Abstract
The missense mutation Y65C in polyglutamine-binding protein 1 (PQBP1) is associated with Renpenning syndrome, characterized by X-linked intellectual disability and microcephaly. However, the pathogenic mechanism underlying the microcephaly induced by the Y65C mutation remains unclear. In this study, we generated Pqbp1Y65C/Y knock-in male mice and discovered that the Y65C mutation impairs the proliferation of apical progenitors and their subsequent transition to basal progenitors, resulting in microcephaly and cognitive deficits like those observed in Renpenning syndrome patients. This Y65C substitution induces PQBP1 misfolding, which reduces PQBP1 protein levels and consequently impedes apical progenitor proliferation. Unexpectedly, the Y65C mutation also induces a gain-of-function that interferes with the transition from apical to basal progenitors by enhancing interactions with the core components of the mRNA 3’ end processing machinery, thereby preserving proliferative alternative polyadenylation (APA) profiles. Our study demonstrates that a combination of loss-of-function and gain-of-function contributes to the microcephaly caused by the Y65C mutation.
Similar content being viewed by others
Introduction
Mutations in the polyglutamine-binding protein 1 gene (Pqbp1), located on chromosome Xp11.23, are linked to Renpenning syndrome. This syndrome is characterized by microcephaly, intellectual deficiency, short stature, small testes, and distinct facial dysmorphism1,2,3,4. To date, 22 mutations in the Pqbp1 gene have been documented: 6 small deletions1,4,5,6,7,8,9,10,11,12,13,14, 10 missense mutations6,15,16,17,18,19,20,21,22, 3 small insertions1,5,23, and 3 gross deletions2. Moreover, microduplications at Xp11.23p11.22 have been reported in patients with syndromic MR24,25,26,27. Notably, the Renpenning syndrome associated with the Y65C missense mutation of Pqbp1 exhibits more severe clinical features than other mutations15,28. The reason for the severity caused by a single amino acid mutation remains elusive.
PQBP1 is predominantly expressed in neural stem progenitor cells and striatal progenitors during mouse embryonic development29,30,31. Several conditional knockout (cKO) mouse models of the Pqbp1 gene have been created, and studies using these Pqbp1-cKO mice have highlighted the crucial roles of PQBP1 in neural development and cognition30,31,32,33,34. The missense Y65C mutation, identified in Renpenning syndrome, is distinct among the reported Pqbp1 mutations. This mutation alters the conserved tyrosine residue at position 65 within the functional WW domain to a cysteine (Y65C)15. In vitro biochemical studies have shown that this Y65C mutation interferes with the interactions between PQBP1 and its corresponding partners in the splicing complex35,36. Several studies have shown normal levels of mutant PQBP1 protein and dysregulated pre-mRNA splicing in the patient’s lymphoblasts36,37, suggesting that abnormal splicing might be a primary feature of Renpenning syndrome with the Y65C mutation. However, no animal models are currently available for Renpenning syndrome with the Y65C mutation, and the exact pathological mechanism by which this mutation causes Renpenning syndrome remains unclear.
In this study, we developed a Pqbp1Y65C/Y-knock-in mouse model to explore Renpenning syndrome further. Our findings reveal that Pqbp1Y65C/Y mice display microcephaly and cognitive deficits. We demonstrate that the Y65C mutation significantly reduces the PQBP1 protein level, with the partial loss of PQBP1 function impairing apical progenitor proliferation. Unexpectedly, we discovered that the Y65C mutation also induced a gain-of-function that disrupts the transition from apical to basal progenitors by enhancing interactions with the core components of the mRNA 3’ end processing machinery, thereby preserving proliferative alternative polyadenylation (APA) profiles. Our results elucidate two previously unrecognized post-transcriptional regulatory mechanisms by which the Y65C mutation causes Renpenning syndrome, offering new insights into the pathogenesis of this disease.
Results
Pqbp1 Y65C/Y mice exhibit microcephaly and cognitive deficits
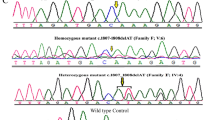
To generate Pqbp1Y65C-knock-in mice, we microinjected Cas9 mRNA, Pqbp1 sgRNA, and a donor oligonucleotide of the Pqbp1-Y65C DNA into zygotes. This procedure resulted in two founder mice with substitutions of tyrosine 65 to cysteine in PQBP1 (Supplementary Fig. 1a, b). The newborn Pqbp1Y65C/Y mice exhibited reduced body weight, brain weight, and decreased cortical area compared to their littermate controls (Fig. 1a–e and Supplementary Fig. 1c–f). Immunostaining analyses revealed a notably thinner neocortex in the newborn PqbpY65C/Y mice compared to their littermate controls (Fig. 1f–h). We further conducted immunostaining using cortical layer-specific markers Tbr1 (Layer VI), Ctip2 (Layer V), and Brn2 (Layer II–IV) to assess the thickness of the cerebral cortex. Although the cortices of newborn Pqbp1Y65C/Y mice showed a normal lamination pattern (Fig. 1f), the thickness of each layer was uniformly reduced in the mutant mice (Fig. 1g–i). Moreover, the proportion of neurons in each layer of the cortex of the mutant mice remained unchanged (Fig. 1j), suggesting no significant abnormalities in the migration and differentiation processes of neurons. The Pqbp1Y65C/Y mice displayed short stature and microcephaly, similar to the major clinical symptoms observed in Renpenning patients.
a General appearance of WT and Pqbp1Y65C/Y mice at P0. Scale bar, 1 cm. b Quantification of body weight for (a). WT: n = 13 littermate wild-type male mice, Y65C: n = 13 Pqbp1Y65C/Y male mice, p < 0.0001. c General morphology of the brains in WT and Pqbp1Y65C/Y mice at P0. The white dotted line represents the cortical area. Scale bar, 2 mm. d Quantification of the brain weights for (c). n = 12 mice in each group, p < 0.0001. e Quantification of the area of the cortex for (c). n = 10 mice in each group, p < 0.0001. f Immunofluorescence staining shows the overall morphology of the cortex at P0. Scale bar, 500 μm. n = 3 biologically independent experiments. g Representative images of WT and Pqbp1Y65C/Y mice cortex stained for Tbr1, Ctip2, Brn2 and DAPI at P0. Scale bar, 100 μm. h, i Quantification of the thickness of cortex (VZ and SVZ are not included) and the thickness (white two-way arrows in g) of Tbr1+, Ctip2+, Brn2+ neurons for (g). n = 9 mice in each group, Thickness: p = 0.0331; Tbr1 + : p = 0.0028; Ctip2 + : p = 0.0076; Brn2 + : p = 0.0137. j Quantification of the proportion of neurons in each layer of the brain for (g). All quantification data are represented as mean ± SD. b, d, e, h, and i, a two-tailed unpaired Student’s t test. *p ≤ 0.05; **p ≤ 0.01; ****p ≤ 0.0001. Source data are provided as a Source Data file.
Next, we performed a behavioral test to evaluate the cognition of Pqbp1Y65C/Y mice at 8–13 weeks. The distances traveled by Pqbp1Y65C/Y mice and the littermate WT controls in the open-field test were comparable (Supplementary Fig. 2a, b), indicating normal locomotion of Pqbp1Y65C/Y mice. However, Pqbp1Y65C/Y mice entered the center less frequently and spent less time in the center than WT controls (Supplementary Fig. 2c). Moreover, the Pqbp1Y65C/Y mice spent significantly less time in the open arms at the elevated plus maze test (Supplementary Fig. 2d–f), suggesting that Pqbp1Y65C/Y mice display increased anxiety. We also performed a novel object-recognition test to evaluate recognition memory (Supplementary Fig. 2g). In the training session, Pqbp1Y65C/Y mice and littermate WT mice explored two identical objects for similar durations (Supplementary Fig. 2h). In the testing session, WT mice spent more time on the novel object, while Pqbp1Y65C/Y mice displayed no preference for either the familiar or novel object (Supplementary Fig. 2i), suggesting impaired recognition memory in Pqbp1Y65C/Y mice. Moreover, we conducted a three-chamber test to evaluate social behavior. We found that the Pqbp1Y65C/Y mice displayed normal social ability, but deficits in social novelty (Supplementary Fig. 2j–n). These social deficits may be associated with underlying cognitive impairments. Our findings suggest that Pqbp1Y65C/Y mice exhibit impaired recognition memory.
Pqbp1 Y65C/Y mice exhibit a reduction in both NSPC proliferation and AP-to-BP transition
The proliferation and differentiation of Neural Stem /Progenitor Cells (NSPCs) drive normal corticogenesis. NSPCs comprise Pax6+ apical progenitors (APs) in the ventricular zone (VZ) and Tbr2+ basal progenitors (BPs) in the subventricular zone (SVZ). To investigate the cellular mechanism underlying microcephaly in Pqbp1Y65C/Y mice, we first examined and quantified APs. Our findings revealed comparable AP numbers during early (E12.5), middle (E15.5), and late (E17.5) neurogenesis phases (Fig. 2a, b). Neural proliferation analysis using 5-bromo-2’-deoxyuridine (BrdU) demonstrated a significant reduction in BrdU+ APs (Pax6+ BrdU+) in the VZ of Pqbp1Y65C/Y cortices compared to the controls (Fig. 2c, d). For a more precise understanding of AP proliferation, we employed a dual labeling strategy with BrdU and EdU to accurately measure the cell cycle parameters of AP progenitors (Fig. 2e–g). We injected BrdU first (2 h before sampling) to label all cells that were in S-phase, and subsequently injected EdU (0.5 h before sampling) to label the cells that entered S-phase later. By analyzing the ratio of labeled cell populations, we can track the progression of a cohort of cells through the cell cycle over time. This analysis revealed an extended cell cycle length for APs in Pqbp1Y65C/Y cortices (Fig. 2e–g). Furthermore, by measuring the proportion of mitotic cells, we found that there is an increase of APs in the M phase in Pqbp1Y65C/Y VZ cortices (Fig. 2h, i), suggesting that the Y65C mutation prolongs the mitotic M phase of APs.
a Representative images of WT and Pqbp1Y65C/Y mice cortex stained for Pax6 at E12.5, E15.5, and E17.5. Scale bar, 50 μm. b Quantification of Pax6+ cell numbers in 200 μm cortex for (a). E12.5: n = 5 mice in WT group, n = 6 mice in Y65C group, p = 0.1443; E15.5: n = 4 mice in WT group, n = 4 mice in Y65C group, p = 0.7153; E17.5: n = 4 mice in WT group, n = 5 mice in Y65C group, p = 0.2775. c APs proliferation in WT and Pqbp1Y65C/Y mice cortex was analyzed by BrdU staining. At 1 h after BrdU injection, the proliferation of cells in the E12.5 embryonic stage was analyzed, and the cells that were undergoing cell proliferation were identified. The left panel shows the overall cortex, scale bar 200 μm, while the right panel is an enlarged image of the area within the white square, scale bar 50 μm. d Quantification of the proportion of the APs undergoing cell proliferation for (c). n = 6 mice in WT group, n = 3 mice in Y65C group, p = 0.024. e Representative images of the cortex in WT and Pqbp1Y65C/Y mice at E12.5, stained to calculate the cell cycle. Scale bar, 50 μm. f, g Time course of dual labeling with EdU and BrdU and quantification of APs cell cycle for (e). Ki67 is detected in all proliferating cells. Ts is the length of the S phase of the cell cycle, Tc is the full length of the cell cycle. Ts= Edu+ /Brdu+Edu- xTi, Tc = Ki67+ /Edu+ xTs, Ti = 1.5 h. n = 5 mice in each group, p = 0.0479. h Representative images of WT and Pqbp1Y65C/Y mice cortex stained for pH3, Ki67, and Tbr2 at E12.5. Tbr2 is used to distinguish between VZ and SVZ. Scale bar, 50 μm. i Quantification of the percentage of APs in the M phase for (h). n = 5 mice in each group, p = 0.0001. All quantification data are represented as mean ± SD. b, d, g, i two-tailed unpaired Student’s t test. ns > 0.05, *p ≤ 0.05; ***p ≤ 0.001. Source data are provided as a Source Data file.
During corticogenesis, APs emerge first and subsequently produce BPs, which differentiate into projection neurons across all cortical layers. In this study, we observed that Pqbp1Y65C/Y mice exhibited a significant reduction in BP numbers (Fig. 3a, b). Further examination following BrdU injection revealed that the proportions of BrdU+ BPs (Tbr2+; BrdU+) in the SVZ were considerably lower in Pqbp1Y65C/Y cortices compared to the controls (Supplementary Fig. 3a, b). Despite the normal number of APs in Pqbp1Y65C/Y mice, APs exhibit slower proliferation, and consequently, BPs are notably reduced. This prompted us to investigate the differentiation potential of APs towards BPs. Our findings indicate that the Y65C mutation decreases the proportion of double-positive Pax6+Tbr2+ cells within the APs (Fig. 3a, c), indicating the compromised ability of APs to transition into BPs. APs possess a unique polarity, characterized by differential protein distribution at their apical and basal poles relative to the epithelial surface. Any deviation in the cell division plane might result in asymmetric cell division and consequently affect cellular lineage. Upon measuring the mitotic angles of APs, we discovered that those from Pqbp1Y65C/Y mice displayed a stronger inclination towards symmetric division (Fig. 3d–f). Using Cadherin to delineate the plasma membrane, we further confirmed the increased tendency for symmetric division in Pqbp1Y65C/Y mice (Fig. 3g, h). Moreover, apoptosis infrequently exists in both groups, with no significant disparities observed (Supplementary Fig. 3c, d). Taken together, these results suggest that the compromised proliferation of NSPCs and the disrupted transition from APs to BPs contribute to a limited BP pool and compromised corticogenesis in Pqbp1Y65C/Y cortices.
a Representative images of WT and Pqbp1Y65C/Y mice cortex stained for Pax6 and Tbr2 at E12.5. Scale bar, 50 μm. b Quantification of Tbr2+ cell numbers in 200 μm cortex for (a). n = 6 mice in WT group, n = 5 mice in Y65C group, p = 0.027. c Quantification of the proportion of the AP-to-BP transition for (a). n = 6 mice in WT group, n = 5 mice in Y65C group, p = 0.0252. d Schematic diagram of mitotic angle measurement and representative pictures for mitotic APs in WT and Pqbp1Y65C/Y mice cortex at E12.5. Scale bar, 2 μm. e, f Distribution and quantification of cleavage plane orientation for mitotic APs in WT and Pqbp1Y65C/Y mice cortex at E12.5. The dots represent individual mitotic cells in a given angle range. N = 3 mice, n = 121 cells in WT group, N = 4 mice, n = 150 cells in Y65C group, p = 0.0014. g Representative images of the symmetric and asymmetric APs in WT and Pqbp1Y65C/Y mice cortex at E12.5. Pan-cadherin (Cadherin) marks the apical plasma membrane, white bars indicate the “cadherin hole”, and the white dashed line represents the split axis. Scale bar, 5 μm. h Quantification of the proportion of the symmetric and asymmetric APs for (g). N = 3 mice, n = 126 cells in WT group, N = 4 mice, n = 148 cells in Y65C group, p = 0.0041. All quantification data are represented as mean ± SD. b, c, h two-tailed unpaired Student’s t-test. f Two-tailed Mann-Whitney rank-sum test. *p ≤ 0.05; **p ≤ 0.01. Source data are provided as a Source Data file.
The Y65C mutation significantly reduces PQBP1 protein levels in Pqbp1 Y65C/Y cortices
To investigate the pathological mechanism underlying the neural progenitor defects caused by the Y65C mutation, we first assessed its impact on PQBP1 protein levels. Immunofluorescence staining and western blot analyses revealed a significant reduction in PQBP1 protein levels in Pqbp1Y65C/Y cortices (Fig. 4a, b). This was evident using both anti-full-length-PQBP1 polyclonal antibodies (anti-PQBP1FL) and polyclonal antibodies specific to the C-terminus of PQBP1 (anti-PQBP1CTD) (Supplementary Fig. 4a–c). A prior study noted a slight reduction in PQBP1 protein levels in lymphoblasts from Renpenning syndrome patients with the Y65C mutation37. Indeed, PQBP1 protein levels were slightly decreased in Pqbp1Y65C/Y mouse lymphocytes (Supplementary Fig. 4d). However, PQBP1 protein levels were markedly reduced in the Pqbp1Y65C/Y heart, lung, and liver (Supplementary Fig. 4e–g), indicating that the decline in PQBP1 levels due to the Y65C mutation is not brain-specific. We further examined PQBP1 protein levels across different tissues to show that PQBP1 protein was highly abundant in the cortex and lung, moderate in the heart and liver, and very low in lymphocytes (Supplementary Fig. 4h). These findings indicate that the decline in PQBP1 levels caused by the Y65C mutation exhibits tissue-specific differences, and the very low basal expression of PQBP1 in lymphocytes likely mask the observation of any further reduction.
a Immunofluorescence staining shows the PQBP1 expression in the cortex at E12.5. Scale bar, 200 μm. Shown are representative results from three independent experiments. b Western blots using anti-PQBP1FL antibodies to detect PQBP1 in P30 WT and Pqbp1Y65C/Y cortices and quantification of the relative PQBP1 level. n = 3 mice in each group, p = 0.0054. c Quantification of Pqbp1 mRNA levels in P30 WT and Pqbp1Y65C/Y cortices. n = 5 mice in each group, p = 0.8461. d Overexpression of Flag-PQBP1 or Flag-PQBP1Y65C protein in SK-N-BE(2) cells. Flag-PQBP1, black line arrows; endogenous PQBP1, black solid arrows, respectively. The right panel is a quantification of the relative Flag-PQBP1 level. n = 4 biologically independent experiments in each group, 12 h: p = 0.3383; 18 h: p = 0.0746; 24 h: p < 0.0001; 36 h: p = 0.0006; 48 h: p = 0.0027; 72 h: p = 0.0096. e Overexpression of Flag-PQBP1 or Flag-PQBP1Y65C protein in 293 T cells for 36 h, 100 μg/ml CHX was added. Flag-PQBP1, black line arrows; endogenous PQBP1, black solid arrows, respectively. The right panel is a quantification of the relative Flag-PQBP1 level. n = 4 biologically independent experiments in each group, 3 h: p = 0.1242; 6 h: p = 0.0703; 9 h: p = 0.0016; 12 h: p = 0.0002. f Overexpression of Flag-PQBP1Y65C in 293 T cells for 36 h, 100 μg/ml CHX, and 10 μM MG132 were added. Flag-PQBP1, black line arrows; endogenous PQBP1, black solid arrows, respectively. The right panel is a quantification of the relative Flag-PQBP1Y65C level. n = 3 biologically independent experiments, 3 h: p = 0.3564; 6 h: p = 0.0765; 9 h: p = 0.0073; 12 h: p = 0.0003. g Overexpression of WT and mutant PQBP1 protein in SK-N-BE(2) cells for 36 h. Flag-PQBP1, black line arrows; endogenous PQBP1, black solid arrows, respectively. The panel below is a quantification of the relative Flag-PQBP1 level. n = 4 biologically independent experiments, WT vs. Y65C (TGC): p < 0.0001; Y65C (TGT) vs. Y65C (TGC): p = 0.1701; Y65F vs. Y65C (TGC): p = 0.0049; Y65S vs. Y65C (TGC): p = 0.0001; Y65W vs. Y65C (TGC): p = 0.0041; Y65C,C60S vs. Y65C (TGC): p = 0.0004. All quantification data are represented as mean ± SD. b–f, g two-tailed unpaired Student’s t test. ns>0.05, *p ≤ 0.05; **p ≤ 0.01; ***p ≤ 0.001; ****p ≤ 0.0001. Source data are provided as a Source Data file.
Next, we examined Pqbp1 mRNA levels in the cortex of Pqbp1Y65C/Y mice. Quantitative RT-PCR revealed that Pqbp1 mRNA levels in control and Pqbp1Y65C/Y mice were comparable (Fig. 4c and Supplementary Fig. 4d–g), indicating that the mutation does not influence Pqbp1 mRNA transcription. To understand how the Y65C mutation reduces PQBP1 protein levels, we transiently expressed flag-tagged full-length PQBP1 (Flag-PQBP1) or its Y65C mutant (Flag-PQBP1Y65C) in SK-N-BE(2) cells and monitored their protein levels. The expression levels of Flag-PQBP1Y65C were notably lower than those of Flag-PQBP1 at every time point (Fig. 4d). Polysome profiling demonstrated that the Y65C mutation did not alter the distribution of Pqbp1 mRNA in ribosomes (Supplementary Fig. 5a, b), implying no significant abnormalities in PQBP1 protein synthesis. Cycloheximide (CHX), a protein synthesis inhibitor, binds to the E-site on the ribosome, inhibiting nascent protein synthesis. Upon monitoring expressed PQBP1 protein levels at various time points post-CHX addition, we observed that the levels of Flag-PQBP1Y65C protein declined swiftly after CHX treatment (Fig. 4e). Moreover, the addition of the proteasome inhibitor, MG132, significantly decelerated the degradation rate of the PQBP1Y65C mutant protein (Fig. 4f). These findings indicate that the Y65C mutated protein undergoes rapid degradation via the ubiquitin-proteasome pathway. We further modulated the expression levels of PQBP1-Y65C in 293 T cells and monitored the corresponding degradation rate of the mutant protein. We did not detect significant degradation of the Y65C protein at very low expression levels, which is consistent with our in vivo observation in lymphocytes, suggesting that low abundance of the Y65C protein may not adequately activate the protein degradation machinery (Supplementary Fig. 5c, d).
Considering that PQBP1 contains a single-cysteine residue at position 60, it is logical to deduce that the Y65C mutation forms anomalous disulfide bonds, subsequently interfering with protein folding and leading to premature degradation. To corroborate this hypothesis, we replaced the tyrosine at position 65 with Phe (Y65F), Ser (Y65S), or Trp (Y65W). These replacements notably elevated the expression levels of PQBP1 (Fig. 4g). Converting the cysteine to Ser at position 60 in the mutant protein (PQBP1C60S, Y65C) also substantially restored PQBP1 expression (Fig. 4g). Conversely, substituting codon TGC with codon TGT, which still encodes the cysteine residue at position 65, did not amplify PQBP1 expression (Fig. 4g). Furthermore, the two mutants, Flag-PQBP1C60S, Y65C and Flag-PQBP1Y65S, which impede disulfide bond formation, exhibited stability comparable to that of Flag-PQBP1 (Supplementary Fig. 5e, f). These findings demonstrate that the Y65C mutation substantially destabilizes PQBP1.
Nestin-cKO mice display microcephaly without deficits in the AP-to-BP transition
Given that the level of PQBP1 protein was significantly decreased in Pqbp1Y65C mice, we asked whether the impaired NSPC proliferation and defective AP-to-BP transition were due to the loss of PQBP1 function. To answer this question, we specifically knocked out PQBP1 in NSPCs by crossing floxed Pqbp1 mice with Nestin-Cre mice38. Through real-time quantitative PCR and immunofluorescence analysis, we confirmed that PQBP1 is significantly decreased at E12.5 and nearly completely knocked out at E14.5 in cKO mice (Supplementary Fig. 6a–c). As predicted, newborn cKO mice (Nestin-Cre; Pqbp1fl/Y) displayed an approximately 6.5% reduction in brain weight compared to their littermate controls (Pqbp1fl/Y) (Fig. 5a). In contrast, Pqbp1Y65C/Y mice exhibited a more severe reduction of approximately 15%. (Fig. 1c, d). Histological analysis demonstrated a significant reduction in the thickness of each layer (Cux1+ for the layer II–IV, Ctip2+ for the layer V, and Tbr1+ for the layer VI) in newborn cKO mice (Fig. 5b–d). Further immunohistochemical analysis revealed a substantial decrease in the number of APs and BPs in cKO mice (Fig. 5e–g). A previous study has shown that the depletion of PQBP1 in NSPCs leads to impaired NSPC proliferation and microcephaly32. These findings suggest that the loss of PQBP1 function impacts NSPC proliferation. However, consistent with the previous observations32, cKO mice show normal AP-to-BP transition (Fig. 5e, h, and Supplementary Fig. 6d, e), indicating that the defective AP-to-BP transition in Pqbp1Y65C/Y mice may be attributed to the Y65C mutation-induced gain of function.
a Pqbp1flox/Y mice as control and Pqbp1flox/Y; Nestin-Cre mice as cKO. The top is the general morphology of the brains in control and cKO mice at P0. Scale bar, 3 mm. The bottom is the quantification of brain weights for the top. n = 7 mice in each group, p = 0.0106. b Representative images of control and cKO mice cortex stained for Tbr1, Ctip2, Cux1, and DAPI at P0. Scale bar, 100 μm. c Quantification of the thickness (white two-way arrows in b) of Tbr1+, Ctip2+, Cux1+ neurons for (b). n = 4 mice in each group, Tbr1+ : p = 0.0005; Ctip2+ : p = 0.0031; Brn2+ : p = 0.0014. d Quantification of the proportion of neurons in each layer of the brain for (b). e Representative images of control and cKO mice cortex stained for Pax6 and Tbr2 at E15.5. Scale bar, 50 μm. f, g Quantification of the Pax6+ cell number, Tbr2+ cell number in 200 μm cortex for (e). n = 4 mice in control group, n = 3 mice in cKO group, Pax6+ number: p = 0.0125; Tbr2+ number: p < 0.0001. h Quantification of the proportion of the AP-to-BP transition for (e). n = 4 mice in control group, n = 3 mice in cKO group, p = 0.361. All quantification data are represented as mean ± SD. a, c, f, g, h two-tailed unpaired Student’s t test. ns>0.05, *p ≤ 0.05; **p ≤ 0.01; ***p ≤ 0.001; ****p ≤ 0.0001. Source data are provided as a Source Data file.
The Y65C mutation induces a gain-of-function to affect the AP-to-BP transition
We next investigated whether the Y65C mutation confers a gain-of-function effect to perturb the AP-to-BP transition in Pqbp1Y65C/WT heterozygous female mice. Although the heterozygous mice expressed normal levels of PQBP1 protein (Supplementary Fig. 7a, b), they showed defective AP-to-BP transition, but not as severe as that in Pqbp1Y65C/Y mice (Fig. 6a, b). We also performed an in utero electroporation experiment by introducing the expression plasmids into the cortex of WT mice at E14.5, which corresponds to the peak period of the AP-to-BP transition (Fig. 6c). Immunofluorescence staining showed that expressing wild-type PQBP1 did not affect the AP-to-BP transition, while expression of PQBP1Y65C mutant proteins caused a significant deficit in the AP-to-BP transition in WT mice (Fig. 6d, e). These results indicate that the Y65C mutation confers a gain-of-function to affect the AP-to-BP transition.
a Representative images of WT, Pqbp1Y65C/WT and Pqbp1Y65C/Y mice cortex stained for Pax6 and Tbr2 at E12.5. Scale bar, 50 μm. b Quantification of the proportion of the AP-to-BP transition for (a). n = 6 mice in WT group, n = 5 mice in Y65C/WT group, n = 6 mice in Y65C group, WT vs Y65C/WT: p = 0.0218; WT vs Y65C/WT: p < 0.0001. c Schematic diagram of in utero electroporation and plasmid construction. The plasmid was mixed with fast green and injected into the ventricles of embryonic mice. + and − represents positive and negative electrodes respectively. d Representative images of the mouse cortex stained for Pax6, Tbr2, and EGFP after electroporation with different plasmids at E15.5. Scale bar, 50 μm. e Quantification of the proportion of the AP-to-BP transition after electroporation with different plasmids for (d). n = 4 mice in EGFP group, n = 6 mice in mPQBP1 group, n = 6 mice in mPQBP1Y65C group, EGFP vs mPQBP1: p = 0.2335; EGFP vs mPQBP1Y65C: p < 0.0001. All quantification data are represented as mean ± SD. b, e A two-tailed unpaired Student’s t-test. ns>0.05, *p ≤ 0.05; ****p ≤ 0.0001. Source data are provided as a Source Data file.
The Y65C mutation preserves the proliferative APA profiles to influence the AP to-BP transition
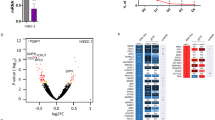
To elucidate the mechanism behind the gain-of-function exhibited by the Y65C mutant protein, we isolated proteins associated with PQBP1 from the embryonic cortices of both wild-type and Pqbp1Y65C/Y mice using co-immunoprecipitation experiments. Subsequently, these proteins were subjected to liquid chromatography-tandem mass spectrometry (LC-MS/MS). Across two independent replicates, 37 proteins displayed increased interactions (Normalized iBAQY65C/iBAQwild-type > 1.5) and 77 proteins displayed decreased interactions (Normalized iBAQY65C/iBAQwild-type < 0.67) with the Y65C protein (Fig. 7a and Supplementary Data 1). Notably, the strengthened interactome was significantly enriched in mRNA processing factors, including known splicing factors such as PRPF19, HNRNPA3, and core components of the mRNA 3’ end processing machinery, such as FIP1L1 (Fig. 7b, c), and the weakened binding proteins related to chromatin remodeling and RNA splicing (Fig. 7b). These results suggest that the gain-of function effects observed with the Y65C mutant protein may be linked to mRNA processing.
a Venn diagram delineating the overlapping distribution of differential binding proteins across parallel IP-MS datasets. n = 3 mice in each group in each independent experiment. b Top 10 enriched biological processes ranked by P-value. Analysis of the overlapping proteins from (a) using a one-sided hypergeometric test with Benjamini–Hochberg FDR correction. Numbers and the size of circles represent counts. c Heatmap visualization reveals differential expression patterns of enhanced binding proteins from (a). IBAQ intensities from duplicate IP-MS experiments were quantile-normalized and averaged. d Scatter plot depicting the genes with APA changes (FDR < 0.05) in E12.5 Pqbp1Y65C/Y cortex. Blue: Transcripts with proximal PAS preference in Y65C. Red: Transcripts demonstrating distal PAS preference in Y65C. n = 3 mice in each group. e The genes with APA changes (FDR < 0.05) in E12.5 Pqbp1-cKO cortex. Blue: transcripts with proximal PAS preference in cKO compared to the control; Red: Transcripts demonstrating distal PAS preference in cKO. n = 3 mice in each group. f Venn diagram shows the overlapped targets between Pqbp1Y65C/Y and Pqbp1-cKO. g The genes overlaping with known ESC self-renewal regulators. h Integrative Genomics Viewer tracks show the representative gene reads of Marcksl1 from RNA-seq. i qRT-PCR measures the ratios of transcripts with the extended 3′ UTR of Marcksl1. The locations of primer sets used to amplify specific regions are marked (top). n = 3 mice in each group, p = 0.0329. j CoIP of PQBP1 and FIP1L1. The cortex of mice at E15.5 was immunoprecipitated with anti-PQBP1 antibodies. n = 3 biologically independent experiments, p = 0.0085. k, l Dual-luciferase reporter assays for Flag, Flag-PQBP1, and Flag-PQBP1Y65C groups. The ratios of R/F (Renilla luciferase 480 nm/firefly luciferase 560 nm) were normalized to the Flag group. CMV, CMV promoter; R luc, Renilla luciferase gene; IRES, internal ribosomal entry site; F luc, firefly luciferase gene. n = 5 biologically independent experiments, Flag vs Flag PQBP1: p < 0.0001; Flag vs Flag PQBP1Y65C: p = 0.0002. All quantification data are represented as mean ± SD. j Two-tailed unpaired Student’s t test. l Two-tailed one-way ANOVA with Tukey’s multiple comparisons test to adjust for multiple comparisons. *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001, ****p ≤ 0.0001. Source data are provided as a Source Data file.
To assess the influence of the Y65C mutation on mRNA processing and pinpoint the targets affecting the AP-to-BP transition, we isolated the cortices at E12.5 and performed high-depth RNA-seq. We initially employed rMATS to analyze alternative splicing and identify target genes impacted by the Y65C mutation. We observed a total of 125 splicing events with significant differences (88 in skipped exons [SE], 14 in retained introns [RI], 9 in alternative 5’ splice sites [A5SS], 6 in alternative 3’ splice sites [A3SS], and 8 in mutually exclusive exons [MXE]) (Supplementary Fig. 8a and Supplementary Data 2). Although these events were statistically significant, we did not identify the critical target genes directly linked to NSPC differentiation among them. Subsequently, we utilized the APAtrap39, a bioinformatic tool that detects the differential polyadenylation site (PAS) usage across experimental conditions, to evaluate the impact of the Y65C mutation on the APA profile of NSPCs. Our analysis revealed 578 genes with significant APA changes (false discovery rate [FDR] <0.05) in Pqbp1Y65C/Y NSPCs compared to the littermate controls. Among these genes, 441 genes (76.3%) exhibited shifts from distal to proximal PAS in Pqbp1Y65C/Y NSPCs (Fig. 7d; Supplementary Data 3), indicating that the Y65C mutation favors proximal PAS selection in NSPCs, generating the shorter isoforms. It has been extensively documented that the proximal PAS-biased APA pattern is correlated with cell proliferation30,40. To further determine whether these PAS shifts are the cause or the consequence of the prolonged M phase observed in Y65C APs, we analyzed APA profiles of E12.5 cortices from Pqbp1-cKO mice that also exhibit prolonged M phase. In contrast to the constitutive Pqbp1Y65C/Y knock-in model, the Pqbp1-cKO model at E12.5 still retains about 25% residual wild-type protein expression (Supplementary Fig. 6b). Only 179 genes showed significant APA changes in Pqbp1-cKO mice, with 56.4% (101) shifting toward proximal and 43.6 % (78) toward distal PAS—an almost balanced pattern (Fig. 7e; Supplementary Data 4). Moreover, merely 39 of the proximal-shifted genes overlapped with the Y65C set (Fig. 7f). We further performed a direct comparative analysis of the APA profiles between the Pqbp1Y65C/Y and Pqbp1-cKO models. This analysis identified 1060 genes with significant APA differences. Among these, 649 genes (61.2%) exhibited a significant preference for the proximal PASs in the Pqbp1Y65C/Y model compared to the cKO, while 411 genes (38.8%) favored distal sites (Supplementary Fig. 8b; Supplementary Data 5). These results underscore substantial differences between the two models and demonstrate that the pronounced proximal-APA bias is a specific molecular signature of the Y65C gain-of-function and is not a consequence of M-phase prolongation. Furthermore, the genes showing distal-to-proximal shifts were significantly enriched in functional categories related to cerebral cortex development and neuronal differentiation (Supplementary Fig. 8c; Supplementary Data 3). Notably, 18 targets were previously identified in genome-wide RNAi screens as potential ESC self-renewal factors41,42 (Fig. 7g, Supplementary Data 6). We further confirmed the APA switches through quantitative real-time PCR (Fig. 7h, i). These findings indicate that the Y65C mutation fosters the selection of proximal PAS, thereby preserving the proliferative APA profiles and influencing the AP-to-BP transition.
To further investigate the molecular mechanism underlying the Y65C mutation-promoted selection of the proximal PAS, we conducted a co-immunoprecipitation experiment. This confirmed the strengthened association between the Y65C mutant protein and FIP1L1, which is essential for activating mRNA 3’ end processing at the cleavage step and facilitating the utilization of the proximal PAS to maintain ESC self renewal43 (Fig. 7j). To assess whether this strengthened association influences the PAS cleavage efficiency, we used the dual luciferase reporter system pPASPORT. In this system, the canonical PAS is inserted between Renilla (Rluc) and Firefly (Fluc) luciferases. An increase in the Rluc/Fluc ratio indicates the cleavage at the inserted PAS. Convincingly, the expression of the Y65C mutant protein significantly increased the Rluc/Fluc ratio, facilitating the PAS cleavage (Fig. 7k, l). Taken together, these findings demonstrate that the Y65C mutation promotes the selection of the proximal PAS to preserve the proliferative APA profiles, thereby influencing the AP-to-BP transition.
Discussion
In summary, we have successfully developed a knock-in mouse model of Renpenning syndrome with the Y65C mutation, thereby uncovering an unanticipated pathologic mechanism. Our findings demonstrate that Pqbp1Y65C/Y mice display microcephaly and cognitive impairments, mirroring the symptoms observed in patients with Renpenning syndrome. Furthermore, we have revealed that the Y65C mutation not only significantly reduces the PQBP1 protein level, thereby impairing AP proliferation, but also enhances the association with the core mRNA 3’ processing components and preserves the proliferative APA profiles that impair the transition from AP to BP.
The Y65C mutation results in a decline in PQBP1 protein level, and loss of PQBP1 function impairs NSPC proliferation
More than a dozen mutations in the Pqbp1 gene have been identified in Renpenning syndrome1,2,3,4,5,11,15. These include frameshifts that produce truncated proteins and introduce a new C-terminal epitope44, as well as missense mutations that change critical residues in the functional domain of the protein, leading to either loss-of-function or gain-of-function35,44,45,46,47. Although Okazawa and our group independently generated Pqbp1-conditional knockout mice using different strategies32, these models do not accurately reflect disease-related mutations, especially those that cause gain of function. This distinction is directly reflected in the behavioral differences observed between Pqbp1Y65C/Y and Pqbp1-cKO mice. While the open field test showed similarly abnormal results in both models, the elevated plus maze test showed Pqbp1Y65C/Y mice spent significantly less time in the open arms, whereas no significant difference was observed in Nestin-Cre;Pqbp1-cKO mice32. Moreover, through the novel object recognition test and three-chamber test, we further demonstrated that Pqbp1Y65C/Y mice exhibit significant cognitive impairments, which have not been reported in cKO models, mimicking the cognitive phenotypes observed in clinical patients. Therefore, it is necessary to generate animal models based on specific disease-related mutations and study their pathological mechanisms individually.
Our primary observation is that the Y65C mutation significantly reduces PQBP1 protein levels in Pqbp1Y65C/Y mouse cortices. The Y65C missense mutation, situated within the WW domain of PQBP1, is unique among the identified PQBP1 mutations15; it affects one of the four conserved amino acids in the WW domain that facilitates specific protein-protein interactions with short proline-rich or proline-containing motifs35,48,49. Previous studies have shown that the Y65C mutation disrupts interactions among splicing complex partners and deregulates pre-mRNA splicing in Renpenning syndrome lymphoblasts35,36. This study showed that the Y65C mutation significantly reduced PQBP1 levels in the Pqbp1Y65C/Y mouse brain. A previous study has shown that depletion of the Pqbp1 gene from mouse NSPCs leads to impaired NSPC proliferation32. Our recent studies also demonstrated that depletion of PQBP1 results in impaired NSPC and striatal progenitor proliferation30,31. Consistent with these findings30,31,32, our results indicate that depletion of PQBP1 in NSPCs results in impaired NSPC proliferation and microcephaly. This data demonstrates that the Y65C mutation causes a significant reduction in PQBP1 levels, and loss of PQBP1 function leads to defective NSPC proliferation.
Unexpectedly, we observed that the tissue-specific differences in the reduction of PQBP1-Y65C protein are associated with its abundance. In this study, we showed a significant reduction in PQBP1 protein levels in the brain, liver, heart, and lung of Pqbp1Y65C/Y mice, all exhibiting high PQBP1 abundance. Conversely, a comparable level of PQBP1 protein was detected in Pqbp1Y65C/Y mouse lymphoblasts, which exhibit very low PQBP1 abundance. This observation is consistent with previous studies with lymphoblasts from patients with Renpenning syndrome carrying the Y65C mutation36,37. These results suggest that lymphoblasts from Renpenning syndrome with the Y65C mutation may not accurately represent the pathological conditions found in the brains of these patients, highlighting the importance of generating a Renpenning syndrome with the Y65C mutation mouse model.
The Y65C mutation inhibits AP-to-BP transition
Our recent studies have demonstrated that PQBP1 plays a critical role in maintaining the balance between proliferation and differentiation in NSPCs and striatal progenitor proliferation30,31. When the PQBP1-mediated alternative polyadenylation process is disrupted, it leads to impaired NSPC proliferation and increased differentiation30. Notably, striatal progenitors deficient in PQBP1 also exhibit reduced proliferation and increased differentiation31. In this study, we found that Pqbp1Y65C/Y mice displayed impaired NSPC proliferation and decreased differentiation, which contrasts with observations in Pqbp1-cKO mice, where there was impaired NSPC proliferation but normal differentiation32. Furthermore, we demonstrated that the expression of PQBP1-Y65C is sufficient to decrease the AP-to-BP transition, suggesting the decline in AP-to-BP transition is due to the gain-of-function induced by the Y65C mutation. The cellular pathological mechanisms causing microcephaly due to the Y65C mutation are clearly distinct from those in Pqbp1-cKO mice. These findings further underscore the necessity of creating animal models based on specific disease-related mutations and studying their pathological mechanisms.
The Y65C mutation preserves the proliferative APA profile to inhibit AP-to-BP transition
APA is a critical post-transcriptional regulatory mechanism governing mRNA isoform diversity during neurodevelopment. Generally, proliferating cells typically favor proximal PAS to generate shorter mRNAs, while differentiating cells shift toward distal PAS usage40,50. In this study, we observed an increased utilization of proximal PAS in Pqbp1Y65C/Y NSPCs, accompanied by a decline in the AP-to-BP transition. Further analysis revealed that Y65C-mediated APA targets are involved in cerebral cortex development and neuronal differentiation, with a subset of these genes previously identified as potential ESC self-renewal factors. These findings suggest that the PQBP1-Y65C mutation preserves a proliferative APA profile and consequently disrupts the AP-to-BP transition.
Our previous study has provided valuable insights into the role of PQBP1 in APA regulation, revealing that PQBP1 directly interacts with the upstream UGUA elements through its C-terminal domain, impeding the recruitment of the CFIm complex and suppressing PAS cleavage. Depletion of PQBP1 in cultured neurospheres leads to pronounced proximal-to-distal APA shifts30. In the current study, however, no strong directional APA shift was detected in E12.5 cKO cortices. We attribute this stage-specific discrepancy to differences in knockout efficiency and neural progenitor cell states. Notably, Nestin-Cre-mediated deletion remains incomplete at E12.5, with approximately 25% residual PQBP1 protein, whereas it is nearly complete by E14.5. Moreover, NSPCs are primarily engaged in self-renewal at E12.5, whereas by E14.5, many have committed to neuronal differentiation. This transition is regulated by dynamic changes in key signaling pathways such as Wnt, which promotes self-renewal at E12.5 but drives differentiation at E14.5. Consistent with these developmental dynamics, PQBP1 function is also stage-dependent. The minimal APA change in cKO cortices at E12.5 suggests that PQBP1 plays a limited role in APA regulation at this stage. By E14.5, however, PQBP1 is essential for gatekeeping NSPC differentiation—its loss leads to a proximal-to-distal APA shift in synapse-related genes, thereby promoting neuronal commitment30. Strikingly, the Y65C mutation produces a fundamentally distinct distal-to-proximal APA profile, enriched in genes involved in nucleosome assembly and Wnt signaling, potently impairing AP-to-BP transition. Direct comparative analysis of APA profiles between Pqbp1Y65C/Y mice and Pqbp1-cKO mice at E12.5 also revealed substantial differences between the two models, despite potential batch effects. These results collectively support a gain-of-function mechanism underlying the Y65C phenotype.
Mechanistically, our co-immunoprecipitation experiments showed the Y65C mutant protein exhibits enhanced associations with FIP1L1, a core component of the mRNA 3’ end processing machinery essential for activating PAS cleavage and promoting proximal PAS utilization to maintain ESC self renewal43. Luciferase reporter assays consistently revealed that the Y65C protein significantly enhances PAS cleavage efficiency, which is in contrast to the inhibitory effect observed with wild-type PQBP1 or a WWD-deleted PQBP1 truncated protein30. These data collectively support that the Y65C mutation exerts a gain-of-function effect, altering APA profiles in NSPC through a distinct mechanism from the wild-type PQBP1. Further systematic studies are required to fully elucidate the molecular mechanisms underlying PQBP1-Y65C’s influence on APA regulation.
Methods
Animals
Mice in the C57BL/6 background were used in this study and bred in the SPF core animal facility of Southeast University at 25 °C with a relative humidity of 40%–60% and a 12 h light/12 h dark cycle. Mice had free access to food and water. Mouse breeding and all animal experiments were approved by the Southeast University Institutional Animal Care and Use Committee (#20211101003, Nanjing, China) and were performed by institutional and national guidelines. Given that PQBP1 is located on the X chromosome and that all reported human cases are male, all experimental groups and control groups in this study consist of male mice, except for the heterozygous female Pqbp1Y65C/WT mice. WT represents Pqbp1WT/Y, Y65C represents Pqbp1Y65C/Y, and Y65C/WT represents Pqbp1Y65C/WT.
Generation of Pqbp1 Y65C mice
Pqbp1Y65C knock-in mice were generated using homologous recombination in zygotes and implanted into pseudopregnant mice51,52. The sgRNA sequences (5S8: 5’-CATGAGTTTTGACCTATA- and 3S1: 5’-CGGACTCCCTTACTAT) were cloned into the CRISPR/Cas9 system and verified by DNA sequencing. The donor vector included the DNA fragments containing exons 3-4; the mutation of Tyr 65 to Cys (codon TAT to TGT) was also performed. After plasmid verification, the donor vector and CRISPR/Cas9 system were injected into zygotes. Then, the zygotes were injected into 0.5 d.p.c. pseudopregnant females for generating chimeric mice (F0). F0 mice were crossed with wild-type (WT) C57BL/6 mice to establish germline-transmitted founders. Genotyping of Pqbp1Y65C mice was performed by amplifying and sequencing a 0.41-kb product from purified mouse tail DNA (forward: 5’-CCATTCTTCCAGAGCCAGAG and reverse: 5’-TTCTTGGCGGATTTGGTAAC).
Generation of Nestin-cre; Pqbp1flox/Y mice
To generate neural progenitor cell-specific Pqbp1 conditional knock-out (cKO) mice, Pqbp1flox/flox mice, which were generated by the Model Animal Research Center of Nanjing University53, were crossed with Nestin-Cre mice38 (Stock #003771, Jackson Laboratory, Bar Harbor, ME, USA) to produce Pqbp1flox/Y (control) and Nestin-cre; Pqbp1flox/Y (Pqbp1-cKO) mice.
Animal mating strategies
1) Mating Strategy for Pqbp1Y65C knock-in mice
Upon obtaining F0-generation mice, we crossed them with wild-type (WT) C57BL/6 mice to generate F1-generation Pqbp1Y65C/WT(Y65C/WT) heterozygous female mice. These heterozygous females were subsequently backcrossed with WT mice to produce experimental Pqbp1Y65C/Y (Y65C) male mice and littermate control WT male mice for phenotypic analysis.
2) Mating Strategy for Pqbp1 Nestin-cKO mice.
We crossed Nestin-Cre heterozygous males with Pqbp1flox/flox females to generate Nestin-Cre; Pqbp1flox/Y males (Pqbp1-cKO) mice and control Pqbp1flox/Y males (control) mice for experimental analysis.
Nissl staining
Mice brains were perfused with cold phosphate-buffered saline (PBS, pH 7.4), fixed in 4% paraformaldehyde (PFA) overnight at 4°C, then processed for paraffin embedding and serially sectioned (10 μm) by a cryostat (Leica, CM 3050S, Germany). Sagittal sections were deparaffinized, ethanol hydrated, and treated with 0.5% cresyl-violet for 10 min, then rinsed with distilled water three times. Slides were ethanol dehydrated and treated with toluene, and finally coverslipped using neutral resin. Images were collected by LAS X 3.3 (Leica, Germany).
Plasmid construction
The full-length mPQBP1 and mPQBP1 Y65C cDNAs, obtained from WT and Y65C mutant mice respectively, were cloned between the Xho I and EcoR I sites of the pCAG-EGFP vector to generate the expression plasmids. The Flag-PQBP1-FL plasmid was were obtained from Zhang et al.44. The pPASPORT plasmids for Luciferase reporter asssys were obtained from Liu et al.30. Other constructs were generated by gene-directed mutation PCR (12351010, Thermo Scientific). A list of primers that were used in all experiments is shown in the Supplementary Data 7.
Cell culture and transfections
293 T (#GNHu17) and SK-N-BE(2) cell (# TCHu200) lines were purchased from Cell bank of Chinese Academy of Sciences and cultured under standard conditions (37°C, 5% CO2) in their respective media: 293 T cells in DMEM and SK-N-BE(2) cells in DMEM/F12, both supplemented with 10% fetal bovine serum (FBS). For transfection experiments, cells were seeded in 6-well plates at a density of 1×10⁵ cells per well. Upon reaching 70-80% confluence, plasmid DNA transfection was performed using cell line-specific reagents: Lipofectamine 3000 (L3000075, Thermo Fisher Scientific) for 293 T cells and X-treme GENE HP DNA Transfection Reagent (06366236001, Roche Diagnostics, Basel, Switzerland) for SK-N-BE(2) cells.
Antibody generation
FL-PQBP1 antibody was raised in rabbits against the full length of the GST-PQBP1 protein. For the production of polyclonal anti-PQBP1-CTD antibody, a specific peptide N-KKATSRKDEELDPMDPSSYSDAPRGTWSTGLPKRNEAKTGADTTAAGPLFQQRPYPSPGAVLRANAEASRTKQQD-C was synthesized. The peptide was injected into rabbits (YUROGEN, Wuhan, China). After three boosts at 1- to 2-week intervals, high-titer sera were collected and then purified using Sulfo Link Coupling Gel (Pierce, Rockford, IL). For Western blots, FL-PQBP1 and CTD-PQBP1 antibodies were used at a 1:500 and 1:100 dilution, respectively.
Western blots
Mice cortices and cell lines were lysed in cold radioimmunoprecipitation assay (RIPA) lysis buffer [consisting of the following: 20 mM Tris-HCl, pH 7.4, 150 mM NaCl, 1 mM EDTA, 1% NP-40, 0.5% sodium deoxycholate, and 0.1% SDS] containing protease and phosphatase inhibitors (04693132001, Thermo Fisher, USA; 04906837001, Thermo Fisher, USA). Lysates were cleared by centrifugation (12,000 × g for 20 min). Normalized volumes of samples (10 μg total protein) were resolved in 10% SDS-PAGE and transferred to PVDF membranes (Millipore, Massachusetts, US). After blocking with 0.5% (w/v) nonfat dry milk in Tris-buffered saline (TBS: 25 mM Tris-HCl, pH 7.4, 150 mM NaCl), membranes were probed with primary antibodies overnight at 4°C. Primary antibodies used were PQBP1 (16264-1-AP, Proteintech, 1:1000 for use); RPS3 (ab128995, abcam, 1:1000 for use); RPL13 (ab134961, abcam, 1:1000 for use); FIP1L1 (83863-2-RR, Proteintech, 1:1000 for use); anti-α-tubulin (T9026, Sigma-Aldrich, 1:5000 for use); anti-actin (T0022, Affinity, 1:5000 for use). The following secondary antibodies were used for western blot, all diluted at 1:1000: Goat anti-rabbit IgG (H + L)-HRP conjugate (401315, Millipore); Goat anti-mouse IgG (H + L)-HRP conjugate (401215, Millipore); Goat anti-Guinea Pig IgG (H + L)-HRP conjugate (AS025, Abclonal). Western blot images were acquired either using an OPTIMAX 2010 automatic film processor for X-ray film development or a Tanon 5200 imaging system for chemiluminescence image capture. To avoid signal saturation, a standardized protocol was strictly followed: an initial short pre-exposure (e.g., 1–2 sec) was performed to estimate signal intensity, followed by a series of graded exposures (e.g., 5 s, 10 s, 30 s, 60 s). Images without overexposed signals were selected for subsequent quantitative analysis. The captured 8-bit TIFF images were analyzed using ImageJ 1.53k software (Fiji). The integrated densities of rectangular regions of interest (ROIs) for target bands and internal control bands were measured, followed by background subtraction. Finally, the normalized ratios of the target proteins to the internal control proteins -Actin were calculated.
Lymphocyte isolation
Mice were euthanized, and the blood was collected through the orbital vein and collected in an anticoagulant tube. Then, it was diluted with PBS buffer and transferred to lymphocyte separation buffer (7211011, DAKEWE, Shenzhen, China). The lymphocytes were isolated after centrifugation and washed twice with PBS buffer. Finally, lymphocytes are used to extract proteins and RNA.
Polyribosome profiling by sucrose gradient ultracentrifugation
For the polyribosome profiling assay, 1-month-old WT and Pqbp1Y65C/Y mouse brains were quickly removed, and whole brain slices (360 μm) were prepared (Leica, VT1000S) in ice-cold artificial cerebrospinal fluid (ACSF) containing (in mM): 120 NaCl, 3.0 KCl, 1.2 MgSO4, 1.0 NaH2PO4, 26 NaHCO3, 2.0 CaCl2, 11 D-glucose saturated with 95% O2/5% CO2. The slices were recovered at 28 °C in ACSF saturated with 95% O2/5% CO2 for at least 2 h before a single slice was transferred to a submersion chamber perfused with 95% O2/5% CO2 saturated ACSF. The brain slices were incubated in ACSF containing 100 mg/ml cycloheximide for 1 h, and then were homogenized in ice-cold PMS buffer containing 20 mM Tris pH 7.5,100 mM KCl, 5 mM MgCl2, 0.3% (v/v) NP-40, 100 U/ml of SUPERase-InTM RNase inhibitor (Invitrogen), 1% w/v protease inhibitor, 1 mM PMSF, and 0.5 mM DTT. The lysates were subjected to centrifugation at 10,000 × g for 10 min at 4 °C, and the resulting supernatants were retained as PMS. PMS was then layered onto 11 mL of 10%-50% (w/v) sucrose gradients and centrifuged at 22,000 × g for 2.5 h at 4 °C using a P40STrotor (HITACHI). The fractions were monitored using an OD254 nm ultraviolet spectrometer (Thermo Scientific, USA). Samples from the fractions were collected for western blot analysis and real-time RT-PCR analysis, respectively.
Quantitative polymerase chain reaction (qPCR)
RNA was isolated using Trizol reagent (9109, TaKaRa, Japan), and 1 µg of the total RNA was reverse transcribed using the Revert Aid First Strand cDNA Synthesis Kit (RR037Q, Takara, Japan). Real-time PCR was performed with 1 μL of the cDNA template added to 10 μL of 2 × TB Green Premix Ex Taq (RR820Q, Takara, Japan) and specific primers (10 pM each). Real-time PCR (Applied Biosystems, USA) was carried out for 40 cycles of denaturation at 95 °C for 5 s, annealing at 60 °C for 30 s. GAPDH was used as an internal control gene. Amplicons were calculated using the 2−ΔΔ Ct method, and data are represented as the mean of three independent experiments ± SD. The primer sequences used for qRT-PCR are shown in the Supplementary Data 7.
BrdU and Edu injection
Since both EdU and BrdU are thymidine analogues that incorporate into DNA during the S-phase, administering them sequentially allows us to track the progression of a cohort of cells through the cell cycle over time. For proliferation analysis, pregnant mice were intraperitoneally injected with BrdU (100 mg/kg, B5002, Sigma) one hour before embryo collection. For cell cycle calculation, we injected BrdU first (2 h before sampling) to label all cells that were in S-phase at that time. This population is referred to as BrdU⁺ cells. Subsequently, we injected EdU (50 mg/kg, A10044, Invitrogen) 0.5 h before sampling to label a later, distinct population of cells that entered S-phase just before collection (Edu⁺ cells). The embryos were then perfused with PBS, fixed in 4% PFA for approximately 8 h, cryo-protected, and sectioned into 14 μm thick slices for immunohistochemistry54.
Immunofluorescence
Paraffin brain sections were deparaffinized and then boiled in 0.01 M sodium citrate buffer solution, pH 6.0. Frozen sections were air-dried and treated with 0.5% Triton in PBS. After blocking in 3% BSA for 1 h, they were immunostained with antibodies raised against Tbr1 (ab183032, Abcam, 1:200 for use); Ctip2 (ab18465, Abcam, 1: 500 for use); Brn2 (sc-6029, SantaCruz, 1:100 for use); Cux1 (sc-13024, SantaCruz, 1:100 for use); Pax6 (901302, biolegend, 1:500 for use); Tbr2 (ab23345, Abcam, 1:500 for use; 14-4875-82, Invitrogen, 1:200 for use); BrdU (ab6326; Abcam, 1:500 for use), Ki67 (ab16667, Abcam, 1:500 for use); pH3 (9701, CST, 1:500 for use); γ-tubulin (ab11316, Abcam, 1:500 for use); Cadherin (ab51034, Abcam, 1:500 for use); cleaved caspase3 (9661, CST, 1:500 for use) and EGFP (A10262, Invitrogen, 1:500 for use). The following secondary antibodies diluted at 1:200 were used for immunofluorescence: Alexa Fluor 555 goat anti-rabbit IgG (ab150078, Abcam); Alexa Fluor 488 goat anti-rabbit IgG (ab150081, Abcam); Alexa Fluor 488 goat anti-mouse IgG (ab150117, Abcam); Alexa Fluor 555 goat anti-mouse IgG (ab150118, Abcam); Alexa Fluor 647 goat anti-rabbit IgG (ab150079, Abcam); Alexa Fluor 647 goat anti-mouse IgG (ab150115, Abcam); Alexa Fluor 488 goat anti- Chicken IgG (ab150169, Abcam); Alexa Fluor 555 goat anti-Rat IgG (ab150158, Abcam); Alexa Fluor 488 goat anti-Rat IgG (ab150157, Abcam). DAPI (A606584, Sangon Biotech, China, 1:5000 for use) was used to stain nuclei. Images were obtained using a Zeiss LSM 700 and LSM 900 confocal microscope (Oberkochen, Germany).
Morphometry
For quantifying cell area and number, images were analyzed and processed using ZEN 2.3 (Zeiss, Germany), LAS X 3.3 (Leica, Wetzlar, Germany), and ImageJ 1.53k (Fiji) software (National Institutes of Health, Bethesda, MD, USA). Three sections from each cortex, spaced 200 μm apart, were used for cell counting.
Behavioral assessments
Age-matched WT littermates of 8–13-week-old healthy male mice were used for all behavioral tests. All behavioral assays in this study were performed during the light cycle and were blinded to genotype.
Open field test
A mouse was positioned adjacent to the wall within a 50 × 50 cm open field arena, and its movement was captured by a camera for a duration of 5 min. The recorded video footage was subsequently analyzed using EthoVision XT 7.0 software (Noldus). Quantitative measures, including the number of entries into and the total time spent in the central area of the arena (15 × 15 cm), as well as the total moving distance, were recorded. Between each trial, the open field arena was thoroughly cleaned with 70% ethanol and dried using paper towels.
Elevated plus maze test
The test mice were placed on the central platform of an elevated plus maze (45 cm above the floor), facing the open arm of the maze, and allowed to explore for 5 min. Their entries into the open and closed arms and the times spent in each location were recorded and analyzed using EthoVision XT 7.0 software (Noldus).
Novel-object recognition test
A test mouse was initially habituated to an open field arena (50 × 50 cm) for a period of 5 min. Following the habituation phase, the mouse was temporarily removed from the arena, and two identical objects, each measuring 6 × 6 × 4 cm and of the same color, were strategically placed on the same side of the arena, positioned 10 cm from the side walls. The mouse was then reintroduced to the center of the arena and allowed to explore the environment, including the two novel objects, for a duration of 5 min. After a 24-hour interval, one of the original objects was replaced with a novel object of similar size but distinct in shape and color. The same test mouse was once again placed in the arena to investigate the arena and the two objects. The mouse’s movements were recorded using a camera for a total of 5 min and subsequently analyzed with EthoVision XT 7.0 software (Noldus).
Three-chamber test
The three-chambered apparatus is designed with three equally sized rectangular compartments, each measuring 20 cm in width, 40 cm in length, and 22 cm in height. The two adjacent compartments are separated by transparent plexiglass walls, each featuring a rectangular door (5 cm wide × 8 cm high) to permit the subject mice to access each compartment. To isolate the stranger mice from the subject mice, wire pencil cups (7.5 cm wide × 7.5 cm long × 10 cm high) are used, with a heavy cup placed on top to prevent the subject mice from climbing. These wire pencil cups are positioned on the outer sides of the apparatus. Before testing, stranger mice are acclimated to the wire pencil cups for 10 min. Between tests, the apparatus, wire pencil cups, and heavy cups are thoroughly cleaned with 75% ethanol and dried with paper towels. Each testing session is composed of three 10-minute stages. During the first stage, subject mice are allowed to freely explore the compartments for 10 min. In the second stage, a stranger mouse (stranger 1) is introduced into one of the wire pencil cups, and the subject mice are given another 10 min to investigate the compartments. Following the sociability test, the subject mice are separated from stranger 1 for 10 min. In the third stage, a novel stranger mouse (stranger 2) is placed into another wire pencil cup, and the subject mice are permitted to explore the three-chambered box. The duration of time that mice spent in each chamber was measured and analyzed for comparison. All stranger mice are male, age-matched, and unfamiliar to the subject mice, with a C57BL/6 J background. The tests are recorded using an overhead camera and Ethovision XT 7.0 software (Noldus). Additionally, a trained experimenter records the sniffing time with a stopwatch to measure sociability and social recognition memory. A “social memory index” is calculated to quantify social recognition memory, using the formula: Sociability index = The time spent with S1/ Total time x100%. Social novelty index = The time spent with S2/ Total time x100%.
Analysis of cleavage plane angle
The mitotic orientation and cleavage plane of neuroepithelial cells in the dorsal telencephalon were analyzed after double staining for γ-tubulin (ab11316, Abcam, 1:500 for use),with DAPI (A606584, Sangon Biotech, China,1:5000 for use) co-stain. The cleavage plane was defined as the perpendicular axis to the mitotic orientation; its angle relative to the apical surface was calculated for each genotype55.
In utero electroporation (IUE)
Pregnant WT mice at E14.5 were anesthetized with 0.8% pentobarbital sodium (80 mg/kg). After anesthesia, a small incision of about 2 cm was made in the lower abdominal region. Embryos were carefully extracted using forceps, and a glass electrode was utilized to inject 1 μg of plasmid (pCAG-EGFP; pCAG-PQBP1; pCAG-PQBP1Y65C) into the lateral ventricle of the embryos, mixed with 0.05% Fast Green (F7252, Sigma). Five electric pulses (36 V, 50 ms duration at 1 s intervals) were generated by using an electroporator (ECM830, BTX) and delivered to the head of embryos using 5 mm forceps-like electrodes (BTX). Following the procedure, the uterus was carefully returned to the abdominal cavity, and the peritoneum and abdominal skin were meticulously sutured. The mice were then placed on a 47 °C heating pad to facilitate recovery. After 24 h (at E15.5), the male embryos were collected for further analysis56.
CO-IP and mass spectrum
Brain tissues from E15.5 embryonic male mice were harvested for cryopreservation, and then lysed in cold HNTG buffer (50 mM HEPES pH7.5, 150 mM NaCl, 1 mM EDTA, 10% glycerol, 0.1% Triton X-100, 1 mM DTT, 1% w/v protein inhibitor, 1% w/v PhosStop) for 30 min on ice. The lysates were subjected to centrifugation at 12,000 g for 10 min at a temperature of 4 °C. Following this, the protein lysates were divided into two equal portions. For each aliquot, either pre-immune serum or the guinea pig polyclonal anti-mPQBP1 antibodies was added accordingly. The mixtures were gently rotated and incubated at 4 °C for 2 h to allow for optimal binding. Following this incubation, an appropriate volume of Protein A agarose beads (10002D, Invitrogen) was introduced to the lysate and allowed to incubate overnight at 4 °C. The next day, the supernatant was carefully removed, and the beads were washed with HNTG buffer to eliminate any unbound material. Following elution with SDS buffer, the samples were denatured in a boiling water bath for 10 min to prepare for protein interaction analysis via Western blot. Before MS characterization, all immunoprecipitation (IP) products underwent Western blot validation. For MS processing, IP products from two biological replicates were co-separated on 10% SDS-PAGE. Specific gel lanes corresponding to each IP sample were excised and submitted to the Proteomics Core Facility at Nanjing Medical University (Nanjing, China) for nano-liquid chromatography tandem mass spectrometry (nano-LC-MS/MS) analysis. Mass spectrum data was collected by Orbitrap Fusion Lumos mass spectrometer (Thermofisher, USA), and the data-dependent acquisition (DDA) mode was applied.The raw files were searched using the MaxQuant (version: 1.5.2.8) software against the mouse UniProt sequence database with default settings.
Luciferase reporter assay
The pPASPORT plasmids inserted with L3 polyA site were co-transfected with pFLAG-CMV2, pFLAG-hPQBP1-WT, and pFLAG-hPQBP1-Y65C, respectively. After 24 h, the transfected cells were lysed with 100 μL of the Passive Lysis Buffer from the Dual-Luciferase Reporter Assay (E1910, Promega), and 30 μL of the lysates were added to 75 μL of Luciferase Assay Reagent. The firefly signals were then quantified by the SpectraMax iD3 multi-function reader (MD, USA). 75 μL of Stop & Glo Buffer was added to each sample before the Renilla signals were quantified. The Rluc/Fluc ratio was then normalized to the control cells.
RNA sequencing and data processing
Total RNA was extracted from the cortices of wild-type and Pqbp1Y65C mice at E12.5. RNA quality was examined using gel electrophoresis and a Nanodrop spectrophotometer (NanoDrop 2000, Thermo Fisher), and sequencing was carried out using the Illumina Novaseq 6000 instrument.
RNA seq data quality was assessed using FastQC v0.11.2, and clean reads were aligned to the mouse reference genome via Hisat v2.2.1. Differential alternative splicing from replicate RNASeq data was detected by rMATs v4.1.2. APA site usage was analyzed via the APAtrap v1.0 tool39, a bioinformatic tool designed to identify novel 3’ untranslated regions (UTRs) and quantify differential alternative polyadenylation (APA) usage between samples. The annotated 3’-UTRs were refined, and the new 3’-UTRs or 3’-UTR extensions were identified using the identifydistal3’UTR module. All potential APA sites were inferred, and their corresponding usages were estimated using the predictAPA module. Significant differences in APA site usage under different conditions were identified using the R package deAPA. A stringent multiple testing correction was applied, and an mRNA was considered to exhibit significantly different 3’-UTR isoforms if the adjusted P value (Padj.) was less than 0.05. The PD index represents the percentage difference in APA site usage between two compared groups, and the Pearson product-moment correlation coefficient (r) quantifies the significant changes in APA site usage within a gene across two samples. A positive r value (r > 0) indicates that the Y65C group preferentially uses distal poly(A) sites (longer 3′-UTRs) compared to the control, whereas a negative r value (r < 0) indicates preferential usage of proximal poly(A) sites (shorter 3’-UTRs) in the Y65C group relative to the control. Gene ontology enrichment analysis was performed with the web-based DAVID v6.8 software (https://david.ncifcrf.gov/). Functional enrichment was determined using a one-sided hypergeometric test with Benjamini and Hochberg correction.
Statistics
Data were analyzed using GraphPad Prism 8.3.0 and are presented as the means ± SD. Statistical significance was determined using the two-tailed, unpaired Student’s t test, the two-tailed Mann–Whitney rank-sum test and two-tailed one-way ANOVA with Tukey post hoc test. P values for each comparison were set as *P ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.001, and ****P ≤ 0.0001. No significance (ns) was set as P > 0.05.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Source data are provided with this paper. All numerical source data and uncropped images of immunoblots are published alongside the paper as a Source Data file. The RNA-seq data generated in this study have been deposited in the NCBI Sequence Read Archive (SRA) under accession codes PRJNA1327650 and PRJNA1327805. The mass spectrometry data have been deposited in the ProteomeXchange Consortium with the dataset identifier PXD068635 (https://www.iprox.cn//page/project.html?id=IPX0013419000). Source data are provided with this paper.
References
Lenski, C. et al. Novel truncating mutations in the polyglutamine tract binding protein 1 gene (PQBP1) cause Renpenning syndrome and X-linked mental retardation in another family with microcephaly. Am. J. Hum. Genet. 74, 777–780 (2004).
Cossee, M. et al. Exonic microdeletions in the X-linked PQBP1 gene in mentally retarded patients: a pathogenic mutation and in-frame deletions of uncertain effect. Eur. J. Hum. Genet. 14, 418–425 (2006).
Kleefstra, T. et al. Genotype-phenotype studies in three families with mutations in the polyglutamine-binding protein 1 gene (PQBP1). Clin. Genet. 66, 318–326 (2004).
Martinez-Garay, I. et al. A two base pair deletion in the PQBP1 gene is associated with microphthalmia, microcephaly, and mental retardation. Eur. J. Hum. Genet. 15, 29–34 (2007).
Kalscheuer, V. M. et al. Mutations in the polyglutamine binding protein 1 gene cause X-linked mental retardation. Nat. Genet. 35, 313–315 (2003).
Germanaud, D. et al. The Renpenning syndrome spectrum: new clinical insights supported by 13 new PQBP1-mutated males. Clin. Genet. 79, 225–235 (2011).
Sheen, V. L. et al. Mutation in PQBP1 is associated with periventricular heterotopia. Am. J. Med. Genet. Part A 152A, 2888–2890 (2010).
Masih, S., Moirangthem, A. & Phadke, S. R. Renpenning syndrome in an Indian patient. Am. J. Med. Genet. Part A 182, 293–295 (2020).
Jeong, H. I. et al. First Korean case of renpenning syndrome with novel mutation in pqbp1 diagnosed by targeted exome sequencing, and literature review. Ann. Clin. Lab. Sci. 48, 522–527 (2018).
Deqaqi, S. C. et al. A gene for non-specific X-linked mental retardation (MRX55) is located in Xp11. Ann. Genet. 41, 11–16 (1998).
Stevenson, R. E. et al. Renpenning syndrome comes into focus. Am. J. Med. Genet. Part A 134, 415–421 (2005).
Mameesh, M. M., Al-Kindy, A., Al-Yahyai, M. & Ganesh, A. Microphthalmos-anophthalmos-coloboma (MAC) spectrum in two brothers with Renpenning syndrome due to a truncating mutation in the polyglutamine tract binding protein 1 (PQBP1) gene. Ophthalmic Genet. 40, 534–540 (2019).
Cho, R. Y. et al. Renpenning syndrome in a female. Am. J. Med. Genet. Part A 182, 498–503 (2020).
Hu, H. et al. X-exome sequencing of 405 unresolved families identifies seven novel intellectual disability genes. Mol. Psychiatry 21, 133–148 (2016).
Lubs, H. et al. Golabi-Ito-Hall syndrome results from a missense mutation in the WW domain of the PQBP1 gene. J. Med. Genet. 43, e30 (2006).
Kurt Colak, F., Eyerci, N., Aytekin, C. & Eksioglu, A. S. Renpenning syndrome in a Turkish patient: de novo variant c.607C>T in PACS1 and hypogammaglobulinemia phenotype. Mol. Syndromol. 11, 157–161 (2020).
Jensen, L. R. et al. Hybridisation-based resequencing of 17 X-linked intellectual disability genes in 135 patients reveals novel mutations in ATRX, SLC6A8 and PQBP1. Eur. J. Hum. Genet. 19, 717–720 (2011).
Abdel-Salam, G. M. H. et al. Phenotypic and molecular insights into PQBP1-related intellectual disability. Am. J. Med. Genet. Part A 176, 2446–2450 (2018).
Morgan, A. et al. Target sequencing approach intended to discover new mutations in non-syndromic intellectual disability. Mutat. Res. 781, 32–36 (2015).
Lopez-Martín, S. et al. A mild clinical and neuropsychological phenotype of Renpenning syndrome: a new case report with a maternally inherited PQBP1 missense mutation. Appl. Neuropsychol. Child 11, 921–927 (2022).
Redin, C. et al. Efficient strategy for the molecular diagnosis of intellectual disability using targeted high-throughput sequencing. J. Med. Genet. 51, 724–736 (2014).
Pan, J., Chia, H., Kusnadi, J., Li, Z. & Yu, L. Renpenning syndrome related to a missense variant in polyglutamine-binding protein 1 (PQBP1): Two pediatric cases from a Chinese family and literature review. Appl. Neuropsychol. Child. 1–9, https://doi.org/10.1080/21622965.2025.2457990 (2025).
Rejeb, I. et al. A novel frame shift mutation in the PQBP1 gene identified in a Tunisian family with X-linked mental retardation. Eur. J. Med. Genet. 54, 241–246 (2011).
Giorda, R. et al. Complex segmental duplications mediate a recurrent dup(X)(p11.22-p11.23) associated with mental retardation, speech delay, and EEG anomalies in males and females. Am. J. Hum. Genet. 85, 394–400 (2009).
Froyen, G. et al. Detection of genomic copy number changes in patients with idiopathic mental retardation by high-resolution X-array-CGH: important role for increased gene dosage of XLMR genes. Hum. Mutat. 28, 1034–1042 (2007).
Flynn, M., Zou, Y. S. & Milunsky, A. Whole gene duplication of the PQBP1 gene in Syndrome resembling Renpenning. Am. J. Med. Genet. Part A 155A, 141–144 (2011).
Nizon, M. et al. Phenotype-genotype correlations in 17 new patients with an Xp11.23p11.22 microduplication and review of the literature. Am. J. Med. Genet. Part A 167A, 111–122 (2015).
Golabi, M., Ito, M. & Hall, B. D. A. new X-linked multiple congenital anomalies/mental retardation syndrome. Am. J. Med. Genet. 17, 367–374 (1984).
Qi, Y. et al. PQBP-1 is expressed predominantly in the central nervous system during development. Eur. J. Neurosci. 22, 1277–1286 (2005).
Liu, X. et al. Dynamic regulation of alternative polyadenylation by PQBP1 during neurogenesis. Cell Rep. 43, 114525 (2024).
Liu, W. et al. PQBP1 regulates striatum development through balancing striatal progenitor proliferation and differentiation. Cell Rep. 42, 112277 (2023).
Ito, H. et al. In utero gene therapy rescues microcephaly caused by Pqbp1-hypofunction in neural stem progenitor cells. Mol. Psychiatry 20, 459–471 (2015).
Shen, Y. et al. PQBP1 promotes translational elongation and regulates hippocampal mGluR-LTD by suppressing eEF2 phosphorylation. Mol. Cell 81, 1425–1438 e1410 (2021).
Huang, X., Cheng, S. & Han, J. Polyglutamine binding protein 1 regulates neurite outgrowth through recruiting N-WASP. J. Biol. Chem. 300, 107537 (2024).
Sudol, M., McDonald, C. B. & Farooq, A. Molecular insights into the WW domain of the Golabi-Ito-Hall syndrome protein PQBP1. FEBS Lett. 586, 2795–2799 (2012).
Tapia, V. E. et al. Y65C missense mutation in the WW domain of the Golabi-Ito-Hall syndrome protein PQBP1 affects its binding activity and deregulates pre-mRNA splicing. J. Biol. Chem. 285, 19391–19401 (2010).
Musante, L. et al. Common pathological mutations in PQBP1 induce nonsense-mediated mRNA decay and enhance exclusion of the mutant exon. Hum. Mutat. 31, 90–98 (2010).
Tronche, F. et al. Disruption of the glucocorticoid receptor gene in the nervous system results in reduced anxiety. Nat. Genet. 23, 99–103 (1999).
Ye, C., Long, Y., Ji, G., Li, Q. Q. & Wu, X. APAtrap: identification and quantification of alternative polyadenylation sites from RNA-seq data. Bioinformatics 34, 1841–1849 (2018).
Shepard, P. J. et al. Complex and dynamic landscape of RNA polyadenylation revealed by PAS-Seq. RNA 17, 761–772 (2011).
Ding, L. et al. A genome-scale RNAi screen for Oct4 modulators defines a role of the Paf1 complex for embryonic stem cell identity. Cell Stem Cell 4, 403–415 (2009).
Hu, G. et al. A genome-wide RNAi screen identifies a new transcriptional module required for self-renewal. Genes Dev. 23, 837–848 (2009).
Lackford, B. et al. Fip1 regulates mRNA alternative polyadenylation to promote stem cell self-renewal. Embo J. 33, 878–889 (2014).
Zhang, X. Y. et al. Mutations of PQBP1 in Renpenning syndrome promote ubiquitin-mediated degradation of FMRP and cause synaptic dysfunction. Hum. Mol. Genet. 26, 955–968 (2017).
Liu, X., Dou, L. X., Han, J. & Zhang, Z. C. The Renpenning syndrome-associated protein PQBP1 facilitates the nuclear import of splicing factor TXNL4A through the karyopherin beta2 receptor. J. Biol. Chem. 295, 4093–4100 (2020).
Mizuguchi, M. et al. Allosteric modulation of the binding affinity between PQBP1 and the spliceosomal protein U5-15kD. FEBS Lett. 590, 2221–2231 (2016).
Mizuguchi, M. et al. Mutations in the PQBP1 gene prevent its interaction with the spliceosomal protein U5-15 kD. Nat. Commun. 5, 3822 (2014).
Chen, H. I. & Sudol, M. The WW domain of Yes-associated protein binds a proline-rich ligand that differs from the consensus established for Src homology 3-binding modules. Proc. Natl. Acad. Sci. USA 92, 7819–7823 (1995).
Sudol, M. et al. Characterization of the mammalian YAP (Yes-associated protein) gene and its role in defining a novel protein module, the WW domain. J. Biol. Chem. 270, 14733–14741 (1995).
Ji, Z., Lee, J. Y., Pan, Z., Jiang, B. & Tian, B. Progressive lengthening of 3’ untranslated regions of mRNAs by alternative polyadenylation during mouse embryonic development. Proc. Natl. Acad. Sci. USA 106, 7028–7033 (2009).
Nakao, K. & Inoue, K. Cryopreservation for broader production of transgenic mice by DNA injection into zygotes. Exp. Anim. 59, 225–229 (2010).
Nakao, H. et al. A possible aid in targeted insertion of large DNA elements by CRISPR/Cas in mouse zygotes. Genes 54, 65–77 (2016).
Shen, Y., Han, J. & Zhang, Z. C. Novel regulation of the eEF2K/eEF2 pathway: prospects of ‘PQBP1 promotes translational elongation and regulates hippocampal mGluR-LTD by suppressing eEF2 phosphorylation. J. Mol. Cell Biol. 13, 392–394 (2021).
Da Silva, F. et al. Mitotic WNT signalling orchestrates neurogenesis in the developing neocortex. Embo J. 40, e108041 (2021).
Konno, D. et al. Neuroepithelial progenitors undergo LGN-dependent planar divisions to maintain self-renewability during mammalian neurogenesis. Nat. Cell Biol. 10, 93–101 (2008).
Li, X. et al. Inhibition of Foxp4 disrupts cadherin-based adhesion of radial glial cells, leading to abnormal differentiation and migration of cortical neurons in mice. Neurosci. Bull. 39, 1131–1145 (2023).
Acknowledgements
We thank the Model Animal Research Center of Nanjing University for generating the Pqbp1Y65C knock-in mice and Pqbp1-flox mice, and members of the Han laboratory for their critical comments on the manuscript. This work was supported by STI2030-Major Projects (2021ZD0202500 to J.H.), the National Natural Science Foundation of China (82071266 to Z.C.Z., 32530041 to J.H. and 31800862 to S.C.).
Author information
Authors and Affiliations
Contributions
L.Y. and S.C. executed experiments and interpreted data. X.L. performed RNA-seq data analyses and conducted experiments. W.L. performed biochemical analyses. T.X. and W.X. contributed to the writing and discussion. Z.C.Z. analyzed data and wrote the manuscript. J. H. designed experiments, interpreted data, and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Yuan, L., Cheng, S., Liu, X. et al. The missense mutation Y65C in PQBP1 causes microcephaly and cognitive deficits through a combination of partial loss-of-function and gain-of-function effects. Nat Commun 17, 1463 (2026). https://doi.org/10.1038/s41467-025-68202-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-68202-5