Abstract
Bone aging compromises skeletal integrity and increases vulnerability to osteoporosis and other age-related disorders, underscoring the need for new therapeutic strategies. Although pharmacological and genetic approaches have been widely explored, how cellular mechanical remodeling contributes to bone aging remains unclear. Here, we find that senescent bone marrow stem cells show markedly reduced intracellular force and impaired mechanical behavior. Moderate mechanical stimulation in cell culture and in mice restores cellular force, increases chromatin accessibility at the FOXO1 locus, activates its expression, and reverses cellular senescence and bone aging. These mechanical interventions also improve physical performance in aged female mice and show a tendency to reduce systemic inflammation, whereas excessive force induces chromatin overextension and DNA damage, indicating the necessity of precise force control. In this work, we show that optimized mechanical stimulation provides a simple and effective strategy to counteract age-related bone deterioration and systemic inflammation, offering potential for clinical translation.
Similar content being viewed by others
Introduction
Age-related bone loss, characterized by an imbalance between bone formation and resorption1,2, is a complex pathological process that leads to osteoporosis and an increased risk of fractures3,4,5,6. This disruption in bone homeostasis is exacerbated by cellular senescence, particularly in bone marrow mesenchymal stem/stromal cells (BMSCs), which exhibit diminished osteogenic potential and self-renewal capacity with aging7,8. As a result, bone remodeling is impaired, contributing to skeletal dysfunction and accelerating the aging process9,10,11. Despite growing interest in delaying bone aging, understanding the mechanisms underlying BMSC senescence remains critical for developing effective therapeutic strategies to counteract aging and its associated skeletal diseases.
Senescent BMSCs exhibit broad transcriptional reprogramming, particularly in genes governing proliferation, differentiation, and extracellular matrix organization12,13. These transcriptional alterations are tightly regulated by diverse biochemical signaling pathways, including metabolic14, transcriptional15, and epigenetic processes16. Dysregulated metabolic signaling, such as impaired mitochondrial function or altered NAD⁺/SIRT axis17,18, perturbs redox balance and energy homeostasis, thereby activating stress-responsive pathways that promote senescence. Concurrently, transcriptional regulators such as p53, FOXO, and NF-κB orchestrate the expression of senescence-associated genes, linking cellular stress to irreversible growth arrest and the senescence-associated secretory phenotype (SASP)15,19. At the chromatin level, epigenetic modifications—including histone acetylation, methylation, and DNA methylation changes—further consolidate the senescent state by repressing proliferation-related genes and altering chromatin accessibility16,20,21. Together, these biochemical processes converge to establish a stable and self-reinforcing senescence program. While these factors are well-documented, identifying straightforward, translatable interventions to reverse senescence presents a significant challenge.
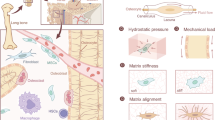
In addition to these biochemical pathways, emerging evidence highlights a critical role for mechanical signaling in modulating transcriptional and epigenetic processes22. The intracellular traction forces, generated by non-muscle myosin II and counterbalanced by cell adhesive environment, can be directly transmitted into the cell nucleus to modulate chromatin structure and gene expression through the linker of nucleoskeleton and cytoskeleton (LINC) complex and lamin skeleton23. Furthermore, various aged cell types, including human dermal fibroblasts, display a marked reduction in chromatin accessibility, leading to impaired transcriptional activity and disrupted gene expression24,25,26. Given that cellular traction force regulates chromatin dynamics and transcriptional activity, it is plausible that it contributes to chromatin condensation and the associated loss of gene expression during cellular senescence. While it is known that cellular mechanotransduction can affect various cell behaviors27, such as cell shape28, differentiation29, and migration30, the precise role of mechanical signals in BMSC senescence remains poorly understood.
BMSCs are embedded in a dynamic microenvironment where they continuously perceive and respond to mechanical cues, affecting cellular traction force and directing cell behavior31. Aging significantly alters the physical properties of the bone microenvironment32,33, disrupting the mechanical signals transmitted to cells. These age-related changes suggest that senescent BMSCs may experience profound shifts in cellular mechanics and mechanotransduction. Exploring the interplay between mechanotransduction and cellular senescence is critical for developing novel strategies to intervene in the aging process and mitigate age-related skeletal disorders.
In this work, we show that aging reduces intracellular traction forces in BMSCs and aged bone tissue, a deficiency that can be reversed through appropriate mechanical stimulation to restore the cell mechanics. Mechanistically, the restoration of cellular traction force enhances chromatin accessibility, leading to the activation of FOXO1 expression. Importantly, FOXO1 knockdown abolished the mechanically rejuvenating effects, underscoring its critical role in mediating cellular responses to mechanical forces. Beyond bone recovery, mechanical interventions improved locomotor activity, alleviated physical frailty, and reduced systemic inflammation in aged mice. These findings highlight both local and systemic benefits of mechanical stimulation, offering a straightforward approach with significant translational potential for combating age-related tissue decline.
Results
Decreased cellular mechanics and mechanotransduction in aged BMSCs
To elucidate the relationship between cellular senescence and mechanotransduction, we isolated human bone marrow stromal cells (BMSCs) from 21 elderly donors (50–80 years) and 17 young donors (18–35 years) for in vitro analysis. The isolated BMSCs were cultured and characterized by flow cytometry, confirming typical MSC surface markers. Compared with young BMSCs, those from elderly donors displayed markedly reduced proliferative and osteogenic potential, indicating age-associated functional decline. In parallel, aged BMSCs exhibited increased p21⁺ and γH2A.X⁺ cells, enhanced SA-β-gal activity34, and upregulated expression of SASP factors (Supplementary Figs. 1–5).
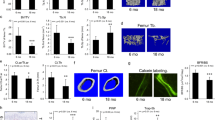
To further delineate the alterations in cellular mechanics associated with aging, we applied a combination of traction force microscopy (TFM), Förster resonance energy transfer (FRET)-based actin tension probes35, and nanoindentation to quantify the mechanical integrity of BMSCs. In TFM assays, cells were cultured on deformable polyacrylamide gels embedded with fluorescent beads, allowing the traction stress generated by cell contraction to be calculated from bead displacement fields. Aged BMSCs produced substantially weaker traction forces than young cells (122 vs. 227 Pa; Fig. 1a), indicating a marked reduction in actomyosin-driven contractility. Consistent with this observation, FRET-based actin tension sensors—which report molecular-scale filament tension by detecting energy transfer between donor and acceptor fluorophores—revealed higher FRET efficiency in aged cells, signifying decreased actin filament tension (Fig. 1b). To complement these findings, nanoindentation analysis was performed to assess the stiffness of individual cells by measuring their resistance to localized deformation; the Young’s modulus of young BMSCs (≈790 Pa) was significantly higher than that of aged cells (≈470 Pa; Fig. 1c), reflecting a more robust cytoskeletal network in the young group. Confocal imaging further illustrated that actin stress fibers36 in aged cells were sparse, disorganized, and randomly oriented, whereas young BMSCs exhibited dense, well-aligned bundles (Fig. 1d). Together, these complementary assays consistently demonstrate that aged BMSCs exhibit markedly diminished traction forces, reduced cytoskeletal tension, lower stiffness, and disrupted actin organization, indicating a comprehensive decline in cellular mechanical competence with aging.
a Schematic illustration, representative images, and quantification of traction force microscopy (TFM) in young and senescent cells. TFM measures the mechanical forces exerted on the substrate by tracking the displacement of embedded fluorescent beads, with larger bead displacements indicating stronger cellular traction forces (n = 11 biologically independent cells per group). b Schematic illustration, representative images, and quantification of traction forces in young and senescent BMSCs measured using FRET-based actin tension probes. A higher FRET/YFP ratio reflects reduced cellular traction force (n = 25 biologically independent cells per group). c Nanoindentation analysis comparing the mechanical properties of young and senescent BMSCs. The schematic illustrates nanomechanical indentation in which stiffness around the nucleus is measured by applying controlled indentation, and the resulting stress–strain curve is used to calculate the Young’s modulus. Higher modulus values indicate a more organized cytoskeletal network. Quantification of nanoindentation measurements is shown (n = 10 biologically independent cells per group). d Analysis of actin stress fiber anisotropy in young and senescent BMSCs. Actin stress fiber anisotropy reflects the directional alignment and organization of cytoskeletal fibers, which are critical for cellular polarization, force transmission, and mechanosensing. Fiber orientation was visualized using a color-coded representation to highlight differences between young and senescent BMSCs (n = 30 biologically independent cells per group). e Western blot analysis (n = 3 biologically independent experiments, uncropped scans of western blots are provided in Supplementary Fig. S34) and f immunofluorescence (n = 50 biologically independent cells per group) of mechanotransduction markers, including Myosin IIa (red), P-myosin IIa (yellow), Lamin A/C (red), and nuclear YAP localization (red) in young and senescent BMSCs. Green indicates F-actin; and blue indicates DAPI. Data are presented as mean ± SD. Scale bar, 50 μm for (a, b, d, f). Two-sided Student’s t-test was used. Exact P values and sample sizes (n) are provided. All raw data supporting the findings of this study are provided in Source data 1 and 10.
To determine whether this mechanical decline was accompanied by alterations in mechanotransduction, we focused on core components of the canonical cytoskeleton–nucleoskeleton–nucleus signaling axis. Specifically, we examined nonmuscle myosin IIa and its active phosphorylated form (P-myosin IIa at Ser1943), which directly drive traction force generation; Lamin A/C, which transmits mechanical stress from the cytoskeleton to the nucleus and maintains nuclear structural integrity; and the transcriptional coactivator YAP, whose nuclear localization serves as a readout of force-dependent mechanosignaling. Western blotting and immunofluorescence analyses revealed consistent reductions in total myosin IIa, P-myosin IIa, Lamin A/C, and nuclear YAP in aged BMSCs (Fig. 1e–f; Supplementary Fig. 6). These findings collectively indicate that the age-related decline in cellular mechanics is tightly linked to impairment of the cytoskeletal–nucleoskeletal mechanotransduction cascade, providing a mechanistic basis for the attenuated force responsiveness and functional deterioration observed in aged BMSCs.
Increased cellular traction force alleviates cellular senescence
Our study identified significant differences in mechanical properties between young and senescent cells, which led us to hypothesize that increased cellular traction force might contribute to functional restoration and alleviate senescence. The decreased mechanotransduction of senescent cells was then restored through mechanical stretching (5% strain and 0.02 Hz frequency). In addition, calyculin A (Supplementary Figs. 26–28 showed that calyculin A exhibited no cytotoxicity and maintained efficiency at the working concentration), the myosin light chain phosphatase inhibitor, was employed to promote the myosin II activity and cell traction force37. After confirming that both methods effectively increased cellular traction forces (Supplementary Fig. 7), we evaluated markers of cellular senescence. Enhanced traction forces resulted in a notable reduction in the expression of p16INK4a (Fig. 2a), p53, and p21WAF1/Cip1, as well as a decrease in the proportion of SA-β-gal-positive cells (Supplementary Figs. 9 and 10). Furthermore, enhancing cellular force in young BMSCs produced a modest reduction in senescence markers, indicating that aged cells are more responsive to mechanical cues than their younger counterparts (Supplementary Fig. 11). These findings provide strong evidence supporting the hypothesis that boosting mechanotransduction can mitigate cellular senescence-related markers.
a Senescent and young BMSCs were subjected to mechanical interventions to alter intracellular traction forces. In senescent cells, traction force was increased using either mechanical stretching (5% strain, 0.02 Hz) or calyculin A treatment (myosin light chain phosphatase inhibitor), resulting in a significant reduction of p16INK4a expression. In young cells, traction force was reduced using either soft hydrogels (1.5 kPa) or blebbistatin treatment (selective myosin II inhibitor), which led to elevated p16INK4a expression (red, n = 50 biologically independent cells per group). Green indicates F-actin; and blue indicates DAPI. b Proliferation analysis using Ki67 immunostaining (red) and EdU incorporation (green). Green indicates F-actin; and blue indicates DAPI. Senescent BMSCs were subjected to enhanced traction force via mechanical stretching, while young BMSCs were exposed to reduced traction force on soft hydrogels (1.5 kPa). Representative images and quantification (n = 6 biologically independent samples per group) are shown. c, d Mechanotransduction and senescence analysis in 3D bone marrow-mimicking microenvironments. Senescent and young BMSCs were cultured in (c) GelMA hydrogels or (d) ECM-mimicking 3D fiber networks (Young’s modulus: 1–4 kPa). Cellular traction forces were pharmacologically modulated: senescent BMSCs were treated with calyculin A to enhance force, whereas young BMSCs were treated with blebbistatin to reduce force. Immunofluorescence analysis of p21 (yellow) and Lamin A/C (red) was performed to assess cellular senescence and mechanotransduction activity, respectively (n = 35 biologically independent cells per group). Data are presented as mean ± SD. Scale bar, 10 μm for p21 in fig. e, f 50 μm for others. One-way ANOVA followed by Tukey’s post hoc test (two-sided) was used, and adjustment for multiple comparisons was applied. Exact P values and sample sizes (n) are provided. All raw data supporting the findings of this study are provided in Source data 2.
Given that replicative depletion is a key feature of aging, we examined the impact of mechanotransduction on cell proliferation using EdU assays, Ki67 staining, and CCK8 assays (Fig. 2b and Supplementary Fig. 13). Our results showed that restoring traction force in senescent cells significantly improved proliferation. These findings highlight the critical role of mechanotransduction in aging, affecting both senescence markers and proliferative capacity.
Furthermore, to investigate the impact of the 3D microenvironment in bone marrow38 (soft tissue) on BMSCs, we employed gelatin methacrylate (GelMA) hydrogels and ECM-mimicking GelMA fiber networks (Young’s modulus of 1–4 kPa, Supplementary Fig. 14), both engineered to replicate bone marrow-like stiffness. Consistent with observations on 2D substrates, mechanotransduction activity, as indicated by Lamin A/C levels, remained higher in young cells compared to senescent cells in both 3D models (Fig. 2c, d). Moreover, enhanced mechanotransduction activity through calyculin A treatment significantly reduced p21 expression in senescent cells, highlighting the pivotal role of mechanotransduction in mitigating cellular senescence even within a 3D bone marrow-mimicking microenvironment.
On the other hand, we explored that weather-reduced mechanical stimuli could induce cellular senescence. To model reduced mechanical stimuli, young BMSCs were seeded on the soft hydrogels with a shear modulus of 1.5 kPa to adjust force balance at the cell adhesion interface, resulting in decreased intracellular traction39. Alternatively, blebbistatin (Supplementary Figs. 26–28 showed that blebbistatin exhibited no cytotoxicity and maintained efficiency at the working concentration), a selective myosin II inhibitor, was used to treat the young cells, which effectively eliminated intracellular traction forces40. Following validation of mechanotransduction disruption (Supplementary Fig. 8), we assessed levels of p16INK4a, p53, p21WAF1/Cip1 and SA-β-gal staining. Both mechanical and pharmacological interventions that attenuated cellular mechanotransduction induced age-associated phenotypes and promoted stem cell senescence (Fig. 2a and Supplementary Figs. 9 and 10). Diminishing cellular force in senescent cells further accelerated aging, albeit less markedly than in young cells, underscoring a reduced but still consequential sensitivity to mechanotransduction impairment (Supplementary Fig. 12). Moreover, EdU assays, Ki67 staining, and CCK8 assays demonstrated a significant reduction in proliferation capacity of young BMSCs subjected to diminished intracellular traction forces, reinforcing the conclusion that reduced mechanotransduction drives cellular senescence and impairs cellular proliferation (Fig. 2b and Supplementary Fig. 13). These effects were notably validated in the 3D GelMA hydrogels and ECM-mimicking fiber networks (Young’s modulus: 1-4 kPa), where reduced cellular forces induced by blebbistatin in young cells promoted senescence phenotypes (Fig. 2c, d).
Overall, cellular mechanotransduction serves as a pivotal regulator in the aging process.
Diminished mechanotransduction activity in aged bone tissues
Considering the decreased cellular traction force in aged BMSCs, we further dissected the mechanotransduction activity in aged tissues. Compared to 2-month-old mice, 20-month-old mice exhibited a significant decline in trabecular number, increased trabecular spacing, and decreased collagen I levels (Fig. 3a, b). These changes in bone microarchitecture severely compromise the niche supporting BMSCs, emphasizing the need to understand mechanotransduction pathways in aging.
a Representative micro-CT assay (n = 6 biologically independent mice per group), b collagen I immunofluorescence staining and Masson staining (n = 8 biologically independent mice per group), c p-FAK immunofluorescence staining (green, n = 10 biologically independent mice per group), d immunohistochemistry for nuclear YAP localization (n = 10 biologically independent mice per group), e Lamin B immunofluorescence staining (red, n = 10 biologically independent mice per group), f immunohistochemistry for p16 (n = 8 biologically independent mice per group), g The p21 (n = 8 biologically independent mice per group), and h rH2A.X (n = 8 biologically independent mice per group) of bone marrow tissue in 20-month-old versus 2-month-old female C57BL/6J mice. Data are presented as mean ± SD. Scale bar, 500 μm for (b), 20 μm for (c–h). Two-sided Student’s t test was used. Exact P values and sample sizes (n) are provided. All raw data supporting the findings of this study are provided in Source data 3.
To assess mechanotransduction activity, we analyzed the phosphorylation of focal adhesion kinase (p-FAK) at Tyr39741 and the nucleus YAP42, which are well-known mechanotransduction markers and are sensitive to intracellular traction force. Immunofluorescence or immunohistochemistry analysis revealed a notable downregulation of both p-FAK (Fig. 3c) and YAP (Fig. 3d) in bone marrow tissue from aged mice (20 months), in stark contrast to the tissue from younger mice (2 months). In addition, the nuclear lamina in aged mice displayed discontinuities and reduced lamin B levels, indicating compromised nuclear integrity and decreased nuclear tension (Fig. 3e). Aging markers, such as p16INK4a (Fig. 3f), P21 (Fig. 3g) and rH2A.X (Fig. 3h), were markedly elevated in older mice compared to younger mice.
These findings collectively highlight an age-associated decline in cellular mechanotransduction activity within the bone marrow, accompanied by compromised nuclear structure and increased senescence markers.
Mechanical stimulation modulates bone aging and remodeling in vivo
To investigate how mechanical cues regulate bone aging, we applied two complementary in vivo paradigms in both aged and young mice: vibrational loading (20 min/day for 30 days with rest days; force+) and tail suspension (30 consecutive days; force−) (Fig. 4a).
a Schematic of in vivo experimental design. Vibrational loading (20 min/day for 30 days, alternating with rest days; force+) and tail suspension (30 consecutive days; force−) were applied to aged (20-month-old) and young (2-month-old) female C57BL/6J mice (n = 8 biologically independent mice per group). b Representative images and (c) quantitative analysis of p-FAK immunofluorescence staining (green, n = 7 biologically independent mice per group), immunohistochemistry for nuclear YAP localization (n = 8 biologically independent mice per group), Lamin B immunofluorescence staining (red, n = 7 biologically independent mice per group) in aged mice, aged mice subjected to vibration treatment (force+), and young mice, young mice subjected to tail suspension (force−). d Immunohistochemistry images of senescence markers p16 (n = 7 biologically independent mice per group) and p21 (n = 8 biologically independent mice per group) in trabecular bone. Quantification shows reductions after vibration in aged mice and increases after tail suspension in young mice. e qPCR analysis of inflammatory cytokines IL-1β, IL-6, CXCL1, and CCL2 in bone marrow (n = 3 biologically independent mice per group). Expression decreased after vibration in aged mice and increased after tail suspension in young mice. f Representative double immunofluorescence images of CD105 (green) and TNF-α (meganta) in trabecular bone. Quantification of TNF-α⁺ CD105⁺ cells shows a decrease after vibration in aged mice and an increase after tail suspension in young mice (n = 5 biologically independent mice per group). H&E (g), Masson’s trichrome (h), and micro-CT (i) images of trabecular bone. Quantification shows recovery of trabecular architecture and mass after vibration in aged mice and trabecular loss after tail suspension in young mice (n = 5 biologically independent mice per group). Data are presented as mean ± SD. Scale bar, 20 μm for (b, d, f), 200 μm for (g, h). Two-sided Student’s t test was used for fig. i One-way ANOVA followed by Tukey’s post hoc test (two-sided) was used for other figures, and adjustment for multiple comparisons was applied. Exact P values and sample sizes (n) are provided. All raw data supporting the findings of this study are provided in Source data 4.
In aged mice, tail suspension did not further reduce trabecular or cortical bone mass (Supplementary Fig. 15), indicating that acute unloading has minimal impact once age-associated bone loss is established. We therefore focused on the effects of vibrational loading. Vibration markedly increased mechanotransduction in femoral trabecular bone, as indicated by elevated p-FAK levels, greater nuclear YAP localization, and improved nuclear envelope integrity reflected by higher Lamin B expression (Fig. 4b, c). These changes were accompanied by reductions in senescence-associated markers (p21, p16, and p53), demonstrated by immunohistochemistry and western blot (Fig. 4d and Supplementary Fig. 16). Bone marrow qPCR likewise showed downregulation of p21, p16, and p19 (Supplementary Fig. 16). Vibration also reduced inflammatory signaling43, with decreased expression of IL-1β, IL-6, CXCL1, CCL2, IL-1α, and TNF-α in bone marrow (Fig. 4e and Supplementary Fig. 17). Double immunofluorescence analysis of CD105 and TNF-α demonstrated a decreased proportion of inflammatory BMSCs following vibrational loading (Fig. 4f), accompanied by reduced expression of CD45 and IL-1β in trabecular bone (Supplementary Fig. 17). Histological staining and micro-CT analysis demonstrated recovery of trabecular architecture and bone mass following vibration (Fig. 4g–i), whereas cortical bone showed no detectable changes (Supplementary Fig. 18). Examination of osteoblast- and osteoclast-associated markers in bone marrow RNA and serum indicated that vibrational loading predominantly enhanced osteoblast activity, with marked increases in Col1a1, Bglap, and serum PINP, while osteoclast markers (Mmp9, Acp5) and CTX were less affected (Supplementary Fig. 19).
In young mice, vibrational loading did not induce detectable changes in trabecular or cortical bone mass (Supplementary Fig. 15). suggesting that younger bone is closer to a homeostatic state and less responsive to additional mechanical input. By contrast, tail suspension produced pronounced trabecular deterioration: mechanotransduction was impaired, as indicated by reduced p-FAK levels, loss of nuclear YAP, and decreased Lamin B expression (Fig. 4b, c), accompanied by upregulation of senescence-associated markers p16, p21, and p53, as shown by immunohistochemistry and Western blot (Fig. 4d and Supplementary Fig. 16). Bone marrow qPCR further confirmed increased expression of p21, p16, and p19 following unloading (Supplementary Fig. 16). Inflammatory cytokines, including IL-1β, IL-6, CXCL1, CCL2, IL-1α, and TNF-α, were significantly elevated in the bone marrow (Fig. 4e and Supplementary Fig. 17), and CD105/TNF-α double staining revealed an increased fraction of inflammatory BMSCs (Fig. 4f). Functionally, tail suspension caused marked trabecular bone loss, as assessed by histology and micro-CT (Fig. 4g–i), while cortical bone remained largely unaffected, as confirmed by micro-CT analysis (Supplementary Fig. 18). Osteoblast and osteoclast analyses confirmed that trabecular loss was primarily due to reduced osteoblast activity, rather than changes in osteoclast function (Supplementary Fig. 19).
Traction forces modulate chromatin accessibility at the FOXO1 locus
To investigate how intracellular traction force regulates aging, we focused on chromatin remodeling, a process linked to chromatin accessibility and gene expression44, and cellular functions. Chromatin remodeling is mediated by mechanotransduction, as previously reported45. Accordingly, we conducted a comprehensive analysis of chromatin accessibility in young and senescent cells. Confocal microscopy imaging of nuclear DNA distribution and subsequent calculation of chromatin condensation parameters (CCP) revealed distinct differences: senescent BMSCs exhibited a more heterogeneous DNA staining pattern compared to young cells, indicating increased chromatin condensation. When traction forces in senescent cells were enhanced using mechanical stretching or calyculin A treatment, DNA distribution became more homogeneous, reflecting decreased chromatin condensation. Conversely, reducing mechanical stimuli through soft hydrogels or blebbistatin treatment in young cells resulted in a DNA distribution resembling that of senescent cells (Fig. 5a, b).
Representative heatmaps derived from DAPI fluorescence images and corresponding chromatin condensation parameter (CCP) maps estimated via edge detection method for cells with (a) traction force modulation (n = 15 biologically independent cells) and (b) treated with different drugs (n = 6 biologically independent cells). Force (+) induced through mechanical stretching or calyculin A treatment, and Force (–) induced by culturing young cells on soft substrates or treating with blebbistatin. Scale bar, 3 μm for a, b. One-way ANOVA followed by Tukey’s post hoc test (two-sided) was used in (a, b), and adjustment for multiple comparisons was applied. c Accessible ATAC-seq peaks at transcription start sites (TSSs) are shown for senescent, Force (+), young, and Force (−) cells. The Force (+) group was induced through mechanical stretching, while the Force (–) group was generated by culturing young cells on soft substrates (n = 2 biologically independent samples per group). d KEGG pathway enrichment analysis reveals differences in aging-related pathways among accessible ATAC-seq peaks in comparisons of Force (+) vs. senescent cells, Force (–) vs. young cells, and senescent vs. young cells. e FOXO signaling pathway analysis of differentially accessible ATAC-seq peaks at TSSs in comparisons of Force (+) vs. senescent cells, Force (–) vs. young cells, and senescent vs. young cells (n = 2 biological independent samples per group). f IGV analysis showing differentially accessible ATAC-seq peaks at FOXO1 among senescent, Force (+), young, and Force (−) cells (n = 2 biologically independent samples). Data are presented as mean ± SD. Differential accessibility was analyzed using DESeq2 (two-sided exact test based on a negative binomial distribution) with Benjamini–Hochberg correction. KEGG enrichment was performed with clusterProfiler using a one-sided hypergeometric test with BH-adjusted P values. Exact P values and sample sizes are provided, and all raw data supporting the findings are included in Source data 5.
We further assessed chromatin accessibility by analyzing histone modifications, which serve as key indicators of chromatin openness and DNA packaging. Chromatin accessibility is closely linked to histone acetylation (H3ac and H4ac), deacetylase 2 and 3 (HDAC2 and HDAC3), and demethylation of lysine 9 and/or 27 in histone H3 (H3K9me3, H3K27me3)46,47. Enhanced traction forces via calyculin A in senescent BMSCs significantly restored histone acetylation levels, whereas reduced traction forces in young cells triggered deacetylation of H3 and H4 (Supplementary Fig. 20). This pattern was corroborated by the expression levels of HDAC2 and HDAC3, which decreased in response to increased cellular forces (Supplementary Fig. 21). Additionally, the demethylation of H3K9me3, but not H3K27me3, was responsive to force modulation (Supplementary Fig. 20).
To further investigate the role of traction forces in chromatin accessibility, we performed ATAC-seq48 to analyze changes in the chromatin landscape. The analysis included senescent cells subjected to mechanical stretching (5% strain, 0.02 Hz), referred to as “senescent, Force (+),” and young MSCs exposed to reduced mechanical stimuli, referred to as “young, Force (-)”. Supplementary Table 1, Figs. 29–33 confirm the high quality of ATAC-seq libraries and reflect the effects of mechanical force modulation on chromatin accessibility. Chromatin accessibility at transcription start sites (TSSs) revealed a significant decrease in accessible peaks in the Force (−) group compared to the young group. In contrast, the Force (+) group exhibited higher TSS accessibility peaks compared to the senescent group, approaching levels observed in the young group (Fig. 5c). These findings align with CCP and histone modification data. KEGG enrichment analysis revealed that force-dependent changes in chromatin accessibility impacted key pathways involved in cellular senescence (Supplementary Fig. 22). To elucidate the mechanisms by which mechanical force regulates cellular senescence, we then concentrated on pathways associated with cellular senescence. Among these, FOXO signaling emerged as one of the most significantly impacted pathways (Fig. 5d), underscoring its pivotal role in the force-regulation of aging49. Chromatin landscape analysis of FOXO signaling genes (Fig. 5e), along with IGV visualization (Fig. 5f), specifically highlighted FOXO1 as a critical force-sensitive target. ATAC-seq data showed that chromatin accessibility at the TSS of FOXO1 was markedly reduced in the Force (−) group compared to the young group, consistent with the global decrease in TSS accessibility under reduced mechanical stimuli. In contrast, the Force (+) group exhibited increased TSS accessibility at FOXO1 compared to the senescent group, approaching levels observed in the young group (Fig. 5e, f). These trends align with the broader ATAC-seq findings, supporting the hypothesis that FOXO1 activation mediates the anti-senescence effects of traction forces.
Traction forces mediate cellular rejuvenation through FOXO1
To test whether intracellular forces regulate FOXO1 expression, we first examined its transcriptional levels under different mechanical conditions. Quantitative PCR revealed that senescent cells subjected to mechanical stretching showed a significant upregulation of FOXO1 expression, whereas young cells cultured on soft substrates (reduced force) displayed marked downregulation (Fig. 6a). We next assessed FOXO1 protein expression. Immunofluorescence staining demonstrated that traction force enhancement, either by mechanical stretching or calyculin A treatment, increased FOXO1 levels in senescent cells, whereas force reduction by blebbistatin treatment or soft substrates suppressed FOXO1 expression in young cells (Fig. 6b).
a Quantitative PCR analysis of FOXO1 expression in senescent cells with or without mechanical stretching, and in young cells cultured on stiff versus soft substrates (n = 3 biologically independent samples per group). b Representative immunofluorescence images showing FOXO1 expression (blue) in senescent cells subjected to mechanical stretching or calyculin A (force+) and in young cells treated with blebbistatin or cultured on soft substrates (force−) (n = 50 biologically independent cells per group). c qPCR validation of FOXO1 knockdown efficiency in senescent cells (n = 3 biologically independent samples per group). d Immunofluorescence staining of p21 (yellow, n = 35 biologically independent cells) and p16 (blue, n = 50 biologically independent cells per group) in FOXO1 knockdown senescent cells following mechanical stretching, demonstrating that enhanced forces failed to reduce senescence marker expression in the absence of FOXO1. e Representative nuclear images and MATLAB-processed chromatin condensation parameters (CCP) from bone marrow cells of aged, aged + vibration, young, and young + tail suspension mice. Quantification of nuclear circularity (n = 19 biologically independent cells) and CCP values (n = 20 biologically independent cells) showed increased nuclear circularity and chromatin openness in aged mice under vibration, whereas tail suspension showed opposite effects in young mice. f Immunohistochemical staining of histone H4 acetylation (H4ac, red) in trabecular bone beneath the growth plate. g Quantification of H4ac staining intensity (n = 8 biologically independent mice per group), showing elevated levels in aged mice with vibration and reduced levels in young mice with tail suspension. h Double immunohistochemistry of CD105 and FOXO1 in trabecular bone. i Quantification of FOXO1⁺ (meganta) CD105⁺ (green) cells, demonstrating increased FOXO1 expression in aged mice subjected to vibration and reduced FOXO1 expression in young mice under tail suspension (n = 6 biologically independent mice per group). Data are presented as mean ± SD. Scale bar, 50 μm for (b, d), 3 μm for (e), 20 μm for f, h. Two-sided Student’s t-test was used for (c). One-way ANOVA followed by Tukey’s post hoc test (two-sided) was used for other figures, and adjustment for multiple comparisons was applied. Exact P values and sample sizes (n) are provided. All raw data supporting the findings of this study are provided in Source data 6.
To determine whether FOXO1 is essential for force-mediated rejuvenation, we silenced FOXO1 in senescent cells (Fig. 6c) and evaluated senescence markers under mechanical stretching. Increased forces failed to reduce p21 and p16 expression upon FOXO1 knockdown (Fig. 6d), indicating that FOXO1 is indispensable for the rejuvenating effects of traction forces.
We then extended these observations in vivo. Aged mice subjected to vibration (Force +) exhibited significantly increased nuclear circularity compared with aged controls, whereas young mice under tail suspension (Force –) displayed reduced nuclear circularity (Fig. 6e). These findings reflect force-dependent alterations in nuclear mechanics and chromatin openness. We further assessed histone acetylation, a well-established marker of chromatin accessibility. Immunostaining for H4ac revealed increased levels in trabecular bone regions beneath the growth plate of aged mice exposed to vibration, whereas young mice subjected to tail suspension showed reduced H4ac expression (Fig. 6f, g).
Finally, to confirm FOXO1 dynamics in bone marrow stem cells, we performed CD105 and FOXO1 double immunohistochemistry. FOXO1 positivity was markedly upregulated in aged mice under vibration, but downregulated in young mice subjected to tail suspension (Fig. 6h, i), consistent with the in vitro findings.
In summary, these results demonstrate that intracellular traction forces modulate senescence through chromatin remodeling and FOXO1 activation.
Mechanical overstimulation accelerates cellular senescence
Moderate levels of mechanical stimulation significantly suppressed aging markers in BMSCs, as described above. We next asked whether intensifying the mechanical input would further augment these rejuvenating effects. To this end, cells were subjected to higher-frequency cyclic stretching (5% strain at 0.1 Hz; Fig. 7a) or to excessive strain amplitude (25% strain at 0.02 Hz; Fig. 7b). Contrary to expectation, these intensified regimens did not enhance rejuvenation. Instead, they provoked a senescent phenotype, as demonstrated by upregulation of senescence markers p16INK4a and p21WAF1/Cip1 (Fig. 7a–c). The pro-senescent effect of force overstimulation was further corroborated in senescent BMSCs exposed to continuous calyculin A for 72 h, both in ECM-mimicking 3D fiber networks (Fig. 7d) and on conventional tissue culture polystyrene (TCPS; Supplementary Figs. 23 and 24). These findings indicate that intracellular traction force is a double-edged sword in the aging process: while moderate levels restore youthful features, excessive force overwhelms cellular tolerance, tipping the balance toward irreversible senescence.
Representative immunofluorescence images of p16 (blue) and p21 (blue) in senescent cells subjected to mechanical stretching with (a) varying frequency (0.02 and 0.1 Hz, at 5% strain) and b varying strains (5% and 25%, at 0.02 Hz). Yellow indicates F-actin; and meganta indicates DAPI. c The quantitative assay of p16 and p21 in senescent cells subjected to varying mechanical stretching. p16 (n = 33 biologically independent cells per group) and p21 (n = 39 biologically independent cells per group) in senescent cells subjected to varying frequency. p16 (n = 64 biologically independent cells) and p21 (n = 80 biologically independent cells) in senescent cells subjected to varying strains. d Representative immunofluorescence images and analysis of p21 (yellow) in cells on the ECM-mimicking 3D fiber networks with calyculin A treatment for 24 or 72 h (n = 32 biologically independent cells). Comet assay and observation of compromised tail DNA percent and tail moment length upon treatment with (e) varying frequency and (f) varying strains. Tail DNA percent (n = 20 biologically independent cells) and tail moment length (n = 12 biologically independent cells) in senescent cells subjected to varying frequency. Tail DNA percent (n = 20 biologically independent cells) and tail moment length (n = 18 biologically independent cells) in senescent cells subjected to varying strains. g Measurement of relative hydrodynamic radius (Rh) of DNA subjected to ultraviolet (UV) irradiation for 150 s. DNA was either unencapsulated or encapsulated in protamine (n = 20 biologically independent DNA. Data are presented as mean ± SD. Scale bar, 50 μm for (a, b, e, f, g). 10 μm for (d). Two-sided Student’s t-test was used for (g). One-way ANOVA followed by Tukey’s post hoc test (two-sided) was used for other figures, and adjustment for multiple comparisons was applied. Exact P values and sample sizes (n) are provided. All raw data supporting the findings of this study are provided in Source data 7.
Given the established role of DNA damage as a central driver of cellular aging, we next investigated whether force-induced senescence was associated with increased genomic instability. Comet assays revealed a pronounced rise in DNA damage under overstimulation, with significantly higher tail DNA content and extended tail moment length in senescent BMSCs subjected to high-intensity stretching, whereas moderate stimulation did not alter these parameters (Fig. 7e, f).
The force-induced DNA damage can be partially explained by the loss of steric effect resulting from highly condensed DNA, which was proven by a molecular model mimicking chromatin. The isolated DNA chains assembled onto protamine, mimicking the condensed chromatin structure within the cell nucleus, exhibit protection against fragmentation induced by ultraviolet irradiation. In contrast, freely dispersed DNA chains, which simulate the dissociation of DNA from histones under intracellular traction force, were fragmented into smaller pieces upon exposure to ultraviolet irradiation (Fig. 7g).
Optimal mechanical stimulation reduces bone aging and inflammation
To evaluate the dose-dependent effects of mechanical stimulation on bone recovery in aged mice, we performed vibrational loading over a 30-day period. Three experimental groups were established: intermittent vibration (force+, 20 min/day with alternating rest days), continual vibration (force++, 20 min/day without rest), and baseline controls (Fig. 8a). Intermittent vibration significantly enhanced trabecular bone formation, as evidenced by increased collagen deposition (Fig. 8b) and improved trabecular architecture observed via micro-CT (Fig. 8c). In contrast, continual vibration led to compromised trabecular structure and reduced collagen levels (Fig. 8b, c), indicating that excessive mechanical stimulation is detrimental to bone integrity.
a Schematic illustration of modulating mechanical force on hindlimbs of aged female C57BL/6J mice (20-month-old), with intermittent vibrational loading (force+) and continual vibrational loading (force++). b Masson staining and collagen I immunofluorescence staining in aged mice subjected to varying doses of vibration (n = 8 biologically independent mice per group). c Representative images and microarchitecture parameters of the distal femoral trabecular metaphysis from aged mice subjected to varying doses of vibration. (n = 5 biologically independent mice per group). d Representative images and quantitative analysis of immunohistochemistry for p21 and p16 in aged mice subjected to varying doses of vibration (n = 8 biologically independent mice per group). e Western blot analysis of p53 levels in aged mice subjected to varying doses of vibration (n = 3 biologically independent mice, uncropped scans of western blots are provided in Supplementary Fig. S35). f Representative images and quantitative analysis of immunohistochemistry for CD45 and IL-1β in aged mice subjected to varying doses of vibration (n = 8 biologically independent mice per group). g Representative images and quantitative analysis of DNA-RNA damage and rH2A.X levels in various experimental groups subjected to varied doses of vibration (n = 8 biologically independent mice per group). h Physical tests, including the grip strength assessment, treadmill running, and water maze were conducted across different experimental groups (n = 5 biologically independent mice). Blood test including serum IL-6, IL-1β levels, and platelet numbers from the related experimental groups (n = 5 biologically independent mice per group). i, j Representative images and quantitative analysis of immunohistochemistry for CD68 and S100A8 in liver (i) and kidney (j) (n = 5 biologically independent mice per group). Data are presented as mean ± SD. Scale bar, 100 μm for (b), 10 μm for (d, f, i, j), 20 μm for (g). Two-sided Student’s t-test was used for (h–j). One-way ANOVA followed by Tukey’s post hoc test (two-sided) was used for other figures, and adjustment for multiple comparisons was applied. All raw data supporting the findings of this study are provided in Source data 8 and 10.
Consistently, analysis of aging markers revealed that intermittent vibration suppressed expression of p16, p21, and p53, whereas continual vibration elevated these markers (Fig. 8d, e and Supplementary Fig. 25).
Inflammatory responses within trabecular bone followed a similar pattern. Intermittent vibration lowered IL-1β and CD45 expression (Fig. 8f), while continual vibration upregulated these markers. Bone marrow PCR analysis further supported these findings, with intermittent vibration reducing IL-1β, CXCL1, CCL2, and TNF-α expression, and continual vibration promoting their expression (Supplementary Fig. 25). DNA damage was elevated following continual vibration but remained unchanged under intermittent stimulation (Fig. 8g), highlighting the importance of force intensity in determining cellular outcomes. Together, these data indicate that judicious mechanical stimulation through intermittent vibration mitigates aging- and inflammation-associated markers while promoting trabecular bone recovery.
Beyond the skeletal effects, intermittent vibration conferred systemic benefits. Mice subjected to intermittent vibration demonstrated improved physical performance across grip strength, treadmill running and water maze tests. Serum analysis revealed reduced levels of IL-6 and IL-1β, as well as lower platelet counts (PLT), consistent with decreased systemic inflammation (Fig. 8h). Furthermore, assessment of key peripheral organs involved in immune modulation—liver and kidney—showed reduced expression of inflammatory markers CD68 and S100A8 following intermittent vibration (Fig. 8i, j), confirming broad anti-inflammatory effects.
Discussion
Our findings delineate a mechanistic axis whereby cellular traction forces orchestrate BMSC aging through modulation of chromatin architecture and activation of FOXO1. Adequate mechanical stimulation promotes localized chromatin decondensation at the FOXO1 locus, enhancing transcriptional activity and facilitating cellular rejuvenation. FOXO1, a transcription factor with established roles in oxidative stress resilience, DNA repair, apoptosis, and metabolic regulation49, functions as a central effector translating mechanical cues into anti-senescent gene expression programs. Conversely, supraphysiological mechanical stimuli induce aberrant chromatin unfolding and DNA damage, while attenuated mechanical input triggers chromatin condensation and FOXO1 repression, thereby accelerating cellular senescence. These observations underscore that both the magnitude and temporal patterning of mechanical cues are critical for preserving genomic integrity and maintaining cellular youthfulness (Fig. 9).
This model highlights how intracellular traction forces modulate cellular senescence by influencing chromatin accessibility. Increased traction force promotes localized chromatin opening, facilitating FOXO1 activation and cellular rejuvenation. Conversely, reduced traction force leads to chromatin condensation, suppressing FOXO1 expression and accelerating senescence. Excessive traction force, however, induces DNA damage within open chromatin regions, exacerbating aging processes.
The interplay between mechanical forces and chromatin architecture is further corroborated by prior studies showing that nuclear deformation induced by intracellular tension remodels chromatin organization and shapes the epigenetic landscape8,50,51. The effects are context-dependent: moderate tension promotes transcriptional competence52, whereas excessive or prolonged stress fosters heterochromatin accumulation, including H3K27me3 enrichment, resulting in transcriptional repression53. Age-related factors, such as citrate carrier degradation14 and increased extracellular matrix stiffness47, synergize with mechanical perturbations to exacerbate chromatin compaction in senescent cells. Our data provide mechanistic insight by demonstrating that precisely controlled mechanical inputs preserve chromatin accessibility and counteract the epigenetic drift characteristic of cellular aging.
The physiological relevance of mechanotransduction in systemic aging is further supported by clinical and experimental evidence. Liu et al.54 showed that exercise elevates systemic anti-senescent and anti-inflammatory factors across multiple tissues, rescuing age-associated phenotypes, while Yue et al.55 identified a population of force-sensitive progenitor cells that mediate cartilage rejuvenation and suppress osteoarthritis progression. Together, these studies substantiate that appropriately applied mechanical cues confer anti-aging benefits at both the cellular and tissue levels. Nonetheless, interindividual variability in senescence status and mechanosensitivity presents challenges for standardizing mechanical interventions, emphasizing the importance of precision mechanotherapy for translational applications.
Despite these advances, several limitations remain. In vivo measurement of cellular forces remains technically challenging, with existing methods offering limited spatiotemporal resolution; developing high-precision, high-resolution techniques will be essential for future studies. While FOXO1 is identified as a key mediator linking mechanical cues to cellular aging, the downstream epigenetic and transcriptional mechanisms remain incompletely understood. The functional relevance of FOXO1-bound target genes to the aging process warrants further systematic investigation. Finally, although this study focused on BMSCs and bone tissue, determining whether targeted mechanical stimulation can similarly rejuvenate other cell populations or tissues will broaden the translational potential of this approach.
In conclusion, our findings highlight the central role of cellular traction forces in regulating chromatin dynamics and activating FOXO1, offering a promising strategy for reversing bone aging. Compared with pharmacological or gene-editing strategies, which face substantial safety concerns and high costs, mechanical stimulation represents a practical, accessible, and translationally viable approach to counteract age-related bone degeneration and systemic inflammatory decline.
Methods
BMSC isolation and culture
Bone marrow aspirates (~1–2 mL) were collected from the posterior superior iliac spine (PSIS) of healthy donors aged 18–35 years (young group) and 60–80 years (elderly group), all of Chinese descent, at West China Hospital of Sichuan University (n = 38 donors). Donors were screened to exclude hematological disorders, and both male and female donors were included, with sex recorded at the time of collection (detailed donor information is provided in the Supplementary Information). Participants gave informed consent to the experiment and the publication of information. All human tissue samples were used exclusively for experimental analyses and were destroyed immediately upon completion of the assays. All procedures involving human participants were reviewed and approved by the Institutional Review Board of West China Hospital, Sichuan University (Approval No. WCHSIRB-D-20220290).
Bone marrow aspirates were processed to isolate mononuclear cells (MNCs) by density gradient centrifugation using Ficoll-Paque, removing red blood cells and other non-nucleated cells. The MNC fraction was plated in T-75 flasks and cultured in low-glucose DMEM supplemented with 10% fetal bovine serum (Gibco) at 37 °C in a humidified atmosphere containing 5% CO2. After 48–72 h, non-adherent cells were removed by medium replacement, and adherent cells were expanded as bone marrow-derived stromal cells (BMSCs). Cells at passages 1–2 were used for all subsequent experiments.
This isolation procedure enriches for BMSCs, as evidenced by their adherence to plastic, fibroblast-like morphology, positive expression of MSC markers CD73 and CD146, negative expression of hematopoietic markers CD34 and CD45, and functional properties including colony-forming ability, proliferation, multilineage differentiation, and mechanosensitive responses.
BMSC treatment and mechanical manipulation
Chemical inhibitors
Blebbistatin (20 μM, B0560, Sigma) or calyculin A (0.1 nM, ab141784, Abcam) was added to the culture medium and maintained throughout the treatment period. Cell viability was assessed using live/dead assays according to the manufacturer’s instructions.
Mechanical stimulation
To enhance cellular traction force, cells were seeded in a stretching device (Cell&Force, Cell Tank, Hangzhou Surface&Force Technology Co., Ltd) with strains of 5% and 25% at frequencies of 0.02 and 0.1 Hz. To reduce cellular traction force, cells were cultured on soft hydrogels with a shear modulus of 1.5 kPa for 24 h.
Genetic manipulation
Electroporation (Celetrix Biotechnologies, China) was used to deliver siRNA targeting FOXO1 (sense: GAGCGUGCCCUACUUCAAG; antisense: CUUGAAGUAGGGCACGCUC). Plasmid DNA constructs were obtained from Hanbio Biotechnology (Shanghai, China).
Cellular mechanics and mechanotransduction in human BMSCs
Nanoindentation
Nano-indentation was used to measure Young’s modulus of young and senescent BMSC. The cells were pressed by a ball-shaped indenter (Piuma Chiaro, Optics11). The diameter of the nano-indentation indenter is 10.5 μm, the stiffness of the indenter is 0.05 N/m, and the indentation depth is 3 μm in this study. The modulus was determined by fitting force-indentation curves to known models according to the manufacturer’s instructions.
Traction force microscopy (TFM)
TFM was used to detect cellular mechanotransduction according to our previous protocol56. The cells were seeded on deformable hydrogels embedded with fluorescent beads. The shifting of the beads, caused by cell adhesion-induced hydrogel deformation, could be recorded to calculate the traction force exerted by the cells. Firstly, RGD-decorated PEG hydrogels containing 0.5 μm fluorescent carboxylated polystyrene beads (Sigma) were fabricated and the micro-beads existed at the surface of the hydrogels. After hydrogels were soaked for 48 h, young and senescent BMSCs were seeded on the hydrogels. A confocal microscope (Leica) was exploited to take images of beads and the spreading area of cells, regarding the shifted position of beads induced by cell migration. Finally, both young and senescent cells were cleared up by dealing with 1% SDS and taken photos afterwards to identify the original position of beads. Images were first aligned to correct for the experimental drift of the samples using the ImageJ plugin “Align slices in stack”. The displacement field in a spread cell region was subsequently calculated by the “particle image velocimetry” plugin in ImageJ. The obtained result was reconstructed using the “Fourier transform traction cytometry” plugin in order to generate the traction force field as a vector plot.
Quantification of fiber’s anisotropy
FibrilTool, a plugin of ImageJ, was introduced to quantify the orientation of fibrillar structures in the studies according to the protocol. After capturing the pictures of β-actin, we calculated the anisotropy of young and senescent cells, respectively, in order to compare the differences in cellular mechanical characteristics. Regarding the anisotropy score, the following convention was used, i.e., 0 for no order (purely isotropic arrays) and 1 for perfectly ordered (purely anisotropic arrays). Another plugin of ImageJ named Orientation J was also used to measure the orientation of the fiber where the color representation reflected different orientations.
Actin tension sensor
Primary extracted young-and senescence-derived human BMSCs were spread on glass cell culture dishes and then cultured for 24 h in the absence of antibiotics. Opti-MEM (Gibco) was used to dilute plasmid Actin-cpstFRET-Actin (80643, Addgene), and X-tremeGENE HP DNA transfection reagent (Sigma) was added at 3 μL/μg to transfect cells for 48 h. Cells expressing β-actin-CFP/YFP-β-actin fusion protein at sufficiently high levels for easy visualization and imaging were selected for imaging. However, cells with excessively high expression levels were avoided. In addition, cells expressing the fusion protein through the whole cell, including the nucleus, were excluded, and only cells expressing the sensor in a discernible cytoplasmic skeleton were imaged.
At 0, 1, 2, 3, 4, 5, 6, 7, 8, 24 h after administration, cells were imaged using a confocal microscope with a ×20 objective (Leica). FRET results (530 nm) were captured with a 405 nm laser to excite β-actin-CFP. At least 20 cells were quantified for each condition. The cell images used for all values shown in the figure were acquired under the same imaging condition. Considering the different expression levels of cytoskeletal proteins in different cells, the YFP fluorescence intensity obtained by fixed 488 nm excitation was standardized to ensure the normal operation of the sensor and FRET assay. Moreover, the ratio of the fluorescence intensity of YFP (530 nm) excited at 405 nm and 488 nm (FRET/YFP) was used as a parameter to reflect the cytoskeletal tension.
Animal models and experimental procedures
Young and aged mice
Female C57BL/6J mice aged 2 months (young group) and 20 months (aged group) were obtained from Chengdu Dossy Experimental Animals Co., Ltd. To investigate the effects of mechanical interventions on bone development, bone aging, and mechanotransduction, both tail suspension and vibrational loading experiments were performed on mice from each age group. All animal procedures were approved by the Ethics Committee of West China Hospital of Stomatology, Sichuan University (approval number: WCHSIRB-D-2022-174), and were conducted in accordance with institutional and national guidelines for the care and use of laboratory animals.
Tail suspension model
Mice were subjected to hindlimb unloading using a tail suspension model for 30 consecutive days. Animals were suspended by taping the tail to a horizontal bar, allowing forelimb contact with the cage floor while keeping the hindlimbs elevated. Mice were housed individually in custom cages and provided with a complete nutritional gel diet during the suspension period. Tail suspension was applied to both young (2 months) and aged (20 months) mice to assess age-dependent responses.
Vibrational loading model
Mechanical stimulation was delivered using vibrational loading for 20 min per day over a 30-day period. Two vibration regimens were applied: (1) intermittent vibrational loading (one day of vibration followed by one day of rest, repeated for 30 days), and (2) daily continuous vibrational loading (20 min per day for 30 consecutive days). Vibrational loading was administered to both young and aged mice to evaluate the effects of increased mechanical stimulation across age groups. A baseline control group without mechanical intervention was included for each age.
All animal procedures were approved by the Ethics Committee of West China Hospital of Stomatology, Sichuan University, and were performed in accordance with institutional guidelines for the care and use of laboratory animals. Multifaceted Assessment of Physiological and Cellular Responses in Animal Models.
Micro CT tomography evaluation
The femurs from mice with different treatments were scanned using micro-CT (SkyScan 1176 desktop X-ray Micro-CT system, Skyscan, Bruker) with 0.5 mm Al filter, image pixel size: 17.75 μm, voltage: 60 kV, electrical current: 400 μA, and exposure time: 1080 ms. The trabecular region of interest (ROI) in the micro-CT image was selected individually and manually to ensure the trabecular bones of the distal femoral metaphysis were included in the ROI as much as possible. The 3D reconstruction was performed using all the outlined slices by NRecon (Skyscan, Bruker) cone-beam reconstruction software. Three-dimensional microarchitecture parameters, including bone volume fraction (BV/TV), structure model index (SMI), trabecular number (Tb. N), trabecular thickness (Tb. Th), and trabecular bone pattern factor (TB. Pf), were calculated using CTAn (CTAn, Skyscan, Belgium).
Histological analyses
Histological analyses involved immunohistochemistry, immunofluorescence, hematoxylin and eosin (HE) staining, as well as Masson staining. The dissected femurs from mice with different treatments were fixed in 4% paraformaldehyde for 2 days and then decalcified in 12% EDTA for 1 week. Slides were subjected to sodium citrate buffer at 99 °C for 20 min for antigen retrieval and then incubated with rabbit polyclonal anti-collagen I (1:800, Servicebio, Cat No: GB11022-3), rabbit anti-phospho-FAK (1:50, Thermo Fisher Scientific, Cat No:700255), anti-Lamin B1(1:100, Abcam, Cat No:ab16048), rabbit polyclonal anti-YAP (1:100, Genxspan, Cat No: GXP539020), rabbit polyclonal anti-p16 (1:100, Invitrogen, Cat No: MA5-17142), anti-p21(1:100, Abcam, Cat No:ab188224), anti-CD45 (1:100, Abcam, Cat No:ab10558), anti-IL-1β (1:100, Santa cruz, Cat No: sc-52012), anti-H4AC(1:100, Abcam, Cat No:ab177790), rabbit polyclonal anti-FOXO1(1:100, Cell signaling, 2880S), rabbit polyclonal anti-phospho-Histone H2A.X (ser139) (1:100, Cell Signaling Technology, Cat No: 9718), anti-DNA-RNA damage (1:100, Abcam, Cat No:ab62623). The anti-rabbit IgG antibody was used as secondary antibody (1:400, SeraCare, Cat No:5220-0336). HE staining was used to assess the histological structure and morphology of the bone tissue. For Masson staining, the tissue was stained by Masson staining solution (Solarbio, China) according to the instruction. Additionally, liver and kidney tissues were stained using immunohistochemistry with anti-S100A8 (1:100, Abcam, Cat No: ab92331) and anti-CD68 (1:100, Abcam, Cat No: ab125212) antibodies to evaluate inflammatory markers. Immunohistochemistry and immunofluorescence slides were scanned in high resolution using the Pannoramic MIDI scanner (3D HISTECH, Hungary) for comprehensive analysis.
Physical test of animals
Grip-strength test
This grip-strength test was performed according to the previously described procedure57. Grip strength of the mouse’s limbs was recorded using a grip strength meter (SA 415, Jiangsu Science Bio-Tech Co., Ltd, Jiangsu, China). Before testing each mouse, the gauge was reset and stabilized to 0 g. The mouse was allowed to grasp a horizontally held metal grid with all four paws, and then it was gently pulled backward by the tail until it could no longer hold onto the grid. The peak pulling force, measured in grams, was automatically recorded on a digital force transducer. Each mouse underwent four trials after adequate rest, and the average value was used to represent the muscle grip strength of the individual mouse.
Treadmill exercise
The treadmill exercise was conducted as previously described58. A treadmill (SA 101B, Jiangsu Science Bio-Tech Co., Ltd, China) with eight lanes (8.5 × 61 cm2 each) and individual moving belts was used to assess exercise endurance in mice. Each lane featured an electric shock grid (0.3 mA for 10 s) to motivate running. Mice underwent pre-training for 1 week at 5 m/min for 10 min, followed by 10 m/min for 30 min, with an acceleration of 70 m/min² on a flat incline. Sessions were conducted twice daily for 40 min with a minimum interval of 3 h. Subsequently, treadmill exercise was performed every morning for 3 days using the same protocol. Exercise tests continued until mice either touched the shock grid 150× or were unable to continue after a maximum shock duration of 10 s. Time to exhaustion, distance covered, and total shocks were recorded.
Morris water maze test
The Morris water maze was used to test spatial learning and memory. The Morris water maze test was performed in an opaque circular pool filled with water (diameter 1.5 m, height 0.6 m, 22−23 °C). An 8 cm diameter escape platform was positioned in the second quadrant of the pool with its top 1 cm below the water surface. A video analysis system (SA201, Jiangsu Science Bio-Tech Co., Ltd, Jiangsu, China) was used to observe and record each mouse’s swimming pattern. All water maze training and testing were conducted during the animals’ active phase of the light cycle (07:00−19:00). Mice were released into the pool from four different starting positions in the pool and allowed to swim for a maximum of 60 s. If a mouse failed to find the platform within 60 s, it was gently guided to the platform and allowed to stay there for 30 s. Before the formal testing began, each animal received two swimming training sessions per day for 5 days, with the second session occurring at least 2 h after the first. During the training trials, the swimming speed, the escape latency and escape distance were recorded. After 5 days of training, mice that still cannot swim and are unable to find the platform within 60 s will be excluded from the study. The formal test was conducted on the 6th day after the training ended, with the platform removed from the pool. The number of crossings of the platform area, average swimming speed, total distance, and the time spent swimming in the second quadrant where the platform had been located were recorded. The swimming pattern of each mouse were recorded using a camera connected to a computer.
Enzyme-linked immunosorbent assay
After the behavioral tests, the mice were sacrificed, and serum was collected from the eyeballs for cytokine analysis. According to the manufacturer’s instructions, the quantitative ELISA kits (Mouse IL-1 beta Uncoated ELISA Kit, 88-7013, Thermo Fisher Scientific Inc, USA; Mouse IL-6 Uncoated ELISA Kit, 88-7064-88, Thermo Fisher Scientific Inc, USA) were used to measure the concentration of IL-1β and IL-6 in the serum. The procedures were done according to the instruction manual: (1) Firstly, coat each well of a Corning™ Costar™ 9018 ELISA plate with 100 µL/well of capture antibody diluted in Coating Buffer. Seal the plate and incubate overnight at 4 °C. Block the wells with 200 µL of ELISA/ELISPOT Diluent (1×) and incubate at room temperature for 1 h. (2) Then, aspirate the wells and wash 3 times with >250 µL/well of Wash Buffer, allowing approximately 1 min of soaking for each wash. Blot the plate on absorbent paper to remove residual buffer. (3) 100 µl of standard, control buffer, and sample were added respectively into wells of the ELISA plate coated with antibody IL-1β or IL-6, then incubated overnight at 4 °C for maximum sensitivity. (4) After that, the wells were washed 5 times. Add 100 µL/well of diluted Avidin-HRP. Seal the plate and incubate at room temperature for 30 min. (5) Washed again, add 100 µL/well of 1× TMB Solution and incubate at room temperature for 15 min in the dark. (6) Add 100 µL/well of Stop Solution. (7) Finally, read absorbance read plate at 450 nm within 30 min of stopping reaction. If wavelength substraction is available, substract the values of 570 nm from those of 450 nm and analyze data.
Western blotting
After euthanising the mice, femurs were quickly dissected, weighed and collected into Eppendorf tubes. Samples were snap-frozen on dry ice and stored at −80 °C for subsequent analysis. Then tissues were lysed in radio immunoprecipitation assay buffer (RIPA) supplemented with 1% protease inhibitor and 1% phosphatase inhibitor and homogenized using a homogenizer at 4 °C. The mixture was spun down at 12,000 × g for 5 min at 4 °C to pellet the insoluble material. The supernatant was lysed through probe sonication on ice to release cytosolic proteins. Proteins were isolated by centrifugation at 12,000 × g for 15 min at 4 °C, resolved by SurePAGE 4–12% Bis-Tris gels, and transferred to PVDF membrane (0.45 µm). The membranes were blocked in block buffer for 1 h at room temperature and incubated with primary antibodies (anti-GAPDH antibody (1:5000, Servicebio, GB15004), anti-p53 antibody (1:1000, Cell Signaling, 2524) in primary antibody dilution buffer (Beyotime, China) overnight at 4 °C. Blots were washed three times with PBST and incubated with secondary antibodies (goat anti-rabbit IgG secondary antibody HRP conjugated (1:10,000, Sianalway Antibody, L3012) and goat anti-mouse IgG secondary antibody HRP conjugated (1:10,000, Sianalway Antibody, L3032) at room temperature for 1 h. Blots were further washed three times with PBST for 5 min. Then blots were treated with an ECL plus detection kit and imaged using ChemiScope 6100 system.
RNA extraction and real-time PCR
Total RNA was isolated from cells using the FastPure Complex Tissue RNA Isolation Kit (RC113-01, Vazyme) following the manufacturer’s instructions. During purification, all centrifugation steps were performed at 12,000 × g for 1 min at 4 °C unless otherwise specified. RNA concentration and purity were assessed by spectrophotometry (NanoDrop One, Thermo Fisher Scientific). For complementary DNA (cDNA) synthesis, 1 μg of total RNA was reverse-transcribed using the HiScript III RT SuperMix for qPCR (R323-01, Vazyme) according to the manufacturer’s protocol. Quantitative PCR was performed using the Taq Pro Universal SYBR qPCR Master Mix (Q712-02, Vazyme) on a CFX96 Real-Time PCR Detection System (Bio-Rad). The thermal cycling conditions were: 95 °C for 30 s, followed by 40 cycles of 95 °C for 10 s and 60 °C for 30 s. Melt curve analysis was carried out to ensure primer specificity. Gene expression was calculated and normalized to GAPDH as the internal reference. Primer sequences used in this study are listed in Supplementary Table 2.
Cellular functional assays
Immunofluorescence and confocal microscopy
Cells were fixed for 15 min at 37 °C with 4% paraformaldehyde, permeabilized with 1% Triton-X100 (Sigma) for 15 min and blocked with 5% BSA diluted in PBS for 15 min at RT. BMSCs were then incubated, overnight at 4 °C with the indicated primary antibodies in 1% BSA. The following antibodies were used: Myosin IIa (1:100, Cell Signaling Technology, Cat No:3403), phospho-Myosin IIa (1:200, Cell Signaling Technology, Cat No:14611), Lamin A/C (1:200, Cell Signaling Technology, Cat No:4777), YAP (1:100, Cell Signaling Technology, Cat No:14074), phospho-Histone H2A.X (ser139) (1:200, Cell Signaling Technology, Cat No: 9718), p16 (1:200, Invitrogen, Cat No: PA1-30670), p21 (1:200, Cell Signaling Technology, Cat No:2947), P53 (1:100, Cell Signaling Technology, Cat No:2524), HDAC2 (1:200, Santa Cruz Biotechnology, Cat SC-9959), HDAC3 (1:200, Cell Signaling Technology, Cat No:3949), Histone H3ac (pan-acetyl) (1:200, Active motif, Cat No: 39040), Histone H4ac (pan-acetyl) (1:200, EMD Millipore, Cat No:06-866), H3K9me3 (1:200, Active motif, Cat No: 39062), H3K27me3 (1:200, Active motif, Cat No: 39055), ki67 (1:200, Abcam, ab15580) and rabbit polyclonal anti-FOXO1(1:100, Cell signaling, 2880S). Following this, cells were incubated for 2 h at 4 °C with the appropriate secondary fluorescent antibodies diluted in 1% BSA, protected from light. At last, nuclei were stained with DAPI (1:1000, Solarbio, China). Images were collected under a confocal laser scanning microscope (CLSM, Olympus, Japan).
Western blot experiments
BMSCs were seeded in six-well plates and treated with different conditions for different times. Then the experiment was finished according to standard protocol. Protein concentration was determined using the BCA Protein Assay Kit (Bio-Rad, U.S.), and then 30 μg of total protein was run on 8–12% SDS-PAGE gel (Beyotime, China), and electro-transferred to polyvinylidene difluoride (PVDF) membranes (Millipore) for 2–3 h at 120 mA in transfer buffer. The following antibodies were used: Myosin II a, phospho-Myosin IIa, Lamin A/C, and GAPDH. The concentration of these antibodies in this experiment was 1:1000. After three 10 min washes with TBST, membranes were incubated with the appropriate secondary antibody (1:5000) for 1 h at room temperature. Immunoreactive bands were visualized by Tanon-5200Multi (Tanon luminous imaging image storage and management software) after staining by a western lightning chemiluminescence detection kit (Zen Bioscience, China).
Senescence-associated β-galactosidase (SA-β-Gal) Staining
Senescent cells were identified by the SA-β-gal staining kit (Solarbio, China) according to standard report. After incubating with the reagent about 8–12 h, senescent cells would be stained with aquamarine blue, whereas young cells were not able to be colored.
Cell proliferation assay
To investigate the effect of cellular mechanotransduction on BMSC proliferation, an EdU incorporation assay (Beyotime, China) was performed. After treating cells under different mechanical stimulation conditions, the assay was conducted according to the manufacturer’s instructions to assess DNA synthesis and proliferation activity.
ECM-mimicking gelatin methacrylate (GelMA) fiber networks
The networks of suspended GelMA fibers were fabricated by a combination of electrospinning and 3D printing technology. GelMA (grafting rate: 90%) was dissolved at 0.1 − 1 g/ml in hexafluoro isopropanol (HFIP, Aladdin) with 0.01% 2-hydroxy-4′-(2-hydroxyethoxy)−2-methylpropiophenone photoinitiator (Irgacure 2959, Aladdin). The electrospinning process was successfully conducted using a conventional roller electrospinning apparatus, which included a high-voltage power source, syringe pump, and grounded roller enclosed within an environmental chamber. Electrospinning was conducted with a flow rate of 0.5−1 h, voltage set at 12 kV, and a gap distance of 10 cm. The fibers are received at rotating roller speeds of 800 r.p.m. Samples were crosslinked for 5 min under ultraviolet light (100 mW cm−2) at 365 nm to stabilize the connection between the fiber and the receiving mold. The samples were then exposed to ultraviolet light for 5 min in an Irgacure2959 ethanol solution (150 mg/ml) to achieve crosslinking. The fibers were collected on polylactic acid (PLA; Tough PLA, Ultimaker, Nederlands) 3D printed square array (side length 1 mm) functionalized with methacrylate to enhance fiber adhesion. Briefly, PLA square arrays are printed using an extruder 3D printer (Ultimaker S5, Ultimaker, Netherlands). PLA mold was immersed in 60 mg/mL hexamethylene diamine/isopropyl alcohol solution at 40 °C for 2 h to produce PLA-NH2 mold. After that, the PLA-NH2 mold was placed in a 10% methacrylic anhydride/ethanol solution at 50 °C for 2 h to obtain a methacrylated PLA mold.
The Young’s modulus of fiber network was measured by a bio-nanoindenter (Optics 11, Piuma Chiaro, Netherlands). The probe with stiffness of 0.025 N/m (Young’s modulus range: 0.1–100 Kpa) was carefully selected, featuring a tip radius of the probe measuring 12.5 μm and applying a pressure depth of 10,000 nm. The obtained data were fitted with the Hertzian contact model and the degree of fitting was 80%.
Fabrication of GelMA hydrogels
The circular glass slides of 9 mm diameter treated with 3-(Trimethoxysilyl) propyl methacrylate (S111153, Aladdin) were sterilized: immersed in 75% alcohol for 30 min, soaked in penicillin-streptomycin solution for 2 h, and then washed three times with sterile PBS for 5 min each time. The modified glass slides were dried in an ultra-clean bench. The 3.5% (w/w) GelMA solution was prepared using GelMA with a grafting rate of 90% and sterilized through a 0.22 μm filter. The 0.25% (w/v) photoinitiator lithium phenyl-2,4,6-trimethylbenzoylphosphinate (LAP) was mixed with the GelMA solution. Young or senescent BMSCs were respectively resuspended in the GelMA solution at a concentration of 1.5 × 105 cells/ml to prepare the final hydrogel precursor. Next, 15 μl hydrogel precursor was evenly spread on the modified glass slides and crosslinked for 30 s under ultraviolet light (100 mW/cm) at 405 nm. After crosslinking, the cell-laden GelMA hydrogels were placed in complete culture medium and incubated in a 37 °C incubator with 5% CO2. After cells extended in the hydrogels (normally 2 days), the young cells were treated with 20 μmol/ml blebbistatin for 24 h, while the senescent BMSCs were treated with 0.1 nmol/ml calyculin A for 24 h.
ATAC-seq methodology for chromatin accessibility analysis
Cell Sample Preparation
Bone marrow aspirates were obtained from patients aged 60–80 years (old group) and 18–30 years (young group). Bone marrow-derived mesenchymal stem cells (BMSCs) were cultured by plating 1 mL of bone marrow into T-75 flasks containing growth medium supplemented with 10% fetal bovine serum (Gibco). BMSCs at passages 1–2 were used for all downstream experiments.
To enhance cellular traction forces, senescent BMSCs were seeded onto a stretching device (Cell&Force, Cell Tank, Hangzhou Surface&Force Technology Co., Ltd) and subjected to cyclic mechanical stimulation at 5% strain and 0.02 Hz for 2 h. Conversely, to attenuate traction forces, young BMSCs were cultured on soft hydrogels with a shear modulus of 1.5 kPa for 24 h.
ATAC-seq library preparation and sequencing
ATAC-seq libraries were generated using the Hyperactive ATAC-Seq Library Prep Kit for Illumina (Vazyme, TD711) in strict adherence to the manufacturer’s protocol. Importantly, intact whole cells were utilized rather than isolated nuclei to preserve the physiological relevance of cellular traction forces, as nuclear isolation disrupts these forces and may alter the chromatin landscape. This approach was critical for accurately examining the effects of mechanical forces on chromatin accessibility.
For each sample, 50,000 cells were used. Cells were washed twice with ice-cold 1× TW buffer (500 × g, 4 °C), resuspended in precooled lysis buffer, and incubated on ice for 5 min. After centrifugation at 500 × g for 5 min, the supernatant was removed, and the cell pellet was resuspended in a precooled Tn5 transposome/transposition reaction mix. The reaction mix was gently vortexed to ensure uniform distribution and incubated at 37 °C for 30 min to facilitate tag mentation. The reaction was terminated by adding a stop buffer, and the fragmented DNA was purified using VAHTS DNA Clean Beads, including two washes with 80% ethanol. The DNA was finally eluted in 26 μL of nuclease-free water.
Library amplification was performed under the following conditions: initial denaturation at 72 °C for 3 min and 95 °C for 3 min, followed by 14 PCR cycles (98 °C for 10 s, 60 °C for 5 s, 72 °C for 1 min), with a final extension at 72 °C for 1 min and a hold at 12 °C. The amplified libraries were further purified with VAHTS DNA Clean Beads, washed twice with 80% ethanol, and eluted in 22 μL of nuclease-free water. Sequencing was conducted on an Illumina NovaSeq X (150-bp paired-end reads), generating approximately 8 Gb of data per library. Read quality was evaluated using FastQC (v0.11.9), and sequencing services were provided by Berry Genomics Co., Ltd (China).
ATAC-seq analysis
ATAC-seq analysis was performed by Touty.io (Shanghai, China). Raw sequencing reads were aligned to the human genome reference sequence (GRCh38) using the BWA-MEM algorithm. To ensure data quality, duplicate reads were removed and reads with a MAPQ score below 30 were excluded. Additionally, genomic regions listed in the ENCODE blacklisted regions were omitted from the analysis.
Peak calling was performed using Genrich (v0.6.1) with the following parameters: PCR duplicates were removed (-r), signals from mitochondrial and Y chromosomes were excluded (-e MT, Y), and ATAC-specific settings were applied (-j). Quantification of identified peaks was performed using BAMscale (v1.0), and the resulting data were normalized to TPM (transcripts per million). BigWig tracks were generated using deepTools (v3.5.3) with a bin size of 10 bp and normalized to TPM to facilitate visualization of ATAC-seq signals in regions of interest.
Differential peak analysis was conducted in R (v4.1.3) using the DESeq2 package (v1.38.0). Statistical significance was determined using a threshold of |log2FC| > 0.5 and p-value < 0.05. Pairwise comparisons were denoted as “A-vs-B,” where “A” represents the experimental group and “B” represents the control group.
DNA protection experiments
DNA sample preparation
T4-DNA, 165.65 kbp was purchased from Nippon Gene, Tokyo, and used without further purification. YOYO-1 fluorescence staining dye was purchased from Invitrogen, Carlsbad, CA. T4-DNA was stained with YOYO-1 with an intercalation ratio of 100 base-pairs per dye molecule. Samples were prepared by dialysing solutions of T4-DNA against 10 mM Tris/HCl (1xT) buffer in micro-dialysers. The Tris is adjusted with HCl to pH 8.5, i.e., 2.9 mM TrisCl and 7.1 mM Tris. The ionic strength of the buffer was calculated with the Davies equation for estimating the activity coefficients of the ions and a dissociation constant pK = 8.08 for Tris. Protamine chloride from salmon was purchased from Sigma-Aldrich and dissolved in the Tris/HCl buffer. For the pre-equilibrated samples, solutions of protamine and DNA were subsequently mixed in equal volumes and incubated for 24 h at 277 K. The final DNA concentration is 3 mg/L. The buffer used for unpacking of DNA inside the channels was of the same composition, but with 100 mM of added NaCl. No anti-photobleaching agent was used, since it might interfere with protein-DNA binding.
Fluorescence and dark-Deld imaging: Bare and YOYO-1-stained DNA molecules dispersed in the relevant buffer were imaged in millimeter-sized droplets from bulk solution. For experiments conducted using FM, at least 10-run series, each of which is 5–10 s in length at 100 fps, was recorded. To prevent photocleavage, the imaging time is determined corresponding to experimental conditions. In DM, ~10 run series of 30 s length were recorded for each experimental condition. Each data point in this study was obtained by tracking and analyzing more than 200 single DNA molecules. The fluorescence of the stained DNA molecules was visualized with an Olympus IX71 inverted fluorescence microscope equipped with a 100 W mercury lamp, a UV filter set, and a 100-oil immersion objective. The exposure time of the YOYO-1 fluorescence was controlled using a UV light shutter. Images were collected with a charge-coupled device camera (Olympus DP 70). For dark-field imaging, DNA molecules were imaged with a Nikon Eclipse 50i microscope with a CytoViva illuminator and a 100× oil immersion objective. Images were collected with a CCD camera (JVC TK-C921EG) connected via an analog-to-digital video converter (ADVC55, Canopus) to a computer.
RNA extraction and real-time PCR
Total RNA was extracted and purified using the FastPure Complex Tissue/Cell Total RNA Isolation Kit (RC113-01, Vazyme) according to the manufacturer’s protocol. Reverse transcription to complementary DNA (cDNA) was performed with the HiScript III RT SuperMix for qPCR (R323-01, Vazyme). Quantitative real-time PCR (qPCR) was conducted using Taq Pro Universal SYBR qPCR Master Mix (Q712-02, Vazyme) on a CFX96 Real-Time System (Bio-Rad). Gene expression levels were normalized to GAPDH expression. Primer sequences used in this study are listed in Supplementary Table 2.
Statistical analysis
All data are presented as mean ± SD. Statistical analyses were performed using Prism 10.2.3 (GraphPad Software). Two-tailed Student’s t-tests were used for comparisons between two groups, and one-way ANOVA followed by appropriate post hoc tests was applied for comparisons involving more than two groups. P value < 0.05 was considered statistically significant. Exact P values and sample sizes (n) are provided in the corresponding figure and figure legends.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The ATAC-seq datasets generated in this study have been deposited in the NCBI Gene Expression Omnibus under accession number GSE287556. All other data supporting the findings of this study, including source data for all figures and supplementary information, are provided with this paper. Source data are provided with this paper.
References
Marie, P. J. & Kassem, M. Extrinsic mechanisms involved in age-related defective bone formation. J. Clin. Endocr. Metab 96, 600–609 (2011).
Kassem, M. & Marie, P. J. Senescence-associated intrinsic mechanisms of osteoblast dysfunctions. Aging Cell 10, 191–197 (2011).
Langdahl, B., Ferrari, S. & Dempster, D. W. Bone modeling and remodeling: potential as therapeutic targets for the treatment of osteoporosis. Ther. Adv. Musculoskelet. Dis. 8, 225–235 (2016).
Saidak, Z. & Marie, P. J. Strontium signaling: molecular mechanisms and therapeutic implications in osteoporosis. Pharmacol. Ther. 136, 216–226 (2012).
Burr, D. B. & Gallant, M. A. Bone remodelling in osteoarthritis. Nat. Rev. Rheumatol. 8, 665–673 (2012).
Cai, Y. et al. Decoding aging-dependent regenerative decline across tissues at single-cell resolution. Cell Stem Cell 30, 1674–1691.e1678 (2023).
Xiao, Y. et al. Splicing factor YBX1 regulates bone marrow stromal cell fate during aging. Embo J. 42, e111762 (2023).
Zhang, X. et al. Harnessing matrix stiffness to engineer a bone marrow niche for hematopoietic stem cell rejuvenation. Cell Stem Cell 30, 378–395.e378 (2023).
Calcinotto, A. et al. Cellular senescence: aging, cancer, and injury. Physiol. Rev. 99, 1047–1078 (2019).
Di Micco, R., Krizhanovsky, V., Baker, D. & d’Adda di Fagagna, F. Cellular senescence in ageing: from mechanisms to therapeutic opportunities. Nat. Rev. Mol. Cell Biol. 22, 75–95 (2021).
van Deursen, J. M. The role of senescent cells in ageing. Nature 509, 439–446 (2014).
Sahu, S. K. et al. Targeted partial reprogramming of age-associated cell states improves markers of health in mouse models of aging. Sci. Transl. Med. 16, eadg1777 (2024).
Yang, J. H. et al. Chemically induced reprogramming to reverse cellular aging. Aging 15, 5966–5989 (2023).
Pouikli, A. et al. Chromatin remodeling due to degradation of citrate carrier impairs osteogenesis of aged mesenchymal stem cells. Nat. Aging 1, 810–825 (2021).
Casella, G. et al. Transcriptome signature of cellular senescence. Nucleic Acids Res. 47, 7294–7305 (2019).
Lu, Y. R., Tian, X. & Sinclair, D. A. The information theory of aging. Nat. Aging 3, 1486–1499 (2023).
Covarrubias, A. J., Perrone, R., Grozio, A. & Verdin, E. NAD+ metabolism and its roles in cellular processes during ageing. Nat. Rev. Mol. Cell Biol. 22, 119–141 (2021).
Katsyuba, E., Romani, M., Hofer, D. & Auwerx, J. NAD(+) homeostasis in health and disease. Nat. Metab. 2, 9–31 (2020).
Ye, X. et al. Alpha-ketoglutarate ameliorates age-related and surgery induced temporomandibular joint osteoarthritis via regulating IKK/NF-κB signaling. Aging Cell 23, e14269 (2024).
Wang, K. et al. Epigenetic regulation of aging: implications for interventions of aging and diseases. Signal Transduct. Target Ther. 7, 374 (2022).
Zhang, D. et al. Spatial epigenome-transcriptome co-profiling of mammalian tissues. Nature 616, 113–122 (2023).
Liu, X. et al. Nanoscale distribution of bioactive ligands on biomaterials regulates cell mechanosensing through translocation of actin into the nucleus. Proc. Natl. Acad. Sci. USA 122, e2501264122 (2025).
Sun, Q., Wei, Q. & Zhao, C. How do the cells sense and respond to the microenvironment mechanics? Chin. Sci. Bull. 66, 2303–2311 (2021).
Pegoraro, G. et al. Ageing-related chromatin defects through loss of the NURD complex. Nat. Cell Biol. 11, 1261–1267 (2009).
Parry, A. J. et al. NOTCH-mediated non-cell autonomous regulation of chromatin structure during senescence. Nat. Commun. 9, 1840 (2018).
Niedernhofer, L. J. et al. Nuclear genomic instability and aging. Annu. Rev. Biochem. 87, 295–322 (2018).
Vining, K. H. & Mooney, D. J. Mechanical forces direct stem cell behaviour in development and regeneration. Nat. Rev. Mol. Cell Biol. 18, 728–742 (2017).
Halder, G., Dupont, S. & Piccolo, S. Transduction of mechanical and cytoskeletal cues by YAP and TAZ. Nat. Rev. Mol. Cell Biol. 13, 591–600 (2012).
Wen, J. H. et al. Interplay of matrix stiffness and protein tethering in stem cell differentiation. Nat. Mater. 13, 979–987 (2014).
Wolf, K. et al. Physical limits of cell migration: control by ECM space and nuclear deformation and tuning by proteolysis and traction force. J. Cell Biol. 201, 1069–1084 (2013).
Iskratsch, T., Wolfenson, H. & Sheetz, M. P. Appreciating force and shape—the rise of mechanotransduction in cell biology. Nat. Rev. Mol. Cell Biol. 15, 825–833 (2014).
Martyniak, K. et al. Do polyunsaturated fatty acids protect against bone loss in our aging and osteoporotic population? Bone 143, 115736 (2021).
Nikopoulou, C. et al. Spatial and single-cell profiling of the metabolome, transcriptome and epigenome of the aging mouse liver. Nat. Aging 3, 1430–1445 (2023).
Cai, Y. et al. Elimination of senescent cells by β-galactosidase-targeted prodrug attenuates inflammation and restores physical function in aged mice. Cell Res. 30, 574–589 (2020).
Guo, J., Wang, Y., Sachs, F. & Meng, F. Actin stress in cell reprogramming. Proc. Natl. Acad. Sci. USA 111, E5252–E5261 (2014).
Boudaoud, A. et al. FibrilTool, an ImageJ plug-in to quantify fibrillar structures in raw microscopy images. Nat. Protoc. 9, 457–463 (2014).
Hou, Y. et al. Surface roughness and substrate stiffness synergize to drive cellular mechanoresponse. Nano Lett 20, 748–757 (2020).
Jansen, L. E., Birch, N. P., Schiffman, J. D., Crosby, A. J. & Peyton, S. R. Mechanics of intact bone marrow. J. Mech. Behav. Biomed. Mater. 50, 299–307 (2015).
Pan, X. et al. Integrin subtypes and lamellipodia mediate spatial sensing of RGD ligands during cell adhesion. Langmuir 40, 24882–24891 (2024).
Geng, J. et al. Microtubule assists actomyosin to regulate cell nuclear mechanics and chromatin accessibility. Research 6, 0054 (2023).
Hamadi, A. et al. Regulation of focal adhesion dynamics and disassembly by phosphorylation of FAK at tyrosine 397. J. Cell Sci 118, 4415–4425 (2005).
Dupont, S. et al. Role of YAP/TAZ in mechanotransduction. Nature 474, 179–183 (2011).
Ma, S. et al. Spatial transcriptomic landscape unveils immunoglobin-associated senescence as a hallmark of aging. Cell 187, 7025–7044.e7034 (2024).
Church, M. C., Price, A., Li, H. & Workman, J. L. The Swi-Snf chromatin remodeling complex mediates gene repression through metabolic control. Nucleic Acids Res. 51, 10278–10291 (2023).
Tajik, A. et al. Transcription upregulation via force-induced direct stretching of chromatin. Nat. Mater. 15, 1287–1296 (2016).
Nava, M. M. et al. Heterochromatin-driven nuclear softening protects the genome against mechanical stress-induced damage. Cell 181, 800–817.e822 (2020).
Koester, J. et al. Niche stiffening compromises hair follicle stem cell potential during ageing by reducing bivalent promoter accessibility. Nat. Cell Biol. 23, 771–781 (2021).
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Martins, R., Lithgow, G. J. & Link, W. Long live FOXO: unraveling the role of FOXO proteins in aging and longevity. Aging Cell 15, 196–207 (2016).
Sladitschek-Martens, H. L. et al. YAP/TAZ activity in stromal cells prevents ageing by controlling cGAS-STING. Nature 607, 790–798 (2022).
Gong, H. et al. Whole body vibration with rest days could improve bone quality of distal femoral metaphysis by regulating trabecular arrangement. Sci. China Life Sci. 62, 95–103 (2019).
Sun, J. et al. LAP2β transmits force to upregulate genes via chromatin domain stretching but not compression. Acta Biomater. 163, 326–338 (2023).
Le, H. Q. et al. Mechanical regulation of transcription controls Polycomb-mediated gene silencing during lineage commitment. Nat. Cell Biol. 18, 864–875 (2016).
Geng, L. et al. Systematic profiling reveals betaine as an exercise mimetic for geroprotection. Cell 188, 5403–5425.e5433 (2025).
Zhu, Q. et al. Procr+ chondroprogenitors sense mechanical stimuli to govern articular cartilage maintenance and regeneration. Cell 188, 5194–5211.e5116 (2025).
Zhang, M. et al. Controllable ligand spacing stimulates cellular mechanotransduction and promotes stem cell osteogenic differentiation on soft hydrogels. Biomaterials 268, 120543 (2021).
Benoit, B. et al. Fibroblast growth factor 19 regulates skeletal muscle mass and ameliorates muscle wasting in mice. Nat. Med. 23, 990–996 (2017).
Chen, K. et al. Exercise training improves motor skill learning via selective activation of mTOR. Sci. Adv. 5, eaaw1888 (2019).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (T2222020 to Q.W.; 32501155 to X.L.) and the Natural Science Foundation of Shandong Province (ZR2024QC308 to X.L.). We thank Ce Zhang for assistance with DNA damage assessment on isolated DNA, and He Gong and Jiazi Gao for their support in the development of the mouse vibration apparatus.
Author information
Authors and Affiliations
Contributions
X.L. designed the study, performed experiments, conducted data analysis, and wrote the manuscript. Y.Y. was responsible for human bone marrow collection. Z.L. contributed to experimental design and discussion of the results. L.L. initiated and co-supervised the project. Q.W. conceived the research, supervised the study, and revised the manuscript. All authors reviewed and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Cecilia Engdahl and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Liu, X., Ye, Y., Li, Z. et al. Mechanical rejuvenation of senescent stem cells and aged bone via chromatin remodeling. Nat Commun 17, 1684 (2026). https://doi.org/10.1038/s41467-026-68387-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-68387-3