Abstract
The emergence of multidrug-resistant and hypervirulent Klebsiella pneumoniae represents a growing threat to global public health. Here we investigate the genomic epidemiology and evolutionary dynamics of emerging K. pneumoniae clonal group 23 (CG23) lineages, with a particular focus on ST218-KL57 (predominantly found in Asia) and ST23-KL57 (circulating in Europe) to understand how genomic recombination and adaptive evolution drive the convergence of hypervirulence and multidrug resistance in CG23. We analyzed 3,118 clinical isolates from China together with 75,954 publicly available genomes, encompassing 122 ST218-KL57 and 45 ST23-KL57 isolates; 13 ST218-KL57 isolates were newly sequenced. Phylogenetic analyses revealed that ST23-KL57 forms a distinct subclade within CG23, more closely related to ST218-KL57 than to the classical ST23-KL1 linage. Neither ST218-KL57 nor ST23-KL57 harbors the chromosomal virulence loci (GIE492 and all island), which are characteristic of hypervirulent ST23-KL1. ST23-KL57 simultaneously harbors dual carbapenemases and key virulence genes (peg-344, iucA, rmpADC, and rmpA2), reflecting its hybrid origin of approximately 80% Asian hypervirulent ST218-KL57 and 20% European multidrug-resistant ST395 linage. The genomic decay of the iroBCDN operon in ST23-KL57 suggests an adaptive trade-off between antimicrobial resistance and virulence. These findings underscore the importance of enhanced genomic surveillance and refined risk assessment for the evolving CG23-KL57 lineage.
Similar content being viewed by others
Introduction
Klebsiella pneumoniae is not only a commensal bacterium of the human gut microbiota but is also a versatile opportunistic pathogen responsible for a wide range of infections, including pneumonia, bloodstream infections, urinary tract infections, and liver abscesses1. It is frequently associated with antimicrobial resistance (AMR) and has been designated by the World Health Organization (WHO) as a priority pathogen for global action due to its escalating resistance to multiple antibiotics2. Two major pathotypes are recognized: classical K. pneumoniae (cKp) and hypervirulent K. pneumoniae (hvKp)3. cKp predominantly causes hospital-acquired infections in immunocompromised individuals4, while hvKp is notably associated with community-acquired infections such as pyogenic liver abscesses, which can spread and lead to complications such as meningitis, necrotizing fasciitis, and endophthalmitis. Owing to its hallmark characteristics of hyperinvasiveness, immune evasion, and rapid progression, hvKp poses a significant threat, even to young and immunocompetent individuals5. In July 2024, the WHO issued a global alert regarding the rising incidence of hvKp sequence type (ST) 23 strains harboring carbapenemase-encoding genes that render most clinically available antimicrobials ineffective6. Similarly, the European Centre for Disease Prevention and Control (ECDC) highlighted the emergence of multidrug-resistant K. pneumoniae ST23 strains in Europe, emphasizing their capacity for rapid evolution and resistance acquisition7. This convergence of hypervirulence and carbapenem resistance poses a severe threat to public health and clinical management.
The hypervirulent phenotype of hvKp is largely plasmid-mediated. These virulence plasmids share structural features with the pK2044 and commonly encode the mucoid regulators (rmpADC and rmpA2)8 that enhance capsular polysaccharide production, leading to a hypermucoviscosity (hmv) phenotype. In addition, they carry the iuc and iro operons responsible for the synthesis of aerobactin, a siderophore strongly linked to enhanced virulence9. Although these classical virulence plasmids are not self-transmissible, they can be mobilized by other conjugative plasmids10. In addition, chromosomal acquisition of ICEKp10, which carries genes for yersiniabactin (ybt) and colibactin (clb), enhances the spread of hvKp8. Core chromosomal features such as enterobactin, as well as adhesins and biofilm-associated genes (mrk and fim), further enhance fitness, persistence, and pathogenicity11.
hvKp has been linked to multiple clonal groups (CGs) and their corresponding STs, notably CG23/ST23, CG65/ST65, and CG86/ST86. Additionally, it is associated with a range of distinct capsular locus (KL) types, including KL1, KL2, KL5, and KL5712. Among these, CG23/ST23 has emerged as the predominant hypervirulent lineage worldwide, with the capacity to acquire multiple carbapenemase genes, including blaKPC-2, blaNDM-1, and blaOXA-4813. Notably, ST23-KL1 (ST23 strains with KL1) has long been considered the prototypical hvKp clone, frequently linked to community-acquired invasive infections in healthy individuals. ST23 strains with the capsular serotype KL57 (ST23-KL57) that also produce carbapenemase have been predominantly reported in Europe14. ST218 and its single locus variant ST23 both belong to CG23, but recent genomic evidence indicates that ST23-KL57 is genetically closer to ST218-KL57 (ST218 strains with KL57) than to the classical ST23-KL1 lineage, despite sharing the same 7-locus MLST profile15,16. These ST23-KL57 strains carry virulence plasmids and harbor key virulence loci such as iuc and ybt, though they often lack clb17. While ST23-KL57 has primarily been reported in Europe14, its close relative ST218-KL57 appears to be more widespread in China18. However, detailed information regarding the evolution, genomic structure, and virulence potential of these emerging lineages remains limited.
In this study, we conducted a systematic genomic and epidemiological investigation of ST218-KL57 isolates collected from multiple hospitals across China. We compared these isolates to the global ST23-KL57 and ST23-KL1 genomic landscape to understand their population structure, resistance and virulence gene profiles, and phylogenetic relationships. In addition, we performed in vivo infection studies using a murine model to explore the association between genomic features and virulence potentials. Our findings reveal the genetic distinctiveness and geographical distribution of ST218-KL57 and related lineages, underscoring the need for enhanced genomic surveillance, particularly for tracking convergence events between AMR and hypervirulence determinants that pose serious clinical and epidemiological challenges.
Results
Divergent geographic prevalence of ST218-KL57 and ST23-KL57 isolates
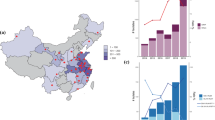
In this study, a total of 3,118 K. pneumoniae isolates, together with all publicly available K. pneumoniae genomes (n = 75,954), were collected to evaluate the prevalence and clonal dissemination of the ST23-KL57 and ST218-KL57 lineages. Based on PCR and ST/KL typing, we found that although ST23-KL57 strains have not yet been detected, this is consistent with current reports indicating that this lineage has not emerged in China. Thirteen ST218-KL57 strains were identified and subjected to whole-genome sequencing. Along with 109 ST218 genomes retrieved from NCBI, our analysis included a total of 122 ST218-KL57 isolates collected from four continents across 18 countries (Supplementary data 1; Supplementary Fig.1). Asia contributed the largest share, accounting for 77.0% (94/122) of the total. Statistical analysis of the number of isolates from each country revealed that China had the highest number, with 72 strains, representing 59.0% (72/122) of all isolates and 76.6% (72/94) of the Asian isolates. The second-highest number of isolates came from Japan (13.1%, 16/122), while the third was from Russia 7.4% (9/122) (Fig. 1A). The collection period spanned from 2009 to 2024, though a significant surge was seen after 2019, making up 54.1% (66/122) of the total (Fig. 1B) In addition, we analyzed all 45 publicly available ST23-KL57 strains (Supplementary data 1). Predominantly found in Europe, these strains are mainly concentrated in Germany, Russia, and Poland. The collection period spanned from 2014 to 2024, presenting a stark contrast to the distribution of ST218-KL57 strains (Fig. 1C).
A Spatial distribution of the 122 ST218-KL57 and 45 ST23-KL57 global isolates. The pie chart represents the number of isolates, with different colors indicating the distribution of isolates within ST218-KL57 and ST23-KL57. B Distribution of the global 122 ST218-KL57 isolates across different years and countries. The heatmap displays the number of ST218-KL57 isolates in each year and country. The total numbers of ST218-KL57 isolates collected in a year or country are shown in the bar charts on the right and top of the heatmap, respectively. C Distribution of the global 45 ST23-KL57 isolates across different years and countries. The heatmap displays the number of ST23-KL57 isolates in each year and country. The total numbers of ST23-KL57 isolates collected in a year or country are shown in the bar charts on the right and top of the heatmap, respectively. ST sequence type, KL capsule locus. Source data are provided as a Source Data file. The world map was generated entirely in R script using the ggplot2 package and geographic coordinates derived from the built-in world map data (via the maps package), which are based on public-domain sources.
Close genetic relationship between ST218-KL57 and ST23-KL57 lineages
To investigate the genetic relationships between ST218 and ST23 strains, we conducted core-genome clustering analysis of ST218-KL57 (n = 122), ST23-KL1 (n = 1,491), and ST23-KL57 (n = 45) strains (Supplementary Fig. 2). The results revealed that ST218-KL57 and ST23-KL57 strains clustered on a deeper branch, with a significant genetic distance from ST23-KL1 strains. The pruned variants derived from this analysis were subsequently used for Principal Component Analysis (PCA) and SNP distribution analysis, with the results further confirming that the genetic distance between ST218-KL57 and ST23-KL57 is relatively small, while both are clearly separated from the traditional hvKp ST23-KL1 strains (Supplementary Fig. 3). Notably, this is unexpected, since isolates of the same ST are usually assumed to be more closely related, yet the PCA revealed the contrary.
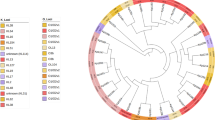
To elucidate the population structure and genetic relatedness between ST218-KL57 and ST23-KL57 strains, while detecting subtle population subdivisions, we performed fine-scale genetic analysis using fineSTRUCTURE complemented by phylogenetic reconstruction. The fineSTRUCTURE results indicated the formation of three discrete clusters, which closely resemble the population structure shaped by geographic isolation, with the clustering closely aligning with regional geographic information (Fig. 2A). The Maximum-likelihood (ML) tree results also showed that ST218-KL57 and ST23-KL57 strains formed two distinct subclades (Fig. 2B). The pruned variants were subsequently used for PCA, and the SNP distance distribution further supported these findings (Fig. 2C; D). Consistent with the above results (Fig. 2A; B), PCA of the ST23-KL57 and ST218-KL57 dataset confirmed their genetic distinction and revealed two divergent groups within ST218-KL57, suggesting intra-lineage heterogeneity. The maximum SNP number between the two was approximately 4000 ( ~ 0.08%), which is significantly smaller compared to the SNP distances observed between other lineages (~0.5%) (Fig. 2)19. The above findings revealed that strains belonging to the same ST exhibited substantial genetic divergence, whereas strains of different STs were genetically closely related. Therefore, we employed a high-resolution core-genome multilocus sequence typing (cgMLST) scheme based on 629 loci20. The results showed that ST23-KL1 and ST23-KL57 were classified as SL23 and SL218, with corresponding LIN codes of 0_0_429_0_37_2_0_0_0_0 and 0_0_115_0_0_0_0_1_*_*, respectively. Notably, ST218-KL57 was also assigned to SL218 (identical to ST23-KL57), with LIN codes of 0_0_115_1_6_*_*_*_*_*. These results indicate that traditional 7-locus MLST is insufficient for accurately resolving the population structure of ST23 strains.
A Geographic origins and ST of isolates are shown by annotation bars. The heatmap gradient indicates a genetic diversity cline rather than discrete clusters. B Maximum-likelihood phylogenetic tree of ST218-KL57 and ST23-KL57 isolates based on core SNPs. Within the innermost ring, orange indicates ST23-KL57 isolates, while purple indicates ST218-KL57 isolates. C Principal Component Analysis (PCA) of ST218-KL57 and ST23-KL57 isolates based on pruned SNPs. The inset scree plot shows the variance explained by the first 20 principal components. D Distribution of pairwise SNP distances between ST218-KL57 and ST23-KL57 isolates. E Nucleotide polymorphisms distinguishing ST218-KL57 isolates from ST23-KL57 isolates. The number of SNPs per 1,000 bp is shown. F Nucleotide polymorphisms distinguishing ST395 isolates from ST23-KL57 isolates. The number of SNPs per 1,000 bp is shown. ST sequence type, KL capsule locus, SNP single-nucleotide polymorphism, cKp classical K. pneumoniae, CRKP carbapenem-resistant K. pneumoniae, hvKp, hypervirulent K. pneumoniae, CR-hvKp carbapenem-resistant hypervirulent K. pneumoniae. Source data are provided as a Source Data file.
Comparative genomic analysis reveals functional differences between ST218-KL57 and ST23-KL57 strains
Given the high genomic similarity between ST218-KL57 and ST23-KL57, we performed pan-genome and comparative genomic analyses to elucidate their shared and distinct genetic features. Both lineages displayed an open pan-genome (Fig. 3A), indicating ongoing gene acquisition and diversification. A total of 12,988 genes were identified across the two lineages, with 3,683 constituting the core genome (present in ≥ 99% of strains) (Supplementary Fig. 4). Functional annotation using Clusters of Orthologous Groups (COG) further demonstrated distinct enrichment patterns in key functional categories between the two strains (Supplementary Fig. 5), suggesting potential differences in metabolic and adaptive strategies. 318 signature genes in ST218-KL57 (odds ratio (OR) > 1, p < 0.05) and 897 signature genes in ST23-KL57 (OR < 1, p < 0.05) were significantly enriched (Fig. 3B; Supplementary data 2). Functional annotation and COG clustering demonstrated that ST218-KL57 signature genes were predominantly associated with amino acid transport and metabolism and inorganic ion transport and metabolism (Fig. 3C), which play crucial roles in bacterial metabolism, nutrient uptake, environmental adaptation, and transcriptional regulation, with a subset linked to virulence-associated functions and horizontal gene transfer (HGT). In contrast, ST23-KL57 strains were significantly enriched in information storage and processing, including DNA replication, recombination, repair, and transcription, mainly associated with antibiotic resistance mechanisms, mobile genetic elements (transposons, plasmids, and prophage-related genes), and stress response systems (Fig. 3B; C; Supplementary data 2).
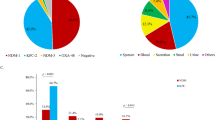
A The pan-genome curves of ST218-KL57 and ST23-KL57 isolates, along with the combined pan-genome curve for both ST218-KL57 and ST23-KL57. The blue, purple, and green colors represent the pangenome curves of ST23-KL57, ST218-KL57, and the combined strains of ST218-KL57 and ST23-KL57, respectively. B Characteristic genes distinguishing ST218-KL57 and ST23-KL57 isolates. Gene presence/absence associations were analyzed using Scoary with two-sided Fisher’s exact tests. Multiple testing was controlled using the Benjamini–Hochberg false-discovery rate (FDR) method, and gene clusters with FDR-adjusted p < 0.05 were considered significant. Genes with odds ratio (OR) > 1 are labeled as ‘Positive’ (enriched in ST218-KL57), whereas genes with OR < 1 are labeled as ‘Negative’ (enriched in ST23-KL57). C Clusters of Orthologous Groups (COG) clustering of characteristic genes for ST218-KL57 and ST23-KL57 isolates, with the size of the circles representing the proportion. D Comparison of virulence gene numbers between ST218-KL57 and ST23-KL57 isolates. E Comparison of resistance gene numbers between ST218-KL57 and ST23-KL57 isolates. For each group, number of genes were summarized using boxplots in which the central line represents the median, the box boundaries indicate the interquartile range (IQR, 25th–75th percentile), and the whiskers extend to 1.5 × IQR. Individual data points were overlaid using jittered dots. Group differences were evaluated using the Wilcoxon rank-sum test (two-tailed), and p-values were reported. Source data are provided as a Source Data file.
ST218-KL57 and ST23-KL57 differ in virulence and antimicrobial resistance genes
A detailed analysis concentrated on the virulence and antimicrobial resistance genes (ARGs). ST218-KL57 contained significantly more virulence genes than ST23-KL57 strains (p = 0.0032; Wilcoxon rank test) (Fig. 3D; Supplementary data 3), while ST23-KL57 had a higher number of ARGs compared to ST218-KL57 (p < 2.2e-16; Wilcoxon rank test) (Fig. 3E; Supplementary data 3). Further analysis focused on key virulence genes (peg-344, iucA, iroN, rmpADC, rmpA2, yptP, clbH) and virulence determinants (ICEKp, GIE492, and all island), as these elements are well-known for their crucial roles in the formation and spread of hvKp21,22,23,24,25,26,27,28. Comparative analysis between the two groups revealed that the virulence genes peg-344, iroN, rmpADC, rmpA2, and ybtP were significantly more prevalent in ST218-KL57 strains than in ST23-KL57 strains (p < 0.05), whereas no significant difference was observed in the distribution of iucA between the two lineages (Table 1). In contrast, the prevalence of ICEKp3 was higher in ST23-KL57 strains compared to ST218-KL57 strains. Regarding carbapenem resistance genes, no significant difference was observed in the prevalence of the blaKPC-2 between ST23-KL57 and ST218-KL57 strains (p = 0.744). However, the prevalence of blaNDM-1 and blaOXA-48 was significantly higher in ST23-KL57 strains compared to ST218-KL57 strains (p < 0.05; Table 1).
Adaptive structural variations of the virulence plasmid in ST218-KL57 and ST23-KL57 strains
Using a set of 9 complete genomes (including 4 newly sequenced in this study and 5 retrieved from NCBI) (Fig. 4), we found that the majority of ST218-KL57 and ST23-KL57 strains harbor the virulence plasmid replicon, repB_KLEB_VIR (Supplementary data 4), which is the predominant plasmid of the sublineage of CG23 (CG23-I)13 and is also present to the hvKp reference strain SGH10. The virulence plasmid backbone structures of both ST218-KL57 and ST23-KL57 strains were identical to the reference plasmid pSGH10, indicating a high degree of conservation (Fig. 4A; B; Supplementary Fig.6).
A Alignment of the ST218-KL57 virulence plasmids with pSGH10. B Alignment of the ST23-KL57 virulence plasmids with pSGH10. The innermost circle shows the GC skew (GC-skew, calculated as (G − C)/(G + C)) with a window size of 500 bp and a step size of 20 bp. Inward indicates that the G content is relatively higher than the C content (G > C), while outward indicates that the G content is relatively lower than the C content (G < C); The next inner circle shows the G and C content. Inward indicates that the G and C content is below the average, while outward indicates that it is above the average; The outermost circle shows the arrangement of genes, where forward genes are displayed in the clockwise direction and reverse genes in the counterclockwise direction. C–E Organization of the deletion regions in the virulence plasmids of ST218-KL57 and ST23-KL57, compared to the corresponding reference regions. Genes are represented by arrows, and shaded regions indicate areas of homology (> 95% nucleotide identity). The accession numbers for pSGH10, pKp_Goe_414-2, and pZ360 are NZ_CP025081.1, NZ_CP018338.1, and NZ_JBQABA010000002.1, respectively. Source data are provided as a Source Data file.
However, we found substantial genotype-specific structural variations between ST218-KL57 and ST23-KL57 and the reference plasmid, characterized by varying extents of genomic deletions as follows: (i) both strains possess a ~ 5 kb deletion, which encompasses the loss of gene fragments related to disulfide bond formation and redox function, mediated by IS1A (Fig. 4C); (ii) both strains showed a significant ~7 kb deletion, including genes associated with nitrogen utilization and secondary metabolite biosynthesis (Fig. 4D); (iii) the ST23-KL57 strain harbors a an additional ~20 kb deletion, which includes iroBCD and iroN, key virulence genes in K. pneumoniae that are crucial for iron acquisition under iron-limited conditions, thereby enhancing pathogenicity29 (Fig. 4E).
ST218 strains divide into two clades that differ in virulence determinants
Phylogenetic reconstruction based on maximum-likelihood phylogeny, fineSTRUCTURE clustering and SNP-based population stratification showed two deeply divergent sublineages within ST218 (designated ST218-Clade 1 and ST218-Clade 2) (Fig. 5; Fig. 2A). When PCA was applied to the ST218-KL57 dataset, the two clades identified in Fig. 5A could be clearly separated along PC1 (Fig. 5B). In contrast, the distribution along PC2 was continuous, indicating no further sub-structuring within this lineage. ST218-Clade 1 and ST218-Clade 2 exhibited striking epidemiological and genomic divergence. While the ST218-Clade 1 disseminated globally, the ST218-Clade 2 remained predominantly restricted to Asia and adjacent regions. The virulence plasmid-associated loci iuc, iro, peg-344, rmpADC, and rmpA2 were present in the vast majority of ST218 genomes (Fig. 5A), with each of these five genes found in over 86% of the cases (Supplementary data 5). Only a small fraction of the strains exhibited deletions in these virulence loci, with such strains predominantly located within the ST218-Clade 1 sublineages (Fig. 5A; D). Other virulence loci, found within the chromosome of ST218, included the ICEKp (the region where the ybt locus resides), which was present in 70.5% of ST218 strains (Supplementary data 5). This locus was predominantly represented by the conserved types ICEKp3 and ICEKp4. However, strains in the ST218-Clade 1 sublineages also exhibited a loss of the ICEKp locus. The genomic island GIE492 and all island were absent in all ST218 genomes.
A Maximum-likelihood phylogenetic tree of ST218-Clade 1 and ST218-Clade 2 isolates based on core SNPs. The orange color represents ST218-Clade 1 isolates, while the pink color represents ST218-Clade 2 isolates. B Principal Component Analysis (PCA) of ST218-Clade 1 and ST218-Clade 2 isolates based on pruned SNPs. The inset scree plot shows the variance explained by the first 20 principal components. C Distribution of pairwise SNP distances between ST218-Clade 1 and ST218-Clade 2 isolates. D The bubble chart displays the differences in virulence factors between the two groups, with the size of the bubbles representing the proportion. ST sequence type, KL capsule locus, SNP single-nucleotide polymorphism, cKp classical K. pneumoniae, CRKP carbapenem-resistant K. pneumoniae, hvKp hypervirulent K. pneumoniae, CR-hvKp carbapenem-resistant hypervirulent K. pneumoniae, VFs virulence factors. Source data are provided as a Source Data file.
Recombination driven emergence of ST23-KL57 lineage
Recombination has been shown to occur in K. pneumoniae13,30. In this study, we conducted an in-depth investigation into the recombination and evolutionary events that have occurred between these two lineages. SNP calling was performed for all 122 ST218-KL57 isolates against the genome of an ST23-KL57 isolate Kp_Goe_154414 (GCA_001902335.1), using MUMmer31. The number of SNPs at each genomic position was then calculated. Analysis of SNP distribution revealed that polymorphisms were concentrated within the 0.5–1.5 Mb region, whereas few or no SNPs were detected in the 0–0.5 Mb and 1.5–5 Mb regions (Supplementary Fig.7). The results revealed that the genome of ST218-KL57 shares a high degree of similarity with approximately 4 Mb of the ST23-KL57 genome (Fig. 2E; Supplementary Fig. 7), with nearly 80% of the genome highly conserved between the two lineages.
We then focused on identifying isolates whose SNP distribution relative to ST23-KL57 occurred outside the 0.5–1.5 Mb interval. We calculated the number of SNPs within this region between ST23-KL57 isolate and all available K. pneumoniae isolates in the public database. We further found that ST395 isolates harbored very few SNPs in the 0.5–1.5 Mb region, whereas abundant SNPs were distributed outside this interval (Fig. 2F; Supplementary Fig. 8). This pattern was complementary to that observed in ST218-KL57 isolates. To further validate our findings, we analyzed the SNP distributions of one representative isolate from each ST included in the public database used in this study. The results revealed that, except for ST395, no other STs exhibited this complementary pattern (Supplementary Fig. 9).
Based on these findings, we propose that ST23-KL57 strains are hybrid strains composed of genomic DNA derived approximately 80% from ST218-KL57—a hypervirulent lineage prevalent in Asia that typically harbors virulence plasmids—and about 20% from ST395, a multidrug-resistant lineage predominantly found in Europe and commonly associated with resistance plasmids. This genomic composition is presumably the result of a major chromosomal replacement event, ultimately giving rise to ST23-KL57 strains that harbor both virulence and resistance plasmids. However, the virulence plasmid carried by ST23-KL57 differs slightly from that of the original ST218-KL57 strains, notably lacking iroBCD and iroN genes (Fig. 4; Supplementary Fig.6; Supplementary data 3).
Virulence phenotypic indicates moderate pathogenicity of ST218 isolates
To gain a deeper understanding of the virulence characteristics of ST218 isolates, we evaluated their virulence phenotypes using the string test, quantification of biofilm formation, and a mouse infection model. The hypermucoviscosity phenotype of all 13 sequenced ST218 isolates was assessed using the string test. Compared to the negative control XWkp27 isolate32, which is non-hypermucoid, the ST218 strains exhibited strong hypermucoviscosity (≥ 5 mm). ST218 isolates displayed robust biofilm formation, indicating their enhanced ability to resist host immune defenses and clinical treatment pressures (Fig. 6A)33. Furthermore, to assess the virulence of ST218 strains, we randomly selected four isolates (Z360, Z362, 7301, and 2022.0044) for in vivo experiments. The virulence of the ST218 isolates was lower compared to hvKp positive control strains 373 (ST23-KL1) and 65 (ST65-KL2), but significantly higher than the negative control, based on survival (Fig. 6B). Taken together, these findings suggest that the ST218 strains exhibit moderate virulence.
A Biofilm formation of strains was quantified by the crystal violet assay at 590 nm after 24 h of incubation, and results are shown as box plots. For each box plot, the central line represents the median, the box spans the interquartile range (25th–75th percentile), and the whiskers indicate the minimum and maximum values; individual points represent independent biological replicates (n = 5). All strains showed significant differences compared with the negative control strain ATCC 13883 (two-sided Wilcoxon rank-sum test); The statistical significance is shown by the number of asterisks as follows: * p < 0.05, ** p < 0.01, *** p < 0.001. B Kaplan-Meier survival analysis of mouse model to assess the virulence of six K. pneumoniae strains (each group consists of 10 mice). The control group was injected with PBS. Source data are provided as a Source Data file.
Discussion
K. pneumoniae CG23, predominantly composed of ST23, constitutes a distinct lineage associated with a unique repertoire of virulence determinants34. The majority of CG23 strains display the K1 serotype35,36,37, and a robust association has been observed with liver abscesses38. A former study analyzed 97 CG23 hvKp strains (all belong to KL1) from humans and horses13, while we included a large collection of CG23 strains. This revealed a genetically divergent subgroup, ST23-KL57, distinguished from the conventional ST23-KL1 lineage, which is currently under scrutiny by the WHO6 and ECDC7 due to its association with carbapenem resistance. Recognizing the emergence of this strain as a potential indicator of multidrug resistance, particularly carbapenem resistance, within the hvKp ST23 lineage, we screened and sequenced K. pneumoniae isolates from hospitals in China. While no ST23-KL57 strains were found in our collection, we identified ST218-KL57 strains, which are prevalent in Asia, particularly in China. These ST218-KL57 strains exhibit a close genetic relationship with ST23-KL57 strains reported in Europe.
High-resolution genotyping (cgMLST, fineSTRUCTURE, and PCA) resolved distinct sublineages within ST23 and ST218, confirming the genetic separation of ST23-KL1, ST23-KL57, and ST218-KL57. These findings underscore the limitations of traditional 7-locus MLST and highlight the necessity of higher-resolution approaches for accurate strain classification. In addition to evolutionary divergence, ST23-KL57 and ST218-KL57 differ from ST23-KL1 in lacking virulence loci such as GIE492 and the all island8,28. Notably, ST23-KL57 harbored multiple carbapenemase genes, whereas ST218-KL57 exhibited intermediate resistance and ST23-KL1 remained largely susceptible. The limited ST-level resolution likely stems from recombination whereby ST23-KL57, originating from an ST218-KL57 background, convergently acquired the classical ST23 allelic profile. However, our in-depth analyses revealed that it differs substantially from the traditional ST23-KL1 lineage, and in fact, shares little to no genetic relatedness with it. To date, many studies investigating ST23 strains have focused primarily on sequence type while neglecting the capsular type16. Such an approach may lead to inaccurate classification and interpretation. Here, we emphasize the importance of incorporating both cgMLST and capsular typing in future analyses of K. pneumoniae, and we suggest that the research community adopt this genotyping strategy.
Virulence plasmids carrying conserved genes (peg-344, iucA, iroN, rmpADC, and rmpA2) are well-documented in hvKp24,39. Structural analysis revealed that both ST23-KL57 and ST218-KL57 plasmids share the same Inc type as the hypervirulent reference strain SGH10 but lack two regions found in SGH10; additionally, the ST23-KL57 plasmid lacking the iroN locus40. The iroN gene, part of the iroBCDN cluster, encodes salmochelin, a glucosylated enterobactin that evades lipocalin-2 binding and inflammatory responses41. Over 90% of K. pneumoniae strains associated with pyogenic liver abscesses harbor salmochelin42, making it a hallmark of hvKp virulence. The loss of iroN suggests an evolutionary trade-off in K. pneumoniae, as strains with both high virulence and multidrug resistance are rare due to the associated metabolic cost43.
Recombination events and the replacement of large chromosomal regions have been observed in numerous bacterial species, with several documented instances where hybrid strains are linked to epidemiological success, particularly within the Enterobacteriaceae family44. In K. pneumoniae, the well-known recombination event is the formation of ST258. ST11 (which differs from ST258 by a single housekeeping gene locus) primarily spreads in Asian countries, particularly in China, while the new ST258 strains, formed by recombination between ST11 and ST442, have led to its spread in the USA30,45. Geographic variation was observed in the distribution of ST218-KL57 and ST23-KL57 lineages, distinguished by only one housekeeping gene. This minimal allelic variation supports the notion that ST23-KL57 likely arose from the ST218-KL57 background through a recent recombination or gene conversion event rather than long-term divergence. The two highly similar recombination events suggest that recombination in K. pneumoniae may be ongoing, underscoring the need for sustained surveillance. In addition, while our results suggest that ST218-KL57 may have originated and initially circulated in China, we acknowledge that the current dataset is constrained by uneven regional sampling intensity across regions. Future studies incorporating broader sampling from diverse sources and geographic areas will be essential to minimize potential biases and to provide a more accurate understanding of the global population structure and transmission dynamics of K. pneumoniae.
To investigate the impact of the newly identified recombination in ST23-KL57 isolates, we performed comparative genomic analyses focusing on multidrug resistance and virulence genes. In ST23-KL57, we examined genomic regions harboring resistance genes, mobile elements, and stress response systems. In contrast, ST218-KL57 strains typically lacked multidrug resistance plasmids such as IncFII and carried only virulence-associated plasmids like repB_KLEB_VIR. In the recombinant ST23-KL57 strains, however, both multidrug resistance and virulence plasmid types were present, suggesting that ST395, a potential recombination donor, contributed not only chromosomal DNA but also AMR plasmids. Consequently, the newly emerged ST23-KL57 strains pose a significant threat, as they combine both hypervirulence and multidrug resistance. The underlying mechanisms driving the fusion of virulence and resistance in these strains are believed to involve chromosomal and plasmid fusion events. Taken together, these observations provide evidence that ST23-KL57 strains are hybrid strains that originated from an ancestral ST218 strain, acquiring a contiguous chromosomal segment from an ST395 strain through DNA recombination/replacement. These recombination events have resulted in the emergence of ST23-KL57, which combines both virulence and antibiotic resistance traits, contributing to its successful spread in Europe.
Phenotypic evaluation of ST218-KL57 isolates revealed string test-positive isolates consistent with rmpADC and rmpA2 gene carriage46,47,48, enhanced biofilm formation, and intermediate virulence in mice, which was greater than that of control strains but lower than that of canonical hypervirulent lineages (ST23-KL1 and ST65-KL2). These results highlight the multifactorial nature of hypervirulence, shaped by both chromosomal and plasmid determinants. Notably, one ST218-KL57 strain also carried blaNDM-1, indicating the concerning convergence of resistance and virulence18. The convergence of multidrug resistance and critical virulence factors in these strains remains concerning, as even moderate virulence in carbapenem-resistant strains also poses a significant public health threat.
In conclusion, our study provides a systematic genomic characterization of the emerging K. pneumoniae lineages ST218-KL57 and ST23-KL57, offering key insights into their evolution, resistance, and virulence. While CG23, primarily comprising ST23, is a recognized hypervirulent K1 serotype, ST23-KL57 represents a distinct, carbapenem-resistant subgroup that cluster with ST218-KL57 within the SL218 lineage. These findings highlight the value of high-resolution genomic typing and demonstrate the dynamic evolution of K. pneumoniae under selective pressure. Enhanced global surveillance is crucial to detect and respond to these emerging threats.
Methods
Ethics statement
All our research complies with the relevant ethical regulations. This study was approved by the Medical Ethics Committee of Sir Run Run Shaw Hospital, Zhejiang University School of Medicine (Approval No. 2022-0335). Written informed consent from the patients was exempted by the Ethics Committee of Sir Run Run Shaw Hospital.
All animal procedures were performed in accordance with the laboratory animal care and use guidelines of the Institutional Animal Care and Ethics Committee of Sir Run Run Shaw Hospital, Zhejiang University School of Medicine. Approval was obtained from the Ethics Committee of Sir Run Run Shaw Hospital, Zhejiang University School of Medicine (Approval No. SRRSH202202075).
Isolation of bacterial strains, genomic DNA extraction, genome sequencing, assembly, and quality control
A total of 3,118 K. pneumoniae isolates were collected between 2021 and 2024 from six hospitals across Zhejiang Province, China. All primers used in this study were designed using SnapGene (version 6.2.2) and synthesized by Sangon Biotech Co., Ltd. (Shanghai, China). Primary screening of ST218-KL57 and ST23-KL57 isolates was performed by PCR using the following primers: forward 5’-TTGGCTTGACGAAGCAGAAATGA-3’ and reverse 5’-CCCGGAGGCTCACATTCTTTC-3’. There were no ST23-KL57 isolates identified in this study. The whole genomes of ST218 strains were sequenced. Genomic DNA was isolated from the strains and used to create shotgun paired-end libraries with an average insert size of around 350 bp49, using the TruSeq DNA Sample Preparation Kit (Illumina, San Diego, CA, USA). The sequencing data were quality-checked, assembled de novo using Shovill v1.1.0 (--trim --minlen 200), and subjected to MLST analysis to confirm STs using mlst v2.19.0. After in silico MLST analysis, 13 high-quality ST218 isolates (1 isolate quality control is not qualified) were identified and selected for further analysis (Supplementary data 1).
Furthermore, five of these ST218 isolates were randomly selected for additional sequencing using the long-read Nanopore MinION to obtain their complete genome. For Nanopore sequencing, a MinION sequencing library was prepared using the Nanopore Ligation Sequencing Kit (Oxford Nanopore, Oxford, UK). The library was sequenced using an R9.4.1 MinION flow cell (FlO-MIN106) for a 24-hour run with MinKNOW v2.0, employing default settings. The raw Nanopore signal data in FAST5 format were base-called and converted to FASTQ format in real-time using Guppy v3.3.0. Subsequently, Porechop v0.2.4 was used to trim barcode and adapter sequences (https://github.com/rrwick/Porechop). To improve assembly, Filtlong v0.2.0, a quality filtering tool for Nanopore reads, was used to remove sequences shorter than 3000 bases and those with mean quality scores below 1250. De novo sequence assembly was performed using Flye 2.8.3 (parameters: --nano-raw).
Additionally, we downloaded a total of 75,954 assembled K. pneumoniae genomes from NCBI (last accessed July 31, 2024). The data quality control criteria required that the following three conditions be met simultaneously: (1) genome size between 5 and 7 Mb; (2) ANI value greater than 95% compared to the hvKp reference genome SGH10, as calculated by pyani v0.2.12; and (3) completeness greater than 95% and contamination less than 5%, as assessed by CheckM v1.2.3. After filtering out low-quality sequencing samples, 69,083 high-quality isolates remained (Supplementary Fig.1). For the quality-controlled strains, we conducted in silico MLST analysis, which identified 109 ST218 isolates. In total, our study included 122 ST218 isolates, 13 sequenced in this study and 109 downloaded from NCBI, for further analysis (Fig. 1; Supplementary data 1).
Multilocus sequence typing and capsule locus typing
Using goeBURST (http://www.phyloviz.net/goeburst), all STs from the Klebsiella MLST database (last accessed December 14, 2024) were grouped into distinct clonal groups (CGs). KL types were determined using Kaptive (https://github.com/klebgenomics/Kaptive). Core genome multilocus sequence typing (cgMLST) was performed via the online platform PathogenWatch (https://pathogen.watch/).
Identification of virulence factors, virulence-associated accessory elements, antibiotic resistance genes, and plasmid replicon types
We used Abricate v1.0.1 (https://github.com/tseemann/abricate) with the parameters set to -minid 95 -mincov 95 to detect ARGs, virulence genes, and plasmid replicon types. ARGs were identified using the ResFinder database51, virulence factors were recognized through the VFDB database52, and plasmid replicon types were determined by the PlasmidFinder database (https://cge.food.dtu.dk/services/PlasmidFinder/) to pinpoint the Inc type.
The ICEKp variants found in all isolates were identified using Kleborate26 (https://github.com/klebgenomics/Kleborate). Due to the high conservation of GIE4928 and all island28 among K. pneumoniae strains, reference sequences for both GIE492 and the all island were obtained from the hvKp reference genome SGH10. These elements were subsequently detected across all isolates by conducting BLAST searches against the extracted reference sequences.
Construction of maximum-likelihood (ML) clustering trees
The strains were aligned to the K. pneumoniae SGH10 reference genome13 using Snippy (https://github.com/tseemann/snippy) to identify core single nucleotide polymorphisms (SNPs), which were then used to construct a ML phylogenetic tree for all strains in the analysis. Pairwise SNP distances between isolates were computed with snp-dists (https://github.com/tseemann/snp-dists). The ML tree was generated with IQ-TREE v2.1.4 (http://www.iqtree.org), utilizing 1,000 bootstrap replicates for reliable phylogenetic inference. The final tree was visualized through iTOL (https://itol.embl.de).
Pangenomic profiling, comparative genomic analysis, and functional enrichment analysis
To further investigate the relationship and differences between these strains, all publicly available ST23-KL1 (n = 1,491) and ST23-KL57 (n = 45) strains were retrieved from NCBI (last accessed July 31, 2024) for pangenome and comparative genomic analyses. To conduct an in-depth analysis of the functional characteristics of the bacterial genomes, we initially utilized Prokka v1.14.6 to predict the open reading frames. The Generic Feature Format (GFF) files generated by Prokka53 were subsequently used as input for Roary54, which facilitated the pangenome analysis of the described isolates. We then applied Scoary55 to identify genes that were significantly enriched in the ST218-KL57 and ST23-KL57 isolates. This approach, which relies on gene presence/absence, is characterized by rigorous and precise statistical properties. Gene clusters were deemed significantly enriched when the Benjamini-Hochberg FDR-adjusted p-value was <0.05. Clusters with an OR > 1 were considered to be linked to ST218-KL57 isolates, whereas those with an OR < 1 were associated with ST23-KL57 isolates. Gene annotation was performed using COG annotation56. COG annotation was performed using EggNOG-mapper v257 (https://github.com/smarted/eggnog-mapper), and we conducted this for all genes and the feature genes of each group.
Population assignment and fineSTRUCTURE analysis
The population structure of ST218/ST23-KL57 was determined using chromosome painting and population clustering analyses. We utilized ChromoPainter, which is part of fineSTRUCTURE (v4.1.0)58, to model the sharing of DNA segments, thereby estimating the genomic fragments inherited by each recipient haplotype from potential donors. This data was then organized into a co-ancestry matrix. Using this matrix, fineSTRUCTURE v4.1.0 clustered the isolates into distinct groups through 100,000 iterations for both the burn-in and Markov Chain Monte Carlo (MCMC) phases.
Principal component analysis
To further substantiate the genomic divergence between ST218 and ST23, we performed a PCA on the ST218 and ST23 dataset. Before PCA, PLINK v1.959 was used to remove loci with a minimum allele frequency (MAF) below 2% from the core SNPs. Subsequently, linkage disequilibrium (LD) pruning was carried out to eliminate highly correlated SNPs. For any SNP pair with an r² value greater than 0.1 within a 50 kb window, one of the SNPs was discarded. The window was then advanced by 10 bp, and the pruning process was repeated to ensure coverage. The pruned variants were then utilized for PCA on the ST218/ST23 dataset, with PLINK v1.959 extracting and reporting the first 20 principal components by default.
Structural comparison and annotation of virulence plasmids
Open reading frames (ORFs) and pseudogenes were identified using RAST 2.060, combined with BLASTP/BLASTN searches61 against the UniProtKB/Swiss-Prot database62 and the RefSeq database63. Annotations for mobile elements and other genomic features were obtained through online resources such as ISfinder64, INTEGRALL65, and Tn Number Registry66. Sequence comparisons, both pairwise and multiple, were performed using MUSCLE 3.8.3167 and BLASTN, respectively. Gene organization diagrams were created using Inkscape 0.48.1 (https://inkscape.org/en/), while the alignment rings for these diagrams, based on the complete genomes, were visualized using BRIG68. The conjugation probabilities of the complete virulence plasmids were evaluated with oriTfinder (https://tool-mml.sjtu.edu.cn/oriTfinder/oriTfinder.html)69.
SNP calling and recombination analysis
MUMmer v3.1 was applied for SNP calling using the ST23-KL57 isolate (GCA_001902335.1) as the reference genome, particularly, nucmer was used for genome alignment, delta-filter for refining the alignments, and show-snps for identifying SNP positions. For each ST218-KL57 isolate, the number of SNPs per 1000 bp was calculated and visualized using custom Python scripts, revealing that SNPs were almost exclusively concentrated within the 0.5–1.5 Mb region (Supplementary Fig. 7). Subsequently, the same approach was applied to calculate the number of SNPs in the 0.5–1.5 Mb interval across all isolates, which showed that ST395 strains exhibited the lowest SNP density in this region (Supplementary Fig. 8). Finally, the SNP distributions of isolates representing each distinct sequence type (apart from ST218 and ST395) were visualized using the same method (Supplementary Fig. 9).
Hypermucoviscosity phenotype identification
The hypermucoviscous phenotypes were determined by the string test, and the isolates that generated strings >5 mm in length after stretching with the tip of a sterile inoculation loop were defined to have a hypermucoviscous phenotype70.
Biofilm formation assay
The quantity of biofilm produced was measured following the method described in a previous study71. Two biofilm-positive isolates (373 and 651) and one negative control (ATCC 13883) were included as controls. Briefly, bacteria were grown overnight in Luria-Bertani (LB) medium, diluted 1:100 in fresh medium and cultured at 37 °C till OD600 values of 0.6–0.8 were reached. Two hundred microliters of culture per well were transferred to 96-well polystyrene microtiter plates at 37 °C for 24 h. After three washes with 200 μL of PBS, the biofilms were fixed with methanol for 15 min. The methanol was removed, and the plates were again washed three times. The biofilms were stained with 200 μL crystal violet (CV) for 30 min, and then the unbound CV dye was removed by washing three times. After destaining with 95% ethanol for 5 min, the biofilms were measured with a SpectraMax® ABS Microplate Reader (Molecular Devices, USA) at 590 nm. Statistical analyses were performed in R (version 4.3.1). The Wilcoxon rank-sum test was used to compare the biofilm-forming ability of all strains with that of the negative control strain ATCC 13883. Statistical significance is indicated by asterisks as follows: p < 0.05, ** p < 0.01, *** p < 0.001.
Mouse intraperitoneal infection model
Specific pathogen-free (SPF) female CD-1 mice (5–6 weeks old) were used for animal experiments. The mice were maintained under SPF conditions in-house at 19–26 °C with 40–70% humidity and a 12 h light/dark cycle, and were provided autoclaved chow, water, and corncob bedding. This study does not involve the influence of sex hormones on K. pneumoniae infection, and the sex of the animals does not affect the experimental outcomes. Female CD-1 mice, aged five to six weeks and weighing approximately 17 g, were randomly divided into groups to assess the virulence of the ST218 isolates. Each group consists of 10 mice. The bacteria were grown in LB broth until the logarithmic phase, and each mouse was injected with 4 × 108 CFU bacteria. The negative control group was treated with PBS, while the positive control group was treated with 373 (ST23) and 651 (ST65). The mortality rates were observed for a week. Survival analysis was performed using the Kaplan-Meier method and compared by the log-rank (Mantel-Cox) test in GraphPad Prism (version 9.2.0).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All sequenced data generated in this study have been deposited in the NCBI database under the BioProject number: PRJNA1286124. Other genome sequences used in this study are available in the NCBI GenBank database, with the accession numbers for the individual sequences provided in Supplementary data 1. Source data are provided with this paper.
Code availability
Code used in this work is available on https://github.com/rzzju/KP.
References
Wu, Y. et al. Global Evolution and Geographic Diversity of Hypervirulent Carbapenem-resistant Klebsiella Pneumoniae. Lancet Infect. Dis. 22, 761–762 (2022).
Wu, Y. et al. Global Phylogeography and Genomic Epidemiology of Carbapenem-Resistant blaOXA-232-Carrying Klebsiella pneumoniae Sequence Type 15 Lineage. Emerg. Infect. Dis. 29, 2246–2256 (2023).
Russo, T. A. & Marr, C. M. Hypervirulent Klebsiella pneumoniae. Clin. Microbiol. Rev. 32, e00001-19 (2019).
Bengoechea, J. A. & Sa Pessoa, J. Klebsiella pneumoniae infection biology: living to counteract host defences. FEMS Microbiol. Rev. 43, 123–144 (2019).
Kochan, T. J. et al. Klebsiella Pneumoniae Clinical Isolates with Features of Both Multidrug-resistance And Hypervirulence have Unexpectedly Low Virulence. Nat. Commun. 14, 7962 (2023).
WHO. Antimicrobial Resistance, Hypervirulent Klebsiella pneumoniae - Global situation. https://www.who.int/emergencies/disease-outbreak-news/item/2024-DON527 (2024).
ECDC. Emergence of hypervirulent Klebsiella pneumoniae ST23 carrying carbapenemase genes in EU/EEA countries, first update. https://www.ecdc.europa.eu/en/publications-data/risk-assessment-emergence-hypervirulent-klebsiella-pneumoniae-eu-eea (2024). https://doi.org/10.2900/993023
Tan, Y. H. et al. Hypervirulent Klebsiella pneumoniae employs genomic island encoded toxins against bacterial competitors in the gut. ISME J. 18, wrae054 (2024).
Russo, T. A., Lebreton, F. & McGann, P. T. A Step Forward in Hypervirulent Klebsiella pneumoniae Diagnostics. Emerg. Infect. Dis. 31, 1–3 (2025).
Xu, Y. et al. Mobilization of the nonconjugative virulence plasmid from hypervirulent Klebsiella pneumoniae. Genome Med. 13, 119 (2021).
Liu, C. et al. An increased prevalence of carbapenem-resistant hypervirulent Klebsiella pneumoniae associated with the COVID-19 pandemic. Drug Resist Updat 77, 101124 (2024).
Choby, J. E., Howard-Anderson, J. & Weiss, D. S. Hypervirulent Klebsiella pneumoniae - clinical and molecular perspectives. J. Intern. Med. 287, 283–300 (2020).
Lam, M. M. C. et al. Population genomics of hypervirulent Klebsiella pneumoniae clonal-group 23 reveals early emergence and rapid global dissemination. Nat. Commun. 9, 2703 (2018).
Nielsen, K. L. et al. Within-patient horizontal transfer of pOXA-48 from a hypervirulent Klebsiella pneumoniae SL218 to Serratia marcescens following spread of the K. pneumoniae isolate among hospitalised patients, Denmark, 2021. Euro Surveill. 28, 2300196 (2023).
Lam, M. M. C., Holt, K. E. & Wyres, K. L. Comment on: MDR carbapenemase-producing Klebsiella pneumoniae of the hypervirulence-associated ST23 clone in Poland, 2009-19. J. Antimicrob. Chemother. 78, 1132–1134 (2023).
Biedrzycka, M. et al. MDR carbapenemase-producing Klebsiella pneumoniae of the hypervirulence-associated ST23 clone in Poland, 2009-19. J. Antimicrob. Chemother. 77, 3367–3375 (2022).
Maguire, M. et al. Genomic and phylogenetic analysis of hypervirulent Klebsiella pneumoniae ST23 in Ireland. Microb. Genom. 11, 001373 (2025).
Yang, P. et al. ST218 Klebsiella pneumoniae became a high-risk clone for multidrug resistance and hypervirulence. BMC Microbiol 24, 56 (2024).
Holt, K. E. et al. Genomic analysis of diversity, population structure, virulence, and antimicrobial resistance in Klebsiella pneumoniae, an urgent threat to public health. Proc. Natl. Acad. Sci. USA 112, E3574–81 (2015).
Hennart, M. et al. A Dual Barcoding Approach to Bacterial Strain Nomenclature: Genomic Taxonomy of Klebsiella pneumoniae Strains. Mol. Biol. Evol. 39, msac135 (2022).
Bulger, J., MacDonald, U., Olson, R., Beanan, J. & Russo, T. A. Metabolite transporter PEG344 is required for full virulence of hypervirulent Klebsiella pneumoniae strain hvKP1 after pulmonary but not subcutaneous challenge. Infect. Immun. 85, e00093–17 (2017).
Nassif, X., Honoré, N., Vasselon, T., Cole, S. T. & Sansonetti, P. J. Positive control of colanic acid synthesis in Escherichia coli by rmpA and rmpB, two virulence-plasmid genes of Klebsiella pneumoniae. Mol. Microbiol. 3, 1349–1359 (1989).
Tang, H. L. et al. Correlation between Klebsiella pneumoniae carrying pLVPK-derived loci and abscess formation. Eur. J. Clin. Microbiol. Infect. Dis. 29, 689–698 (2010).
Russo, T. A. et al. Identification of Biomarkers for Differentiation of Hypervirulent Klebsiella pneumoniae from Classical K. pneumoniae. J. Clin. Microbiol. 56, e00776-18 (2018).
Lam, M. M. C. et al. Genetic diversity, mobilisation and spread of the yersiniabactin-encoding mobile element ICEKp in Klebsiella pneumoniae populations. Microb. Genom. 4, e000196 (2018).
Lam, M. M. C. et al. A genomic surveillance framework and genotyping tool for Klebsiella pneumoniae and its related species complex. Nat. Commun. 12, 4188 (2021).
Marcoleta, A. E., Berríos-Pastén, C., Nuñez, G., Monasterio, O. & Lagos, R. Klebsiella pneumoniae asparagine tDNAs are integration hotspots for different genomic islands encoding Microcin E492 production determinants and other putative virulence factors present in hypervirulent strains. Front. Microbiol. 7, 849 (2016).
Chou, H. C. et al. Isolation of a chromosomal region of Klebsiella pneumoniae associated with allantoin metabolism and liver infection. Infect. Immun. 72, 3783–3792 (2004).
Hsieh, P. F., Lin, T. L., Lee, C. Z., Tsai, S. F. & Wang, J. T. Serum-induced iron-acquisition systems and TonB contribute to virulence in Klebsiella pneumoniae causing primary pyogenic liver abscess. J. Infect. Dis. 197, 1717–1727 (2008).
Chen, L., Mathema, B., Pitout, J. D., DeLeo, F. R. & Kreiswirth, B. N. Epidemic Klebsiella pneumoniae ST258 is a hybrid strain. mBio 5, e01355–14 (2014).
Marçais, G. et al. MUMmer4: A fast and versatile genome alignment system. PLoS Comput. Biol. 14, e1005944 (2018).
Liu, N. et al. Genomic surveillance indicates clonal replacement of hypervirulent Klebsiella pneumoniae ST881 and ST29 lineage strains in vivo. Front. Microbiol. 15, 1375624 (2024).
Beckman, R. L., Cella, E., Azarian, T., Rendueles, O. & Fleeman, R. M. Diverse polysaccharide production and biofilm formation abilities of clinical Klebsiella pneumoniae. npj Biofilms Microbiomes 10, 151 (2024).
Struve, C. et al. Mapping the evolution of hypervirulent Klebsiella pneumoniae. mBio 6, e00630 (2015).
Shon, A. S., Bajwa, R. P. & Russo, T. A. Hypervirulent (hypermucoviscous) Klebsiella pneumoniae: a new and dangerous breed. Virulence 4, 107–118 (2013).
Lin, J. C. et al. Genotypes and virulence in serotype K2 Klebsiella pneumoniae from liver abscess and non-infectious carriers in Hong Kong, Singapore and Taiwan. Gut Pathog. 6, 21 (2014).
Harada, S., Ishii, Y., Saga, T., Aoki, K. & Tateda, K. Molecular epidemiology of Klebsiella pneumoniae K1 and K2 isolates in Japan. Diagn. Microbiol. Infect. Dis. 91, 354–359 (2018).
Siu, L. K. et al. Molecular typing and virulence analysis of serotype K1 Klebsiella pneumoniae strains isolated from liver abscess patients and stool samples from noninfectious subjects in Hong Kong, Singapore, and Taiwan. J. Clin. Microbiol. 49, 3761–3765 (2011).
Jiang, X. et al. The global genomic landscape of hypervirulent Klebsiella pneumoniae from 1932 to 2021. mLife 4, 378–396 (2025).
Müller, S. I., Valdebenito, M. & Hantke, K. Salmochelin, the long-overlooked catecholate siderophore of Salmonella. Biometals 22, 691–695 (2009).
Fischbach, M. A. et al. The pathogen-associated iroA gene cluster mediates bacterial evasion of lipocalin 2. Proc. Natl. Acad. Sci. U.S.A. 103, 16502–16507 (2006).
Bachman, M. A. et al. Klebsiella pneumoniae yersiniabactin promotes respiratory tract infection through evasion of lipocalin 2. Infect. Immun. 79, 3309–3316 (2011).
Chiarelli, A. et al. Diversity of mucoid to non-mucoid switch among carbapenemase-producing Klebsiella pneumoniae. BMC Microbiol 20, 325 (2020).
Bellais, S. et al. Capsular switching in group B Streptococcus CC17 hypervirulent clone: a future challenge for polysaccharide vaccine development. J. Infect. Dis. 206, 1745–1752 (2012).
Gaiarsa, S. et al. Genomic epidemiology of Klebsiella pneumoniae in Italy and novel insights into the origin and global evolution of its resistance to carbapenem antibiotics. Antimicrob. Agents Chemother. 59, 389–396 (2015).
Hsu, C. R., Lin, T. L., Chen, Y. C., Chou, H. C. & Wang, J. T. The role of Klebsiella pneumoniae rmpA in capsular polysaccharide synthesis and virulence revisited. Microbiology 157, 3446–3457 (2011).
Li, W. et al. Increasing occurrence of antimicrobial-resistant hypervirulent (hypermucoviscous) Klebsiella pneumoniae isolates in China. Clin. Infect. Dis. 58, 225–232 (2014).
Wacharotayankun, R. et al. Enhancement of extracapsular polysaccharide synthesis in Klebsiella pneumoniae by rmpA2, which shows homology to NtrC and FixJ. Infect. Immun. 61, 3164–3174 (1993).
Ribeiro, F. J. et al. Finished bacterial genomes from shotgun sequence data. Genome Res. 22, 2270–2277 (2012).
Breese, M. R. & Liu, Y. NGSUtils: a software suite for analyzing and manipulating next-generation sequencing datasets. Bioinformatics 29, 494–496 (2013).
Zankari, E. et al. Identification of acquired antimicrobial resistance genes. J. Antimicrob. Chemother. 67, 2640–2644 (2012).
Liu, B., Zheng, D., Zhou, S., Chen, L. & Yang, J. VFDB 2022: a general classification scheme for bacterial virulence factors. Nucleic Acids Res 50, D912–D917 (2022).
Seemann, T. Prokka: rapid prokaryotic genome annotation. Bioinformatics 30, 2068–2069 (2014).
Page, A. J. et al. Roary: rapid large-scale prokaryote pan genome analysis. Bioinformatics 31, 3691–3693 (2015).
Roder, T. et al. Scoary2: rapid association of phenotypic multi-omics data with microbial pan-genomes. Genome Biol. 25, 93 (2024).
Galperin, M. Y. et al. COG database update 2024. Nucleic Acids Res 53, D356–D363 (2025).
Cantalapiedra, C. P., Hernández-Plaza, A., Letunic, I., Bork, P. & Huerta-Cepas, J. eggNOG-mapper v2: Functional Annotation, Orthology Assignments, and Domain Prediction at the Metagenomic Scale. Mol. Biol. Evol. 38, 5825–5829 (2021).
Lawson, D. J., Hellenthal, G., Myers, S. & Falush, D. Inference of population structure using dense haplotype data. PLos Genet 8, e1002453 (2012).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet 81, 559–575 (2007).
Brettin, T. et al. RASTtk: a modular and extensible implementation of the RAST algorithm for building custom annotation pipelines and annotating batches of genomes. Sci. Rep. 5, 8365 (2015).
Boratyn, G. M. et al. BLAST: a more efficient report with usability improvements. Nucleic Acids Res 41, W29–W33 (2013).
Boutet, E. et al. UniProtKB/Swiss-Prot, the Manually Annotated Section of the UniProt KnowledgeBase: How to Use the Entry View. Methods Mol. Biol. 1374, 23–54 (2016).
O’Leary, N. A. et al. Reference sequence (RefSeq) database at NCBI: current status, taxonomic expansion, and functional annotation. Nucleic Acids Res 44, D733–D745 (2016).
Siguier, P., Perochon, J., Lestrade, L., Mahillon, J. & Chandler, M. ISfinder: the reference centre for bacterial insertion sequences. Nucleic Acids Res 34, D32–D36 (2006).
Moura, A. et al. INTEGRALL: a database and search engine for integrons, integrases and gene cassettes. Bioinformatics 25, 1096–1098 (2009).
Roberts, A. P. et al. Revised nomenclature for transposable genetic elements. Plasmid 60, 167–173 (2008).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32, 1792–1797 (2004).
Alikhan, N. F., Petty, N. K., Ben Zakour, N. L. & Beatson, S. A. BLAST Ring Image Generator (BRIG): simple prokaryote genome comparisons. BMC genomics 12, 402 (2011).
Li, X. et al. oriTfinder: a web-based tool for the identification of origin of transfers in DNA sequences of bacterial mobile genetic elements. Nucleic Acids Res 46, W229–W234 (2018).
Gu, D. et al. A fatal outbreak of ST11 carbapenem-resistant hypervirulent Klebsiella pneumoniae in a Chinese hospital: a molecular epidemiological study. Lancet Infect. Dis. 18, 37–46 (2018).
Heiden, S. E. et al. A Klebsiella pneumoniae ST307 outbreak clone from Germany demonstrates features of extensive drug resistance, hypermucoviscosity, and enhanced iron acquisition. Genome Med. 12, 113 (2020).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (82472335 to Z.R., 32300156 to N.D., 82572580 to N.D., and 82202565 to Y.K.), National Key Research and Development Program of China (2023YFC3603104 to Z.R. and 2024YFC2310905 to N.D.), Zhejiang Provincial Natural Science Foundation of China (LR23H200001 to Z.R.), “Pioneer” and “Leading Goose” R&D Program of Zhejiang Province (2024C03217 and 2026C02A1132 to Z.R.), Key Program of the Zhejiang Medical and Health Science and Technology Project (WKJ-ZJ-2506 to Z.R.), Zhejiang Provincial Medical and Health Science and Technology plan (2022RC185 to G.L.), and São Paulo Research Foundation (23/16216-4 to J.P.F.). We thank Bin Chen for excellent technical assistance with animal experiments.
Author information
Authors and Affiliations
Contributions
Z.R., Y.X. and N.D. conceived the study. S.S.L., M.Z., and J.P.F. performed the bioinformatics analyses. Y.Y., G.L., Q.S., Q.T., and H.J. carried out the experimental analyses. S.S.L., M.Z., S.Q.L., and Y.Y. drew the figures. Z.W., Y.K., and Y.Z. contributed reagents and material. Z.R., M.D. and N.D. wrote the manuscript. All authors read and approved the submission.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Li, S., Yu, Y., Liu, S. et al. Genomic dissection of the clonal background and global dissemination of hypervirulent Klebsiella pneumoniae CG23-KL57 lineage. Nat Commun 17, 1451 (2026). https://doi.org/10.1038/s41467-025-68184-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-68184-4