Abstract
Growing oocytes accumulate maternal mRNA to support subsequent meiotic maturation and maternal-to-zygotic transition. However, the regulatory mechanisms governing the fate of these maternal mRNAs remain largely unknown. Here, we identified heterogeneous nuclear ribonucleoprotein M (hnRNPM) as a critical regulator of pre-mRNA alternative splicing during mouse oocyte development. Genetic ablation of hnRNPM leads to severe cytoplasmic defects, meiotic arrest, and complete female infertility. Using SCAN-seq, we uncovered novel transcript isoforms and systematically characterized hnRNPM-regulated alternative splicing events. Furthermore, LACE-seq revealed hnRNPM-binding sites at single-nucleotide resolution in oocytes, linking its RNA-binding activity to splicing fidelity. Additionally, hnRNPM interacts with BCAS2, a known splicing factor critical for oocyte development, and modulates its binding to pre-mRNA loci to precisely control the alternative splicing. Overall, our study not only uncover an essential role of hnRNPM in mammalian oocyte development and female fertility but also unveils a critical regulatory network governing alternative splicing during oocyte development.
Similar content being viewed by others
Introduction
Oocytes, the largest mammalian cell type, accumulate substantial amounts of maternal transcripts during their growth phase to support subsequent meiotic maturation and early embryonic development1,2,3. The proper production, storage, and surveillance of mRNA are critical for oocyte and early embryonic development. We previously identified nuclear poly(A) domains as critical processing hubs that coordinate the maturation and storage of newly synthesized mRNAs in growing oocytes3. N-acetyltransferase 10 catalyzes the N4-acetylcytidine modification of mRNAs encoding key maternal factors4. RNA helicase mRNA transporter 4 functions as nuclear RNA surveillance machinery to ensure the clearance of aberrant transcripts in oocytes2. However, the post-transcriptional regulation of pre-mRNAs remains largely unknown.
Alternative pre-mRNA splicing represents a classical mechanism for enriching the diversity of the transcriptome and proteome5,6. Through this process, a single pre-mRNA can generate multiple mRNA isoforms, which subsequently translate into distinct protein variants7. mRNA undergoes significant alternative splicing dynamics during maternal-to-zygotic transition, a process referred to as zygotic splicing activation (ZSA)8. ZSA occurs concurrently with zygotic genome activation (ZGA) during early embryonic development in both humans and mice8. Precise expression of transcript isoforms is critical for successful preimplantation embryogenesis. Additionally, genetic studies in mice have identified several key splicing regulators, including RTCB, BCAS2, ESRP1, and PCBP1, that are essential for oogenesis9,10,11,12, further illustrating the significant role of alternative splicing in oocytes.
While pre-mRNA splicing is critical for oocyte and early embryonic development, capturing full-length transcripts and identifying splicing factor-binding sites remains technically challenging because of the limited quantity and scarcity of oocyte samples. Recent technological advances in RNA sequencing, particularly the emergence of third-generation sequencing platforms, have revolutionized our ability to detect and characterize splicing events with unprecedented accuracy13. The development of single-cell amplification and sequencing of full-length RNAs using a nanopore platform (SCAN-seq) based on third-generation sequencing platforms has overcome previous limitations by enabling comprehensive full-length transcriptome analysis from low-input samples and even on a single oocyte14. This method provides a sensitive and accurate approach for detecting full-length transcripts in oocytes. Additionally, linear amplification of cDNA ends and sequencing (LACE-seq), which combines linear cDNA end amplification with RNA-binding protein (RBP)-induced reverse transcription termination, allows precise mapping of RBP interaction sites at a single-nucleotide resolution in oocytes15.
Heterogeneous nuclear ribonucleoproteins (hnRNPs) are a family of RBPs that form complexes with heterogeneous nuclear RNA to regulate pre-mRNA processing16. Although hnRNPs play crucial roles during spermatogenesis17,18,19,20,21, their functions in oogenesis remain largely unknown. In a systematic evaluation of hnRNP expression across multiple tissues, we observed that Heterogeneous nuclear ribonucleoprotein M (hnRNPM) is highly expressed in reproductive system22, suggesting a potentially critical role in germ cell development and reproductive physiology. Moreover, an independent study employing mRNA interactome capture (RIC) analysis on 50,000 fully-grown mouse oocytes identified hnRNPM as one of the core mRNA-binding proteins (RBPs) enriched in oocytes23. This finding indicates that hnRNPM may functionally engage in regulating the fate of maternal mRNAs in oocytes. hnRNPM contains three RNA recognition motif domains that bind to RNAs. Recent studies have demonstrated that hnRNPM preferentially binds to deep introns to repress cryptic splicing, thereby maintaining transcriptome integrity, which is important for regulating interferon-associated pathways and enhancing cancer surveillance24. Additionally, hnRNPM forms a complex with 5′-tRH-GlyGCC to regulate cancer cell proliferation25 and interacts with Fragile X Mental Retardation Protein and PARP4 to influence tumor progression26,27.
In this study, we demonstrated that hnRNPM coordinates with BCAS2 to regulate alternative pre-mRNA splicing during oogenesis and is essential for establishing competent oocytes capable of supporting oocyte maturation and early embryogenesis. We found that hnRNPM exhibited stage-specific nuclear localization throughout folliculogenesis, disappeared during meiotic resumption, and reappeared at the 2-cell stage. Genetic ablation of hnRNPM led to severe defects in cytoplasmic maturation, failure to enter metaphase I, and female infertility. Mechanistically, hnRNPM regulates alternative splicing of key factors involved in cytoplasmic lattice formation (Khdc3 and Nlrp14) and meiotic maturation (Cenph and Zar1l). We identified differential alternative splicing patterns in the novel Nlrp14 and Zar1l isoforms using SCAN-seq, overcoming the limitations of conventional sequencing technologies. Furthermore, we mapped the hnRNPM-binding sites and revealed their preferential association with transcripts governing organelle localization and meiotic regulation using LACE-seq analysis. These findings establish that hnRNPM is an indispensable regulator of oocyte development and provide a comprehensive analysis of hnRNPM-mediated splicing control during mammalian oogenesis.
Results
The dynamic nuclear expression of hnRNPM during oocyte development
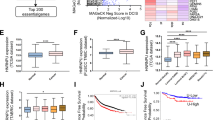
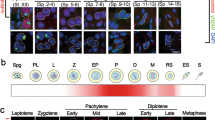
Multi-alignment and phylogenetic analyses of the hnRNPM protein sequences showed that hnRNPM is highly conserved across multiple mammalian species, including human, rhesus monkey, mouse, Norway rat, pig, dog, domestic cattle, and bovine (Fig. 1a and Supplementary Fig. 1a, b). Strikingly, the sequence homology between mouse and human hnRNPM proteins was remarkably high (97.53%), suggesting that murine models may provide valuable insights for understanding and treating human diseases (Supplementary Fig. 1a). To elucidate the role of hnRNPM in female fertility, we systematically examined its spatiotemporal expression patterns during oogenesis. Immunofluorescence analysis revealed that hnRNPM was localized in oocyte nuclei. Its expression was weak in embryonic germ cells, increased after birth, and sustained throughout all subsequent follicular stages, including primordial, primary, secondary, and antral follicles (Fig. 1b and Supplementary Fig. 1c–e). To characterize the dynamics of hnRNPM expression following meiotic resumption, we conducted in vitro maturation (IVM) and in vitro fertilization (IVF). Immunofluorescence co-staining of hnRNPM with α-Tubulin (a marker of the spindle) and DAPI revealed its distinct spatiotemporal localization. hnRNPM was strongly expressed in the nuclei of germinal vesicle (GV) with non-surrounded nucleus (NSN) oocyte, whereas the signal was diminished in GV with surrounded nucleus (SN) oocyte and was completely absent in germinal vesicle breakdown (GVBD), metaphase I (MI), and metaphase II (MII) oocytes (Fig. 1c). Following fertilization, hnRNPM remained undetectable in zygotes but reappeared in the nuclei of 2-cell embryos, persisting through the 4-cell, morula, and blastocyst stages (Fig. 1d).
a The amino acid sequence identity of hnRNPM protein was compared between humans and other mammalian species. Source data are provided as a Source data file. b Immunofluorescence (IF) images showing hnRNPM expression in different follicle types. The enlarged panels below correspond to the areas indicated in the panels above. Scale bars: 50 μm. c, d Spatiotemporal expression and localization of hnRNPM were characterized using immunofluorescence during meiotic maturation of mouse oocytes and subsequent early embryonic stages. Scale bars: 20 μm. Representative results shown in (b–d) were obtained from at least three independent experiments with similar results.
Maternal Hnrnpm loss results in infertility
To illustrate the physiological role of hnRNPM during oocyte development, we conditionally deleted Hnrnpm in germ cells using Stra8-Cre transgenic mice (Stra8-Cre; Hnrnpmflox/Del, herein called Hnrnpm cKO), in which Cre specifically knockout the exon 7–12 of Hnrnpm in female germ cells from embryonic day 12.5 (Fig. 2a–c)28. Both the Hnrnpm mRNA and hnRNPM protein levels in cKO oocytes were significantly lower than those in the litter-matched controls (Hnrnpmflox/+; Stra8-Cre) mice (Fig. 2d, e and Supplementary Fig. 2a–c), indicating that hnRNPM was successfully inactivated in oocytes with high efficiency. The fertility test revealed that conditional deletion of Hnrnpm in germ cells led to complete female infertility (Fig. 2f).
a The strategies employed to construct Hnrnpm conditional knockout mice. Exons 7–12 of Hnrnpm were deleted from the germ cells using Stra8-Cre. Images of scissors were generated by figdraw.com. b The expression of Stra8-Cre commenced from embryonic day 12.5 (E12.5). c Representative genotyping results for wild-type, heterozygous (Stra8-Cre; Hnrnpm+/flox), and cKO (Stra8-Cre; Hnrnpmflox/Del) mice. d RT-qPCR analysis revealed a significant reduction in Hnrnpm mRNA expression in GV oocytes. Two-sided Student’s t-tests. Data are presented as mean ± SEM. n = 3. p < 0.0001. e Representative images of IF with anti-hnRNPM antibody (purple) and DAPI (white) on ovary sections from control and Hnrnpm cKO mice. Primordial, primary, and secondary follicles are highlighted by the white squares. Scale bar: 50 μm. f Comparison of offspring numbers between control and female mice during the 5-month observation period. Two-sided Student’s t-tests. Data are presented as mean ± SEM. n = 5. p = 0.005276 (1), p < 0.000001 (2,3,4,5). g Representative ovaries from 3-month-old control and Hnrnpm cKO mice. Scale bars: 1 mm. h Representative H&E-stained ovarian sections from 3-month-old control and Hnrnpm cKO mice. Scale bars: 500 μm (left), 250 μm (right). i Quantitative analyses of primordial follicles (PrF), primary follicles (PF), secondary follicles (SF), and antral follicles (AF) per ovary at 3 months of age. Two-sided Student’s t-tests. Data are presented as mean ± SEM. n = 3. ns not significant. j Number of oocytes retrieved following PMSG and hCG administration in 3-week-old control and Hnrnpm cKO mice. Two-sided Student’s t-tests. Data are presented as mean ± SEM. n = 10. ns not significant. k Representative images of the zygote, 2-cell, 4-cell, morula, and blastocyst embryos generated by in vitro fertilization (IVF) of oocytes from superovulated 3-week-old control and Hnrnpm cKO female mice using wild-type sperm. Scale bar: 200 μm. l Developmental rates of embryos reaching specific stages at corresponding time points following IVF. Two-sided Student’s t-tests. Data are presented as mean ± SEM. n = 3. p = 0.000002 (Pronucleus, Blastocyst), p < 0.000001 (2-cell, 4-cell, Morula). Representative results shown in (c, e, g, h, k) were obtained from at least three independent experiments with similar results. Source data (d, f, i, j, l) are provided as a Source data file.
To investigate the role of hnRNPM in oocyte development, we conducted comprehensive histological analyses of ovarian tissues from 3-month-old female mice. Comparative evaluation of control and Hnrnpm cKO ovarian tissues revealed no significant differences in ovarian size or maximum cross-sectional area (Fig. 2g, h). Quantitative assessment of follicle populations using hematoxylin-eosin staining demonstrated comparable numbers of follicles at all developmental stages, including the primordial, primary, secondary, and antral follicles (Fig. 2i).
To synchronize follicular growth and promote ovulation, control and Hnrnpm-deficient female mice were injected with pregnant mare serum gonadotropin (PMSG) at postnatal day 21 (P21), followed by human chorionic gonadotropin (hCG) 48 h later. After induction of superovulation, we collected cumulus-oocyte complexes (COCs) and performed IVF. While the number of oocytes obtained was comparable between control and Hnrnpm-deficient mice (Fig. 2j and Supplementary Fig. 2d), Hnrnpm-deficient oocytes failed to complete cleavage and embryonic development after IVF (Fig. 2k, l), indicating that Hnrnpm-deficient oocytes lacked developmental potential after fertilization.
hnRNPM regulates the formation of cytoplasmic lattice
Given the robust expression of hnRNPM during folliculogenesis and its complete absence from GVBD through the zygotic stage, we hypothesized that maternal hnRNPM deficiency causes early developmental impairments that subsequently compromise embryonic competence. To test this hypothesis, we collected GV-stage oocytes from 21-day-old female mice for statistical and morphological analysis. To synchronize follicular growth, control and Hnrnpm cKO female mice at P21 were injected with PMSG, and GV oocytes were collected from the ovaries at 14 h post-injection (Fig. 3a). Strikingly, although the number of GV oocytes collected from the ovaries was comparable between control and Hnrnpm cKO female mice (Fig. 3b), 88.3% of Hnrnpm-deficient oocytes exhibited abnormal dark cytoplasmic granules, in contrast to the homogeneous cytoplasm observed in the controls (Fig. 3c, d).
a Schematic diagram illustrating the methods used to collect GV oocytes from 3-week-old control and Hnrnpm cKO mice. Schematic illustration of mouse was adapted from SciDraw. The oocyte diagram is a modified version of an original Servier Medical Art resource. All modified images are used under a CC BY 4.0 license (https://creativecommons.org/licenses/by/4.0/). b Quantitative analysis of GV oocytes collected from 3-week-old control and Hnrnpm cKO mice. Two-sided Student’s t-tests. Data are presented as mean ± SEM. n = 10. ns not significant. c Representative images of GV oocytes derived from control and Hnrnpm cKO mice. Large dark cytoplasmic granules were observed in Hnrnpm cKO mice. Scale bar: 20 μm. d Quantitative analysis of GV oocytes displaying abnormal cytoplasmic granules in control and Hnrnpm cKO mice. Two-sided Student’s t-tests. Data are presented as mean ± SEM. n = 10. p < 0.0001. e Representative transmission electron microscopy (TEM) images of GV oocytes from control and Hnrnpm cKO mice. Red arrowheads denote cell lattices, blue arrowheads highlight mitochondria, and yellow arrowheads indicate lipid droplets. Scale bars: 2 μm (left), 1 μm (right). f Representative fluorescent images showing mitochondrial distribution (MitoTracker-labeled) in germinal vesicle (GV) oocytes from control and Hnrnpm cKO mice. White arrowheads indicate mitochondrial aggregates. Scale bars: 20 μm. g Nile red staining, indicating the distribution of lipid droplets in control and Hnrnpm cKO mice. White arrowheads indicate lipid aggregates. Scale bars: 20 μm. Representative results shown in (c, e–g) were obtained from at least three independent experiments with similar results. Source data (b, d) are provided as a Source data file.
Cytoplasmic lattices serve as critical structural scaffolds that maintain proper organelle distribution by preventing the aberrant aggregation of cytoplasmic components29,30. To investigate the subcellular differences between control and Hnrnpm knockout oocytes, we performed transmission electron microscopy analysis of GV-stage oocytes. Cytoplasmic lattices were abundant in control oocytes, whereas they were markedly reduced in Hnrnpm cKO oocytes (Fig.3e), demonstrating that hnRNPM is essential for their formation.
Consistent with the observed loss of cytoplasmic lattices, transmission electron microscopy images revealed abnormal aggregation of lipid droplets and mitochondria in Hnrnpm cKO oocytes (Fig. 3e). To further characterize mitochondrial distribution patterns, we conducted MitoTracker staining. Control oocytes exhibited a homogeneous mitochondrial distribution throughout the cytoplasm, whereas Hnrnpm-deficient GV oocytes displayed pronounced mitochondrial clustering and uneven localization (Fig. 3f). Similarly, Nile red staining revealed significant lipid droplet aggregation in Hnrnpm cKO oocytes compared with controls (Fig. 3g). Some lipid droplets colocalized with dark cytoplasmic aggregates in Hnrnpm-deficient oocytes (Fig. 3g and Supplementary Fig. 2e), suggesting that these dark aggregates represented coalesced lipid droplets. Taken together, these findings demonstrate that Hnrnpm deficiency disrupts cytoplasmic lattices, leading to aberrant mitochondrial distribution and lipid droplet aggregation in GV-stage oocytes.
Hnrnpm deficiency impairs the meiotic maturation of oocytes
Acquisition of full developmental competence in oocytes depends on the coordinated progression of nuclear and cytoplasmic maturation31,32. To investigate the role of hnRNPM in oocyte maturation, we performed IVM assays using GV-stage oocytes. While the rates of GVBD were comparable between control and Hnrnpm-deficient oocytes (Fig. 4a, b), only 22.77% of Hnrnpm-deficient oocytes successfully progressed to the MII stage and extruded the first polar body compared with 77.20% of the controls (Fig. 4c). Consistent with IVM results, only ~30% of ovulated oocytes from Hnrnpm cKO mice reached the MII stage following hCG injection (Supplementary Fig. 2f, g), confirming severe maturation defects. Subsequently, we examined the nuclear behavior during oocyte meiosis. In control MI and MII oocytes, chromosomes are neatly aligned at the metaphase plate, accompanied by the establishment of a bipolar spindle, whereas Hnrnpm-deficient oocytes display misaligned chromosomes and exhibit apolar and multipolar spindles (Fig. 4d–g).
a Representative image showing IVM progression of control and Hnrnpm cKO oocytes. Scale bars: 50 μm. b Quantitative analysis of germinal vesicle breakdown (GVBD) rates in control and Hnrnpm cKO mice. Two-sided Student’s t-tests. Data are presented as mean ± SEM. n = 5. ns not significant. c Quantitative analysis of polar body extrusion (PBE) rates in cultured oocytes from control and Hnrnpm cKO mice. Two-sided Student’s t-tests. Data are presented as mean ± SEM. n = 5. p < 0.0001. d Immunofluorescence showing spindle assembly, chromosome alignment, and microtubule-organizing centers (MTOCs) of control and Hnrnpm cKO oocytes after 8 h of in vitro culture. Microtubules were labeled with α-Tubulin (green) and TPX2 (red). The MTOCs are marked with pericentrin (green). Chromosomes were counterstained with DAPI (blue). Scale bars: 10 μm. e Quantitative assessment of meiotic abnormalities in control and Hnrnpm cKO oocytes. Two-sided Student’s t-tests. Data are presented as mean ± SEM. n = 3. p < 0.0001. f Representative images showing spindle assembly, chromosome alignment, and MTOCs in control and Hnrnpm cKO oocytes after 16 h of in vitro culture. Scale bars: 10 μm. g Proportion of oocytes with meiotic defects after 16 h in vitro maturation. Two-sided Student’s t-tests. Data are presented as mean ± SEM. n = 5. p < 0.0001. Representative results shown in (a, d, f) were obtained from at least three independent experiments with similar results. Source data (b, c, e, g) are provided as a Source data file.
To determine whether hnRNPM regulates the assembly of a microtubule-organizing center (MTOC), we analyzed the expression of pericentrin, a key MTOC marker. Interestingly, pericentrin was localized around the spindle poles in control oocytes following 8 h of in vitro maturation, while most Hnrnpm-deficient oocytes failed to show detectable pericentrin expression (Fig. 4d and Supplementary Fig. 2h). This observation suggests that Hnrnpm deficiency may disrupt the timely expression of pericentrin after meiotic resumption. Remarkably, after in vitro maturation for 16 h, the expression of pericentrin can be observed in majority of Hnrnpm-deficient oocytes (Fig. 4f and Supplementary Fig. 2h). These results indicate that hnRNPM is required for the proper temporal regulation of pericentrin expression during oocyte maturation. We next asked whether aberrant spatiotemporal expression of pericentrin stemmed from defects in earlier growth phases. Fluorescence staining of GV oocytes revealed that well-defined puncta of pericentrin were present in controls but undetectable in KO oocytes (Supplementary Fig. 2h, i), indicating an early developmental defect. Collectively, these results demonstrate that hnRNPM is essential for meiotic maturation in oocytes.
hnRNPM regulates alternative splicing during oocyte development
To determine whether hnRNPM regulates alternative splicing in oocytes, we conducted SCAN-seq (single-cell amplification and sequencing of full-length RNAs using a nanopore platform) using control and Hnrnpm-deficient GV oocytes from postnatal day 21 mice (Fig. 5a and Supplementary Fig. 3a). Across all samples, we detected an average of 29,778 transcript isoforms per sample (Supplementary Fig. 3b). Interestingly, through full-length sequencing, novel transcripts in oocytes were detected (Supplementary Data 1). To study the function of these novel transcripts, we performed a Gene Ontology (GO) enrichment analysis using the international standard database (https://www.geneontology.org/) to investigate the biological processes involving these novel transcripts. Strikingly, the top enriched processes were closely related to female infertility—including female meiosis chromosome segregation, regulation of fertilization, and maternal-to-zygotic transition of gene expression (Supplementary Fig. 3c and Supplementary Data 1), suggesting that unique transcripts may exist in oocytes to regulate female fertility. To validate our sequencing results, four unannotated transcripts related to female fertility were amplified using RT-PCR and detected using Sanger sequencing (Fig. 5b–j). Meikin is a conserved regulator of kinetochores during meiosis I33. Nlrp14 regulates the subcellular localization of Dnmt1/Uhrf1 in mouse oocytes and is associated with cell lattice formation, oocyte maturation, and early embryonic development34,35,36. Vrk1 (Vaccinia-related kinase 1) mediates meiotic progression during oogenesis37. Zar1l, also known as Zar2, mediates maternal transcriptome and translational activation and regulates oocyte meiotic maturation38,39,40,41. The Sanger sequencing results for these four transcripts precisely matched the transcripts predicted by SCAN-seq (Fig. 5f–j). The remarkable accuracy in the identification of unannotated transcripts provided evidence that SCAN-seq was successful.
a Schematic illustration of the SCAN-seq method. Briefly, control and Hnrnpm cKO GV oocytes were lysed to release mRNAs. First-strand cDNAs were synthesized using a 24-nt barcode RT primer. Following amplification, the cDNAs derived from multiple cells were combined, pooled together, and purified. Subsequently, the full-length cDNAs were used for the construction of nanopore libraries and sequencing. The mouse, oocyte, and nucleic acid schematics are modified adaptations of original resources from SciDraw and Servier Medical Art, respectively, and are employed under a CC BY 4.0 license. b–j PCR and Sanger sequencing validation of genes (Nlrp14, Meikin, Vrk1, and Zar1l) with novel isoforms identified in GV oocytes. Ref. Reference. Representative results shown in (b–e) were obtained from at least three independent experiments with similar results. Source data (b–e) are provided as a Source data file.
To characterize alternative splicing patterns, we systematically analyzed seven major types of splicing events: Skipping exon (SE), Mutually exclusive exons (MX), Alternative 5’ splice-site (A5), Alternative 3′ splice-site (A3), Retained intron (Rl), Alternative first exon (AF), and Alternative last exon (AL). Among these, the skipping exons and alternative first exons were the most prevalent splicing types (Supplementary Fig. 3d). Subsequently, we compared the splicing patterns between control and Hnrnpm-deficient GV oocytes and identified 1147 differentially spliced events, of which 992 were successfully annotated to known genes (Fig. 6a, Supplementary Fig. 4a, b and Supplementary Data 2). The gene ontology (GO) terms of these differentially spliced genes revealed multiple distinct gene clusters with a particular GO term enriched in the meiotic cell cycle (Fig. 6b). Other pathways included chromatin organization, protein transport, regulation of DNA-templated transcription, protein localization to microtubules, and spliceosomal complex assembly, implying that hnRNPM regulates multiple cellular processes in oocytes (Fig. 6b).
a The numbers and categories of annotated differential alternative splicing (AS) events detected in GV oocytes following Hnrnpm depletion. b GO term analysis of differentially alternative splicing (AS) genes by Metascape. c Diagrammatic representation of alternative splicing quantification methodology using percentage spliced in (PSI) values. The PSI (Percent Spliced In) represents the relative abundance of transcripts containing a specific exon or splice site. d, e Sashimi plots illustrating differential alternative splicing events of cell lattice formation-associated genes (Khdc3 and Nlrp14) and meiotic maturation-related genes (Cenph and Zar1l). PCR validation of these candidate genes confirmed the splicing alterations in Hnrnpm cKO oocytes. Schematic of alternatively spliced exons. Bar plots display the percentage spliced in (PSI) values of control and Hnrnpm cKO oocytes, calculated as PSI = splice_in/(splice_in + splice_out). Two-sided Student’s t-tests. Data are presented as mean ± SEM. n = 3. p = 0.0022 (Khdc3), 0.0012 (Nlrp14), 0.0001 (Cenph), 0.0002 (Zar1l). Representative results shown in (d, e) were obtained from at least three independent experiments with similar results. Source data (b, d, e) are provided as a Source data file.
Among differentially spliced genes, those associated with cytoplasmic lattice formation (Khdc3 and Nlrp14), meiotic maturation (Cenph and Zar1l), and female fertility (Bcl2l1 and Prkaa1) have attracted attention. Khdc3 (also known as FILIA) is a component of the subcortical maternal complex (SCMC) that is essential for the formation of cytoplasmic lattices42. Nlrp14 has been implicated in cell lattice formation, oocyte maturation, and early embryonic development34,35,36. Cenph is a kinetochore component that regulates meiotic resumption in oocytes43. Zar1l mediates maternal transcriptome and translational activation and regulates oocyte meiotic maturation38. Prkaa1 (also known as AMPK) and Bcl2l1 are associated with female infertility44,45. Analysis of RNA sequencing data using sashimi plots revealed distinct alternative splicing patterns, including increased exon skipping in Cenph and Prkaa1, decreased exon skipping in Nlrp14 and Zar1l, elevated intron retention in Khdc3, and differential first-exon splicing in Bcl2l1, all of which were validated using RT-PCR (Fig. 6c–e and Supplementary Fig. 4c, d). These findings support the notion that hnRNPM is crucial for ensuring the fidelity of pre-mRNA splicing in genes that regulate cytoplasmic lattice formation and meiotic maturation. To assess the impact of alternative splicing defects on protein expression, we conducted Western blot analysis for two key downstream targets, CENPH and ZAR1L (Supplementary Fig. 4e). These data demonstrate that splicing defects caused by hnRNPM deficiency leads to a sharp decline in ZAR1L and a mild downregulation of CENPH protein, thereby bridging the observed molecular changes to downstream phenotypic consequences.
Profiling the RNA-binding landscape of hnRNPM in oocytes using Lace-seq
Given that hnRNPM is an RBP that regulates alternative splicing during oocyte development, we investigated the RNA-binding landscape of hnRNPM in oocytes. Lace-seq is a high-resolution method capable of identifying RBP-binding sites with high precision in dozens of cells15. In this technique, the binding site of the RBP is directly retrieved by linearly amplifying the termination signal of the reverse transcriptase precisely at the RBP-binding site (Fig.7a). Compared with conventional methods, this technology achieves remarkable accuracy in identifying RBP-binding sites with single-base resolution at the single-cell level4,15.
a Schematic illustration of the LACE-seq workflow used to profile hnRNPM-binding sites in GV oocytes. The mouse, oocyte, nucleic acid schematics, syringe, and lightning are modified adaptations of original resources from SciDraw, Servier Medical Art, and figdraw, and are licensed under CC BY 4.0 license. b Heatmap visualization of the hnRNPM LACE-seq signal intensity across all identified peaks. Genomic regions are displayed as 6-kb windows centered on peak summits, with hierarchical clustering of similar binding patterns. c Pie chart displaying the genomic distribution of hnRNPM-binding sites identified by LACE-seq. CDS, coding sequence; UTR, untranslated region d RNA-binding motifs of hnRNPM were characterized using de novo motif analysis of LACE-seq peaks using HOMER. e GO term enrichment analysis of hnRNPM-binding transcripts. f Venn diagram showing the overlap between hnRNPM-binding genes and differential alternative splicing (AS) genes. The heatmap on the right displays the GO enrichment results for the overlapping genes. Metascape was used to perform GO term. Source data are provided as a Source data file. g–i Genome browser tracks showing LACE-seq binding peak distributions of cell lattice formation-associated genes (Khdc3 and Nlrp14) and meiotic maturation-related genes (Cenph). The purple and green tracks show hnRNPM binding profiles from LACE-seq performed on control and cKO oocytes, respectively.
To profile the targets of hnRNPM at a single-nucleotide resolution in GV oocytes, we performed hnRNPM LACE-seq using GV oocytes. In parallel, we conducted hnRNPM LACE-seq using Hnrnpm cKO oocytes to eliminate false positive signals (Fig. 7a). Bioinformatics analyses of the LACE-seq results showed that the binding signals of hnRNPM were markedly decreased in Hnrnpm cKO oocytes, suggesting that our antibody exhibited high specificity (Fig. 7b). We identified 1691 hnRNPM-binding sites (corresponding to 1690 genes) in wild-type GV oocytes compared with Hnrnpm cKO oocytes (Supplementary Data 3). Genomic distribution analyses of the reads revealed that 69.67% of the putative binding sites were located in the exons of mRNAs, with 45.02% of reads mapped to the coding sequence (CDS), 9.57% to other exons, 11.75% to the 3′ untranslated region (3′ UTR), and 3.32% to the 5′ UTR, respectively, suggesting that hnRNPM preferentially binds to exons in oocytes (Fig. 7c). Previous eCLIP-seq analyses have demonstrated preferential binding of hnRNPM to the GU-rich hexamer46,47. To determine the consensus-binding sequence of hnRNPM, we performed motif analysis using the findMotifsGenome.pl tool from HOMER on the identified peak regions. Consistent with previous reports, hnRNPM was preferentially crosslinked with GU-rich sequences (Fig. 7d). GO terms indicated that hnRNPM-binding transcripts were enriched in establishment of organelle localization, heterochromatin formation, actin cytoskeleton organization, and regulation of synapse organization (Fig. 7e).
To further investigate the correlation between hnRNPM-targeted mRNA and alternative splicing, we compared the LACE-seq data with differential alternative splicing events identified by SCAN-seq. Among the 722 genes undergoing differential alternative splicing events, the mRNAs of 116 genes were identified as direct targets of hnRNPM based on LACE-seq data, indicating their potential role as direct targets of hnRNPM-alternative splicing regulatory complexes. Gene enrichment analysis revealed that the main functions of these overlapping genes were related to cell growth and mechanisms associated with pluripotency, microtubule nucleation, mitotic prometaphase, and sister chromatid cohesion (Fig. 7f). Given that hnRNPM regulates the alternative splicing of genes associated with cytoplasmic lattice formation (Khdc3 and Nlrp14), meiotic maturation (Cenph and Zar1l), and female fertility (Bcl2l1 and Prkaa1), we investigated whether hnRNPM could bind to the splicing sites of these transcripts. Strikingly, hnRNPM was specifically enriched at the alternative splicing sites of Khdc3, Nlrp14, Cenph, Bcl2l1, and Prkaa1 (but not Zar1l), indicating its direct regulation of these transcripts (Fig. 7g–i and Supplementary Fig. 5a, b). Furthermore, to obtain more direct evidence for hnRNPM’s role in regulating alternative splicing, we analyzed hnRNPM binding signals on differentially alternative spliced transcripts (AS) and unaffected alternative spliced transcripts (Un-AS). Strikingly, hnRNPM binding was significantly enriched on the exons of transcripts that underwent alternative splicing upon its depletion, compared to those that were unaffected (Supplementary Fig. 5c, d). This provides much more direct evidence that hnRNPM binding at these specific regions is functionally responsible for the splicing regulation.
hnRNPM cooperates with BCAS2 to regulate the alternative splicing
To unveil the protein-protein interaction network of hnRNPM in oocytes, we conducted immunoprecipitation-mass spectrometry using approximately 8000 GV oocytes (Fig. 8a and Supplementary Data 4). GO analysis revealed that hnRNPM-interacting proteins were significantly enriched in mRNA processing pathways (Fig. 8b), consistent with the known molecular functions of hnRNPM24. Among the interacting proteins enriched in the RNA splicing pathway, BCAS2 garnered attention because of its role as an RNA splicing factor that regulates cell lattice formation and oocyte maturation12 (Fig. 8c). Western blotting confirmed the robust interaction between hnRNPM and BCAS2, and this interaction does not rely on RNA (Fig. 8d and Supplementary Fig. 6a, b). Protein localization studies further revealed that BCAS2 displayed a nuclear distribution pattern similar to that of hnRNPM throughout follicular development (Fig. 8e). Previous studies reported that BCAS2 participates in pre-mRNA splicing by forming a complex with PRP19 and CDC5L12,48,49,50,51,52. We therefore asked whether hnRNPM associates with this complex. Strikingly, co-immunoprecipitation assays demonstrated strong interactions between hnRNPM and both PRP19 and CDC5L (Supplementary Fig. 6c, d), suggesting that hnRNPM and BCAS2 may function in a highly coordinated manner to regulate alternative splicing.
a Schematic diagram of the hnRNPM immunoprecipitation-mass spectrometry (IP-MS) workflow for profiling hnRNPM-interacting proteins in GV oocytes. Schematic illustration of mice was adapted from SciDraw. The oocyte diagram is a modified version of an original Servier Medical Art resource. All modified images are used under a CC BY 4.0 license (https://creativecommons.org/licenses/by/4.0/). b GO enrichment analysis of the identified hnRNPM-interacting proteins by Metascape. c Protein-protein interaction (PPI) network of hnRNPM-interacting proteins involved in RNA splicing. d In vivo co-immunoprecipitation (Co-IP) assays in mouse oocytes to determine whether the interaction between hnRNPM and BCAS2 is RNA-dependent, using treatments with RNasin (ribonuclease inhibitor) or RNase A. All samples presented within this figure panel derive from the same experiment and were processed in parallel. e The localization of BCAS2 in the primordial, primary, secondary, and antral follicles. The lower panels show magnified views of the indicated regions (boxes above). Scale bars: 50 μm. f Venn diagrams showing the overlap of abnormal AS genes and events in Hnrnpm and Bcas2 cKO oocytes. The overlaps of 98 common genes and 57 identical splicing events were both highly significant by hypergeometric test (p = 2e-27 and p = 4e-27, respectively). g–i Sashimi plots showing the splicing patterns of Cenpf, Prc1, and Hdlbp in control, Hnrnpm cKO, and Bcas2 cKO oocytes. j PCR validation of splicing events in (g–i). Schematic showing the alternatively spliced exons. k Bar plots display the percent spliced- in (PSI) values, calculated as PSI = splice_in/(splice_in + splice_out). Two-sided Student’s t-tests. Data are presented as mean ± SEM. n = 3. For Bcas2 ctrl and cKO group, p = 0.003473 (Cenpf), 0.000519 (Prc1), 0.007735 (Hdlbp). For Hnrnpm ctrl and cKO group, p = 0.009008 (Cenpf), 0.012079 (Prc1), 0.001602 (Hdlbp). l, m RIP-PCR and qPCR analyses of hnRNPM and BCAS2-interacted transcripts in P21 ovaries. RNA immunoprecipitation followed using qPCR (RIP-qPCR) was performed to detect Cenpf, Prc1, and Hdlbp transcripts co-precipitated with anti-hnRNPM antibodies using anti-IgG as a negative control in P21 mouse ovaries. Two-sided Student’s t-tests. Data are presented as mean ± SEM. n = 5. n Western blotting confirmed the high knockdown efficiency of hnRNPM in HEK293T cells. o, p RIP-PCR and qPCR analyses revealed BCAS2-RNA interactions in control and HNRNPM knockdown 293T cells. The association of CENPF, PRC1, and HDLBP transcripts with BCAS2 was examined using RIP-PCR (o) and qPCR (o). Two-sided Student’s t-tests. Data are presented as mean ± SEM. n = 5. p = 0.000037 (PRC1). p < 0.000001 (CENPF, HDLBP). All samples presented within a single figure panel (d, j, n) derive from the same experiment and were processed in parallel. Representative results shown in (d, e, j, l, n, o) were obtained from at least three independent experiments with similar results. Source data (d, j–p) are provided as a Source data file.
To investigate the potential cooperative functions of BCAS2 and hnRNPM, we compared our SCAN-seq data with published SCAN-seq data from control and Bcas2 cKO oocytes12. This analysis revealed that hnRNPM and BCAS2 shared 98 commonly affected alternative splicing genes (Fig. 8f). Of these 98 alternative splicing genes, 57 same alternative splicing events occurred in both Hnrnpm and Bcas2 cKO oocytes (Fig. 8f), demonstrating functional synergy between these two factors. To validate the co-regulation of splicing events by hnRNPM and BCAS2, we selected three representative targets for detailed analysis using sashimi plots and RT-PCR. These analyses confirmed consistent splicing alterations in both Bcas2 and Hnrnpm cKO oocytes, including increased exon skipping in Cenpf, decreased exon skipping in Prc1, and differential first-exon splicing in Hdlbp (Fig. 8f–k). Notably, Cenpf and Prc1 play significant roles in the meiotic maturation of oocytes. Prc1 is associated with spindle bipolarization in MI oocytes53, which is consistent with our observation of defective bipolarized spindle formation in Hnrnpm-deficient oocytes during the GVBD-to-MI transition. Cenpf mediates meiotic regulation by recruiting dynamin-related protein 1 (an atypical spindle assembly checkpoint protein) to the kinetochore during oocyte meiosis I54. To investigate whether a similar regulated role exists in human cells, we knocked down HNRNPM in 293T cells, and confirmed efficient protein depletion by western blotting (Supplementary Fig. 6e). Consistent with a conserved regulatory role, depletion of either hnRNPM or BCAS2 in human cells recapitulated the splicing changes of PRC1 and HDLBP, indicating that hnRNPM and BCAS2 also cooperates to control alternative splicing of human transcripts (Supplementary Fig. 6f).
Considering the ability of hnRNPM to co-regulate alternative splicing with BCAS2 in oocytes, we next asked whether transcripts with altered alternative splicing were bound by hnRNPM and BCAS2 proteins. RNA immunoprecipitation (RIP)-qPCR was performed using anti-hnRNPM and BCAS2 antibodies to immunoprecipitate RNAs from ovaries. Notably, Cenpf, Prc1, and Hdlbp transcripts were significantly enriched in both hnRNPM and BCAS2 immunoprecipitates (Fig. 8l, m).
To elucidate how hnRNPM cooperates with BCAS2 during the alternative splicing process, we analyzed the expression and localization of BCAS2 protein upon hnRNPM depletion. The results showed that hnRNPM loss significantly decreased BCAS2 protein expression but did not affect its nuclear localization (Fig. 8n and Supplementary Fig. 6g). Interestingly, analysis of BCAS2-knockout oocytes revealed that BCAS2 depletion also led to a marked reduction in hnRNPM protein levels, without disrupting its nuclear localization (Supplementary Fig. 6h, i). These findings suggest that hnRNPM and BCAS2 likely form a complex and are mutually dependent for protein stability.
To examine whether the reduction of BCAS2 protein in hnRNPM‑deficient cells compromises its enrichment on pre‑mRNA, we conducted BCAS2 RNA immunoprecipitation assays in control and HNRNPM‑knockdown cells. The results showed a marked decrease in BCAS2 binding to CENPF, PRC1, and HDLBP transcripts upon hnRNPM depletion (Fig. 8o, p), indicating that hnRNPM deficiency impairs BCAS2 association with transcripts co‑regulated by both proteins. To further validate these findings, we analyzed six exon-skipping events (Arfip1, Bod1l, Cep43, Lrrc17, Mis18bp1, and Zcchc10) and seven alternative first-exon splicing events (Adh5, Eloc, Esyt1, Lonp2, Mapk6, Mrps33, and Snx15) co-regulated by hnRNPM and BCAS2. All 13 transcripts showed significant enrichment in both hnRNPM and BCAS2 immunoprecipitates (Supplementary Fig. 7a–d). Strikingly, hnRNPM knockdown led to a significant reduction in BCAS2 binding at 9 of these 13 co-regulated targets (69.2%) (Supplementary Fig. 7e–h). Together, these findings support a model in which hnRNPM cooperates with BCAS2 to regulate pre-mRNA alternative splicing in oocytes (Fig. 9).
hnRNPM cooperates with BCAS2 to regulate alternative splicing during oocyte development. The depletion of hnRNPM results in reduced protein levels of BCAS2. The phenotype of disrupted cytoskeletal organization and impaired meiotic maturation in Hnrnpm-deficient oocytes is likely due to aberrant splicing of key genes governing these processes. The oocyte diagram is a modified version of an original Servier Medical Art resource, licensed under a CC BY 4.0.
Discussion
The functional significance of hnRNPM’s Spatiotemporal regulation during oogenesis
Mammalian oocytes, the largest cells in the human body, undergo extensive deposition of maternal transcripts during their growth phase1,2,3. Following meiotic resumption, oocytes enter a transcriptionally silent state that persists until two-cell stage55. This developmental paradigm highlights the fundamental requirement for appropriate accumulation of maternal determinants during oogenesis to ensure successful embryogenesis1,56. hnRNPM exhibits stage-specific expression during oocyte development, with high levels detected in the primordial, primary, secondary, and antral follicles. The nuclear signal diminishes upon GVBD and remains absent until ZSA occurs at the two-cell stage. The specific absence of hnRNPM from the GVBD to zygote stages implies that its expression may be precisely regulated to ensure proper ZSA. We propose that this transient downregulation serves at least two critical functions. First, the absence of hnRNPM, a known splicing regulator, may protect the pre-deposited maternal mRNA pool from spurious or aberrant splicing. Second, its removal might be essential to “reset” the splicing landscape, allowing the nascent zygotic transcriptome to be established under a new regulatory framework. Conversely, persistent hnRNPM expression potentially causes mis-splicing of both maternal transcripts and early zygotic genes, ultimately leading to failure of ZSA and subsequent embryonic developmental arrest. In addition, our recent studies have shown that ZSA is a previously unnoticed process that plays an essential role during embryonic development8. Restoration of hnRNPM expression coincided with the timing of ZSA occurrence, leading us to speculate that hnRNPM may also play a significant role in ZSA.
hnRNP proteins: emerging master regulators of oocyte development
Alternative splicing of pre-mRNAs plays a crucial role in the regulation of eukaryotic gene expression by enabling a relatively limited number of genes to generate significantly larger proteomes. Additionally, many alternatively spliced genes are regulated in a tissue- and cell-specific manner, allowing protein isoforms to be tailored to the specific needs of different cell types and developmental stages. The hnRNP family, which encompasses over 20 members (hnRNP A-U), plays an important role in alternative mRNA splicing57. We systematically investigated the functions of several hnRNP family members, including hnRNPA2B1, hnRNPC, hnRNPH1, and hnRNPU, during spermatogenesis17,18,19,20,58. However, the role of hnRNP family proteins during oocyte development remains largely unexplored. In this study, we demonstrated that hnRNPM exhibits dynamic expression patterns during oocyte growth and early embryogenesis and is essential for cytoplasmic maturation and meiotic progression.
Molecular mechanisms underlying cytoplasmic lattice defects
Cytoplasmic lattices are highly abundant in mammalian oocytes and serve as RNA and protein reservoirs that are essential for early embryonic development56. Previous studies have identified key proteins required for cytoplasmic lattice formation, including PADI6, and components of the subcortical maternal complex (SCMC)—such as KHDC3, NLRP4f, NLRP5, OOEP, TLE6, and ZBED3. Disruption of either PADI6 or SCMC impairs the cytoplasmic lattice assembly, leading to severe embryonic developmental defects29,30,59,60,61,62. Moreover, ZAR1/2 can bind to MATER/PADI6 and localize to the mRNA-MSY2-ZAR1/2 complex in cytoplasmic lattices (CPLs) to ensure the storage and ordered release of maternal mRNAs during maternal-to-zygotic transition38. In this study, we revealed that hnRNPM plays a crucial role in CPL biogenesis by regulating the alternative splicing of key genes associated with CPL formation. Specifically, Hnrnpm deficiency causes retained introns in Khdc3 and aberrant exon skipping in Nlrp14. KHDC3, an essential component of SCMC, is indispensable for CPL formation42, whereas NLRP14 regulates mitochondrial distribution and CPL organization in the cytoplasm36. These findings establish hnRNPM as a regulator of cytoplasmic maturation that orchestrates CPL formation through the pre-mRNA splicing control of critical maternal-effect genes.
Potential mechanisms of meiotic regulation
Hnrnpm-deficient oocytes exhibited severe abnormalities in meiotic spindle formation, specifically failing to establish a bipolar spindle during meiosis I. SCAN-seq analysis revealed the aberrant splicing of key meiotic regulators in mutant oocytes, including exon 10 skipping of Cenpf, exon 8 skipping of Cenph, exon 14 skipping of Prc1, and exon 4 skipping of Zar1l. These genes play critical roles in meiotic progression: Prc1 mediates acentrosomal spindle bipolarization during meiosis I53; Cenpf recruits the atypical spindle assembly checkpoint protein dynamin-related protein 1 to kinetochores54; Cenph is a component of the kinetochore and regulates meiotic resumption of oocytes43; and Zar1 collaborates with Zar2 to regulate oocyte meiotic maturation38. The loss of Hnrnpm disrupts the splicing of multiple meiotic regulators, leading to spindle defects and meiotic failure. In addition, hnRNPM-mediated dysregulation of pericentrin may disrupt microtubule-organizing center (MTOC) function, leading to severe defects in meiotic spindle assembly and chromosome segregation. Notably, the meiotic abnormalities observed in Hnrnpm-deficient oocytes may also arise from impaired cytoplasmic maturation, which is supported by the following evidence: (i) hnRNPM expression ceases after GVBD, indicating that it does not directly regulate subsequent meiotic events or fertilization, and (ii) proper cytoplasmic maturation is a prerequisite for successful meiotic progression1,56. These findings demonstrate that hnRNPM ensures meiotic fidelity through the direct regulation of spindle-associated transcripts and indirectly supports cytoplasmic competence.
Functional synergy with BCAS2
Bcas2 has been conditionally knocked out at various stages of oocyte development to investigate its function and molecular mechanisms in oocytes12,49,63. Loss of Bcas2 results in the formation of dark cytoplasmic granules and defects in the meiotic maturation of oocytes12, a phenotype similar to that observed in Hnrnpm-deficient oocytes. Furthermore, we revealed that hnRNPM and BCAS2 can co-regulate the alternative splicing of Prc1 and Cenpf, both of which are involved in the meiotic maturation of oocytes. Our study also demonstrated that hnRNPM affects the binding intensity of BCAS2 to RNA. Collectively, these findings established a functional partnership between hnRNPM and BCAS2 to safeguard proper oocyte development and meiotic progression through the coordinated regulation of alternative splicing.
Although the conditional knockout of Bcas2 using Stra8-Cre affects meiotic prophase I in both female and male germ cells49,64, it is interesting to note that no significant changes were observed in the first meiotic division in either male or female germ cells when Hnrnpm was conditionally knocked out using Stra8-Cre65. Furthermore, Bcas2 knockout disrupts primordial follicle pool establishment and follicle development, whereas no such defects are observed in Hnrnpm knockout mice, suggesting that BCAS2 can function independently of hnRNPM in addition to acting in synergy with it. In addition, the complete deficiency of BCAS2 likely abolishes its entire function, leading to a catastrophic failure in spliceosome assembly and mRNA processing, thus manifesting a more severe phenotype. In contrast, the loss of hnRNPM results in a partial reduction, but not a complete ablation, of BCAS2 protein. We propose that the residual BCAS2 in Hnrnpm-deficient oocytes retains a degree of functionality, thereby mitigating the phenotypic severity compared to a full BCAS2 knockout.
Insights of SCAN-seq
SCAN-seq identified novel transcript isoforms that had eluded previously detected using conventional RNA sequencing methods. Importantly, we confirmed hnRNPM-mediated alternative splicing regulation of these newly discovered transcripts (Nlrp14 and Zar1l), underscoring the superior capability of SCAN-seq in revealing such molecular events. We observed that the GV oocytes exhibited a significantly higher proportion of alternative splicing in the first exon. Alternative first-exon splicing regulates gene expression diversity by selecting different promoters, or transcription start sites, generating distinct protein isoforms, and modulating translation efficiency. This mechanism enables tissue-specific expression and dynamic responses to environmental cues and plays crucial roles in development and cellular function. However, it remains unclear whether the high proportion of first-exon alternative splicing is a transcriptional splicing characteristic unique to GV-stage oocytes. Additionally, we identified novel transcripts and their corresponding genes associated with the female infertility phenotype. Whether these novel transcripts are oocyte-specific or have unique functions warrants further investigation.
Methods
Ethics statement
The mice used in this study were bred at the Animal Center of Sir Run Run Shaw Hospital, Zhejiang University School of Medicine. The environment was rigorously controlled to adhere to specific pathogen-free standards, with the temperature maintained between 22 °C and 25 °C, humidity levels kept at 50% to 70%, and a regular 12-h day/night cycle of light. All experimental procedures involving animals were approved by the Animal Ethics Committee of Sir Run Run Shaw Hospital, Zhejiang University School of Medicine (approval number: SRRSH2025-0029).
Mice
Hnrnpmflox/+ mice (strain no. T008736) were generated by GemPharmatech (Nanjing, China). Bcas2flox/+ mice were generously provided by Professor Lei Li at the State Key Laboratory of Stem Cell and Reproductive Biology, Institute of Zoology, Chinese Academy of Sciences63. The Stra8-Cre mouse line was gifted by Professor Minghan Tong of the Center for Excellence in Molecular Cell Science, Chinese Academy of Sciences (Shanghai, China)28. Vasa-Cre mice were obtained from The Jackson Laboratory. According to the structure of the Hnrnpm gene, exons 7–12 of Hnrnpm-206 (ENSMUST00000148178.7) transcript containing a 490-bp coding sequence were selected as the knockout region. Knocking out this region results in the impairment of protein function. To generate Hnrnpmflox/+ mice, loxP sites were inserted into introns 6 and 12 of Hnrnpm. Hnrnpmflox/flox mice were obtained by crossbreeding female Hnrnpmflox/+ and male Hnrnpmflox/+ mice. Hnrnpmflox/+; Stra8-Cre mice were generated by mating Stra8-Cre and Hnrnpmflox/+ mice. To produce Hnrnpmflox/Del; Stra8-Cre (designated Hnrnpm cKO) mice, male Hnrnpmflox/+; Stra8-Cre mice were mated with female Hnrnpmflox/flox or Hnrnpmflox/+ mice. To generate Bcas2flox/Del; Vasa-Cre (designated Bcas2 cKO) mice, we crossed male Bcas2flox/+; Vasa-Cre mice with female Bcas2flox/flox or Bcas2flox/+ mice. To genotype the mice, PCR amplification was performed on genomic DNA extracted from mouse tails. The primer sequences used for genotyping are listed in Supplementary Data 5. All mice used in this study were in C57BL/6J background.
Oocyte collection and IVM
Twenty-one-day-old female mice were intraperitoneally injected with 5 IU of PMSG (Ningbo Sansheng). After 44 to 48 h, the ovaries were punctured using a needle to release the oocytes. GV-stage mouse oocytes were manually retrieved using a mouth pipette. GV oocytes were harvested in M2 medium (Sigma, M7167) supplemented with 2.5 μM milrinone (MCE, HY-14252). Only full-grown GV oocytes were cultured in M16 medium (Sigma, M7292) and covered with mineral oil (Sigma, M5310). MI and MII oocytes were collected after 8 and 14 h of culture, respectively.
In vitro fertilization and culture
Three-week-old female mice were superovulated with 5 IU of hCG (Ningbo Sansheng), followed by 5 IU PMSG (Ningbo Sansheng) 44 h later. After 14 h, the female mice were euthanized, and COCs were collected from the ampulla of the fallopian tubes. Sperms were obtained from the epididymis of wild-type adult male mice and capacitated in TYH medium (Nanjing Aibei Biotechnology, M2035) for 50 min. Subsequently, the capacitated sperms and COCs were combined in HTF medium (Nanjing Aibei Biotechnology, M1135) for fertilization. After 6 h, fertilized oocytes were transferred to KSOM (Nanjing Aibei Biotechnology, M1435) for further culturing. Representative early embryos were photographed at 1, 2, 3, and 4 days post-IVF.
Cell culture, plasmid construction, and transfection
HEK293T cells (Procell, CL-0005) were maintained in Dulbecco’s Modified Eagle Medium supplemented with 1% penicillin–streptomycin and 10% fetal bovine serum (NEST Biotechnology, 209111) at 37 °C in a 5% CO₂ humidified atmosphere. To passage cells, begin by removing the spent medium from the culture dish (Bioland, CCD06) and washing the cell layer gently with PBS. Dissociate the adherent cells by adding pre-warmed trypsin-EDTA and incubating at 37 °C. After cell detachment, inhibit trypsin activity by adding Fetal Bovine Serum (Jin Yuan Kang Biotechnology, FBS-300). Then, transfer the cell suspension to a 15 mL centrifuge tube (Bioland, ATS05-15) and pellet the cells via centrifugation at 300 × g for 3 min. Remove the supernatant and resuspend the cells in fresh complete medium. For cryopreservation, resuspend the cell pellet in Serum-free Cell Freezing Medium (Solarbio, C2950), distribute into cryovials (Bioland, AC05-20), and store at −80 °C.
To construct plasmids, the full-length coding sequences of mouse Cdc5l, Hnrnpm, and Prp19 were assembled into the pCMV-C-Flag, pEGFP-N1, or pCMV-HA-C vectors using the Clon Express Ultra One Step Cloning Kit (Vazyme, C115). Following this, each plasmid construct expressing a tagged protein was transfected into HEK293T cells via Polyethyleneimine. For RNA interference experiments, cells were transfected with human hnRNPM-targeting siRNA (Thermo Fisher Scientific, s9261) or human BCAS2-targeting siRNA (Thermo Fisher Scientific, 135537) using Lipofectamine RNAiMAX transfection reagent (Invitrogen, 13778150) according to the manufacturer’s protocol. Following 48 h of transfection, cells were harvested for subsequent analyses.
RT-PCR and qPCR
Total RNA was isolated from oocytes using the RNeasy Mini Kit (QIAGEN, 74104) following the manufacturer’s protocol. Genomic DNA was removed, and cDNA synthesis was performed using the PrimeScript™ RT Reagent Kit with gDNA Eraser (Takara, RR047A). For conventional PCR amplification, the 2X Accurate Taq Master Mix (Accurate Biotechnology, AG11009) was used. The samples were then subjected to agarose gel electrophoresis with a 100 bp DNA ladder (Amizona Scientific LLC, AMG2001-100) and imaged on a ChemiDoc Touch (Bio-Rad) and analyzed using Image Lab (Bio-Rad) software. qRT-PCR was conducted using TB Green® Premix Ex Taq™ II (Takara, RR820A) on a QuantStudio™ Real-Time PCR System (Thermo Fisher Scientific). All reactions were performed in technical triplicate with appropriate negative controls. The primer sequences are provided in Supplementary Data 5.
Immunofluorescence staining
Freshly collected oocytes were fixed in 2% paraformaldehyde for 30 min. After permeabilization with 0.25% Triton X-100 for 15 min, oocytes were blocked with 0.3% bovine serum albumin (BSA) for 30 min. The oocytes were then incubated with the corresponding primary antibodies overnight at 4 °C. Next, the oocytes were washed three times with washing buffer (0.1% Tween-20 and 0.04% Triton X-100 in PBS (MCE, HY-K1023)). The oocytes were subsequently transferred to the secondary antibody and incubated for 30–60 min. After washing three times with washing buffer containing 0.1% Hoechst 33342 (Invitrogen, H3570), the oocytes were mounted using the Slow Fade Diamond Antifade Mountant (Invitrogen, S36967).
For the immunolabeling of tissue slides, the samples were washed with PBS and blocked with 0.3% BSA (MCE, HY-D0842) for 30 min. They were then incubated with primary antibodies at 4 °C overnight. After washing with PBS three times, the samples were incubated with secondary antibodies for 30 to 60 min and then mounted with ProLong™ Diamond Antifade Mountant (Invitrogen, P36970). The image was captured by Zeiss laser-scanning confocal microscope and processed by Zen 2.3 (blue edition) software (Carl Zeiss). For the antibody information, please refer to Supplementary Data 6.
MitoTracker and Nile red staining
Oocytes were cultured in M2 medium containing MitoTracker Red CMXRos (Beyotime, C1049B) for 30 min or Nile red (MCE, 7385-67-3) for 15 min in a humidified chamber at 37 °C. Following three washes with fresh M2 medium, each lasting 5 min, the samples were imaged immediately using a Zeiss laser-scanning confocal microscope.
Transmission electron microscopy
GV oocytes from control and Hnrnpm cKO mice were collected and fixed overnight at 4 °C in 0.1 M cacodylate buffer (pH 7.4) containing 2.5% glutaraldehyde and 3% paraformaldehyde. After three 10-min washes with 0.1 M PB (pH 7.4), samples were post-fixed in 1% osmium tetroxide for 1.5 h and washed again with 0.1 M PB (3 × 10 min). Dehydration was performed using a graded ethanol series (50%, 70%, and 90%; 15 min each), followed by two 30-min immersions in 100% ethanol, a 30-min treatment with a 1:1 ethanol-acetone mixture, and two 30-min immersions in 100% acetone. The specimens were then infiltrated with mixtures of epon resin and acetone (1:1 for 2 h, 3:1 overnight), embedded, and polymerized at 65 °C for 24–48 h. Ultrathin sections (60 nm thick) were prepared and stained with uranyl acetate and lead citrate. Oocytes were sectioned into 70-nm ultrathin slices, stained with uranyl acetate and lead citrate, and imaged using a JEM-1400 transmission electron microscope (JEOL).
Western blotting
Samples were collected and lysed in RIPA UltraMix lysis buffer (X-blot,100086) to extract proteins. Protein concentration was determined with the Bradford Protein Assay Kit (Absin, abs580304). The loading buffer containing β-mercaptoethanol was added to the lysates, and this mixture was heated at 95 °C for 10 min. The samples and protein standard were subsequently loaded onto 10% sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE). After separation, the proteins were transferred onto PVDF Transfer Membranes (Thermo Fisher,88518) and blocked in Tris-buffered saline with Tween 20 (TBST) containing 5% non-fat milk for 1 h at room temperature (RT). Primary antibodies were incubated overnight at 4 °C. The membranes were washed with TBST and incubated with secondary antibodies for 1 h at RT. Protein bands were visualized using a Light Chemiluminescence Kit (Epizyme, SQ201) and imaged using a chemiluminescence detection device (Bio-Rad). The uncropped membranes of the blots are included in the Supplementary Information.
Immunoprecipitation coupled mass spectrometry
GV oocytes were snap-frozen in liquid nitrogen. A total of 8000 GV oocytes were pooled, lysed, and homogenized in 1 mL of IP buffer (NCM Biotech, P70100) containing a complete protease inhibitor (EDTA-free, Roche) and 25 µg/mL RNase A (Sigma) using a glass douncer, followed by incubation for 2 h at 4 °C. The lysates were cleared at 15,000 × g for 15 min, and the supernatant was incubated with 50 µL of pre-cross-linked antibody-magnetic beads (MCE, HY-K0202) overnight at 4 °C. The beads were washed five times in IP buffer and eluted for 10 min at 100 °C in 1× loading buffer. For immunoblotting analysis, protein complexes were eluted from the beads using 1× SDS loading buffer by heating at 98 °C for 5–10 min. For mass spectrometry, proteins were boiled in SDT buffer (4% SDS, 100 mM DTT, and 100 mM Tris-HCl) for 3 min with gentle agitation. The proteins eluted for mass spectrometry analysis were subjected to in-solution tryptic digestion following standard protocols. Briefly, proteins were reduced with DTT, alkylated with iodoacetamide, and digested overnight with sequencing-grade trypsin (Promega) at 37 °C. The resulting peptides were desalted using C18 Stage Tips and analyzed using liquid chromatography-tandem mass spectrometry on a Q-Exactive Plus mass spectrometer (Thermo Fisher Scientific) equipped with a nano-electrospray ion source.
RNA immunoprecipitation
RIP assays were performed using the RNA Immunoprecipitation Kit (BersinBio, Bes5101), according to the manufacturer’s protocol, with minor modifications. Briefly, P21 mouse ovaries or HEK293T cells were lysed in RIP lysis buffer supplemented with 1× protease inhibitor cocktail (Roche, 04693159001) and 40 U/mL RNase inhibitor (Invitrogen, 10777019). The cleared lysates were incubated with 5 μg of IgG, anti-hnRNPM, or anti-BCAS2 antibodies at 4 °C for 16 h with rotation. After six stringent washes with ice-cold RIP wash buffer (5 min per wash at 4 °C), RNA-protein complexes were extracted using TRIzol™ LS Reagent (Invitrogen, 10296028) following the manufacturer’s protocol. The purity and integrity of RNA were verified using an Agilent 2100 bioanalyzer.
LACE-seq sequencing and data analyses
Ctrl and cKO Hnrnpm GV oocytes were collected into the bottom of 1.5 mL Eppendorf LoBind microcentrifuge tubes and irradiated twice with 400 mJ UV-C light on ice. The oocytes were lysed in 50 μL of wash buffer on ice for 10 min, followed by the addition of 1 μL of ribonuclease inhibitor (Ambion, AM2696) and 4 μL of RQ1 DNase (Promega, M6101), with incubation at 37 °C for 3 min. The tube was then snap-chilled on ice for 3 min, after which 10 µl of hnRNPM antibody-coupled beads were added to the lysate. The mixture was rotated at 4 °C for 1 h and subsequently placed on a magnetic stand for 1 min to separate the beads, with the supernatant being carefully discarded. Three washing steps were performed: washing with wash buffer (repeated thrice), followed by washing with high-salt wash buffer, and finally washing with PNK buffer. Immunoprecipitated RNAs were fragmented using MNase (Thermo Scientific, EN0181) diluted 300,000–600,000-fold in 1× MN reaction buffer. After thorough washing of the beads, 10 µL of diluted MNase was added, and the reaction was conducted at 37 °C for 3 min. Enzyme activity was quenched through sequential washes: twice with 1× PNK + EGTA buffer, twice with wash buffer, and twice with PNK buffer.
The beads were resuspended in 20 μL of FastAP reaction mixture (containing 2 μL 10× FastAP buffer, 1 μL FastAP alkaline phosphatase (Thermo Scientific, EF0651), and 17 μL nuclease-free water) and incubated at 37 °C for 10 min. Following washing, the beads were suspended in 20 μL of ligation mixture (comprising 12.5 μL of water, 2 μL of 10× ligation buffer, 0.5 μL of 1 μM 3′ linker, 1 μL of T4 RNA ligase 2 truncated (NEB, M0242), and 4 μL of 50% PEG8000), then incubated at 25 °C for 2.5 h with periodic vortexing. The 3′ linker was synthesized and high-performance liquid chromatography-purified by Integrated DNA Technologies. The beads were resuspended in 8.5 µL of DEPC-treated water containing 1 µL of biotinylated T7-RT primer. Following denaturation at 65 °C for 5 min and immediately cooling on ice for 2 min, the RT mixture was immediately transferred to a PCR tube and subjected to the following incubation protocols: 42 °C for 50 min, 70 °C for 15 min, followed by maintenance at 12 °C. The cDNA was then treated with 3 μL of Exonuclease I (NEB M0293) in 1× buffer (37 °C, 1 h) followed by heat inactivation (80 °C, 20 min). First-strand cDNAs were eluted from protein A/G beads and subsequently captured on streptavidin C1 beads for pre-PCR amplification. Following the addition of a 3′ cDNA linker, double-stranded DNA templates were generated for in vitro transcription (IVT). The IVT products were purified through sequential treatments: turbo deoxyribonuclease digestion to remove DNA templates, followed by purification using Agencourt RNA Clean beads, according to the manufacturer’s protocol. Linearly amplified RNA was reverse transcribed into cDNA and PCR-amplified using P7 and barcoded P5 index primers. Amplification products (130–300 bp) were size-selected using electrophoresis on a 2% agarose gel and purified using a MinElute Gel Extraction Kit (Qiagen, 28604) following the manufacturer’s protocol. LACE-seq libraries were subjected to single-end sequencing on an Illumina HiSeq 2500 platform by Novogene.
Raw sequence reads were first processed using Fastp (v0.23.4) for quality control66, after which poor-quality reads and adapter sequences were removed. Quality filtered reads were then mapped to the mm39 (Gencode, vM33) reference genome using STAR(v2.7.11b)67 with the parameters “ --alignEndsType Local --alignIntronMin 20 --alignIntronMax 20000 --outFilterMismatchNmax 2 --outFilterMultimapNmax 10 --alignMatesGapMax 700 --alignSJoverhangMin 10 --outFilterIntronMotifs RemoveNoncanonicalUnannotated.” Samtools(v1.19.2)67 was used to calculate the read coverage and depth. Duplicate PCR reads and reads in the blacklist regions68 were filtered using BedTools. The filtered reads were exported as a BED file for peak calling using Piranha (v1.2.1)69. Statistical significance was calculated based on a Zero-Truncated Negative Binomial distribution (parameters: “-s -p 0.001 -b 20”). Specifically, a one-sided (right-tailed) test was employed to identify regions with a significant enrichment of reads. To account for multiple comparisons across genomic bins, p values were adjusted using the Benjamini-Hochberg method, and a False Discovery Rate (FDR) threshold of <0.001 was applied. Subsequently, the peak regions were extensively annotated using the reference annotation dataset Gencode_vM33 with ChIPSeeker (v1.42.0)70. The genomic distribution frequency of annotated peaks across gene body features (including UTRs, 1st exons, and introns) was statistically analyzed using the “TxDb.Mmusculus.UCSC.mm39.refGene” database. Finally, findMotifsGenome.pl from HOMER (v4.11)71 was used with default parameters to identify known motifs within the peak regions. Additionally, the DeepTools(v3.5.4)72 bamCoverage function was used to generate the LACE-seq signal track files in the BigWig format. Subsequently, heatmaps of peak signals across all TSS were generated using the computeMatrix function and plotHeatmap from DeepTools, along with aligned peak signal heatmaps.
SCAN-seq single-cell amplification
GV oocytes from control and Hnrnpm cKO mice were collected using a mouth pipette and placed in a lysis buffer containing 0.1% Triton X-100 (Sangon, A110694-0100), dNTP mixture (Invitrogen, 00575484), barcode oligo (dT) primers, and an RNase inhibitor (Takara, 2313A). Immediately, the tubes underwent vigorous vortexing for 60 s and then incubated at 72 °C for 3 min to promote cell lysis and release of linearized RNA. Then, each sample was mixed with 2.83 μL of reverse transcription mixture, comprising SuperScript IV reverse transcriptase (Invitrogen, 18090050), RNase inhibitor (Takara, Cat. 2313A), SuperScript IV first-strand buffer, 1 M betaine (Sigma-Aldrich, B0300-1VL), 100 mM DTT (Invitrogen, 18090050), 50 mM MgCl₂ (Sangon, A610328-0500), and template-switching oligo primers. The reverse transcription reaction was executed under the conditions detailed below: 25 °C for 5 min, 42 °C for 60 min, 50 °C for 30 min, and 70 °C for 10 min. Subsequently, RT samples were mixed with 12.5 μL of PCR mixture containing ISPCR primers, the first-strand reaction product, 2× KAPA HiFi Hot-Start Ready Mix, and nuclease-free water. Amplification was performed under the following conditions: 98 °C for 20 s, followed by four cycles of 98 °C for 20 s, 65 °C for 30 s, and 72 °C for 5 min, and then 16 cycles of 98 °C for 20 s, 67 °C for 15 s, and 72 °C for 5 min, with a final extension at 72 °C for 5 min. Following PCR amplification, cDNA products from different cell barcodes were pooled and subjected to dual-round purification with 0.6× volume of AMPure XP beads. Finally, 1 μg of cDNA was used for subsequent library construction.
SCAN-seq library preparation and sequencing
The nanopore sequencing library was prepared using the Ligation Sequencing Kit 1D (Oxford Nanopore Technologies, ONT, SQK-LSK114), following the manufacturer’s instructions. Briefly, the cDNA fragments were subjected to end repair and dA-tailing using the Ultra II End Prep module. Subsequently, the dA-tailed cDNA fragments were ligated to the 1D adapter using the Quick Ligation Module. The resulting libraries were precisely quantified, quality-checked, and loaded onto a PromethION flow cell for sequencing on an Oxford Nanopore PromethION platform.
Bioinformatics analyses of SCAN-seq data
MinKNOW (v3.6.3) and Guppy (v3.1.5) were used to convert electrical signals from the Oxford Nanopore PromethION platform into FASTQ sequence data. Quality control was performed using Nanofilt (v2.8.0, https://github.com/wdecoster/nanofilt) and NanoPlot (v1.42.0, https://github.com/wdecoster) to filter out low-quality reads, defined as those with a Q-score <7 or length shorter than 100 bp. The remaining reads were identified, oriented, and trimmed using Pychopper (v2.7.10, https://github.com/nanoporetech/pychopper). Full-length reads were then extracted based on cell barcodes and PCR anchor sequences (AAGCAGTGGTATCAACGCAGAGTAC), followed by alignment to the mouse genome (Ensembl, GRCm39) using minimap2 (v2.25)73 with the parameters “-ax splice -uf --junc-bed.” The resulting BAM file was used for downstream analysis.
Differentially expressed genes in the full-length transcriptome were analyzed using the DESeq2 (v1.46.0)74 package with default parameters, screening thresholds set at padj < 0.05 and |log2(FoldChange)| > 1. GffCompare (v0.12.6)75 was used to detect unannotated novel transcripts across individual samples. The SUPPA2 (v2.4, https://github.com/comprna/SUPPA) was used to identify alternative splicing events in each sample. Differential alternative splicing events across samples were visualized using sashimi plots generated using ggsashimi (v1.1.5; https://github.com/guigolab/ggsashimi).
Statistics and reproducibility
GraphPad Prism 9 and RStudio were used for statistical evaluations. Differences between two groups were assessed using a two-sided unpaired Student’s t-test. Data are presented as mean ± SEM. ns not significant. *p < 0.05, **p < 0.01, ***p < 0.001. Detailed descriptions of the statistical tests used for each experiment are available in the respective figure legends. All experiments, including histology, electron microscopy, immunofluorescence, western blotting, immunoprecipitation, PCR, and qPCR, were executed with a minimum of three biological replicates with similar results. The complete, unprocessed images for all Western blots are provided in the Source data file and Supplementary Information.
Data availability
The LACE-seq (hnRNPM) and SCAN-seq (Hnrnpm Ctrl and cKO) data generated in this study have been deposited in the NCBI SRA database under accession code PRJNA1251287. SCAN-seq (Bcas2 Ctrl and cKO) data was from Prof. Lei Li12. The raw and unprocessed data presented in Figs. 1–8 and Supplementary Figs. 1–7 are provided in the Supplementary Information/Source data file.
References
Cheng, S. et al. Mammalian oocytes store mRNAs in a mitochondria-associated membraneless compartment. Science 378, eabq4835 (2022).
Wu, Y.-W. et al. RNA surveillance by the RNA helicase MTR4 determines volume of mouse oocytes. Dev. Cell 60, 85–100.e4 (2025).
Dai, X.-X. et al. PABPN1 functions as a hub in the assembly of nuclear poly(A) domains that are essential for mouse oocyte development. Sci. Adv. 8, eabn9016 (2022).
Chen, L. et al. NAT10-mediated mRNA N4-acetylation is essential for the translational regulation during oocyte meiotic maturation in mice. Sci. Adv. 11, eadp5163 (2025).
Barbosa-Morais, N. L. et al. The evolutionary landscape of alternative splicing in vertebrate species. Science 338, 1587–1593 (2012).
Keren, H., Lev-Maor, G. & Ast, G. Alternative splicing and evolution: diversification, exon definition and function. Nat. Rev. Genet. 11, 345–355 (2010).
Black, D. L. Mechanisms of alternative pre-messenger RNA splicing. Annu. Rev. Biochem. 72, 291–336 (2003).
Zhang, H. et al. Zygotic splicing activation of the transcriptome is a crucial aspect of maternal-to-zygotic transition and required for the conversion from totipotency to pluripotency. Adv. Sci. 11, 2308496 (2024).
Xia, M. et al. PCBP1 is required for maintenance of the transcriptionally silent state in fully grown mouse oocytes. Cell Cycle 11, 2833–2842 (2012).
Yu, L. et al. Loss of ESRP1 blocks mouse oocyte development and leads to female infertility. Development 148, dev196931 (2021).
Zhang, H. et al. The RNA ligase RNA terminal phosphate cyclase B regulates mRNA alternative splicing and is required for mouse oocyte development and maintenance. Development 149, dev200497 (2022).
Zhang, J. et al. BCAS2 is involved in alternative splicing and mouse oocyte development. FASEB J. 36, e22128 (2022).
Garalde, D. R. et al. Highly parallel direct RNA sequencing on an array of nanopores. Nat. Methods 15, 201–206 (2018).
Fan, X. et al. Single-cell RNA-seq analysis of mouse preimplantation embryos by third-generation sequencing. PLoS Biol. 18, e3001017 (2020).
Su, R. et al. Global profiling of RNA-binding protein target sites by LACE-seq. Nat. Cell Biol. 23, 664–675 (2021).
Dreyfuss, G., Matunis, M. J., Piñol-Roma, S. & Burd, C. G. hnRNP proteins and the biogenesis of mRNA. Annu. Rev. Biochem. 62, 289–321 (1993).
Wang, X. et al. hnRNPA2B1 represses the disassembly of arsenite-induced stress granules and is essential for male fertility. Cell Rep. 43, 113769 (2024).
Xiong, X. et al. hnRNPC functions with HuR to regulate alternative splicing in an m6A-dependent manner and is essential for meiosis. Adv. Sci. 12, e2412196 (2025).
Feng, S. et al. hnRNPH1 recruits PTBP2 and SRSF3 to modulate alternative splicing in germ cells. Nat. Commun. 13, 3588 (2022).
Wen, Y. et al. hnRNPU is required for spermatogonial stem cell pool establishment in mice. Cell Rep. 43, 114113 (2024).
Yin, L. et al. m6A reader hnRNPA2B1 modulates late pachytene progression in male meiosis through post-transcriptional control. Adv. Sci. 12, e06600 (2025).
Wang, X.-L., Li, J.-M. & Yuan, S.-Q. Characterization of the protein expression and localization of hnRNP family members during murine spermatogenesis. Asian J. Androl. 25, 314–321 (2023).
Li, H. et al. LSM14B is an oocyte-specific RNA-binding protein indispensable for maternal mRNA metabolism and oocyte development in mice. Adv. Sci. 10, 2300043 (2023).
Zheng, R. et al. hnRNPM protects against the dsRNA-mediated interferon response by repressing LINE-associated cryptic splicing. Mol. Cell 84, 2087–2103.e8 (2024).
Jin, H. et al. 5′-tRNAGly(GCC) halves generated by IRE1α are linked to the ER stress response. Nat. Commun. 15, 9273 (2024).
Lee, Y. F. et al. PARP4 interacts with hnRNPM to regulate splicing during lung cancer progression. Genome Med. 16, 91 (2024).
Wang, N. et al. FMRP protects breast cancer cells from ferroptosis by promoting SLC7A11 alternative splicing through interacting with hnRNPM. Redox Biol. 77, 103382 (2024).
Lin, Z. et al. Mettl3-/Mettl14-mediated mRNA N6-methyladenosine modulates murine spermatogenesis. Cell Res. 27, 1216–1230 (2017).
Gao, Z. et al. Zbed3 participates in the subcortical maternal complex and regulates the distribution of organelles. J. Mol. Cell Biol. 10, 74–88 (2018).
Qin, D. et al. The subcortical maternal complex protein Nlrp4f is involved in cytoplasmic lattice formation and organelle distribution. Development 146, dev183616 (2019).
Conti, M. & Franciosi, F. Acquisition of oocyte competence to develop as an embryo: integrated nuclear and cytoplasmic events. Hum. Reprod. Update 24, 245–266 (2018).
Jiang, Z.-Y. & Fan, H.-Y. Five questions toward mRNA degradation in oocytes and preimplantation embryos: When, who, to whom, how, and why?†. Biol. Reprod. 107, 62–75 (2022).
Kim, J. et al. Meikin is a conserved regulator of meiosis-I-specific kinetochore function. Nature 517, 466–471 (2015).
Meng, T.-G. et al. NLRP14 safeguards calcium homeostasis via regulating the K27 ubiquitination of Nclx in oocyte-to-embryo transition. Adv. Sci. 10, e2301940 (2023).
Yan, R. et al. Dynamics of DNA hydroxymethylation and methylation during mouse embryonic and germline development. Nat. Genet. 55, 130–143 (2023).
Zhang, W. et al. NLRP14 deficiency causes female infertility with oocyte maturation defects and early embryonic arrest by impairing cytoplasmic UHRF1 abundance. Cell Rep. 42, 113531 (2023).
Schober, C. S., Aydiner, F., Booth, C. J., Seli, E. & Reinke, V. The kinase VRK1 is required for normal meiotic progression in mammalian oogenesis. Mech. Dev. 128, 178–190 (2011).
Rong, Y. et al. ZAR1 and ZAR2 are required for oocyte meiotic maturation by regulating the maternal transcriptome and mRNA translational activation. Nucleic Acids Res. 47, 11387–11402 (2019).
Yk, W. & Hy, F. Revisiting ZAR proteins: the understudied regulator of female fertility and beyond. Cell. Mol. Life Sci. 79, 92 (2022).
Wu, Y.-K. et al. ZAR1 and ZAR2 orchestrate the dynamics of maternal mRNA polyadenylation during mouse oocyte development. Genome Biol. 26, 1–28 (2025).
Rong, Y. et al. ZAR1/2-regulated epigenetic modifications are essential for age-associated oocyte quality maintenance and zygotic activation. Adv. Sci. 12, e2410305 (2025).
Zheng, P. & Dean, J. Khdc3 Role of Filia, a maternal effect gene, in maintaining euploidy during cleavage-stage mouse embryogenesis. Proc. Natl. Acad. Sci. USA 106, 7473–7478 (2009).
Zhang, T. et al. CenpH regulates meiotic G2/M transition by modulating the APC/CCdh1-cyclin B1 pathway in oocytes. Development 144, 305–312 (2017).
Froment, P. et al. At the crossroads of fertility and metabolism: the importance of AMPK-dependent signaling in female infertility associated with hyperandrogenism. Hum. Reprod. 37, 1207–1228 (2022).
Rucker, E. B. et al. Bcl-x and Bax regulate mouse primordial germ cell survival and apoptosis during embryogenesis. Mol. Endocrinol. 14, 1038–1052 (2000).
Ho, J. S. et al. HNRNPM controls circRNA biogenesis and splicing fidelity to sustain cancer cell fitness. eLife 10, e59654 (2021).
Ramesh, N., Kour, S., Anderson, E. N., Rajasundaram, D. & Pandey, U. B. RNA-recognition motif in Matrin-3 mediates neurodegeneration through interaction with hnRNPM. Acta Neuropathol. Commun. 8, 138 (2020).
Yao, X. et al. BCAS2 regulates granulosa cell survival by participating in mRNA alternative splicing. J. Ovarian Res. 16, 104 (2023).
Yao, X. et al. BCAS2 regulates oocyte meiotic prophase I by participating in mRNA alternative splicing. FASEB J. 38, e23361 (2024).
Grote, M. et al. Molecular architecture of the human Prp19/CDC5L complex. Mol. Cell. Biol. 30, 2105–2119 (2010).
de Moura, T. R. et al. Prp19/Pso4 is an autoinhibited ubiquitin ligase activated by stepwise assembly of three splicing factors. Mol. Cell 69, 979–992.e6 (2018).
Chen, P.-H. et al. BCAS2 is essential for Drosophila viability and functions in pre-mRNA splicing. RNA 19, 208–218 (2013).
Yoshida, S. et al. Prc1-rich kinetochores are required for error-free acentrosomal spindle bipolarization during meiosis I in mouse oocytes. Nat. Commun. 11, 2652 (2020).
Zhou, C.-J. et al. CENP-F-dependent DRP1 function regulates APC/C activity during oocyte meiosis I. Nat. Commun. 13, 7732 (2022).
Lee, M. T., Bonneau, A. R. & Giraldez, A. J. Zygotic genome activation during the maternal-to-zygotic transition. Annu. Rev. Cell Dev. Biol. 30, 581–613 (2014).
Jentoft, I. M. A. et al. Mammalian oocytes store proteins for the early embryo on cytoplasmic lattices. Cell 186, 5308–5327.e25 (2023).
Xie, W. et al. Crucial roles of different RNA-binding hnRNP proteins in Stem Cells. Int. J. Biol. Sci. 17, 807–817 (2021).
Wen, Y. et al. hnRNPU in Sertoli cells cooperates with WT1 and is essential for testicular development by modulating transcriptional factors Sox8/9. Theranostics 11, 10030–10046 (2021).
Esposito, G. et al. Peptidylarginine deiminase (PAD) 6 is essential for oocyte cytoskeletal sheet formation and female fertility. Mol. Cell. Endocrinol. 273, 25–31 (2007).
Kim, B., Kan, R., Anguish, L., Nelson, L. M. & Coonrod, S. A. Potential role for MATER in cytoplasmic lattice formation in murine oocytes. PLoS ONE 5, e12587 (2010).
Li, L., Baibakov, B. & Dean, J. A subcortical maternal complex essential for preimplantation mouse embryogenesis. Dev. Cell 15, 416–425 (2008).
Tashiro, F. et al. Maternal-effect gene Ces5/Ooep/Moep19/Floped is essential for oocyte cytoplasmic lattice formation and embryonic development at the maternal-zygotic stage transition. Genes Cells 15, 813–828 (2010).
Xu, Q. et al. Maternal BCAS2 protects genomic integrity in mouse early embryonic development. Development 142, 3943–3953 (2015).
Sun, L. et al. BCAS2 and hnRNPH1 orchestrate alternative splicing for DNA double-strand break repair and synapsis in meiotic prophase I. Cell. Mol. Life Sci. 81, 449 (2024).
Lv, P. et al. HnRNPM modulates alternative splicing in germ cells by recruiting PTBP1. Reprod. Biol. Endocrinol. 23, 3 (2025).
Chen, S. Ultrafast one-pass FASTQ data preprocessing, quality control, and deduplication using fastp. Imeta 2, e107 (2023).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Amemiya, H. M., Kundaje, A. & Boyle, A. P. The ENCODE Blacklist: identification of problematic regions of the genome. Sci. Rep. 9, 9354 (2019).
Uren, P. J. et al. Site identification in high-throughput RNA-protein interaction data. Bioinformatics 28, 3013–3020 (2012).
Wang, Q. et al. Exploring epigenomic datasets by ChIPseeker. Curr. Protoc. 2, e585 (2022).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Ramírez, F. et al. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 44, W160–W165 (2016).
Li, H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics 34, 3094–3100 (2018).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Pertea, G. & Pertea, M. GFF utilities: GffRead and GffCompare. F1000Research 9, ISCB-Comm (2020).
Acknowledgements
The authors sincerely thank Prof. Minghan Tong (Shanghai Institute of Biochemistry and Cell Biology, Chinese Academy of Sciences) for generously providing the Stra8-Cre mice. The authors are also grateful to Prof. Lei Li (Institute of Zoology, Chinese Academy of Sciences) and Prof. Jiali Liu (College of Biological Sciences, China Agricultural University) for sharing the Bcas2 flox mice. The authors thank Prof. Lei Li and Dr. Jiaqi Zhang for sharing the source data of SCAN-seq12. The authors deeply appreciate all members of Prof. Heng-Yu Fan’s laboratory for their valuable discussions and constructive comments. The authors gratefully acknowledge Li Wang at the Center of Electron Microscopy, Zhejiang University, for expert technical assistance with electron microscopy imaging. The authors also thank Bioprofile for providing proteomics sequencing. The authors would like to thank Editage (www.editage.cn) for English language editing. The authors thank Ethan Tyler and Lex Kravitz for graphic support. All related schematics, derived from Servier Medical Art, SciDraw, and Figdraw, are used under a CC BY 4.0 license. This work was supported by the National Natural Science Foundation of China (Grants No. U24A20657 and 82371613 to F.S.) and the Key Research and Development Program of Zhejiang Province (Grant No. 2023C03035 to F.S.).
Author information
Authors and Affiliations
Contributions
S.Z., D.L., S.G., H.Z., H.X., Y.Z., H.Z., Z.D., X.W., C.L., and Y.L. conducted the bench experiments. R.S. constructed the Lace-seq library and performed sequencing. Z.Y. and S.Z. carried out bioinformatic analyses. H.F., S.Y., and J.Z. provided technical support and conceptual guidance. S.Z. wrote the manuscript. H.F. and F.S. revised it. F.S. supervised the study. All authors engaged in discussions regarding the results and conducted a critical review of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhou, S., Liu, D., Gan, S. et al. hnRNPM cooperates with BCAS2 to modulate alternative splicing during oocyte development. Nat Commun 17, 2681 (2026). https://doi.org/10.1038/s41467-026-69176-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-69176-8