Abstract
Immune exclusion is associated with adverse prognosis in tumors, and this phenomenon may be facilitated by a physical barrier constituted by the extracellular matrix (ECM). Hypoxia and acidity promote immune exclusion, however the detailed mechanism remains unclear. This study demonstrate that G-protein-coupled-receptor-4 (GPR4), as a proton sensor, induces immune exclusion in colon cancer by promoting collagen fiber alignment and deposition. Specifically, GPR4 facilitates collagen alignment via LOXL2 regulation and enhances collagen I by modulating TGF-β, both mediated by the JAK2/STAT3 pathway. We further validate our findings in a male animal model, observing that elevated GPR4 expression in colon cancer results in an immune-excluded microenvironment. Inhibition of the JAK2/STAT3 pathway and LOXL2 function effectively reverse immune exclusion and enhance immunotherapy efficacy. Collectively, our findings elucidate a mechanism of immune exclusion and propose a potential target for improving the therapeutic efficacy of immunotherapy through the remodeling of the tumor ECM.
Similar content being viewed by others
Introduction
Colorectal cancer (CRC) is the third most common cancer, with 1.926 million new CRC cases, and the second leading cause of cancer-related deaths, with 904,000 CRC-related deaths, posing an enormous threat to public health. Colon cancer (CC) is the major subtype, accounting for approximately 90% of CRC1. There is a huge difference for CC patients with CC between the early and advanced stages. The 5-year survival rates for patients with CC with localized lesions and distant metastases are 91% and 14%, respectively1. Immunotherapy, especially immune checkpoint inhibitors (ICIs), is highly effective for some advanced tumors, such as lung adenocarcinoma, clear cell renal cancer, and melanoma2,3. However, the prognosis of patients with CC undergoing ICI therapy remains suboptimal, with a median progression-free survival (PFS) of 16.5 months in patients with MSI versus 8.2 months in patients with MSS.4. The therapeutic efficacy of ICIs is mainly induced by promoting the antitumor immune activity of CD8 + T cells in the tumor immune microenvironment (TIME)4,5. Therefore, it is essential to explore the TIME from different perspectives to reveal new potential immunotherapy targets.
Recently, based on the quantity and spatial distribution of immune cells, particularly T lymphocytes, within the TIME, it has been classified into three distinct subtypes: the immune-inflamed, immune-excluded, and immune-desert phenotypes6. The immune-excluded phenotype is specific, characterized by abundant immune cells in the stroma instead of the tumor nest7. The extracellular matrix (ECM) facilitates immune exclusion. The mechanisms can be summarized as follows: (1) establishing a physical barrier through the remodeling of ECM scaffold components (collagen, fibronectin, and chitosan) into denser, more organized peri-tumoral structures that impede immune cell infiltration and (2) creating a chemical barrier by modulating cytokine/chemokine secretion within the tumor microenvironment (TME), thereby suppressing chemotaxis and reducing immune cell recruitment8,9,10,11,12,13. From the perspective of the cancer-immunity cycle, the immune-excluded phenotype exhibits robust generation of tumor-specific immune cells, yet demonstrates impaired intra-tumoral infiltration. The failed penetration of immune cells into tumor nests constitutes a rate-limiting step in the antitumor immune response14,15. Therefore, enhancing T cell infiltration into tumor nests may be a promising strategy to improve immunotherapy efficacy.
Hypoxia and acidity are hallmark features of the TIME in most solid tumors, promoting immune exclusion16. The G protein-coupled receptor (GPCR) family of proton sensors, which comprises four members: GPR4, TDAG8 (GPR65), OGR1 (GPR68), and G2A (GPR132), can detect extracellular hydrogen ion concentrations through histidine residues within their extracellular domains17. Since their initial discovery in 2003, these receptors have been shown to participate in diverse biological processes, including inflammatory responses and the regulation of TIME-associated pathways17,18,19,20. Our team has focused on GPR4, which is widely recognized as an oncogenic driver that plays a significant role in tumor progression and angiogenesis21,22,23,24. Zhong et al. also found that GPR4 can activate RhoA signaling to inhibit Lats kinase phosphorylation, thereby promoting nuclear translocation and transcriptional activation of YAP, ultimately driving tumor progression25. Rogler et al. discovered that simultaneous loss of GRP4 and OGR1 functions exerts an additive effect on inhibiting colonic inflammation26. Wagne et al also found that GPR4 promotes CD4 + T helper cell infiltration and the expression of IFN-γ, iNOS, MCP-1, and CXCL2, eventually inducing mucosal inflammation, via using an inflammatory bowel disease animal model27. This research provided a novel link between tissue pH and the immune system. However, to the best of our knowledge, no study has focused on the role of GPR4 in shaping the TIME of CC.
In the present study, we find that an acidic microenvironment activates GPR4, which subsequently drives ECM remodeling via LOXL2-mediated reorganization of collagen fibrils from disordered to highly ordered spatial arrangements. This effect enhances the physical barrier properties of the ECM, ultimately impairing CD8+ T cell infiltration into tumor nests and fostering immune exclusion in the tumor microenvironment. Furthermore, we demonstrate that GPR4 regulates both LOXL2 and TGF-β through the JAK2/STAT3 signaling pathway and investigate the potential of LOXL2 inhibition to enhance immunotherapy efficacy. In summary, our findings underscore the importance of GPR4 in immune exclusion employed by CC, and suggest that inhibition of GPR4 downstream effectors can reverse immune exclusion, presenting a novel combinatorial strategy for CC patients.
Results
GPR4 is correlated with immune exclusion
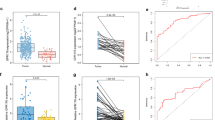
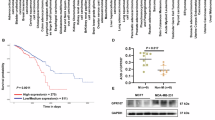
We explored TCGA-CC data to compare the mRNA expression of the GPCRs family of proton sensors between CC tissues and adjacent normal tissues. As shown in Fig. 1A, GPR4 was highly expressed in CC tissues comparing with normal tissues, and GPR65 presented a low expression in CC tissues. No significant differences were observed between GPR68 and GPR132. The IHC results demonstrated that the expression of GPR4 protein was highly regulated in CC tissues compared to adjacent non-cancerous tissues (Fig. 1B and Supplementary Fig. 1A). We then explored the relationship between GPR4 mRNA expression and clinical features, including prognosis. As shown in Fig. 1C, high GPR4 expression was significantly correlated with worse OS. Its expression was associated with poor PFS, although the statistical difference was not significant (Supplementary Fig. 1B). In addition, the expression of GPR4 was significantly related to the T stage of CC patients, and there were no relationships between GPR4 expression and age, sex, N stage, M stage, microsatellite instability (MSI) status, and treatment types (Table 1). Subsequently, we explored the immune cell infiltration levels in CC samples and found a vast difference between CC samples with high expression level of GPR4 and low expression samples (Supplementary Fig. 1C). Different algorithms for the evaluation of CD8+T cells consistently revealed that high levels of CD8+T cell infiltration were present in CC samples with high GPR4 expression (Fig. 1D). However, studies have demonstrated that the infiltration of CD8 + T cells tends to be associated with a better prognosis28,29,30,31. Therefore, we conducted a TIDE evaluation to explain this interesting point and found that the immune exclusion score in CC samples with a high expression of GPR4 was higher than that in CC samples with a low expression of GPR4 (Fig. 1E). Therefore, we speculated that GPR4 is related to immune exclusion.
A mRNA expression difference of the proton sensors (including GPR4, GPR65, GPR68, GPR132) between CC tissues and adjacent normal tissues (Wilcoxon Signed-Rank test, n = 473 in CC and n = 40 in adjacent normal). B Representative images and quantification (two-tailed t test) of GPR4 staining in CC tissues and normal tissues. Scale bars, 200 µm. C Kaplan–Meier curve analysis (Log-Rank test, n = 5 biologically independent samples) for the correlation between GPR4 and OS of TCGA-CC cohort. D Different infiltration of CD8+ T cell in CC samples between high and low expression of GPR4, based on four algorithms (Wilcoxon Signed-Rank test, n = 237 in high-GPR4 and n = 236 in low-GPR4). E Different TIDE scores between high and low expression of GPR4 (Wilcoxon Signed-Rank test, n = 237 in high-GPR4 and n = 236 in low-GPR4). F Representative mIHC images of CD8+ T cell infiltration in CC (left, Scale bar, 500 μm). pan-CK labeled cancer nest. The area within the red line indicates the tumor nest (right, Scale bar, 200 μm). G Quantification of CD8+ T cell infiltration in nest region and stroma region (n = 4 biologically independent samples, two-tailed t test, data are presented as mean ± SD). H proteins expression of TGF-β and MHC-Ⅰ in GPR4-knockdown cell line (HCT116) and GPR4-overexpression cell line (SW480). ns > 0.05.
Moreover, to explore the relationship between GPR4 and immune exclusion, we conducted mIHC for CC tissues, and pan-CK was used to label tumor nests. As shown in Fig. 1F, we observed strong co-localization of GPR4 (green fluorescence) and CK-positive regions. Although some stromal cells also showed limited GPR4 expression, the fluorescence intensity was significantly lower than that in tumor cells. Therefore, we conclude that GPR4 is predominantly expressed by tumor cells in CRC tissues. More importantly, we found that in colon cancer tissues with low GPR4 expression, CD8+ T cells were more abundant within the tumor nests, showing no significant difference in quantity compared to the stromal regions. In contrast, colon cancer tissues with high GPR4 expression exhibited relatively fewer CD8+ T cells in tumor nests, with a more pronounced disparity in distribution between the nests and stromal areas. Quantitative analysis of CD8+ T cells revealed that specimens from patients with relatively high GPR4 expression had significantly lower intratumoral CD8 + T cells infiltration than those with low GPR4 expression. However, no notable difference was observed in the stromal CD8+ T cell infiltration between the high and low GPR4 expression groups (Fig. 1G). These findings indicate that elevated GPR4 expression in colon cancer reduces intratumoral CD8+ T cell infiltration, consistent with the hallmark feature of immune exclusion6.
Melanie et al. also identified two hallmarks about molecular character: (1) Impaired antigen presentation, particularly the downregulation of MHC class I molecules; (2) Upregulation of the TGF-β7. To verify these hallmarks, we first detected the endogenous expression of GPR4 in five human established CC lines and discovered that the highest expression of GPR4 was in the HCT116 cell line, and the lowest expression was in the SW480 cell line (Supplementary Fig. 1D). We then successfully built a GPR4-knockdown cell model based on the HCT116 cell line using sh-GPR4 lentivirus and constructed a GPR4-overexpression cell line based on the SW480 cell line via OE-GPR4 lentivirus (Supplementary Fig. 1E). Subsequently, we found that in SW480 cells, GPR4 overexpression significantly downregulated MHC class I protein expression while markedly increasing the TGF-β levels. Conversely, GPR4 knockdown in HCT116 cells led to elevated MHC class Ⅰ expression and reduced TGF-β levels (Fig. 1H). As antigen presentation and TGF-β are mechanistically linked, we adopted SB-431542 as a TGF-β inhibitor. As shown in Supplementary Fig. 1F, GPR4 overexpression significantly downregulated MHC-Ⅰ protein expression, whereas SB-431542 reversed the major inhibition of MHC-Ⅰ. The flow cytometry results were consistent with the WB results (Supplementary Fig. 1G). We found that the Mean Fluorescence Intensity (MFI) of MHC-Ⅰ in OE-NC cells was higher than that in OE-GPR4 cells. When combined with the TGF-β inhibitor SB431542, the MFI level of OE-GPR4 cells increased. These results proved that the effect induced by GPR4 on antigen presentation is mainly dependent on the TGF–β signaling pathway. These results demonstrate that GPR4 expression suppresses tumor antigen presentation and activates the TGF-β pathway, which aligns with the established molecular signature of immune exclusion reported in the literature. Therefore, GPR4 is related to immune exclusion in CC.
GPR4 induces immune exclusion via remodeling ECM
The occurrence of tumor immune exclusion is primarily associated with physical barriers formed by aberrant ECM composition or reconstruction, as well as chemical barriers resulting from dysregulated cytokine secretion9. In light of the previously mentioned relationship between GPR4 and immune exclusion, we conducted a comprehensive investigation of the biological functions of GPR4 using the Genebridge approach across all human tissues. Our findings indicate that the biological functions of GPR4 not only align with established pathways, such as angiogenesis and G protein-coupled receptor signaling, but also exhibit significant relevance to extracellular matrix organization and ECM structural remodeling (Supplementary Fig. 2A). When focusing specifically on colon tissue, we observed that GPR4’s functional roles extend beyond ECM organization and structural remodeling to encompass collagen fibril formation, collagen degradation, and collagen trimers (Fig. 2A). Gene Ontology (GO) analysis of sequencing data from TCGA-CC patients revealed consistent functional patterns (Supplementary Fig. 2B). Within the Biological Process (BP) category, GPR4 demonstrated strong associations with ECM organization, ECM structural remodeling, and collagen fibril assembly. In terms of Molecular Function (MF), GPR4 exhibited significant enrichment in extracellular collagen structural organization, integrin binding, and collagen extracellular matrix binding activities. Subsequent RNA sequencing analysis of the constructed GPR4-overexpressing SW480 cell lines revealed significantly elevated transcriptional levels of GPR4, with substantial differentially expressed genes (DEGs) identified between the high-expression and control groups (Supplementary Fig. 2C). GO functional enrichment analysis of these DEGs demonstrated that GPR4 was strongly associated with ECM organization and structural remodeling (Fig. 2B). These findings were consistent with the results obtained from the TCGA database analysis. Consequently, we hypothesize that GPR4 induces immune exclusion by promoting physical barriers within the ECM.
A Biological functions of GPR4 via Genebridge approach across colon tissue. B GO functional enrichment analysis of GPR4 related DEGs (Hypergeometric test). C CD4+ T cells in vitro migration activity was impaired by decellularized ECM from GPR4-overexpression SW480 tumor cells (two-tailed t test, n = 3 biologically independent samples). D CD8+ T cells in vitro migration activity was impaired by decellularized ECM from GPR4-overexpression SW480 tumor cells (two-tailed t test, n = 5 biologically independent samples). E CD8+ T cell horizontal migration was impaired by GPR4-overexpression SW480 tumor cells via collagen gel-based model. CD8+ T cell was labeled by Hochest 33342, and SW480 cell mass was labeled by Hochest 33342 and GFP simultaneously. Green circle showing tumor cells mass, and blue circle showing some CD8+ T cells infiltrated toward tumor cells. Rectangle in the third columns showing the position of magnified images in the fourth columns. n = 3. Scale bars 855.1 μm. F The effect and representative images of GPR4 knockdown MC38 tumor growth was evaluated in BALB/c-nude mice (two-way ANOVA, n = 5 mice per group). Scale bars, 1 cm. G The effect and representative images of GPR4 knockdown MC38 tumor growth was evaluated in C57BL/6 J mice (two-way ANOVA, n = 5 mice per group). Scale bars, 1 cm. H Difference of tumor inhibition rate for GPR4 knockdown MC38 cells relative to control cells between inoculation into C57BL/6 J or BALB/c-nude mice (two-tailed t test, n = 5). I The GPR4 knockdown group exhibited increased number of CD4 + T cells (normalized for tumor weight) compared to the NC group (two-tailed t test, n = 5). J The difference of positive proportion of Ki-67 + CD4 + T cells between two groups (two-tailed t test, n = 5). K The GPR4 knockdown group exhibited increased number of CD8+ T cells (normalized for tumor weight) compared to the NC group (two-tailed t test, n = 5). L The difference of positive proportion of Ki-67 + CD8+ T cells between two groups (two-tailed t test, n = 5). M–O Comparion of T cell subset proportions within the total T cell population between two groups (two-tailed t test, n = 5). ns > 0.05.
To assess whether GPR4 inhibits T cell migration and infiltration, a Transwell chamber-based cell model was established (Supplementary Fig. 2D). The results indicated that, in the absence of tumor cell-derived ECM in the upper chamber (Fig. 2C, D), the addition of CCL21 in the lower chamber significantly enhanced the migration of CD8+ T cells and CD4+ T cells, confirming its efficacy as a chemoattractant. Notably, ECM formed by GPR4-overexpressing SW480 cells in the upper chamber markedly impaired the migration capability of CD8+ T cells and CD4+ T cells compared to ECM produced by control (NC) SW480 cells. Furthermore, using a collagen gel-based model to evaluate horizontal CD8+ T cell migration and infiltration (Supplementary Fig. 2E), it was observed that CD8+ T cells exhibited minimal migration in collagen gels embedded with GPR4-overexpressing SW480 cells. Microscopic examination (with the field positioned approximately 2 mm from the tumor edge) revealed no CD8+ T cell infiltration. In contrast, collagen gels containing NC SW480 cells displayed multiple CD8+ T cells migrating toward the tumor mass (Fig. 2E). The number of T cells within a range of no more than 2 mm from the edge of the tumor and the average distance between CD8+ T cells and the tumor margin were utilized for statistical analysis. As shown in Supplementary Fig. 2F and G, the number of infiltrating CD8+ T cells in collagen gel embedded with NC SW480 cells was more abundant than in collagen gel embedded with GPR4-overexpressing SW480 cells. Additionally, the average distance between CD8+ T cells and the tumor margin in the NC group was lower than in the GPR4-OE group. These findings suggest that GPR4 overexpression in tumor cells alters collagen matrix properties and structures, physically impeding CD8+ T cell migration and inducing an immune exclusion phenomenon.
To validate the impact of GPR4 on immune exclusion of CC in vivo, subcutaneous xenograft tumor mouse models were established using immunodeficient or immunocompetent mice. Knocking down GPR4 (sh-GPR4) reduced MC38 cell growth (tumor volume and tumor weight) in both immunodeficient BALB/c-nude mice and immunocompetent C57 mice (Fig. 2F and G, and Supplementary Fig. 2H, I). By comparing the tumor growth inhibition rates between immunodeficient and immunocompetent mice, we revealed that GPR4 knockdown exerted significantly obvious antitumor effects in immunocompetent mice compared to immunodeficient nude mice (86.8% vs. 28.7%, Fig. 2H). The differential antitumor efficacy of GPR4 knockdown indicates immune-dependent tumor suppression mechanisms.
Further flow cytometry analysis of tumor-infiltrating T lymphocytes in C57 immunocompetent mice revealed significant differences between the GPR4 knockdown group and the NC group (Supplementary Fig. 2J). The GPR4 knockdown group exhibited an increased number of both CD8+ and CD4+ T cells (normalized for tumor weight) compared to the NC group (Fig. 2I, K). Additionally, the proportion of Ki-67+ CD8+ T cells remained unchanged between the groups, while GPR4 knockdown tumors demonstrated a reduced percentage of Ki-67+ CD4+ T cells relative to controls, indicating that GPR4 impedes T cell infiltration rather than influencing their proliferation (Fig. 2J, L). We also compared T cell subset proportions and discovered that sh-GPR4 tumors exhibited a significantly higher percentage of CD8+ T cells and a concomitantly lower percentage of CD4+ T cells within the total T cell population compared to sh-NC controls (Fig. 2M–O), indicating that the changes in CD8+ T cells were more pronounced than those in CD4+ T cells.
Quantitative analysis (collagen volume fraction, CVF) of collagen deposition via Masson’s trichrome staining revealed significantly reduced collagen content in both the tumor margin and tumor core of GPR4 knockdown tumors compared to NC tumors (Fig. 3A, B). Moreover, assessment of CD8 + T cell infiltration demonstrated markedly increased CD8+ T cell density within the tumor core of GPR4 knockdown tumors, whereas no significant difference was observed at the tumor core and margin (Fig. 3A, C). To investigate the spatial organization of collagen fibers, we performed SHG imaging and quantitatively analyzed the collagen architectural features using CT-FIRE software for fiber pattern characterization.
A Representative images of collagen deposition via Masson’s trichrome staining and CD8+ T cell staining for tumor margin region and tumor core region of GPR4 knockdown MC38 tumor and control tumor. Rectangle in the second and fourth columns showing the position of magnified images in third and fifth columns. Scale bar: 500 µm (left) and 100 µm (right). B Quantitative analysis of collagen volume fraction (CVF) for GPR4 knockdown MC38 tumor and control tumor (two-tailed t test, n = 5 per group). C Quantitative analysis of CD8+ T cell for GPR4 knockdown MC38 tumor and control tumor (two-tailed t test, n = 5 per group). D Representative images of collagen alignment via SHG (Second harmonic generation) for GPR4 knockdown MC38 tumor and control tumor. Left: SHG imaging, Right: fiber imaging via CT-fire. Scale bar: 147.6 µm. E Quantitative analysis of collagen fiber alignment, collagen fiber width (F), and collagen fiber length (G). Two-tailed t test for all tests in this Figure (n = 5 per group). ns > 0.05.
SHG analysis revealed notable alterations in collagen architecture following GPR4 knockdown. Specifically, the GPR4 knockdown group exhibited a significantly higher coefficient of variation (CV) in the angle of collagen fibers within both the invasive margin and tumor core regions compared to the NC group, indicating a more disordered configuration and poorer fiber alignment (Fig. 3D, E). Regarding fiber width and length, an increase was observed in the tumor core of the GPR4 knockdown group compared to the NC group; however, no differences were noted in the tumor margin between these groups (Fig. 3F, G). Collectively, these findings suggest that GPR4 facilitates immune exclusion through collagen architectural reprogramming, inducing fibrillar densification and disorganized alignment that physically restricts CD8+ T cell infiltration in CC.
GPR4 re-constructs ECM collagen alignment through regulating LOXL2 expression
We have demonstrated the regulation of GPR4 for collagen deposition and collagen fibers alignment. Previous studies have demonstrated that TGF-β upregulates collagen I (COL1A1 and COL1A2) in multiple cancer types through the activation of the SMAD2/SMAD3 pathway32. To investigate the founding of GPR4 regulating TGF-β expression mentioned above in our research, we focused on the regulation of GPR4-mediated collagen architectural remodeling.
Subsequently, a collagen contraction assay was conducted to examine the impact of GPR4 expression levels on collagen remodeling in vitro. The results revealed that SW480 CC cells with GPR4 overexpression demonstrated a significant increase in collagen contraction compared to the NC group (Fig. 4A). In contrast, HCT116 CC cells with GPR4 knockdown exhibited a notable decrease in collagen compaction ability relative to their respective controls (Fig. 4B). These findings indicate that GPR4 may have a regulatory function in modulating collagen structure.
A Representative images (upper-left) and quantitative analysis of collagen contraction assay (Right) for GPR4 overexpression SW480 cells and the cartoon diagram for collagen area (left-lower). Two-tailed t test, n = 4 biologically independent samples. B Representative images (upper-left) and quantitative analysis of collagen contraction assay (Right) for GPR4 knockdown HCT116 cells and the cartoon diagram for collagen area (left-lower). Two-tailed t test, n = 4 biologically independent samples. C Heatmap showing different expression of GPR4-OE cells and NC cells. D Volcano plot for Top different expression genes between GPR4-OE cells and NC cells (Limma R). E The difference of emission fluorescence for GPR4-OE SW480 cell groups NC group (Amplex red method). Two-tailed t test, n = 4 biologically independent samples. F The difference of emission fluorescence for GPR4-KD HCT116 cell groups NC group (Amplex red method). Two-tailed t test, n = 4 biologically independent samples. G quantitative analysis of collagen contraction assay for GPR4-OE SW480 cell groups, GPR4-OE SW480 cell combined with BAPN groups, and NC group. Two-tailed t test, n = 4 biologically independent samples. H The difference of emission fluorescence for these three groups (Amplex red method). Two-tailed t test, n = 4 biologically independent samples. I Representative images and quantification of LOXL2 staining in CC tissues and normal tissues. Scale bars, 200 µm. J The correlation (Pearson test) between GPR4 and LOXL2 in CC tissues by IHC. Data are presented as scatter plots with a linear regression line (n = 8). K Representative images of NHS collagen fluorescence for rLOXL2 function on collagen alignment. Scale bars, 106.9 µm. Polarized model. L Quantitative analysis (Two-tailed t test, n = 3 biologically independent samples) for collagen fiber length, width (M), and fiber alignment (N) of NHS imaging. O Representative images (upper-left) and quantitative analysis of collagen contraction assay (Right) for rLOXL2 function and the cartoon diagram for collagen area (left-lower). Two-tailed t test, n = 3 biologically independent samples. P CD8+ T cell horizontal migration assay for GPR4-OE SW480 cell groups, GPR4-OE SW480 cell combined with BAPN groups, and NC group, via collagen gel-based model. n = 3. Scale bars 855.1 μm. ns > 0.05.
To further investigate the downstream effector genes, we extracted and analyzed genes associated with extracellular matrix (ECM) structure and organization using our RNA-sequencing analysis. The results revealed significant alterations in the expression of multiple ECM-related genes in the GPR4 overexpression (OE) group (Fig. 4C). Among the upregulated genes, LOXL2 exhibited notable increases, suggesting that the lysyl oxidase family (LOXs) may play a crucial role in GPR4 function (Fig. 4D). As previous shows, LOXs can catalyze the oxidative deamination of lysine and hydroxylysine residues within extracellular matrix collagen, thereby facilitating collagen cross-linking33. Alba et al demonstrated that in an animal model of clear cell renal cell carcinoma, inhibition of LOX reduced tumor stiffness and resulted in disorganized collagen fiber alignment13. Therefor we explored the function of LOXs in CC cells. As shown in Fig. 4E, the emission fluorescence of GPR4-OE SW480 cell groups was higher than in the GPR4-NC SW480 group (Amplex Red method). Conversely, the emission fluorescence of the GPR4-sh HCT116 group was lower than in the GPR4-NC HCT116 group (Fig. 4F). These results suggest that GPR4 in CC cells modulates the catalytic activity of LOXs in collagen cross-linking. To determine whether LOXs mediate the effect of GPR4 on collagen contraction, we employed BAPN, a pan-LOX inhibitor. The results indicated that GPR4-OE SW480 colon cancer cells significantly enhanced collagen contraction compared to the NC cells. However, when BAPN was added to the GPR4-overexpressing SW480 cells, the pro-collagen compaction effect was significantly suppressed (Supplementary Fig. 3A and Fig. 4G). Moreover, we performed an Amplex Red assay. Consistent with the mentioned findings, the GPR4-overexpressing group exhibited a significantly stronger fluorescent signal compared to the NC group, indicating elevated LOXs activity. Co-treatment with BAPN in the GPR4-overexpressing cells led to a marked reduction in fluorescence, further confirming the dependency of GPR4-enhanced LOXs activity on its catalytic function (Fig. 4H).
We explored the mRNA expression of LOXL proteins in cervical cancer (CC) tissues and normal tissues using TCGA-CC data. In Supplementary Fig. 3B, LOX, LOXL1, and LOXL2 were found to be higher in CC tissues, while LOXL4 was lower. A correlation analysis showed a strong positive link between GPR4 expression and LOXs, including LOX (r = 0.71), LOXL1 (r = 0.52), LOXL2 (r = 0.76), and LOXL3 (r = 0.69) (Supplementary Fig. 3C). LOXL2 not only has high levels in colorectal cancer but also has the strongest positive link with GPR4 expression among all LOX family members. To confirm the different expression of LOXL2 and its link with GPR4, we did IHC analyses using public databases and clinical samples from our institution. We found that LOXL2 was much higher in colorectal cancer tissues compared to adjacent normal tissues (Fig. 4I and Supplementary Fig. 3D). Also, GPR4 expression was strongly linked with LOXL2 expression (Fig. 4J). We also checked our findings at the cell level. Western blotting showed that in GPR4-overexpressing SW480 CC cells, LOXL2 and COL1A1 levels were much higher than in NC cells. On the other hand, reducing GPR4 in HCT116 cells led to a reduction in LOXL2 and COL1A1 levels, with no significantly change in LOX levels (Supplementary Fig. 3E).
Building upon our finding that GPR4 promotes LOXL2 expression, we further investigated whether the LOXL2 protein exerts a remodeling effect on collagen fiber organization by employing exogenous recombinant LOXL2 protein. As demonstrated in Fig. 4K, L, compared to the PBS control group, the rLOXL2-treated group exhibited a more organized spatial arrangement of collagen fibers, with a significantly reduced CV value of angular and distinct directional alignment (as indicated by arrow directions in the figure). Regarding collagen fiber morphology, the rLOXL2-treated group showed significantly greater fiber length compared to the PBS group, while no statistically significant difference in fiber width was observed between the two groups (Fig. 4L–N). In addition to these findings, when compared with the PBS control group, the rLOXL2-treated group exhibited significantly enhanced collagen contraction (Fig. 4O). Furthermore, during the catalytic process using collagen as a substrate, the rLOXL2 group generated higher levels of H2O2—one of the terminal products—resulting in stronger fluorescence intensity (Supplementary Fig. 3F). In our T-cell horizontal migration assay (Fig. 4P and Supplementary Fig. 3G, H), collagen gel-embedded NC group cells exhibited more CD8+ T cell infiltration and less distance between CD8+ T cells and the tumor margin. This response was abolished in the GPR4-overexpressing group but was rescued by BAPN inhibition. The rescue experiment of CD8+ T cell migration upon rLOXL2 protein confirmed that GPR4 impaired the migration of CD8+ T cells in collagen gels via regulating LOXL2 expression (Supplementary Fig. 3I–K). Collectively, these results indicate that GPR4-mediated LOXL2 expression drives ECM collagen reorganization, which functions as a physical barrier to CD8+ T cell migration, consequently facilitating tumor immune escape.
TGF-β is a well-established master regulator of collagen deposition in cancer. Previous studies demonstrated that it regulates the expression of LOXs in liver, kidney, heart, leading related diseases34,35,36. In light of these considerations, we further examined whether the influence of the GPR4-LOXL2 pathway on the extracellular matrix (ECM) is contingent upon the TGF-β pathway. To this end, we employed the TGF-β inhibitor SB-431542. The results of the Western blot analysis indicated that SB-431542 effectively inhibited the TGF-β signaling pathway and significantly decreased LOXL2 expression in SW480 NC cells (Supplementary Fig. 3L). However, GPR4-OE SW480 cells, although the inhibition of the TGF-β pathway also led to a reduction in LOXL2 expression, this effect was not obvious, and LOXL2 expression remained substantially higher than in the SW480 NC group. These findings suggest that the regulation of LOXL2 protein expression by GPR4 is not predominantly reliant on the TGF-β pathway but is instead mediated by other pathway. Furthermore, the results of the Amplex Red assay indicated that inhibition of the TGF-β pathway partially mitigated the increased collagen catalytic activity of LOXs induced by GPR4 overexpression; however, this mitigation was not statistically significant (Supplementary Fig. 3M). Additionally, collagen contraction assays revealed that inhibiting the TGF-β pathway did not counteract the enhanced collagen contraction capability of SW480 cells following GPR4 overexpression (Supplementary Fig. 3N, O). These findings align with the Western blot data and further substantiate the conclusion that GPR4-mediated extracellular matrix remodeling via LOXL2 operates independently of the TGF-β pathway.
In subsequent in vivo experiments, we utilized immunocompetent C57BL/6 J mice to ascertain whether GPR4 facilitates immune rejection via LOXL2. As illustrated in Fig. 5A–C, the overexpression of GPR4 enhanced the growth of MC38 cells, as evidenced by increased tumor volume and weight in immunocompetent mice. Conversely, treatment with BAPN significantly inhibited the growth of MC38 cells, indicating that the BAPN inhibitor effectively mitigates the tumor-promoting effects induced by GPR4 overexpression. Masson staining revealed an elevation in collagen content within GPR4-overexpressing tumors, both at the tumor margin and core, while BAPN treatment reversed this effect (Fig. 5D, E). Quantification of CD8+ T cell infiltration demonstrated that the GPR4 overexpression group exhibited the lowest CD8+ T cell density in the tumor core, significantly reduced compared to the control group. The GPR4 overexpression + BAPN group displayed intermediate infiltration levels, lower than the control but higher than the GPR4 overexpression group. In the tumor margin, CD8+ T cell numbers were similarly minimal in the GPR4 overexpression group, whereas BAPN treatment significantly restored infiltration compared to GPR4 overexpression alone (Fig. 5D, F). SHG experiments revealed a significant reduction in the CV of angle in collagen fiber orientation in the GPR4 overexpression group compared to controls, observed in both the tumor core and margin. This improved alignment of collagen fibers was partially rescued by BAPN treatment, with the GPR4 overexpression + BAPN group exhibiting increased CV compared to GPR4 overexpression alone (Fig. 5G, H). Although collagen fibers in the tumor margin were consistently thicker than those in the core region across all groups, no intergroup differences reached statistical significance (Fig. 5G, I). Additionally, GPR4 overexpression significantly promoted collagen fiber length compared to controls, an effect attenuated by BAPN (Fig. 5G, J). These findings demonstrate that GPR4 regulates the reconstruction of extracellular collagen through LOXL2 in vivo, thereby contributing to immune exclusion.
A The effect and representative images of GPR4-OE MC38 cell groups, GPR4-OE MC38 cell combined with BAPN groups, and NC group tumor growth were evaluated in C57BL/6 J mice. Scale bars, 1 cm. B The difference of tumor volume for these three groups (two-way ANOVA, n = 5 per group). C The difference of tumor weight for these three groups (two-tailed t test, n = 5 per group). D Representative images of collagen deposition via Masson’s trichrome staining and CD8+ T cell staining for tumor margin region and tumor core region of GPR4 -OE MC38 tumor, GPR4 -OE combined with BAPN MC38 tumor and control tumor. Scale bar: 500 µm (left) and 100 µm (right). Rectangle in the second and fourth columns showing the position of magnified images in third and fifth columns. E Quantitative analysis of CVF for these three groups (two-tailed t test, n = 5 per group). F Quantitative analysis of CD8+ T cell for these three groups (two-tailed t test, n = 5 per group). G Representative images of collagen alignment via SHG for GPR4 -OE MC38 tumor, GPR4 -OE combined with BAPN MC38 tumor and control tumor. Scale bar: 147.6 µm. Left: SHG imaging, Right: fiber imaging via CT-fire. H Quantitative analysis of collagen fiber alignment, collagen fiber width (I), and collagen fiber length (J) for these three groups (two-tailed t test, n = 5 per group). ns > 0.05.
In order to investigate the potential impact of GPR4 and ECM remodeling on T cell activity, we conducted mIHC staining on tumor tissues from three groups of mice: OE-NC, OE-GPR4, and OE-GPR4 + BAPN. As illustrated in Supplementary Fig. 4A, CD8, granzyme B, and perforin were visualized as red, green, and yellow fluorescence, respectively. A statistical analysis was performed on the number of granzyme B⁺ perforin⁺ CD8⁺ T cells. As depicted in Supplementary Fig. 4B, the group with GPR4 overexpression demonstrated a reduced count of granzyme B⁺ perforin⁺ CD8⁺ T cells compared to the NC group. The count of these cells in the GPR4 overexpression + BAPN treatment group was marginally higher than in the GPR4 overexpression group; however, this difference was not statistically significant. Furthermore, statistical analysis of the proportion of granzyme B⁺ perforin⁺ CD8⁺ T cells among total CD8⁺ T cells revealed no significant differences among the three groups (Supplementary Fig. 4C). In conclusion, our findings suggest that GPR4 does not significantly affect T cell activity, and its primary function appears to be in modulating ECM and inhibiting immune cell infiltration.
JAK2/STAT3 pathway mediates the regulation between GPR4 and LOXL2
TGF-β is a well-established upstream regulator of collagens, including COL1A132. In colorectal cancer, we found that GPR4 modulates the expression of TGF-β and LOXL2, promotes collagen deposition, and remodels ECM collagen alignment, ultimately driving immune exclusion. Therefore, we further investigated how GPR4 regulated the expression of LOXL2 and TGF-β. The JAK2/STAT3 signaling pathway plays a critical role in tumorigenesis and cancer progression37. And our previous work demonstrated that GPR4 promotes both STAT3 expression and phosphorylation in ovarian cancer. So, we firstly explored that mRNA expression of GPR4 was positively correlated with a string of gene’s expression (including CCL5, CXCR4, IL6, PTGIS, SOCS1, and CXCL12)37 transcribed by STAT3 in CC tissues via TCGA-CC data, which indicted that GPR4 may be functionally related with the activation of JAK2/STAT3 pathway (Supplementary Fig. 5A).
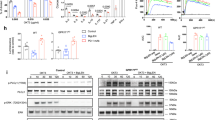
Western blotting analysis indicated that in GPR4-overexpressing SW480 cells, the total protein levels of STAT3 and JAK2 remained unchanged compared to control cells. However, there was a significant increase in phosphorylation levels at STAT3-Y705 and JAK2-Y1007/1008. Conversely, in GPR4-knockdown HCT116 cells, phosphorylation at STAT3-Y705 and JAK2-Y1007/1008 was notably suppressed. These findings suggest that GPR4 modulates JAK2/STAT3 pathway activity by influencing their phosphorylation status (Fig. 6A). Immunofluorescence experiments corroborated these results, indicating that GPR4 regulates phosphorylation at STAT3-Y705 (Fig. 6B–D). Subsequently, Stattic was employed to inhibit STAT3 phosphorylation. Western blot analysis demonstrated that in SW480 cells, GPR4 overexpression significantly elevated pSTAT3-Y705 levels compared to control cells, along with increased expression of LOXL2, COL1A1, and TGF-β. However, treatment with Stattic markedly reduced both pSTAT3-Y705 and the expression of LOXL2, COL1A1, and TGF-β in GPR4-overexpressing cells (Fig. 6E). Rescue experiments in HCT116 cells further substantiated these observations. As illustrated in Supplementary Fig. 5B, GPR4 knockdown significantly downregulated pSTAT3-Y705 levels as well as LOXL2 and COL1A1. Notably, STAT3 overexpression via plasmid transfection restored pSTAT3-Y705 levels and subsequently increased LOXL2 and COL1A1 expression compared to control cells. These results demonstrate that GPR4 regulates the expression of downstream proteins, including LOXL2, through modulation of STAT3 phosphorylation.
A GPR4 regulates JAK2 and STAT3 phosphorylation via western blotting. n = 3. Scale bar: 100 µm. B Representative IF images of STAT3-Y705 phosphorylation regulated by GPR4. C Quantitative analysis of STAT3-Y705 phosphorylation in GPR4 - OE SW480 cell line and (D) GPR4 - KD HCT116 cell lline. Data are presented as mean ± SD. Two-tailed t test, n = 3 biologically independent samples. E Static treatment reduces both pSTAT3-Y705 and the expression of LOXL2, COL1A1, and TGF-β in GPR4 - OE SW480 cells. F RT-PCR analysis for LOXL2 in SW480 cell lines after receiving Stattic treatment. Two-tailed t test, n = 3 biologically independent samples. G RT-PCR analysis for TGF-β in SW480 cell lines after receiving Stattic treatment. Data are presented as mean ± SD.Two-tailed t test, n = 3 biologically independent samples. H STAT3 transcription factor binding site. I In SW480 cells, STAT3 binds to the LOXL2 promoter region confirmed via ChIP assays. J STAT3 binds to the TGF-β promoter region confirmed via ChIP assays. K RT-PCR assesses the transcriptional activity of LOXL2 promoter regions and TGF-β promoter region L. Data are presented as mean ± SD. Two-tailed t test, n = 3 biologically independent samples. ns > 0.05.
Based on these findings, we investigated the mechanism by which STAT3 regulates LOXL2 and TGF-β expression. RT-PCR analysis revealed that Stattic-mediated STAT3 phosphorylation inhibition significantly reduced both LOXL2 and TGF-β mRNA levels in both SW480 and HCT116 cell lines (Fig. 6F, G and Supplementary Fig. 5C, D). These results suggest that STAT3 regulates LOXL2 and TGF-β expression at the transcriptional level. Subsequently, we explored whether STAT3 directly regulates LOXL2 and TGF-β transcription. As shown in Fig. 6H, we first predicted the STAT3 transcription factor binding sites using the JASPAR database (http://jaspar.genereg.net). ChIP assays confirmed that STAT3 could bind to the LOXL2 promoter region, with a specific binding site located at −1377 to −1385 bp relative to the transcription start site (Fig. 6I and Supplementary Fig. 5E). Meanwhile, STAT3 could also bind to the TGF-β promoter region, with a binding site at −611 to −619 bp upstream of the transcription start site (Fig. 6J, Supplementary Fig. 5F). In addition, using RT-PCR, we assessed the transcriptional activity of these promoter regions and found that STAT3 significantly enhanced the transcriptional efficiency of both LOXL2 and TGF-β (Fig. 6K, L and Supplementary Fig. 5G, H).
Our previous study demonstrated that GPR4 directly regulates the phosphorylation of JAK2. As a member of GPCR family, GPR4 has been widely reported to activate downstream small G proteins upon stimulation, leading to the initiation of signaling cascades such as the cAMP/PKA pathway38. However, it is noteworthy that a small subset of GPCRs, including AGTR1 and CXCR4, have been shown to directly interact with JAK2 and promote its phosphorylation39,40. Based on these findings, we hypothesized that GPR4 might also physically associate with JAK2. To test this hypothesis, we performed co-immunoprecipitation (Co-IP) experiments, and found a endogenous interaction between GPR4 and JAK2, which was observed in SW480 and HCT116 cells (Fig. 7A, B). To further validate the exogenous binding between GPR4 and JAK2, we performed co-IP assays in HEK293T cells co-transfected with HA-tagged JAK2 and FLAG-tagged GPR4 overexpression plasmids. As demonstrated in Fig. 7C, D, exogenous binding between GPR4 and JAK2 was clearly detected. Furthermore, we used IF to examine the subcellular localization of GPR4 and JAK2 in CC cells. Strikingly, both HCT116 and SW480 cells exhibited significant colocalization of GPR4 and JAK2 (Fig. 7E), with Pearson’s correlation coefficients of 0.8 and 0.66, respectively (Fig. 7F, G). These results further support the Co-IP results and reinforce the likelihood of direct binding between these two molecules.
A Co-immunoprecipitation (Co-IP) experiments shows an endogenous interaction between GPR4 and JAK2 in SW480 cells and B HCT116 cells. n = 3. C, D Co-IP experiments shows an exogenous interaction between GPR4 and JAK2 in HEK293T cells. n = 3. E Representative IF images of colocalization between GPR4 and JAK2. n = 3. Scale bar: 19.4 µm. F Pearson’s correlation coefficients of colocalization in HCT116 cells (Pearson test) and G SW480 cells (Pearson test). H Full-length HA-JAK2, HA-tagged JAK2 FERM-SH2 domain (amino acids 1-510), HA-tagged JAK2 JH1-JH2 domain (amino acids 540-1124) and empty vector plasmid co-transfected with Full-length FLAG-tagged GPR4 plasmid in HEK293T cells, Co-IP assay was performed to verify their interaction. I Full-length Flag-GPR4 plasmid, the extracellular and 7-transmembrane domain (Flag-GPR4-1-290aa) plasmid, the intracellular C-terminal domain (Flag-GPR4-290-362aa) plasmid, or the extracellular, 7-transmembrane, partial intracellular hydrophobic segment (Flag-GPR4-1-330aa) plasmid, and empty vector plasmid co-transfected with Full-length HA-tagged JAK2 plasmid in HEK293T cells, Co-IP assay was performed to verify their interaction.
To elucidate the specific interaction domains between GPR4 and JAK2, we constructed full-length and truncated overexpression plasmids based on literature-reported functional domains of JAK2. To map the specific interaction domains between GPR4 and JAK2, we performed a series of Co-IP experiments using domain-specific truncation constructs. HEK293T cells were co-transfected with full-length FLAG-tagged GPR4 together with either: (1) full-length HA-JAK2, (2) HA-tagged JAK2 FERM-SH2 domain (amino acids 1-510), or (3) HA-tagged JAK2 JH1-JH2 domain (amino acids 540-1124). Subsequent IP using anti-FLAG antibody followed by immunoblotting revealed that GPR4 specifically interacted with both full-length JAK2 and its FERM-SH2 domain, but not with the JH1-JH2 region (Fig. 7H). These results demonstrate that the FERM-SH2 domain of JAK2 is necessary for its binding to GPR4.
Moreover, based on the spatial architecture of GPR4, we generated a series of truncated overexpression plasmids and the full-length FLAG-tagged GPR4 expression plasmid. By co-transfecting HEK293T cells with the full-length HA-JAK2 plasmid along with either the full-length Flag-GPR4 plasmid, the extracellular and 7-transmembrane domain (Flag-GPR4-1-290aa) plasmid, the intracellular C-terminal domain (Flag-GPR4-290-362aa) plasmid, or the extracellular, 7-transmembrane, and partial intracellular hydrophobic segment (Flag-GPR4-1-330aa) plasmid, we performed Co-IP to pull down HA-JAK2. The results demonstrated that both the 290-362aa and 1-330aa regions of GPR4 could interact with JAK2 (Fig. 7I). These findings suggest that the binding between GPR4 and JAK2 depends on the intracellular domain of GPR4, particularly the 290-330aa region.
Acidic environment promotes the activation of JAK2/STAT3 pathway induced by GPR4
Collectively, we demonstrated the detailed regulatory mechanism of GPR4 on JAK2/STAT3, LOXL2, and TGF-β. GPR4, a member of the hydrogen ion-sensing receptor family, can detect acidic environments through histidine residues in its extracellular domain, triggering changes in downstream signaling pathways18. Therefore, we investigated whether GPR4 activation in an acidic environment promotes activation of the JAK2/STAT3 signaling pathway.
HCT116 and SW480 cells were treated with acidic culture medium to simulate acidic stimulation. Western blotting results demonstrated that an acidic environment significantly upregulated the phosphorylation of JAK2 at tyrosine residues 1007/1008 and STAT3 at tyrosine residue 705, thereby activating the JAK2/STAT3 signaling pathway. Moreover, prolonged acidic stimulation (0–60 min) progressively enhanced the phosphorylation and activation of JAK2 and STAT3 (Fig. 8A, B), suggesting a time-dependent increase in the activation of the JAK2/STAT3 pathway. Furthermore, we found that under neutral pH conditions, GPR4 knockdown suppressed the phosphorylation of JAK2 and STAT3. When GPR4-knockdown HCT116 cells were stimulated with an acidic medium, no obvious increase in JAK2 and STAT3 phosphorylation was observed. Conversely, GPR4 overexpression in SW480 cells enhanced JAK2 and STAT3 phosphorylation at neutral pH. Moreover, acidic stimulation of GPR4-overexpressing SW480 cells significantly elevated the phosphorylation levels of JAK2 and STAT3 (Fig. 8C, D). Therefore, we concluded that an acidic microenvironment promotes JAK2 and STAT3 phosphorylation by activating GPR4. To validate the western blotting results, IF assays were performed on the same samples. Acidic stimulation significantly upregulated STAT3-Y705 expression in HCT116 cells without GPR4 knockdown (Fig. 8E, F). In contrast, it failed to significantly enhance STAT3-Y705 expression in GPR4-knockdown cells (Fig. 8G). Furthermore, acidic conditions promoted STAT3-Y705 expression in SW480 cells without GPR4 overexpression, and this stimulatory effect was markedly amplified in GPR4-overexpressing SW480 cells (Fig. 8H–J).
A The dynamic changes with time for protein level of JAK2-Y1007/1008 and STAT3-Y705 stimulated by acidic environment in SW480 cells and B HCT116 cells. C The protein level of JAK2-Y1007/1008 and STAT3-Y705 in GPR4-KD HCT116 cells stimulated by acidic environment. D The protein level of JAK2-Y1007/1008 and STAT3-Y705 in GPR4-OE SW480 cells stimulated by acidic environment. E–G Representative IF images and quantitative analysis of TAT3-Y705 expression in HCT116 stimulated by acidic environment. Data are presented as mean ± SD. Scale bars, 100 µm. Two-tailed t test, n = 3 biologically independent samples. H–J Representative IF images and quantitative analysis of TAT3-Y705 expression in SW480 stimulated by acidic environment. Scale bars, 100 µm. Data are presented as mean ± SD. Two-tailed t test, n = 3 biologically independent samples. ns > 0.05.
Targeting LOXL2 function or JAK2/STAT3 pathway sensitizes CRC to ICI therapy
Our results demonstrate that GPR4 directly regulates the JAK2/STAT3 pathway, which promotes the transcriptional expression of LOXL2 and TGF-β to remodel collagen components in the extracellular matrix, ultimately facilitating immune evasion in colon cancer. Furthermore, previous studies have revealed that STAT3 activation can upregulate PD-L1 expression in tumor cells, thereby suppressing the cytotoxic activity of CTLs41. Therefore, for colon cancers with GPR4 high expression exhibiting immune-excluded characteristics, a dual-pronged therapeutic strategy may have superior antitumor efficacy: (1) enhancing intratumoral infiltration of CD8+ T lymphocytes by inhibiting LOXL2 activity or JAK2 signaling, and (2) augmenting CD8+ T cell-mediated cytotoxicity through PD1/PD-L1 blockade.
Utilizing MC38 cells and C57BL/6 mice, we developed a subcutaneous xenograft model to assess the therapeutic efficacy of combining LOXL2 inhibition or JAK2 inhibition with immunotherapy in GPR4-overexpressing colon cancer. The administration schedule and dosages of the LOXL2 inhibitor (me-Hyd), anti-PD-L1 antibody, and JAK2 inhibitor (Fedratinib) are depicted in Fig. 9A. Comparative analysis of tumor volume and weight revealed the following: (1) Both the NC + PDL1 and OE-GPR4 + PDL1 groups exhibited significantly slower tumor growth and reduced weight compared to their respective NC and OE-GPR4 control groups, demonstrating that anti-PD-L1 monotherapy effectively suppresses colon cancer progression in immunocompetent C57BL/6 mice; (2) The OE-GPR4+Fedratinib group showed markedly inhibited tumor growth and decreased weight relative to the OE-GPR4 group, indicating the therapeutic potential of JAK2 inhibition against GPR4-high tumors; (3) Notably, both the OE-GPR4 + PDL1+Fedratinib and OE-GPR4 + PDL1+me-Hyd combination therapy groups displayed substantially greater tumor suppression compared to either OE-GPR4 controls or the OE-GPR4 + PDL1 monotherapy group, suggesting that dual blockade of PD-L1 with either LOXL2 or JAK2 inhibition generates superior antitumor effects against GPR4-overexpressing colon cancers (Fig. 9B, C).
A Diagram illustrating the construction and intervention of the CC tumor in C57BL/6 J mice model. B, C The effect and representative images of GPR4-OE MC38 cell group, GPR4-OE MC38 cell combined with Fedratinib group, GPR4-OE MC38 cell combined with anti PD-L1 group, GPR4-OE MC38 cell combined with me-Hyd and anti PD-L1 group, GPR4-OE MC38 cell combined with Fedratinib and anti PD-L1 group and NC group tumor growth were evaluated in C57/BJ6 mice (two-tailed t test, n = 5 per group). Scale bars, 1 cm. D, E Representative images and quantitative analysis (two-tailed t test, n = 4 per group) of PD-L1 staining. Scale bars, 500 µm and 100 µm. F–J Representative images and quantitative analysis of CD8+ T cell staining and Masson’s trichrome staining. Scale bars, 500 µm and 100 µm. Rectangle in the second and fourth columns showing the position of magnified images in third and fifth columns. Two-tailed t test, n = 4 per group. H–O Representative images and quantitative analysis of SHG images. Scale bar: 147.6 µm. Two-tailed t test, n = 4 per group. ns > 0.05.
Analysis of PD-L1 IHC results (Fig. 9D) yielded three key findings: (1) The NC + PD-L1 and OE-GPR4 + PD-L1 groups exhibited significantly lower IHC scores compared to their respective NC and OE-GPR4 control groups, thereby confirming the efficacy of anti-PD-L1 antibody treatment in reducing PD-L1 protein expression in colon cancer cells; (2) The OE-GPR4 group demonstrated the highest levels of PD-L1 expression, which were significantly elevated relative to the NC group, indicating a positive correlation between GPR4 expression and PD-L1 levels. This observation, in conjunction with previous studies that have shown STAT3-mediated transcriptional activation of PD-L1, suggests that GPR4 enhances PD-L1 expression in colon cancer; (3) PD-L1 expression in the OE-GPR4+Fedratinib group was significantly reduced compared to the OE-GPR4 group, illustrating that JAK2 inhibition effectively suppresses GPR4-induced upregulation of PD-L1 (Fig. 9E).
Representative images of Masson’s staining and CD8 IHC are shown in Fig. 9F. Quantitative analysis of CD8+ T cell infiltration in the tumor core region (Fig. 9G, H) demonstrated that (1) The OE-GPR4 group exhibited significantly fewer intra-tumoral CD8+ T cells than the NC group; (2) Among all OE-GPR4 tumors, the OE-GPR4 + PD-L1+me-Hyd group showed the highest CD8+ T cell infiltration, exceeding that of the OE-GPR4 + PD-L1 group, indicating that LOXL2 inhibitor synergizes with PD-L1 blockade to markedly enhance T cell recruitment and may offer superior therapeutic efficacy over anti-PD-L1 monotherapy; (3) While both OE-GPR4+Fedratinib and OE-GPR4 + PD-L1+Fedratinib groups showed increased CD8+ T cell infiltration relative to the OE-GPR4 group, the combination of Fedratinib with anti-PD-L1 (OE-GPR4 + PD-L1+Fedratinib) did not achieve statistically significant improvement over PD-L1 blockade alone (OE-GPR4 + PD-L1). This suggests that JAK2 inhibition may be less effective than LOXL2 inhibition in promoting CD8+ T cell trafficking into GPR4-high tumors. Analysis of the tumor margin revealed consistent CD8+ T cell infiltration patterns with the core region: (1) The OE-GPR4 group showed significantly reduced CD8+ T cell numbers compared to NC controls; (2) The OE-GPR4 + PD-L1+me-Hyd combination therapy group demonstrated enhanced CD8+ T cell accumulation relative to the OE-GPR4 group. This spatial consistency suggests that LOXL2 or JAK2 inhibitors effectively attenuate GPR4-mediated immune exclusion.
Quantitative analysis of collagen content via Masson’s staining revealed (Fig. 9I, J): (1) The collagen volume fraction (CVF) was significantly higher in OE-GPR4 tumors compared to NC controls across both margin and core regions; (2) Both combination therapy groups (OE-GPR4 + PD-L1+me-Hyd and OE-GPR4 + PD-L1+Fedratinib) exhibited reduced CVF values relative to the OE-GPR4 group. Notably, the OE-GPR4 + PD-L1+Fedratinib group showed further collagen reduction compared to PD-L1 monotherapy (OE-GPR4 + PD-L1 group), suggesting JAK2 inhibition may exert additional extracellular matrix remodeling effects.
SHG imaging was employed to analyze the spatial organization of collagen fibers (Fig. 9K). Quantitative comparison of collagen angular variance (CV) revealed the following findings in tumor core regions (Fig. 9L) (1) The OE-GPR4 group exhibited significantly lower CV values than NC controls; (2) Neither NC + PD-L1 nor OE-GPR4 + PD-L1 groups showed significant CV changes compared to their respective controls, demonstrating that PD-L1 blockade alone did not alter collagen architecture; (3) JAK2 inhibition (OE-GPR4+Fedratinib) increased CV values versus OE-GPR4, suggesting effective disruption of ECM organization; (4) both combination therapies (OE-GPR4 + PD-L1+Fedratinib and OE-GPR4 + PD-L1+me-Hyd) substantially elevated CV values compared to OE-GPR4 and OE-GPR4 + PD-L1 groups, confirming that both LOXL2 and JAK2 inhibitors promote collagen disorganization. These patterns were consistently observed in tumor margin (Fig. 9M). Meanwhile, analysis of fiber length (Fig. 9N, O) showed: (1) in tumor cores, only the LOXL2 inhibitor combination (OE-GPR4 + PD-L1+me-Hyd) significantly reduced fiber length compared to OE-GPR4 controls; (2) in margins, both therapeutic combinations (OE-GPR4 + PD-L1+me-Hyd and OE-GPR4 + PD-L1+Fedratinib) demonstrated fiber shortening relative to OE-GPR4.
In conclusion, for GPR4-high colon cancers, the combination of LOXL2 and JAK2 inhibitors with anti-PD-L1 immunotherapy synergistically remodeled the extracellular matrix collagen architecture, thereby enhancing the intra-tumoral infiltration of CD8 + T cells. This approach demonstrates superior therapeutic efficacy compared to PD-L1 monotherapy.
Discussion
GPR4 is widely acknowledged as an oncogene that significantly contributes to tumor progression and angiogenesis. It is highly expressed in various cancers, including ovarian and colorectal cancer, and is negatively correlated with patient prognosis25,42,43,44. Mechanistically, GPR4 enhances the expression and secretion of IL-6, IL-8, and vascular endothelial growth factor (VEGF-A) via a p38-dependent pathway, thereby promoting angiogenesis and tumor proliferation45,46. Furthermore, research by Professor Ehata et al. has demonstrated that GPR4 substantially facilitates lymphoma metastasis by regulating the expression of vascular cell adhesion molecule 1 (VCAM1)47. However, to the best of our knowledge, the role of GPR4 in the TIME remains unexplored. In the present study, we have demonstrated for the first time that in colorectal cancer cells, increased GPR4 expression is associated with poor prognosis and suppresses the intra-tumoral infiltration of CD8+ T lymphocytes, thereby promoting immune exclusion. The immune-excluded phenotype in tumors is associated with poor patient prognosis and diminished efficacy of immune checkpoint inhibitors48,49,50. Its most prominent feature is the impaired intra-tumoral infiltration of CD8+ T lymphocytes. Based on this finding, we investigated the detailed mechanism by which GPR4 contributes to immune exclusion.
Our investigation identified two principal mechanisms by which GPR4 induces immune exclusion. Firstly, GPR4 facilitates immune exclusion by enhancing the expression and deposition of collagen I through the upregulation of TGF-β expression. Collagen serves as the primary structural component of ECM, accounting for approximately 90% of its total mass. At the molecular level, collagen is composed of three left-handed α-polypeptide chains that are stabilized by interchain hydrogen bonds and subsequently assemble into a characteristic right-handed triple-helical conformation51. Among the 28 identified collagen subtypes, type I collagen demonstrates the highest abundance in tumor stroma and serves as the fundamental architectural scaffold. Multiple malignancies, including ovarian cancer, gastric adenocarcinoma, prostate cancer, and colorectal carcinoma, exhibit significantly elevated expression of collagen I. Importantly, higher collagen I expression consistently correlate with poorer clinical outcomes in these cancer types52,53,54,55. The accumulation of type I collagen in the tumor stroma actively promotes immune evasion by physically and functionally restricting antitumor immunity. As a major driver of immune exclusion, excessive type I collagen deposition establishes a fibrotic barrier that impedes T cell migration and intra-tumor infiltration, thereby suppressing effective anti-tumor immune response9,56. The distribution of tumor collagen exhibits significant heterogeneity. However, a general pattern emerges wherein collagen density increases toward the tumor periphery and decreases toward the central region57. Moreover, study in pancreatic cancer revealed that denser collagen networks impede T-cell infiltration into the tumor nest, instead of redirecting their migration toward the stromal compartment. And this localization of T cells is ultimately associated with poorer clinical outcomes in pancreatic cancer patients58. This phenomenon, wherein increased collagen deposition promotes immune exclusion, has been observed not only in pancreatic cancer but has also been consistently demonstrated across multiple tumor types, including breast cancer, ovarian cancer, and cholangiocarcinoma13. Additional we found that TGF-β is an essential role in the process of GPR4 induced collagen deposition. Previous studies have established that elevated collagen expression is closely associated with increased TGF-β levels59. Across multiple tumor types, TGF-β signaling activates its cognate receptors, leading to phosphorylation and subsequent formation of the SMAD2/SMAD3 complex. This activated complex then associates with SMAD4 to form a heterotrimeric complex, which translocates to the nucleus. Within the nuclear compartment, this complex interacts with specific transcription factors to assemble a functional transcriptional machinery that directly upregulates the expression of type I collagen genes (COL1A1 and COL1A2)32.
In the second mechanism, GPR4 drives immune exclusion by orchestrating ECM remodeling through LOXL2 upregulation. This process induces a spatially ordered alignment of collagen fibers, promoting a physical barrier that impedes CD8+ T lymphocyte infiltration into tumor nests, thereby promoting immune exclusion. The re-construction of ECM collagen fibers impairs the infiltration of CD8+ T lymphocytes. Sun et al found that DDR1, a canonical collagen receptor, plays a pivotal role in breast cancer progression by orchestrating ECM reorganization. Upon binding to collagen within the ECM, DDR1 drives the spatial rearrangement of collagen fibers, transforming scattered and disorganized fibrils into densely packed, aligned structures with uniform orientation. Notably, this remodeling function is exclusively dependent on DDR1’s extracellular domain, while its intracellular domain is dispensable for this process12. Weeraratna et al found that hyaluronic and proteoglycan link protein 1(HAPLN1) is lost in aged skin fibroblasts, resulting in a more aligned ECM collagen fibers that eventually promoted metastasis of melanoma cells. Interestingly, aged skin fibroblasts present an opposite phenomenon on the migration of T cells, via inhibiting motility60. Research has found that this is not only related to multiple proteases secreted by tumor cells, such as MMPs, but also to different migration characteristics between immune cells and tumor cells. In vitro 3D culture models have demonstrated that T cell migration is significantly slower through dense collagen matrices than through loose collagen matrices, with this reduced motility correlating with decreased matrix pore size; however, increased collagen density does not impair tumor cell migration and invasion61.
Moreover, we found that LOXL2 mediated immune exclusion induced by GPR4 in CC. In our study, LOXL2 was significantly overexpressed and functionally contributed to collagen alignment, thereby promoting the spatial reorganization of collagen fibers into more ordered and directional architectures. LOXL2, is a member of the lysyl oxidase (LOX) family, and study demonstrated that LOX family can catalyze the oxidative deamination of lysine and hydroxylysine residues within extracellular matrix collagen. This modification initiates covalent crosslinking between collagen fibrils, leading to the promotion of collagen fiber alignment, critically regulating the tensile strength and structural integrity of connective tissues33. Donnadieu et al. demonstrated that in multiple preclinical models of clear cell renal cell carcinoma, pharmacological inhibition of LOX enzyme activity (BAPN) significantly reduced tumor stiffness and disrupted collagen organization, resulting in enhanced T cell infiltration and potentiation of anti-PD1 monoclonal antibody therapy13. LOXL2 is widely recognized as an oncogenic driver in most studies62. Elevated LOXL2 expression promotes tumor progression and is significantly associated with poor prognosis. Mechanistically, increased ECM stiffness and hypoxic conditions have been shown to upregulate LOXL2 transcription. Subsequently, LOXL2-mediated collagen crosslinking further enhances tumor rigidity, creating a feed-forward loop that facilitates tumor invasion and metastasis63. The role of LOXL2 in the TIME remains unexplored. Emerging evidence from a recent study on hepatocellular carcinoma revealed that LOXL2 can interact with IQGAP1 and this interaction appears to modulate lymphocyte infiltration within the tumor microenvironment64. For our best research, this is the first study to focus on the function of LOXL2 in immune exclusion.
In addition, we found that the JAK2/STATE pathway mediated the regulation between GPR4 and LOXL2 or TGF-β. We found that GPR4 activates the JAK2/STAT3 signaling pathway by inducing the phosphorylation of both JAK2 and STAT3. Then p-STAT3 can translocate to the nucleus and binds to specific promoter regions of the LOXL2 and TGF-β genes, directly enhancing their transcriptional activation. STAT3, as a key member of the STAT family, plays a crucial role in tumorigenesis by promoting cell proliferation, inhibiting apoptosis, facilitating angiogenesis, enhancing invasion and metastasis, and enabling immune evasion65. Studies have demonstrated that phosphorylation of the Y705 residue activates STAT3, allowing it to form homodimers that recognize specific promoter regions, recruit RNA polymerase, and initiate the transcriptional expression of target genes66. Current studies have revealed that STAT3 can directly activate the transcriptional expression of CCL5, CXCR4, IL6, MMPs, and other genes67. Additionally, research has shown that STAT3 can influence the expression of TGF-β68,69. However, no study has demonstrated that STAT3 can directly bind to the promoter regions of LOXL2 and TGF-β to promote transcription.
We found that GPR4 directly interacts with JAK2 to promote its phosphorylation. This interaction depends on the 290-330 amino acid (aa) region within the intracellular domain of GPR4 and the FERM-SH2 domain (1-510 aa) of JAK2. JAK2 is an upstream protein of STAT3, which structurally consists of a receptor-binding domain (FERM-SH2), a JH1 kinase domain, and a JH2 pseudo kinase domain. Previous studies have established that JAK2 primarily undergoes phosphorylation and activation through direct binding to cytokine receptors, such as the growth hormone receptor (GHR) and interleukin-6 receptor (IL-6R)70. However, few studies have reported direct binding and activation of JAK2 by GPCRs. To date, only AGTR1 (angiotensin II receptor type 1) and GHRHR (growth hormone-releasing hormone receptor) have been documented in the literature as GPCRs capable of this interaction39,40. Research found that proteins that can directly interact with JAK2 typically contain regions rich in basic amino acid clusters (e.g., lysine, arginine) and hydrophobic residues71,72. As a GPCR, GPR4 binds to JAK2 primarily through its intracellular 290aa-330aa aa segment. Notably, this region (292aa-309aa: VAAKALAHNLALRFAL) contains a characteristic sequence enriched with both basic and hydrophobic amino acids. We hypothesized that this specific motif may provide the structural basis for binding to JAK2’s FERM-SH2 domain.
Another important finding of our study is that elevated GPR4 expression positively correlated with increased PD-L1 levels. In vivo studies further demonstrated that in GPR4-high CC models, both LOXL2 and JAK2 inhibitors enhanced intratumoral infiltration of CD8+ T lymphocytes by inhibiting the alignment of ECM collagen fibers. When combined with anti-PD-L1 monoclonal antibody therapy, these inhibitors exhibit superior therapeutic efficacy compared with anti-PD-L1 monotherapy. The core function of immune checkpoint inhibitors is to block or suppress the expression of immune checkpoint molecules, thereby enhancing the tumor-killing capacity of CD8+ T lymphocytes73. For immune exclusion induced by GPR4 high expression in CC, LOXL2 and JAK2 inhibitors synergistically enhance the anti-tumor efficacy of PD-L1 monoclonal antibody through dual mechanisms: (1) PD-L1 blockade reverses tumor-mediated immunosuppression of CD8+ T lymphocytes in GPR4-high tumors; (2) LOXL2/JAK2 inhibitors disrupt collagen alignment and reduce tumor collagen deposition, thereby promoting CD8+ T cell infiltration into tumor nests and increasing their intra-tumoral density. In lung cancer, elevated collagen expression exhibits a negative correlation with the efficacy of anti-PD-L1 therapy. Resistance to anti-PD-L1 treatment is associated with increased collagen deposition in the tumor region. Further investigations have revealed that collagen promotes CD8 + T lymphocyte exhaustion through LAIR-1 signaling74. Furthermore, reducing local collagen deposition in tumors effectively suppresses FAK protein secretion, which consequently decreases the infiltration of tumor-associated macrophages (TAMs) and Tregs, while enhancing the local accumulation of CD8+ T lymphocytes to improve immunotherapy efficacy75. Therefore, we believe that ICI therapy may present a more effective treatment response when combined with LOXL2 or JAK2 inhibitors for CC.
In conclusion (supplementary Fig. 6), GPR4 facilitates immune exclusion in CC through two distinct collagen-modulating mechanisms: (1) GPR4 upregulates LOXL2 expression, thereby enhancing the alignment of collagen fibers; (2) GPR4 increases TGF-β expression, leading to elevated type I collagen expression and deposition. These synergistic effects create a physical barrier that inhibits CD8+ T cell infiltration into tumor nests, thereby promoting an immune-excluded microenvironment. Furthermore, the intracellular domain of GPR4 directly interacts with the FERM-SH2 domain of JAK2, initiating the activation of the JAK2/STAT3 pathway. Phosphorylated STAT3 binds to the promoter regions of both LOXL2 and TGF-β, driving their transcription. Finally, targeted inhibition of GPR4 downstream effectors (LOXL2 or JAK2) reverses immune exclusion and synergizes with ICI, presenting a novel combinatorial strategy for CC patients.
Methods
Cell culture and plasmid construction
Human CC cell lines, including HT-29 (CAT: CL-0118), HCT116 (CAT: CL-0096), SW-480 (CAT: CL-0223), SW-620 (CAT: CL-0225), and Caco2 (CAT: CL-0050), mouse CC cell line MC38 (CAT: CL-0972), and HEK293T (CAT: CL-0005) cell line were obtained from Procell Life Science& Technology Co Ltd. HT-29 cells were cultured in McCoy’s 5 A medium (Gibco, 16600082). Other cell lines were cultured in DMEM (Gibco, 11965092). All media were supplemented with 10% fetal bovine serum (Lonsera, S711-001S) and penicillin-streptomycin solution (Sigma, P4333). All cell lines were routinely tested for mycoplasma contamination using a Luminescent Mycoplasma Detection Kit (Beyotime, C0297S) to ensure they were free of mycoplasma infection. The LOXL2 inhibitor β-aminopropionitrile (BAPN), JAK2 inhibitor fedratinib (fedratinib hydrochloride hydrate), and STAT3 inhibitor Stattic were purchased from MedChemExpress (CAT: HY-Y1750, HY-10409A, and HY-13818). BAPN and Fedratinib were dissolved in pure water, and Stattic was dissolved in DMSO to prepare working solutions of appropriate concentrations. GPR4 shRNA and negative control shRNA were constructed into the pLV3-U6-MCS-3×FLAG-CopGFP-Puro plasmid. And the target sequences were referenced the research of Zhong et al.25 and shown in supplement Table 1. The full length GPR4 tagged with Flag and truncated isoforms of GPR4 (animo acids 1-290, animo acids 1-330, animo acids 290-362) tagged with Flag were constructed into pLV3-CMV-MCS-3×FLAG-CopGFP-Puro plasmid. Then, the full length JAK2 tagged with HA and truncated isoforms of JAK2 (animo acids 1-510, animo acids 540-1124) tagged with Flag were constructed into the same plasmid as GPR4.
Lentiviral production and cell transduction
Lippo 3000 transfection kit (Thermo, L3000015) was utilized for the transfection of plasmids mentioned above into CC cells. Lentiviral packaging vector kits (Beyotime, L00002), which included REV, pMDLg, and VSVG, were used to transfect HEK293T cells with the abovementioned plasmids. After 48 h of transfection, the cell culture supernatants were collected to obtain lentiviruses. Finally, western blotting was performed to determine the efficiency of target gene expression.
Western blotting
Cell lysates were extracted by radioimmunoprecipitation (RIPA, Beyotime P0013B) buffer. And protein concentration was assayed by BCA Assay Kit (Beyotime, P0010S). Protein lysates were measured by 10% or 12% SDS–PAGE gels and transferred to 0.22 µm PVDF membrane (Millipore, ISEQ00010). Primal antibodies are: anti-GPR4 (dilution: 1:800, SAB, 44927), anti-JAK2 (dilution: 1:500, SAB, 48778), anti-JAK2-Tyr1007/1008 (dilution: 1:500, Zenbio, R381556), anti-STAT3 (dilution: 1:800, Proteintech, 10253), anti-STAT3-Tyr705 (dilution: 1:800, Beyotime, AF1276), anti-TGF-β (dilution: 1:500, Zenbio, R346599), anti-LOXL2 (dilution: 1:500, Zenbio, R382092), anti-COL1A1 (dilution: 1:500, Zenbio, R26615), anti-LOX (dilution: 1:800, SAB, 36747). The second antibodies are: goat anti-rabbit HRP-conjugated IgG (dilution: 1:10 000, SAB, L3012), goat anti-mouse HRP conjugated IgG (dilution: 1:10 000, SAB, L3032). The membrane was incubated in primal antibodies 12 h at 4 °C and in second antibodies 1.5 h at room temperature. Amersham Imager 600 (Cytiva) and ECL West Pico PLUS (Thermo, 34579) was used for visualized protein bands in membrane.
qRT-PCR
RNA extraction kit (Takara, 9767) was utilized for extracting total RNA from CC cells. Then, the RNA was transcribed to cDNA by using PrimeScript RT reagent Kit (Takara, RRO47A). Finally, SYBR Green Fast qPCR Mix (Abclone, RK21204) was used for assessing the RNA expression. GAPDH was used as internal reference. The 2-ΔΔCt method was applied to compare the relative mRNA expression levels of different genes. The primer sequences used in this study were listed in Supplementary Table S2.
Immunohistochemistry (IHC)
Tumor tissues was fixed with 10% formalin buffer and then embedded in paraffin as a standard protocol. Paraffin-embedded samples were deparaffinized in xylene, rehydrated in ethanol solution, washed in PBS buffer. EDTA antigen unmasking solution was used for antigen retrieval at 95 °C for 15 min, and blocked whith 5% BSA buffer 1.5 h subsequently. Sections were incubated with the first antibody overnight at 4 °C. The first antibodies were: anti-CD8 (dilution: 1:200, Abclone, A23305PM), anti-GPR4 (dilution: 1:200, Thermo, 118810), anti-LOXL2 (dilution: 1:150, Zenbio, R26615). Subsequently, sections were incubated with HRP-conjugated rabbit/mouse secondary antibodies (Prointech, RGAU011) and presented with DAB kit (Servicebio, G1212).
CC patient samples
Primary CC specimens and corresponding adjacent normal tissues were obtained from five patients treated at the First Affiliated Hospital of Xi’an Jiaotong University. All patients were received surgery and diagnosed with colorectal adenocarcinoma. The experimental protocol was reviewed and approved by the Ethics Committee of the First Affiliated Hospital of Xi’an Jiaotong University and was performed in accordance with the ethical standards as laid down in the 1964 Declaration of Helsinki and its later amendments or comparable ethical standards. Written informed consent was obtained from all participants prior to the specimen collection.
Multiplex immunohistochemical (mIHC)
Tyramide signal amplification technology was used for mIHC imaging of paraffin-embedded samples. Deparaffinization, rehydration, and antigen unmasking were performed as described in the IHC protocol. The first antibodies were: anti-GPR4 (dilution: 1:500, Thermo, 118810), anti-pan-CK (dilution: 1:500, abcam, ab7753), anti-pan-CK (dilution: 1:200, abcam, ab7753), and anti-CD8 (dilution: 1:500, abcam, ab237709). The second antibodies were used as per the IHC protocol. Sections were then incubated with TSA buffer (Servicebio, G1236) containing iF488-TSA, iF555-TSA, and iF647-TSA. After each staining cycle, the primary and secondary antibodies bound during were removed by heating, followed by the initiation of the next round of staining with the next antibody. Finally, a DAPI kit (Servicebio, G1236) was used to stain the cell nuclei.
Immunofluorescence staining
CC cells were cultured in laser confocal dishes and fixed with −20 °C pre-cooled methanol. The cells were then treated with 1% Triton X-100 for permeabilization and 5% BSA for blocking nonspecific binding sites. The first antibodies were: anti-STAT3-Tyr705 (dilution: 1:100, Beyotime, AF1276), anti-GPR4 (dilution: 1:150, Thermo, 118810), anti-JAK2 (dilution: 1:200, SAB, 44247). The cells were incubated with the primary antibodies overnight at 4 °C. The second IF antibodies were: Coralite488-conjugated goat anti-mouse second antibody (dilution: 1:200, Proteintech, SA00013), Coralite633-conjugated goat anti-rabbit second antibody (dilution: 1:200, Proteintech, SA00014), Cy3-conjugated goat anti-rabbit second antibody (dilution: 1:200, Proteintech, SA00009). The cells were incubated with the second antibodies for 1.5 h at room temperature and then treated with DAPI buffer (Thermo, D1306) for 15 min in the dark. A Leica STELLARIS 5 confocal fluorescence microscopy and Leica SP5 fluorescence microscope.
Second harmonic generation (SHG) imaging
Mouse CC samples were embedded in paraffin, and 10 µm thick sections were cut using a microtome. The sections were imaged using a Lecia SP8 DIVE multiphoton confocal microscope. The excitation wavelength was 810 nm, and a 405 ± 15 nm bandpass emission controlled by a slit was used to detect the SHG signal of collagen12,13. The SHG signal of collagen was detected when two photons of emission light focused on the specific structure of the collagen fibers. CT-fire was used to measure the collagen signal, which assessed collagen length, width, and orientation. In this study, the coefficient of variation (CV) was adopted to assess the collagen orientation.
T cell isolation and culture
Peripheral blood mononuclear cells (PBMC) were isolated from healthy adult donors via density gradient centrifugation using a Ficoll separation solution. This protocol was also reviewed and approved by the Ethics Committee of the First Affiliated Hospital of Xi’an Jiaotong University. The relevant consent was informed to health donors before PBMC collection, and all participants received no compensation from this study. After erythrocyte lysis with RBC lysis buffer, PBMCs were seeded into 6-well plates and incubated at 37 °C for 5 h in a humidified CO₂ incubator. After gentle agitation, the non-adherent cells (enriched for lymphocytes) were collected. Subsequently, these lymphocytes were cultured in 6-well plates coated with anti-CD3 and anti-CD28 antibodies (Elabscience, MIH001A) and abundant IL-2 (MCE, HY-P7037B) to promote T cell proliferation and activation. Finally CD8+ T cell were isolated from PBMCs via using CD8+ T Cell Isolation Kit (Milteniy, 130-094-156) as recommended protocols.
Migration assay of T cells
In this study, there were two types of migration assay for T cells. Based on previous study12, vertical migration assay for T cells was constructed. T cells were incubated with flow antibodies: FITC-conjugated anti-CD3 antibody (dilution: 1:100, BioLegend, 317305), PerCP-conjugated anti-CD8 antibody (dilution: 1:100, BioLegend, 980916), and APC-conjugated anti-CD4 antibody (dilution: 1:100, Biolegend, 980812). About 10,000 CC cells were seeded in an upper chamber with 5 µm pores (Corning, 3421) and cultured 3 days. The chamber was then immersed in a special lysate buffer (0.5% Triton X-100 and 20 mM NH4OH) for 5 min. After washing with PBS buffer for the chamber, a decellularized ECM was obtained. About 106 T cells were added into the upper chamber which contained decellularized ECM. And enriched-CCL21 (200 ng/mL, MCE) medium was supplied to the bottom chamber to promote T cell migration through the ECM into the bottom chamber. After 2 h, the T cells in the bottom chamber were collected and quantified using flow cytometry.
T-cell horizontal migration assay model was constructed by applying collagen type I (Corning, 354236) and four-chamber glass bottom dishes (Cellvis, D35C4). A collagen solution (2 mg/mL) was prepared and evenly pipetted into the bottom recesses of each compartment of a quadripartite confocal dish. Prior to collagen polymerization, 2000 tumor cells were seeded at the apical end of the collagen matrix. Complete culture medium was then added, followed by incubation at 37 °C for 72 h. Subsequently, carefully add 10000 CD8+ T cells pre-incubated with Hoechst 33342 to each partitioned edge region. After one day of culture, the migration efficiency of T cells was observed using fluorescence microscopy. The stably transfected CC cells constitutively expressed GFP (green fluorescence), whereas CD8+ T cells lacked GFP fluorescence, enabling a clear distinction between the two cell types. Hoechst 33342 (Thermo, R37605) staining labels all cell nuclei with bright blue fluorescence, facilitating microscopic field localization.
Amplex red fluorescence measurement for LOX activity
The LOX enzyme family catalyzes the oxidative deamination of peptidyl lysine and hydroxylysine residues on collagen, a process that is accompanied by the formation of hydrogen peroxide. In the presence of horseradish peroxidase, Amplex Red reacts with hydrogen peroxide to form resorufin, which emits fluorescence at 590 nm upon excitation at 530–560 nm. By measuring the fluorescence intensity, the hydrogen peroxide concentration and the catalytic activity of the LOX enzyme family in collagen crosslinking can be determined. In this study, culture supernatant, 0.1 U/mL HRP (Thermo, 31491), and 50 µm Amplex red buffer (Thermo, A12222) were mixed and added to a 96-well plate. The well-mixed sample solution was then incubated at room temperature for 30 min. The fluorescence signal was measured using a multifunctional microplate reader (Biotek), with the excitation wavelength set at 540 nm and the emission wavelength detected at 590 nm.
Collagen gel contraction assay
According to previous research60,76, 1 × 105 CC cells were mixed with 2 mg/mL collagen (Corning, 354236). A total of 300 μL mixture were added into 48-well plate, and incubated at 37 °C for 3 days. For the influence of recombinant protein LOXL2 (MCE, HY-P74770), 10 ng/mL rLOXL2 were added into collagen culture. The collagen area in plate were measured by ImageJ. Images were taken by a camera, and a well area was used as control.
Visualization of collagen structure remodeling by rLOXL2
Collagen solution was uniformly added to a 24-well plate (500 μL/well) and polymerized at 37 °C for 1 h. After adding rLOXL2 (10 ng/mL in PBS) to the treatment groups, the plates were incubated for 48 h at 37 °C. Post-incubation washes (3× PBS) preceded 15 min of NHS-ester labeling. Collagen fiber alignment was visualized using fluorescence microscopy (Leica, polarized model, 20× objective). CT-fire software was used to analyze the images.
Flow cytometry
Fresh tumor tissues were minced into 1 mm³ fragments and digested in a tissue digestion solution containing 2% serum, 1 mg/mL collagenase, and 200 U/mL DNase I at 37 °C on a constant-temperature shaker for 1.5 h. Then, cell suspension passed through 70-µm cell strainer to obtain single-cell suspension, and CD16/CD32 (Elabscience, E-AB-F0997A) were added to block unspecific binding sites. Single-cell samples were stained for viability assess by using Zombie Yellow Fixable Viability Kit (Biolegend, 423103) at 4 °C in the dark for 15 min. Then samples were stained by commercial antibodies for 20 min in dark, including anti-CD45-violent 450 (Elabscience, E-AB-F1136E), anti-CD3-APC (Elabscience, E-AB-F1013Q), anti-CD4-FITC (Elabscience, E-AB-F1097C), anti-CD8-PerCP (Elabscience, E-AB-F1104J). Subsequently, Transcription Factor Staining Kit (Elabscience, E-CK-A108) was applied for cells fixation and Permeabilization as recommended method. And samples were stained by anti-KI67-PE (Biolegend, 151209) for 20 min in dark. Finally, data were acquired on ACEA Novo Cyte flow cytomete.
Animal studies
All animal experiments were approved by the Ethics Committee of Xian Jiaotong University. Male BALB/C nude mice (5–6 weeks old, catalog: D000521, BALB/cNj-Foxn1nu/Gpt) and male immunocompetent C57BL/6 J mice (5–6 weeks old, catalog: C57BL/6JGpt, N000013) were purchased from the Animal Center of Xi’an Jiaotong University. All mice were maintained in a specific pathogen-free (SPF) facility in the animal center with controlled temperature (22 ± 1 °C), humidity (50 ± 10%), and a 12-hour light/dark cycle. Mice were randomly allocated to different groups and bred separately. Each group is guaranteed to have at least five experimental mice. MC38 cells were subcutaneously injected into the right flank (1 × 10⁶ cells/100 μL in PBS). Tumor volume was monitored starting at 7 days post-inoculation and tumor volume was calculated using the modified ellipsoid formula: volume = (length × width²)/2. Measurements were performed every 2–3 days until the ethical endpoint criteria were reached. Different treatments were initiated when the largest subcutaneous tumor reached a threshold volume of 100 mm³. Treatments included: BAPN (100 mg/Kg, twice a day, intraperitoneal injection), (2-Chloropyridin-4-yl) methanamine hydrochloride (10 mg/Kg, once a day, intraperitoneal injection, MCE HY-101771A), Anti-PD-L1 monoclonal antibody (100 µg, every three days, intraperitoneal injection, SELLECK A2115), and fedratinib (50 mg/Kg, once a day, oral gavage). Tumors were measured the maximal and shortest diameters of the tumor using a vernier caliper every 3 to 5 days. Mice were euthanized when tumors exceeded 1.5 cm in maximal diameter, 2000 mm3 in tumor maximum volume or 30 days post-inoculation. The animals will be euthanized immediately if they are unable to stand or if they display agonal breathing, severe weight loss, or uncontrolled bleeding. After tumor sample collection, the tumors were weighed and paraffin-embedded for IHC or minced into fragments for flow cytometry.
RNA-seq
RNA-seq was performed on the OE-GPR4 SW480 cell and OE-NC SW480 cell. The sequencing library preparation in this study was conducted by New Open Medical Group. Total RNA was extracted using the Trioz method, with subsequent procedures primarily encompassing RNA extraction and quality control, library construction, sequencing, and transcriptome assembly. And The gene expression data were normalized and quantified using FPKM (Fragments Per Kilobase of transcript per Million mapped reads). Gnen different expression analysis was performed using the edgeR algorithm, followed by functional enrichment analysis conducted with the ClusterProfil R package.
Bioinformatic analysis
RNA-seq data (Level 3) and corresponding clinical data of 437 colon adenocarcinoma samples, were obtained from the TCGA dataset. Protein-level validation was obtained through immunohistochemistry (IHC) data from The Human Protein Atlas (HPA) database (https://www.proteinatlas.org/), specifically examining expression patterns in colorectal cancer specimens. Bioinformatic analysis were performed with R software 3.6.1. Differential expression analysis between groups was performed using the “limma” R package. For clinical data, prognostic correlation analysis was conducted with the “survival” package, followed by Kaplan-Meier curve visualization. The association between GPR4 expression levels (high vs. low) and clinicopathological characteristics (TNM stage, age, gender, etc.) was statistically evaluated using the “gtsummary” package. To quantify CD8+ T-cell infiltration levels from transcriptomic data, the “IOBR” package77 was employed to implement four computational algorithms: “EPIC78”, “MCPcounter79”, “TIMER80” [105], and “ssGSEA81”. Additionally, the TIDE database was utilized to assess CC patients across three dimensions: (1) immune cell dysfunction, (2) T-cell exclusion, and (3) potential immunotherapy benefit. For patient stratification based on proton-sensing receptor expression, dimensionality reduction was performed via the “Rtsne” package, while consensus clustering was achieved using “ConsensusClusterPlus”. Functional enrichment analysis was performed using the R package clusterProfiler, implementing both Gene Ontology (GO) term enrichment. For cross-species functional annotation, gene universality was assessed via the GeneBridge method82 (https://www.systems-genetics.org/).
Co-immunoprecipitation assay (Co-IP)
IP lysis buffer was used to obtain cell lysates. The antibodies for IP were: anti-GPR4 antibody (dilution: 1:50, Thermo, 118810), anti-JAK2 antibody (dilution: 1:50, SAB, 44247), HA tag antibody (dilution: 1:50, Proteintech, 51064), Flag tag antibody (dilution: 1:50, Proteintech, 66008). The immunoprecipitation kit with protein A + G magnetic beads (Beyotime, P2108) was used to drag out the antigen-antibody complex. Finally, Western blotting was conducted to assay the complex.
Chromatin immunoprecipitation assay (ChIP)
Chromatin Immunoprecipitation (ChIP) Assay Kit with Protein A/G Magnetic Beads (Beyotime, P2179) was used for ChIP assay. CC cells were corss-linked in 1% formaldehyde for 10 min. Cells were lysed on ice using SDS Lysis Buffer supplemented with protease inhibitors and PMSF. Chromatin was sheared into 200–500 bp fragments by sonication. The resulting DNA-protein complexes were incubated overnight at 4 °C with the target antibody. Subsequently, Protein A/G magnetic beads were added, followed by incubation at room temperature for 2 h to facilitate antibody-antigen complex capture. After washing with salt buffer, adding EDTA, Tris, and proteinase K at 45 °C for 1 h and then was purified, DNA was obtained for qPCR analysis. The primer sequences applied for ChIP-qPCR assay in the LOXL2 and TGF-β promotor region were listed in Supplementary Table S3.
Statistics and reproducibility
All statistical data with error bars were presented as the mean ± standard deviation (SD), and all experiments were independently repeated at least three times. All statistical analyses and data visualizations were performed using R and GraphPad Prism 8. Student’s t-tests were used to compare the mean differences between the two groups. ANOVA was used for multiple groups. Comparisons were conducted using Pearson’s χ² test or Fisher’s exact test, as appropriate. Kaplan-Meier curves were generated, and differences were evaluated using the log-rank test. All statistical tests were two-sided, and a P ≤ 0.05 was considered statistically significant.
Ethical statement
This research complies with all relevant ethical regulations. All animal experimental protocols were reviewed and approved by the First Affiliated Hospital of Xi’an Jiaotong University (XJTUAE2025-11).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The RNA-seq data generated in this study have been deposited in the Genome Sequence Archive in China National Center for Bioinformation83 (https://ngdc.cncb.ac.cn/gsa-human) under the accession number: HRA014624. The TCGA COAD data is available in the Genomic Data Commons (GDC) Data Portal. The remaining data are available within the Article, Supplementary Information or Source Data file. Further information and requests for resources are available from corresponding author upon reasonable request. All data are included in the Supplementary Information or available from the authors, as are unique reagents used in this Article. The raw numbers for charts and graphs are available in the Source Data file whenever possible. Source data are provided with this paper.
References
Bray, F. et al. Global cancer statistics 2022: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. Ca-Cancer J. Clin. 74, 229–263 (2024).
Xu, C. et al. Comparative safety of immune checkpoint inhibitors in cancer: systematic review and network meta-analysis. BMJ 363, k4226 (2018).
Lahiri, A. et al. Lung cancer immunotherapy: progress, pitfalls, and promises. Mol. Cancer 22, 40 (2023).
Li, J. et al. Remodeling of the immune and stromal cell compartment by PD-1 blockade in mismatch repair-deficient colorectal cancer. Cancer Cell 41, 1152–1169 e1157 (2023).
Sinicrope, F. A. & Turk, M. J. Immune checkpoint blockade: timing is everything. J. Immunother. Cancer 12. https://doi.org/10.1136/jitc-2024-009722 (2024).
Chen, D. S. & Mellman, I. Elements of cancer immunity and the cancer-immune set point. Nature 541, 321–330 (2017).
Desbois, M. et al. Integrated digital pathology and transcriptome analysis identifies molecular mediators of T-cell exclusion in ovarian cancer. Nat. Commun. 11, 5583 (2020).
Cox, T. R. The matrix in cancer. Nat. Rev. Cancer 21, 217–238 (2021).
Bruni, S., Mercogliano, M. F., Mauro, F. L., Cordo Russo, R. I. & Schillaci, R. Cancer immune exclusion: breaking the barricade for a successful immunotherapy. Front. Oncol. 13, 1135456 (2023).
Bhowmick, N. A., Neilson, E. G. & Moses, H. L. Stromal fibroblasts in cancer initiation and progression. Nature 432, 332–337 (2004).
Quante, M. et al. Bone marrow-derived myofibroblasts contribute to the mesenchymal stem cell niche and promote tumor growth. Cancer Cell 19, 257–272 (2011).
Sun, X. et al. Tumour DDR1 promotes collagen fibre alignment to instigate immune exclusion. Nature 599, 673–678 (2021).
Nicolas-Boluda, A. et al. Tumor stiffening reversion through collagen crosslinking inhibition improves T cell migration and anti-PD-1 treatment. Elife 10. https://doi.org/10.7554/eLife.58688 (2021).
Liu, Y. T. & Sun, Z. J. Turning cold tumors into hot tumors by improving T-cell infiltration. Theranostics 11, 5365–5386 (2021).
Chen, D. S. & Mellman, I. Oncology meets immunology: the cancer-immunity cycle. Immunity 39, 1–10 (2013).
Pietrobon, V. & Marincola, F. M. Hypoxia and the phenomenon of immune exclusion. J. Transl. Med. 19, 9 (2021).
Justus, C. R., Dong, L. & Yang, L. V. Acidic tumor microenvironment and pH-sensing G protein-coupled receptors. Front. Physiol. 4, 354 (2013).
Ludwig, M. G. et al. Proton-sensing G-protein-coupled receptors. Nature 425, 93–98 (2003).
Murakami, N., Yokomizo, T., Okuno, T. & Shimizu, T. G2A is a proton-sensing G-protein-coupled receptor antagonized by lysophosphatidylcholine. J. Biol. Chem. 279, 42484–42491 (2004).
Sisignano, M., Fischer, M. J. M. & Geisslinger, G. Proton-Sensing GPCRs in Health and Disease. Cells 10. https://doi.org/10.3390/cells10082050 (2021).
Ren, J. et al. Human GPR4 and the Notch signaling pathway in endothelial cell tube formation. Mol. Med. Rep. 14, 1235–1240 (2016).
Bai, Z. et al. Downregulation of GPR4 and TCF7 promotes apoptosis and inhibits growth and invasion of ovarian cancer cells. Anticancer Agents Med. Chem. 21, 1544–1550 (2021).
Ren, J. et al. Relations between GPR4 expression, microvascular density (MVD) and clinical pathological characteristics of patients with epithelial ovarian carcinoma (EOC). Curr. Pharm. Des. 20, 1904–1916 (2014).
Xue, C. et al. Association between G-protein coupled receptor 4 expression and microvessel density, clinicopathological characteristics and survival in hepatocellular carcinoma. Oncol. Lett. 19, 2609–2620 (2020).
Yu, M. et al. Increased proton-sensing receptor GPR4 signalling promotes colorectal cancer progression by activating the hippo pathway. EBioMedicine 48, 264–276 (2019).
Foti, F. et al. The Simultaneous Deletion of pH-Sensing Receptors GPR4 and OGR1 (GPR68) Ameliorates Colitis with Additive Effects on Multiple Parameters of Inflammation. Int. J. Mol. Sci. 26. https://doi.org/10.3390/ijms26041552 (2025).
Wang, Y. et al. The Proton-activated Receptor GPR4 Modulates Intestinal Inflammation. J. Crohns Colitis 12, 355–368 (2018).
Lee, S. et al. Phase 2 Open-Label Trial of Brentuximab Vedotin with Pembrolizumab in PD-1 Pretreated Metastatic Non-Small Cell Lung Cancer and Metastatic Cutaneous Melanoma. Clin. Cancer Res. https://doi.org/10.1158/1078-0432.CCR-24-1478 (2024).
Jansen, C. S. et al. An intra-tumoral niche maintains and differentiates stem-like CD8 T cells. Nature 576, 465–470 (2019).
Virassamy, B. et al. Intratumoral CD8(+) T cells with a tissue-resident memory phenotype mediate local immunity and immune checkpoint responses in breast cancer. Cancer Cell 41, 585–601.e588 (2023).
Donne, R. & Lujambio, A. The liver cancer immune microenvironment: Therapeutic implications for hepatocellular carcinoma. Hepatology 77, 1773–1796 (2023).
Cheng, G. et al. Trichinella spiralis-Secreted Products Promote Collagen Capsule Formation through TGF-beta1/Smad3 Pathway. Int. J. Mol. Sci. 24. https://doi.org/10.3390/ijms241915003 (2023).
Li, J. et al. Lysyl Oxidase (LOX) family proteins: key players in breast cancer occurrence and progression. J. Cancer 15, 5230–5243 (2024).
Lu, Y., Li, H., Chen, M., Lin, Y. & Zhang, X. LOX-induced tubulointerstitial fibrosis via the TGF-beta/LOX/Snail axis in diabetic mice. J. Transl. Med. 23, 35 (2025).
Li, H. et al. Single-cell analysis reveals lysyl oxidase (Lox)(+) fibroblast subset involved in cardiac fibrosis of diabetic mice. J. Adv. Res. 54, 223–237 (2023).
Ma, L. et al. Knockdown of LOXL1 inhibits TGF-beta1-induced proliferation and fibrogenesis of hepatic stellate cells by inhibition of Smad2/3 phosphorylation. Biomed. Pharmacother. 107, 1728–1735 (2018).
Cortes-Ballinas, L. & Lopez-Perez, T. V. & Rocha-Zavaleta, L. STAT3 and the STAT3‑regulated inhibitor of apoptosis protein survivin as potential therapeutic targets in colorectal cancer (Review). Biomed. Rep. 21, 175 (2024).
Liu, J. P. et al. Each one of certain histidine residues in G-protein-coupled receptor GPR4 is critical for extracellular proton-induced stimulation of multiple G-protein-signaling pathways. Pharm. Res. 61, 499–505 (2010).
Marrero, M. B. et al. Direct stimulation of Jak/STAT pathway by the angiotensin II AT1 receptor. Nature 375, 247–250 (1995).
Siejka, A., Schally, A. V. & Barabutis, N. Activation of Janus kinase/signal transducer and activator of transcription 3 pathway by growth hormone-releasing hormone. Cell Mol. Life Sci. 67, 959–964 (2010).
Min, H. Y. et al. S100A14: a novel negative regulator of cancer stemness and immune evasion by inhibiting STAT3-mediated programmed death-ligand 1 expression in colorectal cancer. Clin. Transl. Med. 12, e986 (2022).
Sin, W. C. et al. G protein-coupled receptors GPR4 and TDAG8 are oncogenic and overexpressed in human cancers. Oncogene 23, 6299–6303 (2004).
Insel, P. A., Sriram, K., Salmeron, C. & Wiley, S. Z. Proton-sensing G protein-coupled receptors: detectors of tumor acidosis and candidate drug targets. Future Med. Chem. 12, 523–532 (2020).
Klatt, W., Wallner, S., Brochhausen, C., Stolwijk, J. A. & Schreml, S. Expression profiles of proton-sensing G-protein coupled receptors in common skin tumors. Sci. Rep. 10, 15327 (2020).
Wyder, L. et al. Reduced pathological angiogenesis and tumor growth in mice lacking GPR4, a proton-sensing receptor. Angiogenesis 14, 533–544 (2011).
Jing, Z. et al. The proton-sensing G-protein coupled receptor GPR4 promotes angiogenesis in head and neck cancer. PLoS ONE 11, e0152789 (2016).
Nakanishi, M. et al. Acid-sensing receptor GPR4 plays a crucial role in lymphatic cancer metastasis. Cancer Sci. 115, 1551–1563 (2024).
Massi, D. et al. The density and spatial tissue distribution of CD8(+) and CD163(+) immune cells predict response and outcome in melanoma patients receiving MAPK inhibitors. J. Immunother. Cancer 7, 308 (2019).
Verdegaal, E. M. et al. Successful treatment of metastatic melanoma by adoptive transfer of blood-derived polyclonal tumor-specific CD4+ and CD8+ T cells in combination with low-dose interferon-alpha. Cancer Immunol. Immunother. 60, 953–963 (2011).
Mlecnik, B. et al. Histopathologic-based prognostic factors of colorectal cancers are associated with the state of the local immune reaction. J. Clin. Oncol. 29, 610–618 (2011).
Brodsky, B. & Persikov, A. V. Molecular structure of the collagen triple helix. Adv. Protein Chem. 70, 301–339 (2005).
Xiao, X. et al. Col1A1 as a new decoder of clinical features and immune microenvironment in ovarian cancer. Front. Immunol. 15, 1496090 (2024).
Yichao, Y. et al. COL1A1, COL1A2, CHN1, and FN1 Promote Tumorogenesis and Act as Markers of Diagnosis and Survival in Gastric Cancer Patients. Curr. Pharm. Biotechnol. https://doi.org/10.2174/0113892010332329241119104430 (2025).
Cong, G., Shao, J., Xiao, F., Zhu, H. & Kang, P. The role of endothelial cell-related gene COL1A1 in prostate cancer diagnosis and immunotherapy: insights from machine learning and single-cell analysis. Biol. Direct 20, 3 (2025).
Salimian, N., Peymani, M., Ghaedi, K., Hashemi, M. & Rahimi, E. Collagen 1A1 (COL1A1) and Collagen11A1(COL11A1) as diagnostic biomarkers in Breast, colorectal and gastric cancers. Gene 892, 147867 (2024).
Hsu, K. S. et al. Cancer cell survival depends on collagen uptake into tumor-associated stroma. Nat. Commun. 13, 7078 (2022).
Tabdanov, E. D. et al. Engineering T cells to enhance 3D migration through structurally and mechanically complex tumor microenvironments. Nat. Commun. 12, 2815 (2021).
Hartmann, N. et al. Prevailing role of contact guidance in intrastromal T-cell trapping in human pancreatic cancer. Clin. Cancer Res. 20, 3422–3433 (2014).
Lodyga, M. & Hinz, B. TGF-beta1 - a truly transforming growth factor in fibrosis and immunity. Semin. Cell Dev. Biol. 101, 123–139 (2020).
Kaur, A. et al. Remodeling of the collagen matrix in aging skin promotes melanoma metastasis and affects immune cell motility. Cancer Discov. 9, 64–81 (2019).
Wolf, K., Muller, R., Borgmann, S., Brocker, E. B. & Friedl, P. Amoeboid shape change and contact guidance: T-lymphocyte crawling through fibrillar collagen is independent of matrix remodeling by MMPs and other proteases. Blood 102, 3262–3269 (2003).
Wen, B., Xu, L. Y. & Li, E. M. LOXL2 in cancer: regulation, downstream effectors and novel roles. Biochim. Biophys. Acta Rev. Cancer 1874, 188435 (2020).
Cano, A., Eraso, P., Mazon, M. J. & Portillo, F. LOXL2 in cancer: a two-decade perspective. Int. J. Mol. Sci. 24. https://doi.org/10.3390/ijms241814405 (2023).
Zhao, N. et al. LOXL2 serves as a prognostic biomarker for hepatocellular carcinoma by mediating immune infiltration and vasculogenic mimicry. Dig. Liver Dis. 55, 661–672 (2023).
Yuan, J., Zhang, F. & Niu, R. Multiple regulation pathways and pivotal biological functions of STAT3 in cancer. Sci. Rep. 5, 17663 (2015).
Bharadwaj, U., Kasembeli, M. M., Robinson, P. & Tweardy, D. J. Targeting Janus kinases and signal transducer and activator of transcription 3 to treat inflammation, fibrosis, and cancer: rationale, progress, and caution. Pharm. Rev. 72, 486–526 (2020).
Shi, D. et al. Structure, function, signaling pathways and clinical therapeutics: The translational potential of STAT3 as a target for cancer therapy. Biochim. Biophys. Acta Rev. Cancer 1879, 189207 (2024).
Sadrkhanloo, M. et al. STAT3-EMT axis in tumors: modulation of cancer metastasis, stemness and therapy response. Pharm. Res. 182, 106311 (2022).
Celada, L. J. et al. PD-1 up-regulation on CD4(+) T cells promotes pulmonary fibrosis through STAT3-mediated IL-17A and TGF-beta1 production. Sci Transl Med 10 https://doi.org/10.1126/scitranslmed.aar8356 (2018).
Waters, M. J. & Brooks, A. J. JAK2 activation by growth hormone and other cytokines. Biochem. J. 466, 1–11 (2015).
Glassman, C. R. et al. Structure of a Janus kinase cytokine receptor complex reveals the basis for dimeric activation. Science 376, 163–169 (2022).
Abraham, B. G. et al. Molecular basis of JAK2 activation in erythropoietin receptor and pathogenic JAK2 signaling. Sci. Adv. 10, eadl2097 (2024).
Strati, A. et al. Targeting the PD-1/PD-L1 Signaling Pathway for Cancer Therapy: Focus on Biomarkers. Int. J. Mol. Sci. 26. https://doi.org/10.3390/ijms26031235 (2025).
Ramos, M. I. P. et al. Cancer immunotherapy by NC410, a LAIR-2 Fc protein blocking human LAIR-collagen interaction. Elife 10. https://doi.org/10.7554/eLife.62927 (2021).
Jiang, H. et al. Targeting focal adhesion kinase renders pancreatic cancers responsive to checkpoint immunotherapy. Nat. Med. 22, 851–860 (2016).
Bell, E., Ivarsson, B. & Merrill, C. Production of a tissue-like structure by contraction of collagen lattices by human fibroblasts of different proliferative potential in vitro. Proc. Natl. Acad. Sci. USA 76, 1274–1278 (1979).
Zeng, D. et al. IOBR: Multi-Omics Immuno-Oncology Biological Research to Decode Tumor Microenvironment and Signatures. Front. Immunol. 12, 687975 (2021).
Racle, J., de Jonge, K., Baumgaertner, P., Speiser, D. E. & Gfeller, D. Simultaneous enumeration of cancer and immune cell types from bulk tumor gene expression data. Elife 6. https://doi.org/10.7554/eLife.26476 (2017).
Becht, E. et al. Estimating the population abundance of tissue-infiltrating immune and stromal cell populations using gene expression. Genome Biol. 17, 218 (2016).
Li, B. et al. Comprehensive analyses of tumor immunity: implications for cancer immunotherapy. Genome Biol. 17, 174 (2016).
Zhang, Z. et al. Computational principles and practice for decoding immune contexture in the tumor microenvironment. Brief Bioinform. 22. https://doi.org/10.1093/bib/bbaa075 (2021).
Li, H. et al. Identifying gene function and module connections by the integration of multispecies expression compendia. Genome Res. 29, 2034–2045 (2019).
Members, C.-N. & Partners. Database Resources of the National Genomics Data Center, China National Center for Bioinformation in 2025. Nucleic Acids Res. 53, D30–D44 (2025).
Acknowledgements
This manuscript is supported by National Natural Science Foundation of China (J.R., 82473252, 81772793); supported by the Scientific and Technological Research Foundation of Shaan’xi Province, Key Research and Development Project, General project (J.R., 2023-YBSF-666); supported by the Shaanxi Provincial Key Research and Development Plan Project (Key support projects Category A) (J.R., 2021A011); supported by the Basic and Clinical Integration Innovation Project of Xi’ an Jiao tong University (J.R., YXJLRH2022006; YXJLRH2022004). Nanjing Tianqing Scientific Research Fund of First Affliated Hospital of Xi’an Jiaotong University, (J.R., TQ202205).
Author information
Authors and Affiliations
Contributions
S.B. conducted experiments and analyzed data; S.B. wrote the manuscript; J.R. conceived and supervised the study; J.R., S.B., M.C., X.W., N.L., R.M., Y.Y., W.L., W.Y.L., F.W., X.Z., F.H., and R.L. made manuscript revisions. All authors have read and approved the final version of this submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Bai, S., Chen, M., Wang, X. et al. GPR4 promotes immune exclusion in colon cancer through LOXL2-mediated extracellular matrix remodeling. Nat Commun 17, 1196 (2026). https://doi.org/10.1038/s41467-025-67967-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-67967-z