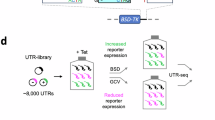
Abstract
Master transcription factors (MTFs) are key regulators in cell fate determination. However, an approach to profile MTF’s direct transcriptional targets together with their associated RNA-binding proteins (RBPs), is still lacking. Here, we applied 5-ethynyluridine RNA metabolic labeling and click chemistry to map the dynamics of the transcriptional targets and the RBPs interacting with the newly transcribed RNAs upon acute NANOG degradation in mouse embryonic stem cells (mESCs). We identified UTP15, a classic rRNA-biogenesis regulator, acts as a key activator of pluripotency-associated gene transcription independently of rRNA biogenesis. Importantly, NANOG-regulated transcription enhances UTP15 binding to transcription start sites (TSSs), associated with increased Pol II binding and more active transcription. Moreover, UTP15 promotes the assembly of Pol II biomolecular condensates, thereby potentially driving pluripotency gene transcription. Collectively, our study uncovers a NANOG-nascent transcript-UTP15 regulatory axis to activate pluripotency gene transcription, providing a distinct approach for studying MTF’s function during cell fate determination.
Similar content being viewed by others
Introduction
Pluripotency represents a specific cellular state featured by the capacity of differentiating into various lineages and propagated infinitely in vitro1. Enormous efforts have been taken to decipher the molecular basis underlying the regulation of pluripotency. Among these, signaling pathways, epigenetics, transcription, translation, and protein degradation orchestrate the regulatory network in pluripotency maintenance or acquisition2,3,4,5,6,7,8,9. Noteworthy, the pluripotency program was gradually recognized to rely heavily on a remarkably small set of transcription factors, which were denoted as master transcription factors (MTFs)5,10,11.
MTFs play fundamental roles in controlling the core transcriptional circuitry of different cell types to maintain their specific identities. For instance, NANOG, SOX2, and OCT4 were shown to maintain the core pluripotency circuitry in embryonic stem cells5, GATA1 in hematopoietic stem cells12, and MyoD in myogenic stem cells13. MTFs bind to and activate the core transcriptional gene expression or typical enhancers and super-enhancers to drive the core transcriptional gene program as identified by chromatin immunoprecipitation followed by sequencing (ChIP-seq)10. However, the genomic binding of MTFs does not ensure a direct transcriptional output, as it is influenced by the context-dependent chromatin state14. In this regard, the identification of direct transcriptional targets of specific factors has been achieved through a combination of rapid protein degradation and newly transcribed RNA sequencing15. The newly transcribed RNAs regulated by specific TFs undergo precisely governed transcription, splicing, modification, as well as degradation to ultimately establish a specific transcriptomic profile. These processes are potentially guided by certain RNA-binding proteins (RBPs)16,17,18,19. Yet, an approach to systematically identify RBPs interacting with newly transcribed RNAs regulated by specific MTFs is still lacking.
Accumulating evidence showed that RBPs play crucial roles in regulating various aspects of RNA metabolism and function20,21. More recently, RBPs are reported to be more enriched in intrinsically disordered regions (IDRs)22,23,24. These IDRs facilitate biomolecular condensation through mechanisms such as liquid-liquid phase separation (LLPS). Specifically, intrinsically disordered prion-like low-complexity domains (PLCDs) enable phase separation coupled to percolation25,26,27, thereby supporting diverse biological processes28,29. In this regard, IDRs mediate autophagosome formation, high-order chromatin organization, immune signaling transduction, chromosome segregation during cell division, and the assembly of transcription machinery30,31,32. However, how RBPs interact with newly transcribed RNA to fulfill and coordinate the functions of specific MTFs in maintaining cell identity remains largely unexplored.
Here, using NANOG as a representative MTF, we profile the direct transcriptional targets of NANOG by leveraging 5-ethynyluridine (EU)-based nascent RNA sequencing. We applied RNA interactome using click chemistry (RICK) technology to map the dynamics of newly transcribed RNA-associated RNA binding proteins (RBPs) upon acute NANOG depletion33,34. This identified 195 candidate NANOG-related RBPs, of which 106 showed a decreased tendency to bind newly transcribed RNAs upon acute NANOG depletion. We verified UTP15, a component of the small subunit processome (SSUP) complex, recognizes a subset of NANOG transcriptional targets to activate cognate gene transcription independent of rRNA biogenesis. We also demonstrated that the IDR of UTP15 has the capacity to form biomolecular condensates, and to support the formation of Pol II clusters to activate gene transcription. Hence, our study establishes a NANOG-nascent RNA-UTP15 regulatory axis to maintain pluripotency in mESCs and provides a novel strategy to study the MTF’s function in diverse cell identities.
Results
Acute depletion of NANOG identifies direct transcriptional targets
To determine the direct transcriptional targets of NANOG, we employed a previously established mouse embryonic stem cell (mESC) line containing an auxin-inducible degron (AID)-tagged NANOG35, which allows a rapid and inducible degradation of NANOG upon indole-3-acetic acid (IAA) treatments (Fig. 1a)15,35. We first measured the level of AID-tagged NANOG protein to ascertain that its expression is comparable to endogenous NANOG in wildtype mESCs (Supplementary Fig. 1a). This is important because high levels of NANOG enforce mESC self-renewal even if leukemia inhibitory factor (LIF) was removed14,36,37. In line with previous reports, we observed upregulation of endoderm/primitive endoderm markers (e.g., Gata6 and Gata4)38,39 and downregulation of pluripotency markers (e.g., Klf4 and Esrrb)40,41 48 hours after the IAA-induced NANOG degradation (Supplementary Fig. 1b). This indicates that the original function of NANOG is not noticeably altered, and the rapid degradation of NANOG provides a foundation for direct transcriptional target analysis.
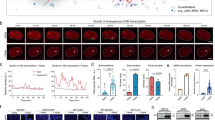
a Schematic overview of nascent RNA-seq in NANOG-AID cells. Cells were incubated with 1 mM EU for 15 minutes at designated time points (0 h, 2 h, 4 h, 8 h, 24 h) after IAA treatment. Total RNA was extracted, and EU-labeled spike-in RNA was added for normalization. The RNA was biotinylated using click chemistry, then purified and used for Next-Generation Sequencing (NGS) library construction. Western blot analysis verifies NANOG protein levels following IAA treatment. Representative western blots are shown from two independent experiments with consistent results. b MA plots show nascent RNA level changes following acute IAA-induced degradation of NANOG protein. 2 h versus 0 h (left), 4 h versus 0 h (right). (FDR < 0.05, fold change > 1.5). c Percentage of transcriptionally changed genes (at 2 h vs. 0 h, identified by nascent RNA-seq; n = 1,812) with NANOG genomic binding. (FDR < 0.05, fold change > 1.5). d Heatmap showing fold change (log2) of transcriptionally changed genes in both nascent RNA-seq and corresponding total RNA-seq (n = 2,091), 2 h versus 0 h, 4 h versus 0 h; genes were divided into six categories (Cluster Ⅰ–Ⅵ) using hierarchical clustering. e Gene Ontology (GO) analysis of Cluster Ⅱ genes related to Fig. 1d (n = 486). f Integrative Genomic Viewer (IGV) snapshot of nascent RNA-seq at Gata6 gene locus after IAA-induced NANOG degradation (0 h, 2 h, 4 h, 8 h, 24 h). Source data for Fig. 1a are provided as a Source Data file.
We performed nascent RNA sequencing (RNA-seq) using 15-minute EU labeling at different time points after the IAA treatments (i.e., 0, 2, 4, 8, and 24 hours) (Fig. 1a). Nascent RNA-seq measures the immediate transcriptional dynamics upon stimulation or ablation of specific factors15,42,43,44. Notably, we identified 1812 genes that showed immediate transcriptional change within as short as two hours after the IAA treatments (Supplementary Data 1). These genes are potential direct transcriptional targets of NANOG in mESCs. Among them, 1,788 genes are downregulated while only 24 genes upregulated (Fig. 1b). These genes include classical NANOG transcriptional targets such as Esrrb and Klf441,45 (Fig. 1b, Supplementary Fig. 1c). Further analysis showed that 1,306 (72.1%) genetic loci of the 1,812 changed genes are physically bound by NANOG as defined by ChIP-seq (Fig. 1c)46,47,48.
To assess the relationship between transcriptional change and total RNA level upon acute NANOG degradation, we analyzed both nascent RNA-seq and total RNA-seq at two hours and four hours after the IAA treatments (Fig. 1d). We noted that of all 2091 transcriptionally changed genes, most of them (1,605, 76.8%: cluster Ⅰ, n = 1,493; cluster Ⅲ, n = 43; cluster Ⅳ, n = 44; cluster Ⅴ, n = 24 and cluster Ⅵ, n = 1) showed a consistent tendency or no obvious change at their total RNA level. Among these genes, the transcriptionally decreased ones (1538, 73.6%: cluster Ⅰ, cluster Ⅳ and cluster Ⅵ) show enrichment of terms related to multicellular organism development and cell differentiation in Gene Ontology (GO) analysis (Supplementary Fig. 1d). In contrast, a small proportion of them (cluster Ⅱ, n = 486) had higher levels of total RNA even though the nascent transcript declined (Fig. 1d, Supplementary Data 1). GO analysis showed that these genes are highly enriched in terms related to regulation of transcription from RNA polymerase II promoter, cell migration, and somatic stem cell population maintenance (Fig. 1e). The different change tendency of transcription and total RNA level upon acute NANOG degradation indicates that unresolved mechanisms including RNA splicing and RNA degradation may regulate the ultimate total mRNA levels.
Further analysis on nascent RNA-seq showed that genes related to pluripotency (e.g., Klf4 and Esrrb) are transcriptionally downregulated upon acute NANOG degradation (Fig. 1b, Supplementary Fig. 1c). In contrast, among the small number of transcriptionally upregulated genes, Gata6 transcription is enhanced as early as two and four hours after IAA treatments (Fig. 1f). Gata6 is a master transcription factor at the top of the hierarchy regulating early primitive endoderm (PrE) specification at blastocyst stage38,49,50. This fast and potentially direct regulation mode51,52 provides an approach to determine the “salt-and-pepper” expression pattern in a short time-window during early embryonic development53. However, we did not observe immediate transcriptional increase in other differentiation markers including Otx2 and Pdgfra upon NANOG degradation (Supplementary Fig. 1e). These markers were upregulated only at eight or 24 hours after the IAA treatments, suggesting they are not direct transcriptional targets of NANOG in mESCs, or their transcriptional changes require prerequisite events such as chromatin remodeling or a more permissive epigenetic state concomitant with differentiation.
Taken together, we have constructed a NANOG acute degradation system, which combines with the EU-based nascent RNA-seq to enable the identification of potential direct transcriptional targets of NANOG in mESCs. This also paves the way for measuring the RBPs interacting with the altered nascent RNAs upon acute NANOG depletion.
Newly transcribed RNA-binding proteome dynamics upon NANOG depletion
We next investigate whether NANOG depletion would induce changes in the binding proteome of newly transcribed RNAs. This is important because the binding proteins of the newly transcribed RNAs may coordinate RNA transcription or processing and translocation shortly after transcription, and thus coordinate the function of nascent RNAs regulated by specific TFs. To achieve this, we performed RICK following a two-hour EU labeling shortly after IAA treatments (at both two and six hours) to identify RBPs that could coordinate and trigger the effects upon NANOG depletion (Fig. 2a). This led to the identification of a combined 786 putative RBPs. Among these, 569 (72.4%) were previously identified as RBPs by different approaches (Fig. 2b, Supplementary Fig. 2a)24,54,55. As expected, GO molecular function analysis of the RBPs showed enriched terms related to mRNA processing and RNA splicing (Supplementary Fig. 2b). Moreover, classical RNA-binding domains (RBDs), non-classical RBDs and many unknown RBDs (e.g., fn3 and HMG box) are overrepresented in these identified RBPs (Fig. 2c, Supplementary Fig. 2c)23,55. As expected, the distribution of the predicted isoelectric points of identified proteins shifts toward a basic pH, which is a typical feature of RNA binding proteome (Fig. 2d)55.
a The schematic diagram showing capture of RNA interactome using click chemistry (RICK) in NANOG-AID cells. We treated the cells with IAA at different time points (i.e., 0 h, 2 h, 6 h), and incubated with 1 mM EU for 2 h. After 254 nm UV crosslinking, the cells were collected and lysed with lysis buffer. The resultant lysates were biotinylated and enriched with streptavidin agarose beads. The obtained RNA-protein complexes were digested for mass spectrometry. b Overlap between 786 RICK proteins and RBP2GO database54 (mouse, n = 2886). c Bar chart showing the number of 786 RICK proteins with classical RNA binding domains (left panel) and non-classical RNA binding domains (right panel). d Density plot showing the distribution of isoelectric point (pI) for 786 RICK proteins and control proteins (RICK searching proteins by MaxQuant, n = 3825). e Line plots showing the fold change (log2) dynamics of 786 RICK proteins at different time points compared to 0 h are represented, with each line corresponding to one protein. Solid lines, the mean values and the average trend of all proteins within the cluster. The 786 RICK proteins were categorized into three groups based on change trends: Increased (n = 89), Decreased (n = 106), and no significant change (n = 591). f GO analysis of increased proteins related to Fig. 2e (n = 89). g GO analysis of decreased proteins related to Fig. 2e (n = 106).
Although the overall levels of the identified RBPs remained unchanged following NANOG degradation (Supplementary Fig. 2d), a subset showed altered binding capacity to newly transcribed RNAs, with 89 (11.3%) exhibiting increased binding and 106 (13.5%) showing decreased binding, while the majority (591, 75.2%) remained stable (Fig. 2e, Supplementary Data 2). Notably, the increased RBPs are mainly associated with chromatin remodeling, chromatin organization, or mitochondrial translation (Fig. 2f). In contrast, the decreased ones were more enriched for mRNA processing, RNA splicing, and ribosome biogenesis (Fig. 2g). The protein-protein interaction (PPI) analysis also showed that chromatin organization and RNA processing are clustered among the increased and decreased RBPs, respectively (Supplementary Fig. 2e, f). These indicate that chromatin reorganization and RNA metabolic switch occur during the ESC differentiation6,7, which could be triggered by NANOG depletion36,37.
Altogether, the application of RICK enables the identification of RNA binding proteome to newly transcribed RNAs regulated by NANOG upon acute protein degradation. This also reveals a global change of newly transcribed RBPs related to chromatin organization (increased RBPs) and RNA processing (decreased RBPs) upon NANOG depletion.
UTP15 is essential for pluripotency and embryonic development
Considering that the majority of NANOG direct transcriptional targets are downregulated after IAA treatment, we expect that mass-spectrometric capture of the RNA-binding proteins that associate with these transcripts will be markedly decreased upon NANOG depletion. Hence, we focused on these decreased RBPs to understand their potential roles in pluripotency regulation. First, we compared them with the chromatin-associated RBPs (chrRBPs) and mRNA interactome datasets in mESCs24,55, which represent nuclear transcription-related RBPs and RBPs binding to polyadenylated RNAs, respectively. The comparison showed that 47 of them overlapped with chrRBPs while 50 overlapped with mRNA interactome dataset (Fig. 3a). We are particularly interested in the 47 overlapped chrRBPs as their chromatin association hints at potential roles in coordinating NANOG-mediated transcription or transcription-associated events. Hence, we performed a small-scale shRNA screen targeting these overlapping chrRBPs. Among the ten selected candidate RBPs, knockdown of three of them resulted in decreased Nanog gene expression and alkaline phosphatase activity, which were indicative of mESC differentiation (Supplementary Fig. 3a–d). In contrast, control shRNAs targeting luciferase (shCtrl) and a ubiquitously expressed RBP HNRNPM had no effect. Among these, we selected UTP15 for further study as it showed a consistent decrease in binding to nascent RNAs transcriptionally regulated by NANOG, as quantified by mass spectrometry (Fig. 3b). UTP15 is a component of the SSUP complex regulating ribosomal RNA (rRNA) transcription and processing56,57,58. However, its role in maintaining pluripotency and its reciprocal regulation with MTFs in ESCs remain unknown. To understand this, we first performed Photoactivatable Ribonucleoside-Enhanced Cross-linking and Immunoprecipitation (PAR-CLIP) chemiluminescent labeling, which also showed a decreased binding of UTP15 to nascent RNAs upon NANOG depletion (Fig. 3c left panel). In contrast, its protein level and binding capacity to steady-state RNA seem not affected (Fig. 3c right panel and Supplementary Fig. 3e).
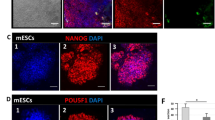
a Overlap of decreased proteins (n = 106) with chromatin-related RNA-binding proteins (chrRBPs, n = 512)24 and poly (A) RBPs (n = 555)55. b Box plots showing the LFQ intensity (log2) of UTP15 in the RICK experiment at different time points (n = 3 RICK experiments, box plots show the median (centre line), 25th–75th percentiles (box bounds), and whiskers extending to 1.5 × the interquartile range (IQR), two-sided Student’s t-test). c Representative PAR-CLIP-biotin chemiluminescent nucleic acid detection showing the newly transcribed RNA (labeled with 4sU) binding activity of UTP15 (left panel). Representative PAR-CLIP-biotin chemiluminescent nucleic acid detection showing the steady-state RNA binding activity of UTP15 (right panel). A Western blot was used to confirm the protein size and comparable loading of samples. Representative western blots are shown from two independent experiments with consistent results. d Relative expression of pluripotency and differentiation marker genes upon Utp15 knockdown. (mean ± SD, n = 3, two-sided Student’s t-test, * p < 0.05, ** p < 0.01, *** p < 0.001). e The schematic diagram showing the strategy of UTP15-AID knock-in. f Wildtype or truncated UTP15 proteins (upper panel) were reintroduced into mESCs after IAA-induced UTP15 degradation. The line graph showing the result of cell proliferation in UTP15-AID cells, which were rescued by UTP15 variants (lower panel). g MA plots showing the fold change (log2) of spike-in total RNA-seq after IAA-induced UTP15 degradation. 24 h versus 0 h (left), 48 h versus 0 h (right). (FDR < 0.05, fold change > 1.5). h Gene Set Enrichment Analysis (GSEA) analysis of embryonic genes in spike-in total RNA-seq after acute UTP15 degradation. 24 h versus 0 h (left), 48 h versus 0 h (right). i Immunostaining of GATA6 (green), NANOG (red) and CDX2 (magenta) at E3.5 embryos. Zygotes were microinjected with control siRNA or Utp15 siRNA and cultured in vitro to E3.5 stage. Nuclei were stained with Hoechst 33342 (blue). Merge means GATA6 merged with NANOG, CDX2, and Hoechst. BF (Bright Field). Scale bar = 50 μm. Representative images are shown from two independent experiments with consistent results. Source data for Fig. 3b, c, d and f are provided as a Source Data file.
We then constructed another shRNA targeting Utp15 to confirm its roles in mESC pluripotency. As expected, knockdown of Utp15 using two different shRNAs showed coherent downregulation of pluripotency genes (e.g., Nanog, Prdm14 and Pou5f1) and upregulation of differentiation genes (e.g., Gata6 and Tfap2c) (Fig. 3d). Moreover, we constructed a homozygous knock-in mES cell line bearing AID, Strep-tag II and FLAG tags in C-terminus of UTP15 (Fig. 3e). Addition of IAA induced acute UTP15 degradation accompanied with downregulation of Nanog, decreased cell proliferation and alkaline phosphatase activity (Fig. 3f, Supplementary Fig. 3f-h). RNA-seq confirmed that UTP15 depletion decreases pluripotent and embryonic gene expression59, leading to pluripotency exit as well as cell differentiation (Fig. 3g, h, Supplementary Fig. 3i and Supplementary Data 3). Importantly, reintroduction of full-length UTP15 rescued such effects while the truncated ones failed to (Fig. 3f).
Next, we asked whether UTP15 regulates embryonic pluripotency under physiological conditions. To achieve this, we performed microinjection of siRNA against Utp15 into mouse zygotes. Interestingly, we observed an obvious arrest of embryonic development compared to that of scramble siRNA control (Supplementary Fig. 3j). Moreover, we noticed a reduction of NANOG level as well as an increase in GATA6 expression in early blastocyst stage upon Utp15 knockdown (Fig. 3i), which is consistent with the observation in cultured mESCs (Fig. 1f, Fig. 3g). However, whether UTP15 coordinates the establishment of “salt-and-pepper” pattern through balancing Nanog and Gata6 expression in early embryonic development requires further investigation.
Taken together, UTP15 plays vital roles in the regulation of pluripotency in both cultured mESCs and embryonic development.
UTP15 is recruited to chromatin to activate gene transcription
Considering the absence of existing UTP15 CLIP-seq data, we performed CLIP-seq on UTP15 to investigate its roles in regulating mESC pluripotency. We observed that UTP15 displays substantial binding to rRNA, with preference for the 5’ETS region (Supplementary Fig. 4a). This finding is consistent with its well-known role as a constituent of SSUP. More intriguingly, our analysis of protein-coding genes revealed that UTP15 is predominantly localized to RNA regions in the vicinity of transcription start sites (TSSs), and its binding does not exhibit sequence specificity (Fig. 4a, Supplementary Fig. 4b and Supplementary Data 3). The latter observation is consistent with what has been observed for many other chrRBPs24, and also points to a possibility that UTP15 may be recruited to chromatin by transcribing RNAs or other factors to activate or inactivate cognate gene transcription24. We first validated the localization of UTP15 to chromatin through Western blot following the chromatin extraction (Supplementary Fig. 4c)24,60. Moreover, we conducted ChIP-seq on UTP15, which also showed a prominent binding to the promoter loci of many genes (Fig. 4b, Supplementary Fig. 4d and Supplementary Data 3). Further analysis showed that the binding sites of UTP15 to chromatin and RNA are enriched around each other, with a significant overlap to each other (Fig. 4c, Supplementary Fig. 4e, f). This highly correlated binding pattern supports the notion that UTP15’s interaction with chromatin may be RNA-dependent. To test this, we performed ChIP-seq on UTP15 with adding actinomycin D (ActD) to block RNA transcription. Indeed, the binding capacity of UTP15 to the TSSs of its chromatin targets is significantly decreased upon ActD treatment (Fig. 4d, Supplementary Fig. 4g). This was also confirmed by ChIP-qPCR for some selected targets including key pluripotency factors (e.g., Esrrb and Lin28a) upon ActD or Triptolide (TPL) treatment (Supplementary Fig. 4h). Hence, our results suggest that UTP15 is potentially recruited to chromatin by transcribing RNAs, which include some pluripotency genes.
a Average density profiles of sense and antisense CLIP read at the transcription start sites (TSSs) of protein-coding genes (n = 22,470). b Distribution of UTP15 ChIP-seq binding sites. c Average reads density (top) and heatmap (bottom) analysis of UTP15 CLIP-seq peaks around the center of UTP15 ChIP peaks (left panel). Average reads density (top) and heatmap (bottom) analysis of UTP15 ChIP-seq peaks around the center of UTP15 CLIP peaks (right panel). d Box plots represent UTP15 ChIP-seq signal intensity (log2) at promoter regions of UTP15 ChIP target genes (n = 7,091 genes) after two hours of treatment with ActD (replicate 1). Box plots show the median (centre line), 25th–75th percentiles (box bounds), and whiskers extending to 1.5 × IQR; data points beyond the whiskers are considered outliers (not shown). Two-sided Mann-Whitney U test (left). Average density profiles of UTP15 ChIP-seq density at the TSSs of UTP15 ChIP target genes after ActD treatment (right). e Volcano plot showing the distribution of proteins captured by UTP15 Co-IP. IgG was used as a control. (p value < 0.05, fold change > 1.5). f Dual-luciferase reporter assay in 293T cells. GAL4 DNA-binding domain was fused to UTP15; 5 × upstream activation sequence (UAS) was added upstream of SV40 promoter. GAL4 DNA-binding domain alone was used as control. (mean ± SD, n = 3 biological replicates, two-sided Student’s t-test, **** p < 0.0001). GSEA analysis of embryonic genes (g) and ESC-high genes (h) in nascent RNA-seq after acute UTP15 degradation (4 h versus 0 h). i Nascent RNA qPCR analysis of pluripotency genes after acute UTP15 degradation. Relative gene level was normalized with spike-in RNA. (mean ± SD, n = 3 biological replicates, two-sided Student’s t-test, * p < 0.05, ** p < 0.01). j Nascent RNA qPCR analysis of pluripotency genes and 5’-ETS upon reintroduction of full-length UTP15 and NoLS mutant during acute UTP15 depletion. Relative RNA level was normalized to spike-in RNA (fold change of IAA 4 h versus 0 h). (mean ± SD, n = 4 biological replicates, two-sided Student’s t-test, ** p = 0.0093). k Nascent RNA qPCR analysis of pluripotency genes and 5’-ETS after acute POLR1A degradation. (mean ± SD, n = 4 biological replicates, two-sided Student’s t-test, * p = 0.0171). l Box plots represent POLR1A-AID PRO-seq signals for 685 UTP15 target genes following POLR1A degradation (n = 685 genes, box plots show the median (centre line), 25th–75th percentiles (box bounds), and whiskers extending to 1.5 × IQR, two biological replicates). Source data for Fig. 4f, i, j and k are provided as a Source Data file.
To further understand the roles of UTP15 on chromatin, we conducted coimmunoprecipitation (co-IP) followed by mass spectrometry using FLAG antibodies in UTP15-AID mESCs (Fig. 3e) to enrich UTP15-interacting proteins. The analysis showed that UTP15 forms strong associations with other SSUP components (e.g., WDR43, UTP6, and UTP18) and several transcriptional activators, including LIN28A and FUS (Fig. 4e, Supplementary Data 2). The interactions with transcriptional activators suggest an activation effect of UTP15 in transcription in ESCs24,61,62. To investigate this, we first constructed a GAL4 reporter luciferase system for transcriptional analysis (Fig. 4f, upper panel)60. Unsurprisingly, tethering of UTP15 to a synthetic promoter directly increases the reporter luciferase activity (Fig. 4f, lower panel). Next, we conducted nascent RNA staining after EU labeling, which also showed significantly decreased signals when UTP15 was depleted (Supplementary Fig. 4i, j). To further identify specific transcriptional targets of UTP15, we also performed nascent RNA-seq upon acute UTP15 degradation. Interestingly, we observed a slight reduction in overall RNA transcription when IAA was added for four hours, while this effect becomes more pronounced after 24-hour IAA treatments (Supplementary Fig. 4k). Gene Set Enrichment Analysis (GSEA) also showed a significant reduction of transcription in embryonic gene set59 and ESC-high gene set60 at four hours of IAA treatments (Fig. 4g, h). More specifically, we identified 685 transcriptionally downregulated genes at four hours of IAA treatments, which exhibited significant enrichment of GO terms related to translation, cell proliferation, and stem cell population maintenance (Supplementary Fig. 4l). These genes are exemplified by Esrrb, Zic5, and Pou5f1, which are consistent with the total RNA-seq upon UTP15 degradation (Fig. 3g). Downregulation of these pluripotency-related genes was also confirmed by RT-qPCR analysis on isolated nascent RNA upon acute UTP15 degradation (Fig. 4i). Moreover, these representative genes are also bound by UTP15 at their promoters, which shows significant decrease upon ActD or TPL treatment (Supplementary Fig. 4h). We then termed these 685 genes UTP15 target genes for further analysis.
Given that UTP15 is a component of the SSUP complex and its binding to 5’-ETS of rRNA, we explored whether its impact on target mRNA transcription relies on rRNA. To this end, we measured both rRNA and representative UTP15 target gene transcription using RT-qPCR analysis on isolated nascent RNA after reintroduction of full-length UTP15 and its variants upon acute UTP15 depletion. Indeed, full-length UTP15 restored both rRNA and pluripotency gene transcription. However, the NoLS mutant, which lacks the nucleolar localization signal (NoLS, amino acids 318 to 337)63,64, rescues the transcription of pluripotency genes but not rRNAs (Fig. 4j). These results indicate that the regulation of the UTP15 target gene transcription by UTP15 is rRNA-independent. To further clarify this, we also constructed a POLR1A-dTAG (the largest subunit of Pol I) cell line. We observed a significant reduction in rRNA transcription upon acute Pol I depletion, while the UTP15 target gene transcription remained unaffected (Fig. 4k, Supplementary Fig. 4m). Consistent with this, a reanalysis of gene transcription using precision run-on sequencing (PRO-seq) datasets65 also showed a significant decrease in rRNA transcription upon Pol I depletion, with no decreased effect on the UTP15 target genes (Fig. 4l).
Taken together, these results suggest that UTP15 is recruited to chromatin to activate a subset of gene transcription, including mESC pluripotency genes, independent of rRNA.
UTP15 regulates a subset of NANOG-target gene transcription
To further elucidate how UTP15 activates gene transcription, we focused on analyzing the 685 UTP15 target genes (Supplementary Data 4). Initially, we observed that these UTP15 targets exhibit more open chromatin accessibility, characterized by significant enrichment of H3K4me3 and H3K27ac marks, and notably, stronger UTP15 as well as NANOG bindings compared to randomly selected genes (Fig. 5a). Furthermore, the transcription of UTP15 targets showed a significant decrease upon the acute degradation of NANOG (Fig. 5b, Supplementary Fig. 5a, b). Sequencing tracks for three representative UTP15 target genes (i.e., Zfp57, Zic5 and Zic2) are depicted in Fig. 5c. Similarly, NANOG transcriptional targets, which exhibit greater chromatin accessibility and higher enrichment of H3K4me3 and H3K27ac, also show stronger UTP15 binding compared to randomly selected genes (Supplementary Fig. 5c). CLIP-seq analysis further revealed that UTP15 binds more extensively at the TSS regions of NANOG transcriptional targets (Supplementary Fig. 5d). Additionally, NANOG transcriptional targets had a more significant overlap with UTP15 ChIP-seq binding genes compared to randomly selected genes (Supplementary Fig. 5e). These findings suggest that UTP15 may work in coordination with NANOG to regulate gene transcription in mESCs.
a Average reads density (top) and heatmap (bottom) analysis of ATAC-seq, H3K4me3, H3K27ac83, UTP15 ChIP rep1, UTP15 ChIP rep2, and NANOG ChIP46 in the region from −2 kb upstream of TSS to +2 kb downstream of the TES across UTP15 target genes (n = 685). Random genes were random selected from outside of UTP15 target genes (n = 685). b Box plots showing normalized read counts (log₂) of UTP15 target genes (n = 685 genes) from nascent RNA-seq in UTP15-AID (left panel) and NANOG-AID (right panel) cells treated with IAA. (two biological replicates). Box plots represent the median (centre line) and 25th-75th percentiles (bounds); whiskers extend to 1.5 × IQR. Two-sided Mann–Whitney U test. c IGV snapshot of UTP15 CLIP-seq, UTP15 ChIP-seq, NANOG ChIP-seq (GSM4231200), Pol II ChIP-seq (UTP15-AID after treatment with IAA) and nascent RNA-seq at Zfp57, Zic5 or Zic2 gene locus. d Average reads density (top) and heatmap (bottom) analysis of UTP15 ChIP-seq at the TSSs across UTP15 target genes (n = 685) after IAA induced NANOG degradation. Random genes were random selected from outside of UTP15 target genes (n = 685). e Box plots represent UTP15 ChIP-seq signal intensity (log2) at promoter regions of random genes (left, n = 685 genes) and UTP15 target genes (right, n = 685 genes) in NANOG-AID cells before (-IAA) and after (+IAA) NANOG degradation. Results are shown for two biological replicates (Rep1 and Rep2). Random genes, equal in number to the UTP15 target genes, were randomly selected from non-target regions. Box plots represent the median (centre line) and the 25th–75th percentiles (bounds); whiskers extend to 1.5 × IQR, and data points beyond the whiskers are considered outliers (not shown). Two-sided Mann-Whitney U test. f Western blot showing FLAG-mediated UTP15 co-IP in UTP15-AID cell lines under formaldehyde crosslinking conditions. The asterisk (*) denotes the purpose band. Representative western blots are shown from two independent experiments with consistent results. g Co-immunostaining of UTP15 and NANOG in UTP15-AID cells. Endogenous UTP15 protein was visualized using an anti-FLAG antibody. A magnified view of the indicated region is shown on the right. Scale bar, 5 μm. Representative immunofluorescence images are shown from two independent experiments with consistent results. Source data for Fig. 5f are provided as a Source Data file.
To further understand the coordination between UTP15 and NANOG, we first performed ChIP-seq for UTP15 following IAA treatment in NANOG-AID mESCs to assess whether NANOG depletion affects UTP15’s binding to its target genes on chromatin. Notably, acute degradation of NANOG reduces the binding of UTP15 at the promoters of its targets along with a decrease in their transcriptional levels (Fig. 5b, d, e and Supplementary Fig. 5f). This raises a possibility that the coordination between NANOG and UTP15 is potentially mediated by NANOG transcriptional targets or other factors rather than NANOG itself, as UTP15 co-IP failed to capture NANOG (Fig. 4e). Supporting this, proximity labeling using UTP15 bait indeed captures NANOG (Supplementary Fig. 5g). In consistence with this, we also found that NANOG can be captured using UTP15 as a bait via co-IP under mild crosslinking conditions with 0.1% formaldehyde (Fig. 5f). Moreover, Immunofluorescence analysis also revealed a partial co-localization of NANOG and UTP15 (Fig. 5g). Addition of ActD to block RNA transcription reduces the binding of UTP15 at the promoters of its targets (Supplementary Fig. 5h). These observations collectively suggest that the coordination between NANOG and UTP15 might be indirect, potentially mediated via components of the transcriptional machinery, or nascent transcripts. Of note, among the 685 UTP15 target genes, 223 (~32.6%) showed significant downregulation upon NANOG depletion (Supplementary Fig. 5i). These genes include classical NANOG targets such as Esrrb (Supplementary Fig. 5j, left). In contrast, the number of genes (17, ~2.5%), which are upregulated upon NANOG depletion was negligible (Supplementary Fig. 5j, right).
Taken together, these results indicate that UTP15 may regulate a subset of NANOG-targeted genes by cooperating with NANOG to enhance their transcription in mESCs.
UTP15 supports the formation of Pol II condensates
Next, we focused on elucidating the role of UTP15 in the transcriptional activation of its target genes in mESCs. Considering that UTP15 interacts with a series of proteins related to chromatin organization (e.g., SMARCA4), chromatin remodeling (e.g., BRD3) as well as component of the Pol II core complex (e.g., POLR2B) (Fig. 4e, Supplementary Fig. 6a), we envisioned that UTP15 may interact with Pol II core complex to directly alter Pol II-mediated transcription in mESCs. To test this, we first performed Western blot following Co-IP, which confirmed the interaction between UTP15 and POLR2A (Fig. 6a). This was further verified by proximity labeling approach (Supplementary Fig. 6b). Next, we aimed to assess the impact of UTP15 degradation on Pol II binding to chromatin through ChIP-seq analysis. Our findings revealed a reduction in Pol II binding at both the transcription start sites (TSSs) and gene bodies of UTP15-targeted genes following UTP15 degradation (Fig. 6b, Supplementary Fig. 6c). This reduction is corroborated by our nascent RNA-seq data, which also showed decreased transcriptional activity at both the TSS and gene body regions (Supplementary Fig. 6d). These results support the notion that UTP15 is a key regulator of Pol II transcription in mESCs. To delve deeper into UTP15’s influence on transcription, we performed ChIP-seq analysis on RNA Pol II Ser5P, an indicator of transcriptional initiation. This analysis showed that UTP15 depletion results in a diminished presence of RNA Pol II Ser5P at the TSSs of its target genes, suggesting its role in the transcriptional initiation (Fig. 6b, Supplementary Fig. 6c). Additionally, the levels of RNA Pol II Ser2P, a marker of transcriptional elongation, were also significantly reduced in the gene body regions of UTP15-targeted genes upon acute UTP15 degradation (Fig. 6b, Supplementary Fig. 6c). While these findings strongly suggest UTP15’s involvement in transcriptional initiation, UTP15 may also involve in transcriptional elongation. Collectively, our data indicate that UTP15 activates the transcription of its target genes by modulating Pol II binding to chromatin in mESCs.
a Western blot showing HA-mediated UTP15 co-IP in POLR2A-FLAG knock-in cell lines. Representative western blots are shown from two independent experiments with consistent results. b Average read density analysis of Pol II ChIP-seq (NTD, Ser5, Ser2) after IAA induced acute UTP15 degradation in the region from −2 kb upstream of TSS to +2 kb downstream of the TES across UTP15 target genes (n = 685). c Prediction of disorder regions for UTP15 using PONDR. The sequence of UTP15 was submitted to the PONDR server, which predicted regions of disorder with the VL-XT, VL3, VSL2 predictor. d Droplet formation assays of varying concentrations of GFP and GFP-UTP15IDR in the presence of 16% dextran. Scale bar, 10 μm. e Quantification of nuclear UTP15 signal intensity in control and 1,6-HD treated cells (Ctrl: n = 18; 1,6-HD: n = 28, n represents individual nuclei analyzed). Cells were treated with 3% 1,6-HD for 10 minutes. Box plots show the median (centre line) and 25th-75th percentiles (bounds); whiskers extend to 1.5 × IQR. Statistical significance was assessed using a two-sided Mann-Whitney U test. f Representative immunofluorescence images of UTP15 (green) in UTP15-AID cells following treatment with 3% 1,6-HD for 10 min or control (untreated) conditions. Scale bar, 5 μm. g, h Droplet formation assays of mCherry-CTD (10 μM) with GFP (10 μM) or GFP-UTP15IDR (10 μM) in the presence of 16% dextran. The incubation time was increased from 30 min to 150 min at room temperature. Quantification and representative pictures are shown in h and g, respectively. In h, y axis shows the sum of fluorescence intensity of mCherry-CTD within droplets in each field (n = 5, n represents individual image analyzed, two-sided Student’s t-test). Scale bar, 10 μm. i Representative wide field and SIM² super-resolution images showing Pol II-NTD (green) and UTP15 (red) in UTP15-AID cells after IAA induced UTP15 degradation. Scale bar, 5 μm. j Quantification of nuclear Pol II clusters per cell using Laplace of Gaussian (LoG) filter method under different durations of IAA treatment (n = 20 per group, n represents individual nuclei analyzed). Box plots show the median (centre line) and 25th-75th percentiles (bounds); whiskers extend to 1.5 × IQR. Statistical significance was assessed using a two-sided Mann-Whitney U test. Source data for Fig. 6a, e, h and j are provided as a Source Data file.
UTP15 features an intrinsic disorder region (IDR) at its carboxyl terminus, as predicted by Predictor of Natural Disordered Regions (PONDR), which may mediate formation of biomolecular condensates66 (Fig. 6c). To test this, we purified a GFP-tagged UTP15 IDR fragment (UTP15IDR, amino acids 314 to 528) from bacteria (Supplementary Fig. 6e). The purified GFP-tagged UTP15 IDR fragment is functionally intact, as evidenced by its binding capacity to RNA probes corresponding to UTP15’s binding site on the 5’ETS (Supplementary Fig. 6f). In contrast, the purified GFP does not bind to RNA. Our further experiments showed that GFP-UTP15IDR forms spherical droplets, with an increase in both number and size as the concentration rises under physiological salt conditions (Fig. 6d). Importantly, the addition of 1,6-Hexanediol (1,6-HD) disrupts the droplet formation (Supplementary Fig. 6g). Moreover, we noticed that UTP15 showed a recovery after photobleaching in mESCs (Supplementary Fig. 6h). Treatment of mESCs with 1,6-HD also affect the condensates of UTP15 (Fig. 6e, f). These suggest UTP15 is capable of forming biomolecular condensates both in vitro and in vivo.
Next, we asked whether UTP15’s capacity of forming biomolecular condensates would affect POLR2A during gene transcription. The C-terminal domain (CTD) of POLR2A composed of heptapeptide repeats, is known to form protein clusters involved in various transcriptional processes67,68,69. We asked whether UTP15 could influence CTD clustering, thereby modulating Pol II transcription through its IDR. Indeed, incubating UTP15IDR with CTD revealed that UTP15IDR enhances both the speed and the size of POLR2A CTD cluster formation (Fig. 6g, h). Moreover, we observed that POLR2A partially colocalizes with UTP15 and its NoLS mutant in mESCs (Supplementary Fig. 6i). More importantly, acute depletion of UTP15 reduces the number of Pol II cluster in mESCs (Fig. 6i, j). Hence, our observations suggest that UTP15 may play a role in transcriptional regulation through facilitating Pol II clustering in mESCs.
In summary, our findings suggest that UTP15 may be recruited by NANOG transcriptional target genes to chromatin, promoting active transcription and sustaining pluripotency in mESCs. This process is potentially facilitated through Pol II-UTP15-mediated phase separation. Thus, loss of either NANOG or UTP15 disrupts pluripotency and predisposes cells towards a differentiation state (Fig. 7).
Acting as a master transcription factor, NANOG initiates the transcription of pluripotency genes. The newly synthesized RNA recruits UTP15 to chromatin at transcription start sites (TSSs). There, UTP15 supports the formation of transcription condensates by RNA polymerase II (Pol II). This UTP15-mediated process further amplifies and sustains the transcription of pluripotency genes.
Discussion
Previous studies on MTF in mESCs predominantly utilized traditional knockout or knockdown methods and relied on steady-state RNA-seq analysis41,70. These approaches were unable to precisely identify the direct transcriptional targets of MTF, as some observed changes in gene expression were attributed to mESC differentiation that occurred concurrently with MTF depletion. Our approach, in contrast, is more advanced by employing EU-based capture of newly transcribed RNAs together with acute protein degradation. This method takes into account that total RNAs could display different changes compared to newly transcribed RNAs in response to specific stimuli71,72 or acute MTF degradation, as observed in our study (Fig. 1d). A possible explanation for the discrepancies between total RNA and newly transcribed RNA could involve specific RBPs that interact with newly transcribed RNAs, participating in processes such as RNA splicing, degradation, or other regulatory functions. In this regard, our study, to the best of our knowledge, first globally captured RBPs interacting with newly transcribed RNAs modulated by acute MTF depletion. Through this, we have identified a series of RBPs that may be engaged in the regulation of pluripotency. Further studies are needed to elucidate the roles of these RBPs in RNA metabolism and pluripotency within mESCs. Taken together, our strategy not only deepens our understanding of the regulation of MTF-regulated transcripts from a different perspective (i.e., transcribing RNA-protein interaction), but also opens up the possibility of applying this strategy to various cellular contexts, such as MTF studies in hematopoietic stem cells, myogenic stem cells, and cancer research.
Our nascent RNA-seq data showed that acute depletion of UTP15 leads to a rapid downregulation of pluripotency gene transcription within as short as four hours. Yet, the immediate impact of UTP15 on transcription appears to be relatively smaller (685 genes versus 1,812 genes) compared to that of NANOG (Fig. 1b, Supplementary Fig. 5a), suggesting that other RBPs may also sense and regulate the NANOG transcriptional targets. In this regard, RBPs are pervasively present on chromatin and activate or repress gene transcription. Of note, Shao et al. have shown that the IDRs, which are generally enriched in RBPs, facilitates the formation of transcriptional condensates to activate gene transcription. Similarly, we observed that UTP15 contains an IDR in its C terminus, which also mediates LLPS and enhance the formation of condensates of POLR2A CTD. However, the observation that UTP15 exhibits relatively slower fluorescence recovery after photobleaching suggests the involvement of complicated networks in biomolecular condensate formation (Supplementary Fig. 6h). This could be mediated by percolation through sticker-and-spacer architectures, rather than by simple liquid-liquid phase separation25,26,27. In this regard, the precise role of UTP15 in gene transcription or rRNA biogenesis remains to be further elucidated. Additionally, the interplay between these RBPs and NANOG in the regulation of gene transcription within mESCs is also a subject of ongoing interest and requires further investigation. In this context, Oksuz et al. have recently demonstrated that NANOG is a potential RBP73. This also raises another important question regarding which RNAs are bound by NANOG, and whether these bindings have a synergistic effect with the identified RBPs in the regulation of pluripotency gene transcription.
UTP15 is a component of the SSUP complex, suggesting a potential role in regulating pluripotency via ribosome biogenesis-related mechanisms. However, our study strongly indicates that UTP15 primarily modulates Pol II-mediated transcription activation independent of ribosome biogenesis. Firstly, UTP15’s binding to nascent RNA diminishes progressively following acute degradation of NANOG, which does not impact rRNA transcription (Supplementary Fig. 6j). This suggests that UTP15’s observed reduction in binding to newly transcribed RNAs are likely due to its association with Pol II-transcribed RNAs rather than rRNAs. Secondly, the acute depletion of UTP15 leads to an immediate reduction in the transcription of a subset of NANOG target genes within as short as four hours, coinciding with an immediate decrease in Pol II’s binding to the target genes of UTP15. Thirdly, we also observed that the NoLS mutant is capable of restoring the transcription of pluripotency-associated genes, but fails to rescue rRNA transcription, suggesting a separation of UTP15’s roles in pluripotency gene transcription and rRNA transcription. Fourthly, UTP15 interacts with POLR2A and facilitates POLR2A CTD cluster formation through its IDR region both in vitro and in vivo. Moreover, WDR43 and DDX21, two other SSUP complex components, have been reported to regulate Pol II transcription upon its depletion60,74. Taken together, these lines of evidence collectively support the hypothesis that UTP15 directly modulates Pol II-mediated transcription. Nevertheless, whether UTP15 functions independently or in concert with WDR43, DDX21 and other UTP15 interacting RBPs as part of a complex remains to be further explored.
In addition, an increasing body of evidence underscores the nucleolus’s role in shaping genome architecture and regulating transcription, thus expanding its importance beyond ribosome biogenesis75. For example, circular chromosome conformation capture sequencing (4C-seq) in the MYC-driven malignant transformation model has shown that genes associated with the nucleolar-associated domain (NAD) are enriched for growth and metabolic gene ontologies among the actively transcribed genes76. This finding suggests that the dynamics of rDNA interactions may also have functional implications for the regulation of Pol II transcriptional activity, highlighting an unknown interplay between Pol I and Pol II-mediated transcription. In this regard, we found that some LINE1 retrotransposons were downregulated upon acute NANOG depletion, while no changes occurred upon UTP15 depletion (Supplementary Fig. 6k). Since LINE1 can associate with Nucleolin to boost rRNA synthesis in mESCs77, this implies NANOG might also act as a bridge to regulate Pol II and Pol I interplay. Hence, further studies may also focus on illustrating the interplay between Pol I and Pol II to reveal unrecognized roles of UTP15 and NANOG in promoting mESC pluripotency or self-renewal.
Methods
Cell culture
Mouse embryonic stem cell line E14TG2a was cultured with serum/LIF medium containing high-glucose Dulbecco’s modified Eagle’s medium (DMEM, HyClone), 15% fetal bovine serum (FBS, Lonsera), 1 × GlutaMAX (Gibco), 1 × nonessential amino acids (NEAA, Gibco), 1 × sodium pyruvate (Gibco), 1 × penicillin-streptomycin (HyClone), 1 mM 2-mercaptoethanol (Gibco), and 1000U ml-1 Leukemia inhibitory factor (LIF, Origene) on plates coated with 0.2% gelatin. HEK293T cells were grown in DMEM (HyClone) containing 10% fetal bovine serum (FBS, Lonsera) and 1 × penicillin-streptomycin (HyClone).
Cell line generation
UTP15-AID cells were generated using CRISPR-Cas9 as described in ref. 78the gRNA sequences and strategy are shown in Supplementary Data 5 and Fig. 3e, respectively. Briefly, the donor vector and vector for expression of gRNA and Cas9 were co-transfected into E14TG2a cells. Cells were selected with hygromycin. POLR2A-FLAG and POLR1A-dTAG cell lines were generated using the same approach. Short hairpin RNA (shRNA) knockdown cells were generated by lentiviral infection and puromycin selection. shRNA constructs and packaging vectors (PMD2.G and psPAX2) were co-transfected into HEK293T cells. The lentivirus particles were harvested at 48 h-72 h after transfection. After filtration through a 0.45 μm filter (Millipore) and concentrated by 100 kDa ultrafiltration centrifuge, the supernatants were stored at −80 °C.
Nascent RNA-sequencing
The cells at 70-80% confluency were treated with 1 mM EU for 15 min; the EU labeling was stopped by direct addition of TRI Reagent (MRC) to cells. Total RNA was isolated and treated with DNase I, EU-labeled spike-in RNAs were added to each sample in equal amounts. The RNA was then biotinylated with click chemistry and fragmented with NaOH. After ethanol precipitation, the fragmented RNA was resuspended in nuclease-free water and separated with Myone Streptavidin C1 Dynabeads (Invitrogen). The enriched EU-labeled RNAs were ligated with pre-adenylated 3’-adapter using T4 RNA ligase 2 (Cat#: M0373L) and reverse-transcribed on a bead. The cDNA was purified for next-generation sequencing library construction. For nascent RNA qPCR, enriched RNAs were reverse-transcribed with random primers (TaKaRa) and ReverTra Ace® (Toyobo) according to the manufacturer’s instructions. The relative gene level was normalized with the spike-in RNA.
Capture of the newly transcribed RNA interactome using click chemistry (RICK)
Cells were cultured with EU for 2 h at different time points after indole-3-acetic acid (IAA) treatments. After washing three times with cold PBS, the plates were placed on ice and irradiated with 0.15 J/cm2 UV light at 254 nm. Cells were then lysed in lysis buffer (20 mM Tris-HCl, pH 7.5, 500 mM LiCl, 0.5% lithium-dodecyl sulfate) supplemented with protease inhibitor cocktail (Roche) and RNase inhibitor (Thermo). Lysates were homogenized using a syringe with a narrow needle (29 G). The cell lysates were diluted 100 times with 50 mM Tris-HCl (pH 7.5) and concentrated by using an Amicon Ultra-15 centrifugal filter unit (Millipore). 100 μM azide-PEG-biotin, 500 μM CuSO4, 2 mM THPTA, 1 mM aminoguanidine and 5 mM sodium ascorbate (fresh) were added to the lysates for click reaction at room temperature with rotation (2 h), followed by adding 5 mM EDTA to stop the reaction. After methanol precipitation, the pellets were dissolved in reconstitution buffer (0.5% sodium dodecyl sulfate, 1 mM EDTA, 1% Brij-97, 150 mM NaCl, 50 mM triethanolamine supplemented with 200 U ml-1 RNase inhibitor). Then the soluble fraction from the lysate was incubated with streptavidin agarose beads at 4 °C overnight. Then the RNA-bound agarose beads were collected and washed twice with buffer Ⅰ (2% SDS in PBS), buffer Ⅱ (8 M urea and 0.25 M NH4HCO3 in water), buffer Ⅲ (2.5 M NaCl in PBS) and 50 mM Tris-HCl (pH 7.5) in a sequential order for 10 min respectively. The resultant proteins eluted from the beads were digested for mass spectrometry. RICK experiments were performed at each time point using no-EU treatment as the control. The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE79 partner repository with the dataset identifier PXD043911.
Western blot
Proteins were loaded onto SDS-PAGE gel for separation. After transferring onto PVDF membrane (Millipore), they were blocked for 1 h with 5% milk in TBST (0.1% Tween-20 in tris-buffered saline), and incubated with the primary antibodies at 4 °C overnight. After three washes with TBST, the membranes were incubated for 2 h at room temperature with HRP-conjugated secondary antibodies. After three washes with TBST again, the protein signals were generated with ECL (Millipore) and detected with FluorChemE (ProteinSimple). The uncropped and unprocessed scans of the most important blots are provided in the Source Data file.
Antibodies for Western blot
NANOG antibody (Abcam, ab80892), UTP15 antibody (Novus, NBP2-58570), FLAG-M2 antibody (Sigma, F1804), GAPDH antibody (Proteintech, 10494-1-AP), ACTB antibody (SANTA CRUZ Biotechnology, sc-47778). Pol II S5P antibody (Abcam, AB5131), Pol II S2P antibody (Abcam, AB5095), Pol II NTD antibody (CST, 14958S).
Reverse transcription quantitative PCR (RT-qPCR)
Total RNA was isolated using TRI Reagent (MRC). Then the RNA was reverse-transcribed with Random primers (TaKaRa) and ReverTra Ace® (Toyobo) according to the manufacturer’s instructions. qPCR was performed on the QuantStudioTM 3 system using 2 × PerfectStart Green qPCR SuperMix (TransGen). All primers for qPCR were listed in Supplementary Data 5. Data were analyzed using the 2-ΔΔCt method.
RNA-seq
Total RNA was isolated using TRI Reagent (MRC). After DNase I treatments, rRNA was removed using QIAseq FastSelect-rRNA HMR Kit (QIAGEN). RNA fragmentation and construction of strand-specific RNA libraries using VAHTS Strand mRNA-seq Library Prep Kit for Illumina (Vazyme, Cat#: NR616). Sequencing was carried out on an Illumina NovaSeq 6000 platform.
Spike-in RNA-seq
UTP15-AID cells were counted after IAA treated 24 hours or 48 hours. Total RNA was isolated by TRI Reagent (MRC) and added ERCC RNA Spike-In Control Mixes (Cat#: 4456740) to RNA samples according to the manufacturer’s instructions. Then the poly (A) RNAs were enriched by NEBNext® Poly (A) mRNA Magnetic Isolation Module (Cat#: E7490S) according to the manufacturer’s instructions. Finally, the libraries were constructed by NEBNext® Ultra II Directional RNA Library Prep Kit for Illumina (Cat#: E7760S). Sequencing was carried out on an Illumina NovaSeq 6000 platform.
CLIP-seq and PAR-CLIP-biotin chemiluminescent nucleic acid detection
UV cross-linking and immunoprecipitation sequencing (CLIP-seq) and PAR-CLIP-biotin chemiluminescent nucleic acid detection were conducted as previously described33. Briefly, ten 10 cm-plates of UTP15-AID cells stably expressing FLAG-tagged UTP15 were irradiated with 254 nm UV light at 0.15 J/cm2. After crosslinking, the cells were collected by cell scraper (Greiner). Then the cell pellets were fully lysed followed by centrifugation. The supernatant was digested with RNase T1 (Thermo) and immunoprecipitated using FLAG M2 Affinity gel (Sigma). After that, the beads were isolated with centrifugation, washed, and digested with RNase T1 again. Then the beads were resuspended and treated with calf intestinal alkaline phosphatase (NEB). The protein-RNA complexes were now ready for RNA labeling or for CLIP-seq library construction. For CLIP-seq library, RNA co-immunoprecipitated with UTP15-FLAG was treated with T4 PNK (NEB), and then the RNAs were ligated with pre-adenylated 3’-adapter using T4 RNA ligase 2. And then the samples were washed, denatured in 1 × loading buffer and resolved on Novex Bis-Tris 4–12% gel, then transferred onto nitrocellulose membrane. Protein-RNA bands were excised from the membrane and digested with proteinase K (Vetec™). RNA was subsequently purified with acidic phenol: chloroform and ethanol precipitation. The resultant RNAs were reverse-transcribed by Hifair® III Reverse Transcriptase (Yeasen) with RT primer. The cDNA was purified and ligated with another adapter with T4 RNA ligase 1. Then the cDNA was purified again for next-generation sequencing library construction. The purified library was evaluated using the Agilent 2100 TapeStation and diluted for cluster generation and high-throughput sequencing (2 × 150 bp) on an Illumina NovaSeq 6000 platform.
Chromatin immunoprecipitation sequencing (ChIP-seq)
ChIP assays for UTP15 were performed as previously described60,80. Briefly, cells were crosslinked by 3% formaldehyde for 10 min at room temperature and quenched by 2.5 M glycine for 15 min. For nuclei isolation, cells were incubated with Cell lysis buffer (10 mM Tris-HCl pH 8.0, 10 mM NaCl, 10% Glycerol, 0.2% NP-40, 1 mM EDTA, 1 mM DTT, 1 × Proteinase inhibitor cocktail)for 20 min. Nuclei pellets were resuspended with Nuclear lysis buffer (50 mM Tris-HCl pH 8.0, 0.5% SDS, 10 mM EDTA, 1 mM DTT, 1 × Proteinase inhibitor cocktail) and incubated for 10 min. Lysate was sonicated by the Bioruptor plus (Diagenode) for 25 cycle (30 s ON/30 s OFF) at high intensity. Following centrifugation at 12,000 × g for 30 min. 50 μL lysate was saved as input, the remaining lysate was incubated with anti-FLAG M2 Affinity gel (Sigma) overnight. The beads were washed with low-salt wash buffer (20 mM Tris-HCl, pH 8.0, 150 mM NaCl, 2 mM EDTA, pH 8.0, 1% Triton X-100, 0.1% SDS supplemented with protease inhibitor), high-salt wash buffer (20 mM Tris-HCl, pH 8.0, 500 mM NaCl, 2 mM EDTA, pH 8.0, 1% Triton X-100, 0.1% SDS supplemented with protease inhibitor) and LiCl wash buffer (10 mM Tris-HCl, pH 8.0, 250 mM LiCl, 1 mM EDTA, pH 8.0, 1% NP-40, 0.1% deoxycholate supplemented with protease inhibitor). The bound chromatin was eluted and purified. The ChIP-seq library was constructed using NEBNext Ultra II DNA Library Prep Kit for Illumina (NEB), and sequenced on an Illumina NovaSeq 6000 platform. For UTP15 ChIP-seq after IAA-induced NANOG degradation and ActD treatments, a total 20% of cross-linked Drosophila S2 chromatin was spiked-in as a reference.
Pol II ChIP-seq
UTP15-AID Cells were treated with IAA for 4 hours and crosslinked with 1% formaldehyde for 10 min at room temperature. Crosslinking was quenched with 125 mM glycine for 5 min. Then cells were scraped and collected into tubes. For nuclei isolation, cells were incubated with Cell lysis buffer (10 mM Tris-HCl pH 8.0, 10 mM NaCl, 10% Glycerol, 0.2% NP-40, 1 mM EDTA, 1 mM DTT, 1 × Proteinase inhibitor cocktail, 1 mM Na3VO4)for 20 min. Nuclei pellets were resuspended with Nuclear lysis buffer (50 mM Tris-HCl pH 8.0, 0.5% SDS, 10 mM EDTA, 1 mM DTT, 1 × Proteinase inhibitor cocktail, 1 mM Na3VO4) and incubated for 10 min. Lysate was sonicated for 20 cycle (30 s ON/30 s OFF) at lower intensity using Bioruptor plus (Diagenode). Nuclear lysate was centrifuged with 18,000 × g for 30 min. Then nuclear lysate was diluted 1:5 with ChIP Dilution Buffer (12.5 mM Tris-HCl pH 8.0, 187.5 mM NaCl, 1.25% Triton X-100, 1 mM DTT, 1 × Proteinase inhibitor cocktail, 1 mM Na3VO4), 20 ng Spike-in Chromatin (Cat#: 53083) was added to 100 μg Chromatin. Take 1% nuclear lysate as input, the remaining lysate was incubated with antibody of interest and H2Av Spike-in antibody (Cat#: 61686) at 4 °C overnight. After adding Protein G Dynabeads (Thermo Fisher) into it, incubated the mixture with beads for another 2 hours. Beads were then washed once with low-salt wash buffer, twice with high-salt wash buffer, once with LiCl wash buffer, and once with TE buffer. For elution, beads were incubated twice with fresh elution buffer (50 mM Tris-HCl pH 8.0, 10 mM EDTA, 1% SDS, 100 mM NaHCO3, and 250 mM NaCl) for 20 min at 65 °C. To reverse crosslink, ChIP IP and input DNA were incubated at 65 °C overnight in elution buffer with RNase A, followed by the addition of Protease K and incubation for 2 hours at 56 °C. Finally, the ChIP-seq library was constructed using NEBNext Ultra II DNA Library Prep Kit for Illumina (NEB), and sequenced on an Illumina NovaSeq 6000 platform
Cell proliferation assay
UTP15-AID cells were infected with lentivirus expressing OsTIR1 and full-length or truncated UTP15. 2 × 105 cells were cultured with 6-well plates, and cells were counted at different time points after addition of IAA (Sigma) at a final concentration of 500 μM.
Alkaline Phosphatase staining (AP staining)
The BCIP/NBT Alkaline Phosphatase Color Development Kit (Vazyme) was used for the detection of the AP activity according to the manufacturer’s instructions. For AP staining, 5000 E14TG2a cells were infected with lentivirus expressing shRNA and cultured in 12-well plates for 2 days before staining. 5000 UTP15-AID cells with OsTIR1 expression were plated and cultured with or without IAA for 2 days before staining.
Co-Immunoprecipitation (Co-IP)
5 × 107 cells were collected and washed with cold PBS. The cells were resuspended and lysed in TNE lysis buffer (50 mM Tris-HCl pH 7.5, 250 mM NaCl, 0.5% NP-40 and 1 mM EDTA pH 8.0) supplemented with 1 × EDTA-free protease inhibitor cocktail (Roche) on ice for 15 min. The lysates were homogenized by a syringe with a narrow needle (29 G) and centrifuged at 13,000 × g for 15 minutes at 4 °C. The supernatant was transferred to a new tube and diluted it with an equal volume of TNEG buffer (50 mM Tris-HCl pH 7.5, 50 mM NaCl, 0.5% NP-40, 20% glycerol, and 1 mM EDTA pH 8.0). The supernatant was incubated with the pre-prepared protein A/G beads (which were conjugated to the selected antibodies or relevant IgG control) for 3 h at 4 °C. The beads were isolated on a magnetic stand, and washed with washing buffer (20 mM Tris-HCl pH 7.5, 125 mM NaCl and 0.5% NP-40) 3 times and subsequently with 1 × PBS once. The final protein complexes were eluted and digested for mass spectrometry. The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE79 partner repository with the dataset identifier PXD043911. For the Co-IP assay after formaldehyde crosslinking, UTP15-AID cells crosslinked with 0.1% formaldehyde, UTP15 were enriched using FLAG M2 Affinity gel (Sigma), followed by detection and analysis via Western blot. For HA-UTP15 co-IP, it was expressed in POLR2A-FLAG knock-in cell line, co-IP was performed using HA-tag Protein IP Assay Kit with Agarose Gel (Beyotime, Cat#: P2206S) according to the manufacturer’s instructions. For UTP15-APEX2, it was expressed in E14TG2a cells. Subsequently, labeling with biotin-phenol and H2O2 was carried out. Thereafter, streptavidin-mediated pull-down was performed, followed by western blot analysis.
Sample preparation for mass spectrometry
Different on-bead protein samples obtained from RICK or co-immunoprecipitation were resuspended with elution buffer (2 M urea, 100 mM Tris-HCl pH 8.5 and 10 mM DTT) and incubated in a shaker at speed of 1200 rpm for 20 min at room temperature, following with alkylation steps by adding with iodoacetamide (IAA, Sigma) at a final concentration of 50 mM for 10 min in the dark. The protein was partially digested with 250 ng of trypsin in the shaker (Promega) for 2 h, followed by an additional digestion with 100 ng of trypsin without shaking in the dark overnight. The digestion was quenched by adding TFA at the final concentration of 1%, followed by desalting on C18 StageTips.
Liquid chromatography-tandem mass spectrometry (LC-MS/MS)
Tryptic and desalted peptides were separated and analyzed by liquid chromatography-tandem mass spectrometry (LC-MS/MS) using an Easy-nLC 1200 system coupled online to an Orbitrap Eclipse mass spectrometer (Thermo Fisher Scientific). Peptides were separated on a 250-mm Acclaim PepMap100 C18 column (75 μm internal diameter; Thermo Fisher Scientific) with a 60-min linear gradient of mobile phase B (0.1% formic acid in 80% acetonitrile) versus mobile phase A (0.1% formic acid in water). The gradient was as follows: 4–20% B in 45 min, 20–36% B in 8 min, 36–100% B in 2 min, and 100% B held for 5 min. The flow rate was maintained at 300 nL/min. The mass spectrometer was operated in data-dependent acquisition mode. Full-scan MS1 spectra were acquired in the Orbitrap at a resolution of 120,000, with a normalized AGC target of 100% and a maximum injection time of 50 ms. Precursor ions with charge states between 2 and 7 were selected for MS/MS sequencing, with a dynamic exclusion window of 60 s. The MS2 spectra were generated by higher-energy collisional dissociation (HCD) at a normalized collision energy of 30% and analyzed in the ion trap, using a normalized AGC target of 100%, a maximum injection time of 10 ms, and an isolation window of 1.6 m/z. FAIMS was employed with compensation voltages (CVs) of −40 V, −60 V, and −80 V, and a cycle time of 1 s per CV.
Subcellular fractionation
UTP15-AID cells were crosslinked with 1% formaldehyde for 10 min, and quenched by 2.5 M glycine for 5 min. Cells were resuspended in 2 × cell pellet volumes of cold hypotonic buffer (10 mM Tris-HCl pH 7.5, 2 mM MgCl2, 3 mM CaCl2, 10% glycerol, 1 mM DTT, 1 × Proteinase inhibitor cocktail). Adding 2 × cell pellet volumes of cytomembrane lysis buffer (10 mM Tris-HCl pH 7.5, 2 mM MgCl2, 3 mM CaCl2, 10% glycerol, 1% NP-40, 1 mM DTT, 1 × Proteinase inhibitor cocktail) and incubated on ice for 10 min. The sample was then centrifuged at 600 × g for 5 minutes, and the supernatant was collected as the cytoplasmic fraction. The nuclei pellet was washed once with nuclear lysis buffer without NP-40, and resuspended and incubated with nuclear lysis buffer (20 mM HEPES, pH 7.5, 10 mM KCl, 350 mM NaCl, 1 mM EDTA, 0.2% NP-40, 10% glycerol, 1 mM DTT, 1 × Proteinase inhibitor cocktail) for 30 min on ice, and lysate was passed through 29 G needle 10 times. The supernatant was taken as nucleoplasm after being spun down. The chromatin pellet was treated by MNase buffer (50 mM Tris-HCl Ph 8.0, 5 mM MgCl2, 5 mM CaCl2, 1 × Proteinase inhibitor cocktail) with Micrococcal nuclease for 20 min. Following high-speed centrifugation, the supernatant was taken as chromatin fraction.
Protein expression and purification
The expression plasmids for HIS tagged GFP-UTP15IDR (amino acids 314-528) and mCherry-CTDC26 (C-terminal half of the CTD of POLR2A, heptad repeats 27–52) were transformed into E. coli BL21 (DE3). Cells were cultured in 6 L LB media at 37 °C until OD 600 reaches 0.6. Followed by adding 0.5 mM IPTG to express protein. Cells were cultured at 16 °C overnight. Cells were collected and resuspended in lysis buffer (50 mM Tris-HCl pH 7.5, 150 mM NaCl, 1 mM PMSF), then lysed under high pressure for 5 min. GFP-UTP15IDR and mCherry-CTDC26 proteins were purified using a Ni-NTA resin, and washed with wash buffer (50 mM Tris-HCl pH 7.5, 150 mM NaCl, 1 mM PMSF, 20 mM imidazole) and eluted with elution buffer (50 mM Tris-HCl pH 7.5, 150 mM NaCl, 1 mM PMSF, 250 mM imidazole). Finally, the purified proteins were concentrated in Millipore concentration tubes (> 30 kDa) in 50 mM Tris-HCl pH 7.5, 150 mM NaCl, and flash-frozen in liquid nitrogen and stored at −80 °C.
Liquid droplet formation
The indicated concentrations of proteins were added to a buffer containing 50 mM Tris-HCl pH 7.5, and 150 mM NaCl supplemented with 16% dextran, and the mixture was then transferred to a 384-well glass-bottomed plate (P384-1.5H-N) and placed at room temperature for the indicated time. Imaging analysis was performed using a confocal microscope (LSM710, Zeiss) immediately. The subsequent statistical analysis of the images was performed using ImageJ software.
Dual luciferase reporter assay
The cDNA of UTP15 or other RBPs was cloned and fused with GAL4 DNA-binding domain (GAL4-UTP15) into pcDNA3.1. A 5 × upstream activation sequence (UAS) was inserted into the upstream of the SV40 promoter of psiCHECK-2 vector. For a 6-well plate, 100 ng of psiCHECK-2 and 1 μg of pcDNA3.1-GAL4-UTP15 vectors were transfected into HEK293T cells by polyethylenimine (PEI). After 24 h of transfection, luciferase activity measurements were performed using the Dual-Luciferase Reporter Assay System (Vazyme) according to the manufacturer’s instructions. The Renilla luciferase activity was measured and normalized with firefly luciferase activity.
EU staining
Cells were treated with 1 mM EU for 30 minutes, then fixed for 30 minutes at room temperature with 4% paraformaldehyde (BOSTER). Cells were permeabilized with 0.5% Triton X-100 (in PBS) for 30 minutes at room temperature. Then, Apollo567 dye was conjugated through click reaction with Cell-Light EU Apollo567 RNA Imaging Kit (RiboBio) following the manufacturer’s instructions. Stained cells were subjected to immunofluorescence imaging on an inverted confocal microscope (LSM STED, ZEISS). The mean EU fluorescence intensity was calculated in the 3D projection procedure.
Immunofluorescence
Cells were cultured on glass coverslips and washed with PBS. Cells were then fixed with 4% paraformaldehyde for 15 min. After fixation, cells were washed three times with PBS and permeabilized with 0.1% Triton X-100 for 10 min. After washing with PBS, cells were blocked with 3% Bovine Serum Albumin (BSA) for 1 hour at room temperature. After blocking, cells were washed three times with PBS. Primary antibodies were applied, and cells were incubated overnight at 4 °C. The following antibodies were used: FLAG (1:200, Sigma M2), Pol II NTD (1:100, CST 14958S), NANOG (1:100, Abcam), HA (1:100, Abcam). Cells were washed with PBS and incubated with Alexa fluor 568 donkey anti-rabbit IgG (1:400, Invitrogen), Alexa fluor 488 donkey anti-mouse IgG (1:400, Invitrogen) for 1 hour in the dark. Then cells were washed three times with PBS and incubated with DAPI (1:1000, Invitrogen) for 5 minutes at room temperature for nuclear counterstaining. Coverslips were subsequently mounted onto slides containing anti-fading reagent. Images were captured using a confocal microscope.
Super-resolution imaging and data analysis
Super-resolution imaging was performed using Zeiss Elyra 7 with Lattice SIM2. Images were captured with a 100× oil immersion objective. Single plane images were preprocessed with consistent parameters in SIM2 before puncta quantification in ImageJ. During the analysis, nuclear regions were delineated based on the DAPI signal. RGB images were converted to 8-bit, processed with a Laplace of Gaussian (LoG) filter, and the threshold for fluorescence channels was set using the ‘Auto Threshold’ function in ImageJ for the image. The number of clusters per cell from independent images was quantified, and statistical comparisons were performed using a non-parametric test (Mann-Whitney U test) to assess significance.
Fluorescence recovery after photobleaching (FRAP) analysis
Mouse E14 cells transduced with EGFP-UTP15 IDR lentivirus were cultured on MEF cells. FRAP experiments were performed on a LEICA STELLARIS 8 confocal microscope equipped with a 63x oil immersion objective (Leica) in a live cell imaging chamber. For each experiment, circular regions of interest (ROIs) were selected, and a total of 10 ROIs were bleached using 50% laser power. The fluorescence recovery of each ROI was monitored over time. Bleaching was performed every 30 seconds for a total of 10 minutes. The resulting recovery curves were plotted as relative fluorescence intensity over time, where the intensity values were normalized to the pre-bleach baseline.
Embryonic microinjection
Zygotes were collected from superovulated female mice. Microinjection of siRNA into zygotes was carried out using the Eppendorf FemtoJet microinjector and Narishige micromanipulators. Approximately 2 pl of siRNA (300 nM) was injected into the cytoplasm of zygotes. siRNA for the microinjection was synthesized by the Guangzhou Ribobio company. Animal experiments followed the National Research Council’s Guide for the Care and Use of Laboratory Animals and were approved by the Guangzhou Institutes of Biomedicine and Health (GIBH) Institutional Animal Care and Use Committee. Mice were maintained under specific pathogen-free (SPF) conditions with a 12-hour light/dark cycle, at 22 ± 2 °C and 50–70% humidity, with ad libitum access to food and water.
Embryonic immunofluorescence
Immunofluorescence was performed using the following primary antibodies: rabbit anti-NANOG (Cosmo Bio, REC-RCAB0002PF), 1/100; goat anti-GATA6 (R&D, AF1700), 1/300; mouse anti-CDX2 (BioGenex, MU 392-UC), 1/200. Embryonic immunofluorescence imaging was performed on an inverted confocal microscope (LSM 900, ZEISS).
Electrophoretic mobility shift assay (EMSA)
EMSA was performed as described before (NAR 2020). In brief, RNA binding reactions were conducted in a 20 μL reaction system (10 mM Tris-HCl pH 7.4, 1 mM MgCl2, 1 mM DTT, 10 mM KCl, 5% glycerol, 1 U/μL RNase inhibitor, 50 nM Cy3-labeled RNA probe (UCGUGUCUCG CUUGACCAUG UUCCCAGAGU CGGUGGAUGU), and with indicated amounts of proteins). After incubation, samples were resolved on a 6.5% native polyacrylamide gel in 0.5 × TBE at 4 °C, 200 V in the dark. Signals were visualized with ChampChemiTM.
Nascent RNA-seq analysis
Raw reads were processed with fastp to remove adapter and low-quality sequences. UMI information was extracted using the parameter set --umi_loc=read1 --umi_len=6. The remaining reads were aligned to the mouse reference genome mm10 or the spike-in reference using Hisat2 under default parameters. UMI-tools were used to identify UMIs and remove PCR duplicates. Subsequently, featureCounts was applied to assign aligned reads to genomic features. Differential expression analysis was conducted using DESeq2 with a threshold of fold change > 1.5 and FDR < 0.05. Data normalization was performed by the estimateSizeFactors function with the controlGenes option. The scale factor derived from spike-in RNA was used to convert BAM files to BigWig format.
To quantify the signals of the 685 UTP15 target genes, we implemented distinct methods according to gene length. For genes longer than 500 bp, signals within ± 200 bp of the transcription start sites (TSSs) on the sense strand were quantified using multiBigwigSummary from deepTools, focusing on the promoter regions. For the gene bodies, signals from 300 bp downstream of the TSS to the transcription end sites (TES) were analyzed using the same tool. For genes shorter than 500 bp, the promoter region was defined as ± 50 bp around the TSS, and the gene body extended from 50 bp downstream of the TSS to the TES.
Total RNA-seq analysis
After quality filtering and trimming with fastp, the resulting reads were aligned to the mm10 reference genome using Hisat2. PCR duplicates were removed by collapsing the sequences with Picard. The count matrix generated by featureCounts served as input for DESeq2 to perform differential expression analysis. (fold change > 1.5, FDR < 0.05). Reads per kilobase per million mapped reads (RPKM) were used to determine relative gene expression levels. For total RNA-seq after UTP15 degradation, reads were also aligned to the ERCC spike-in reference, and a spike-in count matrix was generated. This matrix was subsequently passed to the estimateSizeFactors function of DESeq2 to calculate scale factors.
ChIP-seq analysis
To analyze ChIP-seq data and identify target genes, adapter sequences were removed, and quality filtering was conducted using fastp. Subsequently, the quality of the sequencing data was assessed using FastQC. Paired-end sequencing reads were mapped to either the mouse reference genome (mm10) or the D. melanogaster genome (dm6) using Bowtie 2 with the following parameters: --local --very-sensitive-local --phred33 --no-unal --no-mixed --no-discordant -I 10 -X 700 -S, along with the additional --no-overlap option for mapping spike-in reads. Uniquely mappable reads were filtered from the aligned reads using samtools and Picard. ChIP-seq peaks were identified using MACS2 with a significance threshold of q-value < 0.05. Additionally, to ensure the reproducibility of the results, the Irreproducible Discovery Rate (IDR) method was applied to the replicates with a threshold set at 0.05 to select reliable peaks for downstream analysis. Peaks were then annotated using the HOMER tool to identify potential target genes, and visualization of annotated peaks was performed using ChIPseeker.
To generate metagene plots, BAM files were converted to BigWig format using bamCoverage with the scaleFactor option for normalization based on spike-in reads and the blackListFileName option to filter out mouse ENCODE blacklisted genomic regions. ChIP-seq read density was then calculated with computeMatrix. Heatmaps and meta-profiles were generated using the plotHeatmap function from DeepTools, and motif discovery was performed with the FIMO tool.
CLIP-seq analysis
The raw data were processed to obtain unique CLIP tags using the CTK pipeline, as described previously81. Briefly, the raw data were subjected to a filtration process to eliminate adapters, low-quality reads, and PCR duplicates. Subsequently, reads longer than 15 nt were aligned to the mouse reference genome (mm10) using BWA. CTK scripts were then utilized to parse alignments and call peaks using a valley-seeking algorithm. The annotation of peaks was generated by HOMER, and visualization of annotated peaks was performed using ChIPseeker. Irreproducibility Discovery Rate (IDR < 0.05) was used to identify the reproducible peaks between the replicates. Average profiles were generated using ngs.plot.r.
Processing and analysis of mass spectrometry (MS) datasets
All MS raw data were searched using MaxQuant against the reviewed UniProt mouse proteome database. The output generated by MaxQuant was then analyzed using the DEP package in RStudio. First, the LFQ intensity values were subjected to filtering, normalization, and imputation of missing values. Next, differential enrichment analysis was conducted based on linear models combined with empirical Bayes statistics. Proteins that exhibited a fold change > 1.5 and a p value < 0.05 were considered as enriched in the Co-IP samples. For the analysis of RNA-binding proteins from RICK, enriched proteins were screened out with the following conditions (EU versus non-EU, fold change > 4, p value < 0.01). Subsequently, the differences in enriched proteins within the EU groups at different time points versus 0 h were analyzed. DEP computes p value based on a two-sided moderated t test. The identification of RNA-binding domains was performed using pfam_scan.pl, and classification was conducted following the approach described by Castello et al.23. The values of isoelectric point (pI) were calculated with the ExPASy Compute pI/MW tool. The protein-protein interactions (PPI) were conducted using STRING.
Functional Enrichment Analysis
Gene set enrichment analysis (GSEA) was performed using the GSEA 4.2.3 software. The enrichment p values were obtained by permutation-based tests (gene-set permutations, as implemented in GSEA) Kyoto Encyclopedia of Genes and Genomes (KEGG) and Gene-Ontology (GO) Enrichment analysis was performed using the DAVID Functional Annotation Bioinformatics Microarray Analysis tool. DAVID computes p values based on a modified Fisher’s exact test (EASE score).
Transposable elements analysis (TEs)
Transposable elements were quantified at the locus level using SQuIRE software82. The squire Fetch command downloaded TE annotation from RepeatMasker. squire Clean filtered the RepeatMasker file for repeats of interest and collapsed overlapping repeats. squire Map aligned reads to mm10. squire Count quantified reads aligning to TEs and genes. Differentially expressed TE loci were identified with DESeq2 applying thresholds of fold change >1.5 and FDR < 0.05.
Permutation Test
We split all genes into 20 bins based on their expression levels at NANOG-AID 0 h. Then, we created a random gene set by drawing N genes from each bin, excluding UTP15-target genes, where the N is equal to the number of UTP15-target genes in the given bin. The above process was repeated 5,000 times to build a distribution (expression-matched random sets) with matched expression level of the UTP15-target genes. Subsequently, a permutation test was applied to determine whether the observed expression change of UTP15-target genes (Δ expression = mean at 2 h − mean at 0 h) significantly deviates from the distribution of changes in these expression-matched random sets.
Data visualization
Bedgraphs and bigwig format were obtained for visualization in Integrative Genomics Viewer (IGV). Data were analyzed in RStudio and visualized using the ggplot2 library.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The sequencing data generated in this study have been deposited in the GEO database under accession codes GSE276877, GSE276878 and GSE276879. The mass spectrometry data generated in this study have been deposited in the ProteomeXchange Consortium via the PRIDE partner repository under accession code PXD043911. The main data supporting the findings of this study are available within the article and in Supplementary information. Source data are provided with this paper.
References
De Los Angeles, A. et al. Hallmarks of pluripotency. Nature 525, 469–478 (2015).
Ng, H. H. & Surani, M. A. The transcriptional and signalling networks of pluripotency. Nat. Cell Biol. 13, 490–496 (2011).
Weinberger, L., Ayyash, M., Novershtern, N. & Hanna, J. H. Dynamic stem cell states: naive to primed pluripotency in rodents and humans. Nat. Rev. Mol. Cell Biol. 17, 155–169 (2016).
Lessard, J. A. & Crabtree, G. R. Chromatin regulatory mechanisms in pluripotency. Annu Rev. Cell Dev. Biol. 26, 503–532 (2010).
Boyer, L. A. et al. Core transcriptional regulatory circuitry in human embryonic stem cells. Cell 122, 947–956 (2005).
You, K. T., Park, J. & Kim, V. N. Role of the small subunit processome in the maintenance of pluripotent stem cells. Genes Dev. 29, 2004–2009 (2015).
Corsini, N. S. et al. Coordinated Control of mRNA and rRNA Processing Controls Embryonic Stem Cell Pluripotency and Differentiation. Cell Stem Cell 22, 543–558.e512 (2018).
Vilchez, D. et al. Increased proteasome activity in human embryonic stem cells is regulated by PSMD11. Nature 489, 304–308 (2012).
Buckley, S. M. et al. Regulation of pluripotency and cellular reprogramming by the ubiquitin-proteasome system. Cell Stem Cell 11, 783–798 (2012).
Whyte, W. A. et al. Master transcription factors and mediator establish super-enhancers at key cell identity genes. Cell 153, 307–319 (2013).
Tsankov, A. M. et al. Transcription factor binding dynamics during human ES cell differentiation. Nature 518, 344–349 (2015).
Kulessa, H., Frampton, J. & Graf, T. GATA-1 reprograms avian myelomonocytic cell lines into eosinophils, thromboblasts, and erythroblasts. Genes Dev. 9, 1250–1262 (1995).
Davis, R. L., Weintraub, H. & Lassar, A. B. Expression of a single transfected cDNA converts fibroblasts to myoblasts. Cell 51, 987–1000 (1987).
Heurtier, V. et al. The molecular logic of Nanog-induced self-renewal in mouse embryonic stem cells. Nat. Commun. 10, 1109 (2019).
Muhar, M. et al. SLAM-seq defines direct gene-regulatory functions of the BRD4-MYC axis. Science 360, 800–805 (2018).
Ji, X. et al. SR proteins collaborate with 7SK and promoter-associated nascent RNA to release paused polymerase. Cell 153, 855–868 (2013).
Fu, X. D. & Ares, M. Jr. Context-dependent control of alternative splicing by RNA-binding proteins. Nat. Rev. Genet 15, 689–701 (2014).
Zhao, B. S., Roundtree, I. A. & He, C. Post-transcriptional gene regulation by mRNA modifications. Nat. Rev. Mol. Cell Biol. 18, 31–42 (2017).
Wang, L. et al. The THO complex regulates pluripotency gene mRNA export and controls embryonic stem cell self-renewal and somatic cell reprogramming. Cell Stem Cell 13, 676–690 (2013).
Gerstberger, S., Hafner, M. & Tuschl, T. A census of human RNA-binding proteins. Nat. Rev. Genet. 15, 829–845 (2014).
Hentze, M. W., Castello, A., Schwarzl, T. & Preiss, T. A brave new world of RNA-binding proteins. Nat. Rev. Mol. Cell Biol. 19, 327–341 (2018).
Castello, A. et al. Insights into RNA biology from an atlas of mammalian mRNA-binding proteins. Cell 149, 1393–1406 (2012).
Castello, A. et al. Comprehensive Identification of RNA-Binding Domains in Human Cells. Mol. Cell 63, 696–710 (2016).
Shao, W. et al. Phase separation of RNA-binding protein promotes polymerase binding and transcription. Nat. Chem. Biol. 18, 70–80 (2022).
Mittag, T. & Pappu, R. V. A conceptual framework for understanding phase separation and addressing open questions and challenges. Mol. Cell 82, 2201–2214 (2022).
Farag, M. et al. Condensates formed by prion-like low-complexity domains have small-world network structures and interfaces defined by expanded conformations. Nat. Commun. 13, 7722 (2022).
Farag, M., Borcherds, W. M., Bremer, A., Mittag, T. & Pappu, R. V. Phase separation of protein mixtures is driven by the interplay of homotypic and heterotypic interactions. Nat. Commun. 14, 5527 (2023).
Lin, Y., Protter, D. S., Rosen, M. K. & Parker, R. Formation and Maturation of Phase-Separated Liquid Droplets by RNA-Binding Proteins. Mol. Cell 60, 208–219 (2015).
Maharana, S. et al. RNA buffers the phase separation behavior of prion-like RNA binding proteins. Science 360, 918–921 (2018).
Shin, Y. & Brangwynne, C. P. Liquid phase condensation in cell physiology and disease. Science 357, https://doi.org/10.1126/science.aaf4382 (2017).
Hnisz, D., Shrinivas, K., Young, R. A., Chakraborty, A. K. & Sharp, P. A. A Phase Separation Model for Transcriptional Control. Cell 169, 13–23 (2017).
Elguindy, M. M. & Mendell, J. T. NORAD-induced Pumilio phase separation is required for genome stability. Nature 595, 303–308 (2021).
Bao, X. et al. Capturing the interactome of newly transcribed RNA. Nat. Methods 15, 213–220 (2018).
Huang, R., Han, M., Meng, L. & Chen, X. Transcriptome-wide discovery of coding and noncoding RNA-binding proteins. Proc. Natl Acad. Sci. USA 115, E3879–e3887 (2018).
Li, J. et al. A One-step strategy to target essential factors with auxin-inducible degron system in mouse embryonic stem cells. Front Cell Dev. Biol. 10, 964119 (2022).
Chambers, I. et al. Functional expression cloning of Nanog, a pluripotency sustaining factor in embryonic stem cells. Cell 113, 643–655 (2003).
Mitsui, K. et al. The homeoprotein Nanog is required for maintenance of pluripotency in mouse epiblast and ES cells. Cell 113, 631–642 (2003).
Morrisey, E. E., Ip, H. S., Lu, M. M. & Parmacek, M. S. GATA-6: a zinc finger transcription factor that is expressed in multiple cell lineages derived from lateral mesoderm. Dev. Biol. 177, 309–322 (1996).
Arceci, R. J., King, A. A., Simon, M. C., Orkin, S. H. & Wilson, D. B. Mouse GATA-4: a retinoic acid-inducible GATA-binding transcription factor expressed in endodermally derived tissues and heart. Mol. Cell Biol. 13, 2235–2246 (1993).
Takahashi, K. & Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126, 663–676 (2006).
Festuccia, N. et al. Esrrb is a direct Nanog target gene that can substitute for Nanog function in pluripotent cells. Cell Stem Cell 11, 477–490 (2012).
Core, L. J., Waterfall, J. J. & Lis, J. T. Nascent RNA sequencing reveals widespread pausing and divergent initiation at human promoters. Science 322, 1845–1848 (2008).
Rabani, M. et al. High-resolution sequencing and modeling identifies distinct dynamic RNA regulatory strategies. Cell 159, 1698–1710 (2014).
Sun, Z. et al. Chromatin regulation of transcriptional enhancers and cell fate by the Sotos syndrome gene NSD1. Mol. Cell 83, 2398–2416.e2312 (2023).
Bourillot, P. Y. et al. Novel STAT3 target genes exert distinct roles in the inhibition of mesoderm and endoderm differentiation in cooperation with Nanog. Stem Cells 27, 1760–1771 (2009).
Niu, C. et al. BACH1 recruits NANOG and histone H3 lysine 4 methyltransferase MLL/SET1 complexes to regulate enhancer-promoter activity and maintains pluripotency. Nucleic Acids Res. 49, 1972–1986 (2021).
Avsec, Ž et al. Base-resolution models of transcription-factor binding reveal soft motif syntax. Nat. Genet. 53, 354–366 (2021).
An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57-74 (2012).
Capo-Chichi, C. D. et al. Perception of differentiation cues by GATA factors in primitive endoderm lineage determination of mouse embryonic stem cells. Dev. Biol. 286, 574–586 (2005).
Plusa, B., Piliszek, A., Frankenberg, S., Artus, J. & Hadjantonakis, A. K. Distinct sequential cell behaviours direct primitive endoderm formation in the mouse blastocyst. Development 135, 3081–3091 (2008).
Singh, A. M., Hamazaki, T., Hankowski, K. E. & Terada, N. A heterogeneous expression pattern for Nanog in embryonic stem cells. Stem Cells 25, 2534–2542 (2007).
Zhang, J. et al. Dax1 and Nanog act in parallel to stabilize mouse embryonic stem cells and induced pluripotency. Nat. Commun. 5, 5042 (2014).
Chazaud, C., Yamanaka, Y., Pawson, T. & Rossant, J. Early lineage segregation between epiblast and primitive endoderm in mouse blastocysts through the Grb2-MAPK pathway. Dev. Cell 10, 615–624 (2006).
Caudron-Herger, M., Jansen, R. E., Wassmer, E. & Diederichs, S. RBP2GO: a comprehensive pan-species database on RNA-binding proteins, their interactions and functions. Nucleic Acids Res 49, D425–d436 (2021).
Kwon, S. C. et al. The RNA-binding protein repertoire of embryonic stem cells. Nat. Struct. Mol. Biol. 20, 1122–1130 (2013).
Kawai, J. et al. Functional annotation of a full-length mouse cDNA collection. Nature 409, 685–690 (2001).
Dragon, F. et al. A large nucleolar U3 ribonucleoprotein required for 18S ribosomal RNA biogenesis. Nature 417, 967–970 (2002).
Prieto, J. L. & McStay, B. Recruitment of factors linking transcription and processing of pre-rRNA to NOR chromatin is UBF-dependent and occurs independent of transcription in human cells. Genes Dev. 21, 2041–2054 (2007).
Hutchins, A. P. et al. Models of global gene expression define major domains of cell type and tissue identity. Nucleic Acids Res 45, 2354–2367 (2017).
Bi, X. et al. RNA Targets Ribogenesis Factor WDR43 to Chromatin for Transcription and Pluripotency Control. Mol. Cell 75, 102–116.e109 (2019).
Zeng, Y. et al. Lin28A Binds Active Promoters and Recruits Tet1 to Regulate Gene Expression. Mol. Cell 61, 153–160 (2016).
Kwon, I. et al. Phosphorylation-regulated binding of RNA polymerase II to fibrous polymers of low-complexity domains. Cell 155, 1049–1060 (2013).
Scott, M. S., Troshin, P. V. & Barton, G. J. NoD: a Nucleolar localization sequence detector for eukaryotic and viral proteins. BMC Bioinforma. 12, 317 (2011).
Scott, M. S., Boisvert, F. M., McDowall, M. D., Lamond, A. I. & Barton, G. J. Characterization and prediction of protein nucleolar localization sequences. Nucleic Acids Res 38, 7388–7399 (2010).
Jiang, Y. et al. Cross-regulome profiling of RNA polymerases highlights the regulatory role of polymerase III on mRNA transcription by maintaining local chromatin architecture. Genome Biol. 23, 246 (2022).
Linding, R. et al. Protein disorder prediction: implications for structural proteomics. Structure 11, 1453–1459 (2003).
Boehning, M. et al. RNA polymerase II clustering through carboxy-terminal domain phase separation. Nat. Struct. Mol. Biol. 25, 833–840 (2018).
Lu, H. et al. Phase-separation mechanism for C-terminal hyperphosphorylation of RNA polymerase II. Nature 558, 318–323 (2018).
Guo, Y. E. et al. Pol II phosphorylation regulates a switch between transcriptional and splicing condensates. Nature 572, 543–548 (2019).
Bates, L. E., Alves, M. R. P. & Silva, J. C. R. Auxin-degron system identifies immediate mechanisms of OCT4. Stem Cell Rep. 16, 1818–1831 (2021).
Rabani, M. et al. Metabolic labeling of RNA uncovers principles of RNA production and degradation dynamics in mammalian cells. Nat. Biotechnol. 29, 436–442 (2011).
Crump, N. T. et al. BET inhibition disrupts transcription but retains enhancer-promoter contact. Nat. Commun. 12, 223 (2021).
Oksuz, O. et al. Transcription factors interact with RNA to regulate genes. Mol. Cell 83, 2449–2463.e2413 (2023).
Calo, E. et al. RNA helicase DDX21 coordinates transcription and ribosomal RNA processing. Nature 518, 249–253 (2015).
Gupta, S. & Santoro, R. Regulation and Roles of the Nucleolus in Embryonic Stem Cells: From Ribosome Biogenesis to Genome Organization. Stem Cell Rep. 15, 1206–1219 (2020).
Diesch, J. et al. Changes in long-range rDNA-genomic interactions associate with altered RNA polymerase II gene programs during malignant transformation. Commun. Biol. 2, 39 (2019).
Percharde, M. et al. A LINE1-Nucleolin Partnership Regulates Early Development and ESC Identity. Cell 174, 391–405.e319 (2018).
Shalem, O. et al. Genome-scale CRISPR-Cas9 knockout screening in human cells. Science 343, 84–87 (2014).
Perez-Riverol, Y. et al. The PRIDE database resources in 2022: a hub for mass spectrometry-based proteomics evidences. Nucleic Acids Res. 50, D543–d552 (2022).
Xiao, R. et al. Pervasive Chromatin-RNA Binding Protein Interactions Enable RNA-Based Regulation of Transcription. Cell 178, 107–121.e118 (2019).
Shah, A., Qian, Y., Weyn-Vanhentenryck, S. M. & Zhang, C. CLIP Tool Kit (CTK): a flexible and robust pipeline to analyze CLIP sequencing data. Bioinformatics 33, 566–567 (2017).
Yang, W. R., Ardeljan, D., Pacyna, C. N., Payer, L. M. & Burns, K. H. SQuIRE reveals locus-specific regulation of interspersed repeat expression. Nucleic Acids Res. 47, e27 (2019).
He, J. et al. Transposable elements are regulated by context-specific patterns of chromatin marks in mouse embryonic stem cells. Nat. Commun. 10, 34 (2019).
Acknowledgements
We thank all other members from the Bao and Wu laboratories for their support and active discussion. We thank Dr Rui Xiao (Wuhan University), Dr Andrew P Hutchins (Southern University of Science and Technology of China), Dr Hui Zhang (Guangzhou National Laboratory), Dr Hui Wang (Peking University), Dr Jiangping He (Guangzhou Laboratory), Dr Jiajian Zhou (Southern Medical University), Ying Li (Proteomics and Metabolomics Core Facility, Guangzhou National Laboratory) and Ruhai Chen (Guangzhou Institutes of Biomedicine and Health, Chinese Academy of Sciences) for inputs during this study. We are grateful to Dr Xiaohua Shen (Tsinghua University) for providing the plasmids. This work was supported by National Key Research and Development Program of China (2021YFA1100401, 2024YFA1108201 and 2022YFE0210600 for X.C.B.); National Natural Science Foundation of China (32170745 for X.C.B.); Major project of Guangzhou National Laboratory (GZNL2023A03005 for X.C.B.); Science and Technology Projects in Guangzhou, China (2023A04J0100 for C.M.Z.); Guangdong Basic and Applied Basic Research Foundation (2022A1515110502 for C.M.Z.); Science and Technology Projects in Guangzhou, China (2023A04J0104 for Z.B.M.); Basic Research Project of Guangzhou Institutes of Biomedicine and Health, Chinese Academy of Sciences (No. GIBHBRP24-03 for X.C.B.); National Natural Science Foundation of China (32270625 for H.Z.); Guangdong Province Grant for Belt and Road Joint Laboratory (2022B1212050004 for X.C.B.); Partially supported by Science and Technology Planning Project of Guangdong Province, China (2023B1212060050 and 2023B1212120009 for X.C.B.).
Author information
Authors and Affiliations
Contributions
X.C.B. and M.Q.D. conceived the idea. X.C.B., M.Q.D., and D.M.H. designed the experiments. X.C.B. wrote the manuscript with help from M.Q.D., D.M.H. and X.W.W. M.Q.D., D.M.H., X.W.W., Y.T.Y., C.M.Z., L.S.W., Q.Y.Z., Y.L.S., Y.L.Z. performed the experiments. D.M.H., Z.B.M. performed bioinformatics analysis. Y.L.H. and Z.W.N. conducted embryo injections. Q.F.W., L.Y.P., B.D., D.J.L., D.L., Z.X., J.X.X., D.J.Q., H.Z., Y.Z., P.T., J.K.W., C.P., R.J., G.M.W. and W.S.Z. contributed technical assistance/suggestions. G.M.W. and W.S.Z. contributed to infrastructural support. X.C.B. supervised the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Deng, M., He, D., Wang, X. et al. Monitoring rapid degradation of NANOG reveals UTP15 maintains pluripotency by regulating nascent transcripts. Nat Commun 17, 486 (2026). https://doi.org/10.1038/s41467-025-67018-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-67018-7