Abstract
Insect venoms can cause severe allergic reactions, including anaphylaxis, in sensitized individuals. In this study, we aim at preventing anaphylaxis mediated by the most abundant and dominant honeybee venom allergen phospholipase A2 (Api m 1) by blocking its interaction with allergic patient IgE. Therefore, we characterize selected Api m 1-specific nanobodies and identify two high-affinity binders with non-overlapping epitopes. Crystal structures of Api m 1/nanobody complexes reveal diametrically opposed epitopes, one of which involves the active site of Api m 1. Based on this background, we develop mono- and bispecific nanobody-human IgG1 Fc, which exhibits pronounced blocking of IgE binding and effector cell activation in blood samples from honeybee venom allergic patients and reduces systemic reactions in a mouse model of allergen-induced anaphylaxis. This work provides a rationale for using nanobody-based inhibitors to prevent Api m 1-mediated anaphylaxis in honeybee venom allergy.
Similar content being viewed by others
Introduction
Hymenoptera venom allergy (HVA) is a severe, potentially life-threatening condition and one of the most common causes of anaphylaxis1,2. The prevalence of systemic reactions to Hymenoptera stings is between 0.3 and 7.5% in the European population3. One of the main perpetrators of HVA is the honeybee (Apis mellifera). The response to honeybee venom (HBV) is a type I hypersensitivity reaction caused by allergen-induced cross-linking of IgE bound to the high-affinity receptor FcεRI on mast cells and basophils and subsequent release of mediators, which trigger immediate-type allergic reactions4,5.
The IgE response in HBV-allergic patients is directed against different venom allergens. The most abundant component is the cytolytic peptide melittin, which constitutes 50% of the venom dry weight6. The 16 kDa allergen phospholipase A2 (Api m 1) constitutes 12% of the venom dry weight and has a reported sensitization rate of up to 97% of HBV-allergic patients6,7,8, thus being considered the dominant allergen in HBV. Most of the other allergens are of low abundance, comprising each less than 1–2% of venom dry weight, and have reported sensitization rates of >50%6,9. The individual clinical relevance of the lower-abundance allergens is not yet proven.
Api m 1 is in synergy with melittin, involved in the development of pain and inflammation induced by the sting. Api m 1 is a Ca2+-dependent phospholipase A2, which disrupts cellular membranes, resulting in release of pro-inflammatory mediators10,11,12. It comprises a conserved active site at the core of three α-helices, an N-terminal calcium-binding loop, and an asparagine-linked glycosylation site (N13)13. The carbohydrate moiety itself represents a prominent IgE epitope, which is considered to have rather limited clinical relevance14,15,16,17.
Currently, the only available treatment for HBV allergy that can limit further systemic reactions is venom immunotherapy (VIT)18. VIT can be performed with different treatment regimens, all of which aim to restore immune tolerance through subcutaneous injection of HBV extract over the course of 5 years, or even lifelong under certain circumstances including patients with a severe initial sting reaction or mastocytosis19. VIT has been reported to be effective in 77–94% of HBV-allergic patients19,20. However, VIT is also associated with a risk of systemic adverse effects19,21.
The dominant mechanism by which allergen immunotherapy (AIT) induces tolerance is through the production of allergen-specific IgG that block the interaction between the allergen and allergen-specific IgE22,23,24. Recent studies on passive immunotherapy in cat and birch pollen allergy revealed that a single dose of two or three monoclonal IgG4 antibodies against the major cat allergen Fel d 1 and the major birch pollen allergen Bet v 1, respectively, reduced clinical symptoms in allergic patients25,26,27,28. Passive immunization of HBV-allergic patients with purified IgG from tolerant beekeepers has been demonstrated to protect against systemic reactions when undergoing rush immunotherapy with HBV and increase the tolerated venom dose29,30. Thus, these studies underline the potential of using allergen-blocking IgG for passive immunotherapy as a treatment option for HBV allergy. In HBV allergy, stings occur unpredictably within a well-defined season, and passive immunotherapy with IgG having a half-life of 20–30 days31 could provide seasonal protection against severe symptoms, including anaphylaxis in case of a sting event.
In recent years, single-domain antibody therapeutics have gained ground. Single-domain antibodies, often called nanobodies, are isolated from heavy-chain-only antibodies found in camelid species32. Compared to conventional IgG antibodies, nanobodies are tenfold smaller and thus can easily be linked to other molecules, generating a variety of nanobody-based formats with numerous biotechnological and therapeutic applications33,34. Their slightly longer complementarity-determining region 3 (CDR3) and prolate shape render nanobodies particularly suited for targeting concave or cryptic structures such as active sites of enzymes35.
In this study, we generate Api m 1-specific nanobodies for interfering with the binding of HBV-allergic patient IgE to Api m 1 and asses a potential reduction of Api m 1-mediated anaphylaxis. We characterize the interaction between the nanobodies and the allergen structurally and biochemically and document the blocking potential of individual, combined, and bispecific nanobody-based hIgG1 Fc fusion proteins (nb-hIgG1). The structural and functional data provide insights into the allergenicity of venom allergens as well as a proof-of-concept for using nanobody-based formats for blocking allergen-induced anaphylaxis in the context of HBV allergy.
Results
Establishment of high-affinity Api m 1-specific nanobodies
Api m 1-specific nanobodies were isolated by the establishment and selection of an immune library by phage display as recently described36. Selected nanobodies were initially characterized with respect to expression yield, affinity, and epitope overlap. On this background, two nanobodies, AM1-1 and AM1-4, were chosen for further studies. The two nanobodies are highly diverse in their CDR residues and length, in particular in the CDR3 (Supplementary Fig. 1). Both nanobodies were recombinantly expressed and purified at high yields from the culture supernatant. Sodium dodecylsulfate polyacrylamide gel electrophoresis (SDS-PAGE) confirmed the expected size of 15 kDa (Fig. 1a). Biolayer interferometry (BLI) demonstrated binding to immobilized Api m 1 with dissociation constant (KD) values in the nanomolar to sub-nanomolar range (Fig. 1b). Sequential BLI binding analysis further demonstrated independent, non-competitive binding to Api m 1, irrespectively of the order of nanobody binding, indicating two distinct epitopes of the nanobodies (Fig. 1c).
a Purified nanobodies were assessed by SDS-PAGE. b, c Binding kinetics of the nanobodies (Nb) were obtained with BLI using biosensors loaded with nApi m 1. b Kinetic constants were calculated using a 1:1 binding model (black lines). c Simultaneously binding was assessed by binding of AM1-1 (green) followed by binding of AM1-4 (blue) (left) or vice versa (right).
Structural analysis of Api m 1/nanobody interactions
In order to understand in detail the nanobody-allergen interactions, a ternary complex was constituted by mixing native Api m 1 (nApi m 1), AM1-1 and AM1-4 in equimolar amounts, and crystallized. X-ray diffraction data were obtained with a maximum resolution of 1.76 Å (Table 1). The structure revealed a 1:1:1 complex with the two nanobodies binding to diametrically opposing epitopes of the allergen (Fig. 2a, b). The architecture of the ternary complex corroborates the BLI data.
a, b Cartoon representation with transparent surface of the nanobodies AM1-1 (green) and AM1-4 (blue) binding two distinct epitopes on opposing sides of Api m 1 (orange). c, d Api m 1 with bound nanobody (left). Api m 1 residues that interact with the nanobodies by direct hydrogen bonds are colored green (AM1-1) and blue (AM1-4) (right). e Api m 1 sequence with secondary structures (orange; arrow: β-strand, cylinder: α-helix). Residues interacting with AM1-1 and AM1-4 by direct hydrogen bonds are highlighted.
The structure of Api m 1 in complex with the two nanobodies is in close agreement with the known structure of unbound Api m 113, with a root mean square deviation <1 Å over 99 Cα atoms. Notably, the residues 1–5 and 11–23 in Api m 1, including the calcium-binding loop and the N-linked glycosylation site, could not be modeled in the Api m 1/ nanobody complex, suggesting that they are disordered.
Clear electron density is present for the residues at the Api m 1/nanobody interfaces (Supplementary Fig. 2). According to PISA analysis, the buried surface areas at the interface between Api m 1 and AM1-1 or AM1-4 are 1627.9 Å2 and 1473.3 Å2, respectively, which are common interface areas for antigen/nanobody complexes37. Both AM1-1 and AM1-4 recognize conformational epitopes on Api m 1 (Fig. 2c–e). As expected, the CDRs are responsible for the majority of the interactions (Supplementary Figs. 3 and 4). The relatively long CDR3 of AM1-4 reaches into a pocket between the α-helical segments and the C-terminal β-strands of Api m 1 (Fig. 2a, b, d, and Supplementary Fig. 4c). On the opposite hemisphere, the CDR3 of AM1-1 interacts directly with the active site of Api m 1 (Figs. 2a–c and 3a, and Supplementary Fig. 4a). In addition to the CDR3, residues of the framework regions are also involved in direct and water-mediated contacts with Api m 1 (Supplementary Figs. 3 and 4b).
a Interactions of the AM1-1 complementarity determining region 3 (CDR3) (green) with residues of the catalytic network (yellow) and residues involved in Ca2+ binding (purple) of Api m 1 (orange). Hydrogen bonds are shown as dotted lines. b Phospholipase activity assay of Api m 1 in the presence of the nanobodies. Activity was measured as fluorescence using the phospolipase A2 substrate Red/Green BODIPY®PC-A2. Data are presented as mean of duplicates. Source data are provided as a Source Data file. c Cartoon representation of the nanobody AM1-4 (blue) in complex with a dimer of Api m 1 (orange). d Allergen-induced activation of effector cells was assessed with RBL-SX38 cells expressing the human FcεRI receptor. The effector cells were sensitized with nb-hIgE. Degranulation was induced by nApi m 1 and measured as released Beta-Hexosaminidase activity. Data are presented as mean of duplicates. Source data are provided as a Source Data file. e Cartoon representation of a single molecule of Api m 1 (orange) from the dimer complex. The calcium-binding loop (purple) and the glycosylation site (N13) with the first N-acetylglucosamine (GlcNAc) (cyan) are highlighted. f Superposition of a Api m 1 molecule (orange) from the binary complex with the Api m 1 molecule (gray) and nanobody AM1-1 (green) from the ternary complex. Bound AM1-1 sterically clashes with parts of the N-terminal loop and the glycan linked to N13.
The nanobody AM1-1 binds to the active site of Api m 1 and blocks its enzymatic activity
The residues S102 and R104 of the AM1-1 CDR3 forms hydrogen bonds with the residues H34, D35, and Y87 of Api m 1 (Fig. 3a, and Supplementary Fig. 4a). Together with D64, the residues H34 and Y87 form a catalytic network, while D35 is essential for Ca2+ binding13,38. Hence, our structure suggested that the binding of AM1-1 interferes with the enzymatic activity of Api m 1. Therefore, the enzymatic activity of Api m 1 was assessed by monitoring the cleavage of a fluorogenic substrate in the presence of the nanobodies. AM1-1 reduced the activity of Api m 1 to 50% whereas AM1-4 had no reducing effect (Fig. 3b). A reduction of the inhibitory effect upon combination might rely on minor conformational rearrangements affecting the binding of AM1-1. The observed inhibition supported the finding that AM1-1 interacts directly with the active site of Api m 1 and exhibits the potential to reduce the enzymatic activity of Api m 1, which contributes to the pain and inflammation of a honeybee sting.
The nanobody AM1-4 enables dimer formation and cellular activation
In parallel, an Api m 1/AM1-4 binary complex was crystallized (Table 1). For these crystals, diffraction data extending to a maximum resolution of 4.1 Å was obtained. The structure revealed a 1:1 complex in which an Api m 1 dimer is bound by two molecules of AM1-4 oriented in opposing directions in relation to the Api m 1 dimer axis (Fig. 3c). The position of AM1-4 in this complex is identical to the position in the complex of Api m 1 with both nanobodies. Also in this complex, the structure of Api m 1 is very similar to that of unbound Api m 113, with a root mean square deviation <1 Å over 116 Cα atoms.
Interestingly, in this complex Api m 1 forms a tight dimer in the crystal packing like the dimer observed for unbound Api m 113. The relationship between the two monomers differs by a rotation of 40° (Supplementary Fig. 5). Although the formation of dimers in solution has not been reported, Api m 1 might transiently form dimers at locally high concentrations, e.g., at cellular surfaces. Such propensity to dimer formation has been proposed as a mechanistic rationale for enhanced allergenicity of a large number of allergens39,40,41. Thus, a single IgE antibody clone recognizing one IgE epitope would be sufficient for cross-linking and cellular activation.
In order to investigate the potential formation of transient dimers, nanobody-human IgE Fc formats (nb-hIgE) of AM1-1 and AM1-4 were generated as previously described (Supplementary Fig. 6)36. The ability of the individual and combined nb-hIgEs to activate effector cells upon Api m 1 stimulation was assessed using RBL-SX38 cells. Expectedly, the combination of AM1-1 hIgE and AM1-4 hIgE lead to allergen-dependent degranulation upon stimulation with Api m 1. Remarkably and in contrast to the AM1-1 hIgE, the AM1-4 hIgE alone was able to induce Api m 1-dependent cellular activation and degranulation to the same extent as the combination of two IgE (Fig. 3d). In contrast to cytokinergic IgE, which rely on spontaneous oligomerization of IgE42,43, the cellular activation was strictly allergen-dependent, and could be explained convincingly by the formation of Api m 1 dimers at the surface. This concept is supported by the structure showing that AM1-4 binding is compatible with dimer formation.
The binding of AM1-1 induces conformational rearrangements of Api m 1
Notably, the N-terminal calcium-binding loop and the N-linked glycosylation site of Api m 1 are disordered in the ternary Api m 1/nanobody complex. In contrast, in the Api m 1/AM1-4 binary complex, all Api m 1 residues including the first N-Acetylglucosamine (GlcNAc) of the glycan at N13 could be modeled (Fig. 3e). An overlay of the two Api m 1 complexes reveals a steric clash of the AM1-1 nanobody with the N-terminal loop and the glycan of Api m 1. Since the glycan is significantly larger than the first GlcNAc sugar that can be modeled in our structure, the clash between a bound AM1-1 nanobody and the glycan is likely to be pronounced. This finding suggests a displacement of the N-terminal loop including the calcium-binding loop and the glycan upon binding of AM1-1 to Api m 1 (Fig. 3f).
Generation of nanobody-human IgG1
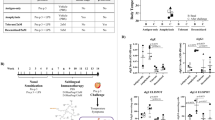
With the purpose of blocking the binding of allergen-specific IgE from HBV-allergic patients to Api m 1, monospecific nb-hIgG1 (AM1-1 hIgG1 and AM1-4 hIgG1) comprising either of the nanobodies were generated by fusion of the nanobodies to the hinge region of human IgG1 Fc (Fig. 4a). In addition, a bispecific nb-hIgG1 format (AM1B-1-4 hIgG1) comprising both of the nanobodies within a single molecule was generated by fusion of the nanobodies to human IgG1 Fc containing DEKK mutations for facilitated heterodimerization of the two different antibody chains44. The IgG1 format was chosen since IgG1 is an early blocking antibody arising in AIT24. The three nb-hIgG1 formats were expressed in mammalian cells, purified and analyzed by SDS-PAGE, which verified the expected molecular mass of a dimer glycosylated within the Fc domains (Fig. 4b). For the bispecific nb-hIgG1, two bands under reducing conditions represented the two monomers of slightly different molecular weight due to the presence of specific epitope tags, suggesting efficient heterodimerization
a Schematic representation of nanobody-human IgG1 (nb-hIgG1) formats generated by linking the nanobody to the hinge region of a human IgG1. Created in BioRender. Spillner, E. (2026) https://BioRender.com/7p62zgg. b Purified nb-hIgG1 formats were assessed by SDS-PAGE under non-reducing (left) and reducing (right) conditions. c Binding kinetics of the nb-hIgG1 were obtained with BLI by loading the nb-hIgG1 to the biosensors and detecting the binding of nApi m 1. Kinetic constants were calculated using a 1:1 binding model (black lines).
ELISA assessment verified the immunoreactivity of the different nb-hIgG1 (Supplementary Fig. 7a). ImmunoCAP analyzes further documented specificity for HBV without cross-reactivity to other Hymenoptera venom from species such as bumblebee, yellow jacket, and fire ant venom (Supplementary Fig. 7b). Notably, the bumblebee phospholipase A2, Bom t 1, shares a sequence identity of 54% and a conserved active site with Api m 1. Only one residue of the AM1-1 epitope is replaced, but no cross-reactivity with bumblebee venom was observed (Supplementary Fig. 7b, c).
BLI binding studies of the three immobilized nb-hIgG1 further revealed agreement of the kinetic data with those of the individual nanobodies, confirming that the Fc fusion has no impact on binding affinity (Fig. 4c). Notably, the dissociation rate of the bispecific nb-hIgG1 was significantly lower (<10−7 s−1), most likely reflecting increased avidity due to simultaneous binding of two Api m 1 epitopes. The change in the dissociation rate yielded an overall KD in the sub-picomolar range (<10−12 M).
Nanobody-hIgG1 formats block the binding of HBV-allergic patient IgE to Api m 1
The ability of the nb-hIgG1 formats to inhibit the binding of IgE from HBV-allergic patients to Api m 1 was assessed in an inhibition ELISA. Immobilized Api m 1 was incubated with the nb-hIgG1 followed by incubation with serum from HBV-allergic patients, and the remaining binding of allergen-specific IgE was analyzed. For this analysis, HBV-allergic patients with elevated specific IgE titers to Api m 1 and anaphylaxis grade ≥1 were chosen (Serological data Supplementary Table 1). Both individual monospecific nb-hIgG1 formats inhibited the binding of patient IgE partially (Fig. 5a, b). Varying ratios of inhibition with AM1-1 and AM1-4 were observed between different patients but in general AM1-4 hIgG1 showed a more pronounced inhibition (mean 69%) than AM1-1 hIgG1 (mean 49%). The inhibition could be significantly increased by using a combination of the two monospecific nb-hIgG1 formats (mean 91%) or the bispecific AM1B-1-4 nb-hIgG1 (mean 94%) (Fig. 5b).
a Binding of serum IgE from HBV-allergic patients to immobilized nApi m 1 was assessed in the presence of the different nb-hIgG1 formats. IgE binding was detected with anti-human IgE antibody conjugated to alkaline phosphatase. IgE binding is indicated as the percentage compared to the binding of patient IgE in the presence of a nb-hIgG1 isotype control calculated from the mean of duplicates. b Data are presented as mean ± SD of sera from the five HBV-allergic patients (n = 5). Statistical analysis was performed by one-way ANOVA with Tukey test (**p < 0.01; ****p < 0.0001; ns:non-significant). Source data are provided as a Source Data file.
These data demonstrate that the individual and combined nb-hIgG1 formats can efficiently block the binding of IgE from HBV-allergic patients to immobilized Api m 1.
Nanobody-hIgG1 reduces Api m 1-induced basophil activation
The nb-hIgG1 formats were further evaluated for their ability to block Api m 1-induced activation of basophils from Api m 1-sensitized HBV-allergic patients (Supplementary Table 2).
It is well established that patient basophils respond to allergen stimulation with varying reactivity and sensitivity. Therefore, the response of basophils from nine HBV-allergic patients after stimulation with nApi m 1 was assessed as an initial step (Supplementary Fig. 8). Subsequently, the response in the presence of the nb-hIgG1 formats was analyzed. For seven out of nine patients, the nb-hIgG1 formats reduced basophil activation although to a varying degree (Fig. 6a, Supplementary Fig. 9, and Supplementary Table 3). For these seven patients, the AM1-4 hIgG1 format resulted in a more pronounced reduction than the AM1-1 hIgG1 format, which only exhibited a minor decrease in basophil reactivity. For most patients, an increased blocking was observed when using a combination of the two nb-hIgG1 formats. Notably, the bispecific format performed equally well as the combination of the two individual antibodies, and significantly reduced basophil activation compared to the isotype control (Fig. 6b). For two out of nine patients, the nb-hIgG1 formats had no inhibitory effect on basophil activation. One of these patients (patient N) showed a very low basophil response to Api m 1 at all, while the other patient (patient M) showed a very high response and highly elevated sIgE to Api m 1 (Fig. 6a, and Supplementary Fig. 8).
a Basophil activation test was performed in order to assess the potential of nb-hIgG1 to reduce the activation of basophils from honeybee venom (HBV) allergic patients upon Api m 1 stimulation. nApi m 1 was pre-incubated with 10-fold dilution series of nb-hIgG1 before incubation with EDTA blood from HBV-allergic patients. Activated basophils were identified as CD63 positive. b Combined data of the nine patient (n = 9) comparing the AM1B-1-4 hIgG1 with the isotype control at the highest molar ratio of nb-hIgG1/Api m 1. Statistical analysis was performed by two-tailed paired t test. **p < 0.01. c Basophils from four patients (patient O, P, Q and R) were stimulated with Api m 1 or HBV in the presence of the bispecific AM1B-1-4 hIgG1 or the isotype control. Statistical analysis was performed by two-tailed paired t test (*p < 0.05). Source data are provided as a Source Data file.
Overall, these data demonstrate the potential of the nanobody formats for blocking effector cell activation by Api m 1 stimulation. The data also confirms a large blocking variation, reflecting diverse IgE repertoires of the HBV-allergic patients.
Inhibition by the bispecific nb-hIgG1 format was further evaluated with HBV-stimulated basophils from four poly-sensitized patients, demonstrating the ability to significantly reduce basophil activation upon HBV stimulation (Fig. 6c, and Supplementary Fig. 10, Serological data in Supplementary Table 2). In three out of four patients, a clear reduction of above 34% was observed with a mean reduction of 35% for all patients. Thus, blocking the dominant allergen Api m 1 can reduce the activation of effector cells upon HBV-stimulation. Expectedly, a larger reduction of basophil activation was observed on Api m 1-stimulated basophils than on HBV-stimulated basophils, supporting a general contribution of other venom allergens to the overall basophil activation in HBV allergy.
Bispecific nanobody-hIgG1 reduces mast cell activation and anaphylaxis in vivo
The therapeutic potential of the bispecific nb-hIgG1 was evaluated in vivo using a mouse model of allergen-induced anaphylaxis. Sensitization with HBV extract resulted in the induction of HBV- and Api m 1-specific IgG and IgE (Fig. 7a and Supplementary Fig. 11). After sensitization, anaphylaxis was induced by injection of nApi m 1 (Fig. 7a). Anaphylaxis was evaluated as a drop in body temperature and mast cell activation was assessed by measuring serum concentrations of mast cell protease-1 (MCPT-1). Challenge with nApi m 1 resulted in an increased MCPT-1 serum concentration and a temperature drop in HBV-sensitized mice compared to non-sensitized mice, validating the model (Fig. 7b–d). Treatment with the bispecific nb-hIgG1 prior to Api m 1 challenge significantly reduced the Api m 1-induced MCPT-1 release and body temperature drop (Fig. 7b–d). The evaluation of treatment with the bispecific nb-hIgG1 prior to challenge with HBV showed, although non-significant, an average reduction of MCPT-1 release of 40%, which is comparable to the 35% reduction observed in HBV BAT (Supplementary Figs. 12 and 13). However, a pronounced body temperature drop (Supplementary Fig. 12) was also observed in non-sensitized mice, suggesting a non-allergic, mast cell-independent response to the venom.
a Schematic representation of the systemic anaphylaxis mouse model. Non-sensitized mice (n = 12) and mice sensitized by three weekly intraperitoneal (IP) injections of Honeybee venom (HBV) extract were challenged with Api m 1 at day 28. Sensitized mice received treatment with either PBS (n = 10) or bispecific nb-hIgG1 (n = 12) four hours prior to the challenge. b Blood was collected after the challenge and serum MCPT-1 concentration was measured by ELISA. Data are presented as mean ± SD and each symbol represents a single mouse. Statistical analysis was performed using one-way ANOVA followed by Tukey multiple comparison test (****p < 0.0001; ns: non-significant). c Body temperature was measured every 15 min for 90 min after the challenge. Data are presented as mean ± SD. d The area under the curve (AUC) of the temperature curves were calculated and statistical analysis was performed using one-way ANOVA followed by Tukey multiple comparison test (****p < 0.0001; ns:non-significant). Data are presented as mean ± SD and each symbol represents a single mouse. Source data are provided as a Source Data file.
Thus, the bispecific nb-hIgG1 format demonstrated the potential to reduce Api m 1-induced mast cell activation and Api m 1-mediated anaphylaxis in vivo in HBV-sensitized mice. The potential of the bispecific nb-hIgG1 format to reduce HBV-mediated anaphylaxis needs further investigation.
Discussion
In this study, we provide proof-of-concept for using nanobody-based formats to inhibit IgE binding to the major HBV allergen Api m 1 and the subsequent activation of effector cells in order to reduce the risk of Api m 1-mediated anaphylaxis.
For HVA with a well-defined season of 3–4 months in which sting events might occur, passive immunotherapy with IgG antibodies with a half-life of 20–30 days31 could elegantly provide transient protection by a single injection and substitute the regular application of VIT vaccines, potentially eliciting anaphylactic side effects. Moreover, imperative is the reduction of symptom severity and thereby the prevention of potentially life-threatening anaphylaxis.
In the present study, we employed allergen-specific nanobodies as building blocks for generation of blocking antibodies. Neutralizing nanobodies have been intensively applied within virology but have only lately gained attention for treatment of allergies45,46. Recently, nanobodies specific for the major birch allergen Bet v 1 were reported to reduce the binding of patient IgE, demonstrating the potential use of nanobodies to block allergens47. Importantly, nanobodies possess several properties that render them powerful alternatives to conventional antibodies34,48. In contrast to conventional antibodies, the single-domain nature of nanobodies facilitates the generation of multi-specific antibody formats targeting different molecules or different epitopes within the same molecule. This has recently been demonstrated by Lunsekimig, a pentavalent anti-TSLP/anti-IL-13 nanobody format, currently in phase 2 for the treatment of asthma49. Furthermore, nanobodies display a prolate shape and frequently extended CDR3 loops and thus exhibit a certain preference for concave epitopes such as the active site of an enzyme35,50, which is also evident from the data in this study.
Currently, only scarce information on Api m 1 IgE epitopes is available. A recent study described two linear Api m 1 IgE epitopes, one of which is located at the N-terminal loop51. However, in HBV allergy, allergens are readily injected as intact molecules, and recombinant, non-refolded Api m 1 has been shown to be of low clinical relevance in intradermal skin testing16. In the present study, X-ray crystallography revealed that both epitopes are strictly conformational without overlap with the previously suggested linear epitopes. However, the binding of the AM1-1 nanobody appears to induce conformational changes in the N-terminal loop and may, thus, interfere with the proposed linear epitope. Both nanobodies blocked the binding of patient IgE and must therefore either bind epitopes overlapping with those of patient IgE or sterically block the binding of these.
In addition to blocking IgE binding, and in line with the idea that nanobodies are suited for targeting enzymatically active allergens, we found that the nanobody AM1-1 also reduces the enzymatic activity of Api m 1. This effect is most likely caused by the direct interaction of AM1-1 with residues of the active site. The enzymatic activity of Api m 1 is known to contribute to the pain and inflammation of a honeybee sting11,12 and a targeted inhibition might lower these adverse reactions upon a sting event. Furthermore, with the emergence of Africanized bees (popularly known as “killer bees”), there is an increasing need for molecules that act as anti-venoms10.
Dimer formation and increased allergenic potency have been demonstrated for e.g., Bla g 2 and Bos d 539,40,41. Likewise, it has been proposed that a large number of allergens, including Api m 1, may form transient dimers at locally high concentrations41. Supported by the structure of the dimer, the nanobody-based IgE provided evidence for efficient activation of effector cells by a single IgE clone, suggesting dimerization as contributor to the allergenicity of Api m 1. We cannot provide evidence for dimerization upon interaction with allergen-specific IgE of HBV-allergic patients, but binding of patient IgE to epitopes in proximity to AM1-4 is likely to enable similar assemblies.
Fusion of the nanobodies to the human IgG1 Fc moiety results in higher avidity as well as larger size and thus increased half-life52. The IgG1 format was chosen as blocking allergen-specific IgG1 occurs naturally during AIT. The functionally monovalent IgG4 is the antibody subclass frequently associated with long-lasting blocking effect; however, it has been recognized decades ago that IgG1 represents the dominant IgE-blocking antibody in the early immune response of AIT24,53,54,55. In fact, for the activity of blocking antibodies the subclass seems to be irrelevant56,57. Instead, high affinity and epitope specificity are crucial for the neutralizing effects58,59. Of note, the affinities of the nb-hIgG1 generated here are comparable to those of recently reported allergen-specific monoclonal antibodies25,27.
The nb-hIgG1 formats were able to inhibit both the binding of patient IgE to Api m 1 and the subsequent basophil activation. In general, AM1-4 hIgG1 reduced the binding of IgE and cellular activation to a larger extent than AM1-1 hIgG1. This indicates that AM1-4 hIgG1 most likely blocks an area of Api m 1 that is more immunodominant than the area blocked by AM1-1. As demonstrated in comparable studies25,27 combining the two nb-hIgG1 formats increased the blocking. Furthermore, the blocking effect of the bispecific nb-hIgG1 was comparable to the combination of the two monospecific nb-hIgG1 formats.
In general, the efficacy of blocking basophil activation by VIT-induced IgG and the change in basophil sensitivity by VIT has been described inconsistently60,61,62. It is, however, well established that basophil activation after successful VIT is only reduced to some extent. Hence, the fact that we could observe a clear reduction in Api m 1- and HBV-induced basophil activation supports the concept of an efficient mode of action of the nb-hIgG1 assessed here.
Expectedly, the degree of blocking varied highly within the patient population in the BAT. These findings indicate that at least some of the HBV-allergic patients have a broad and diverse IgE repertoire in terms of epitope specificity and/or affinity, corroborating our understanding of Api m 1 as a potent allergen. Similar to previous studies, combining two blocking antibodies did not lead to full blocking in all patients27,63. Hence, blocking additional epitopes might be needed for sufficient blocking of effector cell activation in those patients.
In the present study, we focused our approach on the dominant HBV allergen Api m 1. Recent studies on passive immunotherapy in cat and birch pollen allergy have demonstrated the reduction of clinical symptoms in allergic patients by targeting only the dominant allergen Fel d 1 and Bet v 1, respectively25,26,27,28. Furthermore, it has been demonstrated in peanut-sensitized mice treated with Ara h 2-specific antibodies that targeting only the dominant allergen may be sufficient in preventing anaphylaxis even in poly-sensitized individuals63.
However, as also evident from our study, HBV-allergic patients often have IgE directed against additional venom allergens of which the clinical relevance remains unclear. Thus, blocking only one allergen might not be sufficient.
In the comparative assessment of basophil activation, the bispecific nb-hIgG1 reduced the activation of HBV-stimulated basophils with >34% in three out of four patients. Considering the incomplete blocking of Api m 1, these initial data corroborate a dominant role of Api m 1 in effector cell activation and support the selective blocking for reducing the immediate response and prevention of anaphylaxis in case of a sting event. A larger reduction was observed for Api m 1-stimulated basophils compared to HBV-stimulated basophils, suggesting the contributions of other HBV allergens but with limited impact as compared to Api m 1. Thus, further studies are needed to understand and differentiate the impact of the single allergens on the development of anaphylaxis. Importantly, the single-domain nature of the nanobodies could facilitate the generation of multi-specific formats targeting different allergens at once.
In addition to blocking basophil activation in blood samples from HBV-allergic patients, we provided evidence for the in vivo protective efficacy of the bispecific nb-hIgG1 in a mouse model of systemic anaphylaxis, demonstrating the ability of the nanobody format to strongly reduce Api m 1-induced anaphylaxis. A similar, but not significant, tendency was observed when using HBV for the challenge and MCPT-1 for monitoring mast cell degranulation. Although a mast cell independent response to the venom at doses used in our model hampered further analysis, the reduction of mast cell activation appears similar as compared to the picture observed in BAT.
Notably, the inhibitory activity observed in the animal model for Api m 1 is more pronounced as compared to the BAT inhibition, suggesting that the mouse model might reflect the complexity of the allergic response to Api m 1 and HBV differently. Hence, targeting high-abundance, dominant allergens in a mouse model is complementary to cellular analysis on the basis of allergic patients.
The results of this study show proof-of-concept for the use of blocking antibodies, but also suggest potential challenges and limitations, also mentioned above. Firstly, blocking additional epitopes on Api m 1 might be required for increased blocking of effector cell activation for those donors showing limited reduction of activation. Secondly, the antibodies with inhibitory characteristics, such as AM1-1 might have limited blocking capacity. Thirdly, blocking only one allergen might not be sufficient for preventing HBV-mediated anaphylaxis in donors with pronounced sensitization to other allergens than Api m 1. Although Api m 1 apparently is a dominant driver of the effector cell response and anaphylaxis, further studies are needed to understand and differentiate the impact of other allergens on the development of anaphylaxis. Fourthly, the correlation of reduction of effector cell activation with protection against anaphylaxis remains unclear. VIT-treated patients are protected despite having a limited reduction of effector cell activation, and other biomarkers for treatment success are currently not available.
To overcome these factors, a dual strategy could be useful. The development of further blocking antibodies against Api m 1 and other venom allergens could enable a deeper dissection of the impact of the individual allergens and their blocking on the allergic activation. Simultaneously, purified or recombinant allergens might be used to generate a tailor-made “synthetic” venom which would allow for detailed comparison of allergenic activity in absence of selected allergens and a more reliable use in in vivo experiments.
In conclusion, we have structurally defined two important nanobody epitopes on the dominant HBV allergen Api m 1 and described the molecular and functional consequences of their recognition. The nanobody-based inhibitors of both enzymatic and allergenic activity of Api m 1 provide a rationale for future development of broadly blocking antibody combinations for seasonal protection against anaphylaxis in HVA by passive immunotherapy.
Methods
Ethical statement
The use of patient material was approved by the Ethics Committee of the Medical Faculty of Philipps University, Marburg, Germany (Az. 112/17). All patients provided written informed consent to participate in the study, including informed consent for the publication of potentially identifying individual-level data, as seen in Supplementary Tables 1 and 2.
All scientific procedures on living animals were approved by the regional council (RP Giessen Az: V 54 − 19 c 20 15 h 01 GI 20/4 Nr. G 43/2024) in accordance with the German animal welfare law and the European legislation for the protection of animals used for scientific purposes (2010/63/EU).
Materials
Native Api m 1 was obtained from Latoxan.
Selection of Api m 1-specific nanobodies
Api m 1-specific nanobodies were selected previously using a nanobody library generated from peripheral blood mononuclear cells of a llama immunized with nApi m 136.
Recombinant expression of nanobodies, nb-hIgE, and nb-hIgG1 formats
Selected nanobodies were cloned into a pET-22b(+) vector (Novagen) containing a pelB signal sequence, a C-terminal polyhistidine tag, and a C-terminal V5-tag. Expression was performed in Rosetta (DE3) cells (Novagen).
Nanobody-hIgE formats were cloned as previously described36. Briefly, the nanobody sequences were cloned into a pcDNA3.1 vector (Invitrogen) between a human immunoglobulin-derived signal sequence and the human IgE Fc domains (CH2-4 domains) containing a C-terminal polyhistidine-tag. Nanobody-hIgG1 formats were generated similarly by cloning the nanobody sequences into the pcDNA3.1 vector between a human immunoglobulin-derived signal sequence and the hinge region of a human IgG1 Fc (Hinge, CH2, and CH3 domain) containing a C-terminal polyhistidine-tag as previously described64. Expression of the nb-hIgE and nb-hIgG1 formats was performed in transiently transfected mammalian FreeStyle 293-f cells (Gibco, Thermo Fisher Scientific, catalog number R79007). Bispecific nb-hIgG1 formats were generated by introducing DEKK mutations into the human IgG1 Fc as previously described44. Briefly, two vectors were constructed, containing each one chain of the nb-hIgG1 with either a L351D and a L368E mutation or a L351K and a T366K mutation. The two chains contained different tags (polyhistidine-tag or BC2-tag) for purification and assessment of heterodimerization. The two cloning vectors contained genes for either Zeocin or Geneticin (G418) resistance for selection of double-transfected cells. The bispecific nb-hIgG1 formats were expressed in adherent FreeStyle 293-f cells.
Purification of nanobodies, nb-hIgE formats, and nb-hIgG1 formats from expression media was performed by IMAC using HisTrapTM Excel columns (Cytiva). After washing (20 mM sodium phosphate, 500 mM NaCl, 30 mM imidazole, pH 7.4), bound protein was eluted with elution buffer (20 mM sodium phosphate, 500 mM NaCl, 500 mM imidazole pH 7.4). Eluted proteins were further purified by SEC on either a HiLoad 16/600 superdex 200 pg or 75 pg column (Ge Healthcare) equilibrated in PBS pH 7.4. Purified proteins were assessed by SDS-PAGE and immunoreactivity of nb-hIgG1 was assessed by ELISA with immobilized nApi m 1. Bound nb-hIgG1 was detected with AP-conjugated anti-human IgG (Sigma).
Affinity determination and epitope binning
BLI measurements were performed at 30 °C on an Octet RED96 (ForteBio) using amine-reactive second-generation (AR2G) biosensors (Sartorius). Sensors were loaded in 20 µg/ml nApi m 1 in 10 mM sodium acetate pH 6.0 for 2 min. Sensors were quenched for 300 s in 1 M ethanolamine pH 8.5, and the baseline was obtained for 120 s in kinetic buffer (PBS pH 7.4, 0.02% Tween20, 0.1% BSA). Association was performed over 300 s, and the dissociation phase was monitored for 1200 s, both in kinetic buffer. Samples without nanobody were subtracted as background. Curve fitting was performed with Octet software using a 1:1 binding model. For epitope binning 600 s association was followed by 120 s dissociation and then 600 s association of the second nanobody.
Affinity measurements of the nb-hIgG1 formats were performed using anti-hIgG Fc Capture (AHC) biosensors (Sartorius). Sensors were loaded in 2 nM nb-hIgG1 for 420 s, and a baseline was obtained for 300 s in kinetic buffer. Association was performed over 300 s with twofold dilution series of nApi m 1 from 160 to 5 nM (AM1-1 hIgG1), 40 to 1.25 nM (AM1-4 hIgG1) and 50 to 0.8 nM (AM1B-1-4 hIgG1). Dissociation was measured for 1200 s in kinetic buffer. Samples with no Api m 1 were subtracted as background. Curve fitting was performed with Octet software using a 1:1 binding model.
Structure determination
For crystallization of the ternary complex, nApi m 1, and the two nanobodies were concentrated to 14.5 mg/ml in 20 mM HEPES, 150 mM NaCl, 10 mM CaCl2, pH 7.4 with a 1:1:1 molar ratio. The complex was crystallized by vapor diffusion in sitting drops formed by mixing protein solution and reservoir solution containing 0.2 M NaCl, 0.1 M Phosphate/citrate pH 4.2 and 20% w/v polyethylene glycol (PEG) 8000 in a 1:1 ratio at 18 °C. Crystals were cryo-cooled in liquid nitrogen. Data were collected at the beamline P13 operated by EMBL at the PETRA III storage ring (DESY, Hamburg, Germany) and processed with the XDS package65. The structure was determined by molecular replacement with phenix.phaser66, using PDB ID: 1POC as search model for Api m 1 and 7NP9 for the nanobodies. The final structure was obtained by alternating manual model building in COOT67 and refinement with phenix.refine68. The complex was analyzed with the “protein interfaces, surfaces and assemblies” service PISA at the European Bioinformatic Institute69. Figures were prepared with the PyMOL Molecular Graphics System (Schrödinger LLC).
The binary complex of nApi m 1 and the nanobody AM1-4 was crystallized by the same approach. Briefly, crystals were obtained with a reservoir containing 0.1 M sodium citrate pH 5.5, and 15%w/v PEG 6000 at 4 °C. Data were collected at the beamline i24 at Diamond Light Source, UK. The structure was determined by molecular replacement using the subcomplex of Api m 1 and AM1-4 from the ternary complex as search model.
Activity assay
Inhibition of the enzymatic activity of nApi m 1 was evaluated using the EnzChek Phospholipase A2 Assay kit (Invitrogen) according to the manufacturer´s recommendation. Briefly, 2 µg/ml nApi m 1 was pre-incubated with twofold dilution series of the nanobodies and incubated for 1 h at room temperature shaking. The enzymatic reaction was started by adding 1:1 PLA2 substrate-lipid mix. Fluorescence (Excitation 467 ± 7 and Emission 515 ± 10 and 575 ± 10) was measured after 10 min incubation on a ClarioStar Plus plate reader (BMG LABTECH). The fluorescence ratio (515 nm/575 nm) was determined, background (no Api m 1) subtracted, and the data normalized to a positive control (nApi m 1 without nanobody).
RBL assay
In vitro degranulation was analyzed using RBL-SX38 cells expressing the human FcεRI receptor70. RBL-SX38 cells were a generous gift from Jean-Pierre Kinet. Cells were cultured in Minimum Essential Medium (GIBCO) supplemented with Earle’s salts, 2 mM L-glutamine, penicillin/streptomycin, 1 mM sodium pyruvate, 15% fetal calf serum, and 1–2 mg/mL G418 at 37 °C and 5% CO2. For analysis, cells were seeded in 96-well culture plates, 1 × 105 cells per well overnight at 37 °C and 5% CO2. The cells were sensitized with 1 µg/ml nb-hIgE for 1 h at 37 °C. After washing with Tyrode’s buffer (10 mM HEPES pH 7.4, 130 mM NaCl, 5 mM KCl, 1.4 mM CaCl2, 1 mM MgCl2, 5.6 mM glucose, and 0.1% BSA), the cells were incubated with dilution series of nApi m 1 and incubated for 1 h at 37 °C. As a reference, cross-linking was achieved by incubation with a 1:1000 dilution of polyclonal anti-human IgE antibody (Dako). Supernatants were saved before wash and lysis of cells with Tyrode’s buffer containing 0.1% triton-X. β-hexosaminidase release of viable versus lysed cells was assessed with 8 mM p-nitrophenyl N-acetyl-glucosaminide (Sigma). Results were calculated as percentage of total release.
Inhibition ELISA
Inhibition ELISA was performed using serum from 5 HBV-allergic patients sensitized to Api m 1. Clinical data of the patients with regard to HBV-specific allergic symptoms, sensitization profiles, age, and sex are reported in Supplementary Tables 1 and 2. High-binding 384 microwell plates (Greiner Bio-One) were coated o.n. at 4 °C with 5 µg/ml nApi m 1 diluted in PBS. After blocking for 30 min with TBS containing 4% BSA, the wells were incubated for 1 h at room temperature. shaking with 1 µM nb-hIgG1 diluted in TBS + 2% BSA. Serum was then added to the wells, and the plates were incubated for an additional 1 h. After washing, bound IgE was detected by 1 h incubation with AP-conjugated goat anti-hIgE antibody (Sigma) diluted 1:2500 in TBS + 2% BSA. The plates were developed with substrate solution (5 mg/mL 4-nitro-phenylphosphate, Sigma) for 100 min and measured at 405 nm. Background was subtracted, and IgE binding was normalized to the binding observed in presence of a nb-hIgG1 isotype control.
Basophil activation test
BAT was performed using serum from HBV-allergic patients sensitized to Api m 1. Clinical data of the patients with regard to HBV-specific allergic symptoms, sensitization profiles, age, and sex are reported in Supplementary Tables 1 and 2. BAT was performed with the Flow Cast kit (Bühlmann) according to the manufacturers´ recommendations. Peripheral EDTA blood of HBV-allergic patients was incubated with tenfold dilutions series of nApi m 1. Staining with anti-CD63-FITC and anti-CCR3-PE was performed in a 37 °C water bath for 15 min. Erythrocytes were lysed for 7.5 min followed by centrifugation at 600 × g for 5 min. Cell pellets were resuspended in 100 µl washing buffer, and basophil activation was evaluated by flow cytometry on a FACSCalibur (BD Biosciences) using FlowJo software (version 10.8.1) for analysis. Basophils were defined as CCR3+/SSClow cells and activated basophils were distinguished by being CD63 positive.
Based on the assay above, a fixed sub-maximal concentration of Api m 1 was chosen for each patient. Inhibition was assessed by pre-blocking Api m 1 with nb-hIgG1 (tenfold serial dilutions) in stimulation buffer for 1 h at room temperature. Peripheral EDTA blood from HBV-allergic patients was then incubated with the pre-blocked allergen, and activated basophils were stained and detected as mentioned before.
To assess inhibition of basophil activation upon HBV-stimulation versus Api m 1-stimulation, titrations of Api m 1 and HBV extract (Bühlmann) was performed as above. Based on the titration a fixed sub-maximal concentration of Api m 1 and a tenfold higher concentration of HBV was chosen for each patient. The inhibition assay was performed as described using a 20-fold molar excess of the bi-specific nb-IgG1 compared to Api m 1.
ImmunoCAP measurements
Specific IgE and IgG antibody measurements were performed by ImmunoCAPs on a Phadia 200 and 250 instrument (Thermo Fisher Scientific).
Mouse experiments
Female BALB/c mice were purchased from Charles River Laboratories (Erkrath, Germany) and used at age 8–10 weeks. They were housed under specified pathogen-free conditions in filter-top cages and received a standard diet. Housing of single mice was avoided. All cages were provided with nesting material, mouse houses and chew toys. Mice were kept with a constant day/night rhythm of 14 h day and 10 h night (day beginning at 5 am wintertime). Mice were allocated in groups and intraperitoneal (IP) injected three times (day 0, 7, 14) with either Dulbeccos phosphate buffered saline (DPBS) (n = 12) as a control or 25 µg HBV (Latoxan) adsorbed to 2 mg aluminum hydroxide (Imject alum, Pierce, Thermo Fisher Scientific). For evaluation of the HBV- and Api m 1-specific IgE and IgG1 response, peripheral blood was taken on day 0 and 21, and the serum was stored at −20 °C until further processing.
Sensitized mice were challenged on day 28 IP with 75 µg of Api m 1 contained in 100 μl DPBS. The challenge with HBV was performed identically using 30 µg of HBV in DPBS.
Anaphylaxis was monitored by measuring the core body temperature with an infrared thermometer (Voltocraft/Conrad Electronic) on the abdomen every 15 min for a period of 90 min. Temperature-time curves were integrated, and the area under curve (AUC) indicated as a measure for anaphylaxis, the baseline for the AUC curve being the pre-challenge body temperature.
Sensitized mice were treated by IP injection of the bispecific nb-hIgG1 using doses of 1200 µg per animal in a total volume of 200 μl DPBS. Mice injected with DPBS (non-sensitized) and sensitized mice injected with DPBS were used as controls. To test the potential of the bispecific nb-hIgG1 in preventing anaphylaxis, sensitized and treated mice were challenged by IP injection of Api m 1 4 h later as described above.
90 min after provocation mice were sacrificed and bled by heart puncture. The serum was stored at −20 °C for further analysis.
Analysis of mouse IgG1 and IgE antibodies and MCPT-1
Specific IgG1 levels were quantified in mouse serum by ELISA. Briefly, high-binding 384-well plates were coated with 1 µg/ml antigen and blocked with 4% skimmed milk in DPBS. Dilutions of samples were performed in 4% skimmed milk and incubated for 2 h before the addition of biotinylated rat anti-mouse IgG1 (BD Pharmingen; 1:1000). The plates were incubated for 30 min with horseradish peroxidase-conjugated streptavidin (Merck-Millipore) before adding the tetramethylbenzidine (TMB) substrate. The reaction was stopped with sulphuric acid and the absorbance read at 450 nm.
For the detection of specific IgE, plates were coated with 10 µg/ml of antigen. To reduce IgG in the serum, samples were diluted 1:2 in NaCl and pre-incubated with Protein G agarose at a ratio of 1:5 (Santa Cruz Biotechnology, Dallas, TX) while shaking for 1 h at room temperature. Following a centrifugation for 1 min at 12,000 × g the supernatant was used for the ELISA and the remaining Protein G agarose was discarded. Biotinylated rat anti-mouse IgE (BD Pharmingen) was used as secondary antibody. Blocking, detection, and incubation times were done according to the spec. IgG1 ELISA.
In addition, MCPT-1 levels in mice as marker for cellular degranulation and anaphylaxis were determined with a mouse MCPT-1 ELISA kit (Invitrogen). All steps were performed according to the manufacturer’s protocol.
Statistics & reproducibility
The quality of purified proteins was routinely assessed by SDS-PAGE. Kinetic data was fitted with a 1:1 binding model using the Octet System Data Analysis Software, Release 7.1. The activity assay and RBL assay were reproduced in two independent experiments. For experiments using patient serum, no statistical method was used to predetermine sample size. The number of patients was determined based on feasibility criteria and the availability of untreated patients with clinically confirmed HBV-allergy or sensitization in the Department of Dermatology and Allergology at Philipps University Marburg, Germany. The experiments were not randomized. The investigators were not blinded to allocation during experiments and outcome assessment, and no data were excluded from the analyses. Sex- and gender-based analysis of the samples was not performed.
For mice experiments, a temperature difference between two groups of at least 1.5 °C was considered as a clearly measurable difference, and significance was calculated with a one-tailed Wilcoxon–Mann–Whitney test, an alpha-error of 0.05 and an actual power of 0.8. This led to a sample size of 6 mice per group. Two independent experiments with 6 mice per group were performed. The investigators were not blinded to allocation during experiments and outcome assessment. Data from two mice had to be excluded: One mouse lost weight after the 3rd sensitization and had to be sacrificed 12 days before anaphylaxis provocation due to regional animal care regulations. The second mouse died during the anaphylaxis between 30 and 45 min after provocation. Both mice belonged to the group that was HBV sensitized, vehicle pre-treated and Api m 1 challenged.
Statistical analysis was performed using GraphPad Prism 10. Data was tested for normality and statistical analyses were performed accordingly.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Coordinates and structure factors for the two crystal structures are available in the Protein Data Bank entries 8PIH and 8PKC. Other protein structures used in this work are available in the Protein Data Bank as entry 1POC and 7NP9. Source data are provided with this paper.
References
Bilo, M. B. Anaphylaxis caused by Hymenoptera stings: from epidemiology to treatment. Allergy 66, 35–37 (2011).
Worm, M. et al. First European data from the network of severe allergic reactions (NORA). Allergy 69, 1397–1404 (2014).
Bilo, M. B. & Bonifazi, F. The natural history and epidemiology of insect venom allergy: clinical implications. Clin. Exp. Allergy 39, 1467–1476 (2009).
Galli, S. J. & Tsai, M. IgE and mast cells in allergic disease. Nat. Med. 18, 693–704 (2012).
Valenta, R. et al. Molecular aspects of allergens and allergy. Adv. Immunol. 138, 195–256 (2018).
Spillner, E., Blank, S. & Jakob, T. Hymenoptera allergens: from venom to “venome. Front. Immunol. 5, 77 (2014).
Jakob, T., Rafei-Shamsabadi, D., Spillner, E. & Muller, S. Diagnostics in Hymenoptera venom allergy: current concepts and developments with special focus on molecular allergy diagnostics. Allergo J. Int. 26, 93–105 (2017).
Muller, U. R., Johansen, N., Petersen, A. B., Fromberg-Nielsen, J. & Haeberli, G. Hymenoptera venom allergy: analysis of double positivity to honey bee and Vespula venom by estimation of IgE antibodies to species-specific major allergens Api m1 and Ves v5. Allergy 64, 543–548 (2009).
Kohler, J. et al. Component resolution reveals additional major allergens in patients with honeybee venom allergy. J. Allergy Clin. Immunol. 133, 1383–1389 (2014). 1389.e1-6.
Pucca, M. B. et al. Bee updated: current knowledge on bee venom and bee envenoming therapy. Front. Immunol. 10, 2090 (2019).
Elieh Ali Komi, D., Shafaghat, F. & Zwiener, R. D. Immunology of bee venom. Clin. Rev. Allergy Immunol. 54, 386–396 (2018).
Jepson, T. A., Hall, S. C. & Chung, J. K. Single-molecule phospholipase A(2) becomes processive on melittin-induced membrane deformations. Biophys. J. 121, 1417–1423 (2022).
Scott, D. L., Otwinowski, Z., Gelb, M. H. & Sigler, P. B. Crystal structure of bee-venom phospholipase A2 in a complex with a transition-state analogue. Science 250, 1563–1566 (1990).
Kubelka, V. et al. Primary structures of the N-linked carbohydrate chains from honeybee venom phospholipase A2. Eur. J. Biochem. 213, 1193–1204 (1993).
Gattinger, P. et al. Glycosylation enhances allergenic activity of major bee venom allergen Api m 1 by adding IgE epitopes. J. Allergy Clin. Immunol. 147, 1502–1504.e5 (2021).
Muller, U. R. et al. Type I skin reactivity to native and recombinant phospholipase A2 from honeybee venom is similar. J. Allergy Clin. Immunol. 96, 395–402 (1995).
Plum, M. et al. Structural and functional analyses of antibodies specific for modified core N-glycans suggest a role in T(H) 2 responses. Allergy 78, 121–130 (2023).
Sahiner, U. M. & Durham, S. R. Hymenoptera venom allergy: how does venom immunotherapy prevent anaphylaxis from bee and wasp stings? Front. Immunol. 10, 1959 (2019).
Sturm, G. J. et al. EAACI guidelines on allergen immunotherapy: hymenoptera venom allergy. Allergy 73, 744–764 (2018).
Kranert, P. et al. Safety and effectiveness of a 3-day rush insect venom immunotherapy protocol. Int. Arch. Allergy Immunol. 181, 111–118 (2020).
Dhami, S. et al. Allergen immunotherapy for insect venom allergy: a systematic review and meta-analysis. Allergy 72, 342–365 (2017).
James, L. K. et al. Long-term tolerance after allergen immunotherapy is accompanied by selective persistence of blocking antibodies. J. Allergy Clin. Immunol. 127, 509–516.e1-5 (2011).
Shamji, M. H. & Durham, S. R. Mechanisms of allergen immunotherapy for inhaled allergens and predictive biomarkers. J. Allergy Clin. Immunol. 140, 1485–1498 (2017).
Strobl, M. R. et al. The role of IgG(1) and IgG(4) as dominant IgE-blocking antibodies shifts during allergen immunotherapy. J. Allergy Clin. Immunol. 151, 1371–1378 (2023).
Orengo, J. M. et al. Treating cat allergy with monoclonal IgG antibodies that bind allergen and prevent IgE engagement. Nat. Commun. 9, 1421 (2018).
Shamji, M. H. et al. Passive prophylactic administration with a single dose of anti-Fel d 1 monoclonal antibodies REGN1908-1909 in Cat allergen-induced allergic Rhinitis: a randomized, double-blind, placebo-controlled clinical trial. Am. J. Respir. Crit. Care Med. 204, 23–33 (2021).
Atanasio, A. et al. Targeting immunodominant Bet v 1 epitopes with monoclonal antibodies prevents the birch allergic response. J. Allergy Clin. Immunol. 149, 200–211 (2022).
Gevaert, P. et al. Novel antibody cocktail targeting Bet v 1 rapidly and sustainably treats birch allergy symptoms in a phase 1 study. J. Allergy Clin. Immunol. 149, 189–199 (2022).
Bousquet, J., Fontez, A., Aznar, R., Robinet-Levy, M. & Michel, F. B. Combination of passive and active immunization in honeybee venom immunotherapy. J. Allergy Clin. Immunol. 79, 947–954 (1987).
Lessof, M. H., Sobotka, A. K. & Lichtenstein, L. M. Effects of passive antibody in bee venom anaphylaxis. Johns Hopkins Med. J. 142, 1–7 (1978).
Kamal, M. A. et al. REGN1908-1909 monoclonal antibodies block Fel d 1 in cat allergic subjects: Translational pharmacokinetics and pharmacodynamics. Clin. Transl. Sci. 14, 2440–2449 (2021).
Hamers-Casterman, C. et al. Naturally occurring antibodies devoid of light chains. Nature 363, 446–448 (1993).
Muyldermans, S. Nanobodies: natural single-domain antibodies. Annu. Rev. Biochem 82, 775–797 (2013).
Steeland, S., Vandenbroucke, R. E. & Libert, C. Nanobodies as therapeutics: big opportunities for small antibodies. Drug Discov. Today 21, 1076–1113 (2016).
De Genst, E. et al. Molecular basis for the preferential cleft recognition by dromedary heavy-chain antibodies. Proc. Natl. Acad. Sci. USA 103, 4586–4591 (2006).
Aagaard, J. B. et al. Nanobody-based human antibody formats act as IgE surrogate in hymenoptera venom allergy. Allergy 77, 2859–2862 (2022).
Jensen, R. K. et al. A potent complement factor C3-specific nanobody inhibiting multiple functions in the alternative pathway of human and murine complement. J. Biol. Chem. 293, 6269–6281 (2018).
Scott, D. L. et al. Interfacial catalysis: the mechanism of phospholipase A2. Science 250, 1541–1546 (1990).
Li, M. et al. Crystal structure of a dimerized cockroach allergen Bla g 2 complexed with a monoclonal antibody. J. Biol. Chem. 283, 22806–22814 (2008).
Niemi, M. et al. Molecular interactions between a recombinant IgE antibody and the beta-lactoglobulin allergen. Structure 15, 1413–1421 (2007).
Rouvinen, J. et al. Transient dimers of allergens. PLoS ONE 5, e9037 (2010).
Kitaura, J. et al. Evidence that IgE molecules mediate a spectrum of effects on mast cell survival and activation via aggregation of the FcepsilonRI. Proc. Natl. Acad. Sci. USA 100, 12911–12916 (2003).
Kawakami, T. & Kitaura, J. Mast cell survival and activation by IgE in the absence of antigen: a consideration of the biologic mechanisms and relevance. J. Immunol. 175, 4167–4173 (2005).
De Nardis, C. et al. A new approach for generating bispecific antibodies based on a common light chain format and the stable architecture of human immunoglobulin G1. J. Biol. Chem. 292, 14706–14717 (2017).
De Vlieger, D., Ballegeer, M., Rossey, I., Schepens, B. & Saelens, X. Single-domain antibodies and their formatting to combat viral infections. Antibodies 8, 1 (2019).
Flicker, S., Zettl, I. & Tillib, S. V. Nanobodies-useful tools for allergy treatment? Front. Immunol. 11, 576255 (2020).
Zettl, I. et al. Isolation of nanobodies with potential to reduce patients’ IgE binding to Bet v 1. Allergy 77, 1751–1760 (2022).
Jovcevska, I. & Muyldermans, S. The therapeutic potential of nanobodies. BioDrugs 34, 11–26 (2020).
Deiteren, A. et al. A first-in-human, single and multiple dose study of lunsekimig, a novel anti-TSLP/anti-IL-13 NANOBODY(R) compound, in healthy volunteers. Clin. Transl. Sci. 17, e13864 (2024).
Zavrtanik, U., Lukan, J., Loris, R., Lah, J. & Hadzi, S. Structural basis of epitope recognition by heavy-chain camelid antibodies. J. Mol. Biol. 430, 4369–4386 (2018).
Zahirovic, A. et al. Identification of bee venom Api m 1 IgE epitopes and characterization of corresponding mimotopes. J. Allergy Clin. Immunol. 143, 791–794.e5 (2019).
Kontermann, R. E. Strategies for extended serum half-life of protein therapeutics. Curr. Opin. Biotechnol. 22, 868–876 (2011).
Djurup, R., Malling, H. J., Sondergaard, I. & Weeke, B. The IgE and IgG subclass antibody response in patients allergic to yellow jacket venom undergoing different regimens of venom immunotherapy. J. Allergy Clin. Immunol. 76, 46–55 (1985).
Peng, Z. K., Naclerio, R. M., Norman, P. S. & Adkinson, N. F. Jr. Quantitative IgE- and IgG-subclass responses during and after long-term ragweed immunotherapy. J. Allergy Clin. Immunol. 89, 519–529 (1992).
Eckl-Dorna, J. et al. Two years of treatment with the recombinant grass pollen allergy vaccine BM32 induces a continuously increasing allergen-specific IgG(4) response. eBioMedicine 50, 421–432 (2019).
Zinkhan, S., Thoms, F., Augusto, G., Vogel, M. & Bachmann, M. F. On the role of allergen-specific IgG subclasses for blocking human basophil activation. Front. Immunol. 13, 892631 (2022).
Ejrnaes, A. M., Bodtger, U., Larsen, J. N. & Svenson, M. The blocking activity of birch pollen-specific immunotherapy-induced IgG4 is not qualitatively superior to that of other IgG subclasses. Mol. Immunol. 41, 471–478 (2004).
Strobl, M. R. et al. Affinity matters for IgE-blocking activity of allergen-specific antibodies. Allergy 78, 2543–2546 (2023).
Zha, L. et al. An unexpected protective role of low-affinity allergen-specific IgG through the inhibitory receptor FcgammaRIIb. J. Allergy Clin. Immunol. 142, 1529–1536.e6 (2018).
Arzt, L. et al. Immunological differences between insect venom-allergic patients with and without immunotherapy and asymptomatically sensitized subjects. Allergy 73, 1223–1231 (2018).
Erzen, R., Kosnik, M., Silar, M. & Korosec, P. Basophil response and the induction of a tolerance in venom immunotherapy: a long-term sting challenge study. Allergy 67, 822–830 (2012).
Zitnik, S. E. et al. Monitoring honeybee venom immunotherapy in children with the basophil activation test. Pediatr. Allergy Immunol. 23, 166–172 (2012).
Paolucci, M. et al. Targeting Ara h 2 with human-derived monoclonal antibodies prevents peanut-induced anaphylaxis in mice. Allergy 78, 1605–1614 (2023).
Braren, I. et al. Generation of human monoclonal allergen-specific IgE and IgG antibodies from synthetic antibody libraries. Clin. Chem. 53, 837–844 (2007).
Kabsch, W. Xds. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007).
Wiegand, T. W. et al. High-affinity oligonucleotide ligands to human IgE inhibit binding to Fc epsilon receptor I. J. Immunol. 157, 221–230 (1996).
Acknowledgements
Gratefully acknowledged is the excellent technical assistance by Nanna Breum Nielsen, Manuel Schulze-Dasbeck, and Lisa-Marie Kuhnhenne. Figures 4A and S6 were created with BioRender.com. This study was supported by the Novo Nordisk Foundation, grant NNF19OC0058484 (to ES), and the Independent Research Foundation Denmark through grants 9041-00291A (to ES) and 0135-00061B (to GRA) and by the Behring-Röntgenstiftung, Marburg, Germany, grant 66-0004 (to TJ).
Author information
Authors and Affiliations
Contributions
J.B.A., C.M., G.R.A., and E.S. designed the experiments. J.B.A., R.G., A.V.B.L., A.L., M.S., B.K.J., and M.M. performed experiments and analyzed the data. J.B.A. and R.G. performed the crystallization and structure analysis. J.B.A., C.M., W.P., B.D., A.S.R.B., and T.J. designed and performed the basophil activation tests and mouse experiments. J.B.A., C.M., G.R.A., and E.S. wrote the manuscript. All authors critically revised the article and approved the version to be published.
Corresponding authors
Ethics declarations
Competing interests
Aarhus University together with the authors J.B.A., M.M. and E.S. has published a patent (WO2023170282A1) covering the molecules described in this manuscript. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Pål Johansen, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Aagaard, J.B., Gandini, R., Ballegaard, AS.R. et al. Nanobody-based IgG simultaneously inhibit the allergenic and enzymatic activity of the dominant honeybee venom allergen. Nat Commun 17, 1814 (2026). https://doi.org/10.1038/s41467-026-69572-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-69572-0