Abstract
Lung microenvironment controls the homeostasis and function of alveolar macrophages (AMs), the major regulators of lung immunity, but the underlying mechanisms, particularly the role of microbiota, remain unclear. Here, with Lyz2creEi24fl/fl mice, we report that EI24 deficiency in macrophages disrupts AMs homeostasis but enhances their phagocytosis and inflammatory responses via metabolic rewiring. Consequently, Lyz2creEi24fl/fl mice exhibit resistance to viral infection and tumor metastasis in lung. Notably, EI24 expression in AMs is upregulated by commensal microbiota through TLR2/4 signaling. These data demonstrate that microbiota upregulates EI24 in AMs to favor their homeostasis, but it retards their immune surveillance function in the lung. Our study thus indicates that deleting EI24 enhances anti-viral and anti-tumor effects of macrophage-based immunotherapy.
Similar content being viewed by others
Introduction
Alveolar macrophages (AMs) are major immune cells in alveolar airspace. These cells have the capacity of self-renewal and play vital roles in maintaining tissue homeostasis, clearing pathogens and malignant cells, and promoting tissue repair1,2,3,4. It is well established that AMs clear pulmonary surfactant and cell debris under homeostatic condition5, produce proinflammatory cytokines IL-1β, IL-6, and TNF6. In addition to their “big eater” role, AMs also prevent pathogen infection and tumor growth, and release TGF-β and IL-10 to regulate epithelial cell function and participate in immunosuppression7,8. Impaired homeostasis or malfunction of AMs is associated with multiple lung diseases9,10,11.
AMs are derived from fetal liver, and their development is controlled by GM-CSF-PPARγ axis and TGF-β11,12,13. Their self-renewal maintains homeostasis in steady state1. The pulmonary environment is quite unique, with abundant surfactant, unusual nutrients, and exposure to pollutants and microbiota14. Published studies imply key roles of local microenvironment in regulating differentiation, homeostasis, and function of AMs. For example, downregulation of hypoxia-inducible factor 1-alpha (HIF-1α) as postnatal adaptation of AMs to lung microenvironment transition from hypoxia to normoxia is important for their maturation in lung15,16. In line with the low glucose environment in lung, AMs depend on mitochondrial oxidative phosphorylation rather than glycolysis in steady state17, whereas increase glycolysis under hypoxia condition to support their survival via a HIF-1α dependent manner15,18. Lipid enrichment is another feature of lung microenvironment and is in line with high lipid metabolism in AMs19,20. Impaired lipid metabolism, as a result of NK cell-associated receptor NKR-P1B deficiency, leads to death of AMs21. Additionally, binding of hyaluronan to CD44 in lung is crucial for AMs survival and renewal22. In terms of functional control, low glycolysis in AMs is coupled to hypo-responsiveness of AMs to IL-423. Epithelial cells derived IL-10, TGF-β, and CD200-CD200R signaling favor inflammation resolution mediated by AMs7,24. Surfactant proteins A and D suppress phagocytosis of AMs in steady state25,26. In respect of microbiota, not only pulmonary microbiota but also intestinal microbiota has been reported to modulate function of AMs27,28,29. Several microbes have been shown to enhance pathogen clearance function of AMs via inducing trained immunity or via microbial metabolites30,31,32. On the other hand, lung microbiota is known to induce tolerance for maintaining respiratory health. For instance, microbiota dampens anti-viral response of AMs and induces M2 polarization of AMs27,29. Overall, local microenvironment maintains AMs homeostasis, educates them to act as the first line of body defense, and on the other hand, avoids excessive activation upon pulmonary air antigen encounter. The mechanisms underlying precise control of homeostasis and function of AMs by local microenvironment are largely unknown. Particularly, the contribution of microbiota and its impact on lung immunity and lung diseases remain to be explored.
Etoposide-induced protein 2.4 (EI24) is a p53-induced protein and has been previously reported as an essential autophagy protein33,34,35,36. Although EI24 has been suggested as a biomarker for prognosis of cancers and is related to tumor immune microenvironment37,38, its roles in innate immune homeostasis and immune surveillance remain unexplored. Here, we propose that microbiota upregulates EI24 in AMs through Toll-like receptors (TLRs) signaling, and that maintains homeostasis of AMs but retards their immune surveillance function against viral infection and tumor metastasis in lung. Furthermore, EI24 generally inhibits macrophage functions, and thus deleting EI24 in bone marrow derived-macrophages (BMDMs) enhances their efficacy in preventing viral infection and tumor metastasis in lung. Overall, our study reveals adaptations of macrophages to tissue microenvironment, particularly microbiota, and indicates a new strategy for developing macrophage-based immunotherapy.
Results
EI24 is required for homeostasis of mature AMs
To examine EI24 expression in various tissue-resident macrophage populations, we generated Ei24-Gfp reporter mice, and EI24 expression was indicated by GFP level. After autofluorescence subtraction, AMs characterized as CD11bloCD11chiSiglecF+ displayed much higher GFP expression than lung interstitial macrophages (IMs), splenic red pulp macrophages (RPMs) and liver macrophages (Fig. 1a; Supplementary Fig. 1a–c), indicating particularly higher expression of EI24 in AMs. To elucidate the role of EI24 in AMs, we crossed Ei24fl/fl mice with Lyz2cre mice to ablate EI24 in macrophages. Deletion of EI24 was confirmed in primary AMs isolated from Lyz2creEi24fl/fl mice (Fig. 1b). Next, we compared macrophage frequencies and numbers across tissues between Ei24fl/fl and Lyz2creEi24fl/fl mice. We observed a marked reduction (by about 50%) of mature AMs in the lungs of Lyz2creEi24fl/fl adult mice in comparison with littermate controls in steady state (Fig. 1c). In contrast, the frequencies and numbers of lung IMs, liver macrophages and splenic RPMs were unaffected by EI24 deletion (Fig. 1d–f). Additionally, we used another Cre-expressing mice, Cd11ccre mice, to delete EI24 in AMs, RPMs, and liver macrophages11, and confirmed that deletion of EI24 impaired AMs but not RPMs or liver macrophages (Supplementary Fig. 2a–c). These findings suggest that EI24 is required for AMs but dispensable for other tissue-resident macrophages.
a Flow cytometric analysis of GFP expression in RPMs, liver macrophages, AMs, and IMs from Ei24-Gfp mice (n = 6 per group). b Ei24 gene expression of AMs sorted from Ei24fl/fl and Lyz2creEi24fl/fl mice (n = 5 per group). c–f The frequencies (left) and absolute numbers (right) of AMs (c), IMs (d), liver macrophages (e), and RPMs (f) in Ei24fl/fl and Lyz2creEi24fl/fl mice (c, n = 8, 13, 7, 9; d, n = 10, 10, 9, 11; e, n = 15, 13, 9, 9; f, n = 16, 16, 7, 7). g Representative flow cytometry of pre-AMs and mature AMs derived from the lungs of Ei24fl/fland Lyz2creEi24fl/fl mice (PND2). h, i The frequencies and absolute numbers of pre-AMs (h) and mature AMs (i) in PND2 Ei24fl/fl (n = 9) and Lyz2creEi24fl/fl (n = 5) mice. j, k Percentages (j) and absolute numbers (k) of mature AMs at the indicated ages in Ei24fl/fl and Lyz2creEi24fl/fl mice (percentages, n = 6, 6, 13, 16, 5, 6; numbers: n = 6, 6, 8, 9, 5, 6). l Frequencies of Ki67+ AMs in Lyz2creEi24fl/fl and Ei24fl/fl mice (n = 6 per group). m Frequencies of Annexin V+ AMs in adult Ei24fl/fl and Lyz2creEi24fl/fl mice (n = 7 per group). n, o Representative flow cytometry histograms (n) and MFI (o) of Cleaved Caspase-3 expression in AMs from Lyz2creEi24fl/fl (n = 6) and Ei24fl/fl (n = 5) mice. p, q MFI of Cleaved Caspase-3 (p) in AMs and absolute AM numbers (q) in Lyz2creEi24fl/fl mice treated with vehicle- (n = 7) or Z-VAD-FMK (n = 6). Data are pooled from (a–f, h–q) two to three independent experiments and presented as the mean ± SEM. Data were analyzed by one-way ANOVA followed by a Tukey’s test (a), two-tailed unpaired t-test (b–f, h, i, l, m, o–q), and two-way analysis of variance (ANOVA) with Šidák’s multiple comparisons test (j, k). Source data are provided as a Source Data file.
AMs are derived primarily from Ly6ChiCD11bhi fetal liver monocytes, which colonize the developing lung around E16.5 and mature postnatally12. To determine whether EI24 is critical for development of AMs, we assessed the progenitors (pre-AMs) and mature AMs at postnatal day 2 (PND2) and found no differences in the frequencies and numbers of pre-AMs or mature AMs between Ei24fl/fl mice and Lyz2creEi24fl/fl mice (Fig. 1g–i). As the significant reduction of AMs in adult Lyz2creEi24fl/fl mice was independent of developmental defect, we next investigated the impact of EI24 on the homeostatic maintenance of AMs. The kinetic analysis showed that frequencies of AMs decreased dramatically at 4 weeks of age in Lyz2creEi24fl/fl mice (Fig. 1j), and the numbers of AMs reduced at 6 weeks in Lyz2creEi24fl/fl mice (Fig. 1k). These results demonstrate that the observed reduction of AMs in Lyz2creEi24fl/fl mice is not attributable to failures in development, but rather to impaired homeostatic maintenance. AMs self-maintain throughout life with minimal replenishment from circulating monocytes in steady state1. We next examined whether loss of EI24 affected AM survival or proliferation. Ki67 staining showed no differences in proliferation between AMs from Ei24fl/fl mice and from Lyz2creEi24fl/fl mice (Fig. 1l). However, Annexin V staining revealed higher apoptosis in EI24-deficient AMs (Fig. 1m, Supplementary Fig. 2d), and that was corroborated by the significantly increased level of Cleaved Caspase-3 (Fig. 1n, o). Moreover, we found that administration of caspase inhibitor Z-VAD-FMK inhibited Caspase-3 activation in AMs and restored their cell numbers in Lyz2creEi24fl/fl mice (Fig. 1p, q). Together, these data demonstrate that EI24 is essential for maintaining survival of mature AMs.
EI24 inhibits phagocytosis and inflammatory responses of AMs
EI24 deficiency significantly elevated expression of CD11b and CD11c in AMs (Fig. 2a, b, Supplementary Fig. 2e), whereas expression of other AM surface markers, such as F4/80 and SiglecF, were unchanged. Additionally, EI24-deficient AMs exhibited upregulated expression of IA/IE (Fig. 2b, Supplementary Fig. 2e). These results indicated active phenotype of EI24-deficient AMs. Given the role of CD11b and CD11c in phagocytosis39, we then performed a phagocytosis assay using pHrodo-Green-labeled E.coli. EI24-deficient AMs exhibited significantly increased phagocytosis in comparison with control AMs (Fig. 2c, d). To further explore functional changes induced by EI24 deficiency, we sorted lung AMs from Lyz2creEi24fl/fl mice and Ei24fl/fl mice, stimulated them with lipopolysaccharide (LPS) overnight (16 h) in vitro, and measured proinflammatory cytokines production. Comparing with cells from Ei24fl/fl mice, AMs from Lyz2creEi24fl/fl mice secreted higher levels of proinflammatory cytokines IL-1β and IL-6, despite unchanged TNF and IL-10 levels (Fig. 2e), suggesting enhanced inflammatory responses in the absence of EI24. Consistently, AMs from Cd11ccreEi24fl/fl mice also elevated phagocytosis (Supplementary Fig. 2f) and LPS-induced inflammatory responses in comparison with cells from littermate control mice (Supplementary Fig. 2g), confirming the role of EI24 in retarding AMs functionality.
a Representative histograms of indicated molecules on AMs from Lyz2creEi24fl/fl and Ei24fl/fl mice. b MFI of CD11b, CD11c and IA/IE on AMs derived from Lyz2creEi24fl/fl and Ei24fl/fl mice (n = 6 per group). c–d Representative dot plot (c) and summarized data (d) showing phagocytosis of pHrodo-Green-labeled E.coli by AMs from Lyz2creEi24fl/fl and Ei24fl/fl mice (n = 5 per group). e AMs sorted from Ei24fl/fl and Lyz2creEi24fl/fl mice were untreated or stimulated with LPS and ATP in vitro. Levels of IL-1β, IL-6, TNF and IL-10 in the culture supernatant were measured by CBA. IL-1β (n = 9, 9, 10, 11), IL-6 (n = 6, 7, 6, 7), TNF (n = 6, 7, 6, 7), and IL-10 (n = 4, 5, 7, 9). f PCA of RNA-seq normalized gene expression data obtained from AMs of Ei24fl/fl and Lyz2creEi24fl/fl mice (n = 3 per group). g Volcano plots showing genes differentially expressed in AMs from Ei24fl/fl and Lyz2creEi24fl/fl mice (Fold Change ≥ 2 or ≤ 0.5, adjusted P < 0.01). h GO term enrichment of differential expressed genes in AMs sorted from Lyz2creEi24fl/fl mice versus Ei24fl/fl mice. i KEGG pathway enrichment analysis of upregulated genes in Lyz2creEi24fl/fl mice versus Ei24fl/fl mice. j–l RNA-seq heatmaps showing mRNA transcripts significantly upregulated related to antigen presentation process, cytokine-cytokine receptor and phagocytosis in AMs from Lyz2creEi24fl/fl and Ei24fl/fl mice. m–o GSEA analysis of gene transcripts related to receptor-mediated endocytosis, cell killing and regulation of inflammatory response signatures comparing AMs from Lyz2creEi24fl/fl and Ei24fl/fl mice. Data are pooled from two to three independent experiments and presented as the mean ± SEM. Data were analyzed by two-tailed unpaired t-test (b and d), one-way ANOVA followed by a Tukey’s test (e) and the Wald test in DESeq2, P values adjusted by the Benjamini–Hochberg method (g). Source data are provided as a Source Data file.
To comprehensively assess the impact of EI24 on AMs in more detail, we performed bulk RNA sequencing (RNA-seq) with purified AMs from Lyz2creEi24fl/fl mice and Ei24fl/fl mice. Principal component analysis (PCA) showed a clearly separated distribution of AMs from Ei24fl/fl mice and from Lyz2creEi24fl/fl mice, based on their transcriptomes (Fig. 2f). Differential expression analysis identified 1363 differentially expressed gene transcripts, with 604 genes markedly upregulated and 759 genes downregulated in EI24-deficient AMs (Fig. 2g). Gene Ontology (GO) analysis of differentially expressed genes revealed that upregulated genes were related to antigen presentation, inflammatory response regulation, and protein maturation, while downregulated genes were enriched in mRNA processing and RNA splicing pathways (Fig. 2h). Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment further highlighted upregulation of immune activation and effector function pathways in EI24-deficient AMs (Fig. 2i–l), supporting enhanced activity of these cells. Gene set enrichment analysis (GSEA) corroborated these findings, showing enrichment of pathways linked to receptor-mediated endocytosis, cell killing, and regulation of inflammatory response in EI24-deficient AMs (Fig. 2m–o). These data indicate general inhibitory effects of EI24 on AMs activation and functions.
Notably, EI24 deficiency also altered chromatin accessibility in AMs (Fig. 3a), as detected by assay for transposase-accessible chromatin with high throughput sequencing (ATAC-seq). In total, 2151 genes with different chromatin accessibility were identified in AMs from Lyz2creEi24fl/fl mice in comparison with AMs from Ei24fl/fl mice, and 2093 genes among those displayed increased chromatin accessibility in EI24-deficient AMs (Fig. 3b, c). KEGG pathway enrichment of those upregulated ATAC-seq signals highlighted genes related to phagocytosis, apoptosis, macrophage activation and inflammatory responses (Fig. 3d), in line with transcriptional changes observed in EI24-deficient AMs. Motif analysis further indicated an enrichment of binding motifs for AP-1 family transcription factors, including Fos, Atf3, and JunB (Fig. 3e) in EI24-deficient AMs, which were related to activation and inflammatory responses of macrophages40,41. Notably, there was a general concordance between open chromatin regions and upregulated gene transcripts. In total, 224 genes were upregulated in both RNA-seq and ATAC-seq data, representing approximately 60% of genes with enhanced chromatin accessibility in EI24-deficient AMs (Fig. 3f). Pathway enrichment analysis with these overlapping genes revealed enrichment in apoptosis, activation, and inflammatory response pathways (Fig. 3g). Overall, alterations in chromatin accessibility support the enhanced activity of EI24-deficient AMs.
a PCA plots for ATAC-seq of AMs from Ei24fl/fl and Lyz2creEi24fl/fl mice (n = 3 mice per group). b Volcano plot of differentially accessible regions in AMs from Ei24fl/fl and Lyz2creEi24fl/fl mice (Fold Change ≥ 2 or ≤ 0.5, FDR < 0.05). c ATAC-seq heatmaps and metagene plots depicting the genes with gain or loss of chromatin accessibility in AMs from Ei24fl/fl and Lyz2creEi24fl/fl mice. d KEGG pathway enrichment of ATAC-seq chromatin regions with significant gain of chromatin accessibility in AMs from Lyz2creEi24fl/fl mice versus Ei24fl/fl mice (FDR < 0.05). e Homer known motif analysis showing the top 10 significantly enriched motifs for AMs from Lyz2creEi24fl/fl mice over Ei24fl/fl mice. f Plots showing the concordance between ATAC-seq and RNA-seq data. ATAC-seq peaks were assigned to genes according to their genomic locations and their log2FC and the log2FC of gene expression of the RNA-seq analysis in Lyz2creEi24fl/fl AMs relative to Ei24fl/fl AMs. g KEGG pathway enrichment analysis of genes upregulated in EI24-deficient AMs across RNA-seq and ATAC-seq data.
Lyz2 cre Ei24 fl/fl mice exhibit resistance to viral infection and tumor metastasis in lung
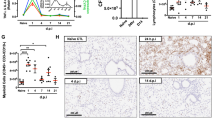
AMs play key roles in lung homeostasis and lung immune defense42. Given the active phenotypes and enhanced functionality of AMs in Lyz2creEi24fl/fl mice, we next explored their lung physiology and lung immunity. Histological analysis revealed no obvious signs of inflammation or injury in the lung (Fig. 4a) or liver (Supplementary Fig. 3) of Lyz2creEi24fl/fl mice in comparison with Ei24fl/fl mice. Consistently, inflammatory cytokine levels in bronchoalveolar lavage fluid (BALF) of Lyz2creEi24fl/fl mice were low and hardly differed from those of Ei24fl/fl mice (Fig. 4b). Furthermore, we performed pulmonary function tests on Lyz2creEi24fl/fl and Ei24fl/fl mice by spirometry, and found that Lyz2creEi24fl/fl mice had increased forced expiratory volumes at 50 ms (FEV50) and forced vital capacity (FVC), and their FEV50/FVC ratio, a key criterion routinely used for chronic obstructive pulmonary disease diagnosis43, remained comparable to Ei24fl/fl mice (Fig. 4c). To evaluate the long-term influence of EI24 deficiency in AMs on lung physiology, we examined lung histology and inflammatory gene expression in 12-month-old Lyz2creEi24fl/fl mice and littermate Ei24fl/fl mice, and found no difference between these mice (Supplementary Fig. 4a, b). Together, these findings demonstrate normal lung function in Lyz2creEi24fl/fl mice under physiological condition.
a Representative lung hispathology from Ei24fl/fl and Lyz2creEi24fl/fl mice. Scale Bar, 50 μm. b Levels of IL-1β, IL-6, and TNF in BALF from Ei24fl/fl and Lyz2creEi24fl/fl mice (n = 12, 12, 8, 11, 10, 9). c Lung function was assessed by measuring FEV50, FVC, and FEV50/FVC (n = 3 per group). d Survival of Ei24fl/fl and Lyz2creEi24fl/fl mice after lethal PR8 infection (n = 8 per group). e, f Body weight changes (e, n = 5, 7), and PR8 loads (f, n = 6 per group) in IAV-infected lungs from Ei24fl/fl and Lyz2creEi24fl/fl mice. g IFNα levels in BALF from IAV-infected Ei24fl/fl and Lyz2creEi24fl/fl mice (n = 11, 12). h, i Survival of IAV-infected Ei24fl/fl, Lyz2creEi24fl/fl, or Lyz2creEi24fl/fl mice with T cell depletion. CD8+ T cell depletion (h, n = 14, 12, 12), CD4+ T cell depletion (i, n = 11, 10, 10). j Schematic of macrophages depletion in IAV-infected mice. k Survival of IAV-infected Ei24fl/fl, Lyz2creEi24fl/fl, or macrophages-depleted Lyz2creEi24fl/fl mice (n = 14, 14, 15). l, m Images of lungs with tumor metastasis (l) and number of tumor nodules (m) in Ei24fl/fl and Lyz2creEi24fl/fl mice (n = 10 per group). n, o Images of lungs with tumor metastasis (n) and number of tumor nodules (o) in Ei24fl/fl and Lyz2creEi24fl/fl mice with or without T cell depletion (n = 5 per group). p, q Images of lungs with tumor metastasis (p) and number of tumor nodules (q) from PBS treated Ei24fl/fl, Lyz2creEi24fl/fl mice and liposome treated Lyz2creEi24fl/fl mice (n = 10 per group). r Schematic of AM phagocytosis of tumor cells. s Flow cytometric analysis of tumor-mCherry cell phagocytosis by AMs from Ei24fl/fl and Lyz2creEi24fl/fl mice (n = 6 per group). Data are representative of (o) or pooled from (b, c, f, g, m, q, s) two to three independent experiments and presented as the mean ± SEM. Data were analyzed using two-tailed unpaired t-test (b, c, f, g, m and s), one-way ANOVA followed by a Tukey’s test (o, q) two-way ANOVA with Šidák’s multiple comparisons test (e), log-rank test (d, h, i and k). Source data are provided as a Source Data file.
When Lyz2creEi24fl/fl mice and Ei24fl/fl mice were infected intranasally (i.n.) with a lethal dose of the A/PR/8/34 (hereafter referred to as PR8) influenza virus strain (IAV), Lyz2creEi24fl/fl mice exhibited a significantly higher survival rate (Fig. 4d). With a sub-lethal dose of PR8 influenza virus challenge, Lyz2creEi24 fl/fl mice displayed reduced body weight loss (Fig. 4e) and lower viral titers in the lung (Fig. 4f), further underscoring their improved resistance to viral infection. In addition to phagocytosis, type I interferons (IFNs) are key molecules for AM-mediated viral control in lungs28,44. Indeed, BALF from PR8-infected Lyz2creEi24fl/fl mice contained higher IFNα levels (Fig. 4g), in comparison with BALF from infected Ei24fl/fl mice. Notably, depleting CD8+ T cells or CD4+ T cells in vivo by administration of either anti-CD8 antibody or anti-CD4 antibody did not impact the survival rate of PR8-infected Lyz2creEi24fl/fl mice (Fig. 4h, i). When AMs and potential monocyte-derived macrophages, including monocyte-derived AMs, in lungs were depleted in PR8-infected Lyz2creEi24fl/fl mice (Fig. 4j), survival of these mice was impaired (Fig. 4k), indicating a key role of macrophages in viral control in lung. Meanwhile, when B16F10 melanoma cells were injected i.v. into Lyz2creEi24fl/fl mice and Ei24fl/fl mice, Lyz2creEi24fl/fl mice exhibited resistance to tumor metastasis in lungs, as indicated by reduced numbers of B16F10 tumor nodules in lung lobes (Fig. 4l, m). Again, T cell depletion did not influence the lung metastasis of B16F10 tumor cells in Lyz2creEi24fl/fl mice (Fig. 4n, o), and AMs and potential monocyte-derived macrophages depletion increased tumor metastasis in lungs of Lyz2creEi24fl/fl mice (Fig. 4p, q). Previous studies demonstrate that macrophages could directly clear tumor cells via phagocytosis45,46. Here, we showed that EI24-deficient AMs displayed significantly increased phagocytosis of mCherry-expressing tumor cells in vitro, suggesting an enhanced capacity to directly eliminate tumor cells (Fig. 4r, s).
Commensal microbiota upregulates EI24 expression in AMs via TLR signaling
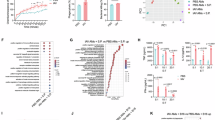
AMs locate in alveolar airspace, where the microenvironment is different from aforementioned other tissues such as spleens and livers. Given the findings that EI24 was highly expressed in AMs but not in RPMs or liver macrophages, it is possible that “open” tissue microenvironment in lungs upregulates EI24 expression. In addition to lung, gut is another “open” tissue in body which is in close contact with microbiota. Notably, macrophages in gut lamina propria displayed higher EI24-GFP expression in comparison with liver macrophages and RPMs in Ei24-Gfp mice (Supplementary Fig. 5a, b), and EI24 deficiency reduced the frequencies of these lamina propria macrophages (Supplementary Fig. 5c). These results imply that microbiota upregulates EI24 expression in macrophages. Next, we investigated the influence of commensal microbiota on EI24 expression. We found that AMs from germ-free (GF) mice showed lower Ei24 mRNA levels than AMs from specific pathogen-free (SPF) mice (Fig. 5a). Consistently, in GF mice with low EI24 expression, EI24 deficiency showed no influence on abundance or phagocytosis of AMs under physiological condition (Fig. 5b, c). When adult GF mice were transferred to SPF environment to reconstitute microbiota to the level of SPF mice (Supplementary Fig. 6), these GF-to-SPF mice successfully upregulated Ei24 expression in AMs (Fig. 5d). These data suggest that commensal microbiota increases Ei24 expression in AMs. To determine whether this effect is mediated by microbial metabolites, we treated AMs with short-chain fatty acids (SCFAs). However, Ei24 expression was not significantly altered (Supplementary Fig. 7). Microbiota derived molecules could modulate macrophage fate and functions through TLR signaling27,47,48. Here, we stimulated sorted AMs from Ei24-Gfp mice with TLR4 agonist LPS or the TLR2 agonist Pam3CSK4 in vitro, and found that both stimuli upregulated GFP expression (Fig. 5e, f). To further validate the role of TLRs, mice were administrated with OxPAPC, an inhibitor of TLR2 and TLR4, and it successfully suppressed the expression of EI24 in AMs at both mRNA level (Fig. 5g) and protein level (Fig. 5h). These results indicate that commensal microbiota upregulates EI24 expression in AMs through TLR2/4 signaling.
a Ei24 expression in AMs sorted from SPF and GF mice (n = 6 per group). b The frequencies (left) and absolute numbers (right) of AMs in GF Ei24fl/fl (n = 7) and Lyz2creEi24fl/fl (n = 9) mice. c Summarized data showing phagocytosis of pHrodo-Green-labeled E.coli by AMs from GF Lyz2creEi24fl/fl and GF Ei24fl/fl mice (n = 8 per group). d Expression of Ei24 in AMs from GF-to-SPF (n = 13) and GF (n = 11) mice. e, f Expression of Ei24 in AMs stimulated with or without LPS (e) or Pam3CSK4 (f) (n = 6 per group). g Schema of PBS or OxPAPC administration and analysis of EI24 expression in WT and Ei24-Gfp mice. h, i Ei24 mRNA level in AMs from WT mice (h, n = 5 per group) and EI24-GFP protein level in AMs from Ei24-Gfp mice (i, n = 4 per group) after PBS or OxPAPC administration. Data are representative of (h, i) or pooled from (a–f) two independent experiments and presented as the mean ± SEM. Data were analyzed using two-tailed unpaired t-test. Source data are provided as a Source Data file.
EI24 deficiency rewires cellular metabolism in AMs to support their enhanced functionality and apoptosis
Cellular metabolism is recognized as a key regulator of immune cell function49. GSEA enrichment analysis and gene expression heatmaps revealed elevated glycolysis and tricarboxylic acid (TCA) cycle in EI24-deficient AMs (Fig. 6a–c). We next assessed the metabolic profile of sorted AMs from Lyz2creEi24fl/fl and Ei24fl/fl mice via metabolic flux assay. In comparison with Ei24fl/fl AMs, EI24-deficient AMs exhibited higher extracellular acidification rate (ECAR) (Fig. 6d), glycolysis, glycolytic capacity (Fig. 6e), and 2-NBDG uptake (Fig. 6f), and increased mitochondrial oxygen consumption rate (OCR) (Fig. 6g), basal respiration, maximal respiration and spare respiratory capacity (SRC) (Fig. 6h). These results demonstrate that EI24 deficiency in AMs causes metabolic reprogramming and increases both glycolysis and oxidative phosphorylation (OXPHOS). Moreover, we found that blocking glycolysis with UK5099 or glutaminolysis with BPTES markedly reduced OXPHOS in EI24-deficient AMs, whereas inhibition of fatty acid oxidation with etomoxir had no effect (Fig. 6g, h). These results indicate that glucose and amino acids rather than fatty acids serve as the primary substrate fueling the increased OXPHOS in EI24-deficient AMs.
a GSEA analysis of gene transcripts related to glycolysis and Citrate cycle in Lyz2creEi24fl/fl AMs versus Ei24fl/fl AMs. b, c RNA-seq heatmaps depict the genes related to the glycolysis (b) and TCA cycle (c) pathways. d ECAR of Lyz2creEi24fl/fl and Ei24fl/fl AMs at resting state (n = 4 per group). e Statistical analysis of glycolysis and glycolytic capacity (n = 8 per group). f Uptake of 2-NBDG by AMs from Lyz2creEi24fl/fl mice (n = 6) and Ei24fl/fl mice (n = 8). g OCR of Ei24fl/fl and Lyz2creEi24fl/fl AMs treated with vehicle or Etomoxir, UK5099, or BPTES (n = 5, 6, 6, 6, 6). h Quantification of basal respiration, maximal respiration, and Spare Respiration Capacity (SRC) of Ei24fl/fl AMs treated with vehicle, Lyz2creEi24fl/fl AMs treated with vehicle or Etomoxir, UK5099, or BPTES (basal respiration, n = 10, 10, 10, 10, 9; maximal respiration, n = 10 per group; SRC, n = 10 per group). i IL-6 production in Lyz2creEi24fl/fl AMs left unstimulated, stimulated with LPS alone, or stimulated with LPS in the presence of 2-DG (n = 6, 6, 8). j IL-6 production in Lyz2creEi24fl/fl AMs left unstimulated, stimulated with LPS alone, or stimulated with LPS in the presence of Oligomycin (n = 7, 6, 8). k Phagocytosis of Lyz2creEi24fl/fl AMs treated with vehicle CTRL, 2-DG or Oligomycin (n = 3 per group). l MFI of MitoSOX in AMs from Ei24fl/fl mice and Lyz2creEi24fl/fl mice (n = 6 per group). m MFI of MitoSOX in Lyz2creEi24fl/fl AMs treated with vehicle, 2-DG, Oligomycin, or mitoTEMPO (n = 8, 6, 6, 6). n Percentage of Cleaved Caspase-3+ Lyz2creEi24fl/fl AMs left alone or exposure to 2-DG, Oligomycin, mitoTEMPO or Z-VAD-FMK (n = 11 9, 9, 8, 9). Data are representative of (d, g, k) or pooled from (e, f, h–j, l–n) two to three independent experiments and presented as the mean ± SEM. Data were analyzed by two-tailed unpaired t-test (e, f, and l), and one-way ANOVA (h–k, m, n). Source data are provided as a Source Data file.
To examine whether altered metabolism contributes to enhanced EI24-deficient AM function and impaired homeostasis, we stimulated sorted EI24-deficient AMs with LPS in the presence of glycolysis inhibitor 2-DG or OXPHOS inhibitor oligomycin. Both of treatments reduced proinflammatory cytokine IL-6 production (Fig. 6i, j), whereas only OXPHOS inhibition suppressed the phagocytic activity in EI24-deficient AMs (Fig. 6k). It has been reported that OXPHOS contributes to the mitochondrial ROS (mtROS) production50, which is known to induce Caspase-3 activation and thus cell apoptosis51,52. Indeed, we detected increase of mtROS in EI24-deficient AMs in comparison with Ei24fl/fl AMs (Fig. 6l). Treatment with 2-DG or Oligomycin efficiently reduced mtROS production in EI24-deficient AMs as same as mtROS scavenger mitoTEMPO did (Fig. 6m). Furthermore, 2-DG, Oligomycin and mitoTEMPO all inhibited Caspase-3 activation in EI24-deficient AMs as efficiently as Caspase inhibitor Z-VAD-FMK did (Fig. 6n). Together, these data indicate that EI24 deficiency enhances glycolysis and OXPHOS in AMs, and this rewired metabolism supports their enhanced inflammatory response, phagocytic activity, and apoptosis.
EI24 has been previously reported to regulate autophagy35,36. Here, we found that EI24-deficient AMs exhibited impaired autophagy, as indicated by increased p62 accumulation and a reduced LC3-II/I ratio (Supplementary Fig. 8a). However, rapamycin administration in Lyz2creEi24fl/fl mice promoted autophagy in EI24-deficient AMs but failed to reverse those phenotypes, including cell frequencies, active state, phagocytic capacity, and IL-6 production (Supplementary Fig. 8b–f). These results exclude the possibility that EI24 deficiency changes AM function and fate via downregulating autophagy.
Deleting EI24 enhances efficacy of BMDMs in preventing tumor metastasis and influenza infection in lung
In addition to AMs, we found that in vitro LPS stimulation upregulated Ei24 levels in other macrophages, such as peritoneal macrophages, RPMs, and liver macrophages (Supplementary Fig. 9a), implying a role of EI24 in controlling their function after activation. Indeed, we confirmed that deficiency of EI24 also enhanced LPS induced inflammatory cytokine response in liver macrophages, peritoneal macrophages, and BMDMs (Supplementary Fig. 9b, c, Fig. 7a). RNA-seq analysis on Ei24fl/fl control BMDMs and Lyz2creEi24fl/fl BMDMs displayed a distinct gene expression profile associated with EI24 deficiency (Fig. 7b). KEGG enrichment analysis of upregulated genes in EI24-deficient BMDMs highlighted pathways associated with phagocytosis, antigen presentation, cytokine interaction and activation, confirming augmented defensive function in these cells (Fig. 7c). Moreover, EI24-deficient BMDMs exhibited increased glycolysis, glycolytic capacity, maximal respiration, and spare respiratory capacity, in comparison with Ei24fl/fl BMDMs (Supplementary Fig. 10a–d), proving the EI24 deficiency-caused metabolic reprogramming in BMDMs. Similar to AMs, inhibition of glycolysis or OXPHOS resulted in decreased production of the proinflammatory cytokine IL-6 in EI24-deficient BMDMs (Supplementary Fig. 10e, f). These results demonstrate a general role of EI24 in inhibiting macrophage function via metabolic control. Given the potential of macrophages in immunotherapy, we evaluated the effect of deleting EI24 on efficacy of macrophage-based therapies. With B16F10 tumor metastasis mouse models and PR8 infection mouse models, we showed that EI24-deficient BMDMs better prevented tumor metastases (Fig. 7d, e) and viral infection (Fig. 7f, g) in lungs than control BMDMs did. Overall, these data suggest that EI24 deficiency enhances efficacy of macrophage-based therapies against tumors and viral infection.
a IL-1β (left) and IL-6 (right) production in BMDMs from Ei24fl/fl untreated, Lyz2creEi24fl/fl untreated, Ei24fl/fl stimulated with LPS and ATP, and Lyz2creEi24fl/fl stimulated with LPS and ATP (n = 9, 9, 8, 9). b Heatmap of mRNA transcripts differentially expressed in BMDMs of Lyz2creEi24fl/fl and Ei24fl/fl mice. c KEGG pathway enrichment of upregulated genes in Lyz2creEi24fl/fl versus Ei24fl/fl mice derived BMDMs. d Images of lungs with tumor metastasis in WT mice transferred with PBS or BMDMs from Ei24fl/fl mice or Lyz2creEi24fl/fl mice. e Lung weight of B16F10-inoculated mice transferred with PBS or BMDMs from Ei24fl/fl or Lyz2creEi24fl/fl mice (n = 5 per group). f Survival curve of IAV infected mice with BMDMs transfer (n = 7, 8, 7 per group). g PR8 loads in IAV-infected lungs from mice that received BMDMs from different groups were assessed by RT-PCR (n = 5 per group). Data are representative of (d–g) or pooled from (a) two independent experiments. Data are presented as the mean ± SEM. Data were analyzed by one-way ANOVA (e, g) and two-way ANOVA with Šidák’s multiple comparisons test (a). Source data are provided as a Source Data file.
Discussion
AMs as major immune cells in lung play important roles in tissue homeostasis and immune defense42. Here, we showed that EI24 did not influence the development of AMs but was essential for their survival. EI24 deficiency led to caspase activation in AMs by increasing OXPHOS and mtROS production, and that caused cell death. Notably, EI24 deficiency did not influence macrophage death in other tissues like liver and spleen. In support of these results, we found that AMs expressed high levels of EI24, whereas liver macrophages and red pulp macrophages expressed low levels. Different from other tissues, lung is exposed to air, and AMs are exposed to airborne stimuli. In GF mice, Ei24 level was low in AMs and was dispensable for survival of AMs. We further proved that commensal microbiota upregulated EI24 in AMs through TLR2/4 signaling. It is likely that AMs are exposed to exogenous stimuli in the “open” microenvironment in lung, and EI24 upregulated by microbiota helps to inhibit Caspase-3 activation and maintain AM survival in this stress condition. In addition to lung, gut is also an “open” tissue with exposure to microbiota. Consistently, EI24 is expressed in lamina propria macrophages in gut and is required for maintaining their homeostasis.
Despite the impaired cell homeostasis, deficiency of EI24 enhanced phagocytosis and inflammatory response of AMs and improved their immune surveillance function. Given the fact that microbiota increased EI24 expression, it is rational that microbiota educates AMs to avoid excessive activation in the “open” microenvironment, and as a consequence dampens their immune surveillance function. Our finding is in agreement with published studies which demonstrate the role of microbiota in inhibiting AMs response and inducing innate immune tolerance29,31. Our study demonstrated that microbiota induced TLR2/4 signaling in physiological condition elevated EI24 expression in AMs, linking the TLR signaling to EI24 mediated macrophage education. In support of our findings, previous studies have shown that LPS stimulation promotes p53 expression in AMs53 which could induce Ei24 expression33,54. Meanwhile, strong TLR2/4 signaling activates macrophages. It is rational that signaling threshold of TLRs determine their influences on AMs. On the other hand, several studies have reported that microbiota could elevate immune surveillance function of AMs30,55,56. Except for TLR signaling that have been reported to induce trained immunity in AMs, metabolites from microbiota also enhance functions of AMs57,58. It is possible that microbiota composition determines signaling types and threshold in AMs, which finely tune their phenotype and function.
In our study, EI24 regulated function of AMs via metabolic control. Although glycolysis and mitochondrial OXPHOS are previously linked to proinflammatory M1 macrophage and anti-inflammatory M2 macrophage respectively59,60, our results, in line with previous study on trained immunity59, demonstrated that the elevated glycolysis and mitochondrial function in EI24-deficient AMs both supported their increased inflammatory responses. These results were in agreement with our findings that OXPHOS in EI24-deficient AMs was fueled by both glycolysis and glutaminolysis. Notably, our results also indicated that phagocytosis of AMs was controlled by mitochondrial OXPHOS, but not glycolysis, deciphering the distinct dependences of AMs functions on metabolic pathways. In line with the crosstalk between metabolism and epigenetics, we also detected epigenetic alteration in EI24-deficient AMs, and that supported their enhanced functionality.
Despite the low expression of EI24 in RPMs, liver macrophages, and peritoneal macrophages at steady state, these macrophages upregulated Ei24 upon activation by LPS, enabling EI24 to control their function. Indeed, our data demonstrate that deficiency of EI24 generally enhances function of different macrophages. Recently, macrophage-based cell therapy has drawn attention, particularly the genetically engineered chimeric antigen receptor-macrophages61. Although EI24 deficiency enhanced defensive functions of AMs, in steady state these cells did not induce lung damage or impair lung function. This result indicates safety of using EI24-deficient macrophages in cell therapy. Here, we showed that deletion of EI24 significantly enhanced anti-viral and anti-tumor efficacy of transferred BMDMs. Notably, macrophages are plastic and typically adopt pro-tumor M2 phenotypes due to the metabolic reprogramming in tumor microenvironment57,62. EI24 deficiency forced metabolic and epigenetic reprogramming which favored anti-tumor effects of macrophages, and that likely would overcome the major hurdle of macrophage-based anti-tumor therapy. Overall, our results propose a new direction to augment the efficacy of macrophage-based cell therapy.
Methods
Mice models
C57BL/6 micewere purchased from the Gempharmatech co., Ltd (stock number: N000013). Cd11ccre mice and Lyz2cre mice have previously been described63 and were provided by Dr Albert Bendelac. Ei24fl/fl mice35 were kindly provided by Dr.Hong Zhang. To generate conditional knockout mice, Ei24fl/fl mice were crossed with mice expressing Cre recombinase. Ei24-Gfp reporter mice were generated at the Laboratory Animal Center of the University of Science and Technology of China using a CRISPR/Cas9-mediated strategy to insert a 3×(GGGGS)-EGFP cassette immediately upstream of the Ei24 stop codon. All mice were on the C57BL/6 background. Mice were housed under specific pathogen-free conditions with 12 h dark/12 h light cycle. Ambient temperature was maintained at 23 °C with 50% humidity. Mice had ad libitum access to food and water. GF mice on a C57BL/6 background were bred and housed in flexible film isolators. Experimental and control mice were co-housed throughout the study. For experiments using adult mice, 4–12-week-old male and female mice were used. For neonatal experiments, 2-day-old mie were used. For aging experiments, 12-month-old mice were used. Animals were grouped randomly in roughly equal numbers within each experiments set, and both sexes were included when appropriate. The number of mice used in each experiment is indicated in the respective figure legends. All experimental procedures involving mice were approved by the Ethics Committee of University of Science and Technology of China. All experiments were conducted in accordance with institutional ethical guidelines. For lung metastasis model, mice were monitored daily and euthanized with cervical dislocation if they lost more than 15% of their initial body weight, developed severe respiratory distress, or exhibited other signs of distress. For influenza virus infection experiments, two types of endpoints were used depending on the experiment. In survival studies, mice were monitored daily and euthanized if they reached humane endpoint criteria, such as sustained weight loss >15%, severe lethargy, or other signs of distress, in accordance with institutional guidelines. In viral load studies, mice were sacrificed at 7 days post-infection to collect lungs for analysis. Euthanasia was performed using cervical dislocation.
For influenza infection studies, mice were anesthetized and infected i.n. with the A/PR/8/34 influenza virus strain (H1N1, PR8; lethal dose, 8 HA; sublethal dose, 4 HA) in 50 μL sterile saline. For tumor metastasis models, 5 × 105 B16F10 cells were resuspended in 100 μL phosphate-buffered saline (PBS) and intravenously injected to establish lung melanoma. In adoptive transfer experiments, BMDMs were collected and resuspended in 100 μL PBS and administered into recipient mice.
To inhibit caspase activation, Z-VAD-FMK was administered i.p. at 5 mg/kg every 2 days for 1 week64. To activate autophagy, mice were administered rapamycin (75 mg/kg) or vehicle i.p. daily for 2 weeks65. To block TLR2 and TLR4 mice were administered i.p. with OxPAPC (2 mg/kg) at day 0 and 366.
Pulmonary function test
Spirometry was performed on Ei24fl/fl and Lyz2creEi24fl/fl mice previously described67. Briefly, mice were anesthetized by intraperitoneal injection of 100 mg/kg of body weight of pentobarbital sodium, and tracheotomy was performed. The mice were intubated with a cannula and placed in a whole-body plethysmograph connected to the eSpira Forced Maneuvers System (EMMS).
Cell isolation, culture, and activation
To isolate peritoneal macrophages, mice were euthanized by cervical dislocation and i.p. with 1 mL PBS, and peritoneal macrophages were harvested by collecting the buffer injected into the peritoneal cavity. To isolate AMs, lungs were cut into small pieces, followed by a digestion in collagenase type IV (2 mg/mL, Sigma, C5138) in Dulbecco’s modified Eagle’s medium (DMEM, Gibco, 12100046) at 37 °C and 150 rpm for 40 min. After digestion, the samples were homogenized through a 100 μm cell strainer and washed, red blood cells were lysed with Erythrocytes Lysate (Biosharp, BL503A), and the samples were washed and then used for staining.
Murine BMDMs were obtained as previously described68, tibiae and femurs were aseptically isolated and bone marrow cells were flushed out with DMEM supplemented with 10% fetal bovine serum (FBS, VivaCell, C2910-0500). BMDMs were differentiated from bone marrow cells in DMEM supplemented with 10% FBS, 50 μM β-mercaptoethanol (Sigma-Aldrich, 444203), 100 U mL−1 penicillin (Sigma, P4333), 100 μg ml−1 streptomycin (Sigma, P4333) and 50 μg ml−1 M-CSF (Novus biologicals, 416-ML).
For ex vivo stimulations of AMs and BMDMs, cells were stimulated with LPS at 50 ng ml−1 and incubated for 16 h (AMs) or 4 h (BMDMs) before supernatants were collected. In some experiments, Z-VAD-FMK (5 μM, MCE, HY-16658B), rapamycin (100 nM, MCE, HY-10219), 2-DG (2 mM, MCE, HY-13966), Oligomycin (50 nM, MCE, HY-N6782), Mito-TEMPO (10 μM, MCE, HY-112879), and BPTES (10 μM, MCE, HY-12683) were added to sorted AMs.
Antibodies and flow cytometry
The fluorochrome-labeled or unlabeled antibodies for flow cytometry analysis including mouse anti-CD16/32 (93), CD45.2 (104), anti-CD11b (M1/70), anti-CD11c (N418), anti-F4/80 (BM8), anti-SiglecF (S17007L), anti-IA/IE (M5/114.15.2), anti-Ly6G (10-3.6), anti-TCRβ (H57.597), anti-CD4 (RM4-5), anti-CD8α (53-6.7), anti-NK1.1 (PK136), and anti-rabbit IgG (Poly4064) were purchased from Biolegend.
Cleaved Caspase-3 Rabbit mAb (#9664), β-Actin Mouse mAb (#3700), EI24 Rabbit mAb (#42328), SQSTM1/p62 Rabbit mAb (#23214), LC3A/B Rabbit mAb (#12741), Cleaved-IL-1β Rabbit mAb (#63124), and IL-6 XP Rabbit mAb (#12912) were purchased from Cell Signaling Technology and were used in our experiments.
For apoptosis analysis, lung single-cell suspensions were isolated and resuspended in PBS, and incubated at 4 °C with Fc receptor-blocking antibody for 15 min. Subsequently, cells were stained with surface marker antibodies for 45 min on ice. Cell death was analyzed later by FITC Annexin V Apoptosis Detection Kit with PI (Biolegend, 640914). For phagocytosis analysis, lung cells were resuspended in 100 μL antibiotic-free complete DMEM and seeded in a 96-well tissue culture plate. Bioparticles were reconstituted as indicated in the pHrodo Green E.coli Bioparticles Phagocytosis Kit (Thermo Fisher Scientific, P35366), and were added to cells, followed by incubation at 37 °C for 2 h with gentle mixing every 15 min. Phagocytosis by lung macrophages was determined by flow cytometer. To measure mtROS, cells were incubated with MitoSOX Red (2 μM) at 37 °C for 30 min. Cytokine levels were assessed using a Cytometric Bead Array Mouse Th1/Th2/Th17 Flex Set (BD, 560485) according to the manufacturer’s instructions. IFNα in BALF from IAV-infected mice was measured by Legendplex Anti-Virus Response Panel (Biolegend, 740621). Flow cytometry analyses were performed on FACSVerse (BD) or CytoFLEX S (Beckman Coulter) flow cytometer. Flow data were analyzed using FlowJo software.
Tissue morphometry
Immediately following mice were euthanized, the whole lung was dissected, and inflated by gravity with 4% paraformaldehyde overnight. The samples were embedded into paraffin and cut into sections. Lung sections were stained with hematoxylin and eosin.
In vivo cell depletion
Mice were first administered i.t. with 100 μL Clodronate or Control liposomes (Liposoma Technology) on day −1 to deplete local AMs, and were subsequently injected i.v. with 100 μL Clodronate or Control liposomes at day 1 and with 5-day intervals to deplete potential monocyte-derived macrophages in lung69. To deplete T cells in vivo, mice were injected i.p. with 200 μg anti-CD4 (clone: GK1.5, Selleck) and 200 μg anti-CD8 (clone: 2.43, Selleck) at 7-day intervals.
Metabolic assays
Real-time cell metabolism of macrophages was measured by using the Seahorse XF Cell Mito stress test kit or a Seahorse XF glycolytic stress test kit (Agilent Technologies) according to the manufacturer’s instructions. Purified macrophages were seeded into XF 96-well plates (Agilent Technologies) at 75,000 cells per well and cultured overnight in complete DMEM medium. The next day, macrophages were washed and incubated for 1 h in Seahorse assay medium (Agilent Technologies) and supplemented with either 2 mM glutamine (glycolytic stress test) or an addition 10 mM glucose and 1 mM pyruvate (mito stress test). The kit compounds were resuspended in assay medium, and the final concentrations of the compounds in culture wells were 1.5 μM oligomycin, 2 μM FCCP and 0.5 μM rotenone in the mito stress test or 10 mM glucose, 1 μM oligomycin and 50 mM 2-DG in the glycolytic stress test. For XF substrate oxidation stress test, 2 μM UK5099, 3 μM BPTES, 4 μM Etomoxir, and medium only were injected to inhibit glycolysis, glutaminolysis, or fatty acid oxidation, respectively. OCRs and ECARs were measured on a Seahorse XFe 96 analyzer (Agilent Technologies), and data were analyzed using Wave Desktop software (v2.6).
Real-time quantitative PCR with reverse transcription (RT-PCR)
Total RNA was extracted from lung tissues or cells using TRIzol reagent (Thermo Fisher). RT-PCR was performed by using a 2× SYBR Green Fast qPCR Mix for RT-PCR, according to the manufacturer’s instructions (Abclonal). See Supplementary Table 1 for primer sequences.
RNA-seq and data analysis
Bulk RNA was collected from purified lung AMs (viable, CD45+CD11bloCD11c+F4/80+SiglecF+ population) from Lyz2creEi24fl/fl mice and Ei24fl/fl control littermates. Then libraries with different indexes were multiplexed and loaded on an Illumina Novaseq instrument for sequencing using a 2 × 150 paired-end (PE) configuration according to manufacturer’s instructions.
In order to remove technical sequences, including adapters, polymerase chain reaction (PCR) primers, or fragments and quality of bases lower than 20, pass filter data of raw RNA-seq fastq format were processed by Cutadapt (v1.9.1, phred cutoff: 20, error rate: 0.1, adapter overlap: 1 bp, min.length: 75, proportion of N: 0.1) to be high quality clean data. Reads were then mapped to mm10 RefSeq genes downloaded from the University of California Santa Cruz Table Browser website. Hisat2 (v2.2.1) was used to index reference genome sequence and align clean data to reference genome. With the file as a reference gene file, HTSeq (v0.6.1) estimated gene and isoform expression levels from the pair-end clean data. Differential expression analysis used the DESeq2 Bioconductor package, a model based on the negative binomial distribution. The resulting P values were adjusted using the Benjamini–Hochberg approach for controlling the false discovery rate (FDR). The RNS-seq was performed by Genewiz (Azenta Life Sciences).
ATAC-seq and data analysis
A total of 50,000 purified AMs per replicate were washed once in cold PBS by centrifugation at 500 × g for 5 min at 4 °C. Post washing, the cell pellets were gently resuspended in 50 μL of resuspension buffer (10 mM Tris-HCl (pH 7.4), 10 mM NaCl, 3 mM MgCl2 and 0.1% IGEPAL CA-630, 0.1% Tween-20, 0.01% Digitonin). The supernatants were discarded, and the nuclei were resuspended in 50 μL of transposition reaction mix (25 μL 2 × TD buffer, 2.5 μL Illumina Tn5 transposase and 22.5 μL nuclease-free water), and incubated at 37 °C on a thermomixer set at 1000 rpm for 30 min. The transposed DNA was purified using a Qiagen MinElute Reaction Cleanup Kit (Qiagen, 28204) and eluted in a 10 μL volume of elution buffer. The transposed DNA was amplified using NEBNext High-Fidelity 2 × PCR Master Mix (New England Biolabs, M0541S) and unique ATAC indexing PCR primers. Post-amplification, the DNA fragments size were verified. The amplified DNA libraries were quantified by quantitative PCR. Paired-end sequencing was performed on an Illumina Novaseq6000 system.
In order to remove technical sequences, including adapters, polymerase chain reaction (PCR) primers, or fragments and quality of bases lower than 20, pass filter data of fastq format were processed by cutadapt (v1.9.1) to be high quality clean data. Clean data were aligned to the mm10 reference genome using bowtie2 (v2.2.6). Samtools (v1.18) was used to remove unmapped, unpaired and mitochondrial reads. PCR duplicates were removed by using sbamba (v1.0.0). Peak calling was performed using MACS2 (2.2.9.1). For each experiment, peaks of all samples were combined to create a union peak list, and overlapping peaks were merged using bedtools (v2.31.1) merge. The number of reads in each peak and differentially accessible peaks were analyzed using R package DiffBind. The tracks were visualized using Integrative Genomics Viewer (v2.16.0). Annotation of peaks relative to genomic features was done using R package CHIPseeker and HOMER.
Data and statistical analysis
The same samples were not repeatedly measured, and no data points were excluded from the analysis. No statistical method was used to predetermine sample size. Results are represented as the mean ± SEM. Statistical analyses were performed using GraphPad Prism v9.5.0 software (GraphPad). Statistical differences were determined using unpaired two-tailed t-test, one-way ANOVA, two-way ANOVA, and log-rank test (survival studies). The confidence interval was set at 95% for statistical analysis.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The RNA-seq and ATACs-seq sequencing data generated in this study have been deposited in the National Center for Biotechnology Information Sequence Read archive (NCBI SRA) under accession code PRJNA1187783. The RNA-seq and ATACs-seq sequencing data available. All data are included in the Supplementary Information or available from the authors, as are unique reagents used in this article. The raw numbers for charts and graphs are available in the Source Data file whenever possible. Source data are provided with this paper.
References
Hashimoto, D. et al. Tissue-resident macrophages self-maintain locally throughout adult life with minimal contribution from circulating monocytes. Immunity 38, 792–804 (2013).
Westphalen, K. et al. Sessile alveolar macrophages communicate with alveolar epithelium to modulate immunity. Nature 506, 503–506 (2014).
Yona, S. et al. Fate Mapping reveals origins and dynamics of monocytes and tissue macrophages under homeostasis. Immunity 38, 79–91 (2013).
Mass, E., Nimmerjahn, F., Kierdorf, K. & Schlitzer, A. Tissue-specific macrophages: how they develop and choreograph tissue biology. Nat. Rev. Immunol. 23, 563–579 (2023).
Trapnell, B. C. & Whitsett, J. A. Gm-CSF regulates pulmonary surfactant homeostasis and alveolar macrophage-mediated innate host defense. Annu Rev. Physiol. 64, 775–802 (2002).
Bhoj, V. G. et al. MAVS and MyD88 are essential for innate immunity but not cytotoxic T lymphocyte response against respiratory syncytial virus. Proc. Natl. Acad. Sci. USA 105, 14046–14051 (2008).
Morris, D. G. et al. Loss of integrin alpha(v)beta6-mediated TGF-beta activation causes Mmp12-dependent emphysema. Nature 422, 169–173 (2003).
Fernandez, S., Jose, P., Avdiushko, M. G., Kaplan, A. M. & Cohen, D. A. Inhibition of IL-10 receptor function in alveolar macrophages by Toll-like receptor agonists. J. Immunol. 172, 2613–2620 (2004).
Roy, M. G. et al. Muc5b is required for airway defence. Nature 505, 412–416 (2014).
Fujimori, T. et al. The Axl receptor tyrosine kinase is a discriminator of macrophage function in the inflamed lung. Mucosal Immunol. 8, 1021–1030 (2015).
Schneider, C. et al. Induction of the nuclear receptor PPAR-gamma by the cytokine GM-CSF is critical for the differentiation of fetal monocytes into alveolar macrophages. Nat. Immunol. 15, 1026–1037 (2014).
Guilliams, M. et al. Alveolar macrophages develop from fetal monocytes that differentiate into long-lived cells in the first week of life via GM-CSF. J. Exp. Med. 210, 1977–1992 (2013).
Yu, X. et al. The cytokine TGF-beta promotes the development and homeostasis of alveolar macrophages. Immunity 47, 903–912 e904 (2017).
Hussell, T. & Bell, T. J. Alveolar macrophages: plasticity in a tissue-specific context. Nat. Rev. Immunol. 14, 81–93 (2014).
Izquierdo, H. M. et al. Von Hippel-Lindau protein is required for optimal alveolar macrophage terminal differentiation, self-renewal, and function. Cell Rep. 24, 1738–1746 (2018).
Woods, P. S. et al. HIF-1alpha induces glycolytic reprograming in tissue-resident alveolar macrophages to promote cell survival during acute lung injury. Elife 11, e77457 (2022).
Woods, P. S. et al. Tissue-resident alveolar macrophages do not rely on glycolysis for LPS-induced inflammation. Am. J. Respir. Cell Mol. Biol. 62, 243–255 (2020).
Roiniotis, J. et al. Hypoxia prolongs monocyte/macrophage survival and enhanced glycolysis is associated with their maturation under aerobic conditions. J. Immunol. 182, 7974–7981 (2009).
Wculek, S. K., Dunphy, G., Heras-Murillo, I., Mastrangelo, A. & Sancho, D. Metabolism of tissue macrophages in homeostasis and pathology. Cell Mol. Immunol. 19, 384–408 (2022).
Caputa, G., Castoldi, A. & Pearce, E. J. Metabolic adaptations of tissue-resident immune cells. Nat. Immunol. 20, 793–801 (2019).
Scur, M. et al. Alveolar macrophage metabolic programming via a C-type lectin receptor protects against lipo-toxicity and cell death. Nat. Commun. 13, 7272 (2022).
Johnson, P., Arif, A. A., Lee-Sayer, S. S. M. & Dong, Y. Hyaluronan and its interactions with immune cells in the healthy and inflamed lung. Front Immunol. 9, 2787 (2018).
Svedberg, F. R. et al. The lung environment controls alveolar macrophage metabolism and responsiveness in type 2 inflammation. Nat. Immunol. 20, 571–580 (2019).
Snelgrove, R. J. et al. A critical function for CD200 in lung immune homeostasis and the severity of influenza infection. Nat. Immunol. 9, 1074–1083 (2008).
Phelps, D. S., Umstead, T. M., Quintero, O. A., Yengo, C. M. & Floros, J. In vivo rescue of alveolar macrophages from SP-A knockout mice with exogenous SP-A nearly restores a wild type intracellular proteome; actin involvement. Proteome Sci. 9, 67 (2011).
Haczku, A. Protective role of the lung collectins surfactant protein A and surfactant protein D in airway inflammation. J. Allergy Clin. Immunol. 122, 861–879; quiz 880-861 (2008).
Wang, J. et al. Bacterial colonization dampens influenza-mediated acute lung injury via induction of M2 alveolar macrophages. Nat. Commun. 4, 2106 (2013).
Ngo, V. L. et al. Intestinal microbiota programming of alveolar macrophages influences severity of respiratory viral infection. Cell Host Microbe 32, 335–348.e338 (2024).
Marquant, Q. et al. The microbiota plays a critical role in the reactivity of lung immune components to innate ligands. FASEB J. 35, e21348 (2021).
Sencio, V. et al. Gut dysbiosis during influenza contributes to pulmonary pneumococcal superinfection through altered short-chain fatty acid production. Cell Rep. 30, 2934–2947.e2936 (2020).
Novakovic, B. et al. beta-glucan reverses the epigenetic state of LPS-Induced immunological tolerance. Cell 167, 1354–1368.e1314 (2016).
Chakraborty, S. et al. Trained immunity of alveolar macrophages enhances injury resolution via KLF4-MERTK-mediated efferocytosis. J. Exp. Med. 220, e20221388 (2023).
Lehar, S. M. et al. Identification and cloning of EI24, a gene induced by p53 in etoposide-treated cells. Oncogene 12, 1181–1187 (1996).
Tian, Y. et al. C. elegans screen identifies autophagy genes specific to multicellular organisms. Cell 141, 1042–1055 (2010).
Zhao, Y. G. et al. The p53-induced gene Ei24 is an essential component of the basal autophagy pathway. J. Biol. Chem. 287, 42053–42063 (2012).
Yuan, L., Liu, Q., Wang, Z., Hou, J. & Xu, P. EI24 tethers endoplasmic reticulum and mitochondria to regulate autophagy flux. Cell Mol. Life Sci. 77, 1591–1606 (2020).
Yang, W. et al. Prognostic role and immune infiltration characteristics of EI24 in multiple cancer types. Expert Rev. Mol. Diagn. 23, 457–469 (2023).
Choi, J. M., Devkota, S., Sung, Y. H. & Lee, H. W. EI24 regulates epithelial-to-mesenchymal transition and tumor progression by suppressing TRAF2-mediated NF-kappaB activity. Oncotarget 4, 2383–2396 (2013).
Barnes, P. J. Inflammatory mechanisms in patients with chronic obstructive pulmonary disease. J. Allergy Clin. Immunol. 138, 16–27 (2016).
He, Y. Y. et al. The Fra-1: novel role in regulating extensive immune cell states and affecting inflammatory diseases. Front Immunol. 13, 954744 (2022).
Mishra, R. K., Potteti, H. R., Tamatam, C. R., Elangovan, I. & Reddy, S. P. c-Jun Is required for nuclear factor-kappaB-dependent, LPS-stimulated fos-related antigen-1 transcription in alveolar macrophages. Am. J. Respir. Cell Mol. Biol. 55, 667–674 (2016).
Aegerter, H., Lambrecht, B. N. & Jakubzick, C. V. Biology of lung macrophages in health and disease. Immunity 55, 1564–1580 (2022).
Bhatt, S. P. et al. FEV(1)/FVC severity stages for chronic obstructive pulmonary disease. Am. J. Respir. Crit. Care Med. 208, 676–684 (2023).
Kumagai, Y. et al. Alveolar macrophages are the primary interferon-alpha producer in pulmonary infection with RNA viruses. Immunity 27, 240–252 (2007).
Lecoultre, M., Dutoit, V. & Walker, P. R. Phagocytic function of tumor-associated macrophages as a key determinant of tumor progression control: a review. J. Immunother. Cancer 8, e001408 (2020).
Gordon, S. R. et al. PD-1 expression by tumour-associated macrophages inhibits phagocytosis and tumour immunity. Nature 545, 495–499 (2017).
Li, C. et al. Gut microbiota-derived GlcNAc-MurNAc is a TLR4 agonist that protects the host gut. Nat. Commun. 16, 5577 (2025).
Li, Y. et al. Gut microbiota dependent anti-tumor immunity restricts melanoma growth in Rnf5(-/-) mice. Nat. Commun. 10, 1492 (2019).
Jung, J., Zeng, H. & Horng, T. Metabolism as a guiding force for immunity. Nat. Cell Biol. 21, 85–93 (2019).
Nolfi-Donegan, D., Braganza, A. & Shiva, S. Mitochondrial electron transport chain: oxidative phosphorylation, oxidant production, and methods of measurement. Redox Biol. 37, 101674 (2020).
An, H. et al. Tetraarsenic hexoxide enhances generation of mitochondrial ROS to promote pyroptosis by inducing the activation of caspase-3/GSDME in triple-negative breast cancer cells. Cell Death Dis. 12, 159 (2021).
Redza-Dutordoir, M. & Averill-Bates, D. A. Activation of apoptosis signalling pathways by reactive oxygen species. Biochim. Biophys. Acta 1863, 2977–2992 (2016).
Gong, L. et al. Nuclear SPHK2/S1P induces oxidative stress and NLRP3 inflammasome activation via promoting p53 acetylation in lipopolysaccharide-induced acute lung injury. Cell Death Discov. 9, 12 (2023).
Gu, Z. et al. The p53-inducible gene EI24/PIG8 localizes to human chromosome 11q23 and the proximal region of mouse chromosome 9. Cytogenet Cell Genet 89, 230–233 (2000).
Brown, R. L., Sequeira, R. P. & Clarke, T. B. The microbiota protects against respiratory infection via GM-CSF signaling. Nat. Commun. 8, 1512 (2017).
Jeyanathan, M. et al. Parenteral BCG vaccine induces lung-resident memory macrophages and trained immunity via the gut-lung axis. Nat. Immunol. 23, 1687–1702 (2022).
Wu, T. et al. Alterations in the gut microbiome and cecal metabolome during Klebsiella pneumoniae-induced pneumosepsis. Front Immunol. 11, 1331 (2020).
Ji, J. J. et al. Probiotics protect against RSV infection by modulating the microbiota-alveolar-macrophage axis. Acta Pharm. Sin. 42, 1630–1641 (2021).
Ferreira, A. V., Dominguez-Andres, J., Merlo Pich, L. M., Joosten, L. A. B. & Netea, M. G. Metabolic regulation in the induction of trained immunity. Semin Immunopathol. 46, 7 (2024).
Huang, S. C. et al. Metabolic reprogramming mediated by the mTORC2-IRF4 signaling axis is essential for macrophage alternative activation. Immunity 45, 817–830 (2016).
Klichinsky, M. et al. Human chimeric antigen receptor macrophages for cancer immunotherapy. Nat. Biotechnol. 38, 947–953 (2020).
Jin, R., Neufeld, L. & McGaha, T. L. Linking macrophage metabolism to function in the tumor microenvironment. Nat. Cancer 6, 239–252 (2025).
Bai, L. et al. Distinct APCs explain the cytokine bias of alpha-galactosylceramide variants in vivo. J. Immunol. 188, 3053–3061 (2012).
Suryawanshi, R. K. et al. mTORC2 confers neuroprotection and potentiates immunity during virus infection. Nat. Commun. 12, 6020 (2021).
Desai, J. V. et al. C5a-licensed phagocytes drive sterilizing immunity during systemic fungal infection. Cell 186, 2802–2822.e2822 (2023).
Zheng, M. et al. TLR2 senses the SARS-CoV-2 envelope protein to produce inflammatory cytokines. Nat. Immunol. 22, 829–838 (2021).
Peh, H. Y. et al. Vitamin E isoform gamma-tocotrienol protects against emphysema in cigarette smoke-induced COPD. Free Radic. Biol. Med. 110, 332–344 (2017).
Xu, H. et al. Notch-RBP-J signaling regulates the transcription factor IRF8 to promote inflammatory macrophage polarization. Nat. Immunol. 13, 642–650 (2012).
Wang, T. et al. Influenza-trained mucosal-resident alveolar macrophages confer long-term antitumor immunity in the lungs. Nat. Immunol. 24, 423–438 (2023).
Acknowledgements
This work was supported by National Natural Science Foundation of China 32325020 (L.B.), 92254304 (L.B.), 82202022 (S.L.), the Strategic Priority Research Program of the Chinese Academy of Sciences (Grant No.XDB0940202, L.B.), the CAS Project for Young Scientists in Basic Research (YSBR-074, L.B.).
Author information
Authors and Affiliations
Contributions
Y.H., D.X., S.L., and L.B. conceived the idea and designed the experiments, Y.H., M.S., and D.C. performed experiments, Y.Z., J.P., S.L., X.C., Y.W., C.T., H.Z., Q.S., B.F., L.Y., and W.J. provided materials, developed methods, or discussed experiments, Y.H., D.X., S.L., and L.B. wrote the manuscript. All authors contributed to the article and approved the submitted version.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Huang, Y., Su, M., Zhang, Y. et al. Microbiota-induced EI24 improves homeostasis but impedes function of alveolar macrophages via metabolic regulation. Nat Commun 17, 2227 (2026). https://doi.org/10.1038/s41467-026-69000-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-69000-3