Abstract
Non-parasite genome encoded RNAs (ngRNAs) are often overlooked in their roles, especially their functions in shaping the biology of organisms. Here we show four non-parasite genome encoded virus-like RNAs that present in Schistosoma japonicum, a human blood fluke responsible for significant morbidity and mortality. These ngRNAs are predominantly localized in the germ cells that play crucial roles in oogenesis and egg production, pivotal processes contributing to host pathology. The ngRNA encode RNA-dependent RNA polymerase (RdRp) that synthesize complementary RNA strands, governing embryo viability and egg production. Notably, similar virus-like ngRNAs are also found in the planarian Dugesia japonica, where RdRp sustains worm survival. In this work, we integrate metatranscriptomics, single-cell RNA sequencing, RNA interference and various molecular methods to comprehensively identify ngRNAs and then characterize their functions. We further demonstrate the mechanism by which ngRNAs shape critical biological traits in parasitic helminths, providing novel insights into potential therapeutic targets for schistosomiasis control and advancing our understanding of ngRNA functions in certain organisms.
Similar content being viewed by others
Introduction
Schistosomiasis is a zoonotic disease caused by schistosome parasites, a serious zoonotic disease with global distribution1. Currently, the primary treatment for schistosomiasis relies heavily on a single drug, praziquantel. However, the long-term usage of a single-drug strategy raises concerns about the development of drug resistance, necessitating further research into alternative treatment approaches. Deeply investigating biological characterizations of schistosomes and identifying novel and effective drug targets is essential. Previous studies among unicellular protozoan parasites such as Trichomonas vaginali2,3, Giardia lamblia4, Leptomonas spp5,6, and Leishmania spp7,8,9. have demonstrated that these organisms are associated with certain viruses and/or virus-like RNA segments. More importantly, these exogenous RNAs play critical roles in the pathogenicity of pathogens and the immune responses of infected hosts10,11,12,13, suggesting that the targeting of functional virus-like RNAs could become an effective strategy against certain diseases.
In this work, we find four non-parasite genome encoded virus-like RNAs (ngRNAs) associated with Schistosoma japonicum. Through in-depth analyzes employing in situ hybridization and single-cell RNA sequencing (scRNA-seq), we demonstrate that these ngRNAs are predominantly localized in the germ cells. Functional analyses reveal that the ngRNAs play crucial roles in oogenesis and egg production, which are pivotal processes contributing to host pathology and disease dissemination. Additionally, we find that ngRNAs encoding RNA-dependent RNA polymerases (RdRp) can synthesize complementary ngRNAs, which are involved in egg production and embryo viability. Similar virus-like ngRNAs are also found in the planarian Dugesia japonica, where RdRp ngRNA contributes to worm survival. Our results indicate the key biological roles of these ngRNAs in certain organisms, which may represent unique targets for controlling certain diseases caused by some pathogens.
Results
Identification of ngRNAs in S. japonicum
We conducted high-throughput sequencing to analyze the libraries generated from total RNA isolated from S. japonicum (Fig. 1a). Briefly, the parasites were collected from the rabbits infected with S. japonicum cercariae at 16-, 22-, and 28-days post-infection (dpi), respectively. Then, the parasites at different time points were subject to independently lysis and treated with DNase and RNase. Subsequently, total RNAs were isolated, and libraries were prepared for sequencing. After removing reads mapped to S. japonicum genomes (NCBI accession: GCA_006368765.1 and GCA_021461655.1) as well as the rabbit host, assembly of the remaining reads yielded 8355, 7246, 7069 contigs, respectively (Fig. 1b). BLASTx analyses indicated that the majority of these assemble sequences are related to four contigs (Fig. 1c). Using the rapid amplification of cDNA end (RACE) technique, we successfully amplified full length sequences for each contig (Supplementary Fig. 1). Further analyses revealed that two long contigs exhibited high similarity to the RNA-dependent RNA polymerase protein (RdRp) of Bunyaviricetes, while two shorter contigs showed high homology to the nucleocapsid protein of Phenuiviridae in Bunyaviricetes. These findings led us to designate these four contigs as rp1/2 and np1/2.
a Scheme illustrating the metatranscriptomics analyses. b Metatranscriptomics analyses of parasites collected from infected rabbits at different stages. “Rabbits” represent reads mapped to the rabbit genome; “S. japonicum” represents reads mapped to the S. japonicum genome; “ngRNAs” represent reads matched to rp1/2 and np1/2; “Unknown” represents reads unmatched to the above genomes and rp1/2 and np1/2. c Proportion of four ngRNAs in reads. R1, R2 and R3 represent RNA samples isolated at 16 dpi, 22 dpi, 28 dpi, respectively. d Metatranscriptomic analyses of S. japonicum collected from infected mice. e Metatranscriptomic analyses of lysates prepared from independently collected S. japonicum. R1 and R2 represent two biological replicates. f PCR analyses of amplification of ngRNAs from total RNAs isolated from Oncomelania hupensis and definitive host livers. Numbers in left side indicate the expective base pair (bp) of PCR products. This is a representative result from three independent experiments. g Bioinformatic analyses of the abundance of rp1/2 and np1/2 sequences from the NCBI datasets. h PCR analyses of amplification of the four ngRNAs using cDNA from S. japonicum and parasite genomic DNA with different treatments. “Total RNA” represents the template of cDNA from S. japonicum total RNA; “Total RNA+RNase” represents the template of cDNA from S. japonicum total RNA treated with RNase; “Total RNA+DNase” represents the template of cDNA from S. japonicum total RNA treated with DNase; “Genomic DNA” represents the template of S. japonicum genomic DNA; “Genomic DNA+RNase” represents the template of S. japonicum genomic DNA treated with RNase; “Genomic DNA+DNase” represents the template of S. japonicum genomic DNA treated with DNase; “Blank” represents PCR reaction without templates genomic material. Numbers in left side indicate the expective base pair (bp) of PCR products. This is a representative result from three independent experiments. Bp means base pair. i Phylogenetic analyses of S. japonicum RP1/2. j Phylogenetic analyses of S. japonicum NP1/2. k Schematic diagram showing the motif and domain in RP1/2 and NP1/2. Source data are provided as a Source data file.
To exclude the possibility these RNAs being detected under opportunistic conditions, we shifted our experimental model from rabbits to mice. Mice were infected with S. japonicum cercariae, and the resulting parasites were collected at 16, 22, and 28 dpi, respectively. These samples were pooled for metatranscriptomic analyses. The results confirmed the consistent presence of these RNAs in the S. japonicum-infected mouse samples (Fig. 1d). Additionally, direct metatranscriptomic analyses of the parasite lysates further corroborated their presence in schistosomes (Fig. 1e). Furthermore, RT-PCR analyses indicated that these RNAs were absent in the livers of uninfected mice and intermediate host Oncomelania hupensis lacking S. japonicum infection (Fig. 1f). To further validate our findings, we conducted a comprehensive search of Schistosoma RNA-seq data deposited in the NCBI database. Among 177 available datasets for various species and geographic regions of Schistosoma, the results indicated that these RNAs are likely specific for S. japonicum (Fig. 1g and Supplementary Fig. 2) (Supplementary data 1). Despite efforts to expand our understanding by exploring potential presence in other Schistosoma species like S. mansoni and S. haematobium, the rules governing biological safety and species availability leave uncertainty regarding the presence of similar RNA in these genera.
Through bioinformatical analyses of metatranscriptomic results (Fig. 1b, d), we removed reads mapped to both S. japonicum and host genomes. The remaining contigs were not theoretically encoded by the parasitic genome, as confirmed by BLAST analyses showing no coding regions for these RNAs in the S. japonicum genome. To further validate the results, we used genomic DNA or cDNA synthesized from reverse transcript of total RNA (both treated and untreated with DNase or RNase) as templates for PCR amplification and also followed sequencing confirmation. These results indicated that these RNAs were not encoded by the S. japonicum genome (Fig. 1h). Therefore, we named them as non-parasite genome encoded RNAs (ngRNAs). Phylogenetic analyses revealed that either RP1 and RP2 or NP1 and NP2 can cluster together, belonging to the Phenuiviridae family (Fig. 1i, j). Generally, they also showed to be closely related to Trichobi phenuili virus (Fig. 1i, j). Protein domain and motif analyses indicated that RP1/2 contained an RNA-dependent RNA polymerase motif related to Bunyaviricetes and NP1/2 contained a Phenuiviridae nucleocapsid protein motif (Fig. 1k). Additionally, the NP1 shows 51.02% identity to NP2 while RP1 displayed 57.84% identity to RP2 (Supplementary Fig. 3).
ngRNAs are enriched in the sexual organs of S. japonicum
Schistosomes undergo multiple developmental stages characterized by distinct morphology. We further evaluated the abundance of these ngRNAs across several key developmental stages, including eggs, schistosomula (7 d, 14 d), and adult worms (21 d, 28 d). The results indicated that the ngRNAs were present in all investigated stages, with the highest abundance in eggs and adult worms (Fig. 2a). This finding aligns with analyses of previous RNA-seq datasets from different development stages14, which confirmed the consistently high abundance of ngRNAs in adult worms (Supplementary Fig. 4a). Given the crucial role of eggs produced by adult worms in disease transmission and host pathology, we conducted Whole mount in situ hybridization (WISH) to localize these ngRNAs within adult females (AF) and males (AM). Since schistosome is diecious, the distribution patterns were analyzed separately for males and females (Fig. 2b). Our findings revealed that the majority of detected ngRNAs were localized in the gonads, including ovaries and testes (Fig. 2b). RT-qPCR analyses of the isolated testes and ovaries from adult worms exhibited consistent results (Fig. 2c).
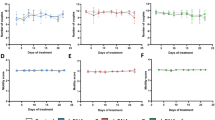
a Expression profiles of four ngRNAs in different stages of schistosomes. Data represent average results derived from three independent experiments with standard deviation calculated from three biological replicates. Each data point represents an individual experiment. b In situ hybridization analyses of the localization of four ngRNAs. An irrelevant RNA probe was used as a control. The number in left lower corner of each picture indicated the number of parasites with similar staining. Scale Bars indicate a length of 50 μm. c RT-qPCR analysis of the abundance of these four ngRNAs in S. japonicum ovaries and testes. Total RNAs isolated from adult females (AF), adult males (AM), ovaries and testes were equalized in RNA amount before reverse transcription. qPCR was performed to determine RNA abundance. Data illustrate representative results and show the mean and standard deviation from three biological replicates. Each data point represents an individual experiment. Statistical significance between two groups (AM vs. Testis and AF vs. Ovary) was determined using an unpaired, two-sided Student’s t-test. d Schematic diagram showing scRNA-seq analyses of cell populations containing rp1/2 and np1/2 in S. japonicum. The image was Created in BioRender (Cheng, G. (2025); https://BioRender.com/rt8qooa). e UMAP plots of the expression profiles of rp1/2 and np1/2 in different cell clusters of adult S. japonicum. f Proportions of different cell clusters expressing rp1/2 and np1/2 in AM and AF. Source data are provided as a Source data file.
To further characterize ngRNA roles, we performed scRNA-seq on males and females (16 and 26 dpi). Since S. japonicum genome lacks the coding regions for these ngRNAs, we manually supplemented the genome with the corresponding ngRNA sequences prior to scRNA-seq analyses. After removing potential doublets and low-quality cells, we obtained 12,076 and 8964 cell populations for AM and AF, respectively. Unsupervised clustering of all integrated datasets resulted in a total of 47 molecularly distinct cell populations (Fig. 2d and Supplementary Fig. 4b, c). This included clusters enriched with specific developmental cell types, such as rp1/2 and np1/2, which showed significant abundance in mature oocytes (female late stage), GSC progeny, neoblast, S1, S1 progeny, and vitellocyte cells in females; and spermatocyte (male late stage), neoblast, GSC, GSC progeny, and tegument progenitors in males (Fig. 2e, f).
ngRNAs are essential to maintain ovarian development and egg production
Interestingly, these ngRNAs were predominantly localized within the gonads of schistosomes, particularly in AF that contribute to egg production. Consequently, we mainly focused on AF in the following study. Next, we employed RNA interference (RNAi) to investigate their functional roles. Optimal siRNA was carefully screened and optimized to achieve maximal knockdown efficiency (Fig. 3a and Supplementary Fig. 5). Following siRNA treatment, significant reductions of ngRNAs were observed (Fig. 3b) alongside severe morphological defects of oocytes and impaired oogenesis (Fig. 3c). Morphological defects were further corroborated by Z-stack scanning, which vividly demonstrated ovarian abnormalities in treated individuals (Supplementary Movie 1–5).
a Scheme of functional analyses of ngRNAs in adult S. japonicum. The image was Created in BioRender (Cheng, G. (2025); https://BioRender.com/jgeel4j). b RT-qPCR analysis of the abundance of ngRNAs in the inhibited worms. Data illustrate representative results indicating mean and standard deviation from an experiment conducted in triplicate. c Confocal microscopy analyses of morphological changes of ovaries treated with siRNAs targeting ngRNAs. io immature oocyte, mo mature oocyte, v vacuole. Data illustrate representative results from three biological replicates. The number in the upper right corner of each image indicates the number of females with similar morphological features. Bars indicate 20 μm. d Effects of ngRNA inhibitions on egg production in vitro cultured parasites. Data illustrate representative results and show the mean and standard deviation from three biological replicates, with each data point representing an individual experiment. e RT-qPCR analyses of ngRNA abundance in eggs produced from inhibited worms. Data illustrate representative results and show the mean and standard deviation from an experiment conducted in triplicate. f Effects of ngRNA inhibition on egg viability in schistosomes as determined by acridine orange staining. Green straining (red arrow) indicates live eggs; red straining (white arrow) indicates dead eggs. Bars indicate 50 μm. g Statistical analysis of egg viability in ngRNA suppressed schistosomes. Data illustrate representative results and show the mean and standard deviation from three biological replicates. h Effects of RdRp inhibitor on egg production in treated parasites. Data illustrate representative results and show the mean and standard deviation from three biological replicates. Each data point represents an individual experiment. i Analyses of egg cell viability in parasites treated with Favipiravir. Data illustrate representative results and show the mean and standard deviation from three biological replicates. For b, d, e, g–i statistical significance between two groups (control vs. target siRNA/mock vs. drug treatment) was determined using an unpaired, two-sided Student’s t-test. Source data are provided as a Source data file.
To assess the impact on egg production, we monitored egg production at various time points of post-siRNA treatments (24, 48, 72, and 96 h). The results indicated a significantly decreased number of egg production following ngRNA suppression, especially for rp1 ngRNA (Fig. 3d). Subsequent RT-qPCR analysis of the eggs also found significantly reduced ngRNA for rp1/2 and np1 (Fig. 3e). Notably, acridine orange staining revealed substantial reductions in egg viability following ngRNA suppression (Fig. 3f, g). The finding was further validated through cell viability assays using dissociated eggs (Fig. 3g). To further corroborate the results, we conducted in silico docking studies of rp1/2 encoding RdRp proteins from commercially available inhibitors. Among these, Favipiravir, an antiviral agent targeting influenza viruses, demonstrated potential inhibition of activities towards RP1/2 RdRp protein (Supplementary Fig. 6). In vivo validation showed that treatment with Favipiravir significantly reduced both egg production and the viability of eggs in the worms compared to untreated controls (Fig. 3h, i).
ngRNA encoding proteins interacting S. japonicum proteins regulate egg production
To investigate the regulatory roles of ngRNAs in egg production, we employed EdU staining to assess cell proliferation in ovaries and Fast Blue BB staining to evaluate vitellaria morphology following rp1 and np1 suppression. The results revealed that suppression of rp1 and np1 led to decreased cell proliferation in ovaries (Fig. 4a), accompanied by reduced Fast Blue BB staining intensity in the vitellarium, suggesting a potential role for these ngRNAs in governing cell proliferation in the gonad. RNA-seq analysis of rp1- and np1-inhibited parasites revealed 56 and 27 down-regulated genes, respectively, of which 5 genes were shown in both (Fig. 4b, c and Supplementary data 2–3), including tyrosinase (EWB00_006115), a known regulator of egg production15. These findings were further corroborated by scRNA-seq data, which demonstrated co-expression of these target genes with RP1-encoded RdRp in specific cell clusters such as S1/S1 progeny, vitellocytes, and female late-stage cells (Fig. 4c).
a EdU staining analyses of cell proliferation from ovaries and Fast Blue BB staining analyses of the vitellaria from rp1 and np1 suppressed parasites at 4 days and 6 days post-treatment. The numbers in the corner indicate the fraction of similar morphological features. Scale bars: 50 μm for ovarian analyses and 100 μm for Fast Blue BB analyses. Arrows in the lower left corner indicate the anterior part of worms. Data are the representative results from two biological replicates. b Volcano plots indicating RNA-seq analyses of differentially expressed genes in rp1 and np1 suppressed worms. c scRNA-seq data analysis of the down regulated genes associated cell clusters. d Identification of RP1 RdRp interacting proteins by Yeast two-hybridization. P1 means the plates containing SD/-leu/-Trp medium; P2 means the plates containing Sd/-Leu/-Trp/-His/-Ade/X-alpha-gal/Aba. e scRNA-seq data analyses of co-expression of RP1 RdRp and its interactors in different cell clusters. f Protein–protein interaction map of RP1-mediated protein interactions identified by yeast two-hybridization analysis and RNA-seq down-regulation of RP1 target genes. STRING analysis of the proteins identified from Yeast two-hybridization combined with the down-regulated proteins from RNA-seq analysis. g RT-qPCR analyses of the transcript levels of Lin-10 and Lp1 in the inhibited worms. Data illustrate representative results indicating mean and standard deviation from an experiment conducted in triplicate. h Fast Blue BB staining analyses of the vitellaria from Lin-10 and Lp1 suppressed parasites at 6 days post-treatment. The numbers in the upper corner indicate the fraction of similar morphological features. Scale bar = 100 μm. i Effects of Lin-10 and Lp1 inhibitions on egg production in cultured parasites. The numbers of eggs produced at 24, 48, and 72 h after dsRNA treatment was counted under microscopy. Data illustrate representative results and show the mean and standard deviation from three biological replicates. For g, i statistical significance between two groups was determined using an unpaired, two-sided Student’s t-test. Source data are provided as a Source data file.
To elucidate the molecular basis underlying these regulatory effects, we utilized the Yeast two-hybrid method to identify the interaction partners of RP1 RdRp. The result indicated RP1 may interact with ribosomal protein LP1 (EWB00_007383), Lin-10 family protein (EWB00_001962), proteasome beta subunit, Peptidase C1A subfamily and Ribosomal protein S5a (Fig. 4 d and Supplementary Fig. 7). Notably, LP1 and Lin-10 have been previously implicated in regulating development in schistosomes and Caenorhabditis elegans16,17. In addition, ribosomal protein RPLP1 is a known host factor in flavivirus infection18. Single cell RNA-seq results further supported the co-expressions of these interactors with RP1 in specific cell clusters, such as vitellocytes and female late-stage cells (Fig. 4e). To integrate these findings into a coherent molecular framework, we constructed an interaction map based on the STRING database and our Yeast two-hybrid results (Fig. 4 f). This map revealed protein–protein interactions mediated by RP1, including tyrosinase, RpS5a, which are known to play roles in egg production within schistosomes15 and Drosophila melanogaster19. Next, we verified the functional roles of these identified interactors using RNAi. The results indicated that suppression of Lin-10 and Lp-1 (Fig. 4g) led to abnormal morphology of vitellocytes (Fig. 4h) and reduced number of egg production (Fig. 4i). Collectively, these results indicated the molecular mechanisms by which RP1 RdRp regulates egg production and ovarian development.
ngRNA encoding RdRp involves the synthesis of complementary ngRNAs
Since these ngRNAs are consistently present in multiple stages of schistosomes, we want to understand the mechanisms of ngRNA replication during cell division. RdRp, an RNA-dependent RNA polymerase found in viral replication machinery, is implied to play a role in the replication of ngRNAs in schistosomes20. To validate this hypothesis, we designed the experiments to determine whether RdRp can generate complementary (positive-sense) RNA and double-stranded RNAs (dsRNAs), which are essential for producing progeny RNA copies of these ngRNAs. We used a strand-specific RT-PCR approach to specifically detect the complementary RNAs (Supplementary Table 1). The PCR results confirmed the presence of complementary RNAs for these ngRNAs (Fig. 5a), suggesting RdRp is capable of synthesizing the complementary strand necessary for ngRNA replication.
a PCR analyses of replication intermediate RNAs for ngRNAs. PCR analysis of replicative intermediates of the ngRNAs in schistosomes. Total RNA was reverse transcribed using specific primers, and PCR products targeting complementary ngRNAs were amplified and analyzed by agarose gel electrophoresis (dotted boxes indicate expected products). Lane 1: cDNA transcribed using specific primer as template for PCR; Lane 2: cDNA transcribed using random primers as template for PCR; Lane 3: Total RNA without a reverse transcriptase as template for PCR; Lane 4: no template. Data illustrate representative results from three independent experiments. b Effect of rp1 inhibition on the expressions of other ngRNAs. Data illustrate representative results from three biological replicates. The results show the mean and standard deviation from an experiment conducted in triplicate. c Analyses of luciferase activities in the cells co-transfected rp1 RdRp plasmid and reversed complementary Gaussia luciferase reporter plasmid or co-added Favipiravir inhibitor in different concentrations. The transfected cells were assayed for luciferase activities after 36–48 h of post-transfection. Data illustrate representative results and show the mean and standard deviation from three biological replicates. d Analyses of the transcript lever of np1 in the cells co-transfected rp1 RdRp plasmid and reversed complementary (RC) np1 plasmids or co-added Favipiravir inhibitor. The transfected cells were subject to RNA isolation after 36–48 h of post-transfection for RT-qPCR analyses. Data illustrate representative results and show the mean and standard deviation from three replicates. e Dot blot analysis of dsRNA abundance in rp1 suppressed parasites. Data illustrate representative results from two independent experiments. f Immunofluorescence analysis of dsRNA expressions in rp1 suppressed parasites. Data illustrate representative results from three independent replicates. g Percentage of cells co-expressing rp1/2 and np1/2 from scRNA-seq data. For b–d statistical significance between two groups was determined using an unpaired, two-sided Student’s t-test. Source data are provided as a Source data file.
To further assess the functional role of RP1 RdRp in ngRNA replication, we suppressed rp1 by siRNA and observed significantly reduced levels of other ngRNAs (Fig. 5b) (Supplementary data 2–3), indicating that the RdRp is critical for maintaining the abundance of ngRNAs. Next, we determined whether that RP1 RdRp is responsible of ngRNA replication and its activity can be inhibited by Favipiravir. Upon co-transfection of recombinant plasmids for expressing RP1 RdRp domain and reversed complementary Gaussia luciferase reporter gene embedding into 5′UTR and 3′UTR of rp1 or reversed complementary np1 into 293 T cells, luciferase assay and RT-qPCR analyses indicated that RP1 RdRp can drive the expression of either Gaussia reporter gene or np1 gene (Fig. 5c, d). When applying Favipiravir in transfected cells, their expressions were significantly suppressed (Fig. 5c, d). Recognizing that dsRNA often serves as an intermediate product during RNA replication, we further performed J2 antibody-based Dot blot analyses to detect its presence. The results indicated that S. japonicum not only contains dsRNA but also shows a reduction in dsRNA levels upon suppression of Rp1 RdRp (Fig. 5e). Collectively, these findings suggest that RP1 RdRp has activity involving ngRNA replication. To ensure that our observations were biologically relevant, we performed Whole-mount immunofluorescence using J2 antibodies in the ovary of female worms. Control animals exhibited robust signals for dsRNA (Fig. 5f), while rp1 RdRp suppressed worms showed undetectable signal intensity, further supporting the important role of RP1 RdRp in maintaining ovarian development. Moreover, scRNA-seq data revealed that rp1/2 and np1/2 are primarily co-expressed in germline cells and vitellocyte (Fig. 5g). These results suggest that RdRp-mediated replication of these ngRNAs is highly active in cell clusters related to egg production, aligning with our experimental findings.
Physical characterization of ngRNA formation in S. japonicum
Since the ngRNAs can be successfully isolated even after DNase and RNase treatment, we hypothesized that these ngRNAs may be physically protected from ribozyme degradation in a micro-sized particle encapsulated form. To this end, we used sucrose density gradient centrifugation to separate the components of worm lysates. The most abundant fraction was collected for RNA isolation and quantification by RT-qPCR, indicating. Electron microscopy (EM) analysis of the fraction with the highest abundance of ngRNAs revealed significant enrichment of micro-sized particles (Fig. 6a), suggesting that these ngRNAs may be present in a particle-like form.
a Transmission electron microscopy (TEM) image showing micro-sized particles separated by sucrose gradient centrifugation from S. japonicum lysates. Arrowheads indicate the presence of micro-sized particles. b TEM analysis of ultrathin sections around S. japonicum ovaries at magnification 49,000×. Arrowheads point to potential micro-sized particles associated with ngRNA formation. c Immune transmission electron microscopy (Immune-TEM) results showing the localization of ngRNA encoding the NP1 protein. Scale bars = 100 nm. d and e are magnified images from the Immune-TEM analysis, with scale bars indicating 100 nm in length. All of the data illustrate representative results from two independent experiments.
To corroborate this result, transmission electron microscopy (TEM) was employed to examine thin sections from the schistosome ovaries. Given the high enrichment of ngRNAs in this tissue region, TEM analysis revealed that the micro-sized particles were indeed abundant in the ovary-associated areas (Fig. 6b). To identify whether these particles are associated with ngRNAs, we cloned the full-length open reading frame (ORF) encoding np1 into the pET-28 vector and expressed the recombinant protein. The purified protein was then used to generate a polyclonal antiserum (Supplementary Fig. 8). When applying the antibodies in an immune TEM assay, some of the micro-sized particles were clearly recognized (Fig. 6c–e), further confirming that these particles are indeed associated with ngRNAs.
Targeting rp1 ngRNA in animals infected with S. japonicum reduces egg production and host pathology
Given the absence of ngRNA coding genes in mammalian genomes21 and certain mammals can serve as definitive hosts for disease dissemination, targeting these ngRNAs may not only represent a unique strategy for disease control with minimal side effects, but also mitigate host pathology. Therefore, we administered rp1 siRNA into the mice infected with S. japonicum to access the effect of ngRNA inhibition on egg production and the pathology of mice livers (Fig. 7a). Upon five injections, at 28 days of post-infection, worms were perfused from the mice. There was a statistically significant reduction in worm recovery [control (RNAi) = 86% versus rp1 (RNAi) = 50%, P < 0.0001, Student’s t test] (Fig. 7b). Additionally, total RNA was extracted from the collected worms, and the transcript levels of rp1 were analyzed by RT-qPCR. The result indicated that the transcript level of rp1 was significantly decreased in remaining worms collected from the mice administered with rp1 siRNA (Fig. 7c). Moreover, mice receiving rp1 siRNA (RNAi) targeting parasites exhibited less eggs and non-significant granuloma in livers, in contrast to the abundance of egg-induced granulomata in livers of control mice (Fig. 7d, f). Subsequently, hatching experiments analyzing the eggs collected from mice livers revealed a significantly lower number of viable eggs [control (RNAi) = 267.8 versus rp1 (RNAi) = 37.17, P < 0.0001, Student’s t test] (Fig. 7e). Overall, these results clearly indicated that targeting rp1 ngRNA in infected animals could reduce egg production and egg viability as well as host pathology, implying the alleviation of disease dissemination.
a Schematic illustrating rp1 siRNA treatment in mice infected with S. japonicum cercariae. Each group contains six mice (n = 6). b Effect of rp1 siRNA treatment on worm burden in infected mice at 28 dpi. Data show the mean and standard deviation. n = 6. c RT-qPCR analyses of the abundance of rp1 in the remain worms collected from administrated mice. Parasites were subjected to RNA isolation and RT-qPCR was performed. Data illustrate representative results indicating mean and standard deviation from an experiment conducted in triplicate. d Effect of rp1 siRNA treatment on egg deposition in the liver of mice infected with S. japonicum cercariae. EPG eggs per gram. Data show the mean and standard deviation. n = 6. e Effect of rp1 siRNA treatment on egg hatching rates from livers of infected mice at 28 dpi. Eggs were isolated from the livers of mice infected with S. japonicum cercariae at 28 dpi. Data show the mean and standard deviation. n = 6. f Histopathological analysis of liver tissues from mice treated with PBS, control siRNA and rp1 siRNA. Arrows indicate egg-induced granuloma formation. Data show representative results from three independent experiments. Scale bar = 200 µm. For b–e statistical significance between two groups was determined using an unpaired, two-sided Student’s t-test. Source data are provided as a Source data file.
RdRp ngRNA is also present in planarian Dugesia japonica that contributes to worm survival
We investigated whether similar features of ngRNAs are also present in other flatworms, such as planarian D. japonica. Additionally, recent studies indicated that RNA viruses are present in Schmidtea mediterranea22,23. Through metatranscriptomic and bioinformatic analyses of RNA libraries from D. japonica, we identified that one ngRNA also encoding RdRp in D. japonica (Fig. 8a), named as rp1 RdRp. Similarly, rp1 ngRNA is not encoded by the D. japonica genome (Fig. 8b). Localization studies by WISH and FISH confirmed that the rp1 ngRNA is intracellularly localized within the worms (Fig. 8c, e). Furthermore, treatment of D. japonica with dsRNA targeting rp1 RdRp resulted in the reduced expression of target ngRNA (Fig. 8d) and also led to compromised epidermis integrity (Fig. 8f) and increased worm mortality (Fig. 8g), highlighting its critical role in maintaining worm survival.
a Metatranscriptomic analyses of ngRNA in Dugesia japonica. “ngRNAs” represent reads mapped to viral database; “D. japonica” represents reads mapped to the D. japonica genome; “Unknown” represents reads unmatched to the above genomes and ngRNAs. b PCR analysis of the RdRp ngRNA by inputting cDNA transcribed from total RNA or worm genomic DNA treated with DNase or RNase. “RNA” represents cDNA reversely transcribed from D. japonica total RNA as template; “RNA+DNase” represents cDNA reversely transcribed from D. japonica total RNA treated with DNase as template; “Genomic DNA” represents genomic DNA as template; “Genomic DNA+DNase” represents genomic DNA treated with DNase as template. This is a representative result from three independent experiments. The numbers in left side indicate the base pair (bp) of DNA marker. c WISH analyses demonstrating the localization of RdRp l in planarian. The number in right upper corner indicates worm counts with similar patterns. Scale bar = 500 μm. d RT-qPCR analyses of the abundance of RdRp in worms treated with RdRp dsRNA and control dsRNA. Data illustrate representative results indicating mean and standard deviation from an experiment conducted in triplicate. e Fluorescent In Situ Hybridization (FISH) analyses showing the localization of RdRp within individual worm. Scale bar = 500 μm. Data illustrate a representative result from two independent experiments. n = 10 for each treatment. f WISH analyses of RdRp abundance in dsRNA treated planarian. The number in right upper corner indicates worm counts with similar features. Scale bar = 500 μm. g Effect of RdRp ngRNA inhibition on worm survival. n = 30 for each treatment or control. Data illustrate representative results and show the mean and standard from three independent experiments. Statistical significance between two groups was determined using an unpaired, two-sided Student’s t-test. Source data are provided as a Source data file.
Discussion
In the present study, we discovered that four RNAs associated in S. japonicum are not encoded by the parasitic genome. These ngRNAs were found to be critically involved in key biological processes of schistosomes, such as germ cell proliferation, ovarian development, and egg production, all critical events related to the pathology in infected hosts and the following disease dissemination. Importantly, these ngRNAs are probably present across different geographical strains and host isolates, suggesting they are broadly relevant to schistosome populations.
A key feature of these ngRNAs is that they lack coding regions within the parasite genome, coupled with their expression across multiple developmental stages, suggesting that there is a unique mechanism for RNA replication. RdRp is one of the most versatile proteins capable of replicating RNA and performing transcription. Our findings reveal that the RdRp encoded by these ngRNAs both synthesizes their complementary RNAs and interacts with key S. japonicum proteins, including ribosomal proteins (LP1, S5a), Lin-10, and the proteasome beta subunit. By using RNA interference, luciferase reporter assay, and a chemical inhibitor, we further demonstrated Rp1 RdRp governs ngRNA replication by interacting with these S. japonicum proteins. More importantly, RdRp ngRNAs are predominantly localized to the germline cells of adult S. japonicum and are essential for ovarian development and egg production in female worms. Given the critical role of eggs in pathogenesis and disease transmission, targeting RdRp ngRNAs or their replication process represents a therapeutic strategy against schistosomiasis. Additionally, C. elegans encodes four distinct RdRps that generate secondary siRNAs24, including EGO-125, which is required for fertility and robust germline RNAi. Consequently, it remains to be determined whether exogenous RdRp can reprogram small RNA functions during germline development and oogenesis in S. japonicum.
Phylogenetic tree and protein motif analyses revealed that these ngRNAs are closely related to the Phenuiviridae family (class Bunyaviricetes). The Phenuiviridae family typically comprises two to eight single-stranded molecules of negative-sense or ambisense encoding RdRp, viral glycoproteins, nucleocapsid protein, and non-structural proteins26. Although we noted that the ngRNAs were significantly enriched, accounting for more than 60% of total reads, our subsequent bioinformatic analyses detected no sequences encoding viral glycoproteins. Additionally, we enriched micro-sized particles by sucrose gradient separation and found that the ngRNAs co-fractionated with the most abundant particle component. Furthermore, sequencing and bioinformatic analyses of the RNA from this component did not detect any segments encoding glycoproteins. While our study was in progress, Shi et al. performed deep transcriptome sequencing on various invertebrate species and reported the presence of similar RNA segments in schistosomes, which also lacked an associated glycoprotein RNA27. Collectively, our findings suggest that these ngRNAs may dispense with a canonical matrix protein for the maintenance of their genomic integrity.
Given the physical presence in micro-sized particles, probably contributing to ngRNA stability, we attempted to inoculate the most abundant fraction from the gradient into a panel of cell lines. These included mammalian lines (Vero, DH82, LX-2, HEK293T, PK-15, MDCK) and insect cell lines (Sf9, C6/36). The cells were then blindly passaged for 3–4 generations. However, RT-qPCR analyses revealed that the ngRNAs were not detectable in passaged cells. Furthermore, the lack of schistosome cell lines and robust genetic tools currently precludes direct evaluation of micro-sized particle replication within the parasite. Moreover, analysis of our small RNA sequencing datasets from extracellular vesicles28 confirmed the absence of significant homology to these ngRNAs. Future studies are needed to determine whether EV-associated longer RNAs contain these ngRNAs that could contribute to the RNA segment stability in parasites.
Phylogenetic analyses of the proteins encoding by these ngRNAs showed that they form a distinct clade. Notably, RP1 and RP2 are most closely related to T. phenuili virus 2 and Wufeng rodent phenuivirus 1, whereas NP1 and NP2 are more closely related to T. phenuili virus 1. The nucleoproteins associated with S. japonicum and Trichobilharzia regenti formed a distinct clade, separate from those of Opisthorchiidae liver flukes (Metorchis orientalis). Interestingly, RP1/2 is also closely related to Wufeng rodent phenuivirus 1, which was identified from rodent internal organs and fecal samples via meta-transcriptomic sequencing29. At present, we are not able to exclude the possibility of flatworm infection in the rodent. Nevertheless, these observations suggest that viral-like RNAs may be widespread among the flatworms. A study on the transmission of tapeworm (Schistocephalus solidus) viruses revealed that most viruses are vertically transmitted from parents to offspring30. This supports our inference that the ngRNAs may be consistently maintained through long-term co-evolution with parasites.
In mammals, cytoplasmic sensors in host cells detect double-stranded RNA (dsRNA), a common byproduct of viral replication. This recognition then triggers the interferon (IFN) system, a first line of defense that limits viral replication. However, how dsRNA triggers an immune-like response in flatworms and what regulatory mechanisms control these processes in helminths remain critical knowledge gaps. In C. elegans, Orsay virus (OrV) is the known natural virus affecting this worm. This nematode employs multiple, orchestrated defenses against OrV infection, including the RNAi pathway, viral RNA uridylation to direct RNA for degradation, and the expression of antiviral proteins such as those involved in ubiquitination31, Conversely, OrV replication also depends on host factors like zinc and lipid metabolism, highlighting a complex interplay between proviral and antiviral mechanisms that maintains dsRNA homeostasis. A recent study demonstrated adult B. malayi worms activate RNA interference as an innate immune response against viral replication32. Although RNAi is widely used to silence target genes in S. japonicum, it remains unknown whether endogenous RNAi is activated during ngRNA replication, particularly within the germline cells.
Taken together, we demonstrate that ngRNAs in S. japonicum are essential regulators of female reproduction—directly controlling ovarian development and egg production—and thereby govern important aspects of parasitic pathogenesis and transmission. The identification of analogous ngRNAs in planarians, which contribute to host survival, underscores that this class of RNA represents a fundamental and conserved regulatory mechanism shaping biology in diverse organisms.
Methods
Animal and parasites
BALB/c mice (6–8 weeks, male) and New Zealand Rabbits (male, three months) were purchased from the Shanghai Experimental Animal Centre of Chinese Academy of Sciences. Mice were housed under specific-pathogen-free, climate-controlled conditions on a 12-h light/dark cycle, with free access to standard chow and water. S. japonicum cercariae were provided by the National Institute of Parasitic Diseases of Chinese Center for Disease Control and Prevention, China. Each mouse and rabbit were percutaneously infected with approximately 200 and 2000 cercariae, respectively. All animal experiments were approved by the Institutional Animal Care and Use Committee (IACUC) of the Shanghai Veterinary Research Institute of Chinese Academy of Agricultural Sciences (Permit number: SV-20170929-01). Dugesia japonica (D. japonica) was obtained from Boshan fountain of China. Animals were reared in spring water at 20 ± 1 °C and fed raw chicken liver once or twice a week. Asexual planarians (a length between 2 and 4 mm) were starved for 1 week before any experiments.
Metatranscriptomics analyses
For metatranscriptomic analyses of parasites collected from rabbits, schistosomes [16, 22, 28 days post-infection (dpi), 1 g in weight for each stage] were homogenized with steel balls in 2 mL phosphate-buffered saline (PBS). The lysates were subjected to three freeze-thaw cycles on dry ice. After centrifugation (4 °C, 10 min, 15,000 × g), the supernatants were collected carefully. To remove cell-sized particles, 300 mL supernatant was filtered through a 0.45 μm filter (Millipore), of which the filtrates were digested using a mixture of DNase (Turbo DNase from ThermoFisher Scientific, Baseline-ZERO from Epicenter, and benzonase from Novagen) and RNase (ThermoFisher Scientific, USA) to remove free nucleic acid at 37 °C for 1.5 h33. Nucleic acids were isolated using the QIAamp RNA Mini Kit (Qiagen, Germany) according to the manufacturer’s protocols. Three libraries were generated using Nextera XT DNA Sample Preparation Kit (Illumina, USA), and then subjected to paired-end (PE) sequenced on the MiSeq Illumina platform (Majorbio, Shanghai)34. For metatranscriptomic analyses of parasites from mice, worms were collected from mice at 16 dpi, 22 dpi, and 26 dpi, respectively. The parasites were pooled and then subjected to metatranscriptomic analyses as described above.
For metatranscriptomic analyses of parasite lysates, adult schistosomes (28 dpi) were amputated and then centrifuged to remove the tissue and cell debris. The supernatants were subject to total RNA isolation for library preparation. The libraries were constructed using the Illumina TruSeq Stranded mRNA LT Sample Prep Kit (Illumina, USA). After screening by an Agilent Bioanalyzer 2100 (Agilent Technologies, USA), the selected libraries were quantified with a Promega QuantiFluor RNA System (Promega, USA), and the samples with a concentration >2 nM were further subjected to PE sequencing. Following serial dilution and mixing, the library was denatured into single strands for sequencing. The Illumina HiSeq X-Ten platform (Illumina, USA) was used for sequencing, along with the PE150 strategy.
Upon the identification of the species of planarian D. japonica by amplifying the Cox gene, worms were carefully washed using PBS (ThermoFisher Scientific) at least three times, and then were subject to total RNA isolation using TRIzol (ThermoFisher Scientific). The quality of isolated RNAs was evaluated by Agilent 2100. The libraries were prepared using the Illumina Stranded Total RNA Prep Kit (Illumina) and then sequenced as described above.
Bioinformatical analyses
Sequencing reads were first adapter- and quality-trimmed using the Trim-galore program (v0.6.10) (-phred33 -length 100 -stringency 3 -paired, https://zenodo.org/records/7598955). The remaining reads were mapped to the S. japonicum reference genomes (NCBI accession: GCA_006368765.1 and GCA_021461655.1), rabbit (NCBI accession: GCA_964237555.1), and mice (NCBI accession: GCF_000001635.27) genomes using the Bowtie2 program (v2.5.0)35. The remaining reads were assembled de novo using MEGAHIT (v1.2.9) with default parameters (-min-contig-len 200) to generate the longest contigs and singlets36. Next, the contigs and singlet reads were analyzed by DIAMOND (v2.1.12) with default parameters. Each sequence was aligned against the NCBI virus reference proteome (ftp://ftp.ncbi.nih.gov/refseq/release/viral/) in addition to viral protein sequences from the NCBI nr Fasta file (based on annotation taxonomy in Virus Kingdom)34,37, and an E-value threshold of <10⁻⁵ was set to screen potential viral sequences. To further eliminate potential false-positive viral sequences, the viral sequences obtained through the above steps were re-aligned against the NCBI Internal Non-Protein Non-Redundant (NVNR) Protein Database. As to planarian D. japonica, the remaining reads were mapped to the planarian reference genome (GCA_001938525.1), and the following analyses were performed for the assembly as described above.
As to metatranscriptomic analyses, the raw sequencing data were trimmed to remove adapters and filtered to remove low-quality reads. The mRNA transcripts were assembled de novo using Trinity (v2.5.1) software and then to obtain unigenes using CD-HIT software (version 4.7). The obtained sequences were then aligned with the bacterial, fungal, and viral sequences from the NCBI nr database using the BLASTX program. For specie annotation, the MEGAN software (v.6.13.1) was employed. QIIME (v.1.7.0) was used to calculate the relative abundance at each classification level. The unigene sets were also aligned with the KEGG and CAZy databases for functional annotation. The threshold value for gene identification was an e-value < 10−5.
The homologous reference sequences from five families within Bunyavirales were downloaded from the NCBI database to analyze phylogenetic relationships. Amino acid sequences of RP/NP were aligned using MUSCLE in MEGA-X38 and trimmed. IQ-TREE (v2.3.6)39 was used to construct phylogenetic trees with 1000 bootstrap replicates (-bb 1000) and the Model Finder function (-m MFP). Phylogenetic trees were visualized and edited using Interactive Tree Of Life (iTOL)40.
Genomic DNA isolation, total RNA isolation, DNase/RNase treatment, and PCR analysis
S. japonicum genomic DNA (adult worms, 28 dpi), murine liver genomic DNA, or planarian D. japonica genomic DNA was isolated using an AxyPrep Mutisource Genomic DNA Miniprep Kit (Axygen, USA) according to the manufacturer’s instructions. The isolated DNA (500 µg) was treated with DNase (1.5 µL) or RNase (1.5 µL) at 37 °C for 30 min. Then, ethylenediaminetetraacetic acid (EDTA) was added to a final concentration of 15 mM and heated at 75 °C for 10 min to inactivate DNase. Next, PCR was performed in a 20 μL reaction mixture containing 1 μL treated DNA, 10 μL 2× SYBR Primer Ex TaqII (TaKaRa, China), 8.5 μL H2O, and 0.5 μL primers (10 μM) using the following thermal cycling profile: 95 °C for 5 min, followed by 40 cycles of amplification (95 °C for 5 s, 57 °C for 30 s, 72 °C for 20 s). The primer sequences that synthesized by Invitrogen were listed in Supplementary Table 1. The PCR products were separated on 2% agarose gels and visualized by staining with Gold View (Yeasen, China).
Total RNAs from S. japonicum, O. hupensis, or murine livers were isolated using TRIzol (ThermoFisher Scientific) according to the manufacturer’s instructions. Isolated RNA (500 µg) was subject to treatment with DNase (1.5 µL) or RNase (1.5 µL) as described above. Treated RNA was further purified using phenol-chloroform extraction, and reverse transcription reactions were performed in a total volume of 10 μL, containing 6 μL of total RNA (100 ng), 2 μL of 5× Prime-Script Buffer (TaKaRa), 0.5 μL of PrimeScript RT Enzyme Mix I (TaKaRa), 1 μL of RNase inhibitor (TaKaRa), and 0.5 μL of random primers (10 μM) or specific strand primer (Supplementary Table 1). PCR and subsequent agarose gel analysis were performed as described above. Similar experiments were performed for planarian D. japonica using the following primers (forward: 5′CGGCACTAGTCAATCAACCCTCT3′, reverse: 5′TGTTTGGTTTACTTGTTGGCTCATTC3′, synthesized by Invitrogen)
5′/3′RACE
To obtain full-length ngRNA sequences, 5′- and 3′-RACE assays were conducted. Total RNA was extracted from adult S. japonicum worms using TRIzol reagent (ThermoFisher Scientific) following the manufacturer’s protocol. Gene-specific primers for 5′- and 3′-RACE were designed based on previously assembled ngRNA sequences, respectively (Supplementary Table 1). 5′- and 3′-RACE reactions were performed using the SMARTer RACE cDNA Amplification Kit (Clontech, USA) in accordance with the official instructions. For 5′-RACE outer PCR, the thermal program consisted the following steps: (i) an initial five cycles of 94 °C for 5 s, 65 °C for 30 s, and 72 °C for 3 min; (ii) followed by five cycles of 94 °C for 5 s, 60 °C for 10 s, and 72 °C for 2 min; then (iii) 30 cycles of 94 °C for 5 s, 55 °C for 10 s, and 72 °C for 2 min; and (iv) a final extension at 72 °C for 10 min. A diluted aliquot of the outer PCR product was subsequently used as a template for nested PCR, which was run under identical thermal conditions. For 3′-RACE outer PCR, the program was as follows: (i) amplification began with an initial denaturation at 94 °C for 3 min; (ii) followed by 20 cycles of 94 °C for 30 s (denaturation), 53 °C for 20 s (annealing), and iii) 72 °C for 3 min (extension), and a final extension at 72 °C for 10 min. Similarly, a diluted fraction of the 3′-RACE outer PCR product was subjected to nested PCR using the same conditions. All primers utilized in this study that synthesized by Invitrogen are listed in Supplementary Table 1. Nested PCR products from both 5′- and 3′-RACE reactions were analyzed by 1.2% agarose gel electrophoresis containing ethidium bromide (EtBr). Target bands were excised, purified, and subcloned into the pMD™19-T vector (TaKaRa) for Sanger sequencing. Full-length ngRNA sequences were reconstructed by in silico assembly of the 5′- and 3′-RACE-derived sequences.
Analyses of the abundance of ngRNAs in different stages of schistosomes
Total RNA was extracted from schistosomes at various developmental stages (eggs, 7, 14, 21, and 28 dpi) using TRIzol reagent (ThermoFisher Scientific) following the manufacturer’s protocol. RNA concentrations were quantified with a NanoDrop 1000 spectrophotometer (ThermoFisher Scientific) for RT-qPCR experiments. Expression levels of each RNA transcript across different stages were determined via RT-qPCR using specific primers (Supplementary Table 1), with the primers targeting S. japonicum nicotinamide adenine dinucleotide dehydrogenase (NADH) gene serving as an internal reference for data normalization. Briefly, RT was carried out in a 10 μL final volume comprising 6 μL of total RNA (400 ng), 2 μL of 5× PrimeScript Buffer (TaKaRa), 0.5 μL of PrimeScript RT Enzyme Mix I (TaKaRa), 1 μL of RNase inhibitor (TaKaRa), and 0.5 μL of 10 μM random primers. RT-qPCR was conducted using SYBR Premix Ex Taq (TaKaRa) in a 15 μL reaction mixture consisting of 1 μL cDNA, 7.5 μL of 2× SYBR Premix Ex TaqII (TaKaRa), 6 μL of nuclease-free H2O, and 0.5 μL of 10 μM primers. Amplification was performed on a Master Cycler Ep Realplex PCR system (Eppendorf, Germany) with the following thermal program: (i) an initial denaturation step at 95 °C for 5 min; (ii) followed by 40 cycles of 95 °C for 5 s, 60 °C for 30 s, and (iii) 72 °C for 20 s. Relative mRNA expression levels were calculated using the 2⁻ΔCt method41.
Gonads (testes and ovaries) from adult S. japonicum were dissected following established protocols42,43. Then, total RNA was subsequently extracted from these tissues using TRIzol reagent (ThermoFisher Scientific) according to the manufacturer’s instructions. The total RNA extracted from testes or ovaries, as well as the total RNAs isolated from AM and AF schistosomes, were reverse transcribed, and qPCR analysis was performed as described above.
Whole mount of in situ hybridization analyses of S. japonicum
Initially, adult males and female schistosomes (28 dpi) were fixed for 15 min in 4% paraformaldehyde (Ted Pella, USA) diluted in PBS containing 0.3% Triton-X 100 (hereafter PBSTx). Following fixation, worms were treated with a solution of 50 mM dithiothreitol, 1% NP-40, and 0.1% sodium dodecyl sulfate at 37 °C for 5–10 min, then dehydrated through a methanol gradient (100% PBSTx, 50% methanol, 100% methanol) with 5-min incubations per step. Bleaching was performed overnight under light using 6% hydrogen peroxide (diluted from a 30% stock in methanol). Subsequently, worms were rinsed twice with 100% methanol, incubated in 50% methanol for 10 min, and then transferred to PBSTx for an additional 10-min incubation. After permeabilization, worms were rinsed with PBSTx twice. Prehybridization began with incubation for 10 min in hybridization buffer (50% deionized formamide, 1.3× SSC (diluted from 20× SSC from Sigma, pH 5.5 with citric acid), 5 mM EDTA, 50 µg/µL torula yeast RNA, 0.2%Tween, 0.5% CHAPS, and 10% Heparin) mixed with PBSTx at a 1:1 ratio for 10 min. Then, the Worms were subjected to a 10-min wash with hybridization buffer at 54–56 °C in a hybridization oven (HBAID). The hybridization buffer was then removed, and the worms were incubated in fresh hybridization buffer for at least 2 h at 54–56 °C. Next, the digoxigenin-labeled riboprobe (5 µM) was added to the hybridization buffer, and the worms were incubated with the mixture overnight at 54–56 °C. Washes were then performed with solutions preheated to 54–56 °C in the following order: prehybridization solution mixed with 2× SSC at a 1:1 ratio, 2× SSC with 0.1% Triton-X 100, and 0.2× SSC with 0.1% Triton-X 100 in the hybridization oven with agitation. All washes were carried out twice for 30 min each at 54–56 °C. After stringency washes, schistosomes were cooled to RT and rinsed twice with maleic acid buffer (100 mM maleic acid, 150 mM NaCl, 0.1% Tween-20, pH 7.5). Blocking was performed with 10% horse serum for 2 h, followed by an overnight incubation at 4 °C with anti-DIG-AP-Fab fragments (Merk, catalog #: 11093274910, Lot #: 65752120) diluted 1:2000 in blocking solution (Roche, Switzerland). Next, worms were washed for 2 h in maleic acid buffer (buffer was changed every 20 min). Then, parasites were incubated in alkaline phosphatase (AP) buffer (100 mM Tris [pH 9.5], 100 mM NaCl, 50 mM MgCl2, 0.1% Tween-20 brought up to the final volume with 10% polyvinylalcohol solution) for 10 min and were then incubated in NBT/BCIP solution for 3 h or overnight. Then, worms were further washed with PBST and post-fixed. Finally, worms were mounted in 80% glycerol under glass coverslips and imaged.
As to probe preparation, cDNA fragments corresponding to each target RNA segment were amplified from cDNA synthesized from total RNA extracted from adult schistosomes, using the primers listed in Supplementary Table 1. PCR products were purified and then utilized to generate digoxigenin-labeled riboprobes using the MEGAscript Kit (ThermoFisher Scientific) and digoxigenin-11-UTP (Roche), following the manufacturer’s instructions.
Single-cell RNA sequencing
S. japonicum worms (16, 26 dpi) (100 worms for each gender) were transferred into 15 mL conical tubes and rinsed twice with PBS pre-warmed at 37 °C and dissociated into single-cell suspensions in 4 mL of 0.25% trypsin in HBSS for 15 min. Cell suspensions were passed through a nylon mesh (Corning, USA) and immediately doused with 10 mL of PBS supplemented with 1% BSA. The sample was then centrifuged at 500 × g for 5 min at 4 °C. Pellets were gently resuspended in PBS with 10% FBS, passed through a 40 μm nylon mesh (Corning, USA), and stained with diamine blue for 30 min, and then were washed and resuspended in PBS with 0.5% BSA and loaded on a SONY SH800S cell sorter. Dead cells were removed using the Dead Cell Removal Kit (Miltenyi Biotec, Germany).
Cell suspension and fluorescent dye AO/PI were mixed at a ratio of 1:1 and incubated for 30 s. The mixture was added to the slide and assessed cell viability using Count Star (Shanghai, China). Viability enrichment was carried out using the Dead Cell Removal Kit (Miltenyi Biotec) according to the manufacturer’s protocol. The cells were resuscitated to a concentration of 700–1200 cells/µl (viability ≥85%) in a final solution of 1× PBS 0.04 BSA before loading on the 10× Genomics Chromium platform. Over 10,000 cells were prepared for scRNA-seq libraries. Chromium Single Cell 3’ Library and Gel Bead Kit V3.1 (10× Genomics, PN1000268) was employed to generate single-cell gel beads in emulsion (GEM). The obtained cells were lysed, and the released RNA was performed for reverse transcription using primers containing poly T, barcode, unique molecular identifiers (UMIs), and read 1 primer sequence in GEMs. Barcoded cDNA was further purified and subsequently amplified by PCR. The adapter was ligated to add the sample index and the read 2 primer sequence. Upon quality control, the libraries were sequenced using the Illumina Novaseq 6000 platform in a 150 bp pair-ended manner (Berry Genomics Corporation, Beijing, China). Raw reads of fastq files were assembled from the Raw BCL files using Illumina’s bcl2 fastq converter. Then, the raw data were firstly processed through primary quality control using the parameters: (i) contain N more than 3; (ii) the proportion of bases with quality value below 5 is more than 20%; (iii) adapter sequence. All the downstream analyses were based on the obtained clean data.
Mapping of single-cell RNA sequencing reads
For reads mapping, a modified reference genome was constructed based on the S. japonicum v2 genome (HuSjv2; from PRJNA520774 WBPS16; https://parasite.wormbase.org), where mitochondrial genes (NCBI Reference Sequence NC_002544.1; naming as “Sj_”) and 4 ngRNA genes (naming as “SjPS”) were manually added. Raw sequencing data were mapped to the constructed reference genome using Cell Ranger v5.0 from 10× Genomics (https://www.10xgenomics.com/support/software/cell-ranger/).
Reference gene information
Orthologues between S. mansoni (v7) and S. japonicum (HuSjv2), protein sequences, and Gene Ontology terms were obtained from WormBase Parasite (https://parasite.wormbase.org/44; using the BioMart tool (access date: 2021-11-25). KEGG mapping was performed using KAAS with the BBH method (https://www.genome.jp/kegg/kaas/; access date: 2021-11-27). Single-cell markers were obtained from published studies for adult S. mansoni (filtered by p_val <10−20)45 and schistosomula46 (filtered by AUC > 0.7). Corresponding S. japonicum gene IDs were generated based on the ortholog information.
Single-cell RNA seq data processing
Cell Ranger filtered matrix files were imported to Seurat (v5.0.1; https://satijalab.org/seurat/) and filtered by nFeature_RNA > 500 & nCount_RNA > 1000 & percent.mt < 5. Each dataset was further processed with doublet detection using scDblFinder (v1.2.0), and only singlets were kept for further analysis. All four data sets were merged using Seurat and integrated using the Harmony Integration method. Briefly, each layer of data was normalized to find the top 2000 variable features, which were scaled (with percentage of mitochondrial expression regressed out) and used for dimensionality reduction using the command RunPCA (npcs = 30), and IntegrateLayer (method=HarmonyIntegration). Neighborhood graphs were constructed on harmony reduction using dims=1:30, and cell clusters were detected using the default algorithm with resolution 1.3. Afterwards, the layers were joined using the command JoinLayers.
Cell type annotation
To facilitate cell type annotation of Seurat clusters, the above-mentioned S. mansoni single-cell marker genes were used to calculate module scores in the current S. japonicum dataset. This was performed using the Seurat command AddModuleScore using cluster marker as features. Cell type information was further assessed by checking the marker genes, which were calculated using the FindAllMarkers command with test.use = ‘roc’. Expression of canonical cell type markers from S. mansoni was also examined. Candidate marker genes were validated using whole amount in situ hybridization, as stated in additional publications.
RNA-seq data processing and differential expression analyses
Parasites treated with rp1 siRNA, np1 siRNA, and control siRNA were subject to total RNA isolation for library preparation. Eight transcriptome libraries (two biological replicates for each treatment or control) were sequenced using the BGISEQ-500 platform. Clean reads were obtained after removing low-quality reads, adapters, as well as any reads containing poly-N from raw reads and mapped to the self-constructed reference genome using STAR_2.6.1 d (alignIntronMin 1047). Read counts for genes were summarized using feature Counts v1.6.448). For downstream analysis, the web tool iDEP 1.12 (http://bioinformatics.sdstate.edu/idep/)49 was used using the DESeq250 pipeline. Differentially expressed genes were selected using padj<0.05 and 1.3-fold change. For functional enrichment of Gene Ontology, only terms with 5–2000 genes in the category “Biological Process” and FDR < 0.05 were selected.
Worm culture and siRNA treatment
S. japonicum were obtained from the infected mice (24–28 dpi), and females were separated and cultured in 12-well flat bottom plates containing 2 mL complete RPMI-1640 medium (ThermoFisher Scientific) supplemented with 2 g/L glucose, 0.3 g/L L-glutamine, 2.0 g/L NaHCO3, 15% fetal bovine serum (heat inactivated), and 5% penicillin/streptomycin (10,000 U penicillin and 10 mg/streptomycin in 0.9% NaCl). The worms were incubated in a humidified 5% CO2 chamber at 37 °C. siRNA duplexes and control siRNAs (3 μg per experiment, chemically synthesized in Shanghai GenePharma, China; Supplementary Table 2) were electroporated (125 V, 20 ms, 1 pulse in 200 μL RPMI 1640 medium) into cultured parasites. Subsequently, worms were then transferred into a 12-well cell culture plate containing 2 mL fresh media. Eggs were collected and counted under an inverted microscope (Olympus, Japan) at the indicated times. Treated worms were collected at 96 h of post electroporation for RT-qPCR analysis and Carmine Red staining. Whole-mount parasites were visualized using confocal microscopy.
Confocal microscopy
At 96 h after electroporation, worms were preserved, stained, and mounted and subjected to morphological observation via a Nikon confocal microscope (Nikon, Japan)51. Briefly, parasites were fixed in a solution containing formalin (10%), alcohol (48%), and glacial acetic acid (2%), stained with Carmine Red, cleared with a 0.5% hydrochloric alcohol solution, and stored as whole mounts. Images were captured with a Nikon CLSI laser confocal microscope (Nikon) under a 488 nm He/Ne laser.
Fast blue BB and EdU staining
Fast Blue BB staining was carried out using the following procedure. Briefly, worms were fixed in 4% formaldehyde in PBSTx for 4 h, then stained for 5 min in freshly prepared, filtered 1% Fast Blue BB solution in PBSTx. Parasites were washed with PBSTx three times (5 min per wash) and mounted on slides with 80% glycerol for microscopic observation.
EdU staining was conducted based on a protocol52. Worms were in vitro labeled with 10 μM EdU (Merck) for 20 h. Subsequently, the parasites were harvested and incubated in 6 M MgCl2 for 1 min to terminate viability. Then, the parasites were fixed in 4% formaldehyde in PBSTx for 4 h, and dehydrated in Methanol. Next, worms were rehydrated in 50% methanol in PBSTx, bleached under bright light for 5–6 h, then treated with 5 µg/ml Proteinase K (ThermoFisher Scientific) in 1× PBSTx for 30 min. The parasites were treated with 4% formaldehyde in PBSTx for 10 min for post-fixation. Worms were stained with EdU Detection solution (789 μL 1× PBS, 10 μL 100 mM CuSO4, 1 μL 10 mM Azide-fluor 488, 200 μL 500 mM Asorbic Acid) for 30 min in the dark. Parasites were then stained with the DAPI solution (1 µg/ml in PBSTx) for 1 h. The worms were further cleared with 80% glycerol and mounted in Vectashield (Yeasen, China). Images were acquired using a Nikon confocal microscope (Nikon) at 456 and 488 nm.
Egg viability
The viability of schistosome eggs was assessed with the acridine orange staining method, as judged by the intensity of the fluorescence of the stained eggs53. Briefly, the eggs were stained with 0.01% acridine orange solution (Merck) at 37 °C and shielded from light for 2 h. The fluorescence of the stained eggs was subsequently recorded and counted under an inverted fluorescence microscope with a CCD camera. Cell viability in eggs was determined using a Cell Titer-Lumi Luminescent cell viability assay kit (Beyotime Biotechnology, China) according to the manufacturer’s instructions. Subsequently, the obtained luciferase activity was normalized with protein concentration using the Pierce BCA protein assay kit combined with the Compat-Able protein assay preparation reagent set (ThermoFisher Scientific).
RT-qPCR analysis of the abundance of ngRNAs in electroporated schistosomes
At 96 h post-electroporation, total RNA was extracted from treated worms using TRIzol (ThermoFisher Scientific) according to the manufacturer’s instructions. First-strand cDNA was produced from 100 ng RNA using a PrimeScript First Strand cDNA Synthesis Kit (TaKaRa) with random primers. PCR was then conducted using the primers listed in Supplementary Table 1. Gene expression was normalized to S. japonicum NADH as an internal control, and relative expression levels were calculated using 2−ΔCt method41.
Screening of a YTH for analyzing Rp1 RdRp interactors
To identify the interactors of RdRp proteins, we cloned the cDNA encoding the conserved domain of rp1 RdRP (forward: 5′gcatatggccatggaggccgaattcCAGTTCCCCATGATAGAGACTC3′, reverse: 5′gcggccgctgcaggtcgacggatccTCAAGGGTTTTTGCATGCCAACATA3′, synthesized by Invitrogen) and fused it to the binding domain of the pGBKT7 vector to serve as the bait. YTH screening was conducted using a previously prepared S. japonicum cDNA library by the Yeastmaker Yeast Transformation System 2 (TaKaRa). The YTH assay was carried out54. The plasmids were isolated from all of the positive clones for sequencing. For the retransformation validation of the YTH assay, the independently prepared positive clone plasmids and RdRp plasmid were co-transformed into YTH Gold yeast competent cells (TaKaRa), respectively. Yeast transformation was performed using the lithium acetate method at 30 °C55. Transformants were first selected on medium lacking Trp and Leu. Positive colonies were then subsequently transferred to medium lacking selection media supplemented with SD/-Trp/-Leu/-His/-Ade with or without X-α-gal.
Inhibition of Rp1 RdRp interactors
Double-stranded RNA (dsRNA) targeting Lin-10 and Lp1 was used to inhibit these Rp1 RdRp interactors. Briefly, PCR was performed using specific primers to generate the templates for in vitro transcription (Supplementary Table 1). The dsRNA was synthesized using the MEGAscript T7 Transcription Kit (ThermoFisher Scientific). The reaction contained 100 ng of PCR product, 2 µL of 10× reaction buffer, 2 µL of each rNTP, 2 µL of T7 RNA polymerase, and nuclease-free H2O to 20 µL. Ten 26 dpi females were cultured in each well using ABC169 medium and then treated with 30 μg/mL dsRNA. This dsRNA was replenished during medium changes at 1, 3, and 6 days post-treatment. Egg number was monitored daily. Fast Blue BB staining was performed at 6 days post-treatment.
As to RT-qPCR analyses, total RNA was isolated from treated worms using TRIzol reagent (ThermoFisher Scientific) and reverse-transcribed using the PrimeScript™ RT Reagent Kit (TaKaRa). PCR reactions were performed in 10 µL volumes containing 5 µL ChamQ Universal SYBR qPCR Master Mix (Vazyme, China), 0.2 µL of each 10 µM primer (Supplementary Table 1), 0.5 µL cDNA, and added nuclease-free H2O to 10 µL under the condition at 95 °C for 30 s, followed by 40 cycles of 95 °C for 10 s and 60 °C for 30 s. Data were analyzed as described above.
PCR analysis for determining the complementarity of RNAs
A strand-specific RT-PCR approach was used to detect the complementary strand of ngRNAs56. Total RNA was isolated from adult S. japonicum as described above. RT was performed in a final volume of 10 μL, containing 6 μL total RNA (400 ng), 2 μL 5× Prime-Script Buffer (TaKaRa), 0.5 μL PrimeScript RT Enzyme Mix I (TaKaRa), 1 μL of RNase inhibitor (TaKaRa), and 0.5 μL of specific primers (10 μM) (Supplementary Table 1, the primers were synthesized by Invitrogen) matching the complementary strand of viral RNA. Next, PCR was performed in a 15 μL reaction mixture containing 1 μL cDNA, 7.5 μL 2× SYBR Primer Ex TaqII (TaKaRa), 6 μL H2O, and 0.5 μL primer mixture (10 μM) (Supplementary Table 1) using the following thermal cycling profile: 95 °C for 5 min, followed by 40 cycles of amplification (95 °C for 5 s, 57 °C for 30 s, 72 °C for 20 s). The PCR products were analyzed using 2% agarose gels and visualized by staining with Gold View (Yeasen).
Plasmid construction, cell culture, and transfection
The reversed complementary Gaussia luciferase reporter gene embedded into 5′UTR and 3′UTR of Rp1 and reversed complementary np1 flanked by BamH I and Hind III sites were synthesized from GENERAY (GENERAY, China). These fragments were subcloned into the pcDNA3.1(+) vector, respectively. Additionally, the conserved domain of Rp1 RdRP was also subcloned into the pcDNA3.1(+) vector.
293T cells were purchased from the American Type Culture Collection (ATCC, USA; catalog# CRL-3216). 293T cells were maintained in Dulbecco’s Modified Eagle’s Medium (DMEM) (ThermoFisher Scientific), supplemented with 10% fetal bovine serum (FBS) (ThermoFisher Scientific) and 1% penicillin/streptomycin (Gibco, MA, USA). Cells were cultured in a humidified incubator at 37 °C with 5% CO2. 293T cells were seeded in a 48-well plate after passaging. When cell density reached 70% confluence, transfection was performed. The rp1 RdRp plasmid and either the embedded Gaussia luciferase plasmid or the np1 plasmid at a 1:4 ratio (50 ng: 200 ng) were transfected into the cells and then incubated for 8 h. Meanwhile, the indicated concentrations of Favipiravir were added to the cells. After 24 h of post-transfection and/or treatment, cells were collected for luciferase assay and RT-qPCR analyses as described below.
Preparation of cell lysate, luciferase assay and RT-qPCR analyses
The cells were rinsed with phosphate-buffered saline (PBS, pH 7.4) and lysed using a lysis buffer from Promega Dual-Luciferase Assay (Promega, USA). The lysates were centrifuged at 16,000 g at 4 °C for 15 min, and the supernatants were carefully transferred to new tubes. Luciferase activity in cell lysates was measured using the Promega Dual-Luciferase Assay, normalized to protein concentrations in worm lysates using the Pierce BCA protein assay kit and Compat-Able protein assay preparation reagent set (ThermoFisher Scientific). Partial cells were collected for total RNA isolation. RT-qPCR was performed to determine the expression of np1 using the specific primers (Supplementary Table 1).
Dot blot assay and immunofluorescent analysis
For the dot blot assay, 500 ng dsRNA was dropped on the nylon membrane and then was cross-linked using a UV cross-linking device (1200 mJ/cm2, 3 min). Upon the blocking of the membrane using 5% skimmed milk powder, the membrane was incubated with primary antibody J2 (1:1000 dilution) (Scicons, Hungary; catalog # 10010200, batch # 72-1710) overnight at 4 °C. The secondary antibody was used HRP conjugated Goat Anti-Mouse IgG(H + L) (1:5000 dilution) (Servicebio, China; catalog# GB23301, lot # AC231007). Imaging signals were obtained by developing with enhanced chemiluminescence (ECL) reagents (ThermoFisher Scientific).
Worms were permeabilized in Tris-Triton X-100 buffer with 5% β-mercaptoethanol and incubated for 20 h in a rocker at 37 °C. Then the worms were washed and put into collagen IV (1 mg/ml) prepared in PBS for 35 min. Worms were quenched in cold antibody buffer composed of 1× PBS, 0.1% BSA, Triton X-100, and 0.05% sodium azide for 15 min. Worms were washed and incubated with J2 antibody (dilution: 1:1000) overnight at 4 °C. After washing, worms were incubated with goat anti-mouse secondary antibody conjugated with Alexa Fluor 568 (dilution: 1:2500; ThermoFisher Scientific; catalog #A-11004, lot # 3141857) for 2 h. Cell nuclei were counter-stained with 4′,6-diamidino-2-phenylindole (1 µg/ml). Finally, worms were mounted with an antifading mounting medium, and images were acquired with a confocal laser scanning microscope (ZEISS)57.
Structural modeling and drug-protein docking of RdRp
To predict the rp1/2 RdRp structure, we employed the alphafold server (https://alphafoldserver.com/) to predict their structures for the drug-protein docking (DPD) using the default settings. The obtained protein molecule PDB file and the small molecules (Favipiravir and its derivative FTP, in mol2 format) were subjected to the DPD program using AutoDock software (https://autodock.scripps.edu/) following the official protocol. The docking result was further processed by AutoDockTools (https://autodocksuite.scripps.edu/adt/) to obtain a drug-protein complex, which was further visualized by PyMol software (https://pymol.org/). The docking affinity of protein and ligands was quantified by the free binding energy.
RdRp inhibitor treatment
S. japonicum were collected from mice 24–28 days of post-infection, and females were separated and cultured in 12-well flat bottom plates containing 2 mL complete RPMI-1640 medium (ThermoFisher Scientific) supplemented with 2 g/L glucose, 0.3 g/L L-glutamine, 2.0 g/L NaHCO3, 15% fetal bovine serum (heat inactivated), and 5% penicillin/streptomycin (10,000 U penicillin and 10 mg/streptomycin in 0.9% NaCl) in a humidified 5% CO2 chamber at 37 °C. Favipiravir was added to the culture medium at the indicated concentrations. The number of egg production was monitored during the time course. Egg viability was determined as described above.
Expression and purification of np1 in Escherichia coli and identification of recombinant protein
The cloning and expression of np1 were carried out based on the standard molecular procedures. In brief, the ORF of np1 was cloned by PCR-based amplification and the insertion of the fragment into a pET28a(+) expression vector (Novagen, Germany). Restriction recognition sites of EcoR І and Xho I were introduced by PCR using the specific primers (forward: 5′CCGGAATTCATGTGTAATAAAGCCAGGTC3′; reverse: 5′CCGCTCGAGTTAAACCATTTTCCTCTTAG3′, synthesized by Invitrogen) with template of prepared cDNA. Recombinant np1 was expressed in Escherichia coli BL21 cells transformed with pET28a (+)-np1 plasmids. The recombinant plasmids were induced under 1.0 mM IPTG at 37 °C. Recombinant NP1(rNP1) was purified by a Ni-NTA His* Band Purification Kit according to the manufacturer’s instructions (Novagen). The purified proteins were analyzed by 15.0% sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS–PAGE), and the recombinant NP1 was verified by mass spectrometry with ESI-Q-TOF (Applied Biosystems, USA). The results of mass spectrometry were searched against NCBInr database using MASCOT under default parameters.
Generation of rabbit antiserum
Two rabbits were intramuscularly immunized with 100 µg recombinant NP1 that was emulsified with 206 adjuvants (Sigma, USA), followed by two boosts at a 3-week interval. Sera were collected after 7 days of the last boost and stored at −20 °C.
Sucrose ultracentrifugation
Sucrose solutions were prepared (w/w) in PBS. Sucrose gradients were prepared by sequentially adding equal amounts of 30% sucrose solution, 45% sucrose solution and 60% sucrose solution. Schistosomes (28 days old, 0.5 g weight) were lysed in 5 mL PBS, and lysates were then centrifuged at 2000 × g for 5 min. The supernatants were carefully transferred into fresh tubes and then centrifuged at 8000 × g for 5 min. The supernatants were also carefully transferred into fresh tubes and centrifuged at 10,000 × g for 10 min. Supernatants were then filtered using a 0.22-µM filter (EMD Millipore) and further centrifuged at 210,000 × g for 2 h to spin down particles.
Electron microscopy
Schistosome and schistosome cell lysates were prepared as described above. Schistosome cell lysates from sucrose ultracentrifugation were deposited onto 200-mesh formvar-coated grids (Agar Scientific) and left to dry at RT. Grids were rinsed with H2O and subsequently treated with 1% uranyl acetate (System Biosciences) for a 5-min incubation for staining. Then, the grids were washed once in 70% ethanol followed by four times of washes with H2O. Then, grids were loaded onto the sample holder of a transmission electron microscope (Hitachi H-7600; Hitachi, Tokyo, Japan) and exposed to an 80 kV electron beam for morphological observation and image capture.
Ultrathin tissue sections for TEM and Immune TEM
Adult schistosomes (28 dpi) were subjected to overnight fixation at 4 °C using 2.5% glutaraldehyde diluted in 0.1 M cacodylate buffer. The worms were washed three times with PBS (3 × 10 min). Subsequently, the parasites were rinsed with 0.1 M PBS (pH 7.4) and post-fixed with 1% osmium tetroxide in PBS (pH 7.4) at 4 °C for 1 h. Dehydration was performed through a graded ethanol series, followed by transfer to propylene oxide, and the samples were embedded in Eponate 812 resin (Ted Pella, USA). Longitudinal ultra-thin sections (60–70 nm thick) were prepared, contrasted sequentially with uranyl acetate and lead citrate, and imaged under a transmission electron microscope with an accelerating voltage of 80 kV.
Ultrathin sections (90 nm) encompassing the regions of interest were harvested on uncoated nickel grids (M200-NI, Electron Microscope Sciences). Sections were etched with 1% sodium periodate (NaIO₄) for 5 min at room temperature (RT), followed by rinsing with H2O and 0.1 mol/L glycine solution. Subsequently, sections were blocked with 1% BSA for 15 min, probed with primary antibodies (1:500 dilution) in 1% BSA at 4 °C for 48 h, and then incubated with Protein A-conjugated 10 nm gold particles (810.111, Aurion) for 2 h at RT. After incubation, the sections were washed 5× 10 min in PBS buffer. Antibody complexes were immobilized via incubation in 2% glutaraldehyde in phosphate buffer for 10 min at RT. The grids were rinsed three times with H2O and stained with 3% uranyl acetate for 10 min and 3% lead citrate (Leica Ultrostain 2) for 6 min. Sections were observed using a JEM-1400 Plus electron microscope, equipped with a Quemsa TEM CCD camera, followed by image capture via iTEM software (TEM Imaging Platform).
In vivo inhibition of rp1 RdRp in mice infected with S. japonicum
BALB/c mice (male, 7–8 week old, mean body weight 25 ± 2 g) were obtained from the Shanghai SLAC Laboratory Animal Co., Ltd and were housed as described above. Mice (n = 18) were randomly allocated into three groups. Each mouse was infected with S. japonicum cercariae (n = 100 ± 5) via abdominal skin penetration. At 18 dpi, mice in each group received a tail vein injection of 1 µg siRNA or scrambled siRNA diluted in 200 μL of nuclease-free H2O. Concurrently, one group was administered an equal volume of PBS via the tail vein. Four subsequent injections were given on days 20, 22, 24, and 26 dpi. At 28 dpi, worms were collected via liver perfusion, and liver worm burden and egg deposition in the liver were counted. The expression of rp1 in collected worms was analyzed by RT-qPCR. Equal weight of liver tissues was collected from each mouse and then subjected to egg purification as described previously58, and the egg hatch experiment was carried out. Liver tissue was fixed and analyzed for Hematoxylin and Eosin staining as described below.
Histopathology of animal liver infected with S. japonicum
To evaluate the pathological changes in livers following in vivo rp1 inhibition, mouse livers were harvested, chopped into small cubes (1.0 × 1.0 × 0.3 cm), and fixed in 10% formalin. Subsequently, the fixed tissues were dehydrated through a graded ethanol series (70, 80, 95, and 100%), cleared with xylene, and embedded in paraffin wax. Finally, the paraffin-embedded specimens were sectioned into 6 µm thick slices and stained using the Hematoxylin and Eosin method according to the standard protocols.
WISH and FISH analyses of RdRp location in D. japonica
WISH analysis of RdRp localizations was performed as described above, except the hybridization at 56 °C using the probe as in vitro transcribed using PCR products amplified by the specific primers forward 5′CGGCACTAGTCAATCAACCCTCT3′, reverse 5′TAATACGACTCACTATAGGTGTTTGGTTTACTTGTTGGCTCATTC3′, synthesized by Invitrogen). WISH was carried out as described above. Fluorescence in situ hybridization (FISH) was performed as described below. Briefly, worms were washed with ice-cold 2% HCl for 5 min before fixation, and then were fixed for 30 min at RT in 4% formaldehyde in PBS. Then, worms were washed using PBSTx (1× PBS + 0.3% Triton X-100). After fixation, worms were dehydrated and rehydrated in methanol and then treated with 1.2% H2O2 under light for 4 h. The worms were then rehydrated in methanol and then permeabilized with 10 µg/mL Proteinase K (ThermoFisher Scientific) for 10 min, and fixed for 10 min immediately. The worms were washed with PBSTx twice and then carried out the hybridization at 56 °C. Then, the worms were washed with TNTx (100 mM Tris pH 7.5, 150 mM NaCl, 0.3% Triton X-100) solution twice and then were incubated in the solution containing FISH blocking solution (5% horse serum (Sigma) and 0.5% Western Blocking Reagent (Merk) in TNTx (100 mM Tris pH 7.5, 150 mM NaCl, 0.3% Triton X-100) overnight at 4 °C. Then, the worms were incubated with anti-DIG-POD (Merk, catalog #11207733910, lot # 77798000) for 4 h at RT. Upon the wash of several times using TNTx, the Cy5 TSA Fluorescence System Kit (YEASEN) was used to stain the worms for 15 min. Then, the worms were subjected to washing twice using TNTx and stained with DAPI (10 μg/ml). The worms were further washed with PBSTx and then cleared with 80% glycerol, and mounted in Vectashield (Yeasen). The images were captured under a confocal microscope (Nikon).
RdRp silencing in D. japonica
The DNA template for generating RdRp dsRNA was obtained by PCR using the specific primers (Supplementary Table 1). The dsRNA was synthesized using the T7 RiboMAX™ Express RNAi kit (Promega, USA), then it was purified with 2.5 volumes of ethanol and 1/10 volume of NH4OAc (pH 5.2) after DNase I treatment (TaKaRa). dsRNA was delivered in planarian (5 μg/ml) based on the protocol with minor modification59. Control animals were treated with S. japonicum Mettl3 dsRNA. After 6 days of post-treatment, worms were collected for WISH and RT-qPCR analyses. The primers of S. japonicum Mettl3 were used to synthesize the dsRNA using the T7 RiboMAX™ Express RNAi kit (Promega, USA) as an irrelevant control dsRNA (Supplementary Table 1). The effect of RdRp inhibition was also evaluated by qPCR using the specific primers (Supplementary Table 1).
Statistical analysis
The data were analyzed using GraphPad Prism 8. Statistical analyses were conducted using Student’s t-test or one-way ANOVA. P ≤ 0.05 was considered significant.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The raw data generated in this study have been deposited in the NCBI Sequence Read Archive database under Bioproject accession: PRJNA1200787 of Sequence Read Archive (SRA) with accession numbers: SRR31784414, SRR31784413, SRR31784412, SRR31784411, SRR31784410, SRR31784409, SRR31784408, SRR31784407, SRR31784406, SRR31784405, SRR31784404, SRR31784403, SRR31784402, SRR31784401, and SRR31784400 [https://www.ncbi.nlm.nih.gov/sra/?term=PRJNA1200787]. The scRNA-seq raw data have also been deposited in the NCBI Sequence Read Archive database under Bioproject accession: PRJNA1417122 of Sequence Read Archive (SRA) with accession numbers: SRR37125568, SRR37125569, SRR37135933, SRR37135934, SRR37125570, and SRR37125571 [https://www.ncbi.nlm.nih.gov/sra/?term=PRJNA1417122]. The four ngRNAs were also deposited in the GenBank with accession number: PX975019-PX975022. All data are included in the article or the Supplementary Information. Source data are provided with this paper.
References
Lewis, F. A. & Tucker, M. S. Schistosomiasis. Adv. Exp. Med. Biol. 766, 47–75 (2014).
Wang, A. L. & Wang, C. C. A linear double-stranded RNA in Trichomonas vaginalis. J. Biol. Chem. 260, 3697–3702 (1985).
El-Gayar, E. K., Mokhtar, A. B. & Hassan, W. A. Molecular characterization of double-stranded RNA virus in Trichomonas vaginalis Egyptian isolates and its association with pathogenicity. Parasitol. Res. 115, 4027–4036 (2016).
Wang, A. L. & Wang, C. C. The double-stranded RNA in Trichomonas vaginalis may originate from virus-like particles. Proc. Natl. Acad. Sci. USA 83, 7956–7960 (1986).
Akopyants, N. S., Lye, L. F., Dobson, D. E., Lukes, J. & Beverley, S. M. A novel bunyavirus-like virus of Trypanosomatid Protist parasites. Genome Announc. 4, e00715–16 (2016).
Kraeva, N. et al. Leptomonas seymouri: adaptations to the dixenous life cycle analyzed by genome sequencing, transcriptome profiling and co-infection with Leishmania donovani. PLoS Pathog. 11, e1005127 (2015).
Widmer, G., Comeau, A. M., Furlong, D. B., Wirth, D. F. & Patterson, J. L. Characterization of a RNA virus from the parasite Leishmania. Proc. Natl. Acad. Sci. USA 86, 5979–5982 (1989).
Zangger, H. et al. Detection of Leishmania RNA virus in Leishmania parasites. PLoS Negl. Trop. Dis. 7, e2006 (2013).
Molyneux, D. H. Virus-like particles in Leishmania parasites. Nature 249, 588–589 (1974).
Ives, A. et al. Leishmania RNA virus controls the severity of mucocutaneous leishmaniasis. Science 331, 775–778 (2011).
Eren, R. O. et al. Mammalian innate immune response to a leishmania-resident RNA virus increases macrophage survival to promote parasite persistence. Cell Host Microbe 20, 318–328 (2016).
Brettmann, E. A. et al. Tilting the balance between RNA interference and replication eradicates Leishmania RNA virus 1 and mitigates the inflammatory response. Proc. Natl. Acad. Sci. USA 113, 11998–12005 (2016).
Rossi, M. et al. Type I interferons induced by endogenous or exogenous viral infections promote metastasis and relapse of leishmaniasis. Proc. Natl. Acad. Sci. USA 114, 4987–4992 (2017).
Wang, J. et al. Dynamic transcriptomes identify biogenic amines and insect-like hormonal regulation for mediating reproduction in Schistosoma japonicum. Nat. Commun. 8, 14693 (2017).
He, Y. et al. siRNA-mediated knockdown of two tyrosinase genes from Schistosoma japonicum cultured in vitro. Exp. Parasitol. 132, 394–402 (2012).
Kim, S. K. & Horvitz, H. R. The Caenorhabditis elegans gene lin-10 is broadly expressed while required specifically for the determination of vulval cell fates. Genes Dev. 4, 357–371 (1990).
Sun, C. et al. MicroRNA-1 targets ribosomal protein genes to regulate the growth, development and reproduction of Schistosoma japonicum. Int J. Parasitol. 53, 637–649 (2023).
Campos R. K. et al. RPLP1 and RPLP2 are essential flavivirus host factors that promote early viral protein accumulation. J. Virol 91, e01706–16 (2017).
Kong, J. et al. A ribosomal protein S5 isoform is essential for oogenesis and interacts with distinct RNAs in Drosophila melanogaster. Sci. Rep. 9, 13779 (2019).
Gong, P. & Peersen, O. B. Structural basis for active site closure by the poliovirus RNA-dependent RNA polymerase. Proc. Natl. Acad. Sci. USA 107, 22505–22510 (2010).
Pinzon, N. et al. Functional lability of RNA-dependent RNA polymerases in animals. PLoS Genet. 15, e1007915 (2019).
Saberi, A., Gulyaeva, A. A., Brubacher, J. L., Newmark, P. A. & Gorbalenya, A. E. A planarian nidovirus expands the limits of RNA genome size. PLoS Pathog. 14, e1007314 (2018).
Burrows, J. T. A., Depierreux, D., Nibert, M. L. & Pearson, B. J. A novel taxon of monosegmented double-stranded RNA viruses endemic to triclad flatworms. J Virol 94, e00623–20 (2020).
Pak, J. & Fire, A. Distinct populations of primary and secondary effectors during RNAi in C. elegans. Science 315, 241–244 (2007).
Maniar, J. M. & Fire, A. Z. EGO-1, a C. elegans RdRP, modulates gene expression via production of mRNA-templated short antisense RNAs. Curr. Biol. 21, 449–459 (2011).
Kapuscinski, M. L. et al. Genomic characterization of 99 viruses from the bunyavirus families Nairoviridae, Peribunyaviridae, and Phenuiviridae, including 35 previously unsequenced viruses. PLoS Pathog. 17, e1009315 (2021).
Shi, M. et al. Redefining the invertebrate RNA virosphere. Nature 540, 539–543 (2016).
Liu, J. et al. Schistosoma japonicum extracellular vesicle miRNA cargo regulates host macrophage functions facilitating parasitism. PLoS Pathog. 15, e1007817 (2019).
Chen, Y. M. et al. Host traits shape virome composition and virus transmission in wild small mammals. Cell 186, 4662–4675 e4612 (2023).
Hahn, M. A., Rosario, K., Lucas, P. & Dheilly, N. M. Characterization of viruses in a tapeworm: phylogenetic position, vertical transmission, and transmission to the parasitized host. ISME J. 14, 1755–1767 (2020).
Castiglioni, V. G. et al. Story of an infection: viral dynamics and host responses in the Caenorhabditis elegans-Orsay virus pathosystem. Sci. Adv. 10, eadn5945 (2024).
Quek, S. et al. Diverse RNA viruses of parasitic nematodes can elicit antibody responses in vertebrate hosts. Nat. Microbiol. 9, 2488–2505 (2024).
Shan, T. et al. The fecal virome of pigs on a high-density farm. J. Virol. 85, 11697–11708 (2011).
Zhang, W. et al. Virome comparisons in wild-diseased and healthy captive giant pandas. Microbiome 5, 90 (2017).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Li, D. et al. MEGAHIT v1.0: a fast and scalable metagenome assembler driven by advanced methodologies and community practices. Methods 102, 3–11 (2016).
Deng, X. et al. An ensemble strategy that significantly improves de novo assembly of microbial genomes from metagenomic next-generation sequencing data. Nucleic Acids Res. 43, e46 (2015).
Kumar, S., Stecher, G., Li, M., Knyaz, C. & Tamura, K. MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 35, 1547–1549 (2018).
Minh, B. Q. et al. IQ-TREE 2: new models and efficient methods for phylogenetic inference in the genomic era. Mol. Biol. Evol. 37, 1530–1534 (2020).
Letunic, I. & Bork, P. Interactive tree of life (iTOL) v6: recent updates to the phylogenetic tree display and annotation tool. Nucleic Acids Res. 52, W78–W82 (2024).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25, 402–408 (2001).
Hahnel, S., Lu, Z., Wilson, R. A., Grevelding, C. G. & Quack, T. Whole-organ isolation approach as a basis for tissue-specific analyses in Schistosoma mansoni. PLoS Negl. Trop. Dis. 7, e2336 (2013).
Giri, B. R., Li, H., Chen, Y. & Cheng, G. Preliminary evaluation of neoblast-like stem cell factor and transcript expression profiles in Schistosoma japonicum. Acta Trop. 187, 57–64 (2018).
Howe, K. L., Bolt, B. J., Shafie, M., Kersey, P. & Berriman, M. WormBase ParaSite—a comprehensive resource for helminth genomics. Mol. Biochem. Parasitol. 215, 2–10 (2017).
Wendt, G. et al. A single-cell RNA-seq atlas of Schistosoma mansoni identifies a key regulator of blood feeding. Science 369, 1644–1649 (2020).
Diaz Soria, C. L. et al. Single-cell atlas of the first intra-mammalian developmental stage of the human parasite Schistosoma mansoni. Nat. Commun. 11, 6411 (2020).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Ge, S. X., Son, E. W. & Yao, R. iDEP: an integrated web application for differential expression and pathway analysis of RNA-Seq data. BMC Bioinform. 19, 534 (2018).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Zhu, L. et al. MicroRNAs are involved in the regulation of ovary development in the pathogenic blood fluke Schistosoma japonicum. PLoS Pathog. 12, e1005423 (2016).
Wang, J., Chen, R. & Collins, J. J. 3rd. Systematically improved in vitro culture conditions reveal new insights into the reproductive biology of the human parasite Schistosoma mansoni. PLoS Biol. 17, e3000254 (2019).
Liu, L., Tang, J. C., Zhang, F., Wang, X. & Xu, B. J. The determination of the viability of Schistosomal eggs by a novel technique: electrorotation. Trop. Biomed. 30, 367–374 (2013).
Zheng, Z. Z. et al. Alternative splicing regulation of doublesex gene by RNA-binding proteins in the silkworm Bombyx mori. RNA Biol. 16, 809–820 (2019).
Daniel Gietz, R. & Woods, R. A. Transformation of yeast by lithium acetate/single-stranded carrier DNA/polyethylene glycol method. In: Guide to Yeast Genetics and Molecular and Cell Biology—Part B (eds Guthrie, C. & Fink, G. R.) (Academic Press, 2002).
Chatterjee, S. N., Devhare, P. B. & Lole, K. S. Detection of negative-sense RNA in packaged hepatitis E virions by use of an improved strand-specific reverse transcription-PCR method. J. Clin. Microbiol. 50, 1467–1470 (2012).
He, J., Mo, D., Chen, J. & Luo, L. Combined whole-mount fluorescence in situ hybridization and antibody staining in zebrafish embryos and larvae. Nat. Protoc. 15, 3361–3379 (2020).
Cheng, G. F. et al. Proteomic analysis of differentially expressed proteins between the male and female worm of Schistosoma japonicum after pairing. Proteomics 5, 511–521 (2005).
Chen, R. et al. A male-derived nonribosomal peptide pheromone controls female schistosome development. Cell 185, 1506–1520 e1517 (2022).
Acknowledgements
We thank Dr. Christoph G. Grevelding from Justus Liebig University Giessen and Dr. Richard E. Davis from University of Colorado for their comments on the manuscript. This study was, in part or in whole, supported by the Fundamental Research Funds for the Central Universities (PA2024000471 to GC, PA2023000254 to GC, PA2022000010 to GC, PA2021000155 to GC, and PA2020000431 to GC) and by National Natural Science Foundation of China (Grants No. 31472187 to GC and 31672550 to GC). The study was also sponsored by the Shanghai Tongji University Education Development Foundation of China.
Author information
Authors and Affiliations
Contributions
G.C. designed the research; T.S., T.X., L.L., X.W., S.L., and Y.Z. performed the experiments; Z.L., T.S., and C.F. analyzed the data; G.C. and Z.L. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Shan, T., Lu, Z., Xia, T. et al. Non-parasite genome encoded virus-like RNAs reprogram the pathogenicity of human blood flukes. Nat Commun 17, 2995 (2026). https://doi.org/10.1038/s41467-025-67822-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-67822-1