Abstract
Accumulating evidence suggests that a compromised intestinal epithelial barrier (IEB) contributes to the progression of metabolic dysfunction-associated steatotic liver disease (MASLD); however, the exact mechanisms remain unclear. Here we reveal that intestinal mucin 1 (MUC1) levels and glycosylation are decreased in both humans and male mice with MASLD. Enterocyte-specific Muc1 knockout aggravates high-fat diet (HFD)-induced IEB impairment and MASLD progression in male mice. Mechanistically, HFD feeding reduces the glycosylation of intestinal epithelial MUC1, triggering its clathrin-mediated endocytosis and NEDD4-mediated lysosomal degradation, which subsequently induces β-Catenin degradation and ultimately impaires the IEB. Notably, enterocyte-specific overexpression of cytoplasmic-tail-deleted MUC1 protects against IEB impairment and mitigates MASLD progression. These findings indicate that reduced intestinal epithelial MUC1 levels facilitate the progression of MASLD. Preserving the glycosylation and levels of intestinal MUC1 to maintain IEB integrity is a potential therapeutic strategy to explore for MASLD.
Similar content being viewed by others
Introduction
Metabolic dysfunction-associated steatotic liver disease (MASLD), previously known as non-alcoholic fatty liver disease, is a global public health challenge that affects one-quarter of the population and poses a significant economic burden on societies worldwide1. MASLD encompasses a spectrum of liver disorders, ranging from simple steatosis to the more severe metabolic dysfunction-related steatohepatitis (MASH). This progression may ultimately result in cirrhosis, liver failure, and hepatocellular carcinoma2. Despite many studies aimed at elucidating the pathogenic mechanisms of MASLD, the current treatment options remain limited. Therefore, it is crucial to investigate the underlying mechanisms of MASLD progression to uncover novel molecular targets for its therapy.
Emerging evidence underscores the pivotal role of intestinal epithelial barrier (IEB) impairment in the progression of MASLD3,4,5. The IEB is a complex structure composed of multiple elements that work together to maintain intestinal integrity and function. The mucus layer serves as the initial defense mechanism, being particularly thicker in the ileum, cecum, and colon, which are the primary habitats for microbes6. The mucus layer physically segregates the microbiota from the underlying physical barrier, which is composed of intestinal epithelia and fortified by cell junctions7. However, disruption of the IEB allows intestinal bacteria and bacterial products to cross the IEB and reach the liver, thereby provoking liver injury and promoting the development of hepatic steatosis8. Patients with MASLD exhibit increased intestinal permeability, which correlates with the severity of hepatic steatosis9. Likewise, mice lacking Jam-A, a crucial component of cell junctions that regulates the paracellular pathway of solutes and prevents the transepithelial translocation of molecules such as lipopolysaccharides (LPS)7, exhibit increased susceptibility to the development of MASLD10. Recent studies have uncovered several novel targets within the IEB that are implicated in MASLD development11,12,13, indicating that genes related to IEB integrity and function may represent promising targets for MASLD treatment.
Mucins comprise a family of highly glycosylated glycoproteins, categorized structurally into secreted forms (e.g., mucin 2, mucin 5AC, mucin 6) and membrane-bound forms (e.g., mucin 1 (MUC1), mucin 3, mucin 4). The mucin family is abundantly expressed in the intestine, where mucin 2, secreted by goblet cells, constitutes the primary component of the intestinal mucus layer, forming a gel-like network. MUC1, expressed on the apical surface of intestinal epithelial cells, participates in cell signaling and intestinal epithelial barrier function maintenance14. Recent studies have highlighted the protective role of MUC2 in MASLD progression15. MUC1, a heterodimeric transmembrane glycoprotein consisting of the N-terminal (MUC1-N) and C-terminal (MUC1-C) subunits, is abundantly present on the surface of intestinal epithelial cells16. MUC1-N is characterized by the presence of sugar chains that are covalently linked to the peptide backbone through O- or N-glycosidic bonds at asparagine or serine/threonine residues. The highly glycosylated MUC1-N significantly contributes to the formation of intestinal epithelial glycocalyx, which connects the intestinal epithelia to the mucus layer17. MUC1-C is situated intracellularly and participates in numerous protein-protein interactions and signaling pathways, regulating various biological processes, including cell adhesion18. Recent studies have demonstrated that intestinal MUC1 plays a crucial physiological role in protecting mucosal cells from infection19. MUC1 deficiency aggravates colitis20, whereas increased MUC1 prevents inflammasome activation in IEB16. Furthermore, the abnormal expression of MUC1 plays a role in the initiation, progression, and metastasis of colonic cancer21. However, the role of intestinal MUC1 in maintaining the integrity of the IEB and contributing the MASLD progression remains unclear.
In this study, we observed that both the levels and glycosylation of MUC1 in colonic mucosal biopsy samples from MASLD individuals were significantly reduced compared to those in healthy controls. Additionally, we employed mouse models with enterocyte-specific Muc1 knockout or overexpression combined with an analysis of glycocalyx primarily formed by MUC1 in intestinal epithelia using transmission electron microscopy (TEM) to demonstrate that the loss of intestinal epithelial MUC1 impairs the IEB and exacerbates MASLD progression. We further identified that the deglycosylation of intestinal epithelial MUC1 during high-fat diet (HFD) feeding triggers its clathrin-mediated endocytosis and NEDD4-mediated lysosomal degradation, which subsequently leads to β-catenin degradation and ultimately impairs the IEB. Notably, we found that enterocyte-specific cytoplasmic-tail-deleted MUC1 (MUC1ΔCT) overexpression preserved intestinal MUC1 levels and alleviated MASLD progression. These findings establish intestinal epithelial MUC1 as a key regulator of the IEB and suggest it as a potential target for MASLD treatment.
Results
Intestinal MUC1 levels are decreased in patients and mice with MASLD
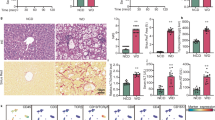
To investigate the potential association between intestinal MUC1 and MASLD, we first determined MUC1 expression in colonic mucosal biopsy samples from 30 individuals diagnosed with MASLD and 30 age- and sex-matched healthy controls. A reduction in the protein levels of MUC1 was observed in MASLD individuals compared to healthy controls (Fig. 1a, b). Furthermore, we found inverse correlations between intestinal MUC1 levels and BMI, serum triglycerides (TG), as well as alanine aminotransferase (ALT) (Fig. 1c). Likewise, intestinal MUC1 levels were also significantly downregulated in the HFD-induced MASLD mouse model, compared to those in the corresponding control mice (Fig. 1d, e). Interestingly, the mRNA levels of Muc1 were not obviously changed in MASLD individuals and mice (Supplementary Fig. 1a, b). Taken together, these data suggest that decreased intestinal MUC1 may be involved in the progression of MASLD.
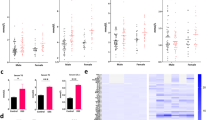
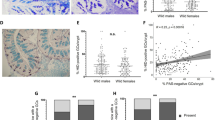
a Relative protein expression of mucin 1 (MUC1) in colon mucosa biopsy samples of the control group and MASLD group (n = 30 per group). b Representative immunofluorescence staining of MUC1 in colon mucosa biopsy samples of control and patients with MASLD. Scale bar, 50 μm. c Correlations between intestinal MUC1 protein expression and BMI, serum TG, TC, ALT, and AST levels in colon mucosa biopsy samples (n = 60). The correlation analysis was performed using Pearson’s product-moment correlation (two-sided without adjustment). d Relative protein expression of MUC1 in colon tissues of mice fed with ND and HFD for 16 weeks (n = 6 per group). e Representative immunofluorescence staining of MUC1 in colon tissues of mice fed with ND and HFD for 16 weeks. Scale bar, 50 μm. f Structural schematic diagram of the membrane MUC1 protein. The diagram of the membrane and MUC1 was created in BioRender. Li, Z. (2025) https://BioRender.com/b1gercd and https://BioRender.com/7jqtxz6. g, h Representative transmission electron microscopy images of the glycocalyx in human colon (g) and mouse colon (h). Scale bar, 200 nm (left), 100 nm (right). i Schematic diagram of peanut agglutinin (PNA)-specific recognition of O-glycan patterns. The diagram of peanut agglutinin was created in BioRender. Li, Z. (2025) https://BioRender.com/txue94i. Ser (Serine), Thr (Threonine). j The expression levels of O-glycans (PNA) on colonic MUC1 were assessed in both the control group and the patient with MASLD group under conditions of equivalent MUC1 levels. k The expression levels of O-glycans (PNA) on colonic MUC1 were assessed in mice fed with ND and HFD under conditions of equivalent MUC1 levels. Data are presented as mean ± SEM. P-values were calculated by a two-tailed unpaired t-test. Western blotting data are representative of three independent experiments. Source data are provided as a Source data file.
Intestinal MUC1 is deglycosylated in humans and mice with MASLD
Given that the intestinal epithelial glycocalyx, mainly composed of the O-glycans of MUC1 (Fig. 1f), connects the intestinal epithelia to the mucus layer, we first analyzed it in colonic mucosal biopsy samples from 30 individuals with MASLD and 30 healthy controls using transmission electron microscopy (TEM). The thickness of the intestinal epithelial glycocalyx layer was decreased in individuals with MASLD (Fig. 1g, Supplementary Fig. 1c), and negatively correlated with BMI, serum TG and ALT (Supplementary Fig. 1e). Consistently, a significant reduction in the thickness of the intestinal epithelial glycocalyx layer of intestinal epithelium was also observed in MASLD mouse model (Fig. 1h, Supplementary Fig. 1d). Further immunoprecipitation of intestinal MUC1 protein showed reduced levels of O-glycosylation as detected by Peanut agglutinin (PNA) in intestinal MUC1 derived from individuals and mice with MASLD (Fig. 1i–k, Supplementary Fig. 1f, g). Taken together, these data suggest that intestinal MUC1 undergoes deglycosylation during MASLD progression (Supplementary Fig. 1h).
Deglycosylation of MUC1 induces its clathrin-dependent endocytosis
Subsequently, we explored why deglycosylation of intestinal MUC1 is associated with reduced MUC1 protein levels. Previous research suggests that glycosylation of transmembrane proteins reduces their endocytosis-mediated removal from the cell membrane22, leading us to hypothesize that deglycosylation might contribute to reduced MUC1 levels. To investigate this, we treated the human enterocyte cell line HT-29 with O-glycosidase and observed a reduction in MUC1 protein levels without obvious effect on Muc1 mRNA levels (Fig. 2a, Supplementary Fig. 2a–c). A recent study reported that the glycosylation of MUC1 is determined by its variable numbers of tandem repeats (VNTRs) of a specific sequence18,22. Thus, we genetically manipulated the glycosylation level of MUC1 by introducing Muc1 vectors carrying varying VNTRs into HT-29 cells and found that MUC1 proteins with five tandem repeats (5TRs) exhibited reduced protein levels compared to MUC1 proteins with high glycosylation, which contained ten tandem repeats (10TRs) (Fig. 2b). Collectively, these data suggest that deglycosylation of MUC1 may contribute to the reduction in MUC1 protein levels.
a Western blot analysis showing the stability of MUC1 after the administration of 50 nM O-glycosidase in complete media for 30 mins follow by cycloheximide (CHX, translation inhibitor) for 24 h in HT-29 cells. b Western blot analysis was performed to detect specified proteins in HT-29 cells that were transfected with deglycosylated MUC1 vectors, specifically MUC1-10TRs or MUC1-5TRs. c Western blots for the indicated proteins from HT-29 cells treated with O-glycosidase for 30 mins follow by 50 μM lysosome inhibitor CQ for 24 h or 10 μM proteasome inhibitor MG132 for 6 h, or DMSO as a control. d Western blots for the indicated proteins from HT-29 cells treated with DMSO, Golgi inhibitor BFA (8 μM), clathrin-dependent endocytosis inhibitor CPZ (10 μM), cavolin-dependent endocytosis inhibitor MβCD (5 mM), or lysosome inhibitor HCQ (50 μM) for 24 h. e Western blots for the indicated proteins in MUC1 of colonic HT-29 cells treated with treated O-glycosidase for 30 mins. f Western blots were used to detect MUC1 protein in co-immunoprecipitation with MUC1 protein or NEDD4 protein in the HT-29 cell line. g Western blots for the indicated proteins from HT-29 cells transfected shNEDD4-#1 and shNEDD4-#2 treated with O-Glycosidase for 30 mins. h A schematic of NEDD4 protein which included C2, WW1/2/3/4 and HECT domain. C2 (Calcium - dependent lipid - binding domain), WW1 (WW domains 1), WW2 (WW domains 2), WW3 (WW domains 3), WW4 (WW domains 4), HECT (Homologous to the E6 - AP Carboxyl Terminus domain), C1046/1197/1286 A (Cysteine residues at positions 1046, 1197, and 1286 were individually mutated to alanine). i Western blots for the indicated proteins from HT-29 cells transfected together with NEDD4WT or NEDD4C1046A, NEDD4C1197A, NEDD4C1286A. j HEK293T cells transfected with the indicated plasmids were treated with CQ (100 μM) for 24 h, and submitted to in vivo ubiquitination assay. k HT-29 cells co-transfected with Flag-MUC1, His-NEDD4(WT or C1046A), HA-Ub (Ub WT or K48, K63) were treated with CQ (100 μM) for 24 h, and submitted to in vivo ubiquitination assay. l Western blots for the indicated proteins from HT-29 cells transfected with MUC1-WT or MUC1-K1231R mutants were treated with glycosidase for 30 mins. Western blotting data are representative of three independent experiments. Source data are provided as a Source data file.
The observation that MUC1 protein levels but not mRNA levels decreased during MASLD raises an intriguing possibility that reduced intestinal MUC1 results from its degradation. Protein degradation typically occurs through either the lysosomal or proteasomal pathways23. To determine the degradation pathway of MUC1, we treated cells with lysosomal inhibitor chloroquine (CQ) and proteasomal inhibitor MG132. The results indicated that MUC1 degradation was significantly inhibited by CQ rather than MG132, suggesting that MUC1 degradation occurs through the lysosomal pathway (Fig. 2c). Furthermore, we employed inhibitors to identify the subcellular location of MUC1 degradation. Chlorpromazine (CZ), an inhibitor of clathrin-dependent endocytosis, was the only compound that significantly blocked MUC1 degradation, indicating the crucial role of clathrin-dependent endocytosis in MUC1 degradation (Fig. 2d). This aligns with previous research demonstrating that a reduced in MUC1 glycosylation alters steric pressure, promoting clathrin-mediated MUC1 endocytosis in breast cancer22,24. Taken together, these data suggest that MUC1 deglycosylation induces its clathrin-dependent endocytosis and lysosomal degradation.
E3 ligase NEDD4 mediates lysosomal degradation of MUC1
Endocytic downregulation serves as a fundamental mechanism that regulates the composition and organization of proteins at the plasma membrane. During this process, integral plasma membrane proteins can undergo ubiquitination, which marks them for (1) internalization via endocytosis, (2) sorting into multivesicular bodies, and (3) subsequent delivery to lysosomes. In certain instances, downregulated proteins directly interact with a specific ubiquitin ligase25. Notably, O-glycosidase treatment of HT-29 cells resulted in reduced O-glycosylation levels, which were associated with increased MUC1 ubiquitination (Fig. 2e). Therefore, we performed immunoprecipitation-mass spectrometry (IP-MS) proteomic analysis and identified E3 ligase Neural Precursor Cell Expressed Developmentally Down-regulated Protein 4 (NEDD4) as an interactor with MUC1, which was confirmed by IP experiment (Supplementary Data 2, Fig. 2f). Since NEDD4 plays a pivotal regulatory role in the ubiquitination and subsequent degradation of various membrane proteins25,26,27, we hypothesized that NEDD4 mediates the ubiquitination and lysosomal degradation of endocytosed MUC1. To investigate this, we knocked down NEDD4 expression in HT-29 cells and observed that, in the presence of O-glycosidase, the knockdown cells exhibited a higher level of MUC1 (Fig. 2g).
The NEDD4C1046A mutant, which is devoid of E3 ligase activity, showed minimal or no impact on MUC1 turnover (Fig. 2h, i), indicating that E3 ligase activity is essential for NEDD4-mediated MUC1 degradation. Accordingly, while the C1046A mutant does not ubiquitinate MUC1, the wild-type NEDD4 does exhibit this capability (Fig. 2j). K48- and K63-linked polyubiquitination are typically recognized as signals for substrate protein degradation. Our findings indicate that MUC1 ubiquitination by NEDD4 is primarily mediated through K63 linkage, rather than K48 linkage (Fig. 2k). To identify potential ubiquitination sites on MUC1, we mutated candidate lysine residues. Notably, the MUC1K1231R mutant was nearly completely resistant to NEDD4-mediated degradation (Fig. 2l, Supplementary Fig. 2d–f). These findings indicate that NEDD4 primarily facilitates K63-linked ubiquitination of MUC1 at the K1231 residue. Taken together, these data suggest that NEDD4 mediates MUC1 degradation via the ubiquitin-mediated lysosomal pathway (Supplementary Fig. 2g).
Enterocyte-specific Muc1 knockout exacerbates MASLD progression in HFD-fed mice
To examine the role of intestinal MUC1 in MASLD progression in vivo, we generated enterocyte-specific Muc1 deletion mice (Muc1EKO) by crossing Muc1Flox/Flox mice with Villin-Cre mice. Western blot analysis confirmed the suppression of MUC1 in the intestine of Muc1EKO mice (Supplementary Fig. 3a). Subsequently, both male Muc1EKO mice and Muc1Flox/Flox mice were fed with either a normal diet (ND) or an HFD for 16 weeks (Fig. 3a, Supplementary Fig. 3b). Muc1EKO mice on an ND showed no differences in body weight, liver weight and hepatic lipid accumulation compared with Muc1Flox/Flox mice (Fig. 3b–h, Supplementary Fig. 3c–k).
a Experimental schematic of Muc1Flox/Flox and Muc1EKO mice after 16 weeks of ND or HFD diet. The diagram of the mouse was created in BioRender. Li, Z. (2025) https://BioRender.com/e2kqno5. Mice were housed at room temperature (RT, 21 °C) or thermoneutrality (TN, 29 °C). CLAMS, Comprehensive Laboratory Animal Monitoring System. Villin-Cre (Villin promoter-driven Cre recombinase), Muc1Flox/Flox (Muc1 with loxP sites flanking the gene), Muc1EKO (Epithelial-Knockout of Muc1), ND (Normal Diet), HFD (High-Fat Diet). b Typical liver H&E staining, oil red O staining, Sirius red staining, and Immunohistochemical staining of F4/80. Scale bars: 25 μm. c Body weight, liver weight. d Glucose tolerance test (GTT). e Insulin tolerance test (ITT). f Quantitative analysis of Serum TG, Hepatic TG, ALT, and AST. g, h Inflammation-related (g) and Fibrosis-related (h) Gene expression in liver tissues. Data are mean ± s.e.m. n = 8 mice per group. P-values were calculated by a two-tailed unpaired t-test. Source data are provided as a Source data file.
After 16 weeks of HFD consumption, the deficiency of intestinal epithelial MUC1 led to increased body weight gain, liver weights and liver-to-body weight ratios without altering food intake or fecal lipid content (Fig. 3c, Supplementary Fig. 3c–f). Furthermore, glucose tolerance tests (GTT) and insulin tolerance tests (ITT) showed that intestinal epithelial MUC1 deficiency worsened HFD-induced glucose intolerance and insulin resistance (Fig. 3d, e). Serum and hepatic levels of TG, and LDL, as well as serum ALT levels, markers of hepatic toxicity, were significantly higher in Muc1EKO mice than those in Muc1Flox/Flox mice after HFD feeding (Fig. 3f, Supplementary Fig. 3h). Consistently, TUNEL, F4/80, and MPO staining demonstrated that inflammation in the livers of HFD-fed Muc1EKO mice (Fig. 3b, Supplementary Fig. 3g, j, k), as well as the mRNA expression of genes related to inflammation and fibrosis were also upregulated in the livers of Muc1EKO mice (Fig. 3g, h). Taken together, these data suggest that enterocyte-specific Muc1 knockout exacerbates MASLD progression in mice.
Enterocyte-specific Muc1 knockout exacerbates HFD-induced IEB impairment
Since MUC1 is localized within IEB and dysfunction of IEB plays a pivotal role in the progression of MASLD, we first examined the impact of enterocyte-specific Muc1 knockout on IEB integrity. Indeed, significantly higher serum LPS and bacteria levels were observed in Muc1EKO mice after HFD feeding (Fig. 4a, b), suggesting an increased IEB permeability. This finding was further corroborated by in vivo intestinal permeability assays using 4KDa FITC-Dextran flux (Fig. 4c). Subsequently, TEM analysis revealed a widening of zonula adherens between intestinal epithelial cells in Muc1EKO mice (Fig. 4d). Significantly elevated LPS levels were observed in HFD-induced MASLD mice and MASLD patients, and these levels were inversely correlated with intestinal MUC1 protein levels (Supplementary Fig. 4a–c). These data support that enterocyte-specific MUC1 deficiency impairs IEB integrity in HFD-fed mice.
a LPS Concentration were measured in the serum of Muc1Flox/Flox and Muc1EKO mice (n = 8 per group). b Bacterial DNA/Whole DNA in blood was measured by RT-qPCR (n = 8 per group). c Plasmatic 4KDa FITC Dextran Flux concentration 4 h after gavage by 750 µg/g bodyweight (n = 8 per group). d Representative transmission electron microscopy images and Width of colonic epithelial cell zonula adherence in mice (n = 8). Scale bars: 1 µm (left), 200 nm (right). e Fluorescent confocal images of colon tissue sections of mice and humans with MASLD and their control group. Scale bars: 40 μm. f Relative protein expression of MUC1 and β-Catenin in colon tissues of the control group and MASLD group (n = 30 per group). g Relative protein expression of MUC1 and β-Catenin in colon tissues of mice fed with ND and HFD (n = 6 per group). h Fluorescent confocal images of colon tissue sections of Muc1Flox/Flox and Muc1EKO mice. Scale bars: 40 μm. i Relative protein expression of MUC1, β-Catenin, and Phospho-β-Catenin in colon tissues of Muc1Flox/Flox and Muc1EKO mice (n = 8 per group). j Western blots for the indicated proteins from HT-29 cells transfected with shCTL or shMUC1 treated with 50 μM lysosome inhibitor CQ for 24 h or 10 μM proteasome inhibitor MG132 for 6 h, or DMSO as a control. CHX (Cycloheximide), DMSO (Dimethyl sulfoxide), CQ (Chloroquine), shCTL (shRNA Control), shMUC1 (shRNA targeting MUC1). Data are means ± SEM. P-values were calculated by a two-tailed unpaired t-test. Western blotting data are representative of three independent experiments. Source data are provided as a Source data file.
MUC1 deficiency leads to degradation of β-Catenin in intestinal epithelia
Previous studies conducted in cancer cells have demonstrated that MUC1 prevents the degradation of β-Catenin28,29,30, a key protein involved in the formation of intestinal epithelial zonula adherens and tumor progression, by sustaining β-Catenin dephosphorylation31,32. We first confirmed a decrease in intestinal β-Catenin levels in both individuals and mice with MASLD (Fig. 4e–g). Therefore, we hypothesized that reduced MUC1 may result in decreased β-Catenin levels in intestinal epithelia during MASLD progression. A similar trend was observed in Muc1EKO mice, with reduced intestinal β-Catenin levels compared to Muc1Flox/Flox mice (Fig. 4h, i). We also noticed an increase in phosphorylated β-Catenin levels in the intestine of Muc1EKO mice (Fig. 4h). However, no difference was observed in β-catenin mRNA levels (Supplementary Fig. 4d). Further in vitro experiments demonstrated that β-Catenin undergoes proteasomal degradation in MUC1-deficient conditions (Fig. 4j). Taken together, these data suggest that MUC1 deficiency facilitates phosphorylation and degradation of β-Catenin in intestinal epithelia.
Intestinal epithelial MUC1 maintains IEB integrity and mitigates MASLD progression via β-Catenin
To further investigate whether intestinal epithelial MUC1 influences MASLD progression via β-Catenin. We generated enterocyte-specific β-catenin knockout mice (β-cateninEKO) through crossing β-cateninFlox/Flox mice with Villin-Cre mice. Subsequently, Muc1 was enterocyte-specific overexpressed (Muc1EOE) in the male β-cateninEKO mice using enterotropic adeno-associated virus 9 (AAV9) carrying an MUC1 recombinase under the control of the enterocyte-specific Villin promoter (AAV-Villin-Muc1), followed by a 16-week HFD feeding (Supplementary Fig. 4f–g). Our findings revealed that enterocyte-specific Muc1 overexpression suppressed HFD-induced glucose intolerance, insulin resistance, hepatic fat accumulation, and IEB impairment. However, these beneficial effects were abolished when β-catenin was knocked out (Supplementary Fig. 4h–n). Taken together, our data suggest that intestinal epithelial MUC1 regulates IEB permeability of IEB and MASLD progression via β-Catenin (Supplementary Fig. 4o).
Overexpression of cytoplasmic-tail-deleted MUC1 in intestinal epithelia inhibits MASLD progression
Previous research has indicated that increased MUC1 promotes cancer development by elevating β-Catenin levels, a process mediated by the cytoplasmic tail of MUC121. Indeed, analysis reveals that enterocyte-specific overexpressing Muc1 results in a significantly elevated β-Catenin levels in mice (Supplementary Fig. 5a), suggesting a potential oncogenic risk. Previous research has also demonstrated that overexpression of cytoplasmic-tail-deleted MUC1 (MUC1ΔCT) in cancer cells reduces the removal of whole-length MUC1 from the plasma membrane by endocytosis22. Our above data showed that intestinal MUC1 downregulation occurs in MASLD progression through clathrin-dependent endocytosis and degradation. Therefore, we enterocyte specifically overexpressed the cytoplasmic-tail-deleted Muc1 (Muc1ΔCTEOE) in mice, followed by a 16-week HFD feeding (Fig. 5a–c, Supplementary Fig. 5b–d), and found MUC1ΔCT overexpression prevented HFD-induced degradation of MUC1 and β-Catenin. Compared with the overexpression of wild-type MUC1 in intestinal epithelia, the overexpression of MUC1ΔCT did not lead to an extremely significant increase in β-Catenin levels, but only maintained the physiological level of β-Catenin (Supplementary Fig. 5e).
a Schematics of mice cytoplasmic-tail-deleted MUC1 constructs. S (Signal Sequence), VNTR (Variable Number of Tandem Repeats), SEA (Sea urchin sperm protein-Enterokinase-Agrin domain), TM (Transmembrane Domain), CT (Cytoplasmic Tail), ΔCT (Cytoplasmic Tail Deletion). b Experimental schematic of Control, Muc1ΔCTEOE, and Muc1EOE mice after 16 weeks of HFD. The diagram of the mouse was created in BioRender. Li, Z. (2025) https://BioRender.com/e2kqno5. Mice were housed at room temperature (RT, 21 °C) or thermoneutrality (TN, 29 °C). CLAMS, Comprehensive Laboratory Animal Monitoring System. Villin-Cre (Villin promoter-driven Cre recombinase), AAV9 (Adeno-Associated Virus Serotype 9), ΔCT (Cytoplasmic Tail Deletion), EOE (Epithelial Overexpression), HFD (High-Fat Diet). c Representative transmission electron microscopy images of the glycocalyx. Scale bar, 100 nm (left), 200 nm (right). d Typical liver H&E staining, oil red O staining. Scale bars: 25 μm. e Body weight, and Liver weight. f Glucose tolerance test (GTT). g Insulin tolerance test (ITT). h Quantitative analysis of Serum TG, Hepatic TG, ALT, and AST. i LPS Concentration was measured in the serum and Plasmatic 4KDa FITC Dextran Flux concentration 4 h after gavage by 750 µg/g bodyweight. Data are means ± SEM, n = 8 mice/group. P-values were calculated by a two-tailed unpaired t-test. Source data are provided as a Source data file.
Further assessment revealed that overexpressing MUC1ΔCT in intestinal epithelia improved hepatic steatosis, which was not significantly different from the effect of MUC1 overexpression (Fig. 5d–h, Supplementary Fig. 5f–h). Additionally, MUC1ΔCT overexpression maintained the integrity of IEB (Fig. 5i). Overall, overexpressing MUC1ΔCT in enterocytes represents an effective and potentially safe method for preserving MUC1 levels and IEB integrity, as well as mitigating the progression of MASLD.
Discussion
Extensive research indicates that impairment of the IEB contributes to the development of MASLD, yet the underlying molecular elements that trigger and drive IEB impairment are not well understood. In the present study, a notable decrease in intestinal MUC1 levels in MASLD patients is observed, with its expression negatively correlating with BMI and hepatic toxicity. The causal role of intestinal epithelial MUC1 in MASLD progression is further investigated in enterocyte-specific Muc1 knockout mice. We observe that a reduction of MUC1 impairs IEB integrity, leading to severe hepatic steatosis. Furthermore, our findings reveal that during MASLD development, MUC1 in enterocytes undergoes deglycosylation, triggers its clathrin-mediated endocytosis of MUC1 and lysosomal degradation via NEDD4-catalyzed ubiquitination. This degradation process further triggers the proteolysis of β-catenin, a protein essential for maintaining the functionality of zonula adherens in intestinal epithelia. The proteolysis of β-catenin increases of IEB permeability, thereby promoting the progression of MASLD. Furthermore, this study proposes a potentially effective approach for MASLD, which involves the overexpression of the cytoplasmic-tail-deleted MUC1 in intestinal epithelia for preserving MUC1 levels and IEB integrity.
Intestinal MUC1 is a transmembrane mucin located in the IEB20,33. Its extracellular domain connects the intestinal epithelia with the mucus layer, while its cytoplasmic tails possess signaling capacities18. Previous research has shown that aberrant expression of intestinal MUC1 contributes to the progression of colitis and colorectal cancer20,21. The glycocalyx, a complex and heterogeneous structure on the cell surface, is formed by highly glycosylated MUC1, along with glycolipids and polysaccharides24. In this study, we observed by TEM that the enterocytes of humans and mice with MASLD have significantly less glycocalyx than those of the control groups, and further noted that deglycosylation of intestinal epithelial MUC1 is associated with reduced MUC1 protein levels.
Recent studies have reported that MUC1 deglycosylation reduces steric pressure and increases MUC1 endocytosis22. Highly glycosylated MUC1 is capable of generating steric pressure, which promotes the formation of spherical and finger-like extensions on the cell surface24, MUC1 deglycosylation leads to a decrease in biopolymers and long-chain polysaccharides within the glycocalyx on the cell surface, potentially reducing the spatial volume of MUC1, which further subdues the limit by MUC1 on transmembrane protein quantity in each clathrin-coated structure, and increases the capacity of endocytotic regoin for transmembrane proteins, ultimately increasing the amount of MUC1 that can be involved in a endocytotic vesicle22,34. We observed that MUC1 deglycosylated by treatment with O-glycosidase can increase MUC1 endocytosis and reduce MUC1 protein levels in an intestinal epithelial cell line. Additionally, we genetically manipulated the glycosylation levels of MUC1 and found that enterocytes with lower MUC1 glycosylation levels exhibit lower MUC1 levels compared to those with higher MUC1 glycosylation levels. This may explain why MUC1 levels are reduced during the progression of MASLD.
Studies have indicated that the composition of gut microbiota and microbial metabolites has an impact on mucin glycans, which involves the regulation of signaling pathways35. Changes in the glycosylation patterns of intestinal mucins have been linked to gut diseases such as irritable bowel syndrome, idiopathic chronic diarrhea, inflammatory bowel diseases, and colorectal cancer, which often occur concurrently with disruptions in the intestinal microbiota36,37,38,39,40. Apart from gut microbiota, environmental factors such as dietary styles and food ingredients have demonstrated the ability to affect mucus secretions and glycosylation profiles in the host intestine, either by regulating mucosal immune signals or by altering the gut microbiota41,42. Of course, the reason why intestinal MUC1 undergoes deglycosylation during the progression of MASLD requires further experimental verification in later stages.
NEDD4, alternatively known as NEDD4-1, is a member of the HECT E3 ubiquitin ligase family and was originally identified for its role in the proteasomal degradation of the epithelial sodium channel (ENaC)43. Previous research has demonstrated that intestinal NEDD4 regulates intestinal stem cell homeostasis44,45 and plays a regulatory role in endocytosis as well as the transportation of membrane proteins into vacuoles46,47. Similarly, other studies have reported that NEDD4 E3 ligase is involved in the endocytogenic degradation of multiple membrane proteins through the ubiquitin system25,26,48,49. This study reveals that endocytosed MUC1 is mediated by NEDD4 for lysosomal degradation. Given that the HECT domain is a functional region of the NEDD4 ubiquitin ligase responsible for protein degradation50, and that NEDD4C1046A mutant lacking E3 ligase activity had no impact on MUC1 degradation, our study suggests E3 ligase activity to be crucial for NEDD4 to degrade MUC1. Overall, we show that NEDD4 is the ubiquitin E3 ligase to mediate MUC1 ubiquitination and degradation.
Next, we investigated the role of decreased MUC1 levels in the progression of MASLD. Previous research has established that IEB disruption is pivotal in MASLD progression12. The IEB comprises both a mucus layer and intestinal epithelia7. The mucus layer forms a dynamic and chemically complex barrier, primarily consisting of secreted mucins51, which create a disulfide cross-linked network that expands to produce an inner layer securely attached to the epithelium52. Transmembrane mucins are highly expressed in the intestinal epithelium, with their extracellular domain acting as a bridge between the intestinal epithelia with the mucus layer, while their cytoplasmic tails possess signaling capacities18,53. Studies have reported that the degradation of mucins within the IEB increases disease susceptibility54. In particular, deficiency in intestinal mucin 2 contributes to steatohepatitis15,55, and loss of MUC13 induces intestinal inflammation53. In this study, we found that deletion of MUC1, a transmembrane mucin, in enterocytes leads to a disruption of IEB integrity and a decrease in β-Catenin levels. The integrity of the intestinal epithelium is heavily reliant on the expression of intercellular junction protein structures, such as tight junctions, zonula adherens, desmosomes, and gap junctions, which physically connect and seal the intercellular spaces within the intestinal epithelial layer56,57. Intestinal β-Catenin is demonstrated by previous studies to be of vital importance in the maintenance of zonula adherens31,32, and a reduction in β-Catenin level contributes to the development of MASLD12. Previous studies find that stabilization of MUC1 protein plays a crucial role in maintaining the stability of β-Catenin in tumor cells29. We revealed for the first time that the MUC1 protein can mitigate the progression of MASLD by stabilizing β-Catenin in enterocytes, thereby maintaining the integrity of IEB. This discovery provides a new approach for the prevention and treatment of MASLD by targeting intestinal MUC1.
Gene editing therapy holds promise in treating cancer and severe metabolic diseases58,59. In this study, we found that specific overexpression of the MUC1 protein in mouse enterocytes alleviated hepatic steatosis caused by HFD feeding. However, this overexpression also increased β-Catenin levels, indicating a potential risk of cancer development, as previous research has linked MUC1 overexpression to cancer development via β-Catenin elevation21. Notably, overexpressing a cytoplasmic-tail-deleted MUC1 in intestinal epithelia did not lead to an extremely significant increase in β-Catenin levels, but only maintained the normal β-Catenin levels after HFD feeding. This effect may be related to changes in cell steric pressure and subsequent inhibition of MUC1 endocytosis22. Interestingly, this modified overexpression mitigated HFD-induced hepatic steatosis, suggesting that overexpressing cytoplasmic-tail-deleted MUC1 in enterocytes could be an effective and potentially safe therapeutic approach for MASLD.
Despite the significant findings of our research, several limitations should be acknowledged. First, a single animal model cannot fully reflect the pathological process of the disease in humans. Therefore, the exploration of the role of MUC1 in MASLD necessitates validation in multiple animal models (e.g, mouse, rat, and monkey). Second, although we validated the interaction between MUC1 and NEDD4 through cellular assays and identified ubiquitination sites on MUC1, the precise binding site between these two molecules remains elusive and necessitates further investigation. Third, all study participants were male and recruited in China, which may introduce gender and ethnic biases and limit the generalizability of our findings to other populations. Future studies should adopt recruitment strategies that ensure broader demographic representation to minimize such selection bias. Fourth, as the Villin-Cre system targets both the small intestine and colon, our findings reflect intestinal-wide Muc1 deletion. However, since the current study primarily examined colonic tissue, future investigations incorporating small intestinal samples and cell lines are needed to delineate segment-specific effects. Fifth, given clinical feasibility limitations, were restricted to using colonic tissues adjacent to polyps from patients with MASLD and colonic polyps for investigating the relationship between MUC1 expression, intestinal epithelial integrity, and MASLD progression. It is not sufficient to rule out that patients may also have intestinal barrier dysfunction in the small intestine. Lastly, despite our results suggesting that overexpression of cytoplasmic-tail-deleted MUC1 attenuates HFD-induced MASLD in mice, the issue of long-term safety still requires further validation. Additionally, translation of these findings from preclinical studies to clinical applications requires rigorous evaluation through human clinical trials.
Methods
Human participants
This study successfully recruited a cohort of participants under strict adherence to ethical norms. All participating subjects voluntarily signed written informed consent forms based on full understanding, including consent for participation in the study and for the publication of potentially identifiable medical data, reflecting respect and protection for their rights and interests. Focusing on the pathological mechanisms of metabolic dysfunction-associated steatohepatitis (MASLD), this study meticulously collected colon mucosal biopsy samples from 30 patients with MASLD and 30 non-MASLD patients as the core materials for investigation. All recruited participants were adult Chinese males. We recruited patients with colonic polyps who were diagnosed with hepatic steatosis by liver ultrasound and had at least one of the following risk factors: 1, body mass index (BMI) ≥ 25 kg/m²; 2, elevated levels of serum alanine aminotransferase (ALT) and aspartate aminotransferase (AST); 3, elevated levels of serum triglycerides or serum cholesterol. The liver ultrasound examination results for the healthy control group were negative. Furthermore, experienced physicians carefully selected small pieces of colonic mucosal tissue from areas distant from the polyps and without obvious lesions, ensuring the representativeness and reliability of the samples. As liver biopsies could not be directly performed, patients could not be specifically classified as having metabolic-associated steatohepatitis (MASH). Therefore, the study cohort is collectively referred to as MASLD. We adopted strict screening criteria to minimize confounding factors: 1, no secondary causes of hepatic steatosis, such as the use of systemic corticosteroids or total parenteral nutrition; 2, no presence of other liver diseases, such as hepatitis B or C infections or autoimmune liver disorders; 3, no history of excessive alcohol consumption within the past 12 months (≥ 140 grams per week for males and ≥ 70 grams per week for females); 4, no use of hepatoprotective agents or exposure to hepatotoxic drugs in recent years.
Detailed clinical characteristics of the participants have been thoroughly documented in Supplementary Table 1, Supplementary Data 1, providing a solid foundation for subsequent data analysis. During sample collection, we strictly adhered to medical norms and operational guidelines. All subjects were diagnosed with colonic polyps, and small pieces of colon mucosal tissue were carefully selected by experienced physicians from areas distant from the polyps and without apparent lesions during colonoscopic polypectomy, ensuring the representativeness and reliability of the samples.
To ensure the accuracy and validity of the research findings, a series of rigorous exclusion criteria were established, excluding individuals with decompensated cirrhosis, moderate-to-severe chronic kidney disease, acute complications of diabetes, inflammatory bowel disease, tuberculosis, HIV/AIDS, malignancy, alcoholism, and those who had recently used medications that could potentially affect gut microbiota (such as antibiotics, probiotics, prebiotics, proton pump inhibitors, laxatives, etc.). Additionally, any subjects deemed unsuitable for biopsy by clinicians were also excluded, further safeguarding the scientific rigor and safety of the study. All clinical information related to the participants, including metadata such as physical characteristics, health status, age, and so forth, was comprehensively and meticulously collected and organized according to established standard procedures, providing abundant background data for subsequent data analysis and result interpretation.
Animals
Male SPF-grade C57BL/6 J mice were randomized into groups, with each group formed by 8 mice. They were housed under standard laboratory conditions in the Laboratory Animal Science Department of the China-Japan Friendship Institute of Clinical Medicine. The mice were housed together in the same cage and had unrestricted access to rodent feed and water. The specific environmental conditions included a 12-h light cycle from 08:00 to 20:00, with the temperature maintained between 21–24 °C and humidity levels kept at 40–70%.
Muc1Flox/Flox and β-CateninFlox/Flox mice were purchased from Cyagen (S-CKO-03832, S-CKO-01559). To induce intestinal Muc1 disruption, Muc1Flox/Flox and β-cateninFlox/Flox mice were crossed with Villin-Cre mice (kindly provided by Cyagen), resulting in the generation of Muc1EKO and β-cateninEKO mice. Villin-Cre mice specifically express Cre recombinase in the villus and crypt epithelial cells of both the small and large intestines.
Cre-dependent overexpression AAV was constructed by inserting the positive and reverse sequences of Loxp and Lox2272 into the polyclonal sites of pAAV-MCS (Public Protein/Plasmid Library, PPL) plasmid, respectively(pAAV-MCS-LOXP). Subsequently, the CDS region of Muc1 and the cytoplasmic-tail-deleted Muc1 were inserted into the polyclonal site of pAAV-MCS-Loxp to generate the plasmids pAAV-Muc1-Loxp and pAAV-Muc1ΔCT-Loxp. These plasmids were then co-transfected with pAAV-RC2/9 and pAAV-Helper vector into AAVpro 293 T cells (Takara) to produce recombinant AAV, designated as AAV-Muc1 and AAV-Muc1ΔCT. AAV-Villin-Muc1 was constructed using Muc1 recombinase driven by the enterocyte-specific Villin promoter based on pAAV-MCS. The original vector was synthesized without any modification as a control group, referred to as AAV9-Control. Eight-week-old male knockout mice fed with ND (Purina Mills 5053) were created by transduction of either AAV-Muc1 or AAV-Muc1ΔCT. A dose of 1.5 × 1011 genome copies per mouse was used for the administration of AAVs.
Eight-week-old male mice were randomized into groups and fed either a high-fat diet (HFD, Catalog No. D12492) or a normal diet (ND, Catalog No. D09100310) obtained from Nanjing Qingzilan Technology Co., Ltd. (China). The HFD composition was as follows: 60% fat, 20% protein, and 20% carbohydrate. Prior to euthanasia, the mice were weighed using a scale to analyze their body weight composition. Immediately after euthanasia, the livers were removed and weighed. To conduct biochemical analysis, serum triglyceride and total liver cholesterol (TC) levels were quantified using kits provided by Wako Diagnostics (Wako Chemicals), in accordance with the manufacturer’s instructions. Alanine aminotransferase (ALT) and aspartate aminotransferase (AST) levels were measured using commercial kits obtained from Catachem, whereas low-density lipoprotein (LDL) and high-density lipoprotein (HDL) levels were assessed using commercial kits from Abcam.
Cell lines and cell culture
Both HT-29 and HEK293T cells originate from the prestigious Cell Resource Center of Peking Union Medical College in Beijing, China, ensuring the reliability and purity of their cellular sources. HT-29 cells were meticulously cultivated in DMEM/F12 medium supplemented with 10% fetal bovine serum (FBS), under rigorously controlled conditions of a constant temperature of 37 °C and a humidified atmosphere containing 5% CO2, to mimic optimal physiological conditions for cell growth. Similarly, HEK293T cells thrived under similar optimal conditions, utilizing high-quality DMEM medium supplemented with 10% FBS sourced from Gibco (Grand Island, NY, USA). This choice not only ensured the nutrient richness of the medium but also maintained consistency and high standards throughout the cell culture process. To preserve the authenticity and integrity of the cell lines, both HT-29 and 293T cells underwent rigorous short tandem repeat (STR) analysis, a molecular biology technique that precisely verifies the identity of cell lines, safeguarding against cross-contamination or misidentification. Furthermore, comprehensive detection for mycoplasma contamination was performed on both cell types, with all results indicating freedom from mycoplasma contamination, further enhancing the purity and safety of the cell culture environment. The intervention method for O-deglycosylation experiments in HT-29 cells was as follows: HT-29 cells were treated with 50 nM O-glycosidase in complete medium for 30 min, followed by incubation with cycloheximide (CHX, a protein synthesis inhibitor) for 24 h.
Plasmids and transfection
For plasmid transfection of HT-29 cells, Lipofectamine™ 3000 transfection reagent (Invitrogen™) was used, while Lipo8000 transfection reagent (Beyotime, China) was used for plasmid transfection of HEK293T cells. Transfection procedures adhered to the procedures of the manufacturers’ instructions.
cDNAs encoding human Muc1 with 5 tandem repeats (Muc1-5TRs), 10 tandem repeats (Muc1-10TRs) generated and cloned into the mammalian expression vector pcDNA3.1, as described previously24. All MUC1 proteins were fused with a Flag tag. The cDNA was amplified using PCR and then cloned into the pcDNA3.1 vector (Invitrogen) to generate the NEDD4 expression plasmid. The MUC1 overexpression vector was synthesized by PPL and cloned into the pcDNA3.1 vector. By amplifying cDNA through PCR and cloning it into the pcDNA3.1 vector with a Flag tag and the pcDNA3.1 vector with a 6 × His tag, respectively, the Flag-MUC1 and His-NEDD4 expression plasmids were constructed. The HA-Ub, HA-K48, and HA-K63 overexpression vectors were provided by the pediatric research team at the China-Japan Friendship Hospital.
NEDD4 knock down cells were established by the NEDD4-specific shRNA sequences shCTL: 5’-CTCGCTTGGGCGAGAGTAA-3’; shNEDD4-#1: 5’- CCTAGGTCAAGTGGATGTTCCACTT-3’; shNEDD4-#2: 5’- GAGATGATTTCCTAGGTCAAGTGGA-3’). MUC1 knock down cells were established by the MUC1-specific shRNA sequences shCTL: 5’-CTCGCTTGGGCGAGAGTAA-3’; shMUC1-#1: 5’-CAGCCTCTCTTACACAAACCCAGCA-3’; shMUC1-#2: 5’-GCCTCTCTTACACAAACCCAGCAGT-3’. NEDD4C1046A, NEDD4C1197A and NEDD4C1286A point mutation overexpression vectors of NEDD4 were constructed based on NEDD4 overexpression vectors using point mutation kits (Vazyme). MUC1K988R, MUC1K1011R, MUC1K1082R, MUC1K1093R, MUC1K1125R, MUC1K1189R, and MUC1K1231R were obtained using Full-Gene Synthesis methods and were then cloned into corresponding vectors by point mutation kits (Vazyme).
Measurement of Glycocalyx layer thickness and Width of zonula adherence
The thickness of the glycocalyx layer and the width of the zonula adherence were measured on each section using transmission electron microscopy (at a magnification of 20,000x), with a focus on the visible glycocalyx layer and zonula adherence. At least 10 measurements were taken for each section, and at least two sections were obtained from each sample.
For each sample, the mathematical mean was calculated based on the readings obtained from all sections, with each control sample represented by a single data point. One sample was collected from each selected site, and sections were prepared for measuring the thickness of the glycocalyx layer and the width of the zonula adherence. The mean of at least 10 measurements was calculated. Each sample was represented by the mathematical mean of several data points obtained from the corresponding anatomical site60.
GTT and ITT
Three weeks prior to euthanasia, glucose tolerance tests (GTT) were performed on the mice, while insulin tolerance tests (ITT) were conducted two weeks before euthanasia. Both tests were carried out after a 6-h fasting period for the mice. In the GTT, the mice were administered 1.25 g/kg of D-glucose via intraperitoneal injection. For the ITT, the mice received an intraperitoneal injection of 1.2 U/kg of insulin (NovoRapid). Blood glucose levels were measured at 0, 30, 60, 90, and 120 min post-injection using a Roche ACCU-CHEK Aviva handheld glucometer for all tests. Blood samples were collected by taking a drop of blood from the tail of the mice. The area under the curve was analyzed using GraphPad Prism software (version 9).
Histological analysis
Mouse liver tissues were preserved in 4% paraformaldehyde solution or embedded in OCT embedding medium. In contrast, human or mouse colonic tissues were fixed using either 4% paraformaldehyde or Carnoy’s fixative, the latter being particularly suitable for mucosal tissues. Liver tissue sections were fixed in formaldehyde, embedded in paraffin, and stained with hematoxylin and eosin (H&E) for assessing liver lipid accumulation and with Sirius Red for evaluating fibrosis. Frozen liver sections were stained with Oil Red O to visualize and assess the presence of lipid droplets. All procedures were rigorously conducted according to established protocols before microscopic examination. Histological scoring assessments were performed based on the reported MASLD scoring system61.
Confocal microscopy for tissues
Human and mouse colonic tissues were initially fixed in 4% paraformaldehyde solution to enhance tissue stability and morphological preservation. Following this, tissue samples were immersed in Triton X-100 for 10 min to promote cell membrane permeabilization, facilitating the penetration of subsequent stains. Subsequently, tissues were blocked for 1 h at room temperature using Beyotime QuickBlock™ Blocking Buffer to reduce non-specific binding. Primary antibodies or Biotin-labeled peanut agglutinin were appropriately diluted in Beyotime Immunostaining Primary Antibody Diluent and co-incubated with tissue samples overnight at 4 °C to ensure sufficient binding between antibodies and target molecules. After completion of primary antibody incubation, tissues were thoroughly washed three times with phosphate-buffered saline (PBS) to remove unbound antibodies. Next, fluorescently labeled secondary antibodies (secondary antibodies) or Streptavidin labeled with SAlexa Fluor 488 were incubated with tissues for 2 h at 37 °C, allowing them to specifically recognize and bind to primary antibodies already bound to target molecules, thereby emitting fluorescent signals. After another three washes with PBS, DAPI (a fluorescent dye commonly used for nuclear staining) was added to the samples. Finally, tissue samples were mounted using mounting medium produced by Sigma-Aldrich (St. Louis, MO, USA) to protect the samples and reduce fluorescence quenching. Upon completion of sample preparation, tissues were observed and imaged in detail using a Zeiss 880 upright confocal microscope equipped with a 40x water immersion objective lens.
Transmission electron microscopy
The transmission electron microscopy (TEM) imaging was accomplished on a cutting-edge technological instrument, the Titan Themis microscope manufactured by Thermo Fisher Scientific in Waltham, Massachusetts, USA. Operating with exceptional 300 kV energy-filtering capabilities, this microscope ensures precision and depth in imaging. This microscope not only incorporates advanced field-emission gun technology but is also equipped with the Gatan K2 Summit direct detection camera from Gatan Inc. in Pleasanton, California, USA. Renowned for its astonishing high resolution of 3838 × 3710 pixels, this camera seamlessly switches between counting mode and dose fractionation mode, enabling the capture of the finest image details. During the image acquisition process, we meticulously set a precise defocus range from −1 mm to −3 mm and skillfully employed a 2x pixel binning technique to further optimize image quality. This processing refined the final image pixel size to between 0.72 and 1.1 nanometers, significantly enhancing imaging resolution and clarity, providing unprecedented detail insights for scientific research.
Quantitative MS
HT-29 cells were lysed, followed by immunoprecipitation with specific antibodies and Protein A magnetic beads (Santa Cruz, CA, USA) in NETN150 buffer containing IP-grade protease/phosphatase inhibitors. Magnetic beads were washed five times to ensure purity. Immunoprecipitated proteins were separated by SDS-PAGE, stained with Coomassie Blue, and target bands were excised.
In-gel digestion: Gel pieces were washed with 50% ACN/water and 50% ACN/100 mM NH₄HCO₃ (≥ 3×) until colorless, dried with ACN, reduced with TCEP, alkylated with IAA, re-washed, dried, and digested with Sequencing Grade trypsin at 37 °C overnight. Peptides were extracted and vacuum-dried.
Samples were analyzed on an Orbitrap Fusion LUMOS mass spectrometer (Thermo Fisher Scientific) coupled with an Easy Spray source and Easy-nLC 1200 system. Loaded onto a PepMap™ Neo trap column, peptides were separated on an in-house packed C18 column (300 nL/min) with a 70-min gradient (4–8% B for 1 min, 8–38% B for 45 min, 38–55% B for 5 min, 55–100% B for 1 min, 100% B for 8 min; B: 0.1% FA in 80% ACN/20% water). MS operated in positive DDA mode with FAIMS Pro (−65/−45 V). Parent scans (350–1500 m/z, 60,000 resolution, 300% AGC) and HCD MS/MS (28/30 NCE for −65/−45 V, 15,000 resolution, 1.6 m/z window, 75% AGC) were acquired.
Data were searched against the Uniprot human library using Proteome Discoverer 2.5 (Thermo Scientific) with trypsin (≤ 2 missed cleavages), fixed (Cys carbamidomethylation) and variable (Met oxidation) modifications, 12 ppm precursor/±0.02 Da fragment tolerance, and Percolator-filtered FDR < 1%.
Quantitative PCR (qPCR)
Human and mouse colonic tissues were promptly immersed in liquid nitrogen for instant freezing and stored appropriately at −80 °C to ensure the integrity and stability of biological samples. Subsequently, following the strict operational guidelines provided by Invitrogen (Carlsbad, California, USA), total RNA was efficiently extracted from these tissues using Trizol reagent. Next, the extracted RNA was precisely reverse-transcribed into cDNA using the M-MLV Reverse Transcriptase Kit manufactured by Promega (Madison, Wisconsin, USA), laying a solid foundation for subsequent experiments. During the PCR amplification phase, we adhered strictly to the protocols established by Applied Biosystems (Warrington, UK), employing the SYBR Green PCR Master Mix Kit and performing the operations on the advanced ABI Prism 7900 HT Fast Real-Time PCR System. This ensured precision and efficiency in the amplification process. By monitoring the cycle threshold (Ct) values, we were able to accurately assess the expression levels of each target mRNA. To enhance the reliability and reproducibility of the experimental results, the aforementioned procedures were meticulously repeated three times. Furthermore, all primer sequences involved in this study are detailed in Supplementary Table 2 for future reference and validation by other researchers.
Western blot analysis
Tissue and cell extracts were first meticulously homogenized in an ice-cold RIPA buffer containing protease and phosphatase inhibitors to ensure complete lysis of samples and uniform distribution of components. Subsequently, these homogenized samples were carefully loaded onto 6% SDS-PAGE gels for electrophoretic separation, arranging them according to their molecular weights. Upon completion of electrophoresis, proteins were efficiently transferred onto polyvinylidene fluoride (PVDF) membranes to facilitate subsequent immunodetection procedures. Next, the PVDF membranes were placed on an orbital shaker and subjected to blocking treatment using Beyotime QuickBlock™ Blocking Buffer, which is fully compatible with lectins. This step aimed to reduce non-specific binding and enhance the accuracy of subsequent antibody recognition. Following blocking, the membranes were incubated overnight at 4 °C with primary antibodies or biotin-labeled peanut agglutinin, precisely diluted in Beyotime Western blot Primary Antibody Dilution Buffer that is also compatible with lectins, ensuring thorough binding of primary antibodies to target proteins. On the following day, the membranes underwent three thorough washes with TBST buffer at room temperature, each lasting 5 min, to remove unbound primary antibodies and impurities. Subsequently, the membranes were transferred to a fresh buffer system and incubated with secondary antibodies, appropriately diluted in Beyotime Western blot Secondary Antibody or streptavidin labeled with Horseradish Peroxidase Dilution Buffer (also compatible with lectins), for 1 h at room temperature with gentle agitation to promote specific binding between secondary antibodies and primary antibodies. Finally, the secondary antibody solution was removed, and the membranes were subjected to three additional washes with TBST buffer at room temperature, each lasting 5 min, to thoroughly eliminate unbound secondary antibodies, preparing them for subsequent visualization or imaging steps. For Western blot visualization, protein bands were detected using the BIO-RAD ChemiDoc™ MP Imaging System with enhanced chemiluminescence (ECL) substrate, and band intensities were quantitatively analyzed by Image Lab™ software (BIO-RAD, Hercules, CA, USA).
Antibodies
Lectin from Arachis hypogaea (peanut) (Sigma Aldrich, L6135, Biotin conjugate), Rabbit anti-MUC1 (Abclonal, A21726), anti-MUC1 (Invitrogen, PA5-95487), anti-MUC1 (Santa cruz, sc-53381), anti-NEDD4 (Proteintech, 21698-1-AP), anti-GAPDH (Proteintech, 10494-1-AP), anti-LC3 (Proteintech, 14600-1-AP), anti-Ub (Wanleibio, WL01368), anti-HA Tag (Proteintech, 81290-1-RR), anti-His-Tag (Proteintech, 66005-1-Ig), anti-DYKDDDDK tag (Proteintech, 80010-1-RR), Mouse anti-β-Catenin (Santa Cruz Biotechnology, sc-7963), anti-p-β-Catenin (Santa Cruz Biotechnology, sc-57533) were used in this study.
Fecal lipid content
During the 12-week high-fat feeding period, feces were collected from the 10th to 12th week of the feeding regimen and subsequently dried at 60 °C for 3 days. Afterward, 0.5 grams (dry weight) of feces were dissolved in water overnight, followed by lipid extraction using the Folch method62. The extracted lipids were placed in pre-weighed glass tubes, and a 2:1 chloroform/methanol (volume/volume) solution was added. The samples were then completely dried under a nitrogen stream, and the recovered lipid mass was determined by calculating the weight difference. To determine the percentage of fecal lipids, the weight of the extract was divided by the initial dry weight of the feces.
Quantification of bacterial DNA in blood
The measurement of bacterial DNA levels in blood was performed based on a previously published method with certain adjustments63. The total DNA was extracted from 50 μL of blood using a commercial kit supplied by Tiangen Biotech (Beijing) Co., Ltd., China. DNA concentrations were determined using a Nanodrop 2000 spectrophotometer (Thermo Fisher Scientific Inc., USA), and the amount of bacterial DNA in the blood samples was quantified utilizing the FemtoTM Bacterial DNA Quantification Kit from Zymo Research, USA.
Serum LPS detection
Serum LPS levels were measured using the Lonza QCL-1000® LAL chromogenic assay (Lonza, Basel, Switzerland) following the manufacturer’s instructions. Fasting serum samples were diluted 1:5 in endotoxin-free water, heat-inactivated at 56 °C for 30 min, and assayed in duplicate. Standards (0.005–5 EU/mL) and controls were included in each run. Absorbance was read at 405–410 nm, and LPS concentrations were calculated from a 4-parameter logistic standard curve. Values below the limit of detection (0.005 EU/mL) were reported as ND. Results were expressed in EU/mL.
To assess the reliability and accuracy of the Limulus Amebocyte Lysate (LAL) assay, endotoxin recovery rates were assessed. All tested samples were subjected to endotoxin spiking and recovery analysis. Prior to analysis, samples were diluted appropriately (1:5) using endotoxin-free water to minimize potential interference from sample constituents. Endotoxin standards (CSE) were obtained from Lonza, and spiking was performed using a known concentration of endotoxin (0.1 EU/mL and 1 EU/mL). Each sample was spiked in parallel with the same volume of CSE and incubated under assay conditions. The percent recovery was calculated using the formula: Recovery (%) = (Measured Endotoxin in Spiked Sample − Measured Endotoxin in Unspiked Sample) / Spiked Endotoxin × 100. Acceptable recovery rates were defined as 50% to 200%, in accordance with the manufacturer’s instructions for the LAL assay kit. All spiked recovery tests were performed in duplicate. If recovery fell outside the acceptable range, the sample was considered to exhibit matrix interference, and further dilution or additional validation was performed until satisfactory recovery was achieved. In this study, recovery rates for all tested sample types ranged from 85% to 110%, indicating minimal matrix interference and confirming the suitability of the assay conditions. These data ensured the quantitative reliability of endotoxin measurements reported herein.
In vivo intestinal permeability assessment
Following a 4-h fast, mice were orally gavaged with 4 kDa FITC-dextran (Merck, Cat# FD4). After 90 min, blood was collected via cardiac puncture, and plasma was separated. FITC concentration was measured using a fluorospectrometer (excitation/emission: 485/535 nm) against a standard curve (0.01–10 μg/mL).
Statistical analysis
All data were comprehensively and rigorously analyzed using the advanced statistical software GraphPad Prism 9. For statistical comparisons, we rigorously employed the unpaired Student’s t-test method to ensure the accuracy and scientific validity of the results. In data presentation, we adopted the format of mean ± standard error of the mean (SEM), aiming to clearly and intuitively convey the central tendency and dispersion of the data. To investigate the associations between variables, we further conducted a two-tailed, uncorrected analysis of the Pearson product-moment correlation coefficient, which effectively evaluates the degree of linear correlation between two variables. In calculating P-values, we adhered to the principles of the two-tailed unpaired t-test, setting p > 0.05 as the statistical criterion for determining non-significant differences. This decision-making process ensures the rigor and reliability of our conclusions.
Ethics statement
This study used human samples under strict adherence to ethical norms and obtained formal approval from the Ethics Committee of China-Japan Friendship Hospital (Authorization Number: 2021–168), ensuring the legality and ethicality of the research process. All animal experiments were carried out in compliance with the animal use, handling, and euthanasia protocols granted approved by the China-Japan Friendship Institute of Clinical Medicine (License Number: 2022065 M).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All the data supporting the findings of this study are included in the Article and Supplementary Information. The protein mass spectrometry raw data generated in this study have been deposited in the ProteomeXchange database under accession code PXD066810. Source data are provided with this paper.
References
Matchett, K. P., Paris, J., Teichmann, S. A. & Henderson, N. C. Spatial genomics: mapping human steatotic liver disease. Nat. Rev. Gastroenterol. Hepatol. 21, 646–660 (2024).
Powell, E. E., Wong, V. W. & Rinella, M. Non-alcoholic fatty liver disease. Lancet 397, 2212–2224 (2021).
Wu, Q. et al. Intestinal hypoxia-inducible factor 2α regulates lactate levels to shape the gut microbiome and alter thermogenesis. Cell Metab. 33, 1988–2003.e1987 (2021).
Kuang, J. et al. Hyodeoxycholic acid alleviates non-alcoholic fatty liver disease through modulating the gut-liver axis. Cell Metab. 35, 1752–1766.e1758 (2023).
Zhang, Z. et al. Deficiency of ASGR1 promotes liver injury by increasing GP73-mediated hepatic endoplasmic reticulum stress. Nat. Commun. 15, 1908 (2024).
Johansson, M. E., Sjövall, H. & Hansson, G. C. The gastrointestinal mucus system in health and disease. Nat. Rev. Gastroenterol. Hepatol. 10, 352–361 (2013).
Buckley, A. & Turner, J. R. Cell biology of tight junction barrier regulation and mucosal disease. Cold Spring Harb. Perspect. Biol. 10, a029314 (2018).
Arab, J. P., Arrese, M. & Trauner, M. Recent insights into the pathogenesis of nonalcoholic fatty liver disease. Annu Rev. Pathol. 13, 321–350 (2018).
Miele, L. et al. Increased intestinal permeability and tight junction alterations in nonalcoholic fatty liver disease. Hepatology 49, 1877–1887 (2009).
Rahman, K. et al. Loss of junctional adhesion molecule a promotes severe steatohepatitis in mice on a diet high in saturated fat, fructose, and cholesterol. Gastroenterology 151, 733–746.e712 (2016).
Hou, Z. et al. Intestinal epithelial β Klotho is a critical protective factor in alcohol-induced intestinal barrier dysfunction and liver injury. EBioMedicine 82, 104181 (2022).
Mouries, J. et al. Microbiota-driven gut vascular barrier disruption is a prerequisite for non-alcoholic steatohepatitis development. J. Hepatol. 71, 1216–1228 (2019).
Natividad, J. M. et al. Impaired aryl hydrocarbon receptor ligand production by the gut microbiota is a key factor in metabolic syndrome. Cell Metab. 28, 737–749.e734 (2018).
Paone, P. & Cani, P. D. Mucus barrier, mucins and gut microbiota: the expected slimy partners?. Gut 69, 2232–2243 (2020).
Hartmann, P. et al. Deficiency of intestinal mucin-2 protects mice from diet-induced fatty liver disease and obesity. Am. J. Physiol. Gastrointest. Liver Physiol. 310, G310–G322 (2016).
Ng, G. Z. et al. The MUC1 mucin protects against Helicobacter pylori pathogenesis in mice by regulation of the NLRP3 inflammasome. Gut 65, 1087–1099 (2016).
Kufe, D. W. Mucins in cancer: function, prognosis and therapy. Nat. Rev. Cancer 9, 874–885 (2009).
Martínez-Sáez, N., Peregrina, J. M. & Corzana, F. Principles of mucin structure: implications for the rational design of cancer vaccines derived from MUC1-glycopeptides. Chem. Soc. Rev. 46, 7154–7175 (2017).
McAuley, J. L. et al. MUC1 cell surface mucin is a critical element of the mucosal barrier to infection. J. Clin. Invest 117, 2313–2324 (2007).
Nishida, A. et al. The membrane-bound mucin Muc1 regulates T helper 17-cell responses and colitis in mice. Gastroenterology 142, 865–874.e862 (2012).
Ahmad, R. et al. Targeting MUC1-C inhibits the AKT-S6K1-elF4A pathway regulating TIGAR translation in colorectal cancer. Mol. Cancer 16, 33 (2017).
Gollapudi, S. et al. Steric pressure between glycosylated transmembrane proteins inhibits internalization by endocytosis. Proc. Natl Acad. Sci. USA 120, e2215815120 (2023).
Zhu, J. et al. The deubiquitinase USP11 ameliorates intervertebral disc degeneration by regulating oxidative stress-induced ferroptosis via deubiquitinating and stabilizing Sirt3. Redox Biol. 62, 102707 (2023).
Shurer, C. R. et al. Physical principles of membrane shape regulation by the glycocalyx. Cell 177, 1757–1770.e1721 (2019).
Lin, C. H., MacGurn, J. A., Chu, T., Stefan, C. J. & Emr, S. D. Arrestin-related ubiquitin-ligase adaptors regulate endocytosis and protein turnover at the cell surface. Cell 135, 714–725 (2008).
Shao, G. et al. The E3 ubiquitin ligase NEDD4 mediates cell migration signaling of EGFR in lung cancer cells. Mol. Cancer 17, 24 (2018).
Staub, O. et al. Regulation of stability and function of the epithelial Na+ channel (ENaC) by ubiquitination. Embo J. 16, 6325–6336 (1997).
Baldus, S. E. et al. MUC1 and nuclear beta-catenin are coexpressed at the invasion front of colorectal carcinomas and are both correlated with tumor prognosis. Clin. Cancer Res 10, 2790–2796 (2004).
Bozkaya, G. et al. Cooperative interaction of MUC1 with the HGF/c-Met pathway during hepatocarcinogenesis. Mol. Cancer 11, 64 (2012).
Huang, L. et al. MUC1 oncoprotein blocks glycogen synthase kinase 3beta-mediated phosphorylation and degradation of beta-catenin. Cancer Res. 65, 10413–10422 (2005).
Grimson, M. J. et al. Adherens junctions and beta-catenin-mediated cell signalling in a non-metazoan organism. Nature 408, 727–731 (2000).
Spadoni, I. et al. A gut-vascular barrier controls the systemic dissemination of bacteria. Science 350, 830–834 (2015).
van der Post, S. et al. Structural weakening of the colonic mucus barrier is an early event in ulcerative colitis pathogenesis. Gut 68, 2142–2151 (2019).
Lu, C. H. et al. Membrane curvature regulates the spatial distribution of bulky glycoproteins. Nat. Commun. 13, 3093 (2022).
Desantis, S., Mastrodonato, M., Accogli, G., Rossi, G. & Crovace, A. M. Effects of a probiotic on the morphology and mucin composition of pig intestine. Histol. Histopathol. 34, 1037–1050 (2019).
Crouch, L. I. et al. Prominent members of the human gut microbiota express endo-acting O-glycanases to initiate mucin breakdown. Nat. Commun. 11, 4017 (2020).
Da Silva, S. et al. Stress disrupts intestinal mucus barrier in rats via mucin O-glycosylation shift: prevention by a probiotic treatment. Am. J. Physiol. Gastrointest. Liver Physiol. 307, G420–G429 (2014).
Kudelka, M. R., Stowell, S. R., Cummings, R. D. & Neish, A. S. Intestinal epithelial glycosylation in homeostasis and gut microbiota interactions in IBD. Nat. Rev. Gastroenterol. Hepatol. 17, 597–617 (2020).
Reily, C., Stewart, T. J., Renfrow, M. B. & Novak, J. Glycosylation in health and disease. Nat. Rev. Nephrol. 15, 346–366 (2019).
Westreich, S. T. et al. Fecal metatranscriptomics of macaques with idiopathic chronic diarrhea reveals altered mucin degradation and fucose utilization. Microbiome 7, 41 (2019).
Birchenough, G., Schroeder, B. O., Bäckhed, F. & Hansson, G. C. Dietary destabilisation of the balance between the microbiota and the colonic mucus barrier. Gut Microbes 10, 246–250 (2019).
Schroeder, B. O. et al. Bifidobacteria or fiber protects against diet-induced microbiota-mediated colonic mucus deterioration. Cell Host Microbe 23, 27–40.e27 (2018).
Staub, O. et al. Regulation of the epithelial Na+ channel by Nedd4 and ubiquitination. Kidney Int 57, 809–815 (2000).
Bae, S. J. et al. NEDD4 controls intestinal stem cell homeostasis by regulating the Hippo signalling pathway. Nat. Commun. 6, 6314 (2015).
Novellasdemunt, L. et al. NEDD4 and NEDD4L regulate Wnt signalling and intestinal stem cell priming by degrading the LGR5 receptor. Embo J. 39, e102771 (2020).
Léon, S., Erpapazoglou, Z. & Haguenauer-Tsapis, R. Ear1p and Ssh4p are new adaptors of the ubiquitin ligase Rsp5p for cargo ubiquitylation and sorting at multivesicular bodies. Mol. Biol. Cell 19, 2379–2388 (2008).
Ren, J., Kee, Y., Huibregtse, J. M. & Piper, R. C. Hse1, a component of the yeast Hrs-STAM ubiquitin-sorting complex, associates with ubiquitin peptidases and a ligase to control sorting efficiency into multivesicular bodies. Mol. Biol. Cell 18, 324–335 (2007).
Fang, D. F. et al. NEDD4 ubiquitinates TRAF3 to promote CD40-mediated AKT activation. Nat. Commun. 5, 4513 (2014).
Yang, Y. et al. Nedd4 ubiquitylates VDAC2/3 to suppress erastin-induced ferroptosis in melanoma. Nat. Commun. 11, 433 (2020).
Maspero, E. et al. Structure of a ubiquitin-loaded HECT ligase reveals the molecular basis for catalytic priming. Nat. Struct. Mol. Biol. 20, 696–701 (2013).
Hansson, G. C. & Johansson, M. E. The inner of the two Muc2 mucin-dependent mucus layers in the colon is devoid of bacteria. Gut Microbes 1, 51–54 (2010).
Li, H. et al. The outer mucus layer hosts a distinct intestinal microbial niche. Nat. Commun. 6, 8292 (2015).
Sheng, Y. H. et al. The MUC13 cell-surface mucin protects against intestinal inflammation by inhibiting epithelial cell apoptosis. Gut 60, 1661–1670 (2011).
Desai, M. S. et al. A dietary fiber-deprived gut microbiota degrades the colonic mucus barrier and enhances pathogen susceptibility. Cell 167, 1339–1353.e1321 (2016).
Hartmann, P. et al. Deficiency of intestinal mucin-2 ameliorates experimental alcoholic liver disease in mice. Hepatology 58, 108–119 (2013).
Landy, J. et al. Tight junctions in inflammatory bowel diseases and inflammatory bowel disease-associated colorectal cancer. World J. Gastroenterol. 22, 3117–3126 (2016).
Zhu, L. et al. Claudin family participates in the pathogenesis of inflammatory bowel diseases and colitis-associated colorectal cancer. Front Immunol. 10, 1441 (2019).
Bönnemann, C. G. et al. Dystrophin immunity after gene therapy for Duchenne’s muscular dystrophy. N. Engl. J. Med 388, 2294–2296 (2023).
Schambach, A. et al. A new age of precision gene therapy. Lancet 403, 568–582 (2024).
Pullan, R. D. et al. Thickness of adherent mucus gel on colonic mucosa in humans and its relevance to colitis. Gut 35, 353–359 (1994).
Kleiner, D. E. et al. Design and validation of a histological scoring system for nonalcoholic fatty liver disease. Hepatology 41, 1313–1321 (2005).
Folch, J., Lees, M. & Sloane Stanley, G. H. A simple method for the isolation and purification of total lipides from animal tissues. J. Biol. Chem. 226, 497–509 (1957).
Yang, H. et al. A common antimicrobial additive increases colonic inflammation and colitis-associated colon tumorigenesis in mice. Sci. Transl. Med. 10, eaan4116 (2018).
Acknowledgments
This study was supported by the National High-Level Hospital Clinical Research Funding (2023-NHLHCRF-DJZD-01 and 2024-NHLHCRF-JBGS-WZ-08). National Natural Science Foundation of China (82170817 and 81970713), Beijing Municipal Natural Science Foundation of China (7222160). Beijing Research Ward Construction Clinical Research Project (2022-YJXBF-04-02).
Author information
Authors and Affiliations
Contributions
Liang Peng and Zecheng Li conceptualized and designed the study. Zecheng Li, Runchuan Gu, Donghai Liu, Yijia Zhang, Yijin Qin, Xinyu Li, Yuan Qiao, Ziyao Li, and Jing Guo performed the experiments and analysed the data. Jun Wang, Hua Meng, and Liang Peng supervised the study. Zecheng Li, Runchuan Gu, Donghai Liu, and Liang Peng wrote the manuscript with input from all the authors. All the authors edited the manuscript and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Li, Z., Gu, R., Liu, D. et al. Lack of intestinal Mucin-1 impairs intestinal epithelial barrier and promotes metabolic dysfunction-associated steatotic liver disease in male mice. Nat Commun 17, 321 (2026). https://doi.org/10.1038/s41467-025-67034-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-67034-7