Abstract
Non-contrast MRI, routinely used for the preoperative diagnosis of glioma tumors and establishing treatment strategies, provides the potential for assessing blood-brain barrier (BBB) status without using gadolinium-based contrast agents (GBCA) which could cause adverse events. Additionally, generative artificial intelligence (AI) models enable the synthesis of contrast-enhanced images from non-contrast images. Despite this potential, the heterogeneity of GBCA-induced features in tumor areas and error accumulation from inaccurate synthesis largely limit the efficacy of conventional generative models. To address these limitations, we introduce a contrast-free BBB status identification model (CBSI) that can identify BBB status with high accuracy using non-contrast MR images and generative diffusion AI networks. Trained and validated on a multi-center dataset of 1,535 patients, CBSI achieves an area under the curve (AUC) of 81.31%, surpassing the performance of the model using only non-contrast MR (AUC = 72.76%) and demonstrating comparable performance to the T1Gd MR model (AUC = 88.68%) in an external test set. Furthermore, validation on two public datasets (BraTS-Africa and BraTS-GLI) supports the generalizability of CBSI in BBB status identification. Notably, with accurate BBB status of synthetic T1Gd, the performance of glioma segmentation and grading is improved significantly compared to existing methods. Generalizability analysis indicates that CBSI can facilitate BBB status identification using synthetic T1Gd findings, avoiding GBCA adverse effects and streamlining clinical workflows.
Similar content being viewed by others
Introduction
The blood-brain barrier (BBB) consists of microvascular endothelial cells that create a barrier between the blood and the brain’s interstitial fluid, while also interacting with brain cells such as astrocytes and pericytes. Tight junctions and adherens junctions between adjacent endothelial cells strengthen the BBB, forming small pores (approximately 1.4–1.8 nm in diameter) that permit only a limited passage of small-sized particles. Thus, the status of BBB significantly influences treatment decisions and prognostic assessment of glioma patients1. In clinical practice, the BBB can be distinguished through gadolinium-enhanced T1-weighted (T1Gd) magnetic resonance (MR) images. Specifically, after injecting the gadolinium-based contrast agents (GBCA), glioma patients with disrupted BBB present as enhanced features in tumor areas2,3 (Fig. 1a). In contrast, an intact BBB restricts GBCA delivery to invasive margins, which results in non-enhancing features in glioma tumor areas. However, the use of GBCA carries potential risks, including nephrogenic systemic fibrosis, allergic-like hypersensitivity reactions, and gadolinium deposition in the brain4. Additionally, patient movement due to discomfort after GBCA injection may increase the likelihood and severity of image motion artifacts. Moreover, T1Gd scans are frequently not performed in many clinical scenarios; for example, neuroimaging may commence with non-contrast MRI during the initial diagnostic stage; the preoperative surveillance of asymptomatic low-grade gliomas often relies on T2-FLAIR sequences5. Unavailable T1Gd images result in the lack of crucial BBB status information, which may restrict critical tasks such as treatment planning or precise target delineation for radiation therapy planning. Therefore, accurately identifying BBB status using non-contrast MR images is essential for avoiding GBCA-induced adverse effects, assessing prognosis, and streamlining clinical workflow. This method can compensate for the lack of contrast-enhanced information in radiation therapy, enlarge training datasets for improved feature engineering and model development, and be combined with other data streams from the same patient to enhance biological understanding of the BBB.
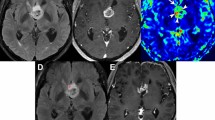
a Two types of glioma BBB status: intact (upper) and disrupted (lower). After GBCA injection, the tumor region represents an enhancement type (ET), classified as enhancing or non-enhancing on the T1Gd scan. b Sources of contrast enhancement, including the traditional method to identify BBB status with GBCA injection and our GBCA-free method to identify BBB status.
The identification of BBB status through non-contrast MR images remains challenging due to the lack of GBCA-induced features. While T1-weighted (T1) MR and T2 fluid-attenuated inversion recovery (T2-FLAIR) MR images cannot provide clear BBB status information, they can reveal changes in tumor micro-environments and tissue structures. Due to the limited BBB status information in non-contrast MR images, existing AI-based classification models6 achieve only modest performance in identifying BBB status. Since AI-based image generative models7,8,9,10 have shown promise in generating contrast-enhanced images from non-contrast images11,12,13,14, synthesizing T1Gd images from non-contrast MR images could enhance the identification of BBB status by providing clear features in tumor areas. However, most T1Gd generative methods are based on pixel-to-pixel translation15,16,17, which overlooks the heterogeneity of enhanced features in tumor areas. Notably, T1Gd MR images of glioma patients with different BBB statuses often exhibit significant intensity variations. This heterogeneity necessitates that a T1Gd generative model be capable of discerning different BBB statuses. To increase the focus on the heterogeneity of enhanced features in tumor areas, some models incorporate auxiliary segmentation tasks18 or directly input lesion masks into the synthesis model19 to improve performance, whereas synthetic images have not been closely integrated with the downstream identification task. Incorporating BBB status knowledge into an AI-based generative model may help advance the identification of BBB status using only non-contrast MR images.
The cascaded synthesis-identification strategy is a common solution to identify BBB status in non-contrast MR images. This method first employs an AI-based generative model for T1Gd synthesis, followed by using the synthetic T1Gd in a classification model. However, error propagation and accumulation resulting from incorrect enhancement type (ET) in the synthetic T1Gd MR images largely limited the accuracy of identifying BBB status. To reduce error accumulation, several studies have enhanced the robustness of downstream models by propagating uncertainty estimates between stages20. Besides, some studies adopt joint learning that leverages multi-task complementarity to enhance overall performance21,22,23. Yet, these strategies may provide limited support for the identification task due to potential model overfitting and imbalances in multi-task optimization.
In this work, we develop a contrast-free BBB status identification model (CBSI) based on a generative diffusion AI network and non-contrast MR images (Fig. 1b). To synthesize T1Gd-like images with accurate BBB status, we introduce an ET-guided generative diffusion AI model to learn the heterogeneous enhancements. Unlike most synthesis methods that rely solely on pixels and contours as conditional information, our diffusion model synthesizes T1Gd images with specific BBB status based on the given ET label. Besides, we incorporate an auxiliary tumor segmentation task in the diffusion model to enhance its semantic perception capabilities. To reduce error accumulation, we develop a contrastive maximum learning-based identification network for classifying BBB status through the synthetic T1Gd and non-contrast MR images. The contrastive maximum learning module distinguishes synthetic T1Gd images with pseudo-positive and negative BBB status labels to determine the realistic BBB status of glioma patients. We train and validate our CBSI model on 1535 patients from two centers and three public datasets. To evaluate the generalizability of our CBSI for accurate synthesis and identification of BBB status, we also assess the performance of our CBSI in glioma tumor segmentation and grading.
Results
Overview
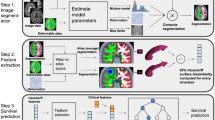
CBSI, as illustrated in Fig. 2a, is a synthesis-identification framework comprising two main steps. First, it synthesizes T1Gd images with pseudo-labels for BBB status (intact or disrupted) from T1 and T2-FLAIR inputs using an ET-guided generative diffusion model (see Fig. 2b). Second, it determines the realistic BBB status via an identification network with contrastive maximum learning. To evaluate its effectiveness, we trained and validated CBSI on a combined dataset (designated as the MR-1 dataset) of 721 glioma patients from Nanfang Hospital of Southern Medical University and The Cancer Genome Atlas Program (TCGA). The model was then tested on three independent external datasets: a dataset of 94 subjects from Zhujiang Hospital of Southern Medical University (designated as the MR-2 dataset), the MICCAI-CAMERA-Lacuna Fund BraTS-Africa 2023 Challenge dataset (BraTS-Africa Dataset)24 comprising 95 cases from Sub-Saharan Africa, and the Brain Tumor Segmentation (BraTS) 2023 Glioma dataset (BraTS-GLI Dataset)25 comprising 625 international cases.
a BBB status identification via contrastive maximum learning. The non-contrast MR images, including T1 and T2-FLAIR, are input into the generator to synthesize two T1Gd images with disrupted and intact BBB under the guidance of pseudo ET labels (enhancing \({\delta }^{+}\) and non-enhancing\({\,\delta }^{-}\)). The synthesized images are then fed into the identification network to obtain self-consistency scores \({\hat{S}}_{{{{\rm{c}}}}}^{+}\) and \({\hat{S}}_{{{{\rm{c}}}}}^{-}\), and contrastive maximum learning is used to obtain the final BBB state \({\hat{S}}_{{{{\rm{BBB}}}}}\). b ET-guided conditional diffusion. The non-contrast MR input is concatenated with noisy T1Gd images \({{{{\bf{x}}}}}_{t}\) to generate T1Gd images and segment tumors. ET labels \(\delta\) are embedded through an ET-guided module and fused with time-step embeddings to condition the features of the generator.
Performance on the identification of BBB status
A set of experiments was conducted to evaluate the performance of our CBSI in BBB status identification compared with state-of-the-art (SOTA) methods. The related performances of BBB status identification are presented in Table 1. Specifically, we trained an identification model using real T1Gd images and non-contrast MR to represent the optimal performance of BBB status identification. Similarly, the AI models based on non-contrast MR images were trained with the lowest performance for all model comparisons. The results were obtained by the 3D ResNet1826 network, which performed best among four widely used classification networks (ResNet18, MobileNet27, DenseNet12128, and ShuffleNet)29. As shown in rows 1–5 of Table 1, our CBSI achieves 14.82%, 12.30%, 13.16%, and 11.11% improvements in the area under the curve (AUC), accuracy (ACC), sensitivity (SEN), and specificity (SPE), respectively, compared with the model trained using non-contrast MR images, and it is close to the model trained using real T1Gd images. Besides, we evaluated the performance of BBB status identification using non-contrast MR images and synthetic T1Gd images from different synthesis methods, as shown in rows 6–11 of Table 1. Compared to the suboptimal Hi-Net model (Delong test: p < 0.05), our CBSI achieves 11.91%, 8.96%, 10.27%, and 7.14% improvements in AUC, ACC, SEN, and SPE, respectively. These impressive performances demonstrate the effectiveness of the proposed CBSI for improving the performance of BBB status identification using only non-contrast MR.
The receiver operating characteristic (ROC) and precision-recall curves (PRC) in Fig. 3a further illustrate the superior performance of CBSI. Additionally, Fig. 3b shows three confusion matrices of BBB status identification. The CBSI reduces the FPR from 32.50% to 25.00% and the FNR from 32.10% to 23.20% for identification using only non-contrast MR, approaching the 20.00% FPR and 21.40% FNR achieved by the real T1Gd-based model. In Fig. 3c, t-SNE visualization was employed to analyze the features extracted from the last convolutional layer of the direct-identification model trained with real T1Gd images. This visualization clearly illustrates that the BBB status-related feature synthesized by CBSI is close to the ground truth.
All analyses were based on 96 independent subjects with no technical replicates. a The receiver operating characteristic (ROC) and precision-recall curves (PRC) for BBB status identification using different methods. b Confusion matrices comparing our CBSI with two direct-identification methods—one using (w) real T1Gd and the other without (w/o). c t-SNE scatter plot of T1Gd features extracted during BBB status identification from real and synthetic images generated by CBSI; orange denotes disrupted and blue intact BBB. d Two representative visual comparisons of T1Gd images generated by the SOTA synthesis models. All evaluations use postprocessed images with pixel values mapped to [0, 255]. Tumor heatmaps within MRI images depict CA-induced enhancement (non-negative component of (T1Gd minus T1)), displayed within 0–150 to highlight subtle differences. Heatmaps below each MRI image show pixel-wise differences between synthetic and real T1Gd (synthetic minus real) within [−150, 150]; warm colors indicate higher positive, while cool colors indicate higher negative differences. e Quantitative evaluation of synthetic T1Gd images across the whole brain, grouped by BBB status (Intact n = 40; Disrupted n = 56). Each data point represents the mean metric from one patient’s brain region (96 independent biological replicates). Violin plots show MAE, SSIM, PSNR, and LPIPS performance. Each violin shows the 25th, 50th (median), and 75th percentiles as dotted lines, with boundaries spanning the full kernel density and all data points overlaid. f As in (e), but with a statistically analyzed tumor region in the form of box plots. Each box shows the median (solid line), the 25th and 75th percentiles (box bounds), and whiskers extending to values within 1.5 times the interquartile range. Statistical significance was assessed using two-sided Wilcoxon signed-rank tests, with exact p-values provided.
Performance on T1Gd synthesis
We compared our proposed CBSI with SOTA image synthesis methods on the internal testing set, including Pix2pix30, SwinUNETR31, CycleGAN32, Hi-Net33, and DDPM34. Figure 3d visualizes the synthetic T1Gd images from different models, and the T1Gd image generated by our CBSI presents richer details and clearer edges than those obtained by other methods, particularly in the tumor regions. The difference maps of our CBSI appear lighter in color, particularly in the tumor regions, indicating that the synthetic image has the fewest errors from the ground truth. Moreover, the tumor area marked by the boxes underscores the effectiveness of our ET-guided module in boosting the performance of enhanced feature synthesis. Figures 3e and f show that our CBSI outperforms all comparison methods in quantitative synthesis performance. Within the whole brain region (Fig. 3e), our CBSI achieves the lowest mean absolute error (MAE), highest peak signal-to-noise ratio (PSNR), highest structural similarity index measurement (SSIM), lowest Learned Perceptual Image Patch Similarity (LPIPS)35×100, and lowest Fréchet Inception Distance (FID)36 of 13.78 ± 2.60, 26.51 ± 2.30, 0.79 ± 0.04, 8.96 ± 1.45, and 22.73, respectively. In the tumor region (Fig. 3f) specifically, CBSI shows superior performance with the lowest MAE (22.38 ± 10.87), highest PSNR (24.33 ± 4.41), highest SSIM (0.72 ± 0.10), lowest LPIPS×100 (0.76 ± 0.60), and lowest FID (19.04), demonstrating its enhanced ability to capture detailed lesion-specific features.
Ablation study of T1Gd synthesis
We conducted an ablation experiment to verify the contribution of each component of our proposed CBSI to the T1Gd image synthesis and BBB status identification on the MR-1 testing set (as shown in Table 2). The baseline model is the image-guided diffusion model without the ET-guided module, contrastive maximum learning (CML), and auxiliary segmentation (SEG) module. Compared with the baseline, the whole CBSI framework shows overall improvement, with MAE reduced by 11.80%, PSNR increased by 3.85%, SSIM increased by 2.60%, LPIPS×100 reduced by 10.21%, and FID reduced by 39.41% in synthesis. In BBB status identification, AUC, ACC, SEN, and SPE increased by 12.58%, 8.96%, 10.27%, and 7.14%, respectively. In particular, adding CML achieved the greatest improvements in synthetic quantitative indicators and identification performance. These results highlight the importance of each component in CBSI in improving the BBB status identification performance.
Generalization to the external testing set
To assess the generalizability of the CBSI, we conducted experiments on the MR-2 dataset using the synthesis and identification sub-networks trained on the MR-1 training set. Compared to direct identification using non-contrast MR images, CBSI improved AUC (95% confidence interval, CI) from 72.76% (61.51%–83.63%) to 81.31% (71.36%–89.84%), ACC from 71.28% to 74.47%, and SEN from 69.84% to 74.60%, as shown in Table 1. Figures 4a and b present ROC, PRC, and confusion matrices, which show CBSI’s superior performance over other SOTA methods (Fig. 4a). As shown in Fig. 4b, CBSI reduced FPR and FNR, leading to higher accuracy in identifying BBB status. For T1Gd synthesis, CBSI demonstrated superior performance with the lowest MAE of 21.78 ± 7.45, highest PSNR of 24.99 ± 3.59, highest SSIM of 0.71 ± 0.09, lowest LPIPS×100 of 10.25 ± 1.28, and lowest FID of 31.80 for the whole brain and tumor regions (Table 3), and it also presented more realistic enhancement (Fig. 4c) in the tumor region. CBSI also delivered the strongest overall performance across three glioma subregions, including the non-enhancing tumor core, non-enhancing FLAIR hyperintensity, and enhancing tumor. These results indicate that CBSI remains effective on external data, demonstrating its strong generalization capabilities.
All evaluations utilize post‑processed images with pixel values mapped to [0, 255]. a ROC and PRC of BBB status identification by different methods. b Confusion matrices for BBB status identification, comparing our proposed CBSI with two direct-identification methods—one with (w) real T1Gd images as input and the other without (w/o). c Visual example comparison with SOTA synthesis models. Heatmaps overlaying the original MRI images illustrate CA-induced signal intensity increments (T1Gd minus real T1; scale 0–150). Corresponding heatmaps below illustrate pixel-wise differences between synthetic and real T1Gd images (synthetic minus real T1Gd; scale −150 to 150), where warm colors indicate positive errors and cool colors indicate negative errors. d Subjective evaluations, Task I: Visual quality assessment (n = 94 samples). Left: Histograms comparing quality scores assigned to real and synthetic T1Gd images by two radiologists using a three-point scale. Right: Inter-reader comparison of ratings on the same image set. e Subjective evaluations, Task II: ET annotation. Left: Sankey diagram comparing ET labels (enhancing or non-enhancing) assigned to real versus synthetic images. Right: Inter-reader comparison of ET annotation on the same image set. Performance was analyzed using standard agreement statistics (Fleiss’ κ, Gwet’s AC1; both reported with 95% confidence intervals and two-sided p-values for testing κ = 0) and comparative tests (two-sided Fisher’s exact test when the expected cell count <5; otherwise, two-sided Chi-square test reporting χ² statistic, degrees of freedom, effect size, and exact p-values). For small p-values that cannot be precisely represented, values are reported as p < \({10}^{-10}\). Panels d and e demonstrate no statistically significant differences between synthetic and real T1Gd images (all independence tests p > 0.05), and show strong inter-reader agreement in both evaluation tasks (all agreement coefficients >0.66, p < 0.01).
Clinical subjective evaluation
To evaluate clinical applicability, we introduced two complementary subjective evaluation tasks. Two radiologist readers (experienced radiologists with over 10 years of expertise in interpreting brain MR images) independently reviewed a randomized, blinded set of 94 real and 94 synthetic 3D T1Gd volumes from the external test set. Readers were asked to (1) rate the overall image quality using a three-point diagnostic scale (poor, satisfactory, high quality) and (2) determine the presence or absence of abnormal enhancement within the tumor region, i.e., conduct ET annotation. The scoring system categorized images into three levels: ‘Poor’ (score = 1) for non-diagnostic images with severely impaired quality, ‘Satisfactory’ (score = 2) for images with acceptable diagnostic quality despite some limitations, and ‘High quality’ (score = 3) for images with excellent diagnostic clarity. Detailed diagnostic evaluation criteria and representative cases were provided in the Supplementary Materials (Supplementary Table 1 and Supplementary Fig. 1). Performance was analyzed using standard agreement statistics (Fleiss’ kappa, Gwet’s AC1) and comparative tests (Fisher’s exact test when the theoretical frequency was less than 5; otherwise, the \({{{{\mathscr{X}}}}}^{2}\) test)37. As shown in Fig. 4d, over 90% of synthetic images were rated as ‘High quality’ (score = 3). For both readers, the quality ratings for synthetic and real images showed strong consistency (Fig. 4d left, AC1 = 0.916 and 0.907, both p < 0.01), with no significant difference in score distributions (both p = 0.06). Inter-reader agreements were also high (Fig. 4d right, all coefficients \(\ge\) 0.661 with p < 0.05) and score differences were not significant (all p > 0.79). For ET annotation (Fig. 4e), the synthetic image-based performance achieved strong diagnostic metrics. AUC, ACC, SEN, and SPE are 84.77%, 86.17%, 88.89%, and 80.65% for Reader 1, and 88.74%, 91.49%, 96.83%, 80.65% for Reader 2, respectively, with substantial inter-reader consistency (κ > 0.69; AC1 > 0.75; all p < 0.01) and no significant differences from real-image performance (p > 0.53). These results demonstrate that the synthesized T1Gd images maintain high visual quality and accurately reconstruct enhancement features critical for BBB status identification.
Generalization to other data protocols
To evaluate the generalization capability of CBSI, we conducted a set of experiments on the BraTS-Africa Dataset and the BraTS-GLI Dataset. Due to the lack of intact BBB samples in the BraTS-Africa Dataset, the AUC and SPE are not applicable; yet, ACC and SEN metrics can be provided. CBSI demonstrated robust T1Gd synthesis performance on the BraTS-Africa Dataset, achieving the lowest MAE (16.04 ± 2.36), highest PSNR (22.58 ± 1.82), highest SSIM (0.80 ± 0.03), lowest LPIPS×100 (5.92 ± 1.04), and lowest FID (37.49, Supplementary Table 2), along with more realistic contrast enhancement (Supplementary Fig. 2). Compared to direct BBB status identification using non-contrast MR images, CBSI improved ACC from 71.58% to 80.00% (Table 1), approaching the performance achieved with real T1Gd scans (85.26%). For the BraTS-GLI Dataset, CBSI exhibited consistent generalization performance. Compared with other synthesis methods, CBSI achieved the best performance across all synthesis metrics (Supplementary Table 2), and its enhancement patterns aligned well with clinical expectations (Supplementary Fig. 2). In BBB status identification, CBSI improved AUC (95% CI) from 60.23% (43.82–79.10%) for baseline non-contrast methods to 91.47% (89.97–92.90%), ACC from 73.60% to 83.04%, SEN from 73.91% to 82.93%, and SPE from 25.00% to 100% (Table 1), demonstrating statistically significant improvements (DeLong test: p < 0.05). In contrast, no significant difference was observed when comparing CBSI-based identification with that using real T1Gd images (DeLong test: p = 0.4707). These results underscore CBSI’s capacity to generalize across geographic, demographic, and technical variations.
Generalization to downstream glioma segmentation
To evaluate the generalizability of different synthesis methods to synthesize accurate glioma anatomical structures, we employed the pre-trained nnU-Net network for glioma segmentation on synthetic T1Gd images. As shown in Table 4, the T1Gd images synthesized from our proposed CBSI achieve higher Dice Similarity Coefficient (DSC) and Jaccard Index (Jaccard) and lower 95th percentile of Hausdorff Distance (HD95), Average Surface Distance (ASD), and Average Symmetric Surface Distance (ASSD). When using only synthetic T1Gd images for segmentation, CBSI outperforms Pix2pix, CycleGAN, SwinUNETR, Hi-Net, and DDPM by 15.49%, 2.50%, 20.59%, 1.23%, and 6.49% in DSC; 20.69%, 2.94%, 27.27%, 1.45%, and 9.37% in Jaccard; and achieves reductions of 75.89%, 45.05%, 76.09%, 15.56%, and 57.27% in HD95; 85.75%, 64.38%, 87.44%, 12.61%, and 61.9% in ASD; and 80.35%, 45.41%, 81.26%, 11.02%, and 49.55% in ASSD, respectively. When non-contrast MR images and synthetic T1Gd are used for segmentation, our CBSI achieves 1.12% increase in Jaccard, 7.83% reduction in HD95, 19.35% reduction in ASD, and 14.00% reduction in ASSD compared with suboptimal performance (Hi-Net). On the MR-2 dataset, the T1Gd image synthesized by CBSI also achieved the best segmentation performance compared with four SOTA methods. The segmentation results visualization of T1Gd images synthesized by different methods were shown in Fig. 5a. Further Wilcoxon tests were conducted, and the statistical results in Table 4 demonstrated that CBSI’s segmentation improvements were statistically significant in many comparisons, with p < 0.05. However, when combining non-contrast MR images with synthetic T1Gd for segmentation, differences across methods became less pronounced. This may be attributed to the fact that whole tumor masks were annotated based on T2-FLAIR images and include multiple components beyond the enhancing tumor region. In addition, while not all p-values in segmentation were significant, it is important to emphasize that our synthesis approach is primarily designed for BBB status identification, where methods like Hi-Net are less effective in capturing critical contrast-enhanced features. To better assess the contrast-specific synthesis quality of CBSI, we introduced an enhanced tumor segmentation task using public datasets with available subregion annotations (the BraTS-Africa Dataset and the BraTS-GLI Dataset). As shown in Supplementary Table 3, CBSI consistently achieved superior performance across all metrics. These results further support CBSI’s capability in preserving contrast-enhancing tumor features and improving segmentation of clinically relevant regions.
a Visual examples of glioma segmentation on the MR-2 dataset (n = 94 samples) obtained by different SOTA methods. In the magnified view, green indicates tumor ground truth, red shows segmentation masks on real or synthetic T1Gd images from our pre-trained glioma segmentation model, and blue marks their overlap, with greater overlap signifying better segmentation. b Performance of glioma grading obtained by the existing five synthesis models and our CBSI on samples from the MR-1 dataset with available glioma grade labels (n = 41 samples). Both overall and grade-specific (Grades 2–4) statistical analyses. The radar chart illustrates classification metrics with the four vertices representing AUC, ACC, SEN, and SPE, the center of the circle indicating 40% for the corresponding metric, and the outermost circle representing 100%.
Generalization to downstream glioma grading
Considering the BBB status of T1Gd lesions is correlated with the grades of glioma, we evaluated the generalizability of the CBSI in the downstream glioma grading task. Figure 5b presents a radar map that visually compares the comprehensive performance of glioma grading on T1Gd images synthesized by different synthesis methods. Our proposed CBSI shows the best grading performance, with the largest overall area covered by the polygon in the radar map. Specifically, the mean AUC (95% CI), ACC, SEN, and SPE for our CBSI are 86.55% (77.20–94.53%), 80.49%, 83.02%, and 79.63%. Among different grades of gliomas, the scores of AUC (95% CI), ACC, SEN, and SPE are 86.52% (73.67–97.25%), 82.93%, 83.33%, and 82.35% for grade 2 gliomas, 83.23% (70.56–94.52%), 75.61%, 80.00%, and 74.19% for grade 3 gliomas, and 89.92% (78.10%–98.18%), 82.93%, 85.71%, and 82.35% for grade 4 gliomas, respectively. Compared to Hi-Net, which has suboptimal overall performance, the mean AUC, ACC, SEN, and SPE of our method are improved by 22.09%, 23.75%, 25.73%, and 23.19%, respectively. This indicates that the T1Gd images synthesized by our method with accurate BBB status have high-level semantic information closer to the real T1Gd images.
Evaluation of label quality impact
To systematically investigate the effects of BBB status label quality, we perturbed the labels with various levels of noise, representing 0%, 40%, 60%, and 100% noise levels, corresponding to a progressive reduction in label quality. Using these perturbed labels, we synthesized T1Gd images and assessed their performance in downstream glioma segmentation and grading tasks. The experimental results presented in Supplementary Table 4 revealed that highly accurate labels (0% noise) achieved an AUC of 86.55% in glioma grading tasks, closely aligning with the 87.56% obtained using real T1Gd images. However, as noise increased to 40%, 60%, and 100%, the AUC significantly declined to 78.27%, 47.58%, and 35.36%, respectively. Similarly, segmentation metrics such as HD95, ASD, and ASSD exhibited worsening trends with increasing noise, indicating diminished segmentation accuracy as label quality deteriorated. These results highlight the critical importance of precise BBB status labels in ensuring the reliability and generalizability of synthesized T1Gd images for downstream tasks. By distinguishing the impacts of image synthesis and label quality, this experiment provides robust validation of CBSI’s performance under varying label conditions.
Discussion
Our study aims to predict BBB status from non-contrast MR images without the need for GBCA. Unlike directly synthesizing and identifying images, the proposed CBSI provides a perspective on synthesis-identification that synthesizes all potential images with different enhanced features and applies the contrastive maximum learning to distinguish synthetic T1Gd images with pseudo-positive and negative BBB status labels to determine the realistic BBB status of glioma patients.
Glioma T1Gd MR image synthesis aims to synthesize contrast information from available non-contrast MR images, thus providing a high reference value for BBB status identification while avoiding the risk of GBCA. Benefiting from the powerful ability of feature extraction, the convolutional neural network (CNN) has been widely used to synthesize glioma T1Gd images. Several U-Net-based38 and FCN-based39 methods demonstrate the feasibility of synthesizing T1Gd MR images from non-contrast MR images. Sharma et al.15 proposed a Pix2Pix-based approach, utilizing zero-filled images to represent missing ones and progressively learning to complete MR images. In addition to the CNN, the Transformer has been introduced in image synthesis to capture long-distance spatial dependencies. For instance, Liu et al.40 proposed a transformer-based model to capture intra- and inter-contrast dependencies and local- and global-context information for accurate synthesis efficiently. Recently, with the excellent performance of diffusion models in computer vision tasks, they have also been employed for synthesizing T1Gd images41,42. For example, Kim et al.42 adopted a conditional latent diffusion model to learn the mapping between source and target modalities in the latent space. However, these methods overlook disease-related semantic information, failing to accurately synthesize lesion-enhanced features. To address this, Huang et al.18 incorporate tumor segmentation as an auxiliary task in the synthesis model, which helps keep the representations closely related to tumor appearance for high-quality image synthesis. Despite advances, these methods focus on improving synthesis indicators, which neglect the heterogeneity of BBB status induced by GBCA in different patients.
Ensuring the realism of synthesized results has always been fundamental to synthesis-identification strategies, particularly when tackling the challenge of generating T1Gd images with heterogeneous lesion-enhanced features indicative of BBB disruption. As the conditional input of the synthesis model, the T1 image can provide nearly identical anatomical structural information to the target T1Gd image (except enhancement components), and the T2-FLAIR image contributes complementary features, such as peritumoral edema, necrosis, and mass effect, that are strongly associated with BBB integrity. For instance, tumor progression in gliomas often leads to necrosis, edema, and structural alterations that manifest as heterogeneous T1 signals and hyperintensity on T2-FLAIR images. These imaging patterns reflect microstructural heterogeneity, vascular permeability changes, and water diffusion properties, which are indirect indicators of BBB integrity. Besides, multi-task learning of contrast synthesis and tumor segmentation can strengthen the model’s attention to the tumor area. Thus, the task of the synthesis model is correspondingly simplified to learning and visualizing specific enhancement types (ET) in the tumor region rather than a pixel-to-pixel fine mapping of the entire T1 to T1Gd15,18,38,40. Our study is based on the hypothesis that non-contrast MRI images inherently contain latent information about the contrast differences between tumors and surrounding tissues. This hidden information can be learned and extracted by an AI model, which can then be used by an image generative network to synthesize reasonable T1Gd images. Recently, many studies have demonstrated the potential of synthetic T1Gd images as an auxiliary task in various downstream tasks, such as tumor segmentation16,18,40 and lesion detection22. These methods leveraged synthetic images to enrich engineering applications, improve model performance, or extend datasets, underscoring their value in complementary roles. Our CBSI framework differentiates itself from traditional intensity transformation methods by utilizing diffusion models, which excel in capturing complex, non-linear relationships between non-contrast and contrast-enhanced signals. Unlike pixel-wise mapping, diffusion models operate through iterative denoising, progressively reconstructing enhancement-relevant features from noise based on high-dimensional latent representations. Our ET-guided mechanism of GBSI incorporates the real ET labels (i.e., enhancing or non-enhancing) as additional guidance during model training. This ET label acts as a reliable and sufficient proxy for the presence or absence of BBB disruption, providing the model with essential diagnostic knowledge. This design principle transforms the synthesis task from an implicit prediction into a targeted reasoning process. The model is trained not to guess the absence and appearance of enhancement, but to bridge the anatomical features of non-contrast images and the BBB status with the given ET in each case. In this way, our framework transforms the synthesis task from solving a blind image pixel-to-pixel translation problem into learning contrast enhancement with real diagnostic and tumor area knowledge.
The contrastive maximum learning of our CBSI also plays a crucial role in ensuring the accurate identification of BBB status. Since the true ET is unknown at inference time, we synthesize all potential T1Gd images (i.e., enhancing and non-enhancing) with the pseudo-positive and negative BBB status labels and use them as the input of identification. Accordingly, the BBB status identification model is converted to determine the realistic BBB status of glioma patients. The identification network learns to predict self-consistency from the synthetic T1Gd images combined with the non-contrast MR images. Notably, the two synthesized images from the same case ultimately produce only one BBB status identification result. Therefore, based on the self-consistency of each synthetic T1Gd image, we adopt the optimal contrastive maximum learning mechanism to identify the image with higher self-consistency as realistic. The overall identification mechanism of our CBSI is fault-tolerant, effectively mitigating the accumulation of errors even when the synthesis quality is limited. The experimental results in Table 2 show the effectiveness of our contrastive maximum learning.
To balance the high quality of synthetic T1Gd images by the diffusion model, the shorter inference time, and the limited GPU memory, we chose the 2D architecture for T1Gd synthesis and BBB status identification. Specifically, existing 3D models adopt patch-based or resizing-based image preprocessing methods to obtain more inter-slice information for medical image synthesis within limited GPU memory. However, resizing the images will affect the resolution of tumors. Moreover, due to the lack of global context in the entire slice, the patch-based 3D model may negatively impact tasks such as whole-tumor segmentation and precise volumetric analysis, where comprehensive anatomical context is critical. In the inference stage, the 3D model usually requires a longer inference time due to overlapping, predicting each patch, and integrating them into a complete image. Although 2D models do not contain inter-slice information, they have the advantage of effectively capturing global information within slices and less inference time. Moreover, recent winning solutions in major public competitions have demonstrated the great potential of 2D-based models in medical imaging segmentation, identification, and detection43. These methods show the potential of 2D models in medical image analysis.
The drive to develop GBCA-free BBB status identification stems from legitimate clinical concerns: gadolinium-based agents carry risks like allergic reactions, tissue retention, and nephrogenic systemic fibrosis in renal-impaired patients. Presently, the clinical translation of these AI models remains questionable to some people. Fundamentally, the ‘black box’ nature of deep learning prevents rigorous pathophysiological validation; feature dimensions lack biological interpretability, and scale mismatches (e.g., voxel size vs. cellular pathology) preclude meaningful image-pathology correlation. Consequently, all hypotheses in enhanced image generation remain theoretical, lacking definitive in vivo validation. Such limitation is pervasive across deep learning-based diagnostic/prognostic studies, including high-impact publications44,45,46 (e.g., Ma et al.)44. Currently, it is common practice to prioritize models for their robustness and generalization, rather than their ability to replicate verified disease mechanisms. Crucially, clinical utility doesn’t require pixel-perfect synthetic enhancement in our study. Radiologists focus on tumor boundaries, size, and enhancement type (ET) derived from T1Gd, which are vital for surgical planning and prognosis. Enhancement patterns (e.g., ring vs. solid) hold secondary value. Therefore, the goal of our generative model should be iterative optimization toward diagnostic sufficiency—reliably visualizing tumor enhancement non-invasively—not mechanistic replication. Their value lies in approximating clinical decision-making utility, not replacing contrast imaging itself, while mitigating GBCA risks.
To validate the performance of our CBSI, we conducted a comprehensive set of experiments. First, we display the overall BBB status identification performance. As the basis for identification, higher-quality synthesis can effectively improve the performance of BBB status identification compared with the model using only non-contrast MR images. Therefore, we carried out both quantitative evaluations using image similarity metrics (MAE, PSNR, SSIM, LPIPS, and FID) and two subjective evaluations by radiologists: one assessed image visual quality using clinically meaningful criteria (enhancement localization, vascular realism, and pattern authenticity), and the other involved ET annotating. Then, we performed an ablation study to assess the effectiveness of each component. The auxiliary segmentation task allows the model to focus more on the lesion area, where the BBB could be disrupted. Improved segmentation performance implies a richer semantic understanding of lesions, thereby enhancing BBB status identification capabilities. Consequently, we evaluated glioma segmentation on synthetic T1Gd images using segmentation metrics (DSC, HD95, ASD, ASSD, and Jaccard). In clinical practice, glioma grade is also related to BBB status, with higher-grade tumors typically causing more significant BBB disruptions. The strong performance in downstream glioma grading suggests that the model may also be effective in identifying BBB status. Therefore, we finally display the downstream performance of glioma grading on synthetic T1Gd images using classification metrics (AUC, ACC, SEN, and SPE).
Despite its promising results, our study has several limitations. First, both the sampling mechanism of the diffusion model and the optimal selection strategy of CBSI incur greater inference time costs. Second, we used the entire tumor mask for segmentation training without dividing it into more detailed enhancing and non-enhancing tumor regions. This relatively coarse mask limits the effect of the auxiliary segmentation task. In subsequent research, we will introduce a fine lesion mask to acquire more precise regional attention. Third, the use of the ET label as a surrogate may introduce bias. Weakly enhancing cases are prone to inter-observer disagreement, leading to potential label noise and reduced ability to capture true enhancement. Although we employed multiple annotators and reported inter-rater consistency, mislabeling cannot be completely avoided. Illustrative failure cases are shown in Supplementary Fig. 3. Future studies may benefit from adopting more fine-grained or soft labels to better capture heterogeneity. Finally, we only verified the effectiveness of T1 and T2-FLAIR sequences in BBB status identification and did not evaluate other non-contrast MR sequences or functional MRI sequences. In this study, we use conventional non-contrast sequences and focus on T1 and T2-FLAIR images. These two modalities are widely available and provide a sufficiently large and consistent sample size for robust analysis. Although additional functional sequences (e.g., diffusion-weighted imaging, diffusion tensor imaging) are sometimes acquired based on institutional protocols, their inclusion would significantly reduce the sample size and introduce heterogeneity that may confound the interpretation of downstream performance. By relying on T1 and T2-FLAIR images, the CBSI benefits from consistent input data that supports its iterative denoising and latent feature reconstruction processes. In future work, we plan to collect multi-sequence data prospectively through multi-center collaborations. This will allow us to more comprehensively evaluate the impact of scan variability on our results.
Methods
Ethical statement
Data collection protocols and the use of images in this study were approved by the ethics committee of Zhujiang Hospital and Nanfang Hospital (NFEC-2021-430). Informed consent was waived for retrospectively collected MR images. The study followed the Declaration of Helsinki and complies with all relevant ethical regulations.
Dataset and annotation
A total of 1535 glioma patients were included in this study, sourced from two clinical centers and three publicly available datasets. Specifically, 815 patients were collected from Nanfang Hospital of Southern Medical University, the TCGA, and Zhujiang Hospital of Southern Medical University. The first two datasets were mixed to construct the MR-1 dataset, which included T1, T2-FLAIR, and T1Gd images of the brain from 721 patients before glioma surgery. MR volumes were obtained using Philips, Siemens, and GE scanners at 1.5 T and 3 T field strengths. The in-plane pixel spacings of the MR images ranged from 0.36 mm to 1.0 mm with an average of 0.55 mm, and the slice thicknesses ranged from 2.0 mm to 10.4 mm with an average of 6.0 mm. The in-plane resolution was 512\(\,\times\) 512 with the number of slices ranging from 12 to 34. The dataset from Zhujiang Hospital of Southern Medical University, labeled as the MR-2 dataset, contained 94 subjects. Acquisitions were performed on multi-vendor (Philips and GE) MR scanners, including T1, T2-FLAIR, and T1Gd sequences. The in-plane pixel spacings ranged from 0.43 mm to 0.53 mm with an average of 0.46 mm. The in-plane resolution was 512 \(\times\) 512 with the number of slices from 16 to 42. The slice thicknesses ranged from 3.3 mm to 7.97 mm with an average of 6.81 mm. Data division was performed by randomly selecting 96 cases from the MR-1 clinical dataset as an internal validation set and the remaining 625 cases as a training set. The MR-2 clinical dataset was used as an independent external validation set. Data division was performed by randomly selecting 55 cases, 96 cases, and the remaining 570 cases from the MR-1 dataset as the internal validation set, internal testing set, and training set. The MR-2 dataset was used as an independent external test set. The tumor masks and ET labels were automatically generated using our enhancement identification and segmentation networks and then manually adjusted by two experienced radiologists with over 10 years of expertise in interpreting brain MR images. The adjustment is based on the relative enhancement appearance on T1Gd, using adjacent white matter as a reference. To assess inter-annotator reliability, we randomly selected 50 cases (6%) of the MR-1 and MR-2 datasets, and their ET annotations were assessed using inter-class correlation coefficients (ICC). The calculated ICC is 0.917, which indicates excellent agreement and supports the reliability of the annotation process. Among these 815 cases, 514 were labeled as having disrupted BBB and 301 as having intact BBB.
To further verify CBSI’s generalizability across other countries and healthcare jurisdictions, two publicly available datasets obtained as part of the Brain Tumor Segmentation (BraTS) Challenge project (through Synapse ID: syn51156910) were additionally included: the BraTS-Africa Dataset24 and the BraTS-GLI Dataset25. Each case features standardized 1 × 1 × 1 mm³ resolution, consistent multi-sequence MRI (T1, T2, T2-FLAIR, and T1Gd), and tumor subregion masks. BBB status was annotated by experienced radiologists. The BraTS-Africa dataset contains 95 cases from Sub-Saharan Africa, all labeled as exhibiting disrupted BBB, consistent with the enhancing tumor annotations from the original challenge. The BraTS-GLI dataset comprises 625 cases from international cohorts, with 621 cases labeled as having disrupted BBB and 4 as having intact BBB.
Metrics
MAE, SSIM, and PSNR were used to evaluate pixel-level similarity and image quality, while FID and LPIPS were employed to measure the distributional and perceptual consistency between synthetic and real images. These five quantitative metrics collectively assess the clinical relevance of synthetic T1Gd images by capturing contrast, anatomical fidelity, and enhancement patterns. To evaluate the accuracy of lesion semantics in synthetic T1Gd images, we calculated the DSC, HD95, ASD, ASSD, and Jaccard index between the ground truth and the masks segmented from synthetic images by a pre-trained nnU-Net47 glioma segmentation network. For BBB status identification and glioma grading performance, we used AUC (with 95% CI), ACC, SEN, and SPE. The formulas for all metrics are provided in Supplementary Equations (1)–(14). Statistical significance was assessed using the DeLong test for the identification task and the Wilcoxon signed-rank test for synthesis and segmentation tasks to compare our CBSI against other models. Unless specified, all tests were two-sided, with a p-value < 0.05 considered significant, and no adjustment for multiple comparisons was made.
Statistics and reproducibility
Statistical analyses were conducted using Python (version 3.8). All statistical tests were two-sided, and p-values < 0.05 were considered statistically significant. The sample size was determined by the availability of eligible cases in the dataset, and no statistical method was used to predetermine the sample size. No data were excluded from the analyses. Data were randomly divided into training, validation, and test sets to ensure balanced distribution and to avoid sampling bias. The investigators were not blinded during data processing or analysis.
Image preprocessing and postprocessing
The image preprocessing included offset correction, registration, skull stripping, and grayscale normalization. The N4ITK48 corrected magnetic field inhomogeneities. T1 and T1Gd volumes were aligned to T2-FLAIR using FSL49 and ANTs50. Skull stripping used the BET in FSL. A binary brain mask excluded the background, and images were cropped to 424 × 424 in the plane. Grayscale normalization51 adjusted gray values to compensate for intensity variability, and images were then linearly transformed to the range [-1, 1]. Specifically, given an MRI scan \({{{\bf{I}}}}\), and each voxel has intensity \({v}_{k},{k}=1,2,...{{{\rm{N}}}}\), the preprocessing grayscale normalization procedure is as follows:
Step 1. Calculate the intensity histogram with 254 bins and obtain the gray value of the highest histogram bin \(\hat{v}\).
Step 2. Calculate the robust deviation \(\phi=\sqrt{\frac{{\sum }_{k=1}^{{{{\rm{N}}}}}{\left(\hat{v}-{v}_{k}\right)}^{2}}{N}}\).
Step 3. Subtract \(\hat{v}\) and divide by the robust deviation \(\phi\).
Step 4. Multiply a constant \(\varphi\) and add a constant \({v}_{0}\). For the T2-FLAIR, T1, and T1Gd, \(\varphi\) is 30, 31, and 31, and \({v}_{0}\) is 75, 99, and 99, respectively.
Step 5. Set the intensities that are below 0 or above 255 to 0 and 255, respectively.
Step 6. Linearly transform the voxel’s value range from [0, 255] to [-1, 1] by \({{{{\bf{I}}}}}_{{{{\rm{normalized}}}}}=\frac{{{{{\bf{I}}}}}_{{{{\rm{original}}}}}}{255}\times 2-\)1. In postprocessing, reverse normalization is applied to map the values of the synthetic T1Gd image from [-1, 1] back to [0, 255] as follows:
Step 7. Reverse normalization by \({{{{\bf{I}}}}}_{{{{\rm{original}}}}}=\frac{\left({{{{\bf{I}}}}}_{{{{\rm{normalized}}}}}+1\right)}{2}\times 255\).
Experimental setting
The experimental setup was conducted on a Linux operating system deployed on a server boasting a 20-core Intel Xeon Gold 6133 CPU @ 2.50 GHz. An NVIDIA GeForce RTX 3090 graphical processing unit (GPU) with 24GB of memory was utilized. The input size of the synthesis model and identification model is 424 × 424. The synthesis model was trained for 200 epochs using the Adam optimizer (weight decay of 1 × 10-4) with a step-decay learning rate, starting at 1 × 10-4 and reduced tenfold at the 100th and 160th epochs. The identification model was trained for 100 epochs using the Adam optimizer (weight decay of 5 × 10-4) with a warm-up cosine annealing schedule. The learning rate rose from 1 × 10-5 to 5 × 10-4 in the first 5 epochs and then decayed cosinely back to 1 × 10-5 over the remaining epochs. The learning rate was linearly warmed up and followed by cosine annealing. The parameter settings are \({\lambda }_{{{{\rm{c}}}}}\) = 0.5, \({\lambda }_{{{{\rm{seg}}}}}\) = 0.1, and \({\lambda }_{{{{\rm{r}}}}}\) is set to achieve the best F1 on the MR-1 validation set.
Algorithm development
The architecture of our CBSI framework is depicted in Fig. 2. Our goal is to identify BBB status SBBB from non-contrast MR images \({{{{\bf{I}}}}}_{{{{\rm{NC}}}}}\) that include T1 and T2-FLAIR MR images. Let \({{{\mathscr{D}}}}\) be the identification model, and the identification is formulated as:
However, directly identifying BBB status from non-contrast images remains challenging due to the limited BBB status information in these images. An intuitive solution is to develop a framework that combines the T1Gd synthesis model \(G\) with the identification model, which visualizes enhancements to improve the identification performance. We formulate this synthesis-identification strategy as:
Due to the insufficient learning of enhanced features heterogeneity in the lesion area and the propagation and accumulation using the cascaded synthesis-identification strategy, the synthetic T1Gd images from the commonly used medical image synthesis method in Eq.(2) largely impact the accuracy of the BBB status identification. To address these issues, we propose our CBSI framework to compulsorily synthesize two T1Gd MR images by enhancing ET \({\delta }^{+}\) and non-enhancing ET \({\delta }^{-}\). Moreover, synthetic two T1Gd images \({{{{\bf{I}}}}}_{{{{\rm{CE}}}}}^{+}\) and \({{{{\bf{I}}}}}_{{{{\rm{CE}}}}}^{-}\) are fed into the identification network, which enhances the learning of heterogeneous enhancements for better BBB status identification. The paradigm is summed up as:
Conditional diffusion
To learn the contrast-enhancement distribution, our CBSI adopts a conditional diffusion model as the synthesis backbone network, which is designed based on the DDPM. The diffusion model contains two Markov chain processes: a forward process and a reverse process. In the forward process, a sequence of noisy images \({{{{{\bf{x}}}}}_{1},\ldots,{{{\bf{x}}}}}_{T}\) is created by gradually adding mean-zero Gaussian noises \({{\epsilon }_{1},\ldots,\epsilon }_{T}\) with a constant variance \({\sigma }^{2}\) to the initial T1Gd image \({{{{{\bf{I}}}}}_{{{{\bf{CE}}}}}={{{\bf{x}}}}}_{0}\sim q\left({{{{\bf{x}}}}}_{t}\right)\) in T time steps. Given a variance schedule \({{\beta }_{1},\ldots,\beta }_{T}\) of the T time steps, the forward process is defined as:
The noisy image \({{{{\bf{x}}}}}_{t}\) can be easily sampled from \({{{{\bf{x}}}}}_{0}\) in a closed form by exploiting the additivity of independent Gaussian noise:
where \({\alpha }_{t}=1-{\beta }_{t},{\bar{\alpha }}_{t}={\prod }_{i}={1}^{t}{\alpha }_{i},\epsilon \sim {{{\mathscr{N}}}}\left({{{\bf{0}}}},{\sigma }^{2}\right)\), and the conditional probability distribution is:
In the reverse process, the diffusion model learns a Markov chain process to convert the Gaussian distribution into the real data distribution by predicting the parameterized Gaussian transition \(p\left({{{{\bf{x}}}}}_{t-1}|{{{{\bf{x}}}}}_{t}\right)\). Some studies52 propose that it is feasible to estimate noise, clean target images, or the mean term to represent this transition. To speed up convergence and reduce the training time, we use the clean target image generator \({\mu }_{\theta }\) to predict \({{{{\bf{x}}}}}_{0}\) directly at each time step. Accordingly, the image \({{{{\bf{x}}}}}_{t-1}\) can be formulated as:
ET-guided mechanism
The heterogeneity of enhanced features is a key characteristic of T1Gd images. Therefore, we designed an ET-guided synthesis strategy to map the non-contrast MR images into the enhancing or non-enhancing T1Gd feature space with distinction. The ET-guided module controls the reverse process in our diffusion model, taking the ET label as input and gradually synthesizing the target image over \(T\) time steps. The whole reverse process can be formulated as:
We follow the diffusive sampling strategy of the denoising diffusion implicit model53 to accelerate the reverse process. Specifically, we transform the ET label to an ET conditional representation \({{{\bf{w}}}}\) using a class embedding layer, and modulate feature map m by affine transformation \({{{\mathscr{A}}}}\left(\cdot \right)\):
where \({{{\bf{e}}}}\) is a time steps-conditional representation in the diffusion model, \({{{{\mathscr{M}}}}}_{{{{\rm{shift}}}}}\) and \({{{{\mathscr{M}}}}}_{{{{\rm{scale}}}}}\) denote the embedding layers for obtaining shift and scale vectors, respectively, and \(\oplus\) and \(\odot\) present element-wise addition and multiplication operations, respectively.
Auxiliary segmentation
Besides the ET-guided, we introduce whole tumor segmentation as an auxiliary task to produce lesion region attention. The segmentation network is composed of an encoder \({{{\mathscr{H}}}}\) shared with the synthesis network and a specific decoder \({{{{\mathscr{F}}}}}_{{{{\rm{seg}}}}}\). The output probability map \(\hat{y}\) can be formulated as:
Contrastive maximum learning
The goal of the contrastive maximum learning is to determine a more rational lesion enhancement in synthetic T1Gd images and then obtain the corresponding BBB status. To achieve this, the identification network is trained to predict the self-consistency score \({S}_{{{{\rm{c}}}}}\) of each synthetic T1Gd image first. On account of CBSI obtaining two synthetic T1Gd images with heterogeneous enhancements from the same case in the synthesis stage, the self-consistency score label was converted from the matching of the ET-guided label \(\delta {\prime}\) and the real ET label \(\delta\):
where \({\mathbb{I}}\left(\cdot \right)\) is the indicator function, which outputs a value of 1 when the condition is met and 0 otherwise. The predicted self-consistency score \({\hat{S}}_{{{{\rm{c}}}}}\) is presented as:
The identification network predicts self-consistency through the realism of synthetic details and the matching between non-enhanced MR and synthetic T1Gd images within tumor slices. In the inference phase, given our 2D network and the 3D nature of the image, we integrate the identification network’s predictions for each 2D slice containing the glioma region to obtain a total self-consistency score for each synthetic 3D T1Gd image. We formulate this as:
where \(i\) and n represent the slice index and the slice numbers of the input image within the tumor region, and \({\lambda }_{{{{\rm{c}}}}}\) denotes the probability threshold parameter. Then the optimal selection strategy is used to obtain the rationality score \({\hat{S}}_{{{{\rm{r}}}}}\) and the final BBB status identification result \({\hat{S}}_{{{{\rm{BBB}}}}}\):
where \({{{\mathscr{J}}}}\left(\cdot \right)\) is a softmax function and \({\lambda }_{{{{\rm{r}}}}}\) denotes the probability threshold parameter. This contrastive maximum learning process integrates different enhanced features related to BBB status to improve identification performance.
Overall loss function and optimization
The training of the proposed CBSI consists of two stages. In the first stage, the synthesis and segmentation modules are optimized simultaneously. The combined loss \({L}_{G}\) is defined as:
where \({\lambda }_{{{{\rm{seg}}}}}\) is the weight factor of segmentation loss. Specifically, the two losses \({L}_{{{{\rm{syn}}}}}\) and \({L}_{{{{\rm{seg}}}}}\) are defined as a cross-entropy loss and a Dice loss:
where N is the number of pixels in the input image, \({a}_{j}\) and \({b}_{j}\) represent the \(j\)-th pixel value in ground-truth annotation y and the probability map \(\hat{{{{\bf{y}}}}}\), respectively.
The loss function of the identification model in the second stage is a binary cross-entropy loss, which can be formulated as:
Network architecture
The sharing encoder \({{{\mathscr{H}}}}\) includes four steps, each with two residual blocks (ResBlock) and a self-attention module. All steps except the final one also have a downsample block. Each ResBlock has two convolutional blocks with a 3 \(\times\)3 convolutional layer, instance normalization, and ReLU activation. The decoder of the synthesis network has a similar architecture to the encoder but includes an upsampling block with nearest neighbor upsampling and a 1 \(\times\)1 convolutional layer. Two ResBlocks and a self-attention module are in the synthesis network bottleneck. In encoding and decoding, embeddings from the ET and time step t are fed into the first ConvBlock of each ResBlock to guide the generation. Each ET-guided module contains a ReLU activation and a fully connected (FC) layer with a specific embedding length depending on the level of the image generator. The decoder \({{{{\mathscr{F}}}}}_{{{{\rm{seg}}}}}\) has three convolutional blocks followed by a 1 \(\times\)1 convolutional layer and a sigmoid layer. Each convolutional block has an upsampling, a 3\(\times\)3 convolutional layer, and ReLU activation.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The sample subdataset of fully anonymized brain MR images with expert annotations in this study has been deposited in the GitHub repository at https://github.com/SMU-MedicalVision/CBSI-master. The remaining datasets used in this study are currently not permitted for public release by the respective institutional review boards (IRBs). Access to the full clinical dataset for non-commercial academic research requires approval from the aforementioned IRB. Researchers seeking access pertaining to the study may submit a formal request with a detailed research proposal to the corresponding author at [Wei Yang; weiyanggm@gmail.com]. Requests will be reviewed within 15 working days. In addition, Source data are provided with this paper.
Code availability
All source code of the CBSI is available under an open-source license at https://github.com/SMU-MedicalVision/CBSI-master.
References
Tan, A. C. et al. Management of glioblastoma: state of the art and future directions. CA Cancer J. Clin. 70, 299–312 (2020).
Steeg, P. S. The blood–tumour barrier in cancer biology and therapy. Nat. Rev. Clin. Oncol. 18, 696–714 (2021).
Arvanitis, C. D., Ferraro, G. B. & Jain, R. K. The blood–brain barrier and blood–tumour barrier in brain tumours and metastases. Nat. Rev. Cancer 20, 26–41 (2020).
Gulani, V., Calamante, F., Shellock, F. G., Kanal, E. & Reeder, S. B. Gadolinium deposition in the brain: summary of evidence and recommendations. Lancet Neurol. 16, 564–570 (2017).
Ius, T. et al. The benefit of early surgery on overall survival in incidental low-grade glioma patients: a multicenter study. Neuro-Oncol. 24, 624–638 (2022).
He, K., Zhang, X., Ren, S. & Sun, J. Deep residual learning for image recognition. In 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Las Vegas, NV, USA 770–778 (IEEE, 2016).
Lee, S. et al. Emergency triage of brain computed tomography via anomaly detection with a deep generative model. Nat. Commun. 13, 4251 (2022).
Yao, Z. et al. Virtual elastography ultrasound via generative adversarial network for breast cancer diagnosis. Nat. Commun. 14, 788 (2023).
Chang, Q. et al. Mining multi-center heterogeneous medical data with distributed synthetic learning. Nat. Commun. 14, 5510 (2023).
Li, Y. et al. Virtual histological staining of unlabeled autopsy tissue. Nat. Commun. 15, 1684 (2024).
Xu, C., Zhang, D., Chong, J., Chen, B. & Li, S. Synthesis of gadolinium-enhanced liver tumors on nonenhanced liver MR images using pixel-level graph reinforcement learning. Med. Image Anal. 69, 101976 (2021).
Müller-Franzes, G. et al. Using machine learning to reduce the need for contrast agents in breast MRI through synthetic images. Radiology 307, e222211 (2023).
Chung, M. et al. Deep learning to simulate contrast-enhanced breast MRI of invasive breast cancer. Radiology 306, e213199 (2022).
Zhang, T. et al. Synthesis of contrast-enhanced breast MRI using T1-and multi-b-value DWI-based hierarchical fusion network with attention mechanism. In Medical Image Computing and Computer Assisted Intervention – MICCAI 2023 79–88 (Springer, 2023).
Sharma, A. & Hamarneh, G. Missing MRI pulse sequence synthesis using multi-modal generative adversarial network. IEEE Trans. Med. Imaging 39, 1170–1183 (2019).
Xin, B., Hu, Y., Zheng, Y. & Liao, H. Multi-modality generative adversarial networks with tumor consistency loss for brain MR image synthesis. In 2020 IEEE 17th International Symposium on Biomedical Imaging (ISBI) 1803–1807 (IEEE, 2020).
Li, H. et al. DiamondGAN: unified multi-modal generative adversarial networks for MRI sequences synthesis. In Medical Image Computing and Computer Assisted Intervention – MICCAI 2019 (eds Shen, D. et al.) 795–803 (Springer International Publishing, Cham, 2019).
Huang, P. et al. Common feature learning for brain tumor MRI synthesis by context-aware generative adversarial network. Med. Image Anal. 79, 102472 (2022).
Guo, P., Wang, P., Zhou, J., Patel, V. M. & Jiang, S. Lesion mask-based simultaneous synthesis of anatomic and molecular mr images using a gan. In Medical Image Computing and Computer Assisted Intervention – MICCAI 2020 104–113 (Springer, 2020).
Mehta, R. et al. Propagating uncertainty across cascaded medical imaging tasks for improved deep learning inference. IEEE Trans. Med. Imaging 41, 360–373 (2021).
Xu, C. et al. Contrast agent-free synthesis and segmentation of ischemic heart disease images using progressive sequential causal GANs. Med. Image Anal. 62, 101668 (2020).
Zhao, J. et al. Tripartite-GAN: synthesizing liver contrast-enhanced MRI to improve tumor detection. Med. Image Anal. 63, 101667 (2020).
Brown, A. et al. Detecting shortcut learning for fair medical AI using shortcut testing. Nat. Commun. 14, 4314 (2023).
Adewole, M. et al. The brain tumor segmentation (BraTS) challenge 2023: Glioma segmentation in sub-Saharan Africa patient population (BraTS-africa). Preprint at https://arxiv.org/abs/2305.19369 (2023).
Karargyris, A. et al. Federated benchmarking of medical artificial intelligence with MedPerf. Nat. Mach. Intell. 5, 799–810 (2023).
He, K., Zhang, X., Ren, S. & Sun, J. Deep residual learning for image recognition. In Proc. IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Las Vegas, NV, USA 770–778 (IEEE, 2016).
Howard, A. G. et al. Mobilenets: efficient convolutional neural networks for mobile vision applications. Preprint at https://arXiv.org/abs/1704.04861 (2017).
Huang, G., Liu, Z., Van Der Maaten, L. & Weinberger, K. Q. Densely connected convolutional networks. In 2017 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Honolulu, HI, USA 4700–4708 (IEEE, 2017).
Zhang, X., Zhou, X., Lin, M. & Sun, J. ShuffleNet: an extremely efficient convolutional neural network for mobile devices. In 2018 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), Salt Lake City, UT, USA 6848–6856 (IEEE, 2018).
Isola, P., Zhu, J.-Y., Zhou, T. & Efros, A. A. Image-to-image translation with conditional adversarial networks. In 2017 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Honolulu, HI, USA 5967–5976 (IEEE, 2017).
Hatamizadeh, A. et al. Swin UNETR: swin transformers for semantic segmentation of brain tumors in MRI images. In Brainlesion: Glioma, Multiple Sclerosis, Stroke and Traumatic Brain Injuries 272–284 (Springer, 2021).
Zhu, J.-Y., Park, T., Isola, P. & Efros, A. A. Unpaired image-to-image translation using cycle-consistent adversarial networks. In 2017 IEEE International Conference on Computer Vision (ICCV), Venice, Italy 2242–2251 (IEEE, 2017).
Zhou, T., Fu, H., Chen, G., Shen, J. & Shao, L. Hi-Net: hybrid-fusion network for multi-modal MR image synthesis. IEEE Trans. Med. Imaging. 39, 2772–2781 (2020).
Ho, J., Jain, A. & Abbeel, P. Denoising diffusion probabilistic models. In Advances in Neural Information Processing Systems 33 (NeurIPS 2020), Vancouver, Canada 6840–6851 (Curran Associates, Inc., 2020).
Zhang, R., Isola, P., Efros, A. A., Shechtman, E. & Wang, O. The unreasonable effectiveness of deep features as a perceptual metric. In 2018 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), Salt Lake City, UT, USA 586–595 (IEEE, 2018).
Heusel, M., Ramsauer, H., Unterthiner, T., Nessler, B. & Hochreiter, S. Gans trained by a two time-scale update rule converge to a local Nash equilibrium. In Advances in Neural Information Processing Systems 30 (NIPS 2017), Long Beach, CA, USA6626–6637 (Curran Associates, Inc., 2017).
Doi, M., Takahashi, F. & Kawasaki, Y. Bayesian noninferiority test for 2 binomial probabilities as the extension of Fisher exact test. Stat. Med. 36, 4789–4803 (2017).
Kleesiek, J. et al. Can virtual contrast enhancement in brain MRI replace gadolinium? Invest. Radiol. 54, 653–660 (2019).
Chen, C. et al. Synthesizing MR image contrast enhancement using 3D high-resolution ConvNets. IEEE Trans. Biomed. Eng. 70, 401–412 (2023).
Liu, J. et al. One model to synthesize them all: multi-contrast multi-scale transformer for missing data imputation. IEEE Trans. Med. Imaging. 42, 2577–2591 (2023).
Jiang, L., Mao, Y., Wang, X., Chen, X. & Li, C. CoLa-Diff: conditional latent diffusion model for multi-modal MRI synthesis. In Medical Image Computing and Computer Assisted Intervention – MICCAI 2023 398–408 (Springer, 2023).
Kim, J. & Park, H. Adaptive latent diffusion model for 3D Medical image to image translation: multi-modal magnetic resonance imaging study. In 2024 IEEE/CVF Winter Conference on Applications of Computer Vision (WACV) 7589–7598 (IEEE, 2024).
Wang, X. et al. A deep learning algorithm for automatic detection and classification of acute intracranial hemorrhages in head CT scans. Neuroimage Clin. 32, 102785 (2021).
Ma, D., Pang, J., Gotway, M. B. & Liang, J. A fully open AI foundation model applied to chest radiography. Nature 643, 488–498 (2025).
Xiang, J. et al. A vision-language foundation model for precision oncology. Nature 638, 769–778 (2025).
Lu, M. Y. et al. AI-based pathology predicts origins for cancers of unknown primary. Nature 594, 106–110 (2021).
Isensee, F., Jaeger, P. F., Kohl, S. A., Petersen, J. & Maier-Hein, K. H. nnU-Net: a self-configuring method for deep learning-based biomedical image segmentation. Nat. Methods 18, 203–211 (2021).
Tustison, N. J. et al. N4ITK: improved N3 Bias Correction. IEEE Trans. Med. Imaging 29, 1310–1320 (2010).
Jenkinson, M., Beckmann, C. F., Behrens, T. E. J., Woolrich, M. W. & Smith, S. M. FSL. Neuroimage 62, 782–790 (2012).
Avants, B., Tustison, N. J. & Song, G. Advanced normalization tools: V1.0. The Insight Journal. 2, 1–35 (2009).
Zhao, X. et al. A deep learning model integrating FCNNs and CRFs for brain tumor segmentation. Med. Image Anal. 43, 98–111 (2018).
Nichol, A. Q. & Dhariwal, P. Improved denoising diffusion probabilistic models. In Proc. 38th International Conference on Machine Learning 8162–8171 (PMLR, 2021).
Song, J., Meng, C. & Ermon, S. Denoising diffusion implicit models. In International Conference on Learning Representa tions (ICLR), Vienna, Austria1–21 (Curran Associates, Inc., 2021).
Acknowledgements
This work was partially supported by grants from the National Natural Science Foundation of China (No. 82472050, No. 82172020), Outstanding Young Talents of Guangdong Special Support Program (Health Commission of Guangdong Province) (0820250210), Guangdong Basic and Applied Basic Research Foundation (No. 2023A1515011453), Special Program for Clinical Research of Nanfang Hospital, Southern Medical University (2025CR004), Guangdong Provincial Key Laboratory of Medical Image Processing (No. 2020B1212060039), and National Key Research and Development Program of China (No. 2024YFA1012000).
Author information
Authors and Affiliations
Contributions
K.Z. and Y.Z. made significant intellectual contributions to theoretical development, data analysis, and manuscript drafting. H.S. and R.X. contributed to reviewing and revising the article for intellectual content. X.L. and Y.W. supported data curation and analysis, and backed the argumentation and conclusions. J.M. played a pivotal role in drafting and refining the manuscript. Q.F. reviewed references and ensured compliance with publication standards. L.Z. and W.Y. contributed to theoretical development, system design, and data analysis, and also reviewed the article. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Andra Krauze, Sushant Patkar, and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zheng, K., Zhang, Y., Shu, H. et al. Contrast-free identification of glioma blood-brain barrier status via generative diffusion AI and non-contrast MRI. Nat Commun 17, 2162 (2026). https://doi.org/10.1038/s41467-026-69578-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-69578-8