Abstract
Focal cortical dysplasia (FCD), a major cause of drug-resistant epilepsy, involves abnormal neuronal migration and cortical architecture, yet its molecular basis remains poorly defined. Here, we identify FOXJ3 pathogenic variants in patients with autosomal dominant focal epilepsy and FCD. In the developing mouse cortex, FOXJ3 declines sharply in neural progenitors after embryonic day 15.5. In utero electroporation-mediated Foxj3 knockdown in mouse brains impairs neuronal migration, disrupts cortical lamination, and alters neuronal specification, promoting upper-layer neuron production at the expense of deeper-layer neurons. ChIP-seq and scRNA-seq analyses identify Pten as a key FOXJ3 target. Notably, Pten overexpression rescues cortical defects caused by FOXJ3 deficiency. FCD-associated variant fails to upregulate Pten, leading to dysregulated mTOR signaling and enlarged neuronal soma, a hallmark of FCD. These findings suggest that mutations in FOXJ3 may cause epilepsy and FCD and define a transcriptional program that regulates the PTEN-mTOR pathway for neuronal specification and cortical lamination.
Similar content being viewed by others
Introduction
In the cerebral cortex, the largest structure of the mammalian central nervous system (CNS), diverse excitatory neurons with distinct functions are organized into six laminar layers. Cortical neurogenesis follows an inside-out pattern, with earlier-born neurons forming deeper layers and later-born neurons populating the superficial layers. The development of this cytoarchitecture involves precise regulation across multiple stages1,2. Initially, neuroepithelial cells self-amplify, producing the first excitatory neurons that migrate to form the preplate, which later splits into Layer 1 and the subplate. Neuroepithelial cells then transform into radial glial cells (RGCs) and intermediate progenitor cells (IPCs)3,4, which generate the majority of cortical excitatory neurons in Layers 5/6, 4, and 2/3 sequentially5,6,7,8. These postmitotic neurons migrate through the intermediate zone (IZ) toward the cortical plate (CP) along radial fibers9,10, eventually reaching their designated layers and undergoing maturation to establish distinct connections and molecular profiles11. Despite advancements in understanding cortical development, the mechanisms underlying the specification of deep-layer versus upper-layer neurons remain a key unresolved issue.
The development of the cerebral cortex relies on a precisely regulated sequence of events, and disruptions in this complex process can lead to malformations of cortical development (MCD). Among these, focal cortical dysplasia (FCD) is one of the most frequently diagnosed forms in pediatric and young adult patients with epilepsy. FCD is characterized by abnormal neuronal organization and cortical architecture and classified based on histopathological criteria established by the International League Against Epilepsy (ILAE)12,13,14. FCD type II is defined by disrupted cortical lamination and distinctive cytological abnormalities, distinguishing type IIa (dysmorphic neurons without balloon cells) from type IIb (dysmorphic neurons with balloon cells)13,14. Magnetic resonance Imaging (MRI) remains the primary noninvasive diagnostic tool for FCD; however, subtle lesions may be overlooked, and FCD type II is still the most common etiology of drug-resistant epilepsy (DRE), even in MRI-negative cases13,15,16,17.
Advances in molecular genetics have revealed that epileptogenesis in FCD II is strongly linked to aberrant hyperactivation of the PI3K (phosphatidylinositol-3-kinase)-PTEN (phosphatase and tensin homolog)-mTOR (mechanistic target of rapamycin) pathway18. The PTEN-mTOR Pathway is a master regulator of cell proliferation, survival, and growth and plays a critical role in cerebral cortex development and synaptic plasticity. Genetically, FCD type II has been linked to mutations in genes that regulate the PI3K-PTEN-mTOR pathway, with PTEN acting as a key inhibitory factor. Dysregulation of this pathway, which can be due to pathogenic variations of PTEN or other genes involved in mTOR signaling, disrupts normal cortical development, leading to abnormal neuronal migration and differentiation19,20,21,22,23.
Transcription factors are pivotal regulators of cortical development, orchestrating neurogenesis, migration, and maturation. Among them, members of the forkhead box (FOX) family have been implicated in diverse biological processes, including neural development. The FOX family consists of approximately 50 genes in humans and 44 in mice, encoding transcription factors with a conserved winged-helix DNA binding domain, known as the FOX domain, and are divided into 19 subfamilies, from FOXA to FOXS24. Previous studies have shown that pathogenic variations in FOX genes can impact embryogenesis, including neural development25. For example, FOXG1 loss of function leads to FOXG1 syndrome, characterized by severe intellectual disability and abnormal movements26,27,28,29,30. The FOXJ family, comprising FOXJ1, FOXJ2, and FOXJ3 (Gene ID: 22887; OMIM:616035), is involved in various biological processes. FOXJ1 regulates ciliary and spinal cord development and controls left-right asymmetry31,32. FOXJ2 promotes meiosis in spermatogenesis, affects cell migration in glioma, and influences embryo development, leading to embryonic lethality33,34,35. FOXJ3 is primarily expressed in the neuroectoderm, neural crest, and telencephalic vesicle of mice during early development, but its precise function remains unclear36. Studies suggest FOXJ3 prevents embryonic stem cells from neural differentiation and regulates cell cycle in different cell lines37,38,39. FOXJ3 also plays a role in cell proliferation, migration, and the regulation of Snail expression and has been implicated in various cancers, including lung, prostate, breast, and colorectal cancer40,41,42. These findings highlight FOXJ3’s role in early embryonic development, particularly in regulating differentiation and proliferation.
Here, we identified heterozygous variants in FOXJ3 associated with autosomal dominant focal epilepsy linked to FCD. We found that FOXJ3 plays a crucial role in specifying deep-layer versus upper-layer neurons in mice. Foxj3 (Gene ID: 230700) is expressed in both NPCs within the VZ and neurons in the CP, with expression in NPCs showing a sharp decline after embryonic day (E) 15.5. FOXJ3 appears to regulate transcriptional programs and cell cycle dynamics, likely by modulating genes involved in cortical development. Notably, Pten emerged as a key downstream target of FOXJ3 in regulating neuronal fate determination during corticogenesis. Together, our data suggest that FOXJ3 orchestrates transcriptional programs and cell cycle regulation to control neuronal migration and deep-layer neuron specification via the PTEN-mTOR pathway, representing a pathogenic mechanism underlying focal epilepsy and FCD.
Results
Heterozygous FOXJ3 missense variants are associated with focal epilepsy related to focal cortical dysplasia
The index patient (DNA No. 50) presented with focal seizures at the age of 6 (Fig. 1A), with brain MRI revealing FCD in the right frontal gyrus (Fig. 1B). Due to DRE, the patient underwent resection surgery, and histopathology confirmed FCD type IIa (Fig. 1C). Notably, the patient’s mother (DNA No. 51) and cousin (DNA No.55) also had focal epilepsy and suspected FCD on MRI (right frontal FCD in the mother and right temporal FCD in the cousin). To identify the underlying genetic cause, whole-genome sequencing (WGS) was performed on the index patient (DNA No. 50), her cousin (DNA No.55), and their asymptomatic grandmother (DNA No.57). Through WGS and Sanger sequencing for intrafamilial segregation, we identified a heterozygous FOXJ3 variant (NM_014947.5:c.1052 A > G, p.Asn351Ser [p.N351S]) shared by the index patient, her mother and her cousin, which was considered the candidate causative variant (Fig. 1D, E). According to the SNP database (https://www.ncbi.nlm.nih.gov/snp/), the FOXJ3:p.Asn351Ser variant (rs1471785278) has a very low minor allele frequency (MAF) in public databases (0.000008/2 in TOPMED, and C = 0.00002/5 in GnomAD_exomes). In the Taiwan View database, which is specific to the Han-Taiwanese population (https://taiwanview.twbiobank.org.tw/), the MAF is 0.0003 in 1,492 WGS cases. In ClinVar (https://www.ncbi.nlm.nih.gov/clinvar/), it has been submitted only once, classified as a variant of uncertain significance, without associated clinical details.
A Pedigree of a family with focal epilepsy and focal cortical dysplasia harboring a FOXJ3 missense variant (c.1052 A > G; p.Asn351Ser). Filled symbols represent individuals affected by epilepsy (DNA #50, 51, and 55), while gray-filled symbols indicate carriers of the FOXJ3 p.N351S variant without epilepsy (DNA #56). The asterisk symbol denotes the index patient. Individuals selected for WGS are marked with green circles (DNA #50, 55, and 57). B Coronal section from serial brain MRIs of the 6-year-old FCD type II index patient (IV-4). The imaging highlights the blurring of the grey/white matter boundary (red box). C Representative histopathological images of the index patient showing FCD type IIa with cortical dyslamination (left, H&E stain, scale bar = 100 μm). Dysmorphic neurons with enlarged somata and nuclei, along with aggregates of Nissl substance, are evident (middle, H&E stain, scale bar = 50 μm). No balloon cells were identified. NeuN immunohistochemistry highlights the abnormal neuronal morphology (right, NeuN stain, scale bar = 50 μm). Similar histopathological features were consistently observed across all five independent sections examined. D Variants identified from WGS include 65,536 SNPs and 49,526 indels. Filtering for exonic, splicing, and non-synonymous variants retained 185 SNPs. Subsequent filtering for population frequency (<0.001) in public databases yielded 2 SNPs and 13 indels. Intrafamilial segregation analysis identified the FOXJ3 c.1052 A > G variant (p.N351S) as segregating with the epilepsy phenotype. E Electropherograms confirmed the heterozygous FOXJ3 variant (c.1052 A > G; p. Asn351Ser) in the index patient (DNA No. 50), and affected family members (DNA No. 51, 55, 56), but not in unaffected individuals (DNA No. 52). This finding was validated by Sanger sequencing.
We further screened FOXJ3 mutations in 170 patients with MCD and 76 patients with epilepsy without MCD on brain MRI at Taipei Veterans General Hospital in Taiwan. Additionally, we analyzed 85,000 WGS datasets from the Genomics England dataset, which primarily includes patients with rare diseases and their parents (https://www.genomicsengland.co.uk/initiatives/100000-genomes-project), as well as 30,000 whole-exome sequencing (WES) datasets from neurodevelopmental or neuromuscular patients registered at the Neurogenetics lab of UCL Institute of Neurology. The p.N351S variant was not detected in these datasets, but two additional heterozygous FOXJ3 missense variants were identified in two unrelated patients with focal epilepsy. FOXJ3 (NM_014947.5:c.1861A>G, p.Ile621Val [p.I621V]) was found in a male patient with focal epilepsy of unknown etiology, while FOXJ3 (NM_014947.5:c.757 C > A, p.Pro253Thr [p.P253T]) was identified in a male patient who developed focal epilepsy in his second decade. Brain MRI at 1.5 T in both patients was reported as essentially normal; however, it is important to note that subtle FCD lesions can be overlooked on standard MRI16.
Although in silico prediction tools provided conflicting evidence regarding the pathogenicity of these three FOXJ3 missense variants (Supplementary Fig. 1A), all are rare and affect highly conserved residues across multiple species (Supplementary Fig. 1B). These clinical-genetic observations suggest that heterozygous pathogenic variants in FOXJ3 are associated with focal epilepsy, potentially related to FCD. This prompted us to investigate the functional role of FOXJ3 in cortical development and its contribution to FCD.
FOXJ3 is expressed in cortical progenitors and neurons during cortical development
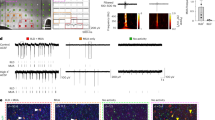
To investigate FOXJ3 expression during neocortical development, we collected protein lysates from mouse brain tissue at E11.5 to E18.5. FOXJ3 expression was consistently detected across these developmental stages, with a progressive increase from E11.5 to E18.5 (Fig. 2A, B). To further delineate its spatial expression, we performed immunofluorescence staining (IF) of FOXJ3 in mouse brain slices from E12.5 to E18.5, followed by confocal microscopy (Fig. 2C). FOXJ3 was localized to the nuclei of cells within the CP and the VZ. Notably, in the CP, FOXJ3 was highly expressed from E12.5 to E18.5. In contrast, in the VZ, FOXJ3 expression was strong from E12.5 to E14.5 but markedly declined after E15.5. Quantitative analysis of fluorescence intensity corroborated these observations, demonstrating sustained FOXJ3 expression in the CP and a significant reduction in the VZ post-E15.5 (Fig. 2D). Additionally, FOXJ3 immunostaining of primary cortical neurons from E14.5 mice at 8 days in vitro (DIV) confirmed the nuclear localization of FOXJ3 in neurons (Fig. 2E). Considering the pathogenesis of FCD mainly in early corticogenesis43, these findings suggest that FOXJ3 may play a crucial role in cortical progenitors during cortical development.
A Western blot analysis of FOXJ3 protein expression in the developing mouse cortex from E11.5 to E18.5. FOXJ3 expression was detected at all examined stages and increased progressively. B Bar graph showing relative FOXJ3 protein expression across developmental stages. Error bars represent S.E.M. (n = 3 independent experiments). C Immunostaining of cortical cryosections reveals FOXJ3 expression (gray, red) in the VZ and CP from E12.5 to E18.5. FOXJ3 levels in the VZ was high before E14.5 and declined sharply after E15.5. Sparse expression of FOXJ3 was observed in the IZ, while CP expression persisted. Brain slices were counterstained with DAPI (blue) to show cell nuclei. Scale bars: 50 μm (top panels), 10 μm (lower panels). D Bar graph showing normalized FOXJ3 fluorescence intensity in VZ. Error bars represent S.E.M. **: p < 0.01, ****: p < 0.0001 (n = 3 independent experiments; one-way ANOVA with post-hoc Dunnett test; p = 0.0025, p < 0.0001, 0.0001, 0.0001). E Primary culture cortical neurons from E14 mouse embryos were infected with GFP (green) and immunostained with anti-FOXJ3 antibody (magenta) and DAPI (blue). Scale bar: 10 μm.
Foxj3 knockdown at E14.5 mildly impaired neuronal migration but did not affect cortical lamination or cell fate determination
To elucidate the roles of FOXJ3 in cortical development, especially in cortical progenitors, we utilized RNA interference (RNAi) to knock down Foxj3 expression (Fire et al., 1998). Short hairpin RNA (shRNA) targeting the Foxj3 coding region (shFoxj3) was delivered via lentivirus to primary cortical cells isolated from E16 mouse embryos at DIV3. After 6 days in culture, shFoxj3-infected cortical neurons exhibited significantly reduced FOXJ3 protein levels compared to control neurons infected with scrambled shRNA (shCtrl) (shFoxj3: 29.1 ± 6.8 %, n = 3; p < 0.001, Student’s t-test), indicating effective (> 70 %) Foxj3 knockdown (Supplementary Fig. 2A).
Next, we assessed the impact of Foxj3 knockdown on the developing neocortex using in utero electroporation (IUE) of shRNAs together with a GFP reporter into mouse embryonic neocortex at E14.5 (Supplementary Fig. 2B). Four days post electroporation, the majority of the GFP+ cells were distributed in the CP (69.4% ± 2.4%, n = 5), with only a few cells in the IZ (19.3% ± 1.4%, n = 5) and VZ/SVZ (11.3% ± 1.5%, n = 5) in control brains electroporated with shCtrl. However, in the brains with Foxj3 knockdown, a significant migration delay was observed, with fewer GFP+ cells in the CP (53.1% ± 1.3%, n = 3; p < 0.01, Student’s t-test) and more in the IZ (38.0% ± 2.2%, p < 0.001; n = 3, Student’s t-test), while the proportion of GFP+ cells in the VZ/SVZ remained unchanged (8.8% ± 0.9%, n = 3; Supplementary Fig. 2B).
To further evaluate FOXJ3’s role during late cortical development, we analyzed brains at postnatal day (P) 7 following IUE at E14.5 (Supplementary Fig. 2C). In control brains electroporated with shCtrl, the majority of GFP+ cells localized to Layer 2/3. Similarly, GFP+ cells in shFoxj3 brains were positioned in Layer 2/3 (Supplementary Fig. 2D, E). Since neurons born at E14.5 are destined for Layer 2/3 and primarily differentiated as BRN2+ and SATB2+ callosal projection neurons8, we examined whether Foxj3 knockdown affected this molecular profile. Immunostaining revealed that the majority of GFP+ cells expressed upper-layer gene BRN2 and SATB2 in both control and Foxj3 knockdown brains (BRN2 + : 93.0% ± 1.5%, n = 3 control vs. 97.3% ± 0.9%, n = 3 shFoxj3; SATB2 + : 84.3% ± 6.7%, n = 3 control vs. 98.0% ± 1.1%, n = 3 shFoxj3) (Supplementary Fig. 2D, E). These results suggest that while FOXJ3 dysfunction may impair neuronal migration during early cortical development, it does not appear to influence the generation or fate of upper-layer callosal projection neurons.
FOXJ3’s critical role in cell fate determination and cortical lamination in deep-layer and Layer 4 neurons
Given that FOXJ3 dysfunction affects neuronal migration without altering the fate of later-born neurons, we explored whether similar phenotypes manifest in earlier-born neurons. Therefore, we conducted IUE of shFoxj3 along with GFP at E12.5. Samples were collected at P7 to examine the identity of the targeted cells (Fig. 3A). In control brains, GFP+ cells electroporated at E12.5 predominantly populated Layer 5/6 of the neocortex at P7. Strikingly, in Foxj3 knockdown brains, most GFP+ cells were found in the superficial layers of the CP. To assess the fates of these GFP+ cells, we performed immunofluorescence staining using cortical layer markers, including TBR1 (Layer 6 neuron marker), CTIP2 (layer 5 neuron marker), and SATB2 (upper layer neuron marker). In control brains, ~40% of GFP+ cells expressed TBR1 (38.1 ± 0.5 %, n = 3), and around 20% expressed CTIP2 (17.8 ± 1.8 %, n = 3; Fig. 3A, B). Additionally, GFP signals in the ventral cerebrum indicated the presence of subcortical projections (Fig. 3C), confirming the deep-layer neuronal identity of these GFP+ cells. Foxj3 knockdown resulted in a significant reduction in TBR1+ and CTIP2+ cells compared to control brains (TBR1: 16.4 ± 2.6 %, n = 3, p < 0.01; CTIP2: 0.8% ± 0.4%, n = 3, p < 0.001, Student’s t-test). Conversely, the SATB2+ cell population significantly increased under Foxj3 knockdown conditions (Ctrl: 26.7 ± 1.0 %, n = 3; shFoxj3: 46.6 ± 3.3 %, n = 3; p < 0.01, Student’s t-test; Fig. 3A, B). Supporting these molecular expression changes, subcortical projections were reduced (Fig. 3C). These findings suggest that FOXJ3 dysfunction impairs the acquisition of deep-layer neuronal fate and induces the production of SATB2+ callosal projection neurons.
A Electroporation of shCtrl or shFoxj3, together with the GFP (green) construct, was performed at E12.5; brains were harvested at P7. Immunostaining of P7 cortices for cortical neuron markers TBR1 (magenta), CTIP2 (red), and SATB2 (red). Enlarged views show colocalization of GFP+ cells with these markers. Brain slices were counterstained with DAPI (blue) to indicate nuclei. Scale bars: 200 μm (top panels), 50 μm (lower panels). B Bar graphs show the percentage of GFP+ cells colocalized with TBR1, CTIP2, or SATB2. Error bars represent S.E.M. **: p < 0.01, ***: p < 0.001 (n = 3 independent experiments; two-tailed unpaired t-test; p = 0.0012, 0.0008, 0.0043). C Axon projection patterns of GFP+ cells (green) electroporated with shCtrl or shFoxj3. DAPI (blue) was used to counterstain cell nuclei. Scale bars: 400 μm (top panels), 100 μm (lower panels). D Electroporation of shCtrl or shFoxj3 with GFP at E13.75 with fixation at P7. Immunostaining for cortical layer markers RORβ (red) and BRN2 (red) in P7 cortices. Enlarged panels show colocalization of GFP+ cells (green) with layer markers. Brain slices were counterstained with DAPI (blue) for nuclei. Scale bars: 200 μm (top panels), 50 μm (lower panels). E Quantification of GFP+ cells colocalized with RORβ, and BRN2. Error bars represent S.E.M. ***: p < 0.001, ****: p < 0.0001 (n = 3 independent experiments; two-tailed unpaired t-test; p < 0.0001, p = 0.0006) F Callosal projection patterns of GFP+ cells (green) after electroporation. Scale bars: 750 μm (top panels), 100 μm (lower panels). G Quantified intensity of callosal projections of GFP+ cells. Error bars represent S.E.M. *: p < 0.05 (n = 3 independent experiments; two-tailed unpaired t-test; p = 0.0184).
To further investigate the effects of FOXJ3 dysfunction on Layer 4 neurons, we performed IUE to knock down Foxj3 at E13.75. Samples were collected at P7 and subjected to immunofluorescence staining using RORB (Layer 4 neuron marker) and BRN2 antibodies (Fig. 3D). In control brains, the majority of GFP+ cells localized in Layer 4 and expressed RORB (69.9 ± 3.1 %, n = 3), consistent with previous findings44. In brains electroporated with shFoxj3, the GFP+ cells shifted to the superficial Layer 2/3, with a significant reduction in the RORB+ cell population (9.5 ± 1.0 %, n = 3; p < 0.0001, Student’s t-test). Instead, most of these Foxj3-knockdown GFP+ cells expressed BRN2 (84.1 ± 2.2 %, n = 3), compared to control brains (21.9 ± 5.9 %, n = 3; p < 0.001, Student’s t-test; Fig. 3D, E). Consistent with these molecular expression changes, we observed a 3-fold increase in GFP signals in the corpus callosum of brains with Foxj3 knockdown (Fig. 3F, G). These results indicate that FOXJ3 dysfunction impairs Layer 4 cell fate acquisition and promotes the production of BRN2+ callosal projection neurons. Collectively, our findings suggest that FOXJ3 dysfunction promotes callosal neuron production, regardless of the timing the genetic manipulation.
FOXJ3 deficiency leads to severe delays in neuronal migration
Cortical lamination defects often arise from dysregulated neuronal migration in postmitotic neurons45,46. Given that Foxj3 knockdown induces the formation of upper callosal projection neurons (Fig. 3), we investigated whether Foxj3 knockdown also impairs neuronal migration, thereby affecting cell fate. We collected brain sections 3 days after electroporation at E12.5 and stained them with TBR1 antibody (Fig. 4A). In control brains, most GFP+ cells had entered the IZ and reached the CP (CP: 50.7% ± 1.9%, IZ: 23.6% ± 1.5%, VZ: 25.7% ± 0.9%, n = 3). However, in Foxj3-knockdown brains, a significant proportion of GFP+ cells failed to enter the CP and remained restricted to the IZ (CP: 24.7 ± 1.7%, p < 0.001; IZ: 47.9 ± 3.6%, p < 0.01, VZ: 27.4% ± 1.9%; n = 3, Student’s t-test). This migration defect persisted in the brains 5 days after electroporation (E12.5-E17.5; Fig. 4B), where the majority of GFP+ cells in control brains had entered the CP (CP: 49.6 ± 0.8%; IZ: 29.5 ± 1.1%; VZ: 20.9 ± 1.9% in shCtrl vs. CP: 28.8 ± 3.9%, p < 0.01; IZ: 37.3 ± 2.9%, VZ: 33.9 ± 6.7% in shFoxj3; n = 3 in each group, Student’s t-test). Similar migration defects were observed in E16.75 brains following electroporation at E13.75 (Fig. 4C). In control brains electroporated with shCtrl, most GFP+ neurons had entered the CP (CP: 84.1 ± 3.4%, IZ: 12.0 ± 1.8%, VZ: 3.9 ± 1.7%, n = 3). In contrast, GFP+ cells in Foxj3-knockdown brains were still confined to the IZ and VZ (CP: 22.1 ± 4.8 %, p < 0.001; IZ: 67.6 ± 6.6 %, p < 0.01; VZ: 10.3 ± 1.9 %; n = 3, Student’s t-test). Collectively, these results indicate that FOXJ3 deficiency induces significant neuronal migration delays, ultimately leading to changes in cell fate during early cortical development.
A Neuronal migration following Foxj3 knockdown was analyzed in brains 3 days post-electroporation (E12.5-E15.5) with the cortical neuron layer marker TBR1 (red). DAPI (blue) counterstaining was used to visualize cell nuclei. In control brains, most GFP+ cells (green) migrated through the IZ and reached the CP. In Foxj3-knockdown brains, a significant proportion of GFP+ cells failed to enter the CP, remaining restricted to the IZ. The stacked bar graph quantifies the distribution of GFP+ cells in the CP, IZ, and SVZ/VZ. Error bars represent S.E.M. **: p < 0.01, ***: p < 0.001 (n = 3 independent experiments; two-tailed unpaired t-test; p = 0.0033, 0.0005). B Neuronal migration was assessed 5 days post-electroporation (E12.5-E17.5) with TBR1 (red) and DAPI (blue) staining. In control brains, most GFP+ cells (green) migrated into the CP, whereas in Foxj3-knockdown brains, many GFP+ cells remained in the IZ and SVZ/VZ. The stacked bar graph quantifies GFP+ cell distribution. Error bars represent S.E.M. **: p < 0.01, ***: p < 0.001 (n = 3 independent experiments; two-tailed unpaired t-test; p = 0.00124, 0.00044). C Neuronal migration was analyzed 3 days post-electroporation (E13.75-E16.75). In control brains, most GFP+ neurons (green) migrated into the CP, but in Foxj3-knockdown brains, GFP+ cells were predominantly located in the IZ and SVZ/VZ. The stacked bar graph quantifies GFP+ cell distribution. Scale bars: 100 μm. Error bars represent S.E.M. **: p < 0.01 (n = 3 independent experiments, two-tailed unpaired t-test; p = 0.006394).
The essential role of FOXJ3 in neural stem cells for cell fate determination and cortical lamination
Given that FOXJ3 is expressed in both progenitors and cortical neurons (Fig. 2), we sought to determine whether FOXJ3 exerts its regulatory effects during the progenitor stage or the postmitotic neuron stage. To address this question, we employed a doxycycline-inducible knockdown system to temporally control FOXJ3 expression during cortical development (Supplementary Fig. 3A, B). By electroporating inducible shRNA constructs into mouse brains at E12.5 or E13.75 and subsequently administering doxycycline at E15 or E15.75, respectively (Supplementary Fig. 3C, D), we selectively knocked down Foxj3 once the targeted cells had exited the cell cycle. Samples were harvested at P7 for analysis (Supplementary Fig. 4A, C). As anticipated, GFP+ cells in control brains differentiated into deeper-layer or Layer 4 neurons expressing TBR1 (47.0 ± 3.0%, n = 3) and RORB (86.0 ± 1.4%, n = 3). Induced Foxj3 knockdown in postmitotic cells did not alter their cell fate, as these cells maintained their identities as deeper-layer (48.5 ± 1.6%, n = 3) and Layer 4 (85.4 ± 2.2%, n = 3) neurons similar to controls (Supplementary Fig. 4). These findings indicate that FOXJ3 regulates cell fate acquisition during the progenitor stage.
Foxj3 knockdown increases neural stem cell maintenance and delays cell cycle exit
Previous studies have highlighted the critical role of FOXJ3 in regulating cell cycle progression38,39,47. Nonetheless, its specific role in controlling the cell cycle of cortical progenitors during cortical development remains unclear. To explore this, we electroporated shCtrl or shFoxj3 constructs into embryonic brains at E12.5 and analyzed the proportion of SOX2+ cortical progenitors among GFP+ cells two days post-electroporation (Fig. 5A). In control brains electroporated at E12.5, approximately 25% of GFP+ cells remained undifferentiated SOX2+ progenitors at E14.5 (18.4 ± 0.6 %, n = 4). However, in the Foxj3 knockdown brains, we observed a significant increase in the proportion of SOX2+ cells (30.1 ± 4.6 %, n = 3; p < 0.05, Student’s t-test; Fig. 5A). A similar increase in the SOX2+ progenitor population was observed in brains electroporated with shFoxj3 at E13.75 and collected at E15.75 (shCtrl: 24.6 ± 1.1 %, n = 3; shFoxj3: 34.2 ± 2.5%, n = 4; p < 0.05, Student’s t-test; Fig. 5B). In addition, we access whether this effect of delayed neuronal differentiation extended to IPCs by performing EOMES/TBR2 immunostaining at E15.5 in embryonic brains electroporated at E12.5 (Supplementary Fig. 5). Foxj3 knockdown significantly increased the proportion of EOMES/TRB2+ IPCs. These results together indicate that FOXJ3 delays neuronal differentiation not only in RGCs but also in IPCs.
A Electroporation of shCtrl or shFoxj3 with GFP (green) at E12.5, followed by fixation at E14.5 and immunostaining for the neural stem cell marker SOX2 (red). The bar graph quantifies SOX2 + /GFP+ cells, showing that Foxj3 knockdown significantly increased the proportion of SOX2+ NSCs. Scale bars: 50 μm. Error bars represent S.E.M. *: p < 0.05 (shCtrl: n = 4, shFoxj3: n = 3 independent experiments; two-tailed unpaired t-test; p = 0.0241) B Similar analysis was performed at E13.75, with fixation at E15.75 and immunostaining for SOX2 (red). Scale bars: 50 μm. Error bars represent S.E.M. *: p < 0.05 (shCtrl: n = 3, shFoxj3: n = 4 independent experiments; two-tailed unpaired t-test; p = 0.0259). C Cell cycle analysis following electroporation at E12.5 and EdU labeling at E13.5, with fixation at E14.5. Ki67 (red) and EdU (blue) immunostaining identified cells exiting the cell cycle (EdU + /Ki67- cells). The bar graph quantifies EdU+ki67- as a proportion of EdU+ cells, showing that Foxj3 knockdown significantly reduced cell cycle exit. Scale bars: 50 μm. Error bars represent S.E.M. **: p < 0.01 (n = 3 independent experiments; two-tailed unpaired t-test; p = 0.0075) D Similar analysis was performed in brains electroporated at E13.75 and fixed at E15.75. Scale bars: 50 μm. Error bars represent S.E.M. **: p < 0.01 (n = 3 independent experiments; two-tailed unpaired t-test; p = 0.0075).
Given that an increased progenitor population could result from cell cycle re-entry, we next assessed this possibility by administering EdU intraperitoneally one day post-electroporation and collecting brains one day later for EdU and Ki67 staining (Fig. 5C, D). As the cortical progenitor cell cycle lasts approximately 14 hours48, the presence of EdU+ cells that are negative for Ki67 one day after EdU administration suggests that these cells have exited the cell cycle. In control brains electroporated at E12.5 and E13.75, we found that most GFP+ cells were EdU- (GFP+EdU-/GFP + : E12.5, 60.75 ± 7.2 %, n = 3; E13.75, 58.0 ± 2.6 %, n = 3), and most GFP+EdU+ cells were ki67- (GFP+EdU+Ki67-/GFP+EdU + : E12.5, 62.6 ± 5.1 %, n = 3; E13.75, 57.0 ± 1.8 %, n = 3), consistent with previous reports indicating cell cycle exit48,49. In contrast, shFoxj3-electroporated brains exhibited a significant decrease in the proportion of GFP+EdU+Ki67- cells (GFP+EdU+Ki67-/GFP+EdU + : E12.5, 36.6 ± 1.2 %, n = 3; E13.75, 41.4 ± 2.5 %, n = 3; p < 0.01, Student’s t-test), indicating that Foxj3 knockdown impairs cell cycle exit. In contrast, Foxj3 knockdown at E14.5 did not lead to significant changes in neuronal differentiation or cell cycle re-entry (Supplementary Fig. 6), suggesting a specific role for FOXJ3 in progenitors during earlier corticogenesis. Furthermore, FOXJ3 overexpression in progenitors electroporated at E12.5 and E13.75 significantly reduced the percentage of SOX2+ cells (Supplementary Fig. 7), indicating its influence on neuronal differentiation. Collectively, our results suggest that FOXJ3 regulates progenitor cell fate by modulating cell cycle progression.
FOXJ3 is involved in transcriptional programs and cell cycle regulation
The FOX family of transcriptional factors is known to play critical roles in regulating cortical development through their transcriptional regulatory functions27,29,50. To further understand FOXJ3’s role in this process, we sought to identify potential downstream genes directly targeted by FOXJ3 that could regulate cortical development. Using chromatin immunoprecipitation followed by high-throughput sequencing (ChIP-seq) in FOXJ3-stable expressing Neuro2A cell lines (clones #1 and #2; Supplementary Fig. 8), we identified 39,576 and 41,853 peaks in FOXJ3 clones #1 and #2, respectively (Fig. 6A). To identify potential FOXJ3-targeted genes active in NPCs, we applied two criteria: First, we cross-referenced FOXJ3-occupied genomic loci with potential active regulatory elements marked by H3K27ac51. Second, we focused on FOXJ3 binding sites close to RNA polymerase II (Pol-II) targeted transcription start site (TSS, from −500 bp to +500 bp; Supplementary Fig. 8). We identified 14,341 FOXJ3-H3K27ac co-occupied peaks (Fig. 6B), of which 6,244 peaks were located near TSS regions (Supplementary Fig. 8C).
A Overlapping binding regions shared between FOXJ3 clone #1 peaks and FOXJ3 clone #2 peaks. B Overlapping binding regions shared between FOXJ3-overlapping peaks and H3K27ac peaks. C GO enrichment analysis of potential FOXJ3 target genes was performed using DAVID, with terms ranked by -log(p-value). D scRNA-seq data were used to identify potential FOXJ3 downstream target genes based on expression patterns similar to Foxj3. Identified genes include Pten, Tsc1, Pafah1b1, Cep120, Dicer1, Nde1, Wdr62 and Mcph1. E Enriched peaks for the candidate genes Pten, Tsc1, Pafah1b1, Cep120, Dicer1, Nde1, Wdr62, and Mcph1 were co-localized with FOXJ3 #1, FOXJ3 #2, H3K27ac, and RNA Pol-II. DNA input was used as the control. F Bar graph showing significant upregulation of Pten, Tsc1, and Pafah1b1 mRNA levels in cortical tissue overexpressing human FOXJ3. In contrast, overexpression of the p.N351S variant failed to upregulate these downstream target genes, indicating a loss-of-function effect. Error bars represent S.E.M. **: p < 0.01 ***: p < 0.001 (n = 3 independent experiments; one-way ANOVA with post-hoc Fisher’s LSD test; p = 0.0065, 0.0025, 0.0007, 0.0007, 0.002, 0.0004). G qPCR analysis of transcriptional changes in downstream genes Pten and Tsc1 upon FOXJ3 expression. CRISPR/Cas9-mediated Foxj3 knockout in mouse embryonic stem (mES) cells resulted in decreased expression of Pten but not Tsc1. Error bars represent S.E.M. *: p < 0.05, (n = 3 independent experiments; one-way ANOVA with post-hoc Fisher’s LSD test; p = 0.0123, 0.0143, 0.0294, 0.0172, 0.0309, 0.0406). H UMAP clustering from scRNA-seq from the human fetal brain dataset57 visualized cell clusters based on cell type, with expression patterns of FOXJ3, PTEN, and TSC1 mapped and visualized.
To explore the functions of these 6,244 FOXJ3-targeted genes, we conducted a gene ontology (GO) term analysis using the Database for Annotation, Visualization, and Integrated Discovery (DAVID; https://david.ncifcrf.gov/). The top 10 GO terms in biological processes included Cell cycle; Protein transport; Transcription, DNA-templated; Cellular response to DNA damage stimulus; mRNA processing; DNA repair; Cell division; Regulation of transcription, DNA-templated; RNA splicing; and Mitotic nuclear division (Fig. 6C). These findings suggest that FOXJ3 predominantly regulates transcriptional programs and cell cycle, both of which are crucial for cortical development.
Potential FOXJ3 target genes are involved in cerebral cortex development
Our findings suggest that FOXJ3 plays a role in regulating the cell cycle during cortical development. To identify potential FOXJ3 target genes implicated in this process, we curated a candidate gene list encompassing genes annotated to both the Cell cycle (GO:0007049) and Pallium development (GO:00219543) Gene Ontology terms. This identified 29 genes associated with cerebral cortex development, including Tsc1 (tuberous sclerosis 1), Pten (phosphatase and tensin homolog), Wdr62 (WD repeat domain), Cep120 (centrosomal protein 120), Mcph1 (microcephaly, primary autosomal recessive 1), Pafah1b1 (platelet-activating factor acetylhydrolase, isoform 1b, subunit 1; also known as Lis1), Dicer1 (dicer 1, ribonuclease type III), Bax (BCL2-associated X protein), and Nde1 (nude neurodevelopment protein 1) among others (Supplementary Table 1).
If these genes are indeed regulated by FOXJ3 and involved in the FOXJ3-mediated pathway during cortical development, their expression profiles should theoretically be co-expressed in similar cell populations. To access this, we utilized our single-cell RNA sequencing (scRNA-seq) dataset on cells isolated from the dorsal pallium at E13.5 and E15.5 (Supplementary Fig. 9)52. Cells were categorized into progenitors (i.e., RGC), precursors (i.e., IP), and neurons (Supplementary Fig. 9A) based on the expression of neural cell markers, such as Pax6, Emx1, Eomes, Neurod6, Tbr1, and Satb2 (Supplementary Fig. 9B). Examination of FOXJ3-targeted gene expression profiles revealed conserved expression patterns in either progenitor or neuron cell types for several genes, including Pten, Tsc1, Pafah1b1, Cep120, Dicer1, Nde1, Wdr62, and Mcph1 (Fig. 6D, Supplementary Fig. 10). Taken together, these analyses refined the list of potential FOXJ3 target genes to the 5 most confident candidates: Pten, Tsc1, Pafah1b1, Cep120, and Dicer1. These findings suggest that FOXJ3 may play a significant role in cortical development through the regulation of its targeted genes involved in cell cycle control, providing insights into the FOXJ3-deficient phenotypes observed at the progenitor stage (Figs. 4, 5).
To validate that FOXJ3 binding sites from ChIP-seq align with active promoter regions, we visualized the peaks using Integrative Genomics Viewer (IGV) software (Fig. 6E). The sharpness of the peaks indicated specific DNA binding, as observed when transcription factors interact with their binding sites53. Active promoter and enhancer regions, marked by histone modification H3K27ac, typically exhibit sharp peaks54, while active promoter regions bound by RNA Pol-II appear as broader regions55,56. Applying these criteria, we identified peaks within the same loci as potential FOXJ3 target genes. Among the 29 candidate FOXJ3 targets selected using GO analysis, Pten, Tsc1, Pafah1b1, Cep120, Dicer1, Nde1, Wdr62, and Mcph1 met these criteria (Fig. 6E). To prevent potential mislabeling, we further analyzed whether FOXJ3 directly binds to genes commonly used as markers for cell identity, including Tbr1, Ctip2, Satb2, Rorb, Brn2, Sox2, and Mki67. Our analysis revealed no significant peaks in the promoter regions of these genes (Supplementary Fig. 11A), indicating that FOXJ3 does not directly regulate their expression.
To further confirm the association between FOXJ3 and the candidate downstream genes PTEN and TSC1 in developing human neocortical cells, we performed an in silico scRNA-seq analysis using a published dataset57. The analysis revealed that the expression pattern of FOXJ3 in human cortical cells mirrors that observed in mouse cortical cells, with enriched expression in progenitors and specific neuron cell types (Fig. 6H). Additionally, PTEN and TSC1 displayed conserved expression patterns consistent with those identified in mice (Fig. 6H). These findings suggest that FOXJ3 plays conserved roles in regulating cortical development in both mice and humans.
PTEN’s role in FOXJ3-mediated functions during cortical development
To confirm FOXJj3’s actual regulation of gene transcription within the identified target list, we employed RT-qPCR to assess the mRNA expression of Pten, Tsc1, Pafah1b1, Cep120, and Dicer1 following FOXJ3 overexpression in vivo. IUE of FOXJ3 fused with mCherry was conducted at E12.5, and the mCherry+ region in the dorsal pallium was isolated at E14.5 for RNA extraction. A significantly increased mRNA level of FOXJ3 was confirmed (Supplementary Fig. 11B). RT-qPCR analysis of the candidate FOXJ3 target genes revealed significant upregulation in Pten, Tsc1, and Pafah1b1 upon FOXJ3 overexpression, suggesting that FOXJ3 can activate the expression of Pten and Tsc1 (Fig. 6F). Notably, the FCDII-associated variant p.N351S abolished this transcriptional activation, indicating a loss of regulatory function (Fig. 6F). To further validate FXOJ3’s role in regulating Pten and Tsc1 expression, we performed RT-qPCR following CRISPR Cas9-mediated Foxj3 knockdown in mouse embryonic stem cells. Under these conditions, the mRNA level of Pten, but not Tsc1, showed a significant decrease under Foxj3 knockdown conditions (Fig. 6G).
Given that PTEN is a critical inhibitor of the mTOR pathway19,20,21,22,23, we next investigated whether FOXJ3 loss influences mTOR signaling. We assessed the phosphorylation of ribosomal protein S6 (pS6), a key downstream effector of the mTOR pathway (Supplementary Fig. 12). Neuronal cells transfected with shFoxj3 exhibited reduced PTEN expression accompanied by a robust increase in S6 phosphorylation. These findings demonstrate that FOJX3 regulates the mTOR pathway through transcriptional control of Pten, and that FOXJ3 dysfunction results in aberrant mTOR hyperactivation.
Next, we investigated the involvement of PTEN and TSC1 in FOXJ3-mediated mechanisms regulating cortical development. We performed a rescue experiment to determine whether the expression of Pten or Tsc1 could mitigate the phenotypes caused by FOXJ3 dysfunction. First, we evaluated the maintenance of progenitor identity and the percentages of cell cycle exit two days after IUE at E12.5 (Fig. 7A, B). Co-electroporation with shFoxj3 and a Pten-expressing construct resulted in a reduced proportion of cells remaining as SOX2+ cortical progenitors (shFoxj3: 30.9 ± 4.6 %; shFoxj3+Pten: 12.3 ± 0.9%; Fig. 7A, C) and an increased proportion of cells exiting the cell cycle at E14.5 (shFoxj3: 36.6 ± 1.2 %; shFoxj3+Pten: 68.3 ± 2.3 %; Fig. 7B, D), compared to non-rescued Foxj3 knockdown brains.
A Electroporation of shCtrl or shFoxj3 or shFoxj3 + Pten constructs along with GFP (green) was performed at E12.5, with brains fixed at E14.5, and Immunostaining for the neural stem cell marker SOX2 (red) in E14.5 cortices. Scale bar: 50 μm. B EdU was injected at E13.5, and brain tissues were fixed at E14.5. Immunostaining for Ki67 (red) and EdU (blue) to assess cell cycle exit in E14.5 cortices. EdU + /Ki67- cells were considered exited from the cell cycle. Scale bar: 50 μm. C Bar graph showing the percentage of SOX2 + /GFP+ cells among GFP+ cells, indicating enhanced NSC maintenance with Foxj3 knockdown, and restoration with Pten rescue. Error bars represent S.E.M. *: p < 0.05, **: p < 0.01 (shCtrl: n = 4, shFoxj3: n = 3, shFoxj3+Pten: n = 3 independent experiments; one-way ANOVA with post-hoc Tukey test; p = 0.0179, 0.0032). D The bar graph shows that Foxj3 knockdown reduced the cell cycle exit (EdU + /Ki67-) percentage, which is restored by PTEN expression. Error bars represent S.E.M. **: p < 0.01, ***: p < 0.001 (shCtrl: n = 3, shFoxj3: n = 3, shFoxj3+Pten: n = 4 independent experiments; one-way ANOVA with post-hoc Tukey test; p = 0.002, 0.0004) (E) Foxj3 knockdown disrupted neuronal migration (GFP+ cells in the IZ), while co-expression with PTEN restored cortical layer distribution, confirmed by TBR1+ immunostaining (red). Scale bar: 100 μm. F Bar graph showing GFP+ cells distributed across the CP, IZ, and SVZ/VZ regions. PTEN expression increased the proportion of cells in the CP compared to the shFoxj3 group. Error bars represent S.E.M. **: p < 0.01 (n = 3 independent experiments; one-way ANOVA with post-hoc Tukey test; p = 0.0023). G At P7, Foxj3 knockdown resulted in a shift toward SATB2+ (red) cell fate. PTEN expression effectively rescued this phenotype, restoring TBR1 (magenta) and CTIP2 (red) marker expression. Scale bar: 200 μm. H Bar graph showing the percentage of SATB2, CTIP2, and TBR1 markers colocalized with GFP+ cells. Error bars represent S.E.M., **: p < 0.01, ***: p < 0.001, ****: p < 0.0001 (TBR1: shCtrl: n = 3, shFoxj3: n = 3, shFoxj3+Pten: n = 3 independent experiments; one-way ANOVA with post-hoc Tukey test; p = 0.0001, p < 0.0001; CTIP2: shCtrl: n = 4, shFoxj3: n = 3, shFoxj3+Pten: n = 3 independent experiments; one-way ANOVA with post-hoc Tukey test; p = 0.0023, 0.0032; SATB2: shCtrl: n = 4, shFoxj3: n = 3, shFoxj3+Pten: n = 3 independent experiments; one-way ANOVA with post-hoc Tukey test; p = 0.0004, 0.0004).
Moreover, the neuronal migration defect observed in Foxj3-knockdown brains was dramatically reversed by Pten overexpression, with significantly more GFP+ cells reaching the CP by E15.5 (CP: 24.7 ± 1.7%; IZ: 48.5 ± 3.0%; VZ: 27.4 ± 1.9% in shFoxj3 versus CP: 37.3 ± 0.5%; p < 0.01, one-way ANOVA, IZ: 42.2 ± 1.5%, VZ: 20.6 ± 1.0% in shFoxj3 + Pten; n = 3 in each group; Fig. 7E, F). Additionally, Pten overexpression in the Foxj3 knockdown condition reversed the cell fate alterations observed at P7 (Fig. 7G, H). In Pten-rescued Foxj3 knockdown brains, the proportion of GFP+ cells expressing TBR1 was significantly increased compared to Foxj3-knockdown brains (shFoxj3: 16.4 ± 2.6%, n = 3; shFoxj3 + Pten: 41.2 ± 0.4%, n = 3; p < 0.0001, one-way ANOVA), while the proportion of GFP+ cells expressing SATB2 was significantly decreased (shFoxj3: 46.6 ± 3.3%, n = 3; shFoxj3 + Pten: 25.8 ± 1.3%, n = 3; p < 0.001, one-way ANOVA; Fig. 7G, H).
Consistent with these findings, Pten knockdown alone recapitulated the neuronal migration delay observed in Foxj3-deficient brains (Supplementary Fig. 13A, B), further supporting that FOXJ3 and PTEN act within the same pathway during cortical development. In contrast, overexpression of Tsc1 in the Foxj3-knockdown condition did not result in significant changes in cell fate (Supplementary Fig. 13C,D). Taken together, these results demonstrate that Pten, rather than Tsc1, is a critical downstream gene of FOXJ3, playing a pivotal role in regulating cortical progenitors during cortical development.
p.N351S variant increased neuronal somal size in adolescent mice
Because FCDII pathology is characterized by dysmorphic neurons and balloon cells, we next investigated the effects of the FCDII-associated FOXJ3 variant on neuronal morphology. We electroporated embryonic brains at E14.5 with either wild-type (WT) FOXJ3 or the p.N351S variant and allowed the mice to develop until P28, corresponding to adolescence (Fig. 8). Neuronal morphology was then examined, and somal dimensions were quantified. Neurons expressing the p.N351S mutant exhibited significantly enlarged somata, as evidenced by increased somal area, diameter, and perimeter, compared with WT FOXJ3-expressing neurons (Fig. 8B, C), while both WT and mutant FOXJ3 localized properly to the nucleus (Fig. 8D). These findings suggest that the p.N351S variant disrupts FOXJ3 function, potentially through dysregulation of the mTOR pathway, thereby driving abnormal neuronal hypertrophy reminiscent of the dysmorphic neurons and balloon cells observed in FCDII.
A Electroporation of WT FOXJ3 or mutant FOXJ3 (N351S) expressing constructs with GFP was performed at E14.5, and brains were harvested at P28. B Representative confocal images of GFP-expressing neurons in the cortical plate, with corresponding segmentation masks generated using Cellpose. Scale bars: 200 μm. Higher-magnification views (yellow boxes) showed detailed cell morphology. Scale bars: 30 μm. C Quantification of neuronal soma morphology. Violin plots comparing the soma area (µm²), diameter (µm), and perimeter (µm) of neurons expressing WT FOXJ3 and mutant FOXJ3 (N351S). The N351S mutation leads to a significant increase in all three morphological parameters. Error bars represent S.E.M. **p < 0.01, ***p < 0.001, ****: p < 0.0001 (n = 3 independent experiments; two-tailed unpaired t-test). D Nuclear localization of WT and mutant FOXJ3 (N351S) in cortical neurons at P28. High-magnification confocal images of cortical neurons labeled with GFP (green, electroporated cells), mCherry–FOXJ3 (red; WT or N351S mutant), and DAPI (blue; nuclei). Both WT and N351S FOXJ3 proteins were predominantly localized to the nucleus, indicating that the N351S substitution does not affect nuclear targeting. Scale bars: 20 μm.
Discussion
In this study, we identified FOXJ3 as a potential causative gene for focal epilepsy and FCD. We also report the pivotal role for the transcription factor FOXJ3 in the regulation of corticogenesis, particularly through its control of the PTEN-associated pathway (Fig. 9). FOXJ3 is expressed in progenitor cells within the VZ/SVZ and in postmitotic neurons within the CP during cortical development in both humans and mice. Notably, there is a significant decrease in FOXJ3 expression in the VZ/SVZ after E15.5. Our findings reveal that the downregulation of FOXJ3 at E12.5 and E13.75 causes sustained proliferation and delays neurogenesis. Additionally, we observed delayed neuronal migration at E15.5 and E16.75, followed by altered neuronal cell fate at P7 under Foxj3 knockdown conditions. This phenotype appears to be primarily driven by FOXJ3’s direct regulation of genes involved in cell cycle control during brain development, as indicated by our combined ChIP-seq and transcriptomic analyses. Furthermore, we identified Pten as a FOXJ3-targeted gene capable of rescuing the phenotypes induced by FOXJ3 deficiency in the developing neocortex. Knockdown of Pten in the developing mouse cortex phenocopied the effect of Foxj3 knockdown. Importantly, Foxj3 knockdown resulted in reduced PTEN expression and increased pS6 levels, further supporting its role in regulating the mTOR signaling pathway. Moreover, the FCD-associated variant p.N351S failed to induce Pten expression and led to enlarged neuronal soma size in adolescent mice, consistent with FCDII pathology. Together, these findings provide compelling evidence that FOXJ3 is a critical regulator of cortical lamination and neuronal specification during brain development and contributes to the pathogenesis of FCD.
A During early cortical development, FOXJ3 is highly expressed in RGCs (blue cells), promoting cell cycle exit and differentiation into deep-layer neurons (layers IV-VI, red and green cells). After E15.5 in mice, FOXJ3 expression declines in RGCs, coinciding with the developmental switch to the generation of upper-layer neurons (layer II/III, yellow cells). Mechanistically, FOXJ3 transcriptionally enhances PTEN expression in progenitor cells, thereby facilitating cell cycle exit, neuronal differentiation, and migration. B In contrast, FOXJ3 dysfunction disrupts this process, resulting in impaired migration and premature production of upper-layer neurons (yellow cells) at the expense of deep-layer neurons (red and green cells). Loss of FOXJ3 reduces PTEN expression in progenitors, leading to hyperactivation of the mTOR pathway and increased phosphorylation of ribosomal protein S6. These changes culminate in delayed cell cycle exit, altered neuronal fate specification, and defective migration (dark green cells). Pathogenic mutations in FOXJ3 are therefore associated with FCD and epilepsy, mediated by impaired regulation of PTEN and consequent dysregulation of the PTEN-mTOR pathway, which also contributes to enlarged neuronal soma resembling dysmorphic neurons in FCD patients. Together, this model highlights FOXJ3 as a key transcription factor orchestrating neuronal specification, laminar organization, and cortical integrity during brain development.
Maintaining a balance between the progenitor pool and neuron production is crucial during cortical development to establish a mature neocortex with a sufficient number and proper composition of projection neurons for the six-layered structure58. Various signaling pathways, such as the PI3K-AKT pathway, have been identified as key regulators of progenitor behavior59,60. PI3K converts phosphatidylinositol 4,5-bisphosphate (PIP2) into phosphatidylinositol 3,4,5-trisphosphate (PIP3), activating AKT at the cell membrane61. Activated AKT promotes cell survival and growth by inhibiting pro-apoptotic signals such as GSK3, which in turn derepresses mTOR signaling62. PTEN dephosphorylates PIP3 back to PIP2, thereby inhibiting AKT activation and balancing proliferation and differentiation63. In humans, mutations in PTEN are associated with cortical dysplasia64 and autism spectrum disorder65, while its overexpression in human brain organoids leads to microcephaly-like phenotypes66. In mice, Pten deficiency induces brain enlargement, seizures, and ataxia67. These data indicated that the precise expression level of PTEN is crucial in cortical development.
In our study, both gain- and loss-of-function experiments demonstrate that FOXJ3 is necessary and sufficient to activate Pten expression in cortical progenitors. In addition, Pten overexpression rescues the Foxj3-knockdown phenotypes, such as cell proliferation, further ensuring the proper production of neuronal subtypes. Taken together, our data identify FOXJ3 as an upstream regulator of PTEN, crucial for modulating PI3K-AKT signaling in cortical progenitors, thereby maintaining the balance between progenitor proliferation and differentiation to support neocortical development.
Projection neurons within the six-layered neocortex exhibit distinct projection patterns and molecular profiles. For example, corticothalamic projection neurons (CThPN) express TBR1, subcerebral projection neurons (SCPN) express CTIP2, Layer 4 granular neurons express RORB, and callosal projection neurons (CPN) express SATB2 and/or BRN268,69. While it is known that these neuronal subtypes are produced sequentially during neocortical development8, the precise molecular mechanisms governing the timing of neuronal production and fate determination remain not fully understood. Neuronal fate can be determined either at the progenitor stage or postmitotically through intrinsic and/or extrinsic pathways70,71,72. In this study, we identified FOXJ3 as a critical regulator of cortical development. FOXJ3 is expressed both in cortical progenitors and postmitotic neurons. In the mouse neocortex, its expression is elevated in cortical progenitors within the VZ/SVZ from E12.5 to E14.5, followed by a marked decline after E15.5. Knocking down Foxj3 in progenitor cells at E12.5 and E13.75, which are expected to generate deep-layer and Layer 4 neurons, induced cell cycle re-entry, causing these cells to remain as progenitors. This delayed their differentiation at later stages, ultimately resulting in the production of upper-layer neurons. Nonetheless, the same manipulation in E15.5 progenitors, which are destined to produce upper-layer neurons, had no significant effect. Together with FOXJ3’s endogenous expression pattern, our data suggest that FOXJ3 levels in progenitors are important for the temporal switch from deep-layer to upper-layer neuron production, highlighting its stage-specific functions in corticogenesis.
The identification of FOXJ3 mutations in patients with FCD and epilepsy prompted us to utilize IUE to mimic FOXJ3 loss of function during cortical development. In Foxj3 knockdown brains, we observed an increase in the number of GFP+ cells expressing the upper-layer neuronal marker BRN2, resulting in the forced generation of upper-layer neurons. These alterations in layer specification may contribute to the characteristic abnormal cortical lamination observed in FCD. However, seizures were not observed in electroporated mice, likely due to the limited proportion of cells affected by Foxj3 knockdown inherent IUE experiments. Future studies using knockout mouse models may help further elucidate the physiological consequences of FOXJ3 dysfunction in vivo. Mutations in genes involved in PI3K-AKT signaling have been associated with FCD in clinical reports, and activation of the PI3K-AKT pathway has been observed in FCD patients. Given that FOXJ3 regulates corticogenesis through its downstream target PTEN, it is plausible that FOXJ3 mutations could lead to FCD. Previous studies have identified somatic point mutations in PIK3CA, which induce FCD and hemimegalencephaly (HME) in clinical cases73,74. A point-mutant mouse model of PIK3CA showed increased expression of upper-layer neuronal markers, such as CUX1 and SATB2, in the mutant cortex75, consistent with our findings in FOXJ3 knockdown. The pathogenic mechanisms of PIK3CA, which involve PI3K-AKT signaling76, bear similarities to our current hypothesis regarding FOXJ3, suggesting that these pathways may be implicated in the pathogenesis of FCD.
Some carriers of heterozygous FOXJ3 variants exhibited non-lesional brain MRI or did not present with clinical seizures. These observations suggest that FOXJ3-associated FCD may be underdiagnosed due to incomplete penetrance of the autosomal-dominant variants and the presence of invisible lesions on conventional MRI. Notably, FCD represents the most common etiology among individuals with focal epilepsy and negative MRI findings15. Advances in imaging, including optimized sequences, post-processing approaches, and ultra-high-field 7 T MRI, have improved detection of subtle FCD lesions, though many remain undetectable16. A deeper understanding of the FOXJ3-mediated transcriptional regulation on cortical development may help explain epileptogenesis in cases where epileptogenic lesions are not visible on MRI. Our study proves convergent lines of evidence linking FOXJ3 dysfunction to the PTEN-mTOR pathway, a central pathogenic mechanism in FCD. Given that somatic mosaic second-hit mutations are frequently implicated in FCD type II, we performed paired high-depth WES on DNA extracted from resected FCD tissue and peripheral blood. However, we did not identify significant somatic variants in FOXJ3 or other genes within the PTEN-mTOR pathway (Supplementary Table 2).
In conclusion, this study identifies FOXJ3 as a critical regulator of cortical lamination and neuronal fate specification during early neurogenesis. FOXJ3 variants are associated with focal epilepsy and FCD, providing a genetic etiology for these disorders. Through the regulation of the PTEN-mTOR pathway, FOXJ3 controls the migration and differentiation of deeper-layer and Layer 4 neurons, and its dysfunction leads to cortical malformations that likely underlie the pathogenesis of FCD. These findings enhance our understanding of cortical development and offer potential avenues for the diagnosis and treatment of epilepsy and cortical malformations.
Methods
Ethical approval
The human study was performed under the approval of the Institutional Review Board (IRB) of Taipei Veterans General Hospital (IRB 2018-01-003 C, 2020-02-007 A, 2023-02-014B). All animal experiments and maintenance were carried out under the guidelines and protocols approved by the Institutional Animal Care and Use Committee (IACUC) of National Yang Ming Chiao Tung University.
Human genomic DNA extraction and purification
Genomic DNA was extracted from the peripheral blood with the approval of the patients and their family members. 10 c.c. of blood was drawn from each subject into 10 c.c. sodium citrate tubes containing Monovettes (Sarstedt, Germany). Whole blood samples were gently mixed under room temperature on the same day of blood draw. Then, the blood samples were submitted for DNA extraction and genotyping. Genomic DNA will be isolated by using the PUREGENE DNA purification systems (Gentra Systems, Minnesota, USA).
Whole genome sequencing and Sanger sequencing
The library preparation of WGS was performed using Nextera DNA Flex Library Prep Kit (Illumina) according to the manufacturer’s protocol. The quality and size distribution of each exome-enriched DNA library were analyzed by Bioanalyzer (Agilent). The amount of each library was quantified using both Qubit Fluorometer (Invitrogen) and quantitative PCR. The enriched DNA library was subjected to massively parallel sequencing with 500-bp paired-end reads. The WGS was performed by using the Illumina NovaSeq 6000 sequencers at a read depth of 30x at the Institute of Molecular and Genomic Medicine, National Health Research Institutes. Sanger sequencing was repeated to validate the identified variants.
DNA extraction from fresh frozen brain tissues
Thin sections (10–30 μm) were cut from fresh frozen brain tissue using a microtome, ensuring a total tissue weight of no more than 25 mg. DNA was extracted with the QIAamp DNA Mini Kit (QIAGEN) according to the manufacturer’s instructions with minor modifications. Briefly, tissue samples were homogenized with a pestle in a 1.5 ml microcentrifuge tube containing 180 μl Buffer ATL, followed by the addition of 20 μl Proteinase K (Thermo Scientific, Waltham, MA). Samples were vortexed and incubated at 56 °C until complete is lysis. Subsequently, 200 μl Buffer ATL was added, and samples were incubated at 70 °C for 10 min. After addition of 200 μl ethanol (96-100%), the mixture was applied to a QIAamp Mini Spin Column and centrifuged at 6000 x g for 1 min. Columns were washed subsequently with 500 μl Buffer AW1 (6000 x g, 1 min) and 500 μl Buffer AW2 (20,000 x g, 3 min). DNA was eluted with 200 μl Buffer AE after 1 min incubation at room temperature for 1 min, followed by centrifugation at 6000 x g for 1 min.
Identifying somatic mutations by WES
WES was performed on DNA extracted from both fresh frozen brain tissue (lesion) and peripheral blood leukocytes (matched normal control). Sequencing data were analyzed using the Sentieon® Genomics– software suite (Sentieon Inc.), following a Lesion-Normal analysis pipeline. Raw FASTQ files were aligned to the human reference genome (GRCh38) using DRAGEN (v4.2.4) with the corresponding reference bundle (hg38 v3, Illumina). Duplicate reads were marked, and base quality scores were recalibrated as part of the DRAGEN pipeline. Somatic single nucleotide variants and structural variants were called using DRAGEN’s tumor-normal analysis mode, restricted to the target regions defined by the SureSelect Clinical Research Exome V2 (GRCh38) BED file. Annotation was performed with VEP (v112.0), incorporating ClinVar, gnomAD, dbSNP, and the Taiwan Biobank. Variants were filtered and prioritized based on functional impact and allele frequency. All genes were considered in the analysis, with focused interrogation of FOXJ3 and genes within the PTEN-mTOR pathway (Supplementary Table 2).
Protein extraction and western blot
The samples were treated with RIPA lysis buffer containing 10% protease and 10% phosphatase inhibitor, followed by sonication to lyse the cells and brain tissues. The resulting protein lysate was then centrifuged at 4 °C, 15,000 x g for 15 min, and the supernatant was collected. Protein quantification was performed using the BCA assay kit (Thermo Fisher Scientific, USA), after which equal amounts of protein were separated by electrophoresis on 8% SDS-PAGE and transferred to a PVDF membrane (Immobilon, Millipore). The membrane was incubated in the blocking buffer (0.1% T-TBS containing 5% skim milk and 3% Bovine Serum Albumin, BSA) for 1 h. To detect specific proteins, the membrane was then incubated with primary antibodies overnight at 4 °C. The antibodies used, along with their concentrations, were as follows: FOXJ3 (Affinity Biosciences, AF0602, 1:1000, https://www.affbiotech.com/goods-1139-AF0602-FOXJ3_Antibody.html), PTEN (Cell Signaling Technology, 9559S, 1:3000, https://www.cellsignal.com/products/primary-antibodies/pten-138g6-rabbit-mab/9559), TSC1 (Cell Signaling Technology, 4906S, 1:3000, https://www.cellsignal.com/products/primary-antibodies/hamartin-tsc1-antibody/4906), S6 (Cell Signaling Technology, 2217S, 1:3000, https://www.cellsignal.com/products/primary-antibodies/s6-ribosomal-protein-5g10-rabbit-mab/2217), pS6(235/236) (Cell Signaling Technology, 4858S, 1:3000, https://www.cellsignal.com/products/primary-antibodies/phospho-s6-ribosomal-protein-ser235-236-d57-2-2e-xp-rabbit-mab/4858), GAPDH (Proteintech, 60004-1-Ig, 1:10,000, https://www.ptglab.com/products/GAPDH-Antibody-60004-1-Ig.htm) and Beta-actin (Proteintech, 66009-1- Ig, 1:10,000, https://www.ptglab.com/products/Pan-Actin-Antibody-66009-1-Ig.htm). After washing the membrane three times, it was incubated with horseradish peroxidase (HRP)-conjugated secondary antibodies: anti-mouse (GeneTex, 1:10,000) and anti-rabbit (Sigma Aldrich, 1:10,000). Protein signals on the membrane were visualized using ECL-Plus reagent (Millipore) and the Amersham Imager 680 (GE). The intensities of protein bands were analyzed using ImageJ software (version 2.17.0, NIH).
Animal model and in utero electroporation (IUE)
IUE was performed following established protocols in previous reports52,77,78,79,80,81,82,83. Timed pregnant ICR mice (BioLASCO, Taiwan) at E12.5, E13.75, and E14.5 were anesthetized with isoflurane. The skin and muscular tissue were incised to expose the uterine horns from the abdomen. Plasmid DNA was injected into one side of the lateral ventricles of the embryos using a glass micropipette. Forceps-type electrodes (5 mm in diameter, Harvard Apparatus) were used for electroporation. Electric pulses delivered at different embryonic stages were as follows: 4 × 30 V pulses, 50 ms duration, 950 ms interval at E12.5; 5 × 35 V pulses, 50 ms duration, 950 ms interval at E13.75; 5 × 40 V pulses, 50 ms duration, 450 ms interval at E14.5. Following electroporation, embryos were carefully returned to the abdominal cavity, and the incision was closed using a surgical suture. Brains of electroporated mice were harvested at E14.5, E15.5, E15.75, E16.75, E17.5, E18.5 or P7. In each animal, multiple embryos were electroporated to minimize individual variations. Thus, even with n = 3 animals, the dataset provides sufficient statistical power, while minimizing animal use is consistent with ethical guidelines. The welfare of the pregnant mice was carefully monitored in accordance with the guidelines from the Institutional Animal Care and Use Committee (IACUC) of National Yang Ming Chiao Tung University (No. 1080626).
Mouse brain sectioning
To obtain brain samples, mice were first perfused with phosphate-buffered saline (PBS, PH7.4) and then fixed with 4% paraformaldehyde (PFA). The brains were collected immediately and immersed in 1% or 2% PFA at 4 °C overnight. For vibratome sectioning, the brains were embedded in 4% low-melting agarose (AMRESCO) dissolved in PBS. The brains were then coronally sectioned into 100 μm-thick slices using a vibrating microtome (Leica, VT1000S) and stored in PBS with 0.05% sodium azide (Sigma) at 4 °C. For cryostat sectioning, the brains were placed in 30% sucrose at 4 °C overnight before being embedded in Tissue-Tek® O.C.T. Compound (Sakura Finetek). The brains were then sectioned coronally into 12 μm-thick slices using a cryostat machine (Thermo Fisher Scientific) and stored at -80 °C.
Immunofluorescence staining
For immunostaining of cryosections, the slices underwent three 10 min wash each in 0.05% PBST. Brain sections were then blocked by incubating them in 0.1% PBST containing 10% normal goat serum (NGS) for 1 h. The sections were then immersed with primary antibodies in the blocking solution at 4 °C overnight. After three washes with 0.05% PBST, the sections were incubated with secondary antibodies in the blocking solution at room temperature for 2 h. Nuclear staining with 4’,6-diamidino-2-phenylindole (DAPI, Molecular Probes) was conducted at room temperature for 15 min. All steps were carried out in the dark. The coverslips were mounted with antifade mounting medium (VECTASHIELD® Vibrance™) and stored at 4 °C until imaging. For immunostaining of vibratome sections, the slices were permeabilized in PBS containing 0.2% Triton X-100 (PBST) for 30 minutes. Then, the brain sections were soaked in 0.2% PBST with 10% NGS and 5% BSA, followed by incubation with primary antibodies in 0.2% PBST containing 5% NGS and 5% BSA at 4 °C for 2–3 days. After six PBS washes, the sections were incubated with secondary antibodies in 0.2% PBST containing 2.5% NGS and 2.5% BSA at room temperature for 2 hours. DAPI was used for nuclear staining, and the sections were incubated overnight at 4 °C. All steps were performed in the dark, and the coverslips were mounted with antifade mounting medium (VECTASHIELD® Vibrance™) and stored at 4 °C until imaging.
The primary antibodies used for staining were: FOXJ3(1:500; Affinity Biosciences, AF0602, https://www.affbiotech.com/goods-1139-AF0602-FOXJ3_Antibody.html), PTEN (1:500; CST, 9559S, https://www.cellsignal.com/products/primary-antibodies/pten-138g6-rabbit-mab/9559), TSC1 (1:500; CST, 4906S, https://www.cellsignal.com/products/primary-antibodies/hamartin-tsc1-antibody/4906), TBR1 (1:500; Abcam, ab31940, https://www.abcam.com/tbr1-antibody-ab31940.html), TBR2 (1:500; Abcam, ab23345, https://www.abcam.com/tbr2--eomes-antibody-ab23345.html), CTIP2 (1:500; abcam, ab18465, https://www.abcam.com/ctip2-antibody-25b6-ab18465.html), SATB2(1:500; Santa Cruz, sc-81376, https://www.scbt.com/p/satb2-antibody-satba4b10?srsltid=AfmBOoplp-XvgR2LrqcLV7-bOVwlS5fcz1YvopqXU5SArKzYQdh3kwRG), RORB (1:500; R&D, PP-N7927, https://www.rndsystems.com/products/human-ror-beta-antibody-clone-n7927-n7927_pp-n7927-00), BRN2 (1:500; Santa Cruz, sc-393324, https://www.scbt.com/p/brn-2-antibody-b-2?srsltid=AfmBOoqF0knd1AuP6tdlihlYYGTy9nZs42L90W0Uig6ejDTOejhy7rG3), Ki-67 (1:500; Abcam, ab15580, https://www.abcam.com/ki67-antibody-ab15580.html), SOX2 (1:500; Millipore, AB5603, https://www.merckmillipore.com/TW/zh/product/Anti-Sox2-Antibody,MM_NF-AB5603?ReferrerURL=https%3A%2F%2Fwww.google.com%2F), GFP (1:1000; Abcam, ab13970, https://www.abcam.com/gfp-antibody-ab13970.html). The secondary antibodies used were as follows: goat anti-mouse, anti-rabbit, anti-chicken or anti-rat Alexa Fluor-546, Alexa Fluor-488, and Alexa Fluor-647 conjugated antibodies (1:500; Invitrogen).
Cell culture and transfection
The Neuro-2a (N2A, purchased from Taiwan Bioresource Collection and Research Center, BCRC) cell line was cultured in Dulbecco’s Modified Eagle Medium (DMEM) supplemented with 10% fetal bovine serum (FBS) and low glucose (Life Technologies, USA) and maintained at 37 °C in a humidified atmosphere with 5% CO2. For transfection of constructs, N2A cells were seeded in 3.5 cm Petri dishes at 70% density. Transfection was performed using the T-pro Non-liposomal Transfection Reagent II (T-Pro Biotechnology, Taiwan) following the manufacturer’s instruction, and cells were then incubated for 24 to 48 h to conduct further experiments.
To assess the knockdown efficiency of shRNA, primary cultured cortical neurons were isolated from the cerebral cortex of ICR mouse embryos at E16. These cultured neurons were infected by lentivirus packaged with shRNA constructs designed and produced by the RNAi core of Academia Sinica at DIV 2, and the virus was removed after 24 h by replacing the medium. Proteins from primary cultures were extracted at DIV 6 based on a previous report84.
Constructs
Knockdown constructs of Foxj3 were based on the pLKO-TRC011 vectors. A sequence targeting Foxj3 CDS (shFoxj3; CCACAGTACAACCTTCCAGAA) was used for Foxj3 knockdown and a non-targeting sequence scramble (shCtrl; CCTAAGGTTAAGTCGCCCTCG) as a control. All knockdown constructs were purchased from the Academia Sinica RNAi core (Taiwan). For conditional Foxj3 knockdown experiments, we utilized an inducible knockdown system set up in a previous study85. Control (shCtrl) and Foxj3 (shFoxj3) shRNA sequence was subcloned into pInducer10-mir-RUP-PheS (addgene #44011) vector. The US2-GFP construct was a generous gift from Dr. Jenn-Yah Yu (National Yang Ming Chiao Tung University, Taiwan). For overexpression of FOXJ3, we subcloned the human FOXJ3 gene (NCBI ref seq: NM_014947.5) from a CMV-promoter driven plasmid (Sino Biological Inc.) into the pCAGGS vector (Addgene #41583). The pCAGGS vector is driven by the CMV enhancer chicken β-actin promoter, with mCherry or Flag tagged on the N terminus of FOXJ3. Site-directed mutagenesis was performed using the QuikChange Site-Directed Mutagenesis Kit (Agilent Technologies, USA) to generate the human FOXJ3 p.N351S variant. The forward primer (5’-CAA CCA AAG CAG CCT GTC CAG CAG TCA TGG CAG TGG CCT CA-3’) and reverse primer (5’-GA GGC CAC TGC CAT GAC TGC TGG ACA GGC TGC TTT GGT TG-3’) were designed to introduce the nucleotide substitution.
To disrupt Foxj3 gene expression in mouse embryonic stem cells, we induced double-strand breaks in the Foxj3 gene locus using the CRISPR/Cas9 system through the following sgRNAs (http://chopchop.cbu.uib.no)86: two pairs of sgRNAs for Foxj3 knockout: gRNA1: GCTAGACAGATAAATCACATAGG, gRNA3: CTCATCTGTTAAATATTATGCGG targeting sequence before Foxj3 start codon and gRNA2: GAGTCTCACCTTGAAACTCACGG, gRNA4: AGTGCAGTTTGGTGGAGTAGGGG targeting sequence after Foxj3 start codon. All sgRNAs were cloned into the backbone from the Multiplex CRISPR/Cas9 Assembly System (addgene, #1000000055) following the manufacturer’s instructions87. To generate Pten overexpression constructs, we subcloned the mouse Pten gene (NCBI ref seq: NM_008960.2) from the TRE promoter plasmid (add gene #135663) into the pCAGGS vector. For the overexpression of Tsc1 (NCBI ref seq: NM_001289575.1), we extracted genomic DNA from mouse tissues and used RT-PCR to amplify the Tsc1 coding sequence (CDS). The full-length Tsc1 CDS was then purified by gel extraction and cloned into the pCAGG vector. The sequences were verified by the protein BLAST algorithm from the National Center for Biotechnology Information (NCBI).
Cell-cycle exit analysis
24 h after the IUE experiment, EdU (5-ethynyl-2’-deoxyuridine, Thermo Fisher Scientific) was intraperitoneally (IP) injected into the maternal mice, with a dosage of 50 mg per kilogram of mouse body weight. Embryonic brains were then harvested and fixed. After immunostaining for Ki-67, the brain slices were stained with Click-iT™ EdU Alexa Fluor™ 647 imaging Kit (C10339, Thermo Scientific) for 30 minutes. The cell-cycle exit index (%) was calculated as the ratio of Ki67-/EdU + /GFP+ signals to the number of electroporated GFP + /EdU+ cells.
Image acquisition and quantification
Fixed brain slices were imaged using Zeiss LSM880 or LSM700 inverted confocal microscopes equipped with a 20× Plan-Apochromat NA 0.8 air objective for tissue imaging and a 100× NA 1.4 oil-immersion objective for cultured cells or high-resolution imaging. DAPI, GFP/Alexa Fluor 488, Alexa Fluor 546, and Alexa Fluor 633 were excited at 405, 488, 555, and 633 nm lasers, respectively. z-stacks were acquired at intervals determined by the microscope’s optimal acquisition settings, and maximum-intensity projections were generated for figure preparation. Images were processed using ZEN (version 3.7, Zeiss) and ImageJ (version 2.17.0, NIH) for contrast adjustment, montage assembly, and pseudocolor rendering. Quantification was performed on at least three slices per animal, with manual cell counting using the ImageJ Cell Counter plugin.
Chromatin immunoprecipitation (ChIP)
108 cells were fixed with 0.95% formaldehyde at room temperature for 9 min and quenched with glycine at a final concentration of 0.125 M at room temperature for 10 min. The cells were then rinsed three times and then centrifuged in ice-cold PBS with 0.05% Triton X-100 and protease inhibitor at 350 x g at 4 °C for 5 min. Cell pellets were resuspended in 3 mL FA lysis buffer (50 mM HEPES-KOH [pH 7.5] /140 mM NaCl /1 mM EDTA [pH 8.0] /1% (v/v) Triton X-100/0.1% (w/v) Sodium Deoxycholate /0.1% (w/v) SDS /0.5% Sarcosyl /Protease inhibitor). Lysates were sonicated for 12 min (40% amplitude, 20 sec-ON/30 sec-OFF) for chromatin shearing. Then, 10 μg of rabbit anti-HA (Abcam: ab9110, https://www.abcam.com/ha-tag-antibody-chip-grade-ab9110.html) and rabbit anti-RNA Polymerase II, and 1 μg rabbit anti-H3K27ac (Abcam: ab4729, https://www.abcam.com/en-us/products/primary-antibodies/histone-h3-acetyl-k27-antibody-chip-grade-ab4729) antibodies were conjugated to protein G beads (PureProteome™ Protein A/G Mix Magnetic Beads, Millipore), which had been washed with antibody blocking reagent (0.5% BSA /0.05% Triton X-100/PBS). The mixtures were incubated with rotation at 4 °C for 2 h. 2 mL of chromatin solutions after centrifugation were diluted with 8 mL dilution buffer (50 mM HEPES [pH 7.5]/1 mM EDTA/150 mM NaCl/1% (v/v) Triton X-100/Protease inhibitor). 1% diluted chromatin was taken as input. The diluted chromatins were incubated with antibody-protein G matrix at 4 °C overnight. The beads were washed 6 times with high salt buffer (20 mM Tris [pH 8.0] /2 mM EDTA/0.1% SDS/1% Triton X-100/500 mM NaCl) and 1 time with TE buffer. The beads were transferred to a new tube after every wash. To elute DNA, beads were incubated with elution buffer (90 μL TE buffer [pH 8.0]/10 μL 10% SDS) at 65 °C for 15 min. Then, solutions were incubated while shaking at 650 rpm at 65 °C overnight after adding 4 μL 5 M NaCl to reverse the covalent histone-DNA bonds. RNase A and Protease K (Sigma) were treated for 2 h at 45 °C and 1 h at 55 °C, respectively. After adding 4 μL 1 M Tris pH 6.5, DNA purification was proceeded following the protocol of the purification kit (Qiagen PCR Purification kit, Cat. no. 28106). The DNA concentrations were measured by using a Qubit fluorometer.
ChIP-sequencing and bioinformatics analysis
High-throughput sequencing was performed on the Illumina NextSeq 550 platform following the manufacturer’s protocols by the Sequencing Core Facility of National Yang Ming Chiao Tung University Genome Center. Library size construction was targeted for 250-350 bp. The experiments achieved an average depth of 40 million reads, with a minimum of 32 million. Raw reads were exported in FASTQ formats and subsequently subjected to bioinformatics analysis of ChIP-seq data. FASTQ files were converted into BAM files, and alignment to mouse reference genome mm9 was carried out by Bowtie 2. Peak calling was executed with MACS version 1.4.2. (P ≤ 1e−5). Enriched peaks were annotated to identify their nearest genes using Hypergeometric Optimization of Motif EnRichment (HOMER) software, with peaks assigned to a gene if they spanned from -500 to +500 bp from its transcription start site. The quality score of ChIP-seq is defined as -10log10 (P). For gene enrichment analysis, the Database for Annotation, Visualization and Integrated Discovery (DAVID) was used to conduct Gene Ontology (GO) analysis.
Single‑cell RNA sequencing (scRNA-seq) and analysis
Mouse scRNA‑seq data from E13.5 and E15.5 mouse embryos were acquired from a previous study52 and human scRNA-seq data from embryonic week 8 to week 28 were from a previous study57. The dataset was analyzed using Seurat v4.0 R toolkit88 with RStudio software. The number of principal components was calculated using the ElbowPlot function. In mouse data, three developmental states, i.e., progenitors, transient stage, and post-mitotic neurons, were designated based on marker genes for dorsal pallium cells (Emx1), progenitors (Pax6), transient stage (Eomes), and post-mitotic cells (Neurod6), as well as cortical genes (Tbr1 and Satb2). In human data, cell identities were defined according to the gene expression profiles based on previous reports57,89.
Cell body size quantification
To quantify cortical neuronal soma size, segmentation was performed using Cellpose (version 2.0)90, a deep learning-based segmentation tool. The cyto model was applied to delineate soma boundaries, with the estimated cell diameter set at 23.1 μm based on visual inspection. Both the graphical user interface (GUI) and command-line interface (CLI) modes were used depending on batch size and computational requirements. Segmentation masks were manually corrected when necessary to improve accuracy. Following segmentation, regions of interest (ROIs) were exported from Cellpose and imported into ImageJ (version 2.17.0, NIH) for quantitative analysis. Morphological parameters, including soma area, diameter, and perimeter, were measured using the ImageJ Measure function. Only neurons with complete boundaries fully contained within the field of view were included in the analysis.
Statistical analysis
Statistical analyses were conducted using Excel (Office 365, version 3511, Microsoft) and Prism (version 10.6.1, Graphpad). Student’s t-test and one-way ANOVA were used to compare differences between groups. The significant difference levels were denoted as follows: * for P < 0.05, ** for P < 0.01, *** for P < 0.001, and **** for P < 0.0001. Error bars in the bar graphs represent means ± S.E.M.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data and materials are contained within the paper and the files or available on reasonable request. The mouse scRNA-seq data52 is acquired from https://doi.org/10.57770/IWASQL. The spatial gene transcriptome data52 from mouse E15.5 is acquired from https://doi.org/10.57770/P3TK72. The human scRNA-seq data57 is acquired from https://tanglab.shinyapps.io/Human_Fetal_Brain_Cell/. Further information and requests for resources and reagents should be directed to and will be fulfilled by Jin-Wu Tsai (tsaijw@nycu.edu.tw). Source data are provided with this paper.
References
Fernández, V., Llinares-Benadero, C. & Borrell, V. Cerebral cortex expansion and folding: what have we learned?. EMBO J. 35, 1021–1044 (2016).
Subramanian, L., Calcagnotto, M. E. & Paredes, M. F. Cortical malformations: lessons in human brain development. Front. Cell. Neurosci. 13, 576 (2020).
Arai, Y. & Taverna, E. Neural progenitor cell polarity and cortical development. Front. Cell. Neurosci. 11, 384 (2017).
Malatesta, P. et al. Neuronal or glial progeny: regional differences in radial glia fate. Neuron 37, 751–764 (2003).
Fietz, S. A. et al. OSVZ progenitors of human and ferret neocortex are epithelial-like and expand by integrin signaling. Nat. Neurosci. 13, 690–699 (2010).
Hansen, D. V., Lui, J. H., Parker, P. R. L. & Kriegstein, A. R. Neurogenic radial glia in the outer subventricular zone of human neocortex. Nature 464, 554–561 (2010).
Noctor, S. C., Martínez-Cerdeño, V., Ivic, L. & Kriegstein, A. R. Cortical neurons arise in symmetric and asymmetric division zones and migrate through specific phases. Nat. Neurosci. 7, 136–144 (2004).
Hevner, R. F. et al. Beyond laminar fate: toward a molecular classification of cortical projection/pyramidal neurons. Dev. Neurosci. 25, 139–151 (2003).
Kwan, K. Y., Šestan, N. & Anton, E. Transcriptional co-regulation of neuronal migration and laminar identity in the neocortex. Development 139, 1535–1546 (2012).
Valcanis, H. & Tan, S. S. Layer specification of transplanted interneurons in developing mouse neocortex. J. Neurosci. 23, 5113–5122 (2003).
Greig, L. C., Woodworth, M. B., Galazo, M. J., Padmanabhan, H. & Macklis, J. D. Molecular logic of neocortical projection neuron specification, development and diversity. Nat. Rev. Neurosci. 14, 755–769 (2013).
Blumcke, I. et al. Histopathological findings in brain tissue obtained during epilepsy surgery. N. Engl. J. Med. 377, 1648–1656 (2017).
Najm, I. et al. The ILAE consensus classification of focal cortical dysplasia: An update proposed by an ad hoc task force of the ILAE diagnostic methods commission. Epilepsia 63, 1899–1919 (2022).
Palmini, A. et al. Terminology and classification of the cortical dysplasias. Neurology 62, S2–S8 (2004).
Chiarello, D. et al. Negative MRI and a seizure onset zone close to eloquent areas in FCD type II: Application of MRg-LiTT after a SEEG re-evaluation in pediatric patients with a previous failed surgery. Epilepsy Behav. 153, 109694 (2024).
Urbach, H., Kellner, E., Kremers, N., Blumcke, I. & Demerath, T. MRI of focal cortical dysplasia. Neuroradiology 64, 443–452 (2022).
Balestrini, S., Barba, C., Thom, M. & Guerrini, R. Focal cortical dysplasia: a practical guide for neurologists. Pr. Neurol. 23, 293–302 (2023).
Rossini, L. et al. FCD Type II and mTOR pathway: Evidence for different mechanisms involved in the pathogenesis of dysmorphic neurons. Epilepsy Res 129, 146–156 (2017).
Nguyen, L. H. & Bordey, A. Convergent and Divergent Mechanisms of Epileptogenesis in mTORopathies. Front. Neuroanat. 15, 664695 (2021).
Man A., et al. The genetics of tuberous sclerosis complex and related mTORopathies: current understanding and future directions. Genes 15, (2024).
Liu, G. Y. & Sabatini, D. M. mTOR at the nexus of nutrition, growth, ageing and disease. Nat. Rev. Mol. Cell Biol. 21, 183–203 (2020).
Moloney, P. B., Cavalleri, G. L. & Delanty, N. Epilepsy in the mTORopathies: opportunities for precision medicine. Brain Commun. 3, fcab222 (2021).
Muhlebner, A., Bongaarts, A., Sarnat, H. B., Scholl, T. & Aronica, E. New insights into a spectrum of developmental malformations related to mTOR dysregulations: challenges and perspectives. J. Anat. 235, 521–542 (2019).
Hannenhalli, S. & Kaestner, K. H. The evolution of Fox genes and their role in development and disease. Nat. Rev. Genet. 10, 233–240 (2009).
Carlsson, P. & Mahlapuu, M. Forkhead transcription factors: key players in development and metabolism. Dev. Biol. 250, 1–23 (2002).
Wong, L.-C. et al. FOXG1-related syndrome: from clinical to molecular genetics and pathogenic mechanisms. Int. J. Mol. Sci. 20, 4176 (2019).
Hanashima, C., Li, S. C., Shen, L., Lai, E. & Fishell, G. Foxg1 suppresses early cortical cell fate. Science 303, 56–59 (2004).
Florian, C., Bahi-Buisson, N. & Bienvenu, T. FOXG1-related disorders: from clinical description to molecular genetics. Mol. Syndromol. 2, 153–163 (2012).
Vezzali, R. et al. The FOXG1/FOXO/SMAD network balances proliferation and differentiation of cortical progenitors and activates Kcnh3 expression in mature neurons. Oncotarget 7, 37436 (2016).
Hou, P.-S., HAilin, D. O., Vogel, T. & Hanashima, C. Transcription and beyond: delineating FOXG1 function in cortical development and disorders. Front. Cell. Neurosci. 14, 35 (2020).
Zhang, Y. et al. Purification and characterization of progenitor and mature human astrocytes reveals transcriptional and functional differences with mouse. Neuron 89, 37–53 (2016).
Kim, H. et al. RNA demethylation by FTO stabilizes the FOXJ1 mRNA for proper motile ciliogenesis. Dev. Cell 56, 1118–1130. e1116 (2021).
He, M. et al. Strategies and Tools for Combinatorial Targeting of GABAergic Neurons in Mouse Cerebral Cortex. Neuron 91, 1228–1243 (2016).
Qiu, F. et al. miR-29a/b enhances cell migration and invasion in nasopharyngeal carcinoma progression by regulating SPARC and COL3A1 gene expression. PloS one 10, e0120969 (2015).
Martín-de-Lara, F., Sánchez-Aparicio, P., Arias de la Fuente, C. & Rey-Campos, J. Biological effects of FoxJ2 over-expression. Transgenic Res. 17, 1131–1141 (2008).
Landgren, H. & Carlsson, P. FoxJ3, a novel mammalian forkhead gene expressed in neuroectoderm, neural crest, and myotome. Dev. Dyn. 231, 396–401 (2004).
Wu, H. et al. Retinoic acid-induced upregulation of miR-219 promotes the differentiation of embryonic stem cells into neural cells. Cell Death Dis. 8, e2953 (2017).
Alexander, M. S. et al. Foxj3 transcriptionally activates Mef2c and regulates adult skeletal muscle fiber type identity. Dev. Biol. 337, 396–404 (2010).
Grant, G. D. et al. Live-cell monitoring of periodic gene expression in synchronous human cells identifies Forkhead genes involved in cell cycle control. Mol. Biol. Cell 23, 3079–3093 (2012).
Seo, Y., Lee, J., Kim, J., Shim, S. & Jang, S. W. FOXJ3 regulates cell proliferation and motility through modulating snail expression in breast cancer cells. Anticancer Res 43, 2995–3001 (2023).
Zhang, J. Y., Su, X. P., Li, Y. N. & Guo, Y. H. MicroRNA-425-5p promotes the development of prostate cancer via targeting forkhead box J3. Eur. Rev. Med Pharm. Sci. 23, 547–554 (2019).
Jin, J. et al. MiR-517a-3p accelerates lung cancer cell proliferation and invasion through inhibiting FOXJ3 expression. Life Sci. 108, 48–53 (2014).
Marin-Valencia, I., Guerrini, R. & Gleeson, J. G. Pathogenetic mechanisms of focal cortical dysplasia. Epilepsia 55, 970–978 (2014).
Hevner RFJJoN, Neurology E. Layer-specific markers as probes for neuron type identity in human neocortex and malformations of cortical development. 66, 101-109 (2007).
Gleeson, J. G. & Walsh, C. A. Neuronal migration disorders: from genetic diseases to developmental mechanisms. Trends Neurosci. 23, 352–359 (2000).
Park, S. M. et al. Brain somatic mutations in MTOR disrupt. Neuronal Ciliogenesis, Lead. Focal Cortical Dyslamination. Neuron 99, 83–97.e87 (2018).
Ni, L., Xie, H. & Tan, L. Multiple roles of FOXJ3 in spermatogenesis: a lesson from Foxj3 conditional knockout mouse models. Mol. Reprod. Dev. 83, 1060–1069 (2016).
Cai, L., Hayes, N. L. & Nowakowski, R. S. Local homogeneity of cell cycle length in developing mouse cortex. J. Neurosci. 17, 2079–2087 (1997).
Calegari, F., Haubensak, W., Haffner, C. & Huttner, W. B. Selective lengthening of the cell cycle in the neurogenic subpopulation of neural progenitor cells during mouse brain development. J. Neurosci. 25, 6533–6538 (2005).
MacDermot, K. D. et al. Identification of FOXP2 truncation as a novel cause of developmental speech and language deficits. Am. J. Hum. Genet. 76, 1074–1080 (2005).
Ramanand, S. G. et al. The landscape of RNA polymerase II-associated chromatin interactions in prostate cancer. J. Clin. Investig. 130, 3987–4005 (2020).
Tsai, M. H. et al. Novel lissencephaly-associated NDEL1 variant reveals distinct roles of NDE1 and NDEL1 in nucleokinesis and human cortical malformations. Acta Neuropathol. 147, 13 (2024).
Peng, W. & Zhao, K. An integrated strategy for identification of both sharp and broad peaks from next-generation sequencing data. Genome Biol. 12, 120 (2011).
Nakato, R. & Sakata, T. Methods for ChIP-seq analysis: a practical workflow and advanced applications. Methods 187, 44–53 (2021).
Park, P. J. ChIP-seq: advantages and challenges of a maturing technology. Nat. Rev. Genet. 10, 669–680 (2009).
Pepke, S., Wold, B. & Mortazavi, A. Computation for ChIP-seq and RNA-seq studies. Nat. Methods 6, S22–S32 (2009).
Fan, X. et al. Single-cell transcriptome analysis reveals cell lineage specification in temporal-spatial patterns in human cortical development. Sci. Adv. 6, eaaz2978 (2020).
Jiang, X. & Nardelli, J. Cellular and molecular introduction to brain development. Neurobiol. Dis. 92, 3–17 (2016).
Brunet, A., Datta, S. R. & Greenberg, M. E. Transcription-dependent and -independent control of neuronal survival by the PI3K–Akt signaling pathway. Curr. Opin. Neurobiol. 11, 297–305 (2001).
Hur, E.-M. & Zhou, F.-Q. GSK3 signalling in neural development. Nat. Rev. Neurosci. 11, 539–551 (2010).
Ersahin, T., Tuncbag, N. & Cetin-Atalay, R. The PI3K/AKT/mTOR interactive pathway. Mol. Biosyst. 11, 1946–1954 (2015).
Inoki, K., Li, Y., Zhu, T., Wu, J. & Guan, K. L. TSC2 is phosphorylated and inhibited by Akt and suppresses mTOR signalling. Nat. Cell Biol. 4, 648–657 (2002).
Lyu, J. et al. The protein phosphatase activity of PTEN is essential for regulating neural stem cell differentiation. Mol. Brain 8, 26 (2015).
Shelkowitz, E. et al. Variants in PTEN are associated with a diverse spectrum of cortical dysplasia. Pediatr. Neurol. 147, 154–162 (2023).
Frazier, T. W. et al. Molecular and phenotypic abnormalities in individuals with germline heterozygous PTEN mutations and autism. Mol. Psychiatry 20, 1132–1138 (2015).
Dhaliwal, N., Choi, W. W. Y., Muffat, J. & Li, Y. Modeling PTEN overexpression-induced microcephaly in human brain organoids. Mol. Brain 14, 131 (2021).
Backman, S. A. et al. Deletion of Pten in mouse brain causes seizures, ataxia and defects in soma size resembling Lhermitte-Duclos disease. Nat. Genet. 29, 396–403 (2001).
Alcamo, E. A. et al. Satb2 regulates callosal projection neuron identity in the developing cerebral cortex. Neuron 57, 364–377 (2008).
Arlotta, P. et al. Neuronal subtype-specific genes that control corticospinal motor neuron development in vivo. Neuron 45, 207–221 (2005).
Baek, S. T. et al. Off-target effect of doublecortin family shRNA on neuronal migration associated with endogenous microRNA dysregulation. Neuron 82, 1255–1262 (2014).
Toma, K. & Hanashima, C. Switching modes in corticogenesis: mechanisms of neuronal subtype transitions and integration in the cerebral cortex. Front. Neurosci. 9, 274 (2015).
Kohwi, M. & Doe, C. Q. Temporal fate specification and neural progenitor competence during development. Nat. Rev. Neurosci. 14, 823–838 (2013).
D’Gama, A. M. et al. Mammalian target of rapamycin pathway mutations cause hemimegalencephaly and focal cortical dysplasia. Ann. Neurol. 77, 720–725 (2015).
Kriegstein, A. & Alvarez-Buylla, A. The glial nature of embryonic and adult neural stem cells. Annu Rev. Neurosci. 32, 149–184 (2009).
D’Gama, A. M. et al. Somatic mutations activating the mTOR pathway in dorsal telencephalic progenitors cause a continuum of cortical dysplasias. Cell Rep. 21, 3754–3766 (2017).
Ranieri, C. et al. In vitro efficacy of ARQ 092, an allosteric AKT inhibitor, on primary fibroblast cells derived from patients with PIK3CA-related overgrowth spectrum (PROS). Neurogenetics 19, 77–91 (2018).
Saito, T. & Nakatsuji, N. Efficient gene transfer into the embryonic mouse brain using in vivo electroporation. Dev. Biol. 240, 237–246 (2001).
Nakajima HTaK. Efficient in utero gene transfer system to the developing mouse brain using electroporation. Neuroscience 103, (2001).
Taniguchi, Y., Young-Pearse, T., Sawa, A. & Kamiya, A. In utero electroporation as a tool for genetic manipulation in vivo to study psychiatric disorders: from genes to circuits and behaviors. Neuroscientist 18, 169–179 (2012).
Tsai, M.-H. et al. Impairment in dynein-mediated nuclear translocation by BICD2 C-terminal truncation leads to neuronal migration defect and human brain malformation. Acta Neuropathol. Commun. 8, 106 (2020).
Tsai M. H., et al. A lissencephaly-associated BAIAP2 variant causes defects in neuronal migration during brain development. Development 151, (2024).
Huang, C.-W. et al. Muscleblind-like 2 knockout shifts adducin 1 isoform expression and alters dendritic spine dynamics of cortical neurons during brain development. Neuropathol. Appl. Neurobiol. 49, e12890 (2023).
Cheng H.-Y., Nian F.-S., Ou Y.-W., Tsai J.-W. Assessment of Dynein-Mediated Nuclear Migration in the Developing Cortex by Live-Tissue Microscopy. In: Dynein: Methods and Protocols (ed Markus S. M.). Springer US (2023).
Nian, F.-S. et al. Rab18 collaborates with Rab7 to modulate lysosomal and autophagy activities in the nervous system: an overlapping mechanism for warburg micro syndrome and Charcot-Marie-Tooth neuropathy type 2B. Mol. Neurobiol. 56, 6095–6105 (2019).
Meerbrey, K. L. et al. The pINDUCER lentiviral toolkit for inducible RNA interference in vitro and in vivo. Proc. Natl. Acad. Sci. USA 108, 3665–3670 (2011).
Labun, K. et al. CHOPCHOP v3: expanding the CRISPR web toolbox beyond genome editing. Nucleic Acids Res. 47, W171–W174 (2019).
Sakuma, T., Nishikawa, A., Kume, S., Chayama, K. & Yamamoto, T. Multiplex genome engineering in human cells using all-in-one CRISPR/Cas9 vector system. Sci. Rep. 4, 5400 (2014).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587.e3529 (2021).
Pollen, A. A. et al. Molecular identity of human outer radial glia during cortical development. Cell 163, 55–67 (2015).
Stringer, C., Wang, T., Michaelos, M. & Pachitariu, M. Cellpose: a generalist algorithm for cellular segmentation. Nat. methods 18, 100–106 (2021).
Acknowledgements
The authors would like to thank all the affected individuals and their families participating in this study. This study was supported by grants from the National Science and Technology Council (NSTC, 108-2638-B-010-001-MY2, 108-2321-B-010-011-MY2, 110-2628-B-A49A-506, 111-2628-B-A49-016, 112-2628-B-A49-005, and 112-2314-B-A49-075), National Health Research Institutes (NHRI-EX109-10904NI), and the Brain Research Center, National Yang Ming Chiao Tung University (NYCU) through the Featured Areas Research Center Program within the framework of the Higher Education Sprout Project by the Ministry of Education (MOE) to JWT; NSTC (111-2314-B-A49-053-MY3), National Health Research Institutes (NHRI-EX113-11007NC) to PSH; and NSTC (107-2314-B-075-020, 109-2314-B-075-055, 110-2628-B-075-014, 112-2314-B-075-078, 113-2314-B-075-003, 113-2314-B-075-072-MY3), Taipei Veterans General Hospital (V108C-075) to YTL. HMC and SMS were supported by the Epilepsy Society; HMC was supported by the Amelia Roberts Fund. HYC is a Postdoctoral Researcher of Fund for Scientific Research – FNRS and supported by the Grant No. 40032806. The authors acknowledge the technical services provided by the National Genomics Center for Clinical and Biotechnological Applications of the Cancer and Immunology Research Center (NYCU), the National Core Facility for Biopharmaceuticals (NCFB), NSTC (113-2740-B-A49-002, 112-2740-B-A49-001, 111-2740-B-A49-001)
Author information
Authors and Affiliations
Contributions
Conceptualization: H.Y.C., Chen Liu, Y.T.L., P.S.H., and J.W.T.; methodology: H.Y.C., Chen Liu, C.W.N., H.C., H.J.Z., F.S.N., Chien Lu, C.S.H., W.C.P., C.C.C., J.S.H., P.L.C., C.Y.T.; clinical and genetic assessments: C.C., H.M.C., S.M.S., H.H.C., S.A., F.P.C., H.H., and Y.T.L.; data analysis: H.Y.C., Chen Liu, C.W.N., H.C.H., H.J.Z., F.S.N., Chien Lu, C.S.H., W.C.P., C.C.C., J.S.H., P.L.C., C.Y.T., W.Y.C., Y.T.L., and P.S.H.; paper writing: H.Y.C., Chen Liu, C.W.N., H.C.H., H.J.Z., S.M.S., Y.T.L., P.S.H., and J.W.T.; paper review and editing: all authors; supervision: P.L.C., S.J.C., S.A., H.H., W.Y.C., Y.T.L., P.S.H., and J.W.T. C.W.N., H.C.H., H.J.Z. also contributed equally to this work.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Cheng, HY., Liu, C., Nien, CW. et al. Epilepsy-associated FOXJ3 variants link a transcriptional program of the PTEN-mTOR pathway to neuronal specification and cortical lamination. Nat Commun 17, 1815 (2026). https://doi.org/10.1038/s41467-026-69241-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-69241-2