Abstract
Genomics, metabolomics, and proteomics offer complementary insights into cardiovascular disease (CVD) risk. Leveraging UK Biobank data, we introduce the CardiOmicScore, a multitask deep learning framework, to learn disease-specific proteomic (ProScore) and metabolomic (MetScore) risk scores for the six most common CVDs by profiling 2920 proteins and 168 metabolites. Experiments demonstrate that ProScore and MetScore are strong sole CVD risk predictors (C-index range: 0.69–0.82 for ProScore and 0.64–0.74 for MetScore), and can significantly enhance risk prediction across CVDs up to 15 years prior to disease onset when combined with clinical data, increasing the C-index by 0.005–0.102. These findings suggest that incorporating multiomics profiling into clinical practice can improve personalized risk assessments at early stages. CardiOmicScore also identifies important CVD-related proteins and metabolites, which represent promising data-driven pathways, calling for further external validation, to develop novel biomarkers and targeted therapies, facilitating precision medicine for primary prevention of CVDs.
Similar content being viewed by others
Introduction
Cardiovascular diseases (CVDs) are the leading cause of mortality worldwide, claiming an estimated 19.8 million lives in 2022 alone1,2. This staggering figure is projected to rise further with the aging global population, underscoring the urgent need for innovative strategies to identify individuals at high risk and intervene early3,4,5.
Accurate CVD risk stratification is challenging since cardiovascular risk is determined by the complex interplay of genetic, environmental, and lifestyle factors6. Several approaches, such as the ASCVD and SCORE2, though widely used7,8,9, rely heavily on conventional clinical risk factors and thus exhibit limited discrimination and calibration10. Recent endeavors to incorporate polygenic risk scores (PRSs) have yielded only modest improvements and fail to capture the pathophysiological processes underlying disease progression11,12,13,14. Blood-based biomarkers offer a promising, non-invasive, and cost-effective alternative, but their full potential remains untapped11.
We are now entering the era of high-throughput metabolomics and proteomics. These cutting-edge technologies further enable the measurement of a wide range of proteins and metabolites in the blood, offering a comprehensive snapshot of the molecular mechanisms driving CVDs11,15. While recent studies have suggested the potential of metabolomics16,17,18,19 and proteomics20,21,22,23,24,25,26,27,28,29,30,31,32,33 data for risk prediction, a holistic integration of genetics, omics-based biomarkers, and traditional clinical risk factors has yet to be fully realized due to the scarcity of longitudinal multiomics cohort data and the limitations in scalable computational frameworks to handle heterogeneous data modalities13,14,16,17,20,21,22,23,24,25,26,27,28,29,30.
Metabolomics and proteomics offer complementary insights into CVD risk, with metabolomics providing a broad profile of metabolites largely involved in lipoprotein metabolism34, while proteomics focuses on a detailed set of proteins related to coagulation, inflammation, oxidative stress, and vascular remodeling35,36. Given the demonstrated effectiveness of multiomics approaches in improving risk identification for diseases such as diabetes and osteoarthritis37,38, integrating these multiomics data with genetic risk and clinical information may further enhance the predictive performance for cardiovascular risk. Moreover, existing prediction models often directly input high-dimensional omics data into linear regression to predict disease risk, which are constrained by assumptions of non-correlation between predictors and linear exposure-outcome associations16,18,19,20,23,25,27,28,29,30,31,32. In fact, high-order correlations exist within and across different omics data, highlighting the need for artificial intelligence (AI) algorithms to capture these complex, nonlinear relationships and interactions, and to generate informative low-dimensional representations for downstream disease risk prediction. Additionally, previous studies incorporating multiomics data have focused on a narrow set of CVDs, such as myocardial infarction and atrial fibrillation18,19,31,32,33, restricting the understanding of the shared pathways at the metabolomic and proteomic levels across a broad spectrum of cardiovascular outcomes.
Our study aims to bridge this gap by proposing a groundbreaking AI-based CardiOmicScore framework. Leveraging the unparalleled depth of data in the UK Biobank (UKB), we introduced two advanced deep learning models, MetNet and ProNet, designed to simultaneously predict the risk of six CVDs using 168 metabolites and 2920 proteins, respectively (Fig. 1). Unlike traditional models, both MetNet and ProNet utilize a multitask architecture to learn high-dimensional representations from metabolomics and proteomics data, generating disease-specific risk scores, namely MetScore and ProScore, for a comprehensive range of CVDs. These CVDs include coronary artery disease (CAD), stroke, heart failure (HF), atrial fibrillation (AF), peripheral artery disease (PAD), and venous thromboembolism (VTE)—conditions with the highest global burden1. We further developed Cox proportional hazards (CPH) models on various combinations of PRS, MetScore, ProScore, and clinical risk factors to predict each CVD onset. We evaluated the predictive performance, as well as the potential clinical utility of all the models to investigate the additive value of multiomics profiling in CVD risk prediction.
a We extracted data from the UK Biobank, including clinical, genomic, metabolomic, and proteomic predictors, as well as six cardiovascular diseases (CVDs) defined by self-reported diagnoses, hospital episode statistics, and death records. b A two-stage data partition strategy was applied: the Multiomics cohort, which remained untouched throughout model training and was used as an independent validation set to evaluate the added value of integrating multiple omics data types, and separate training, validation, and geographic testing sets within the Metabolomics-only and Proteomics-only cohorts for model development. Our CardiOmicScore framework utilized two multitask deep neural networks to predict the risk of six CVDs from 168 metabolites and 2920 proteins separately, generating MetScore and ProScore for each CVD. We trained Cox proportional hazards (CPH) models on various combinations of polygenic risk score (PRS), MetScore, ProScore, and three predefined clinical predictor sets (i.e., AgeSex, Clin, and PANEL). c Model performance was first evaluated for MetScore and ProScore in their respective geographic testing sets, and subsequently for all feature combinations in the Multiomics cohort. Performance metrics included Harrell’s C-index, calibration plots, and net benefit curves, with confidence intervals estimated using 1000 bootstrap resamples. Icons were designed by Freepik from Flaticon (www.flaticon.com).
Results
Study population
A two-stage, individual-level data partitioning strategy was employed to rigorously evaluate our model’s performance and generalizability (see sub-section “Data partition and imputation” and Supplementary Fig. 1). First, we excluded participants who had neither metabolomics nor proteomics data available. Then, we divided the remaining participants into the Metabolomics-only (N = 220,859; those with metabolomics but not proteomics data), Proteomics-only (N = 19,086; those with proteomics but not metabolomics data), and Multiomics (N = 24,287; those with genomics, metabolomics, and proteomics data) cohorts based on omics availability. The Metabolomics-only and Proteomics-only cohorts were used within our CardiOmicScore framework to develop two deep learning models, MetNet and ProNet, respectively. The Multiomics cohort was reserved as an untouched validation set to assess the added predictive value of integrating different omics data.
Following best practices in recent UKB studies39,40,41, each of these development cohorts was further split into training/validation sets (England and Wales) and a geographic testing set (Scotland). Specifically, the Metabolomics-only cohort included 187,272 participants in the training set, 20,808 in the validation set, and 12,779 in the geographic testing set, while the Proteomics-only cohort comprised 15,579, 1732, and 1775 participants in the training, validation, and geographic testing sets, respectively. Baseline characteristics were comparable between training and validation sets in both cohorts, whereas significant differences were observed between training and geographic testing sets (Supplementary Data 2 and 3), thus allowing a robust assessment of the models’ regional generalizability.
Baseline characteristics and incident cases during follow-up were broadly consistent across all study datasets. The median age of participants at baseline was between 56.0 and 58.0 years. The proportion of males ranged from 42.8% to 45.8%. The prevalence of baseline medication use was comparable, with 15.8–18.1% of participants receiving lipid-lowering medication and 10.5–11.2% on antihypertensive medication. The distribution of outcomes was also similar, with 83.2–85.7% of participants remaining free of any incident CVD, 10.6–11.8% developing one CVD, and 3.7–5.0% developing multiple CVDs. Detailed baseline characteristics and follow-up information for each cohort are provided in Supplementary Data 1–3 and Supplementary Fig. 2.
Omics information is the key driver of cardiovascular risk stratification
We derived the MetScore and ProScore for each CVD using the trained MetNet and ProNet with 168 metabolic and 2920 proteomic biomarkers as input. We observed strong correlations among the MetScores for the six different CVDs (Spearman correlation coefficients = 0.78–0.96) and among the ProScores (Spearman correlation coefficients = 0.87–0.94) (Supplementary Fig. 3). Additionally, we constructed PRSs for all CVDs.
Participants were stratified into three risk groups (low, medium, and high) based on the tertiles of the PRS, MetScore, and ProScore. Kaplan-Meier survival curves showed significant differences across risk groups defined by MetScore and ProScore for all CVDs (all log-rank P < 0.001) (Fig. 2a and b). In contrast, PRS risk groups exhibited significant differences in survival curves only for CAD, stroke, AF, and VTE (all log-rank P < 0.001) (Supplementary Fig. 5a). Similar patterns were observed in the correlation between observed event rates and the percentiles of the three omics scores (Supplementary Fig. 4).
Survival probabilities over time for six CVDs, stratified by the tertiles of MetScore (a) and ProScore (b). Solid lines represent survival probabilities (measure of center), estimated using the Kaplan–Meier method, and shaded areas represent 95% exponential Greenwood confidence intervals (error bands). c Associations of CVD risk with MetScore and ProScore. Each score was analyzed both as a categorical variable (tertile groups: low, medium, high; low as reference) and as a continuous variable (per 1-standard deviation [SD] increase after standardization). Data are presented as hazard ratios (measure of center) estimated by Cox proportional hazards models with error bars representing 95% confidence intervals. All models were adjusted for demographic information, healthy lifestyles, family disease history, disease and medication history, physical measurements, and blood count. CAD coronary artery disease, HF heart failure, AF atrial fibrillation, PAD peripheral artery disease, VTE venous thromboembolism. Source data are provided as a Source Data file.
After adjusting for demographic information, healthy lifestyles, family history of disease, disease and medication history, physical measurements, and blood count, both categorical and continuous analyses of ProScore, MetScore, and PRS showed significant associations with incident CVDs (Fig. 2c and Supplementary Fig. 5b). Participants in the top tertile of ProScore had over a 1.68-fold higher risk of all CVDs ranging from stroke (hazard ratio [HR] = 1.68, 95% confidence interval [CI] = 1.40–2.01) to CAD (HR = 2.18, 95% CI = 1.88–2.52) compared with those in the bottom tertile. The top tertile group of MetScore was associated with an increased risk of all CVDs, with the highest HRs observed in HF (HR = 1.64, 95% CI = 1.39–1.92) and CAD (HR = 1.70, 95% CI = 1.49–1.94). Participants in the high PRS risk group had a higher risk of stroke (HR = 1.25, 95% CI = 1.08–1.46), VTE (HR = 1.54, 95% CI = 1.33–1.78), CAD (HR = 1.65, 95% CI = 1.49–1.83), and AF (HR = 1.89, 95% CI = 1.69–2.11) than those in the low PRS risk group. When modeled as continuous variables (per 1-standard deviation [SD] increase), ProScore, MetScore, and PRS remained significantly associated with CVD risk, with HRs per SD increase ranging from 1.07 (95% CI = 1.01–1.13) for PRS of stroke to 1.57 (95% CI = 1.49–1.66) for ProScore of AF.
Advancing cardiovascular risk prediction with the power of omics information
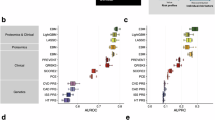
We investigated the predictive performance of three omics scores, two published algorithms for CVD risk assessment (ASCVD and SCORE2)7,8,9, and three clinical predictor sets (AgeSex, Clin, and PANEL). The Clin set contained 25 questionnaire-collected variables and physical measurements relevant to cardiovascular risk, while the PANEL set built upon the Clin set by additionally incorporating blood count information17,26. ASCVD and SCORE2 were calculated using the published regression equations (see sub-section “Clinical predictors and cardiovascular risk scores”). Omics scores were treated as continuous variables, while clinical predictors were included as either continuous or categorical variables. Discriminative performance of all models, evaluated by Harrell’s C-index, and differences in the C-index across models are presented in Fig. 3. Among the three omics scores, ProScore showed the highest performance for all CVDs with the C-index ranging from 0.69 (95% CI = 0.67–0.71) for VTE to 0.82 (95% CI = 0.80–0.84) for PAD (Fig. 3a). The C-index of MetScore spanned from 0.64 (95% CI = 0.61–0.66) for VTE to 0.74 (95% CI = 0.71–0.76) for PAD. Importantly, the strong performance of ProScore and MetScore was replicated in their respective geographic testing sets, underscoring their generalizability across distinct UK populations (Supplementary Fig. 6). In contrast, PRS provided the most limited predictive capacity (C-index range = 0.52–0.60). Finally, the discriminative performance increased with more clinical predictors included in the CPH models for all diseases (Fig. 3a).
a Discriminative performance of models trained on various predictor sets. Vertical dashed lines indicate the performance of the three clinical predictor sets. Discriminative performance was evaluated by Harrell’s C-index. b Differences in discriminative performance (ΔC-index) by adding omics information to clinical predictor sets. In both panels, data are presented as point estimates (measures of center) with error bars representing 95% confidence intervals estimated from 1000 bootstrap resamples. PRS polygenic risk score, CAD coronary artery disease, HF heart failure, AF atrial fibrillation, PAD peripheral artery disease, VTE venous thromboembolism. Source data are provided as a Source Data file.
Compared with baseline clinical-based models (i.e., AgeSex, Clin, and PANEL), these models had significantly greater performance (i.e., 95% CIs of delta C-index not including zero) in all CVDs after adding omics scores (Fig. 3b). For the AgeSex set, a significant increase in the C-index was observed with a range of 0.005 (95% CI = 0.001–0.013, +PRS for stroke) to 0.102 (95% CI = 0.081–0.124, +PRS +ProScore for PAD). The addition of ProScore achieved the greatest improvement in model performance (0.040 for AF to 0.101 for PAD), followed by MetScore (0.009 for AF to 0.051 for PAD). Although the improvements were more modest than those observed with the AgeSex-based model, adding omics scores to the Clin- and PANEL-based models still resulted in significant performance improvements, with the delta C-index ranging from 0.003 (95% CI = 0.001–0.006) to 0.049 (95% CI = 0.036–0.061) for the Clin set and 0.003 (95% CI = 0.001–0.005) to 0.049 (95% CI = 0.035–0.061) for the PANEL set. Even for the PANEL set that already contained predictive information, the inclusion of omics signatures resulted in better discriminative performance for all CVDs.
We then used models combining the PANEL set with one omics score (i.e., +PRS, +MetScore, or +ProScore) as baselines to demonstrate the added value of incorporating additional omics features. Model performance improved for all CVDs when additional omics features were integrated into models already containing PRS (i.e., +PRS) or MetScore (i.e., +MetScore) (Supplementary Fig. 7). Furthermore, when using +ProScore as the baseline, adding PRS (either alone or in combination with MetScore) improved prediction for CAD, stroke, AF, and VTE, while incorporating MetScore specifically improved performance for CAD and stroke.
PANEL-based models achieve optimal calibration and net benefit
We assessed the potential clinical utility of our models by two indicators: calibration and net benefit. Calibration measures the extent to which predicted risks align with observed risks42, while net benefit quantifies the trade-off between true positives and false positives at various decision thresholds43. To provide a summary measure of overall clinical utility, we further calculated the area under the net benefit curve (A-NBC)44. Models based on the PANEL set were well calibrated for all outcomes as their calibration curves closely followed the diagonal line (Fig. 4a). The consistency between the observed risks and the predicted risks was greater in the PANEL-based models than in the AgeSex- and Clin-based models (Fig. 4a, Supplementary Fig. 8a, and Supplementary Fig. 9a). The inclusion of omics information, especially ProScore, improved standardized net benefits across a range of decision thresholds compared with clinical-based models. Specifically, the maximum improvement in A-NBC, achieved by the full multiomics model (i.e., +PRS +MetScore +ProScore), ranged from 35.39 for stroke to as high as 212.36 for AF. The incremental benefits diminished as the number of clinical predictors in the baseline models increased (Fig. 4b, Supplementary Figs. 8b and 9b).
a Calibration curves for Cox proportional hazards models based on the clinical predictor set (PANEL) and its combination with multiomics information. b Net benefit curves for Cox proportional hazards models based on the clinical predictor set (PANEL) and its combination with multiomics information. Horizontal solid gray lines indicate “treat none” and vertical solid gray lines indicate “treat all”. Differences in the area under the net benefit curve (A-NBC) represent improvements in net benefit compared with the PANEL-based models. PRS polygenic risk score, CAD coronary artery disease, HF heart failure, AF atrial fibrillation, PAD peripheral artery disease, VTE venous thromboembolism. Source data are provided as a Source Data file.
AI identifies disease-specific metabolites and proteins
We further utilized the Shapley Additive exPlanations (SHAP) method to examine the relationship of metabolites and proteins with each CVD. To capture global importance, we first calculated the mean absolute SHAP value for each feature and normalized these values within each disease. Figure 5a and b display the top 65 metabolites and 77 proteins ranked by their global importance.
Importance of biomarkers in cardiovascular disease-specific MetScore (a) and ProScore (b). Importance was measured by mean absolute SHAP values. The color represents normalized mean absolute SHAP values, ranging from blue (low) to red (high). For each disease, the 50 most important metabolites and the 25 most important proteins were identified. The union of these features across all diseases yielded a total of 65 metabolites and 77 proteins. SHAP Shapley Additive exPlanations, CAD coronary artery disease, HF heart failure, AF atrial fibrillation, PAD peripheral artery disease, VTE venous thromboembolism. Source data are provided as a Source Data file.
We focused on the top five metabolites and proteins to identify the most impactful biomarkers for each disease. For metabolites, we observed a partial overlap across the six CVDs. The commonly impactful metabolites included creatinine, albumin, glutamine, fatty acids such as linoleic acids (LA) and monounsaturated fatty acids (MUFA), and lipoprotein components such as free cholesterol in intermediate-density lipoprotein (IDL_FC), free cholesterol in very large high-density lipoprotein (XL_HDL_FC), cholesteryl esters in chylomicrons, and extremely large very-low-density lipoprotein (XXL_VLDL_CE). Meanwhile, other biomarkers were important for specific diseases, such as tyrosine and leucine for HF and AF, and glycoprotein acetyls (GlycA) for PAD and VTE. For proteins, the impactful biomarkers were more distinct among CVDs. We found that NT-proBNP and NPPB were important for CAD, HF, and AF, while GDF15 was important for CAD, HF, AF, and VTE. Other key associations included MMP12 for CAD and PAD, CDCP1 for CAD, HF, and VTE, EDA2R for CAD and stroke, NEFL for stroke and PAD, WFDC2 for HF and PAD, and FASLG for stroke and VTE. The most important disease-specific proteins also included KLK4 and CRIP2 for stroke; BCAN and ELN for AF; PLB1 and ENDOU for PAD; and HPGDS and ADGRG2 for VTE.
The SHAP analysis also revealed that while the direction of the top biomarker-disease associations was highly consistent across all six CVDs, the magnitude of their contributions varied. We used CAD as an example to illustrate the associations between plasma biomarkers and disease risks. Among metabolites, higher plasma levels of creatinine, GlycA, MUFA, tyrosine, glutamine, IDL_FC, and XL_HDL_FC were associated with an increased risk of CAD, while lower levels of albumin, LA, leucine, and XXL_VLDL_CE were associated with CAD risk (Fig. 6a). For proteins, NT-proBNP, NPPB, GDF15, MMP12, CDCP1, EDA2R, NEFL, WFDC2, KLK4, CRIP2, and ELN were positively associated with disease risk, while a negative association was observed between CAD risk and elevated levels of FASLG, BCAN, PLB1, ENDOU, HPGDS, and ADGRG2 (Fig. 6b). SHAP beeswarm plots of metabolites and proteins for other diseases are provided in Supplementary Figs. 10 and 11. A summary of the top-ranking metabolites and proteins is presented in Supplementary Data 4.
SHAP beeswarm plots illustrating the associations of metabolites (a) and proteins (b) with cardiovascular disease risk, using coronary artery disease as an example. Each individual is assigned a SHAP value for every protein and metabolite. To simplify the presentation, SHAP values for each biomarker were divided into percentiles, and the mean SHAP value and biomarker level were calculated for each percentile. Each dot represents one percentile. All 168 metabolites and the top-ranked 157 proteins are shown. Dark gray lines indicate zero SHAP value (i.e., no contribution). The farther a dot from this line, the stronger the absolute contribution to the MetScore or ProScore. Deviations toward the center and periphery represent negative and positive contributions. Red and blue colors indicate high and low levels of metabolites and proteins, respectively. SHAP Shapley Additive exPlanations, CAD coronary artery disease. Source data are provided as a Source Data file.
Sensitivity analyses
We evaluated the performance of 25 key individual biomarkers (8 metabolites and 17 proteins) that were selected for either their high global importance in our CardiOmicScore framework (see sub-section “AI identifies disease-specific metabolites and proteins”) or their routine use in clinical practice. Overall, whether used individually or combined with clinical risk factors, these key biomarkers’ discriminative ability was much weaker than that of MetScore or ProScore (Supplementary Fig. 12). Additionally, the incremental benefit of adding omics scores to clinical models was consistent across age groups and sexes (Supplementary Fig. 13), as well as among participants stratified by lipid-lowering and antihypertensive therapy (Supplementary Fig. 14). We also found that the predictive value of the omics scores remained unchanged after excluding incident cases that occurred within the first two years of follow-up (Supplementary Fig. 15) or accounting for the competing risk of death (Supplementary Fig. 16).
Moreover, our CardiOmicScore framework proved superior to conventional machine learning methods for generating omics scores. Specifically, MetScore and ProScore derived from MetNet and ProNet demonstrated generally superior performance across multiple CVDs compared to analogous scores developed using XGBoost, LightGBM, random forest, and logistic regression (Supplementary Fig. 17). Finally, after refitting the MetNet model in the subgroup of participants not taking lipid-lowering medications, the most impactful metabolites included MUFA, glycine, creatinine, albumin, extra-large high-density lipoprotein free cholesterol, glutamine, IDL_FC, tyrosine, LA, and GlycA (Supplementary Fig. 18). Seven of these metabolites overlapped with those highlighted in the main analysis. Furthermore, although the individual predictive performance of some traditional lipid-related biomarkers for CAD improved slightly after excluding participants on lipid-lowering medication, their addition still provided only limited improvement in predictive performance to the PANEL-based model. This finding was consistent in both the full Multiomics cohort and the subgroup excluding baseline users of lipid-lowering medication (Supplementary Figs. 19 and 20).
Discussion
In a large-scale prospective population-based cohort, we developed our CardiOmicScore framework, leveraging multitask deep learning models and multiomics data to derive metabolomic (MetScore) and proteomic (ProScore) risk scores for six CVDs. Our results showed that both ProScore and MetScore improved risk stratification for all CVDs, providing predictive information complementary to traditional clinical predictors. Additionally, the inclusion of omics signatures, particularly ProScore, significantly enhanced CVD risk predictions, showing superior discriminative performance compared to clinical-based models. Importantly, these improvements in discrimination translated to clinical utility across all CVDs.
First, our study demonstrates the clinical benefits of integrating multiomics in CVD risk assessment. Accurate risk stratification is essential for designing tailored interventions to prevent adverse cardiovascular outcomes. Traditional risk assessment methods often focus on predictive factors one disease at a time, which is resource-intensive and potentially limits feasibility in clinical settings17. Omics data offer a novel approach, enabling simultaneous risk stratification for multiple diseases through a single measurement that captures a broad spectrum of molecular predictors17,19,21,26,27,28,29,30. This is particularly relevant for CVDs given their shared metabolomic and proteomic profiles, which reflect common biological pathways and overlapping risk factors16,25. Furthermore, the integration of various omics data types—each capturing distinct disease-associated markers—can provide complementary value and facilitate a more holistic understanding of biological systems45. Specifically, genetic data offer long-term predictive information, while metabolomic and proteomic profiles provide dynamic insights into physiological processes11,15,17,19,26,27,28,29.
By systematically comparing models with different combinations of clinical predictors and omics features, we demonstrated that the integration of multiomics data significantly improved predictive performance across several CVDs, beyond conventional clinical predictors or single omics measurements. The performance of our models was stable across age groups, biological sexes, and participants stratified by lipid-lowering and antihypertensive therapy, suggesting their robustness and broad applicability in diverse subpopulations. This robustness was further confirmed by the findings that the predictive value of the omics scores remained consistent after excluding early incident cases or accounting for the competing risk of death. Our models demonstrated not only strong discriminative performance but also robust calibration. Importantly, they showed clinical net benefits over different decision thresholds. These results further support the clinical utility of multiomics data in CVD risk prediction, which allows for simultaneous risk assessment for common CVDs even up to 15 years before disease diagnosis, and thus informs early and targeted primary prevention strategies11,30.
Second, we find that proteomics contributes most to enhancing CVD risk prediction among the three omics. Consistent with previous studies utilizing proteomics or metabolomics, we demonstrated their high predictive value for multiple CVDs16,17,18,19,20,21,22,23,24,25,26,27,28,29,30,31,32,33. We further compared the predictive performance of the three omics features for CVD risk, finding that ProScore provided the greatest discriminative performance, followed by MetScore. This aligns with findings from a previous study29.
The superior performance of proteomics can be attributed to its dynamic nature, which captures the influences and interactions of genetic and environmental factors, thereby better elucidating biological pathways leading to CVD onset11,15,27,29. In contrast, PRS represents only a static measure for an individual’s lifetime risk of disease and provides no direct therapeutic insights11. Another explanation for the superior performance of proteomics over metabolomics is the larger number of input features in proteomics (2920 proteins vs. 168 metabolites). Although the nuclear magnetic resonance (NMR) panel in the UKB, a targeted metabolomics technique, includes hundreds of metabolites, the majority are lipoproteins and lipid-related markers. This limitation may render the panel insufficient for capturing complex metabolomic profiles associated with CVDs, especially non-lipoprotein metabolites46. In comparison, proteomics may cover broader CVD-related pathological processes, including inflammation, immune regulation, coagulation, and vascular functions24,25,35,36.
Given these findings, our ProScore holds promise as an effective and standardized tool to be integrated into clinical risk assessments to enhance CVD risk prediction. Although the proteomics platform used in this study is relatively expensive and not yet widely accessible, the integration of multiomics into clinical workflows could become increasingly cost-effective over time. As technology advances and costs decrease, the use of omics data and AI models in routine clinical practice could provide a more efficient, precise, and individualized approach to cardiovascular risk assessment. By enabling early identification of patients at high risk of CVD, clinicians may be able to reduce the overall burden of CVD and improve patient outcomes in a cost-effective manner.
Third, evaluating the contributions of important proteins and metabolites to each CVD reveals underlying disease mechanisms and potential therapeutic targets. We confirmed the role of proteins such as NT-proBNP and NPPB, as well as metabolites such as albumin and creatinine, in CVD risk assessment. These biomarkers are well-established in epidemiological studies and are already part of routine clinical care47,48,49,50. However, our analyses revealed that ProScore outperformed any single protein, such as NT-proBNP and NPPB, both as an individual predictor and when added to clinical models. Similarly, individual metabolic biomarkers such as creatinine, albumin, and traditional lipid-related biomarkers (e.g., total lipids or cholesterol) yielded lower predictive performance compared to MetScore. These findings suggest that the MetScore and ProScore may provide a more comprehensive representation of disease-related molecular mechanisms than single biomarkers, and thus highlight the importance of high-throughput metabolomics and proteomics in personalized CVD risk prediction. Moreover, our CardiOmicScore framework outperformed conventional machine learning algorithms, underscoring the unique strength of the multitask deep learning architecture to model complex, non-linear interactions in high-dimensional omics data.
In addition to the established biomarkers, we identified several less-studied circulating metabolites and proteins associated with CVD risk, which were consistent with previous omics-based modeling studies17,22,23,24,25,26,27,28,29,30,31,32,33. We observed a positive association between future CVD and GlycA, a composite NMR-based signal and biomarker of systemic inflammation51. In line with earlier findings, increasing levels of MUFA52 and tyrosine53 were associated with a higher risk of CVDs, while we also confirmed the protective role of LA52. Interestingly, while remnant cholesterol is recognized as a causal risk factor for atherosclerotic cardiovascular disease (ASCVD)54, we found that its components may have inconsistent associations with CVD risk: higher IDL_FC levels were associated with increased risk, whereas higher XXL_VLDL_CE levels were associated with lower risk. These opposing associations warrant future investigation, as they suggest that measuring IDL_FC and XXL_VLDL_CE may further stratify risk beyond traditional LDL-cholesterol levels.
Regarding proteins, our analysis identified a diverse spectrum of protein biomarkers, revealing pathways that are shared across CVDs and highly specific to individual diseases. Several proteins were associated with multiple CVDs, suggesting common underlying pathological processes. For example, the stress- and inflammation-induced cytokine GDF15 showed strong predictive value for CAD, HF, AF, and VTE, acting as a general marker of systemic stress and broad cardiovascular pathology55,56. Similarly, MMP12, an enzyme involved in extracellular matrix degradation and inflammatory responses, was associated with both CAD and PAD57,58. The apoptosis-related ligand FASLG, a key mediator of extrinsic cell death and vascular inflammation, was important for stroke and VTE59. In addition, the neuronal damage marker NEFL, reflecting systemic neuroaxonal injury, was predictive of both stroke and PAD, suggesting a mechanistic link between neural injury and peripheral vascular events60. Beyond these shared pathways, we also found several disease-specific proteins, underscoring the distinct biology of each condition. Key specific associations included such as CRIP2, a key regulator of cellular responses to ischemia, with stroke61, and PLB1, involved in metabolic regulation, with PAD62. Taken together, the identification of biomarkers from such diverse pathways—covering cardiac stress, inflammation, neuronal injury, and matrix remodeling—suggests that our CardiOmicScore framework can capture a multi-system biological signature of cardiovascular risk that extends far beyond traditional risk factors.
These key molecules not only serve as potential targets for new therapeutic interventions to prevent or reverse CVD progression, but also illuminate druggable pathways for repurposing existing medications—such as anti-inflammatory colchicine63. Our findings also suggest that, based on individual omics profiling, clinicians may be able to tailor treatments targeting the underlying biological mechanisms of specific CVDs. For example, patients with high levels of inflammation markers like GlycA may benefit from treatments aimed at reducing systemic inflammation, while those with elevated proteins like MMP12, which is involved in extracellular matrix degradation, could be targeted with therapies focused on preventing the vascular remodeling that leads to atherosclerosis. Notably, our study provides only preliminary evidence for the predictive importance of circulating biomarkers; therefore, it must be clearly stated that these signatures are not clinically generalizable or applicable until they have undergone external validation.
The main strength of our study is the development of the CardiOmicScore framework, which integrates multiomics data and deep learning models to predict multiple CVDs within a large community-based prospective cohort. We implemented a rigorous, two-stage evaluation process, fully utilizing the UKB data to ensure our models’ generalizability by validating the MetScore and ProScore first on a geographic testing set and subsequently on an untouched Multiomics cohort. Additionally, we have deployed a demo of our model on a user-friendly platform (see “Code availability” section), where clinicians can input clinical information and omics measurements to predict individual CVD risk. With further validation in future research, this platform may be embedded into electronic health record systems, increasing its clinical utility and accessibility.
Our study has several limitations. First, the current metabolomics panel focuses on lipid-related markers. The predictive performance of metabolomics techniques that cover more metabolic biomarkers should be explored. Second, the proteomic and metabolomic data were measured at baseline. This static snapshot may not capture the dynamic changes in a person’s molecular profile over time, which could also influence their long-term disease risk. Third, since the majority of the UKB population is of European ancestry and white ethnicity, the generalizability of our findings needs to be validated in ethnically diverse populations to ensure broader applicability. Finally, although we conducted geographic validation to assess model generalizability, our study still lacks an independent external validation.
Taken together, our study demonstrates that integrating genetic, metabolomic, and proteomic data with traditional risk factors significantly improves CVD risk stratification. The findings highlight the potential of high-throughput omics technologies to uncover novel biomarkers and mechanistic pathways associated with CVDs. By leveraging these insights, clinicians can adopt a more personalized approach to risk assessment and treatment planning, ultimately improving patient outcomes. This comprehensive framework paves the way for more precise and individualized cardiovascular care, with the potential to incorporate additional diagnostic modalities in the future.
In future work, our framework has the potential to be expanded into a foundation model capable of incorporating additional important information for CVD risk prediction, such as electrocardiograms and coronary artery imaging. This foundation model could be adapted for a wide range of downstream tasks, including individualized risk assessment and personalized treatment planning. Large language models (LLMs) could play a crucial role in this expansion by enhancing the model’s ability to process and interpret complex, multimodal medical data (e.g., medical records, imaging, and omics data). For example, LLMs could help patients and healthcare professionals by providing real-time insights on patient risk profiles, generating easy-to-understand explanations, and suggesting personalized intervention options based on integrated data. This integration could provide a powerful tool for clinicians, enabling them to make data-driven decisions more effectively and ultimately improve patient outcomes.
Methods
Study population
The UKB is a prospective cohort study that has collected extensive genetic and phenotypic data from ~500,000 participants who were recruited from 22 centers across the UK during 2006–201064. Metabolomics and proteomics profiling were performed in plasma samples collected at baseline from a randomly selected subset of ~280,000 and ~54,000 UKB participants. UKB has approval from the North West Multi-Centre Research Ethics Committee as a Research tissue biobank (REC reference 11/NW/0382). Participants provided written informed consent. This study received approval from the UKB under application number 79146.
Data partition and imputation
To rigorously evaluate model performance and ensure generalizability, we adopted a two-stage individual-level data partition strategy, designed to approximate external validation while fully leveraging UKB data (Supplementary Fig. 1). First, we excluded participants who lacked both metabolomics and proteomics data. The remaining participants were then split into three cohorts based on omics data availability: a Metabolomics-only cohort (N = 220,859; those with metabolomics but not proteomics data), a Proteomics-only cohort (N = 19,086; those with proteomics but not metabolomics data), and a Multiomics cohort (N = 24,287; those with genomics, metabolomics, and proteomics data). In our CardiOmicScore framework, we used the Metabolomics-only and Proteomics-only cohorts to develop the respective deep learning models (MetNet and ProNet) and to generate the corresponding omics scores (MetScore and ProScore). To enable geographic validation, we further split each of these two cohorts into a training/validation set (England and Wales) and a geographic testing set (Scotland) according to the recruitment regions, following approaches commonly used in recent UKB studies39,40,41. The Multiomics cohort was kept untouched throughout model training and was subsequently used as an additional validation cohort to assess the incremental benefit of integrating multiple omics data types for risk prediction.
Categorical variables were one-hot encoded, and continuous variables were standardized by the mean and standard deviation. We used the K-nearest neighbors algorithm (scikit-learn v1.3.2 package)65, setting the number of neighbors to five, to impute missing values for continuous variables. Categorical variables were imputed with the mode. For metabolomics or proteomics data, imputation and standardization of continuous variables were performed using parameters derived exclusively from the training set. The fitted preprocessing models were then applied to the corresponding validation and geographic testing sets, as well as the Multiomics cohort. To derive imputation models and standardization parameters for subsequent application to the Multiomics cohort, we constructed two cohorts from all available individuals with relevant data: one for clinical information (N = 427,225) and one for genomics (N = 412,797).
Ascertainment of cardiovascular diseases
The six CVDs analyzed in this study included coronary artery disease (CAD), stroke, heart failure (HF), atrial fibrillation (AF), peripheral artery disease (PAD), and venous thromboembolism (VTE). All diseases were ascertained based on self-reported diagnoses and operations, hospital episode statistics, and death records. Self-reported information was used only to determine the presence of CVDs at baseline. Detailed definitions are provided in Supplementary Data 5. Follow-up duration was calculated from the date of baseline assessment to the date of incident outcomes, death, loss to follow-up, or end of available registry follow-up (November 30, 2023 for England & Wales and December 31, 2023 for Scotland66, whichever came first).
Multiomics data
Genetic data
Central quality control and imputation of genotypic data are available in previous literature64. In brief, genotypes were generated based on two closely related purpose-designed arrays (UK BiLEVE Axiom and UK Biobank Axiom). Genotypes were imputed using the Haplotype Reference Consortium and the merged UK10K and 1000 Genomes as reference panels.
PRSs were calculated for all CVDs based on the following equation, implemented via PLINK 1.9 software within the ukbrapR v0.3.7R package67,68.
where \(n\) is the number of single-nucleotide polymorphisms (SNPs), \({\beta }_{i}\) is the effect size of SNP \(i\), and \({{{{\rm{dosage}}}}}_{{ij}}\) is the number of risk alleles of SNP \(i\) for individual \(j\). Effect sizes of SNP-disease associations were collected from published PRSs available in the PGS Catalog13,69,70,71,72,73. We included 235 SNPs for CAD (PGS003438)13, 63 SNPs for stroke (PGS005230)69, 38 SNPs for HF (PGS003969)70, 154 SNPs for AF (PGS004905)71, 19 SNPs for PAD (PGS005158)72, and 297 SNPs for VTE (PGS000753)73. Full lists of SNPs are provided in Supplementary Data 6–11.
Metabolomics data
Metabolite biomarkers from EDTA-plasma samples were measured using Nightingale’s high-throughput NMR-based platform34,74. A total of 249 metabolites were quantified, with 168 in absolute concentrations and 81 ratios, covering lipoprotein lipids in 14 subclasses, apolipoproteins, fatty acids, cholesterol, cholesteryl esters, amino acids, ketone bodies, glycolysis metabolites, fluid balance-related metabolites, and inflammation biomarkers. Technical variation in these NMR metabolic biomarkers was removed using the ukbnmr v2.2R package75. We finally included 168 metabolic biomarkers in absolute concentrations in the present study (Supplementary Data 12)17.
Proteomics data
Detailed information on sample selection, data processing, and quality control is provided in published documents76. Briefly, proteomic profiling was performed in EDTA-plasma samples using the Olink Explore 1536 and Explore Expansion platforms, which measured 2923 unique proteins targeted by 2941 assays across four panels containing cardiometabolic, inflammation, neurology, and oncology. Protein measurements were expressed as normalized protein expression values on a log2 scale. After excluding three proteins with over 50% missingness across samples (GLIPR1, NPM1, PCOLCE), we included 2920 proteins in this study (Supplementary Data 13).
Clinical predictors and cardiovascular risk scores
Clinical predictors (n = 34) were collected at baseline, including demographic information (age, sex, ethnicity, and Townsend deprivation index), healthy lifestyles (current smoking, daily alcohol intake, sleep duration, physical activity, diet, and social connection), family disease history (family history of heart disease, stroke, hypertension, and diabetes), disease and medication history (hypertension, diabetes, lipid-lowering medication, and antihypertensive medication), physical measurements (height, weight, waist circumference, waist-hip ratio, body mass index, and systolic/diastolic blood pressure), and blood count (hemoglobin concentration, hematocrit percentage, and the count of leukocyte, lymphocyte, monocyte, neutrophil, eosinophil, basophil, and platelet). We defined three clinical predictor sets—AgeSex, Clin, and PANEL. Detailed definitions are listed in Supplementary Data 14. The Clin set consisted of 25 clinical predictors, covering sociodemographic factors, lifestyle, health history, and physical measurements, and the PANEL set extended the Clin set by adding blood count data17,26. Additionally, we calculated two sex-specific cardiovascular risk scores for primary prevention, ASCVD and SCORE2, that are recommended by the American College of Cardiology and European Society of Cardiology7,8,9 (Supplementary Methods).
Deep learning models
Model development
We developed two multitask deep neural networks, named MetNet and ProNet, to derive the MetScore and ProScore for six CVDs, with 168 metabolites and 2920 proteins as the input, respectively. MetNet was trained on the training set from the Metabolomics-only cohort, and ProNet on the training set from the Proteomics-only cohort, while their respective validation sets were used to monitor model fitting and select the optimal checkpoints. The final model was first evaluated on the geographic testing set to assess generalizability across regions. Subsequently, the Multiomics cohort, which remained untouched throughout model development, was used for a further performance evaluation.
MetNet and ProNet had similar model architectures, consisting of a shared network and six parallel, disease-specific networks (one for each CVD) (Supplementary Fig. 21). The shared neural network (denoted as “shared multilayer perceptron [MLP]”) included multiple fully connected layers, each with a nonlinear activation function, dropout, and batch normalization. The output of the shared network (i.e., shared representation) is a high-dimensional representation, capturing the common features for all CVDs. The disease-specific network, comprising a disease-specific MLP and a predictor MLP, was designed to learn disease-specific features and predict the risk of individual diseases (i.e., whether the disease will occur or not). The original metabolomic/proteomic biomarkers were fed into the disease-specific MLP with nonlinear activation functions, dropout, and batch normalization to obtain a disease-specific representation. The shared and disease-specific representations were then concatenated and passed on to the predictor MLP that included linear layers followed by nonlinear activation functions, dropout, and batch normalization before the final single-output layer to generate the MetScore or ProScore for each disease. These scores were subsequently used as predictors in CPH models to assess their utility for cardiovascular risk prediction. Binary cross-entropy was used as the loss function for each disease. To address the class imbalance in multitask prediction, we applied both sample-level and task-level weighting (Supplementary Methods). At the sample level, the pos_weight parameter was calculated as the ratio of the number of negative samples to positive samples for each task, increasing the penalty for misclassifying positive cases and encouraging the model to pay more attention to rare events. At the task level, each disease was assigned a weight equal to the inverse of its number of positive samples, ensuring that tasks with fewer positive cases contribute proportionally more to the overall loss. The final loss was computed as the mean of the weighted losses across all tasks.
Our multitask design enables the simultaneous modeling of multiple CVDs, accounting for individuals who experience several events over time (e.g., CAD followed by stroke or HF). In this setup, each participant simultaneously contributes to the prediction tasks for all six diseases. If a participant experiences multiple incident diseases, they generate multiple non-zero losses but are only included once in the dataset. This design avoids data duplication while fully leveraging all available incident disease information17,26. It also reflects the clinical reality in which multimorbidity is increasingly common, and where the most useful risk prediction models are those capable of discriminating the outcome of interest in the presence of co-occurring diseases27.
We performed random hyperparameter searches separately for MetNet and ProNet using their respective training and validation sets. Each model was trained and evaluated across 100 randomly sampled hyperparameter configurations. Hyperparameters included the hidden layer architecture, dropout rate, and activation function for all MLPs (Supplementary Data 15). The optimal hyperparameter combination for each model, provided in Supplementary Data 15, was selected based on the highest average Harrell’s C-index across all six CVDs on the validation set. Once the optimal architecture was determined, we trained the final models five times using different random seeds to account for stochastic variability in the training process. The ultimate MetScore and ProScore for each participant represent the average of the predictions from these five independent runs, ensuring the robustness of the final scores. The final models were trained using the Adam optimizer. All deep learning models were developed in Python v3.8.5 using PyTorch v1.11.0 package with hyperparameter tuning using Optuna v4.0.0 package77,78.
Model interpretation
Since our deep learning models, MetNet and ProNet, are black-box models, we further employed the SHAP method (shap v0.46.0 package), a post-hoc explanation algorithm, to explore the relationships of metabolomic and proteomic biomarkers with each outcome. SHAP values, computed based on a game-theoretical approach, can quantify the contribution of each predictor to the prediction for each participant and explain the final prediction as the sum of the SHAP value of each predictor79. Higher (or lower) SHAP values imply large positive (or negative) contributions to the MetScore or ProScore, where both scores were positively related to CVD risk. To ensure stable and reliable interpretations, we generated SHAP values for each of the five models trained with different random seeds. The final SHAP value for each biomarker was calculated by averaging these results across the five runs. We then calculated the mean absolute SHAP values across all participants to obtain the global impact of each predictor. We used all participants in the Multiomics cohort to explain our models.
Statistical analyses
Baseline characteristics were summarized using median (IQR) for continuous variables or numbers (percentages) for categorical variables. Chi-squared tests were used for categorical variables, and Mann–Whitney U tests were used for non-normally distributed continuous variables to compare baseline characteristics between the training and validation sets, and between the training and geographic testing sets. Incidence proportion for each specific CVD was calculated as the number of new cases for that disease divided by the total number of participants at risk at baseline. All statistical analyses were conducted using Python v3.8.5, and visualizations were performed in R v4.3.3. A two-sided P < 0.05 was considered statistically significant.
We conducted model performance evaluations on the Multiomics cohort, including risk stratification, Harrell’s C-index, calibration plot, and net benefit curve. For the risk stratification, we classified the participants into tertile groups (low, medium, and high) based on PRS, MetScore, and ProScore, respectively. Then, we used the Kaplan–Meier curve (lifelines v0.27.8 package)80 to visualize the cumulative survival probability over time and compared the survival functions across groups using the log-rank test (lifelines v0.27.8 package). Finally, we fitted CPH models for each score to estimate HRs with 95% CIs. Each score was analyzed both as a categorical variable (tertiles, with the low group as the reference) and as a continuous variable (per 1−SD increase). All models were adjusted for the following covariates from the PANEL set: demographic information, healthy lifestyle factors, family history of disease, disease and medication history, physical measurements, and blood count.
CPH models were fitted on various predictor sets. Specifically, we first developed models with only PRS, MetScore, or ProScore as continuous variables; next, we fitted models using clinical predictor sets of AgeSex, Clin, and PANEL; finally, we added omics information to clinical predictor-based models to assess the combined predictive power of PRS, MetScore, and ProScore when integrated with traditional clinical variables. For example, AgeSex +PRS +MetScore +ProScore means we included age, sex, and three omics scores as the predictors in the CPH model. All CPH models were fitted on the Multiomics cohort using the lifelines v0.27.8 package, with a penalizer of 0.03 to facilitate model convergence. We calculated the Harrell’s C-index to assess model discriminative performance. The C-index was reported with a point estimate and 95% confidence intervals (CIs) calculated via 1000 bootstrap resamples of the Multiomics cohort. Discriminative performance was considered significantly different when the 95% CIs of the C-index deltas did not include zero17,27.
Additionally, we predicted the probability of incident diseases at 15 years to draw calibration curves and net benefit curves, two recommended metrics for calibration and clinical utility of prediction models42,43. Calibration curves can measure the concordance between observed event rates and predicted risks by CPH models, while net benefit curves (dcurves v0.5.0 R package) are used to evaluate the net benefit of prediction models compared with two strategies of treat all patients and treat no patients. Since the net benefit was difficult to interpret, we further calculated the standardized net benefit by dividing it by its maximum value81. We also computed the area under the net benefit curve (A-NBC) using trapezoidal numerical integration, providing a summary metric of model performance44.
To test the robustness of our findings, we conducted six sensitivity analyses. First, to evaluate the performance of key individual biomarkers, we developed separate CPH models based on clinical predictors in combination with one biomarker at a time. The biomarkers selected for this analysis included those with high global importance in our deep learning models (i.e., the top five non-overlapping biomarkers by mean absolute SHAP value for each CVD, resulting in 8 metabolites and 17 proteins) or routinely used in clinical practice. Second, since age, sex, and lipid-lowering and antihypertensive therapy may influence metabolomic or proteomic profiles82,83,84, we refitted CPH models by age, sex, and baseline use of lipid-lowering or antihypertensive medications to assess whether the predictive performance of the models was consistent across subgroups. Third, to mitigate the potential for reverse causation, we repeated the primary analyses after excluding incident cases that occurred within the first two years of follow-up for each respective CVD. Fourth, to account for the competing risk of death, we fitted Fine–Gray subdistribution hazard models, treating all-cause death as a competing event (cmprsk v2.2.11 R package). Fifth, to demonstrate the advantage of our CardiOmicScore framework for high-dimensional omics data, we developed alternative MetScore and ProScore using several statistical machine learning algorithms, including Extreme Gradient Boosting (XGBoost, xgboost v2.0.3 package), LightGBM (lightgbm v4.4.0 package), random forest (scikit-learn v1.3.2 package), and logistic regression (scikit-learn v1.3.2 package), following an identical development pipeline (Supplementary Methods and Supplementary Data 16). We then compared the predictive utility of these scores against those generated by MetNet and ProNet. Finally, to investigate the potential effect of lipid-lowering treatment, we refitted the MetNet using the same architecture but trained only on participants not taking lipid-lowering medication. We then calculated the mean absolute SHAP values to identify the important metabolites within this subgroup. Additionally, using the same analytical procedure as our first sensitivity analysis, we evaluated the predictive performance of traditional lipid-related biomarkers in the Multiomics cohort after excluding baseline users of lipid-lowering medication, and compared it with their performance in the full set.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
This research has been conducted using data from the UK Biobank (Application Number 79146). Researchers may apply for access to the UK Biobank data through the UK Biobank website (https://www.ukbiobank.ac.uk/). Source data are provided with this paper.
Code availability
Code associated with our paper is available at https://github.com/YanLuoCityU/cardiomicscore. An online demo is available at https://yanluocityu.github.io/cardiomicscore-website/.
References
Mensah, G. A., Fuster, V., Murray, C. J. L., Roth, G. A. & Global Burden of Cardiovascular Diseases and Risks Collaborators. Global burden of cardiovascular diseases and risks, 1990–2022. J. Am. Coll. Cardiol. 82, 2350–2473 (2023).
Chong, B. et al. Global burden of cardiovascular diseases: projections from 2025 to 2050. Eur. J. Prev. Cardiol. zwae281 (2024) https://doi.org/10.1093/eurjpc/zwae281.
World Health Organization. HEARTS: Technical Package For Cardiovascular Disease Management in Primary Health Care: Risk-based CVD Management (World Health Organization, 2020).
Arnett, D. K. et al. 2019 ACC/AHA guideline on the primary prevention of cardiovascular disease: a report of the American College of Cardiology/American Heart Association Task Force on Clinical Practice Guidelines. Circulation 140, e596–e646 (2019).
Visseren, F. L. J. et al. 2021 ESC Guidelines on cardiovascular disease prevention in clinical practice. Eur. Heart J. 42, 3227–3337 (2021).
Sing, C. F., Stengård, J. H. & Kardia, S. L. R. Genes, environment, and cardiovascular disease. Arterioscler. Thromb. Vasc. Biol. 23, 1190–1196 (2003).
Goff, D. C. et al. 2013 ACC/AHA guideline on the assessment of cardiovascular risk. Circulation 129, S49–S73 (2014).
Yadlowsky, S. et al. Clinical implications of revised pooled cohort equations for estimating atherosclerotic cardiovascular disease risk. Ann. Intern. Med. 169, 20–29 (2018).
SCORE2 working group and ESC Cardiovascular risk collaboration. SCORE2 risk prediction algorithms: new models to estimate 10-year risk of cardiovascular disease in Europe. Eur. Heart J. 42, 2439–2454 (2021).
Siontis, G. C. M., Tzoulaki, I., Siontis, K. C. & Ioannidis, J. P. A. Comparisons of established risk prediction models for cardiovascular disease: systematic review. BMJ 344, e3318 (2012).
Giugni, F. R., Berry, J. D., Khera, A., Shah, A. M. & de Lemos, J. A. Precision medicine for cardiovascular prevention and population health: a bridge too far?. Circulation 150, 1720–1731 (2024).
Elliott, J. et al. Predictive accuracy of a polygenic risk score–enhanced prediction model vs. a clinical risk score for coronary artery disease. JAMA 323, 636–645 (2020).
Marston, N. A. et al. Predictive utility of a coronary artery disease polygenic risk score in primary prevention. JAMA Cardiol. 8, 130–137 (2023).
Li, L., Pang, S., Starnecker, F., Mueller-Myhsok, B. & Schunkert, H. Integration of a polygenic score into guideline-recommended prediction of cardiovascular disease. Eur. Heart J. ehae048 https://doi.org/10.1093/eurheartj/ehae048 (2024).
Nurmohamed, N. S. et al. Proteomics and lipidomics in atherosclerotic cardiovascular disease risk prediction. Eur. Heart J. 44, 1594–1607 (2023).
Lv, J. et al. Plasma metabolomics reveals the shared and distinct metabolic disturbances associated with cardiovascular events in coronary artery disease. Nat. Commun. 15, 5729 (2024).
Buergel, T. et al. Metabolomic profiles predict individual multidisease outcomes. Nat. Med. 28, 2309–2320 (2022).
Ritchie, S. C. et al. Combined clinical, metabolomic, and polygenic scores for cardiovascular risk prediction. Eur. Heart J. ehaf947, https://doi.org/10.1093/eurheartj/ehaf947 (2025).
Nightingale Health Biobank Collaborative Group. Metabolomic and genomic prediction of common diseases in 700,217 participants in three national biobanks. Nat. Commun. 15, 10092 (2024).
Helgason, H. et al. Evaluation of large-scale proteomics for prediction of cardiovascular events. JAMA 330, 725–735 (2023).
Garg, M. et al. Disease prediction with multi-omics and biomarkers empowers case-control genetic discoveries in the UK Biobank. Nat. Genet. 56, 1821–1831 (2024).
Hoogeveen, R. M. et al. Improved cardiovascular risk prediction using targeted plasma proteomics in primary prevention. Eur. Heart J. 41, 3998–4007 (2020).
Shelbaya, K. et al. Large-scale proteomics identifies novel biomarkers and circulating risk factors for aortic stenosis. J. Am. Coll. Cardiol. 83, 577–591 (2024).
Royer, P. et al. Large-scale plasma proteomics in the UK Biobank modestly improves prediction of major cardiovascular events in a population without previous cardiovascular disease. Eur. J. Prev. Cardiol. zwae124 (2024) https://doi.org/10.1093/eurjpc/zwae124.
Schuermans, A. et al. Integrative proteomic analyses across common cardiac diseases yield mechanistic insights and enhanced prediction. Nat. Cardiovasc. Res. 1–15 https://doi.org/10.1038/s44161-024-00567-0 (2024).
You, J. et al. Plasma proteomic profiles predict individual future health risk. Nat. Commun. 14, 7817 (2023).
Carrasco-Zanini, J. et al. Proteomic signatures improve risk prediction for common and rare diseases. Nat. Med. 1–10 https://doi.org/10.1038/s41591-024-03142-z (2024).
Carrasco-Zanini, J. et al. Proteomic prediction of diverse incident diseases: a machine learning-guided biomarker discovery study using data from a prospective cohort study. Lancet Digit. Health 6, e470–e479 (2024).
Gadd, D. A. et al. Blood protein assessment of leading incident diseases and mortality in the UK Biobank. Nat. Aging 1–10 (2024) https://doi.org/10.1038/s43587-024-00655-7.
Ho, F. K. et al. A proteomics-based approach for prediction of different cardiovascular diseases and dementia. Circulation https://doi.org/10.1161/CIRCULATIONAHA.124.070454 (2024).
Liu, M. et al. Enhanced prediction of atrial fibrillation risk using proteomic markers: a comparative analysis with clinical and polygenic risk scores. Heart 110, 1270–1276 (2024).
Gustafsson, S. et al. Markers of imminent myocardial infarction. Nat. Cardiovasc. Res. 3, 130–139 (2024).
Mazidi, M. et al. Risk prediction of ischemic heart disease using plasma proteomics, conventional risk factors and polygenic scores in Chinese and European adults. Eur. J. Epidemiol. https://doi.org/10.1007/s10654-024-01168-8 (2024).
Julkunen, H. et al. Atlas of plasma NMR biomarkers for health and disease in 118,461 individuals from the UK Biobank. Nat. Commun. 14, 604 (2023).
Gong, M. et al. Proteomics in Cardiovascular disease. Clin. Chim. Acta 557, 117877 (2024).
Mokou, M., Lygirou, V., Vlahou, A. & Mischak, H. Proteomics in cardiovascular disease: recent progress and clinical implication and implementation. Expert Rev. Proteom. 14, 117–136 (2017).
Carrasco-Zanini, J. et al. Multi-omic prediction of incident type 2 diabetes. Diabetologia 67, 102–112 (2024).
Nielsen, R. L. et al. Data-driven identification of predictive risk biomarkers for subgroups of osteoarthritis using interpretable machine learning. Nat. Commun. 15, 2817 (2024).
Argentieri, M. A. et al. Proteomic aging clock predicts mortality and risk of common age-related diseases in diverse populations. Nat. Med. 1–11 https://doi.org/10.1038/s41591-024-03164-7 (2024).
Zhang, S. et al. A metabolomic profile of biological aging in 250,341 individuals from the UK Biobank. Nat. Commun. 15, 8081 (2024).
Julkunen, H. & Rousu, J. Comprehensive interaction modeling with machine learning improves prediction of disease risk in the UK Biobank. Nat. Commun. 16, 6620 (2025).
Efthimiou, O. et al. Developing clinical prediction models: a step-by-step guide. BMJ e078276 https://doi.org/10.1136/bmj-2023-078276 (2024).
Vickers, A. J., Calster, B. V. & Steyerberg, E. W. Net benefit approaches to the evaluation of prediction models, molecular markers, and diagnostic tests. BMJ 352, i6 (2016).
Zhang, Z. et al. Decision curve analysis: a technical note. Ann. Transl. Med. 6, 308 (2018).
Hasin, Y., Seldin, M. & Lusis, A. Multi-omics approaches to disease. Genome Biol. 18, 1–15 (2017).
Lee, J. et al. Integrative metabolomics differentiate coronary artery disease, peripheral artery disease, and venous thromboembolism risks. Arterioscler. Thromb. Vasc. Biol. 44, 2108–2117 (2024).
McKie, P. M. & Burnett, J. C. NT-proBNP: the gold standard biomarker in heart failure. J. Am. Coll. Cardiol. 68, 2437–2439 (2016).
Volpe, M., Rubattu, S. & Burnett, J. Natriuretic peptides in cardiovascular diseases: current use and perspectives. Eur. Heart J. 35, 419–425 (2014).
Wannamethee, S. G., Shaper, A. G. & Perry, I. J. Serum creatinine concentration and risk of cardiovascular disease. Stroke 28, 557–563 (1997).
Ronit, A. et al. Plasma albumin and incident cardiovascular disease. Arterioscler. Thromb. Vasc. Biol. 40, 473–482 (2020).
Connelly, M. A., Otvos, J. D., Shalaurova, I., Playford, M. P. & Mehta, N. N. GlycA, a novel biomarker of systemic inflammation and cardiovascular disease risk. J. Transl. Med. 15, 219 (2017).
Borges, M. C. et al. Circulating fatty acids and risk of coronary heart disease and stroke: individual participant data meta-analysis in up to 16 126 participants. J. Am. Heart Assoc. 9, e013131 (2020).
Hu, S. et al. Causal relationships of circulating amino acids with cardiovascular disease: a trans-ancestry Mendelian randomization analysis. J. Transl. Med. 21, 699 (2023).
Ginsberg, H. N. et al. Triglyceride-rich lipoproteins and their remnants: metabolic insights, role in atherosclerotic cardiovascular disease, and emerging therapeutic strategies—a consensus statement from the European Atherosclerosis Society. Eur. Heart J. 42, 4791–4806 (2021).
Wollert, K. C., Kempf, T. & Wallentin, L. Growth differentiation factor 15 as a biomarker in cardiovascular disease. Clin. Chem. 63, 140–151 (2017).
Kato, E. T. et al. Growth differentiation factor 15 and cardiovascular risk: individual patient meta-analysis. Eur. Heart J. 44, 293–300 (2023).
Goncalves, I. et al. Elevated plasma levels of MMP-12 are associated with atherosclerotic burden and symptomatic cardiovascular disease in subjects with type 2 diabetes. Arterioscler. Thromb. Vasc. Biol. 35, 1723–1731 (2015).
Yuan, S. et al. Circulating proteins and peripheral artery disease risk: observational and Mendelian randomization analyses. Eur. Heart J. Open 3, oead056 (2023).
Kong, L. et al. Downregulation of let-7e-5p contributes to endothelial progenitor cell dysfunction in deep vein thrombosis via targeting FASLG. Thromb. Res. 138, 30–36 (2016).
Khalil, M. et al. Neurofilaments as biomarkers in neurological disorders - towards clinical application. Nat. Rev. Neurol. 20, 269–287 (2024).
Park, S. et al. Hypoxia-mediated CRIP2 activation via NICD1 translocation regulates glycolysis and cell death. Genes Dis. 101704, https://doi.org/10.1016/j.gendis.2025.101704 (2025).
Hughes, E., Clayton, J. C. & Middleton, D. A. Cytoplasmic residues of phospholamban interact with membrane surfaces in the presence of SERCA: a new role for phospholipids in the regulation of cardiac calcium cycling?. Biochim. Biophys. Acta 1788, 559–566 (2009).
Abdelsayed, M., Kort, E. J., Jovinge, S. & Mercola, M. Repurposing drugs to treat cardiovascular disease in the era of precision medicine. Nat. Rev. Cardiol. 19, 751–764 (2022).
Bycroft, C. et al. The UK Biobank resource with deep phenotyping and genomic data. Nature 562, 203–209 (2018).
Pedregosa, F. et al. Scikit-learn: machine learning in Python. J. Mach. Learn. Res. 12, 2825–2830 (2011).
UK Biobank. Data Providers and Dates of Data Availability https://biobank.ndph.ox.ac.uk/ukb/exinfo.cgi?src=Data_providers_and_dates.
Collister, J. A., Liu, X. & Clifton, L. Calculating Polygenic Risk Scores (PRS) in UK Biobank: a practical guide for epidemiologists. Front. Genet. 13, 818574 (2022).
Pilling, L. ukbrapR: R Functions to Use in the UK Biobank Research Analysis Platform (RAP) https://lcpilling.github.io/ukbrapR, https://github.com/lcpilling/ukbrapR (2025).
Ma, Y. et al. Genetic susceptibility modifies relationships between air pollutants and stroke risk: a Large Cohort Study. Stroke 55, 113–121 (2024).
Rasooly, D. et al. Genome-wide association analysis and Mendelian randomization proteomics identify drug targets for heart failure. Nat. Commun. 14, 3826 (2023).
Ma, Y. et al. Air pollution, genetic susceptibility, and the risk of atrial fibrillation: a large prospective cohort study. Proc. Natl. Acad. Sci. USA 120, e2302708120 (2023).
Klarin, D. et al. Genome-wide Association Study of peripheral artery disease in the Million Veteran Program. Nat. Med. 25, 1274–1279 (2019).
Zhu, K. et al. Modifiable lifestyle factors, genetic risk, and incident peripheral artery disease among individuals with type 2 diabetes: a prospective study. Diabetes Care 47, 435–443 (2024).
Würtz, P. et al. Quantitative serum nuclear magnetic resonance metabolomics in large-scale epidemiology: a primer on -omic technologies. Am. J. Epidemiol. 186, 1084–1096 (2017).
Ritchie, S. C. et al. Quality control and removal of technical variation of NMR metabolic biomarker data in ~120,000 UK Biobank participants. Sci. Data 10, 64 (2023).
Sun, B. B. et al. Plasma proteomic associations with genetics and health in the UK Biobank. Nature 622, 329–338 (2023).
Akiba, T., Sano, S., Yanase, T., Ohta, T. & Koyama, M. Optuna: a next-generation hyperparameter optimization framework. In Proc. of the 25th ACM SIGKDD International Conference on Knowledge Discovery & Data Mining 2623–2631 (Association for Computing Machinery, 2019).
Paszke, A. et al. PyTorch: an imperative style, high-performance deep learning library. In Proc. of the 33rd International Conference on Neural Information Processing Systems 8026–8037 (Curran Associates Inc., 2019).
Lundberg, S. M. et al. From local explanations to global understanding with explainable AI for trees. Nat. Mach. Intell. 2, 56–67 (2020).
Davidson-Pilon, C. Lifelines: survival analysis in Python. J. Open Source Softw. 4, 1317 (2019).
Kerr, K. F., Brown, M. D., Zhu, K. & Janes, H. Assessing the clinical impact of risk prediction models with decision curves: guidance for correct interpretation and appropriate use. J. Clin. Oncol. 34, 2534–2540 (2016).
Darst, B. F., Koscik, R. L., Hogan, K. J., Johnson, S. C. & Engelman, C. D. Longitudinal plasma metabolomics of aging and sex. Aging (Albany, NY) 11, 1262–1282 (2019).
Altmaier, E. et al. Metabolomics approach reveals effects of antihypertensives and lipid-lowering drugs on the human metabolism. Eur. J. Epidemiol. 29, 325–336 (2014).
Enroth, S. et al. Systemic and specific effects of antihypertensive and lipid-lowering medication on plasma protein biomarkers for cardiovascular diseases. Sci. Rep. 8, 5531 (2018).
Acknowledgements
This research has been conducted using the UK Biobank Resource under Application Number 79146. This work was supported by the General Research Fund of the Research Grants Council of Hong Kong (17209225 to Q.Z.) and the Seed Fund for Collaborative Research of The University of Hong Kong (2407102490 to Q.Z.); National Natural Science Foundation of China (82370332 and 82570390 to T.L.), Tianjin Key Medical Discipline Construction Project (TJYXZDXK-3-006B to T.L.). We acknowledge the computational resources provided by the HKU Shanghai Intelligent Computing Research Center. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
Y.L.: Conceptualization, data collection, model development, data analysis, manuscript writing, and manuscript editing. N.Z., J.Y., and M.C.: data collection, data analysis, and manuscript editing. K.K.F.T. and G.Y.H.L.: data analysis and manuscript editing. T.L.: Project supervision, funding acquisition, manuscript writing, and manuscript editing. Q.Z.: Conceptualization, model development, project supervision, funding acquisition, manuscript writing, and manuscript editing.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Luo, Y., Zhang, N., Yang, J. et al. AI-based multiomics profiling reveals complementary omics contributions to personalized prediction of cardiovascular disease. Nat Commun 17, 2269 (2026). https://doi.org/10.1038/s41467-026-68956-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-68956-6