the migration and invasion abilities of the cells, but also decreasing the expression of mesenchymal proteins and p-Smad2 (Fig. 3g, h). KY216 attenuated TGF-β-induced migration and invasion of NCI-H460 cells and adjusted the expression of related proteins within the concentration range of 0-30 nM. In a similar manner, KY216 inhibited the migration and invasion in 95-D cells with high metastatic potential, leading to decreased levels of Vimentin, N-cadherin, and ZEB1, while simultaneously increasing the expression of E-cadherin (Fig. 3i–k and Supplementary Fig. 3c–f). KY216 also attenuated TGF-β-induced wound healing in NCI-H460 cells (Supplementary Fig. 4a). Furthermore, KY216 demonstrated a more pronounced inhibitory effect on NSCLC migration and invasion than paclitaxel and CA-4 (Supplementary Fig. 4b–e). E-cadherin plays a crucial role in intercellular adhesion, and its loss can facilitate cancer cell metastasis, making it a key marker for EMT42. Immunofluorescence results showed that treatment with 20 nM KY216 increased E-cadherin fluorescence at the H460 cell membrane borders (highlighted by white arrows), enhancing intercellular adhesion and reducing metastatic potential (Fig. 3l). Overall, these findings strongly support the inhibitory effects of KY216 on migration and invasion in highly metastatic lung cancer cells.
KY216 inhibits EMT by reducing VASH2
To further elucidate the mechanism of action of KY216, we collected lung cancer cells treated with or without KY216 and performed RNA-seq analysis to identify differentially expressed genes. In a previous study, we have shown that KY216 has the ability to disrupt tumor angiogenesis and inhibit the metastasis of lung cancer cells. Among the genes displaying substantial differential expression, in addition to the well-recognized metastasis-promoting factor ZEB1, we specifically observed a pronounced down-regulation of VASH2 (Fig. 4a). VASH2 has been documented as a pivotal promoter in the processes of angiogenesis and cell migration17,19. We found that KY216 decreased VASH2 levels in a concentration-dependent manner (Supplementary Fig. 5a).
a Volcano plot displaying differentially expressed miRNAs following KY216 treatment. Significantly downregulated and upregulated miRNAs are highlighted in blue and red, respectively; non-significant miRNAs are shown in gray. The p-values were calculated using a two-tailed Student’s t-test and were subsequently adjusted for the false discovery rate (FDR) using the Benjamini–Hochberg method. b Conditioned medium from NCI-H460 and 95-D cells transfected with siVASH2 inhibited the formation of capillary-like tubular networks in HUVECs. Representative images are shown (independant biological replicates n = 3). Scale bar: 50 μm. c–e Transfected with siVASH2 reduced the migration and invasion abilities (c) of TGF-β-induced NCI-H460 and 95-D cells (independant biological replicates n = 3) and the protein blotting analysis (independant biological replicates n = 3) of epithelial and mesenchymal markers, ZEB1 and p-smad2, in TGF-β-induced NCI-H460 (d) and 95-D cells (e). Scale bar: 50 μm. f, g Overexpression of VASH2 increased the migration and invasion abilities (f) of NCI-H460 and 95-D cells (independant biological replicates n = 3) and the protein blotting analysis (independant biological replicates n = 3) of epithelial and mesenchymal markers and ZEB1 in NCI-H460 and 95-D cells (g). Scale bar: 50 μm. h–j Overexpression of VASH2 reversed the inhibitory effect of KY216 on migration and invasion (h) of TGF-β-induced NCI-H460 and 95-D cells (independant biological replicates n = 3) and the protein blotting analysis (independant biological replicates n = 3) of epithelial and mesenchymal markers, ZEB1 and p-smad2, in TGF-β-induced NCI-H460 (i) and 95-D cells (j). Scale bar: 50 μm. All data are shown as means ± SEM. Values shown are P values and were calculated using a two-tailed Student’s t-test (f, i) and one-way ANOVA with Tukey’s comparisons test (b, h) in GraphPad Prism 9.5.0. Source data are provided as a Source Data file.
The study investigated the biological function of VASH2 and its role in inhibiting EMT by KY216 through transfecting lung cancer cells with small interfering RNAs (siRNAs) and overexpression plasmids of VASH2. Three sets of siRNAs were designed to effectively reduce VASH2 gene expression (Supplementary Fig. 5b). Results showed that knockdown of VASH2 notably decreased the angiogenic capacity of both NCI-H460 and 95-D cells (Fig. 4b), as well as reduced the levels of VEGF (Supplementary Fig. 5c). Conversely, overexpression of VASH2 markedly enhanced the angiogenic capacity and VEGF secretion in both cell lines (Supplementary Fig. 5d, e). Migration and invasion assays demonstrated that reducing VASH2 expression observably decreased the migration and invasion capabilities of 95-D cells and TGF-β-treated H460 cells (Fig. 4c and Supplementary Fig. 5f). Western blot analysis demonstrated that VASH2 knockdown led to elevated levels of E-cadherin in TGF-β-induced H460 cells, accompanied by decreased levels of vimentin, N-cadherin, ZEB1, and p-Smad2 (Fig. 4d). Furthermore, the expression of epithelial markers was enhanced in 95-D cells, while the mesenchymal markers exhibited a decrease (Fig. 4e). Conversely, excessive expression of VASH2 notably enhanced the migration and invasion abilities of 95-D and NCI-H460 cells (Fig. 4f). Additionally, these cells exhibited higher levels of mesenchymal markers and ZEB1, along with reduced levels of E-cadherin (Fig. 4g). VASH2 overexpression remarkably reversed the inhibitory effect of KY216 on the migration and invasiveness of TGF-β-induced H460 and 95-D cells (Fig. 4h). In addition, VASH2 overexpression also counteracted the decrease in mesenchymal markers’ levels induced by KY216 and reduced the levels of epithelial markers that were increased by KY216 (Fig. 4i, j and Supplementary Fig. 5g). More importantly, the knockdown of VASH2 significantly enhanced the inhibitory effects of KY216 on TGF-β-induced migration and invasion, markedly suppressed the expression of the mesenchymal marker, ZEB1, and promoted the expression of E-cadherin (Supplementary Fig. 6a, b). These findings provide strong evidence supporting the crucial role of VASH2 in inhibiting the metastasis of NSCLC by KY216.
KY216 facilitates the binding of VASH2 to α-tubulin, consequently promoting tubulin detyrosination
VASH2 functions as a pivotal tubulin tyrosine carboxypeptidase in the post-translational modification of α-tubulin. The catalytic triad Cys158-His193-Ser210 in VASH2 is essential for the detyrosination of the C-terminal tyrosine on α-tubulin24,25. Considering KY216’s ability to decrease VASH2 expression, we explored its potential as an inhibitor of tubulin detyrosination. However, Western blotting and immunofluorescence experiments revealed that KY216 actually led to increased detyrosination of tubulin in NCI-H460 and 95-D cells (Fig. 5a, b and Supplementary Fig. 7a). This intriguing phenomenon prompted further investigation. VASH2 interacts electrostatically with the carboxy-terminal tyrosine residues of α-tubulin via its substrate-binding groove, which comprises Ser210 and a series of nearby positively charged basic amino acids27,43,44. To further explore this interaction, we employed an immunoprecipitation (IP) assay to study the binding between VASH2 and α-tubulin. Excitingly, KY216 obviously increased the interaction between VASH2 and α-tubulin (Fig. 5c and Supplementary Fig. 7b). A more precise protein-protein interaction technique, Duolink in situ proximity ligation assay (PLA), was used to study the interaction between VASH2 and α-tubulin45. The presence of KY216 resulted in a significant increase in PLA fluorescence dots, indicating an enhanced interaction between VASH2 and α-tubulin (Fig. 5d). Furthermore, both α-tubulin and detyr-tubulin co-precipitated with VASH2 showed a significant increase upon KY216 treatment, thereby confirming the binding capacity of VASH2 to α-tubulin in the presence of KY216 and supporting the detyrosination of tubulins (Fig. 5e).
a deTyr-tub protein blot analysis in NCI-H460 and 95-D cells (independant biological replicates n = 3). b Immunofluorescence analysis of deTyr-tub (green), cell nuclei were stained with DAPI (blue), and images were acquired by the confocal microscopy system (independant biological replicates n = 3). Scale bar: 20 μm. Data are shown as means ± SEM. Values shown are P values and were calculated using the one-way ANOVA with Tukey’s comparisons test. c Co-Immunoprecipitation (Co-IP) showed that the binding of VASH2 to α-tubulin in H460 cells was enhanced by KY216 (20 nM) (independant biological replicates n = 3). d The interactions between VASH2 and α-tubulin under KY216 treatment were detected using the PLA. Representative fluorescence images are shown (independant biological replicates n = 3). Scale bar: 20 μm. e Co-IP results showed that KY216 (20 nM) promoted the interaction of VASH2 with α-tubulin and enhanced the process of α-tubulin detyrosination (independant biological replicates n = 3). f Schematic depiction of VASH2. The Cys-His-Ser catalytic triad is highlighted in yellow circles. g Protein blotting was used to analyze changes in the expression of ZEB1 and deTyr-tub following transfection of VASH2 wild-type and mutant plasmids in 95-D cells (independant biological replicates n = 3). h Changes in the interaction of VASH2 (eGFP-tagged) with α-tubulin after transfection of VASH2 wild-type and mutant plasmids in 95-D cells were investigated by Co-IP analysis (independant biological replicates n = 3). i Protein blotting was utilized to detect alterations in the levels of ZEB1 and deTyr-tub in 95-D cells following transfection of VASH2-WT and VASH2-S210A mutant plasmids in the presence of 20 nM KY216 (independant biological replicates n = 3). j Co-IP assay was used to study changes in the interaction between VASH2 (eGFP-tagged) and α-tubulin after transfection of VASH2-WT and VASH2-S210A mutant plasmids in the presence of 20 nM KY216 in 95-D cells (independant biological replicates n = 3). Source data are provided as a Source Data file.
Isothermal titration calorimetry (ITC) experiments further demonstrated KY216’s modulatory effect on the VASH2-tubulin interaction. Ligand titration experiments showed specific high-affinity binding of KY216 to tubulin (Kd = 0.15 ± 0.01 μM), with no significant binding signal observed between KY216 and VASH2 (Supplementary Fig. 7c, d). Subsequent experiments compared thermodynamic parameter changes in VASH2-tubulin binding within a system maintaining KY216 concentration balance (synchronously loaded into both the syringe and the cell). In the absence of KY216, the Kd for VASH2-tubulin binding was 1.53 ± 0.22 μM (ΔG = −33.27 ± 0.35 kJ/mol); in the presence of KY216, the binding affinity increased about 2.9-fold to Kd = 0.52 ± 0.10 μM, accompanied by a more significant decrease in Gibbs free energy (ΔG = −35.97 ± 0.50 kJ/mol). The increased binding affinity and reduced binding free energy (ΔΔG = −2.70 kJ/mol) confirmed that KY216 enhances VASH2-tubulin interaction and stabilizes the complex (Supplementary Fig. 7e, d and Supplementary Table 1).
We investigated whether other MTAs exhibited similar effects. Paclitaxel was found to inhibit VASH2 expression, promote the detyrosination of α-tubulin, and enhance the interaction between VASH2 and α-tubulin (Supplementary Fig. 8a, b). In contrast, CA-4 significantly reduced VASH2 levels, decreased detyrosination of α-tubulin, and had minimal impact on the interaction between VASH2 and α-tubulin (Supplementary Fig. 8c, d). Given the functional differences between CA-4 and KY216, we further analyzed the binding characteristics of CA-4 using ITC (Supplementary Fig. S8e). No binding interaction was detected between CA-4 and VASH2. The binding affinity of CA-4 to tubulin (dissociation constant, Kd = 0.4 ± 0.05 μM) was lower than that of KY216, indicating that the latter exhibits a higher binding propensity for tubulin (Supplementary Fig. 8f and Supplementary Table 1). In the presence of CA-4, the binding affinity between VASH2 and tubulin (Kd = 1.49 ± 0.08 μM) showed no significant difference compared to the baseline level (Kd = 1.53 ± 0.22 μM) (Supplementary Fig. 8g and Supplementary Table 1). In contrast, when KY216 was present, this binding affinity was enhanced, with the Kd value decreasing to 0.52 ± 0.10 μM (Supplementary table S1). These results suggest that, in comparison to CA-4, KY216 can induce a conformational change in tubulin that is more favorable for VASH2 binding. The aforementioned findings demonstrate that although MTAs can all effectively reduce VASH2 expression, they may exert distinct regulatory effects on α-tubulin detyrosination and the VASH2-tubulin interaction, likely due to differences in their chemical structures.
KY216 inhibits EMT by promoting VASH2 binding to α-tubulin
Based on crystal structure analysis combined with co-IP and ITC experimental results, we revealed that KY216 significantly enhances the interaction between VASH2 and tubulin, and promotes the detyrosination of α-tubulin. Notably, how this mechanism influences the EMT process in NSCLC warrants further investigation. Previous research has indicated that VASH2 plays a role in facilitating the EMT process through its regulation of the EMT transcription factors ZEB1/221. We further hypothesize that KY216-mediated VASH2-tubulin binding might alter VASH2’s intracellular localization and functional state, consequently affecting tumor cell migration and invasion capabilities. KY216 treatment significantly promoted tubulin detyrosination while suppressing VASH2 and ZEB1 expression. Conversely, VASH2 knockdown inhibited both tubulin detyrosination and ZEB1 expression. Intriguingly, KY216 reversed the inhibitory effect of siVASH2 on tubulin detyrosination and synergistically enhanced ZEB1 suppression with siVASH2 (Supplementary Fig. 9a). In VASH2-overexpressing models, KY216 synergistically elevated tubulin detyrosination levels with overexpressed VASH2 and effectively counteracted the aberrant ZEB1 upregulation caused by VASH2 overexpression (Supplementary Fig. 9b), indicating a close correlation between tubulin detyrosination and ZEB1 expression under KY216 regulation.
To further clarify the temporal sequence of KY216 binding, VASH2 recruitment, tubulin detyrosination, and changes in ZEB1 expression, we conducted a time-course analysis. The results showed that the optimal duration of action for 20 nM KY216 was 48 h (at this time, cell viability decreased to 50%) (Supplementary Fig. 9c, d). After 24 h of KY216 treatment, an enhanced interaction between VASH2 and α-tubulin was observed, indicating that VASH2 began to be recruited to the complex; at this point, the expression of VASH2 in the Input samples was slightly decreased, but the changes in ZEB1 expression and the level of detyrosinated tubulin were not significant (Supplementary Fig. 9e, f). After extending the KY216 treatment to 48 h, the interaction between VASH2 and α-tubulin was further enhanced, more detyrosinated tubulin was detected in the precipitates, and an increase in detyrosinated tubulin and a significant decrease in ZEB1 expression were also observed in the Input samples; the above effects peaked at 72 h (Supplementary Fig. 9e, f). The above results suggest that under the regulation of KY216, VASH2 binding, tubulin detyrosination, and ZEB1 expression are closely associated.
To further elucidate this expression relationship, we developed a VASH2 plasmid containing an eGFP-tagged variant with a catalytic triplex site mutation known as VASH2 (C158A/H193A/S210A) (Fig. 5f), which is known to hinder VASH2 detyrosination24,25,26,27. Small vasohibin binding protein (SVBP) stabilizes VASH2 and plays a chaperone-like role by enhancing VASH2 solubility25,27. The VASH2/SVBP complex demonstrates higher and more targeted detyrosination activity compared to VASH2 alone24,25,26,27. We created an SVBP expression plasmid with a Flag tag and introduced it into cells as a chaperone factor. Mutation of all three binding sites resulted in a decrease in detyrosination in 95-D cells (Fig. 5g). Immunoprecipitation experiments indicated that mutations in the binding sites significantly weakened the interaction between VASH2 and α-tubulin. Among the mutations, those at the S210 site caused the most significant decrease (Fig. 5h). Interestingly, even with a mutation in the binding site of VASH2 to α-tubulin, its ability to up-regulate ZEB1 remained unaffected (Fig. 5g). Treatment with KY216 was found to block the activation of ZEB1 by VASH2, consequently reducing the migration and invasion capabilities of NCSLC cells. However, this inhibitory effect was reversed when the S210 site of VASH2 was mutated. Moreover, mutating the S210 locus impaired KY216’s capacity to enhance the detyrosination of tubulin (Fig. 5i and Supplementary Fig. 10a). Co-IP experiments revealed the key mechanistic details: mutation at the S210 site impaired the facilitation of VASH2 binding to α-tubulin by KY216 (Fig. 5j), hindered the detyrosination activity of VASH2, and thus enhanced its activating effect on ZEB1. To determine whether the aforementioned mechanism is specific to KY216, CA-4 was employed as a control. Experimental results revealed that CA-4 had no effect on the interaction between VASH2 and tubulin, regardless of whether the S210 mutation was present or not (Supplementary Fig. 10b). Furthermore, treatment with CA-4 alone was found to inhibit α-tubulin detyrosination, and this inhibitory effect remained unaffected following the S210 mutation (Supplementary Fig. 10b). In conclusion, KY216 effectively reduces the activity of VASH2 in promoting EMT by recruiting VASH2, enhancing its interaction with α-tubulin, promoting tubulin detyrosination, and inhibiting ZEB1 expression.
KY216 inhibits EMT, at least partially, through the miR-429/VASH2/ZEB1 axis
KY216 not only reduced the protein levels of VASH2 and ZEB1, but also markedly decreased their mRNA levels (Supplementary Fig. 11a). We wanted to explore whether KY216 regulates VASH2 gene levels through miRNAs. Through miRNA-seq, we identified obvious alterations in miRNA expression profiles following KY216 treatment of NCI-H460 cells (Fig. 6a). KEGG and GO enrichment analyses indicated that the differentially expressed miRNAs were notably enriched in the TGF-β signaling pathway and were involved in the EMT process (Supplementary Fig. 11b, c). Among the various differentially expressed miRNAs, we focused on miR-663b, miR-5699-3p, and miR-429, which are associated with EMT (Fig. 6b). The qRT-PCR assay confirmed that miR-429 expression was significantly upregulated after KY216 treatment compared with miR-663b and miR-5699-3p (Fig. 6c). Notably, KY216 markedly enhanced the expression of miR-429 in TGF-β-induced H460 cells (Fig. 6d). Furthermore, Kaplan–Meier survival analysis demonstrated that high expression levels of miR-429 were positively correlated with the OS of patients with NSCLC (Supplementary Fig. 11d), suggesting that miR-429 may serve as a potential therapeutic target for KY216 in inhibiting the EMT process of NSCLC.
a The heatmap illustrates the significant differential expression of miRNAs in H460 cells following KY216 treatment. b Volcano plot of differentially expressed miRNAs in H460 cells after KY216 treatment. The P-values were calculated using a two-tailed Student’s t-test and were subsequently adjusted for the FDR using the Benjamini–Hochberg method. c Expression levels of miR-663b, miR-5699-3p, and miR-429 in NCI-H460 cells following 20 nM KY216 treatment, as determined by qRT-PCR (independant biological replicates n = 3). d qRT-PCR was used to detect changes in miR-429 gene levels in TGF-β-induced NCI-H460 cells under KY216 treatment (independant biological replicates n = 3). e Schematic representation of five positions of miR-429 binding with the 3’ UTR of VASH2 mRNA. f 293T cells were co-transfected with NC, miR-429, pmiR-GLO-VASH2-WT-C/D, and pmiR-GLO-VASH2-Mut-C/D, the value of firefly luciferase (F-luc) was normalized to that of the renilla luciferase (R-luc) (independant biological replicates n = 4). g, h Transwell assays (g) were used to investigate the effects of transfection with miR-429 mimics or inhibitors, as well as their corresponding negative controls, on the migration and invasion of TGF-β-induced H460 and 95-D cells (independant biological replicates n = 3). Scale bar: 50 μm. Protein blotting (h) was performed to assess the expression of epithelial/mesenchymal markers, p-Smad2, and ZEB1 (independant biological replicates n = 3). i, j Transfection with miR-429 inhibitors reversed the inhibitory effect of KY216 on migration and invasion (i) of TGF-β-induced NCI-H460 and 95-D cells (independant biological replicates n = 3) and the protein blotting analysis (independant biological replicates n = 3) of epithelial and mesenchymal markers, ZEB1 and p-smad2, in TGF-β-induced NCI-H460 and 95-D cells (j). Scale bar: 50 μm. All data are shown as means ± SEM. Values shown are P values and were calculated using two-tailed Student’s t-test (c) and one-way ANOVA with Tukey’s comparisons test (d, f) in GraphPad Prism 9.5.0. Source data are provided as a Source Data file.
MiR-429, a member of the miR-200 family, typically targets ZEB1 and ZEB2 to inhibit E-cadherin, thereby maintaining the epithelial phenotype and suppressing EMT46. We then assessed the effect of KY216 on the expression of members of the miR-200 family and found that miR-429 was the most remarkably upregulated (Supplementary Fig. 12a). In contrast, paclitaxel and CA-4 were more likely to increase the level of miR-141-3p, indicating the specificity of KY216 in promoting miR-429 expression (Supplementary Fig. 12b, c). More importantly, through in-depth analysis of miRNA and mRNA sequencing data, we have revealed a significant negative correlation between miR-429 and VASH2. This finding suggests that miR-429 may be involved in the regulatory mechanism of VASH2 (Supplementary Fig. 12d, e). Overexpression of miR-429 led to a significant down-regulation of VASH2 and ZEB1 mRNA levels, indicating a potential regulatory relationship between miR-429 and VASH2 transcript (Supplementary Fig. 12f). We predicted five potential miR-429-binding sites in the VASH2 3’ UTR using the miRDB47, TargetScan48, and starBase49 databases (Fig. 6e). The A/C/D sites contain eight base complements, while the B/E sites only have seven. To study the functions of these binding sites, mutations were made in the A/C/D sites (CAGUAUUA to GUCAUAAU) (Fig. 6e). Two pairs of plasmids were created due to the length of the VASH2 gene’s 3’ UTR sequence and the distance between the loci: pmiRGLO-WT-A, pmiRGLO-mut-A, and pmiRGLO-WT-C + D, pmiRGLO-Mut-C, pmiRGLO-Mut-D, and pmiRGLO-Mut-C + D. The results from the dual luciferase reporter gene assay revealed that miR-429 overexpression notably reduced the activity of the wild-type plasmid at the C/D site. The three mutant plasmids at the C/D site were able to reverse the decrease in luciferase activity induced by miR-429 (Fig. 6f). However, the activity of the wild-type reporter gene at the A site was not affected by miR-429 overexpression (Supplementary Fig. 12g). Our experimental results fully demonstrated that VASH2 is a target gene of miR-429 and identified the C/D site as the most important binding site. Next, we conducted an in-depth study of the role that miR-429 fills in the pharmacological mechanism of KY216.
The migration and invasion ability of TGF-β-treated H460 and 95-D cells were distinctly restrained by the overexpression of miR-429, whereas miR-429 inhibitors led to the opposite trend (Fig. 6g and Supplementary Fig. 13a). Overexpression of miR-429 improved the expression of epithelial markers and lessened the levels of mesenchymal markers, ZEB1, and p-Smad2. On the other hand, transfection with the miR-429 inhibitor down-regulated the expression of epithelial markers and up-regulated the levels of mesenchymal markers (Fig. 6h and Supplementary Fig. 13b). We used the miR-429 inhibitor to counteract the KY216-induced elevation of miR-429. Migration and invasion assays results showed that the miR-429 inhibitor was effective in reversing the inhibitory effect of KY216 on the migration and invasion of TGF-β-induced NCI-H460 and 95-D cells (Fig. 6i and Supplementary Fig. 13c). Furthermore, inhibitors of miR-429 also reversed the expression of mesenchymal markers and epithelial markers in the presence of KY216 (Fig. 6j and Supplementary Fig. 13d). In addition, the overexpression of miR-429 exhibited a synergistic inhibitory effect in conjunction with KY216, together inhibiting VASH2 expression, suppressing mesenchymal markers, and promoting epithelial markers, thereby impeding the EMT process of NSCLC (Supplementary Fig. 14a, b). These findings indicate that KY216 suppresses NSCLC migration and invasion partly through the miR-429/VASH2/ZEB1 axis. Notably, integrated bioinformatic analysis of miR-429 target prediction data and KY216-induced differentially expressed mRNA profiles identified 20 potential miR-429 target genes involved in the regulation of the EMT process (Supplementary Fig. 15). Among these, SEMA6D, ZNF217, and CREB5 have been confirmed to promote EMT in breast, prostate, and HCCs, respectively, while MAP3K1 is linked to malignant EMT transformation in NSCLC50,51,52,53. These findings suggest that KY216 might inhibit EMT not only through the core regulatory axis but also via a multi-target synergistic regulatory network mediated by miR-429. This multi-target regulatory mechanism warrants further experimental validation.
The inhibitory effect of KY216 on NSCLC metastasis in vivo is dependent on VASH2
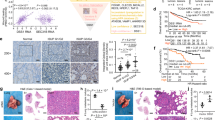
To examine the impact of VASH2 on NSCLC metastasis, we conducted immunofluorescence analysis on 75 lung cancer patient samples. These samples consisted of 35 lymph node metastases and 40 primary tumors. In the fluorescence analysis of tumor tissues, we observed a notable correlation between the green fluorescence intensity of VASH2 and the red fluorescence intensity of N-cadherin. Specifically, in patient samples exhibiting lymph node metastasis (e.g., Patient 15), the expression levels of both VASH2 and N-cadherin were markedly increased. Conversely, in patient samples without lymph node metastasis and with less malignant tumors (e.g., Patient 36), the expression levels of both VASH2 and N-cadherin were observably decreased. This suggests that the coexpression pattern of VASH2 and N-cadherin may be closely related to the invasiveness and metastatic potential of the patients’ tumors (Fig. 7a). Our findings revealed a significantly positive correlation between VASH2 expression and N-cadherin levels, a marker of EMT (Fig. 7b). This association was notably stronger in samples with lymph node metastasis (Fig. 7c). Importantly, patients exhibiting high levels of VASH2 expression experienced shorter survival times (Fig. 7d), suggesting a potential functional and clinical relevance of VASH2 in the regulation of lung cancer metastasis.
a Immunofluorescence staining was used to analyze the association of VASH2 (green) with N-cadherin (red) in patient tissues. The images presented are representative results of selected patient tissues. Scale bar: 50 μm. b, c Correlation analysis was conducted on the expression of VASH2 and N-cadherin in NSCLC patient tissues (b) (n = 75) and in tissue samples from patients with lymph node metastasis (c) (n = 35). Pearson’s correlation coefficient was used for linear correlation analysis. d Survival analysis of VASH2 in 75 tumor tissues. OS was evaluated by Kaplan–Meier plots and the log-rank (Mantel–Cox) test. e–g Bioluminescence imaging was utilized to demonstrate in vivo metastasis in mice (n = 8 mice per group) (e), along with strategy maps (f) detailing the intracardiac injection of 95-D-Luc cells and treatment with AAV-OE-VASH2 and KY216. Quantitative analysis of brain and bone metastasis in mice was conducted using Tanon software (n = 8 mice per group) (g). Data are shown as means ± SEM. Values shown are P values and were calculated using the one-way ANOVA with Tukey’s comparisons test in GraphPad Prism 9.5.0. h, i Representative H&E-stained images (n = 8 mice per group) of brain metastasis (h), scale bar: 50 μm, and bone metastasis (i), scale bar: 200 μm. The formation of metastasis is indicated by black arrows and black dashed boxes. Source data are provided as a Source Data file.
A lung cancer metastasis model was established in nude mice by intracardiac injection of 95-D-luc lung cancer cells. The following day, systemic recombinant adeno-associated virus (AAV) was administered to induce overexpression of VASH2. Subsequent treatment with KY216 (20 mg/kg) was given every two days. In vivo imaging showed the increased brain and bone metastasis in the AAV-OE-VASH2 group compared with the control group, but KY216 administration reduced bioluminescent signals in these tissues. However, the inhibitory effect of KY216 was weakened by VASH2 overexpression (Fig. 7e–g). Histological analysis with H&E staining confirmed that VASH2 overexpression augmented the number of metastatic foci in the brain and bone. Although KY216 treatment reduced these foci, its impact was negated by VASH2 overexpression (Fig. 7h, i). These findings demonstrate that overexpression of VASH2 promotes the metastasis of NSCLC in vivo and mitigates the suppressive effect of KY216 on the metastasis.
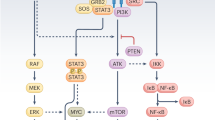
In summary, our results suggest that KY216 binds to tubulin and forms a complex that disrupts the microtubule structure, thereby blocking NSCLC cells at the G2/M cycle and effectively inhibiting their proliferation. More importantly, the KY216-tubulin complex enhances the interaction of VASH2 with α-tubulin to promote the latter detyrosination, inhibits the function of ZEB1, and effectively suppresses the metastasis of NSCLC. Additionally, these effects were partially enhanced by the KY216-induced miR-429 (Fig. 8).
The KY216-tubulin complex disrupts the structure of microtubules, enhances the interaction of VASH2 with α-tubulin to promote the latter detyrosination, inhibits the function of ZEB1, and effectively suppresses the metastasis of NSCLC. These effects were enhanced by the KY216-induced miR-429.
Discussion
KY216 inhibits microtubule assembly by occupying the microtubule colchicine site and effectively preventing the conversion of the αβ-heterodimer to the active straight conformation. Compared to its predecessor DPT, KY216 has been structurally modified to reduce toxicity and improve anti-cancer efficacy38. The lead compound podophyllotoxin binds to tubulin through hydrophobic interactions and lacks hydrogen bonding stabilization54. However, our study shows that KY216 forms four hydrogen bonds and a halogenated bond in addition to hydrophobic interactions with β-tubulin, resulting in enhanced binding stability and superior efficacy. This detailed information on the crystal structure provides valuable insight for designing and developing anti-cancer agents that target the colchicine site of the microtubule protein.
MTAs represent a significant approach in cancer treatment, as they bind to microtubules or free tubulin and modulate their dynamic instability, thereby inducing cell cycle arrest or apoptosis5,6,7. Despite the widespread clinical use of MTAs such as paclitaxel and vincristine, their physicochemical properties, bioavailability, toxicities, and the development of multidrug resistance (MDR) have imposed limitations on their therapeutic efficacy55. Cancer cells exhibit overexpression of P-glycoprotein (P-gp), which mediates the efflux of chemotherapeutics such as paclitaxel and vincristine, ultimately leading to the development of MDR56. Colchicine has been shown to overcome paclitaxel and vincristine resistance by directly binding to and inhibiting P-gp function57. As a member of the colchicine-binding site inhibitors, KY216 may also exert its resistance-reversing effects through binding to P-gp. Additionally, drug resistance is associated with expression alterations or specific mutations in β-tubulin58. For instance, the substitution of serine with arginine at position 277 of β-tubulin induces steric repulsion between βArg277 and the adjacent βArg278, thereby altering the conformation of the tubulin M-loop. This conformational change precludes paclitaxel binding to the M-loop, thereby conferring drug resistance59. Crystal structure analysis reveals that KY216 forms a halogen bond with βVal236 via the chlorine atom on its trisubstituted phenyl, while its carbonyl establishes hydrogen bonds with βAsp249 and βAla248. The dioxane modification enables additional hydrogen bonding with βVal313. Notably, these interactions circumvent known paclitaxel resistance-associated mutation sites, which may underlie KY216’s capacity to overcome paclitaxel resistance. Conversely, studies have demonstrated that β-tubulin mutations at positions A248T and M257V attenuate colchicine binding, representing a mechanism of resistance to colchicine-class drugs60. Although the βAla248 mutation may potentially impact KY216 binding, KY216 retains overall binding capability through multiple hydrogen and halogen bonds formed with βAsp249, βVal313, and βVal236. Consequently, KY216 may exhibit diverse binding sites and reduced propensity for resistance development compared with colchicine. Our investigations confirm that KY216 demonstrates significant efficacy in NSCLC, potently inhibiting cell proliferation, inducing cell cycle arrest, and suppressing tumor growth in xenograft models. Notably, KY216 illuminates lower toxicity and enhanced therapeutic efficacy compared to paclitaxel. Collectively, KY216’s high efficacy, low toxicity, and potential to overcome paclitaxel resistance position it as a promising candidate for NSCLC therapy. However, the resistance profile of this compound in NSCLC still needs to be clarified through systematic in vitro and in vivo studies. In addition, our results indicate that KY216 not only strongly inhibits the proliferation and cycle progression of lung cancer cells but also notably reduces NSCLC metastasis through the VASH2-mediated signaling pathway. This highlights a perspective on the application of microtubule inhibitors in cancer therapy.
The metastatic spread of NSCLC is the primary cause of mortality in affected patients, and the process of metastasis involves intricate molecular mechanisms61. Among the various signaling pathways that regulate NSCLC metastasis, the transforming growth factor-β (TGF-β) signaling pathway is particularly pivotal62. Following binding to its type II and type I transmembrane kinase receptors (TGFβR2 and TGFβR1) on the cell membrane, TGF-β initiates the phosphorylation of Smad2 and Smad3 proteins. The phosphorylated Smad2/3 then forms a complex with Smad4 and translocates into the nucleus to modulate the expression of EMT-related genes, including Snail, Slug, ZEB1, ZEB2, and Twist, thereby inducing the EMT process41. Various reports have shown that non-coding RNAs and cytokines interfere with the TGF-β pathway, affecting EMT in lung cancer. Natural compound HpA inhibits TGF-β-induced EMT in A549 cells through the NF-κB pathway63. PTBP3 gene knockout diminishes Smad2 and Smad3 expression and phosphorylation levels in lung cancer cells, suppressing invasion and metastasis of lung adenocarcinoma64. In lung adenocarcinoma cells, miR-454-3p downregulates TGFβR2 expression, consequently inhibiting EMT65. In this study, low-invasive H460 cells were treated with TGF-β1 to enhance metastatic ability. It was observed that KY216 could inhibit the TGF-β signaling pathway partially via the miR-429/VASH2 axis. Confirmation of this effect was obtained through the detection of p-Smad2 protein and EMT markers. KY216 was found to directly suppress the metastatic potential of highly invasive 95-D cells, with the absence of p-Smad2 protein in 95-D cells when TGF-β activation was not present. Both cell types demonstrated that KY216 effectively inhibited NSCLC metastasis, at least in part, through modulation of the miR-429/VASH2/ZEB1 pathway. KY216 may inhibit EMT through the synergistic action of multiple targets. However, our current research has limitations in the functional verification of multiple targets. In the KY216/miR-429 regulatory network constructed based on bioinformatic analysis, the molecular functions of genes such as SEMA6D, ZNF217, CREB5, and MAP3K1 urgently need to be verified. This provides an important research direction for comprehensively analyzing the multi-dimensional action mechanism of this regulatory axis.
Post-translational modifications of tubulin involve chemical alterations to tubulin heterodimers or free tubulins. One specific modification is the cyclic removal and re-addition of tyrosine to the C-terminus of α-tubulin, a process known as the de-tyrosine/tyrosine loop66. This cycle regulates the polymerization of α- and β-tubulin, with functional defects of CENP-E and MCAK resulting from an imbalance in the cycle leading to chromosome instability, promoting tumor metastasis and contributing to brain diseases and heart failure9,67. Furthermore, detyrosination of tubulin stabilizes microtubules, causing their accumulation in prostate and breast cancer cells9,68. KY216 promotes α-tubulin detyrosination, which appears to contribute to lung cancer growth. However, our findings indicate that in the presence of KY216, the compound binds to α- and β-tubulin dimers, thereby hindering the transition from the curved conformation to the straight conformation, which dramatically impedes microtubule assembly. Based on ITC experimental evidence, KY216 is capable of stabilizing the VASH2–tubulin complex, and compared with CA-4, VASH2 exhibits a higher preference for recognizing the KY216-tubulin complex. In systems where microtubule assembly is impaired, VASH2 can still specifically recognize tubulin bound to KY216 and catalyze de-tyrosination. However, since tubulin dimers are unable to form the basic microtubule structure, this modification cannot be translated into a microtubule-stabilizing effect. Overall, KY216 not only exerts its pharmacological effects by obstructing microtubule assembly but also inhibits cell migration by enhancing the binding of VASH2 to α-tubulin and reducing VASH2’s promoting effect on ZEB1. Notably, although CA-4, paclitaxel, and KY216 all belong to MTAs and can inhibit VASH2, they exhibit differences in their regulatory effects on detyrosination: CA-4 has no impact on the interaction between VASH2 and tubulin; paclitaxel can promote this interaction and increase detyrosination, yet whether its regulatory effect on ZEB1 is consistent with that of KY216 requires further investigation. In conclusion, our study reveals the specific mechanism of action of KY216 in regulating tubulin detyrosination and ZEB1 expression; however, due to differences in compound structures, other MTAs may exert their effects through distinct mechanisms, which await further investigation.
Interestingly, the role of miR-429 in various cancers is intricate and diverse. For instance, in breast cancer, Zhang discovered that miR-429 hindered the Wnt/β-catenin pathway by suppressing FN1, leading to reduced proliferation and migration of MDA-MB-231 cells69. Conversely, Claudia observed that miR-429 actually boosted proliferation and migration in HER2 + BC SKBR-3 cells70. In NSCLC, a study revealed that miR-429 facilitated the proliferation and migration of A549 cells71, and elevated levels were detected in NSCLC patient serum72, contradicting previous reports31,32,33,34,35,36,37. Our research demonstrated that miR-429 inhibited EMT by suppressing VASH2 and ZEB1 expression in NCI-H460 and 95-D cells. The Kaplan–Meier analysis indicated an improved survival in lung cancer patients with higher miR-429 levels. These discrepancies underscore the need for further studies to clarify the precise role of miR-429. Additionally, our study observed the upregulation of miR-429 by KY216, as well as the upregulation of the miR-200 family by microtubule-targeting drugs. The mechanisms underlying the specific upregulation of miRNAs by different drugs warrant further in-depth investigation in future research.
Methods
Protein purification and crystallization
RB3-SLD (stathmin 4, Mus musculus) and tubulin tyrosine ligase (TTL, Gallus gallus) proteins were expressed using pET28a-6His and pET22b-6His vectors, and protein expression was induced in Escherichia coli BL21 (DE3). The gene fragment encoding 6His-RB3-SLD_49-189 was synthesized with NdeI and XhoI restriction sites introduced at the 5’ and 3’ ends, respectively. Subsequently, the synthesized gene fragment and the pET-28a(+) vector were subjected to double digestion with NdeI and XhoI. The digestion reaction was carried out at 37 °C for 2 h. The digested vector and gene fragment were separated by agarose gel electrophoresis, and the target bands were excised and purified using a DNA purification kit (D0056, Beyotime, China). The purified vector and gene fragment were ligated with T4 DNA ligase at a molar ratio of 1:4 and incubated at 16 °C overnight. The ligation product was transformed into competent Escherichia coli BL21 (DE3) cells. After a 90 s heat shock at 42 °C, the cells were immediately cooled on ice for 2 min and then plated onto LB agar plates containing kanamycin. The plates were incubated at 37 °C for 12–16 h to select for positive clones. For the construction of the pET22b-TTL_1-378-6His plasmid, the gene was synthesized and inserted into the pET-22b(+) vector (Ampicillin) following the same procedure described above.
The BL21 was cultured at 37 °C to an OD600 of approximately 0.8, then cooled to 18 °C and induced for 16 h with the addition of 0.1 mM IPTG. BL21 was crushed by high pressure and centrifuged to remove the supernatant, which was gradually purified by a gravity column of IMAC Ni beads, Q-HP column, size exclusion chromatography (SEC), and then analyzed by SDS-PAGE73. The α/β-tubulin extracted from porcine brain was incubated with TTL and RB3 proteins at a molar ratio of 2:1.3:1.2 on ice, the obtained tubulin complex crystallized at a concentration of 20 mg/mL in the presence of 4-8% PEG, 4-5% glycerol, 0.1 M MES, 30 mM CaCl2, and 30 mM MgCl2 (pH 6.7) at 18 °C using the hanging drop method. The obtained Tubulin complex crystals were co-crystallized by incubation with 1 mM KY216, the co-crystals were finally immersed in a 10% DMSO solution containing 10 mM KY216 (buffer: 60 mM MgCl2, 60 mM CaCl2, 0.1 M Imidazole and MES (acid) pH 6.5, 18% ethylene glycol and PEG 8 K). After incubating overnight, the crystals were quickly frozen in liquid nitrogen for data collection.
X-ray data collection, structure solution, refinement, and analysis
X-ray diffraction data were obtained at the Shanghai Synchrotron Radiation Facility (SSRF) beamline BL02U1. The collected data reduction and scale were processed using XDS74 (https://xds.mr.mpg.de/html_doc/downloading.html) and Aimless software75,76. Molecular replacement was performed using Phaser, the model was built using winCoot 9.8 software, and refined using Phenix (https://phenix-online.org). The quality of the models was assessed using the PDB library. The molecular replacement model was generated from the tubulin-podophyllotoxin: stathmin-like domain complex (PDB code: 1SA1). All the structural diagrams presented in Fig. 1 were generated using PyMol (https://pymol.org/).
ITC assay
The recombinant VASH2 protein (cat# YHK84701) and the recombinant Tubulin protein (cat# CSK-T240-B-T) were both provided by AntibodySystem (USA). The ITC experiments were conducted using an ITC instrument manufactured by Malvern (UK). The sample cell had a capacity of 280 μL, and the titration syringe had a capacity of 40 μL. The injection volume per step was 3 μL, with an interval of 150 s between injections. A total of 13 titration steps was carried out. The reaction temperature was set at 25 °C, and the stirring speed was 750 rpm. In the experiment, all samples were prepared in a tubulin buffer containing 1 mM GTP (80 mM PIPES, pH 6.9; 2 mM MgCl₂; 0.5 mM EGTA), supplemented with 30 μM/3 μM VASH2, 3 μM tubulin, and 30 μM KY216 (dissolved from DMSO stock via buffer dilution). Prior to experiments, the instrument underwent a standardized cleaning protocol, and sample chambers were pre-rinsed three times with buffer prior to loading test samples. The experimental design comprised four independent experimental groups: (a) detection of KY216-tubulin interaction: Titration of 30 μM KY216 + buffer (syringe) into 3 μM tubulin + buffer (cell), with a control titration of 30 μM KY216 + buffer into buffer. (b) Detection of KY216-VASH2 interaction: Titration of 30 μM KY216 + buffer (syringe) into 3 μM VASH2 + buffer (cell), with a control titration of 30 μM KY216 + buffer into buffer. (c) Baseline binding detection of VASH2-tubulin: Titration of 30 μM VASH2 + buffer (syringe) into 3 μM tubulin + buffer (cell), with a control titration of 30 μM VASH2 + buffer into buffer. (d) Detection of the interaction between VASH2 and tubulin in the presence of KY216: Titration of 30 μM VASH2 + 30 μM KY216 + buffer (syringe) into 3 μM tubulin + 30 μM KY216 + buffer (cell), with a control titration of 30 μM VASH2 + 30 μM KY216 + buffer into 30 μM KY216 + buffer. The experimental system maintained a constant drug concentration by synchronizing the concentration of KY216 (30 μM) between the syringe and the sample chamber. In the subsequent ITC experiments targeting CA-4, all experimental conditions, including solvent composition, concentrations of the protein and the compound, and group design, were strictly maintained consistent with those employed in the aforementioned KY216 experiments. All experimental groups strictly maintained consistency in cosolvent concentration between the syringe and the sample chamber. The measured data were analyzed using the MicroCal PEAQ-ITC Analysis Software (MAN0576-01-EN-00) and fitted using the one set of sites model.
Synthesis of (6R,10aS)-6-(3,5-dichloro-4-methoxyphenyl)-2,3,10a,11-tetrahydro-[1,4]dioxino[2,3-g]imidazo[1,5-b]isoquinoline-8,10(6H,9H)-dione (KY216)
(1) Synthesis of (S)-2-((tert-butoxycarbonyl)amino)-3-(2,3-dihydrobenzo[b][1,4]dioxin-6-yl)propanoate (compound 1): to a solution of L-DOPA (10.0 g, 50.0 mmol) in MeOH (100 mL) under an ice-bath, SOCl₂ (10.88 mL, 150 mmol) was added dropwise. The reaction mixture was warmed to room temperature and stirred for 18 h. After concentration under reduced pressure, the crude methyl ester was obtained. This unpurified crude product was dissolved in 1,4-dioxane (29.4 mL). Aqueous NaOH solution (1 M, 29.4 mL) and (Boc)₂O (6.3 g, 29.04 mmol) were added successively. The mixture was stirred at room temperature for 6 h, acidified with 1 M HCl, and extracted with EtOAc to afford the crude Boc-protected intermediate, which was used directly in the next step without further purification. This crude material was dissolved in MeCN. K₂CO₃ (10 g, 72.6 mmol) and 1,2-dibromoethane (3.13 mL, 36.3 mmol) were added. The reaction mixture was heated under reflux at 80 °C for 18 h under a N₂ atmosphere. After cooling, the mixture was worked up and purified by silica gel column chromatography (16%EtOAc in PE) to afford pure compound 1 (34% yield, 5.7 g).
(2) Synthesis of (S)-5-((2,3-dihydrobenzo[b][1,4]dioxin-6-yl)methyl)-imidazolidine-2,4-dione (compound 2): compound 1 (3.4 g, 10 mmol) was dissolved in DCM (7.7 mL). TFA (7.7 mL, 100 mmol) was added dropwise. The reaction was allowed to warm to room temperature and stirred for 1 h. The mixture was diluted with DCM (150 mL) and basified with saturated Na₂CO₃ solution. The layers were separated. The aqueous layer was extracted with DCM (50 mL × 3). The combined organic layers were dried over anhydrous Na₂SO₄ and concentrated under vacuum to give the crude amine product. This crude product (2.37 g, 10 mmol) was dissolved in AcOH (50 mL). KCNO (1.62 g, 20 mmol) was added, and the mixture was stirred at room temperature for 18 h. The reaction was neutralized with saturated NaHCO₃ solution and extracted with EtOAc to give the crude cyclized product (hydantoin intermediate). This unpurified crude material was heated under reflux in 1 M HCl for 4 h. After cooling, the mixture was neutralized with saturated NaHCO₃, extracted with DCM, and crystallized to afford pure compound 2 (72% yield, 1.8 g).
(3) Synthesis of KY216: a mixture of compound 2 (62.06 mg, 0.25 mmol) and 3,5-dichloro-4-methoxybenzaldehyde (2 equiv., 0.5 mmol) in anhydrous DCM (10 mL), with a small amount of i-PrOH added to aid dissolution, was cooled in an ice-bath. Concentrated H₂SO₄ (0.027 mL, 0.5 mmol) was added dropwise. The reaction was allowed to warm to room temperature and stirred until completion, as monitored by TLC. The mixture was adjusted to pH 7 with saturated NaHCO₃ solution and extracted with EtOAc. The combined organic layers were dried over anhydrous Na₂SO₄. After concentration under reduced pressure, the crude product was purified by silica gel column chromatography (25% EtOAc in PE) to yield pure KY216 (51% yield, 55.5 mg). The ¹H and ¹³C NMR spectra of the new compound are presented in Supplementary Table 2. The high-resolution mass spectrum (HRMS) of KY216 is shown in Supplementary Fig. 16.
This experimental procedure involves highly hazardous materials, including strongly corrosive and water-reactive reagents (SOCl₂, TFA, concentrated H₂SO₄) and toxic substances (KCNO, 1,2-dibromoethane). Operations such as acid-base neutralization and heated reflux pose risks of strong exothermy, bumping, and fire. All manipulations must be conducted within a fume hood with comprehensive personal protective equipment.
Cell lines and cell culture
NCI-H460 (Cat# SCSP-584), A549 (Cat# SCSP-503), NCI-H1299 (Cat# TCHu160), HOS (Cat# TCHu167), U2OS (Cat# SCSP-5030), 143B (Cat#TCHu264), HUVECs (Cat# PSC-01), MCF 10A (Cat# SCSP-575), BEAS-2B (Cat# GNHu27), MCF-7 (Cat# SCSP-531), MDA-MB-231 (Cat# TCHu227), HCT 116 (Cat# SCSP-5076), SW480 (Cat# SCSP-5033), THLE-2 (Cat# SCSP-5068) and 95-D (Cat# TCHu 61) cells lines were kindly provided by the Stem Cell Bank of the Chinese Academy of Sciences (Shanghai, China). NCI-H460, NCI-H1299, U2OS, and A549 cells were cultured in RPMI 1640 medium (31800-014; Gibco, Waltham, MA, USA) supplemented with 10% fetal bovine serum (FCS500; ExCell Bio, Shanghai, China) and 100 U/mL penicillin-streptomycin (C100C5; New Cell and Molecular Biotech, Suzhou, China). Similarly, HUVECs, MCF 10A, BEAS-2B, MCF-7, HCT 116, HOS, MDA-MB-231, SW480, 143B, and 95-D cells were cultured in DMEM medium (10566016; Gibco, Waltham, MA, USA) with the same concentration of serum and antibiotics. THLE-2 cells were cultured in BEGM medium (CC-3170, Lonza, Suzhou, China) with the same concentration of serum and antibiotics. NCI-H460 cell culture medium was induced with TGF-β1 (10 ng/mL) (New Cell and Molecular Biotech, Suzhou, China), and subsequent experiments were performed 48 h later. All the cell lines were cultured at 37 °C in a 5% CO2 atmosphere. Mycoplasma contamination was tested by PCR assay (C0301S, Beyotime, China) with a negative result.
Colony formation assay
NCI-H460, A549, and 95-D cells were inoculated into 6-well plates at a density of 2 × 103 cells per well, and treated with drugs or DMSO 24 h later. After 48 h of KY216 treatment, the medium was replaced every 3 days with fresh medium and incubated in a 37 °C, 5% CO2 incubator for an additional seven days. Finally, cells were then fixed with 4% paraformaldehyde for 10 min and colonies were stained with crystal violet and counted using ImageJ software.
Cell proliferation
Inoculating NCI-H460, A549, and 95-D cells in 96-well plates (2 × 103 cells per well). After treatment with KY216, 10 μL of CCK-8 was added at the designated time points. After incubation at 37 °C for 2 h, use a Molecular Devices SpectraMaxPlus384 microplate reader to measure the absorbance at 450 nm (optical density [OD] 450). Cell proliferation was assessed using the ethynyldeoxyuridine (EdU, Beyotime) assay. Cells were seeded in 96 wells (1.0 × 104 cells per well). After 5 h of incubation with EdU, the subsequent steps were performed according to the manufacturer’s protocol. DAPI fluorescent dye was added to the plate for 10 min, visualized using the Molecular Devices ImageXpress High Content Confocal Imaging System, and quantified using ImageJ software.
Flow cytometric analysis of the cell cycle
NCI-H460 or 95-D cells were collected and digested with trypsin. The resulting cell suspensions were transferred to centrifuge tubes. Cells were washed twice with pre-cooled PBS, and after centrifugation, the supernatant was carefully removed, leaving the cell pellet. The cells were gently resuspended in 1 mL of pre-cooled 75% ethanol and fixed overnight at 4 °C. After fixation, the ethanol fixative was removed by centrifugation, and the cells were washed once more with PBS to eliminate residual ethanol. The cells were resuspended in 100 μL of binding buffer, and propidium iodide (PI) staining solution (20 μg/mL) was added to achieve a final staining volume of 500 μL. The mixture was incubated at room temperature for 30 min in the dark. Analysis was performed using a BD Accuri C6 flow cytometer. Initially, a gate (P1) was set on the forward scatter (FSC) vs side scatter (SSC) dot plot to identify the target cell population and exclude debris and aggregates. To further eliminate doublet cells, gating (P2) was applied using FSC-A vs FSC-H to ensure analysis of single cells. At least 10,000 valid cellular events within the P2 gate were collected for each sample. Finally, the PI fluorescence intensity histogram of the final gated population (P2) was analyzed using MFLT32 software to assess the cell cycle distribution across the G0/G1, S, and G2/M phases. The gating strategy for cell population identification is provided in Supplementary Fig. 17.
Plasmid and RNA construction and transfection
The OE-VASH2 plasmid in the pcDNA vector was constructed by Sangon Bioengineering Co., Ltd. (Shanghai, China). Wild-type and C158A/H193A/S210A mutant plasmids of VASH2 were inserted into the PEGFP-N1 vector, provided by Genomeditech Co., Ltd. (Shanghai, China). The SVBP-3× Flag plasmid was constructed by Jiman Bio and placed on the pcDNA vector. All VASH2 sequences were obtained from the NCBI for Biotechnology Information database (https://www.ncbi.nlm.nih.gov/datasets/gene/79805/#transcripts-and-proteins) transcript NM_001301056.2.
RiboBio Co., Ltd. (Guangzhou, China) provided siRNA targeting VASH2 and its negative control. The target sequences of si-VASH2-1, si-VASH2-2, and si-VASH2-3 were: 5’-GCGAUCCAGAAUUACAUGA-3’, 5’-CGAGAGUCCUUGCCUAUCA-3’, 5’-GGGCUGAGCUGAUGGACAA-3’. The miR-429 mimic sequence is 5’-UAAUACUGUCUGGUAAAACCGU-3’, and the miR-429 inhibitor sequence is 5’-ACGGUUUUACCAGACAGUAUUA-3’. The negative controls for the mimic (cat# miR1N0000001-1-5) and inhibitor (cat# miR2N0000001-1-5) were obtained from RiboBio Co., Ltd. (Guangzhou, China).
Lipofectamine™ 3000 (Invitrogen) was used to transfect siRNA or miRNA according to the manufacturer’s instructions. Plasmid transfection was performed using Lipofectamine™ 3000 plus P3000™ in Opti-MEM™. The Cells were collected at 48 h after transfection and used for further experiments.
Tube formation assay
Conditioned medium was prepared from NCI-H460 or 95-D cells treated with KY216 or transfected with siRNA or plasmids to alter VASH2 expression levels. HUVEC cells were seeded into six-well plates at a density of 1.5 × 105 cells per well during the exponential growth phase and incubated with 2 mL of conditioned medium for 48 h. After digestion, the cells were re-seeded at a density of 1.5 × 104 cells per well onto angiogenesis-specific slides coated (81506, Ibidi, Germany) with 10 μL of matrix gel (356324, Corning). The slides were incubated at 37 °C for 6 h. Tube formation and morphological changes were recorded using an inverted fluorescence microscope (Nikon Ts2R, Japan).
Enzyme-linked immunosorbent assay (ELISA)
Supernatants from NCI-H460 or 95-D cell cultures treated with KY216 or transfected with siVASH2 or VASH2 overexpression plasmids were collected. The secretion of VEGF was measured using the AuthentiKine™ Human VEGF ELISA Kit (KE00216, Proteintech, China) according to the manufacturer’s guidelines. The final optical density of each well was measured at 450 nm.
Wound healing assay
NCI-H460 or 95-D cells were seeded in a 6-well plate at a density of 1 × 106 cells per well. Once the cells reached the desired confluence, a wound was created by scraping the cells with a pipette. Subsequently, a specific concentration of KY216 was added for further culturing. Wound healing progress was monitored and recorded at 0, 24, and 48 h using a microscope (IX81, Olympus, Japan). The migration percentage was then calculated using the ImageJ software.
Cell migration and invasion assay
The migration and invasion of NCI-H460 and 95-D cells were evaluated using 8-μm pore size Transwell chambers (Corning). For migration experiments, 1 × 105 pretreated cells were placed in the upper chamber of Transwell chambers in 24-well plates with 150 μL of serum-free medium, while the lower chamber contained medium with 10% fetal bovine serum. After 48 h of incubation, the cells in the upper chamber were removed, and those that had migrated to the lower layer were fixed with 4% paraformaldehyde and stained with crystal violet. For invasion experiments, Transwell membranes were coated with a matrix gel (356324, Corning) with 2 × 105 cells per well before following the same steps as in the migration experiments. Subsequently, the cells were imaged using an inverted fluorescence microscope (Nikon Ts2R, Japan) and counted using ImageJ software.
Western blotting analysis
The cells were collected and sonicated using NP-40 lysate. The supernatant protein solution was obtained by centrifugation to remove the precipitate and quantified with a BCA kit (Beyotime, Shanghai, China). After normalization, proteins within 2× SDS-PAGE loading buffer containing 0.3% DTT were boiled for 10 min at 95 °C prior to electrophoresis. Subsequently, the proteins were separated via SDS-PAGE and transferred onto a PVDF membrane. The membrane was blocked with 5% skim milk in 1× TBST for 2 h at room temperature, and then incubated overnight at 4 °C with primary antibodies. Following membrane washing 3 times with 1× TBST, secondary antibodies were added for 2 h at room temperature. Finally, the membrane was washed with 1× TBST, visualized using an enhanced chemiluminescence detection system (Bio-Rad, CA, USA), quantified with ImageJ software, and the results were normalized to either the total protein amount or an internal reference level.
Immunofluorescence
NCI-H460 cells were cultured in 96-well plates (1.2 × 104 cells per well) or in confocal dishes (2 × 105 cells per well) and treated with KY216. Subsequently, cells were fixed (4% paraformaldehyde), permeabilized (0.1% Triton X-100), and blocked (0.5% bovine serum albumin). Next, the cells were incubated with a primary antibody and then with a fluorescence-coupled secondary antibody of the same species. After DAPI staining, images were captured using the ImageXpress Micro® system or Olympus FV3000 and quantified using the ImageJ software.
Co-Immunoprecipitation
The cells were centrifuged, sonicated, and lysed on ice for 20 min, and the protein concentration in the supernatant was determined using a BCA kit, then adjusted to 1 mg per 300 μL with lysate. Pre-cleaned magnetic beads (Selleck, B23202) were incubated with the protein samples overnight at 4 °C. Subsequently, either VASH2 (Proteintech, 13303-1-AP) or α-tubulin (Proteintech, 11224-1-AP) primary antibody was added, with IgG serving as a control, and gentle shaking was continued for 4-6 h. The supernatant was discarded, the magnetic beads were washed 4 times with PBS, and boiled for 10 min at 95 °C in 20 μL of 2× SDS-PAGE loading buffer for gel blot analysis. For cells labeled with GFP, GFP magnetic beads (ABclonal, AE079) were used, following the same procedure but omitting the primary antibody step.
PLA
This study utilized two custom-designed PLA probes to detect protein interactions through synthetic oligonucleotides attached to antibodies. When the distance between the two PLA probes was less than 40 nm, the fluorescently labeled oligonucleotides triggered amplification of the red signal. NCI-H460 cells were cultured in 96-well plates at a density of 1 × 104 cells per well and treated with 10 nM, 20 nM KY216, and DMSO for 48 h. Subsequently, cells were fixed, permeabilized, and blocked for 1 h at 37 °C. The cells were incubated overnight at 4 °C with VASH2 and α-tubulin primary antibody, then exposed to diluted anti-mouse MINUS and anti-rabbit PLUS probes (Sigma-Aldrich, SLCH3021) for 1 h at 37 °C. This was followed by the addition of ligase solution for 30 min at 37 °C and amplification of the polymerase solution for 100 min. Imaging using the ImageXpress Micro® system with 500 nm and quantitative analysis using ImageJ software were performed after restaining the cells with DAPI.
MiRNA sequencing and bioinformatic analysis
After cells were treated with DMSO and 20 nM KY216 for 48 h, total RNA was extracted by Trizol reagent kit (Invitrogen, Carlsbad, CA, USA), and the RNA molecules in a size range of 18-30 nt were enriched by polyacrylamide gel electrophoresis (PAGE). Then the 3’ adapters were added, and the 36–44 nt RNAs were enriched. The 5’ adapters were then ligated to the RNAs as well. The ligation products were reverse transcribed by PCR amplification, and the 140–160 bp size PCR products were enriched to generate a cDNA library and sequenced using Illumina Novaseq6000 by Gene Denovo Biotechnology Co. (Guangzhou, China).
The heatmap of identified miRNAs with log2FC > 3 or <−2 was drawn to display miRNA expression levels in different samples and to cluster miRNAs with similar expression patterns. MiRNAs differential expression analysis was performed by the edgeR software between two different groups or samples. MiRNAs with a fold change ≥ 2 and P value < 0.05 were considered statistically significant miRNAs. Miranda (Version 3.3a) and TargetScan (Version 7.0) were for predicting target genes of miRNAs, and omicshare tools (https://www.omicshare.com/tools/Home/Soft/getsoft/type/report) were for enrichment analyses.
mRNA sequencing
The extracted mRNA was enriched using mRNA Capture Beads (Vazyme, N401-01/02). After purification with the magnetic beads, the mRNA was fragmented at high temperature. Using the fragmented mRNA as a template, the first-strand cDNA was synthesized in a reverse-transcriptase mixture system. While synthesizing the second-strand cDNA, end repair and A-tail addition were completed. Subsequently, adapters were ligated, and the target fragments were purified and selected using Hieff NGS® DNA Selection Beads (Yeasen, 12601ES56). Then, PCR amplification of the library was carried out. After the quality inspection of the constructed sequencing library passed, it was sequenced on the sequencer. The sequencing service was provided by Gene Denovo Biotechnology Co. (Guangzhou, China).
Quantitative real-time reverse transcription PCR (qRT-PCR)
Total RNA was extracted using the RNA-Quick Purification Kit (ES Science, Shanghai, China) and converted to cDNA for mRNA analysis. miRNAs were extracted using the Tissue RNA Purification Kit Plus (ES Science, Shanghai, China) and transcribed into cDNA using the stem-loop method. Real-time fluorescence quantitative PCR was performed using SYBR Green Real-Time PCR Master Mix (Vazyme, Nanjing, China) on a LightCycler 480 system, with the primer sequences provided in Supplementary Table 3. The relative expression levels of the target genes were calculated based on the Ct values.
Dual-luciferase reporter assay
The wild-type plasmid pmiRGLO-WT-A was created by inserting the A site of the VASH2 3’ UTR region and its adjacent 300 bases into the dual luciferase pmiRGLO vector. The pmiRGLO-Mut-A plasmid was generated by altering the A site. Plasmids for sites C and D were constructed in a similar manner. All plasmids were constructed by Sangon Bioengineering (Shanghai, China). In 24-well plates, 8 × 104 293 T cells were seeded per well and co-transfected with miR-429 mimics, NC, or miR-429 mimics with wild-type or mutant plasmid of VASH2 3’ UTR using LipofectamineTM 2000 (Invitrogen). The cells were harvested after 48 h and assayed using the Dual Luciferase Reporter Assay Kit (Vazyme) to determine firefly and renilla luciferase activities in cell lysates. The results were expressed as the ratio of the two luciferase activities.
Patient tissues
A total of 75 NSCLC patient tissues were included (48 males and 27 females, aged from 27 to 76), with 37 having lung squamous cell carcinoma and 38 having lung adenocarcinoma. Sex was not considered a factor in the study design, as there is currently no established evidence linking it specifically to the growth and metastasis of NSCLC. Sex was classified based on biological characteristics, including chromosomes, genitalia, and endogenous hormone levels. All patient data were published after obtaining informed consent. Supplementary Table 4 provides specific information on the 75 patients. Patient tissues were exposed to primary antibodies against VASH2 (1:200) and N-cadherin (1:200) at 4 °C overnight. The horseradish peroxidase (HRP) labeled secondary antibody was incubated at room temperature for 60 min. After co-staining with DAPI, the images were obtained using a Nikon Eclipse C1 fluorescence microscope. Fluorescence quantification was completed by Wuhan Servicebio Biotechnology Co., Ltd (Wuhan, China) using Aipathwell software.
Animal studies
We fully adhere to SAGER guidelines, and our study design was not related to sex. Female SPF grade BALB/c nude mice (n = 30; 6 mice per group), aged 5–6 weeks and weighing 18–20 g, were utilized for xenograft tumor experiments and were procured from Shanghai Sippr-BK Laboratory Animal Co., Ltd. (Shanghai, China). The animals were kept in a specific pathogen-free environment with a temperature range of 20–24 °C, humidity between 40% and 60%, and a 12-h day/night cycle. NCI-H460 cells in logarithmic growth phase (3 × 106 cells) were suspended in 200 μL PBS and injected into the left flank of the mice. Tumor volume was calculated using the formula (length × width2)/2. When the tumor volume reached approximately 200 mm3, 30 mice were randomly assigned to five groups using a randomization procedure (http://www.randomizer.org/, Research Randomizer). KY216 efficacy was evaluated by administering doses of 5, 10, or 20 mg/kg every two days for 14 days, while the positive group received 10 mg/kg of paclitaxel. Regular measurements of body weight and tumor volume were taken, and the animals were euthanized by cervical dislocation at the end of the treatment period. Tumors and internal organs were collected, fixed in 4% paraformaldehyde, and analyzed using GraphPad Prism software for data on tumor volume, weight, and mouse body weight.
Male SPF grade BALB/c nude mice aged 6–8 weeks were maintained in a sterile environment (n = 32, 8 mice per group). To create a mouse model of lung cancer cell metastasis, the mice were first anesthetized with 1.5% sodium pentobarbital at a dose of 50 mg/kg. Subsequently, a 100 μL PBS suspension containing 1 × 106 luciferase-labeled 95-D (95-D-Luc) cells was injected into the left ventricle using a 26 G needle. AAV-OE-VASH2 virus or a control virus (Hanheng Biotechnology Co., Ltd., Shanghai, China) was then administered via the tail vein, with KY216 treatment at 20 mg/kg administered every two days, while the control group received saline. After 30 days, in vivo metastasis was confirmed using bioluminescence imaging. Specifically, the mice were injected intraperitoneally with 150 μL of 1.5% D-luciferin and anesthetized with 1.5% sodium pentobarbital. Image acquisition and analysis were conducted using the small animal in vivo imaging system (ABLX6, Shanghai Tianneng Technology Co., Ltd., Shanghai, China) in conjunction with Tanon Analysis Software.
Histology and IHC
Xenograft tumor tissues and major organs of mice were fixed in paraffin and sectioned for analysis. Pathological sections of major organs such as the heart, liver, and kidney were assessed for drug effects and tumor growth through Hematoxylin and Eosin (H&E) staining. Subcutaneous graft tumor sections underwent IHC analysis and were labeled with Ki-67 or CD31 antibodies for ABC-DAB staining. Antigenicity was restored using Vector Laboratories’ H-3300 repair solution. Microscopic images were captured at 20× magnification using a digital camera and an EVOS XL Core microscope. Finally, quantitative analysis was carried out using ImageJ software.
Data and statistical analysis
A blinded design was implemented for intervention and data analysis in both in vitro and in vivo studies. In vitro tests were performed in at least three independent biological replicates. Statistical significance was assessed using GraphPad Prism 9.5.0 (GraphPad Software, San Diego, CA, USA). Quantitative data were presented as mean ± SD or mean ± SEM. Survival curves were generated using the Kaplan–Meier method. Statistical analysis involved Student’s two-tailed t-test and one-way analysis of variance (ANOVA) with Tukey’s comparisons test. Results with P < 0.05 were considered statistically significant, while P ≥ 0.05 were considered as non-significant (ns).
Materials
Paclitaxel (CAS: 33069-62-4) and CA-4 (CAS:117048-59-6) were obtained from Shanghai Yuanye Bio-Technology (Shanghai, China) and confirmed to be more than 98% pure by performance liquid chromatography analysis. These compounds were dissolved in DMSO to yield a stock concentration of 50 mM, and the DMSO content in the final concentration of the experiment was less than 1‰. Duolink® In situ Red Starter Kit Rabbit/Mouse (DUO92002/DUO92004) was obtained from Sigma-Aldrich (Darmstadt, Germany). The following antibodies were used for the western blot analysis, immunofluorescence staining or Co-Immunoprecipitation: from Proteintech (Wuhan, China), β-Actin (Cat# 66009-1-Ig, Clone: 2D4H5, diluted at 1:50000 for WB), CDK1 (Cat# 10762-1-AP, Polyclonal, diluted at 1:2000 for WB), GAPDH (Cat# 10494-1-AP, Polyclonal, diluted at 1:20000 for WB), CDC25C (Cat# 16485-1-AP, Polyclonal, diluted at 1:2000 for WB), Cyclin B1 (Cat# 28603-1-AP, Polyclonal, diluted at 1:2000 for WB), N-cadherin (Cat# 22018-1-AP, Polyclonal, diluted at 1:2000 for WB, 1:200 for IF), Vimentin (Cat# 10366-1-AP, Polyclonal, diluted at 1:2000 for WB), E-cadherin (Cat# 20874-1-AP, Polyclonal, diluted at 1:2000 for WB, 1:200 for IF), ZEB1 (Cat# 21544-1-AP, Polyclonal, diluted at 1:2000 for WB), VASH2 (Cat# 13303-1-AP, Polyclonal, diluted at 1:2000 for WB, 1:200 for IF, IP: 0.5–4.0 ug for 1.0–3.0 mg of total protein lysate), α-Tubulin (Cat# 11224-1-AP, Polyclonal, diluted at 1:2000 for WB, 1:200 for IF, IP: 0.5–4.0 ug for 1.0–3.0 mg of total protein lysate), GFP Tag (Cat# 50430-2-AP, Polyclonal, diluted at 1:2000 for WB), Ki-67 (Cat# 27309-1-AP, Polyclonal, diluted at 1:5000 for IHC), CD31(Cat# 11265-1-AP, Polyclonal, diluted at 1:2000 for IHC); from Cell Signaling Technology (Boston, MA, USA), p-Smad2 (Cat# 18338, Clone: E8F3R, diluted at 1:1000 for WB), Smad2 (Cat# 5339, Clone: D43B4, diluted at 1:1000 for WB), Flag Tag (Cat# 14793, Clone: D6W5B, diluted at 1:1000 for WB); and from Abcam (Cambridge, England), Detyrosinated α-Tubulin (Cat# ab254154, Clone: AA12, diluted at 1:1000 for WB, 1:200 for IF).
Ethics statement
All research procedures involving human participants strictly adhere to the ethical guidelines of the institutional and national research committees and are in compliance with the Declaration of Helsinki. The NSCLC patient tissues and profiles were obtained from Shanghai Pulmonary Hospital and approved by the Ethical Review Committee of Shanghai Pulmonary Hospital (K23-334). Informed written consent was obtained from all participants involved in the research. All animal care and experimental protocols were approved (2023-10-014) by the Institutional Animal Care and Use Committee (IACUC) of the Laboratory Animal Center of China Pharmaceutical University. Tumor burden was strictly monitored and did not exceed the maximum permitted size (2.0 cm in diameter or 2000 mm³ in volume for mice) as per institutional guidelines.
Data availability
The atomic coordinates and structure factors for the reported crystal structures of the KY216-tubulin complex have been deposited with the Protein Data Bank (PDB) under accession code 8YRK. The raw RNA-sequencing data generated in this study have been deposited in the Genome Sequence Archive in the National Genomics Data Center, China National Center for Bioinformation/Beijing Institute of Genomics, Chinese Academy of Sciences, under accession code HRA010164. The raw miRNA-sequencing data generated in this study have been deposited in the Genome Sequence Archive in the National Genomics Data Center, China National Center for Bioinformation/Beijing Institute of Genomics, Chinese Academy of Sciences, under accession code HRA010082. Source data are provided with this paper.
References
Siegel, R. L., Miller, K. D., Wagle, N. S. & Jemal, A. Cancer statistics, 2023. CA A Cancer J. Clin. 73, 17–48 (2023).
Mehlen, P. & Puisieux, A. Metastasis: a question of life or death. Nat. Rev. Cancer 6, 449–458 (2006).
Wood, S. L., Pernemalm, M., Crosbie, P. A. & Whetton, A. D. The role of the tumor-microenvironment in lung cancer-metastasis and its relationship to potential therapeutic targets. Cancer Treat. Rev. 40, 558–566 (2014).
Dela Cruz, C. S., Tanoue, L. T. & Matthay, R. A. Lung cancer: epidemiology, etiology, and prevention. Clin. Chest Med. 32, 605–644 (2011).
Sebastian, J. & Rathinasamy, K. Microtubules and cell division: potential pharmacological targets in cancer therapy. Curr. Drug Targets 24, 889–918 (2023).
Garcin, C. & Straube, A. Microtubules in cell migration. Essays Biochem. 63, 509–520 (2019).
Gudimchuk, N. B. & McIntosh, J. R. Regulation of microtubule dynamics, mechanics and function through the growing tip. Nat. Rev. Mol. Cell Biol. 22, 777–795 (2021).
Roll-Mecak, A. The tubulin code in microtubule dynamics and information encoding. Dev. Cell 54, 7–20 (2020).
Lopes, D. & Maiato, H. The tubulin code in mitosis and cancer. Cells 9, 2356 (2020).
Khwaja, S., Kumar, K., Das, R. & Negi, A. S. Microtubule associated proteins as targets for anticancer drug development. Bioorg. Chem. 116, 105320 (2021).
Pérez-Pérez, M.-J. et al. Blocking blood flow to solid tumors by destabilizing tubulin: an approach to targeting tumor growth. J. Med. Chem. 59, 8685–8711 (2016).
Krzywik, J. et al. An insight into the anticancer potential of carbamates and thiocarbamates of 10-demethoxy-10-methylaminocolchicine. Eur. J. Med. Chem. 215, 113282 (2021).
Steinmetz, M. O. & Prota, A. E. Microtubule-targeting agents: strategies to hijack the cytoskeleton. Trends Cell Biol. 28, 776–792 (2018).
Shao, Y.-Y. et al. Synthesis and biological evaluation of novel shikonin-benzo[b]furan derivatives as tubulin polymerization inhibitors targeting the colchicine binding site. Eur. J. Med. Chem. 190, 112105 (2020).
Wordeman, L. & Vicente, J. J. Microtubule targeting agents in disease: classic drugs, novel roles. Cancers 13, 5650 (2021).
Kaul, R., Risinger, A. L. & Mooberry, S. L. Microtubule-targeting drugs: more than antimitotics. J. Nat. Prod. 82, 680–685 (2019).
Du, H. et al. The roles of vasohibin and its family members: beyond angiogenesis modulators. Cancer Biol. Ther. 18, 827–832 (2017).
Norita, R. et al. Vasohibin-2 is required for epithelial–mesenchymal transition of ovarian cancer cells by modulating transforming growth factor-β signaling. Cancer Sci. 108, 419–426 (2017).
Zhang, Y. et al. Vasohibin 2 promotes malignant behaviors of pancreatic cancer cells by inducing epithelial-mesenchymal transition via Hedgehog signaling pathway. Cancer Med. 7, 5567–5576 (2018).
Shayimu, P., Yin, C., Zeng, X. & Jiapaer, R. Knockdown of VASH2 inhibits the stemness and EMT process by regulating ZEB2 in colorectal cancer. Curr. Stem Cell Res. Ther. 19, 126–132 (2024).
Xue, X. et al. MiR200-upregulated Vasohibin 2 promotes the malignant transformation of tumors by inducing epithelial-mesenchymal transition in hepatocellular carcinoma. Cell Commun. Signal 12, 62 (2014).
Liu, P. Study on the mechanisms of VASH2/TGF-β/VEGF-D in promoting the lymph node metastasis of lung squamous cell carcinoma. J. Thorac. Oncol. 13, S1075 (2018).
Overby, A., Takahashi, T., Matsui, H. & Nakamura, M. Angiogenesis and MALT lymphoma in stomach, liver and salivary gland: interaction with Vash2 and VEGF. Gastroenterology 150, S37 (2016).
Nieuwenhuis, J. et al. Vasohibins encode tubulin detyrosinating activity. Science 358, 1453–1456 (2017).
Aillaud, C. et al. Vasohibins/SVBP are tubulin carboxypeptidases (TCPs) that regulate neuron differentiation. Science 358, 1448–1453 (2017).
Ramirez-Rios, S. et al. VASH1–SVBP and VASH2–SVBP generate different detyrosination profiles on microtubules. J. Cell Biol. 222, e202205096 (2022).
Zhou, C., Yan, L., Zhang, W. & Liu, Z. Structural basis of tubulin detyrosination by VASH2/SVBP heterodimer. Nat. Commun. 10, 1–8 (2019).
Bartel, D. P. Metazoan MicroRNAs. Cell 173, 20–51 (2018).
Sun, L., Chen, T., Li, T. & Yu, J. LncRNA IUR downregulates ZEB1 by upregulating miR-200 to inhibit prostate carcinoma. Physiol. Genomics 51, 607–611 (2019).
Wen, B., Zhu, R., Jin, H. & Zhao, K. Differential expression and role of miR-200 family in multiple tumors. Anal. Biochem. 626, 114243 (2021).
Title, A. C. et al. Genetic dissection of the miR-200–Zeb1 axis reveals its importance in tumor differentiation and invasion. Nat. Commun. 9, 4671 (2018).
Gibbons, D. L. et al. Contextual extracellular cues promote tumor cell EMT and metastasis by regulating miR-200 family expression. Genes Dev. 23, 2140–2151 (2009).
Pecot, C. V. et al. Tumour angiogenesis regulation by the miR-200 family. Nat. Commun. 4, 2427 (2013).
Zhu, W. et al. Expression of miR-29c, miR-93, and miR-429 as potential biomarkers for detection of early stage non-small lung cancer. PLoS One 9, e87780 (2014).
Xue, B. et al. miR-200 deficiency promotes lung cancer metastasis by activating Notch signaling in cancer-associated fibroblasts. Genes Dev. 35, 1109–1122 (2021).
Garinet, S. et al. Clinical assessment of the miR-34, miR-200, ZEB1 and SNAIL EMT regulation hub underlines the differential prognostic value of EMT miRs to drive mesenchymal transition and prognosis in resected NSCLC. Br. J. Cancer 125, 1544–1551 (2021).
Cao, L., Zhou, X., Ding, X. & Gao, D. Knockdown of circ‑PVT1 inhibits the progression of lung adenocarcinoma and enhances the sensitivity to cisplatin via the miR‑429/FOXK1 signaling axis. Mol. Med. Rep. 24, 1–11 (2021).
Leng, J. et al. Discovery of novel N-heterocyclic-fused deoxypodophyllotoxin analogues as tubulin polymerization inhibitors targeting the colchicine-binding site for cancer treatment. J. Med. Chem. 65, 16774–16800 (2022).
Liu, Y.-N. et al. Design, synthesis, and biological evaluation of 1-methyl-1,4-dihydroindeno[1,2-c]pyrazole analogues as potential anticancer agents targeting tubulin colchicine binding site. J. Med. Chem. 59, 5341–5355 (2016).
Chen, Q. W. et al. LncRNA BC promotes lung adenocarcinoma progression by modulating IMPAD1 alternative splicing. Clin. Transl. Med. 13, e1129 (2023).
Cantelli, G., Crosas-Molist, E., Georgouli, M. & Sanz-Moreno, V. TGFΒ-induced transcription in cancer. Semin Cancer Biol. 42, 60–69 (2017).
Kaszak, I. et al. Role of cadherins in cancer—a review. Int. J. Mol. Sci. 21, 7624 (2020).
Wang, N. et al. Structural basis of tubulin detyrosination by the vasohibin–SVBP enzyme complex. Nat. Struct. Mol. Biol. 26, 571–582 (2019).
Li, F., Hu, Y., Qi, S., Luo, X. & Yu, H. Structural basis of tubulin detyrosination by vasohibins. Nat. Struct. Mol. Biol. 26, 583–591 (2019).
Gullberg, M. & Andersson, A.-C. Visualization and quantification of protein–protein interactions in cells and tissues. Nat. Methods 7, v–vi (2010).
Bracken, C. P. et al. A double-negative feedback loop between ZEB1-SIP1 and the microRNA-200 family regulates epithelial-mesenchymal transition. Cancer Res. 68, 7846–7854 (2008).
Liu, W. & Wang, X. Prediction of functional microRNA targets by integrative modeling of microRNA binding and target expression data. Genome Biol. 20, 18 (2019).
Agarwal, V., Bell, G. W., Nam, J.-W. & Bartel, D. P. Predicting effective microRNA target sites in mammalian mRNAs. eLife 4, e05005 (2015).
Li, J.-H., Liu, S., Zhou, H., Qu, L.-H. & Yang, J.-H. starBase v2.0: decoding miRNA-ceRNA, miRNA-ncRNA and protein–RNA interaction networks from large-scale CLIP-Seq data. Nucleic Acids Res. 42, D92–D97 (2014).
Gunyuz, Z. E. et al. SEMA6D differentially regulates proliferation, migration, and invasion of breast cell lines. ACS Omega 7, 15769–15778 (2022).
Chang, Y. et al. miR-145-5p inhibits epithelial-mesenchymal transition via the JNK signaling pathway by targeting MAP3K1 in non-small cell lung cancer cells. Oncol. Lett. 14, 6923–6928 (2017).
Xu, L.-M. et al. MicroRNA-135 inhibits initiation of epithelial-mesenchymal transition in breast cancer by targeting ZNF217 and promoting m6A modification of NANOG. Oncogene 41, 1742–1751 (2022).
Wang, A. et al. Endoplasmic reticulum stress-related super enhancer promotes epithelial-mesenchymal transformation in hepatocellular carcinoma through CREB5 mediated activation of TNC. Cell Death Dis. 16, 73 (2025).
Ravelli, R. B. G. et al. Insight into tubulin regulation from a complex with colchicine and a stathmin-like domain. Nature 428, 198–202 (2004).
von Eiff, D. et al. Paclitaxel for treatment of advanced small cell lung cancer (SCLC): a retrospective study of 185 patients. J. Thorac. Dis. 12, 782–793 (2020).
Tian, Y. et al. Mechanism of multidrug resistance to chemotherapy mediated by P-glycoprotein (Review). Int J. Oncol. 63, 119 (2023).
Yang, J. et al. Replacing the tropolonic methoxyl group of colchicine with methylamino increases tubulin binding affinity with improved therapeutic index and overcomes paclitaxel cross-resistance. Drug Resist. Updates 68, 100951 (2023).
Wiesen, K. M., Xia, S., Yang, C.-P. H. & Horwitz, S. B. Wild-type class I β-tubulin sensitizes Taxol-resistant breast adenocarcinoma cells harboring a β-tubulin mutation. Cancer Lett. 257, 227–235 (2007).
Mitra, A. & Sept, D. Taxol allosterically alters the dynamics of the tubulin dimer and increases the flexibility of microtubules. Biophys. J. 95, 3252–3258 (2008).
Sahakyan, H., Abelyan, N., Arakelov, V., Arakelov, G. & Nazaryan, K. In silico study of colchicine resistance molecular mechanisms caused by tubulin structural polymorphism. PLoS One 14, e0221532 (2019).
Xie, S., Wu, Z., Qi, Y., Wu, B. & Zhu, X. The metastasizing mechanisms of lung cancer: recent advances and therapeutic challenges. Biomed. Pharmacother. 138, 111450 (2021).
Eser, P. Ö & Jänne, P. A. TGFβ pathway inhibition in the treatment of non-small cell lung cancer. Pharmacol. Ther. 184, 112–130 (2018).
Feng, H.-T., Zhao, W.-W., Lu, J.-J., Wang, Y.-T. & Chen, X.-P. Hypaconitine inhibits TGF-β1-induced epithelial–mesenchymal transition and suppresses adhesion, migration, and invasion of lung cancer A549 cells. Chin. J. Nat. Med. 15, 427–435 (2017).
Wu, Q. et al. PTBP3 promotes migration of non-small cell lung cancer through regulating E-cadherin in EMT signaling pathway. Cancer Cell Int. 20, 172 (2020).
Liao, H. et al. miR‑454‑3p inhibits non‑small cell lung cancer cell proliferation and metastasis by targeting TGFB2. Oncol. Rep. 45, 1–12 (2021).
Nieuwenhuis, J. & Brummelkamp, T. R. The tubulin detyrosination cycle: function and enzymes. Trends Cell Biol. 29, 80–92 (2019).
Chen, C. Y. et al. Suppression of detyrosinated microtubules improves cardiomyocyte function in human heart failure. Nat. Med. 24, 1225–1233 (2018).
Stemberger, M. B. et al. Hydrogen peroxide induces α-tubulin detyrosination and acetylation and impacts breast cancer metastatic phenotypes. Cells 12, 1266 (2023).
Zhang, L. et al. MiR-429 suppresses proliferation and invasion of breast cancer via inhibiting the Wnt/β-catenin signaling pathway. Thorac. Cancer 11, 3126–3138 (2020).
Cava, C. et al. Theranostic application of miR-429 in HER2+ breast cancer. Theranostics 10, 50–61 (2020).
Lang, Y. et al. MicroRNA-429 induces tumorigenesis of human non-small cell lung cancer cells and targets multiple tumor suppressor genes. Biochem. Biophys. Res. Commun. 450, 154–159 (2014).
Halvorsen, A. R. et al. A unique set of 6 circulating microRNAs for early detection of non-small cell lung cancer. Oncotarget 7, 37250–37259 (2016).
Tan, L. et al. Design, synthesis, and biological evaluation of heterocyclic-fused pyrimidine chemotypes guided by X-ray crystal structure with potential antitumor and anti-multidrug resistance efficacy targeting the colchicine binding site. J. Med. Chem. 66, 3588–3620 (2023).
Kabsch, W. XDS. Acta Cryst D Biol. Crystallogr. 66, 125–132 (2010).
Evans, P. R. & Murshudov, G. N. How good are my data and what is the resolution? Acta Cryst. D 69, 1204–1214 (2013).
CNCB-NGDC Members and Partners Database resources of the national genomics data center, China National Center for Bioinformation in 2022. Nucleic Acids Res. 50, D27–D38 (2022).
Acknowledgements
This work was supported by the Guangxi Key Laboratory of Early Prevention and Treatment for Regional High Frequency Tumor (GKE-KF202402) to Y.-Z.X., and the Basic Research Project for the Development of Modern Industrial College of Traditional Chinese Medicine and Health at Lishui University. National Natural Science Foundation of China (grant numbers 81973524) to Y.-Z.X., special funds for science and technology development under the guidance of the central government (2021Szvup163, ZY20198020) to Y.-Z.X. and L.-Y.K., and the “Double First-Class” University Project (CPU2018GF03) to Y.-Z.X. and L.-Y.K. We are grateful for the technical guidance provided by Yu-chen Ye from the Public Experimental Platform, China Pharmaceutical University.
Author information
Authors and Affiliations
Contributions
X.Y. He: conceptualization, data curation, investigation, software, visualization, methodology, and writing—original draft. C.Y. Liu: experimental validation, methodology, and visualization. X.Y. Ding: clinical cohort, resources, data curation, writing—review, and supervision. Q. Lin: formal analysis, investigation, and visualization. J.F. Leng: material support and methodology. C.M. Xiao and Xin Zang: methodology and software. M.Y. Wang: investigation and visualization. Y. Yin: material support, resources, methodology, validation, funding acquisition, and writing—review and editing. L.Y. Kong: supervision, resources, funding acquisition, and writing—review and editing. Y.Z. Xia: conceptualization, design of the project, supervision, funding acquisition, resources, methodology, project administration, experimental validation, and writing—review and editing.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks J Richard McIntosh and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
He, XY., Liu, CY., Ding, XY. et al. KY216-tubulin complex captures VASH2 to inhibit NSCLC metastasis. Nat Commun 17, 191 (2026). https://doi.org/10.1038/s41467-025-66817-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-66817-2