Abstract
Pathogenic CD4+ memory T cells (Tm) sustain chronic inflammation, but mechanisms remain undefined. Here, we identify four donor-type CD4+ Tm subsets in the target tissues of autoimmune-like chronic graft-versus-host disease in mice: Ly108+CD69− stem-like memory T cells (Tsm), Ly108+CD69+ resident memory progenitor T cells (Trmp), Ly108−CD69+ terminally differentiated tissue-resident T cells (Trm), and Ly108−CD69− intermediate T cells (Tint). Trm are terminally differentiated but not exhausted and show highly biased clonotypes with high proinflammatory cytokine expression. Tsm cells require TCR-MHCII interactions for their maintenance and expansion and show greater capacity than Trmp cells in self-renewal/expansion, generation of Trm, and pathogenicity in adoptive recipients. The transcription factors TCF1/BCL6 and BHLHE40 differentially regulate the stemness and differentiation of Tsm into Trm, respectively, and their selective targeting reduces the number of Trm in tissues and ameliorates inflammation. Thus, our findings indicate that targeting the Tsm subset, involved in the maintenance of the pathogenic Tm pool, offers an attractive approach to treat T cell-mediated chronic inflammation.
Similar content being viewed by others
Introduction
Allogeneic hematopoietic cell transplantation (HCT) is a curative therapy for hematological malignancies and certain non-malignant lymphohematopoietic disorders1,2,3,4, but acute graft-versus-host disease (aGVHD) and chronic GVHD (cGVHD) remain the major obstacles4,5. cGVHD is a systemic lupus-like autoimmune syndrome mediated by donor-type CD4+ T cells in the graft or generated de novo in the thymus damaged by aGVHD, as indicated in publications from our group and others during the past two decades6,7,8,9,10,11,12,13. Although murine model of C57BL/6 donor to MHC-mismatched BALB/c recipient had been considered as a model of aGVHD14, we reported that this model could also develop into cGVHD when aGVHD was not lethal9,13. cGVHD in murine models and in many humans is a sequela of aGVHD2,15. During GVHD pathogenesis, naïve donor CD4+ T cells in the graft are activated by recipient antigen-presenting cells (APC) and differentiate into effector helper T cells to mediate tissue damage16,17,18,19. Some of the donor-type CD4+ T cells that possess cross-reactive TCRs that recognized both donor- and host-type MHC-peptide complexes become autoreactive memory T cells8 and long-term tissue-resident Tm (Trm) cells in the target tissues of cGVHD20, but how those Trm cells are maintained remains unclear.
Stemness potential of CD8+ Tm cells has been reported21,22. CD8+ Tsm are characterized by expression of TCF1, Ly108, and CXCR5, with transcription factors (i.e., TCF1 and BCL6) and other factors (i.e., IFN-I, IL-27, ST2, and Bach2)23,24,25. TCF1 plays a fundamental role in maintaining immune response against infections and cancer by sustaining T cell capacity in proliferation, regeneration, and differentiation23,24,26. The TCF1-BCL6 axis counteracts type I interferon to repress exhaustion and maintain T cell stemness27. CD69 is transiently expressed by early activated T cells but is maintained on tissue-resident T memory effector cells in association with markers such as P2RX7 and CD10328,29,30. Four subsets characterized by expression of Ly108 versus CD69 have been identified in a pathway leading to CD8+ exhausted Tm (Tex) in tissues of chronic infection and tumors. These include progenitor-like Ly108+CD69+ (Texprog1) and Ly108+CD69− (Texprog2), intermediate Ly108−CD69− (Texint), and terminally differentiated Ly108−CD69+ (Texterm) cells31. CD4+TCF1+ T memory progenitors play essential roles in sustaining acute inflammation in allograft rejection and colitis32,33, but their role in sustaining autoimmune-like cGVHD pathogenesis and the mechanisms of their maintenance and differentiation in the tissues remain largely unexplored.
To understand the mechanisms that perpetuate chronic inflammation, in this study, we characterize CD4+ Tm subsets in cGVHD target tissues via combining transcriptomics, epigenomics, flow cytometry analysis, and adoptive transfer experiments. We identify four CD4+ Tm subsets: Ly108+CD69− stem-like T memory (Tsm), Ly108+CD69+ tissue-resident progenitor-like T memory (Trmp), Ly108−CD69+ tissue-resident T memory (Trm), and Ly108−CD69− intermediate T (Tint). Trm cells are the key pathogenic effector cells, and the Tsm are required to maintain enough Trm to mediate tissue inflammation. The Tsm, Trmp, and Trm cells are also observed in the liver tissues of cGVHD patients. Thus, Tsm play a critical role in perpetuating chronic inflammation, and the specific targeting of Tsm cells may represent an attractive approach in treating chronic inflammatory diseases.
Results
Four Tm cell subsets in cGVHD target tissues
BALB/c recipients engrafted with C57BL/6 donor cells developed cGVHD 30–60 days after HCT with infiltration and fibrosis in aGVHD overlapping targets such as the liver and lung and in salivary and lacrimal glands as prototypic cGVHD targets9. cGVHD and controls with no cGVHD were produced by injecting T cell-depleted bone marrow (TCD-BM) cells with or without splenic T cells from C57BL/6 donors. On day 60 after HCT, donor-type CD4+ T cells in the liver were sorted for scRNA-seq analysis. With unsupervised hierarchical clustering, we identified 7 clusters (C0-6) among CD4+ T cells (Fig. 1a). As shown in Fig. 1a, b and Supplementary Fig. 1a, b, C0 contained stem-like memory T cells (Tsm) with high expression of stemness markers25 (i.e., Tcf7, Slamf6, Id3, Lef1) and some migration markers (Cd62l and S1pr1). C1 contained naïve T cells (Tn) with high expression of stemness-related markers and resting markers (i.e., Cd62l, S1pr1 and Klf2). C2 contained intermediate T (Tint) cells34 with low expression of TCR signaling markers (i.e., Batf, Lat and Zap70) and high expression of NK markers [i.e., Klrk1 (NKG2D), Klrc1 (NKG2A) and Klrd1 (CD94)]. C3 contained tissue-resident memory T progenitors (Trmp) with high expression of stemness markers and tissue-residency markers (i.e., Bhlhe40, P2rx7). C4 contained terminally differentiated tissue-resident memory T cells (Trm) with high expression of tissue-resident markers and proinflammatory Ifng/Gzm and low expression of Tcf7. In addition, progenitor signatures were present in C0 Tsm, C1 Tn, and C3 Trmp (Fig. 1c), and strong proinflammatory signaling was present in C3 Trmp and especially in C4 Trm (Fig. 1d). The recently identified tissue-resident T helper cells (Trh)35 were mainly observed in C3 and C4, which had high expression of Pdcd1(PD1), Icos, Maf and Il21 with low expression of Cxcr5 (Supplementary Fig. 1c). C5 and C6 T cells did not express typical markers related to Trm or Tsm and were not considered in the current studies. As compared to No-GVHD recipients, cGVHD recipients had a much lower proportion of C1 Tn, a slightly lower proportion of C0 Tsm, and higher proportions of C2 Tint, C3 Trmp, and C4 Trm (Fig. 1e). These results suggest that Tm subsets in cGVHD target tissues include Tsm, Trmp, Trm, and Tint subsets.
Donor-type CD4+ T cells from livers of No GVHD or cGVHD mice on day 60 after HCT were sorted and subjected to scRNA-Seq (N = 3). a Uniform manifold approximation and projection (UMAP) plots. Workflow: Created in BioRender. (Kong, W. (2026) https://BioRender.com/6hbegtm). b Feature plots show expression of genes related to quiescence, activation, migration, stemness, tissue residency and proinflammation. c Feature plots display T cell progenitor signature. d KEGG pathway of DEGs from each cluster. e Proportions of CD4+ T cell subsets defined by clustering. f Representative panel showed four Tm subsets with TCF1-EGFP (Tcf7-EGFP) versus CD69 in the liver of No GVHD and cGVHD mice at day 60 after HCT. g Comparison of protein expression among four CD4+ Tm subsets identified with Ly108 vs CD69, N = 4–6. h %IFN-γ+ GM-CSF+ cells in each Tm subset after in vitro stimulation with PMA and ionomycin. N = 6. i Percentages and yields of four Tm subsets, N = 5.Data presented as mean ± SEM from two repeated experiments. *P < 0.05, **P < 0.01, ***P < 0.001, as determined by one-way ANOVA (g,h) and two-way ANOVA (i).
Alloreactive memory T cells were identified as CD44hiCD62L− in the tissues more than 30 days after HCT, and the tissue resident Tm cells were identified with expression of CD69 as described in our and others’ previous publications20,32,36. We used flow cytometry analysis of CD69 (tissue-residency marker) versus TCF1 or Ly108 (a surrogate for TCF1 expression)31,37 to validate the donor-type CD62L−CD44+CD4+ Tm subsets in cGVHD target tissues, of which > 95% were derived from the injected CD4+ T cells and < 5% from the de novo-generated CD4+ T cells that contained few Trm cells (Fig. 1f and Supplementary Fig. 1d). The Ly108+CD69− Tsm showed the highest expression of stemness-related markers (TCF1, Ly108). The Ly108+CD69+ Trmp showed lower expression of stemness-related markers but high expression of tissue-residency-related markers and exhaustion markers (CXCR6, PD1, TOX, TIGIT)38. The Ly108−CD69+ Trm and Ly108−CD69− Tint subsets showed low expression of TCF1 and ICOS (Fig. 1g). The Tint subset showed uniquely high expression of the NK-related-marker NKG2D (Fig.1g). Trm cells showed high expression of terminal differentiation and effector markers (i.e., T-bet, CD38 and CD39) with higher percentages of IFN-γ+GM-CSF+ cells that are proinflammatory and pathogenic39 (Fig. 1g, h).
The presence of the Tm subsets in the tissues was validated with an assay of 5-min intravenous aCD4 mAbs to label cells in the circulation40. Most ( ≥ 85%) of Trmp and Trm but only a small portion ( ~ 35%) of Tsm were unlabeled tissue resident cells (Supplementary Fig. 1e). We have reported that CD4+ Tm cells in the cGVHD target tissues liver and lung formed lymphoid-like structures (TLS) consisting of T, B, DC and other cells20,35. We observed the four CD4+Tm subsets in the TLS (Supplementary Fig. 1f). cGVHD recipients had higher percentages of Trmp, Trm and Tint but lower percentages of Tsm as compared with No-GVHD controls, but the yields of all 4 Tm subsets were higher in cGVHD recipients than in controls (Fig. 1i). The four CD4+Tm subsets were also observed in the MHC-haploidentical model of C57BL/6 donor to (C57BL/6 x BALB/c) F1 recipients and MHC-matched model of C57BL/6 donor to C3H.SW recipients (Supplementary Fig. 2a–d). These results suggest that while Ly108/TCF1 expression reflects stemness and CD69, CD39 expression reflects the terminal differentiation of Tm, the terminally differentiated Ly108−CD69+/CD39+ CD4+ Tm subset in the GVHD target tissues is not anergic/exhausted. Instead, they contain highly pathogenic T effectors with high production of IFN-γ and GM-CSF.
Transcriptomic and epigenetic remodeling of the four Tm subsets
We performed combined RNA-Seq with ATAC-Seq to identify the transcriptomic and epigenomic relationships among four CD4+ Tm subsets. Principal-component analysis (PCA) confirmed that each of the four subsets was distinct (Fig. 2a). The sample distance heatmap showed that Tsm had a closer relationship with Trmp than with Trm or Tint, while Trm had a closer relationship with Tint than with other subsets (Fig. 2b). The heatmap and KEGG pathways of differentially expressed genes (DEG) among Tm subsets were grouped into six clusters (Fig. 2c, d and Supplementary Data 1). Tsm and Trmp both had higher expression of Cluster 2 DEGs that regulate DNA metabolic and mitotic cell cycle processes, while Trmp had higher expression of Cluster 4 DEGs that regulate inflammatory immune responses. Trm had higher expression of Cluster 3 DEGs that regulate lymphocyte activation and Cluster 6 DEGs related to defense responses. Tint had higher expression of Cluster 1 DEGs that regulate cell adhesion, Cluster 5 DEGs that regulate cell migration, and Cluster 6 DEGs. Compared to the Trmp subset, the Tsm subset showed higher expression of S1pr1, Nav1, E2f2 and Cdkn3 with lower expression of Cd244a (2B4), Il2, Slc7a8 and Gzma. In comparison to Trm subset, Tint subset showed higher expression of Nav1, S1pr1 and Itgax with lower expression of Il17rb, Deptor, Il9r and Ccl6 (Fig. 2e). These results support the stemness features of Tsm and Trmp and the circulatory potential of Tsm and Tint.
Pooled donor-type CD4+ T cells from the lung at day 60 after HCT were sorted and processed for bulk combined RNA-Seq and ATAC-Seq. N = 3. a PCA plot of the four Tm subsets. b Heatmap of sample distances showing the gene profile similarity between the four Tm subsets. c Heatmap shows all DEGs in the four Tm subsets with clusters of Pearson correlation. d Bubble graph displays the top five enriched KEGG pathways for each cluster from (c). e Volcano plot shows top twenty upregulated or downregulated DEGs annotated in the comparison between Tsm and Trmp and between Tint and Trm cells. f Duplicate tSNE maps show the chromatin accessibility profiles of the four Tm subsets. g Volcano plots comparing specific peaks of transcription factor motif enrichment of Tm subsets: Tsm vs Trmp, Trmp vs Trm, Tsm vs Tint and Trm vs Tint. h ATAC-seq tracks at the loci of Slamf6 (Ly108), Tcf7, Cxcr5, Bhlhe40, Tbx21 (T-bet) and Klrk1 (NKG2D).
ATAC-seq analysis showed differences in chromatin accessibility in distal intergenic regions (31.59%), 1st introns (18.43%), other introns (26.94%), and promoter regions (15.96%) (Supplementary Fig. 3a). The Tsm and Trmp subsets showed high similarity in chromatin accessibility, as did the Trm and Tint subsets (Fig. 2f). Enriched open chromatin regions (OCRs) per Tm subset were analyzed by heatmap (Supplementary Fig. 3b) and opening or closing chromatin regions were compared at the transition stages of Tm cell subsets. More opening and closing chromatin regions were observed during the transition from Trmp to Trm than during the transitions from Tsm to Trmp or from Tint to Trm (Supplementary Fig. 3c). We used HOMER to compare the enrichment of TF binding sequences in chromatin-accessible regions. As compared with Tsm, TF binding motifs CTCF and CTCFL were enriched, but KLF1 was depleted in Trmp (Fig. 2g, the first panel). As compared with Trmp, TF binding motifs Gata and X.box were enriched, but BATF, Nur77 and Tcf7 were depleted in Trm (the second panel). As compared with Tsm, TF binding motifs Gata were enriched, but Tcf7 and BATF were depleted in Tint (the third panel). As compared with Trm, TF binding motif TCFL2 was enriched but BATF and X.box were depleted in Tint (the final panel). These results suggest that stemness-related TF binding sites closed sequentially from Tsm to Trmp and from Trmp to Trm and Tint, while differentiation and function-related TF binding sites opened sequentially across the four Tm subsets in the same order.
Specific TF binding sites were selected for comparison. Consistently, the stemness-related loci Slamf6, Tcf7 and Cxcr5, and exhaustion-related loci Tox and Eomes were more accessible in Tsm and Trmp than in Trm or Tint. The cell migration related loci Klf2 and S1pr1 were more accessible in Tsm and Tint than in Trmp and Trm. The T cell activation-related loci Tbx21 (T-bet), Bhlhe40, Prdm1 (Blimp1), Ifng and Gzmb were more accessible in Trm and Tint than in Tsm or Trmp. The NK cell-related genes loci Klrk1 (NKG2D) and Klrd1(CD94) were more accessible in Tint than in other subsets (Fig. 2h and Supplementary Fig. 3d). Taken together, these results have unraveled the key transcriptomic and epigenetic changes (i.e., Tcf7 and Bhlhe40) during Tsm cell activation and differentiation.
T cell lineage development pathways and clonal expansion among Tm subsets
To unravel the differentiation pathways among the Tm subsets in the cGVHD target tissues, we performed pseudotime single-cell trajectory analysis, using combined scTCR-CDR3-Seq with scRNA-Seq. The results show a predicted T cell differentiation pathway starting from Cluster 0 Tsm to Cluster 3 Trmp, and finally to Cluster 4 IFN-γ+/Gzm+ Trm or Cluster 2 Tint (Fig. 3a). The apparent direct pathway from Cluster 1 Tn to Cluster 0 Tsm represents an artifact reflecting the long delay from the initial alloantigen stimulation of injected donor T cells on day 0 to the analysis on day 60 after HCT. Tm from the livers of No-GVHD and cGVHD recipients showed qualitatively similar differentiation pathways (Fig. 3a). But clonal expansion of TRA and TRB was observed mainly in cGVHD target tissues (Fig. 3b and Supplementary Fig. 4a). Consistent with clonal expansion in cGVHD, the size of clonotypes of cGVHD Tm cells consisted of small, medium, and large, while the Non-GVHD Tm cells were with dominant small size (Supplementary Fig. 4b, c). Tm from the livers of No-GVHD or cGVHD recipients showed little clonal overlap (Fig. 3c). The clonality index of Tsm, Trmp, and Trm were similar and lower than Tint or Tn (Fig. 3d). Trm showed a much lower proliferation or apoptosis than Tsm and Trmp (Supplementary Fig. 4d, e). These results indicate that the clonality of Trm cells in cGVHD target tissues is decided by Tsm cells, and the strongly biased clonal expansion of Trm may result from accumulation of Tsm progeny.
a Single-cell trajectory of the liver CD4+ T cell subsets with pseudotime analysis. b UMAP plot displaying clonal TRB counts of donor-type CD4+ T cells from the liver. c Scatter plot comparing TRB clonotype overlap and expansion. d Clonality index values were analyzed and compared among CD4+T cell subsets from the liver of cGVHD. e UMAP plot shows cluster distribution for six representatives TCR clonotypes. f Bar plots show numbers of twelve expanded TCR clonotypes in different Tm clusters (Black) and total numbers of cells with each clonotype (Blue). g Heatmap shows differentially expressed genes among the top 20 expanded T clonotypes.
Based on combined TRA and TRB, we found top 20 expanded clonotypes with two representatives shown (Supplementary Fig. 4f, g). Expanded clonal types that were shared across Tsm, Trmp, and Trm, with a few Tint clusters, represented ~1800 counts among the 5000 analyzed cells (Fig. 3e, f). The highly expanded clonotypes showed dominant TRBV19 and TRBJ1-5 preferences (Supplementary Fig. 4h), and they were associated with higher expression of proinflammatory genes (i.e., chemokines, granzymes B and K) (Fig. 3g). These results suggest that clonally biased Tsm give rise to clonally expanded proinflammatory Trm cells in the cGVHD tissues.
Alloreactive donor T-derived Tsm differentiate into Tm subsets that mediate cGVHD
To validate the suggestion from pseudotime single-cell trajectory analysis that Tsm could differentiate into Trmp and Trm in the GVHD target tissues, we evaluated the percentages and yields of the four Tm subsets among CD62L−CD44hi Tm cells in the spleen, liver, and lung on days 7, 14, 30, and 60 after HCT. On day 7, ≥80% of the Tm cells in all tissues were Ly108+ Tsm-like (Supplementary Fig. 5a). At 14–30 days after HCT, the kinetics of Tm subsets in the spleen diverged from those in the liver and lung. While the percentage of Tsm-like cells in the spleen remained > 60% from days 7–60, the yields of all subsets decreased to < 5×104 after day 14 (Supplementary Fig. 5a), likely reflecting lymphopenia caused by aGVHD. In the liver and lung, the percentages of Tsm-like cells decreased from days 7 to 30 and remained at 10–20% from 30 to 60 days after HCT, while the percentages of other subsets slowly increased (Supplementary Fig. 5a). The yields of Tsm, Trmp, and Trm cells in the liver and lung increased and generally reached a peak at day 30, coinciding with the onset of clinical cGVHD in our murine models9. The yields of these subsets then remained stable or gradually decreased but persisted at far higher numbers ( > 10–50×104) than those in the spleen (Supplementary Fig. 5a). These results suggest that Ly108+ Tsm cells may give rise to the other 3 subsets in GVHD target tissues over time.
To assess the precursor-product relationships among the Tm subsets from GVHD target tissues, we performed adoptive transfer experiments. As depicted in Fig. 4a and Supplementary Fig. 5b, sorted congenic CD45.1+CD62L−CD44+ Tsm, Trmp, and Trm subsets from the liver and lung of primary cGVHD recipients 30 days after HCT were injected into adoptive recipients with GVHD induced by CD45.2+ donor cells on day 14 after HCT. The adoptively transferred CD45.1+ Tm subsets were analyzed 14 days later. The yields of cells derived from the injected CD45.1+ Tsm in the liver and lung were higher than those derived from Trmp, and those from Trmp were higher than from Trm (Fig. 4b). Tsm and Trmp gave rise to all four subsets, while Trm maintained themselves, and Trmp may have a potential to become Tsm (Fig. 4c). The adoptively transferred Tsm and Trmp cells also expressed high levels of TOX (Fig. 4d), consistent with its role in T cell persistence41. These results indicate that Ly108+ Tsm and Trmp, especially Tsm, have strong progenitor capacity and give rise to Trm in the GVHD target tissues, indicating that Tsm play an essential role in maintaining the Tm pool in the GVHD target tissues.
a Tsm, Trmp and Trm subsets were sorted from the liver and lung of cGVHD mice at day 30 after HCT and adoptively transferred into cGVHD recipients at day 14 after HCT. Adoptively transferred CD45.1+ CD4+ Tm subsets were analyzed 14 days later, as shown in the diagram. b Recovery of each injected Tm subset in the liver and lung 14 days after adoptive transfer, N = 4–8. c Percentages of Tsm, Trmp and Trm among cells derived from each injected Tm subsets are shown and calculated, N = 4–5. d %Tox positive T cells among injected CD45.1+ Tm subsets, N = 4 per group. e CD4+ Tm subsets were sorted from the liver and lung of primary cGVHD mice at day 30 after HCT and adoptively transferred into secondary cGVHD recipients established with Rag1−/−-BM plus 0.1 M CD8+ T cells at day 15 after HCT, %baseline body weight and clinical GVHD score are shown, N = 8. f Representative Masion’ Trichrome panel (magnification, x100, N = 4) and pathology score are displayed. g Yields of Tsm, Trmp and Trm cells in the liver and lung derived from injected CD45.1+ Tm subsets, N = 5 or 6 per group. h %baseline bodyweight and %survival among cGVHD mice treated with anti-CD4 mAbs or control IgG weekly from day 29 to 50 after HCT. N = 7 or 10 per group. i Representative Masion’ Trichrome panel (magnification, x200) and pathology score on day 60 of IgG control and aCD4 mAbs group, ~day75 in the cGVHD recurrence group after HCT, N = 9, 4 or 5 per group. j Yields of Tm subsets in the liver and lungs of cGVHD recipients treated in (h). k %baseline bodyweight and %survival among cGVHD mice treated with aCD4 mAbs or control IgG weekly from day 29 to 99 after HCT. N = 13 or 14. Data presented as mean ± SEM from at least 2 replicates. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001, as determined by one-way ANOVA (b,d,f), unpaired two-tailed t-test (e,h,k), log-rank test comparison of survival (h,k) and two-way ANOVA (g,i,j).
Furthermore, we evaluated the pathogenic capacity of Tsm, Trmp, and Trm subset from GVHD target tissues liver and lung. Before tissue harvesting, the mice were treated with Treg Protector to block NAD-induced death of Trm cells42. The Tm subsets were sorted with purity ≥95% (Supplementary Fig. 5b). To avoid the influence of de novo developed T cells, the adoptive recipients were reconstituted with BM (2.5 × 106 cells) from Rag-1−/− C57BL/6 donors with CD8+ T cells (0.1 × 106) from WT C57BL/6 donors to facilitate complete donor chimerism. 15 days after HCT, sorted Tsm, Trmp, and Trm subsets (0.25×106 each) from primary day 30 recipients were respectively transferred into adoptive recipients, which were monitored for clinical GVHD development for up to day 70 after HCT.
At ~45 days after HCT (30 days after transfer), the adoptive recipients given Tsm or Trmp but not Trm cells started to show bodyweight loss and clinical GVHD manifestations (Fig. 4e). Histopathology scores in Tsm recipients were significantly higher than in Trmp recipients (Fig. 4f), although clinical scores showed no significant difference (Fig. 4e). The histopathology scores in Tsm and Trmp recipients were significantly higher than in Trm recipients (Fig. 4f). The yields of CD4+ Tm cells in the liver and lung were higher in Tsm recipients than in Trmp recipients, while the yields in Trm recipients were very low (Fig. 4g). These results indicate that among Tm subsets, Tsm have the strongest capacity to produce pathogenic Trm cells and cause the most severe cGVHD pathology.
In addition, we tested whether donor-type CD8+ T cells play a role in GVHD via a CD4+ T-dependent mechanism. First, we compared the expansion of donor CD8+ T cells in the adoptive recipients with or without infusion of donor-type CD4+ Tm subset. Adoptive transfer of Tsm but not Trmp or Trm induced significant expansion of the injected CD8+ T cells in the liver and lung (Supplementary Fig. 5c). Second, we tested whether donor CD4+ T cells affected the induction of cGVHD by donor CD8+ T cells. In the absence of donor-type CD4+ T cells, donor CD8+ T cells did not induce cGVHD or show expansion of CD8+ Trm cells (Supplementary Fig. 6a–d). Therefore, cGVHD is primarily maintained by donor-type CD4+ Tm especially Tsm, although donor-type CD8+ T cells may contribute some synergistic effect.
Finally, we evaluated the role of CD4+ Tm cell pool in sustaining cGVHD pathogenesis. The cGVHD mice were treated with depleting aCD4 (clone GK1.5) mAbs. Accordingly, cGVHD mice were treated weekly with i.p. injection of aCD4 mAbs (500 µg/mouse) from days 29 to 50 after HCT. Amelioration of cGVHD was indicated by higher bodyweight and prolonged survival compared to IgG-treated controls (Fig. 4h). At ~14 days after the last aCD4 mAbs treatment, however, the recipients showed rapidly decreasing bodyweight and died with recurrent pathologic evidence of GVHD (Fig. 4h, i). Further analysis showed that aCD4 mAbs treatment could completely deplete CD4+ T cells in the blood and dramatically reduce the pathogenic CD4+ Tm subsets in the cGVHD target tissues, particularly the Tsm subset (Fig. 4j and Supplementary Fig. 6e). Mice with recurrence of cGVHD at day 25 after the last aCD4 mAbs treatment was associated with reappearance of CD4+ Tsm and their progeny in the tissues (Fig. 4j and Supplementary Fig. 6e). Continuous aCD4 mAbs treatment prevented recurrent cGVHD (Fig. 4k). These results indicate that aCD4 mAb-treatment can significantly deplete the four Tm subsets and ameliorate cGVHD. Recurrence of cGVHD after the end of aCD4 mAbs treatment is associated with recovery of Tsm, Trmp, and Trm. Taken collectively, Tsm plays a significant role in maintaining the pool of pathogenic Tm.
Impairment of Tsm formation and amelioration of cGVHD by Tcf1 or Bcl6-deficiency in donor T cells
pySCENIC analysis of the scRNA-seq dataset showed that, among the top 50 regulons, Lef1, Tcf7 (TCF1), Bach2 and Klf3 had high regulon activity in Tn and Tsm, while Tbx21, Nfatc1 and Bhlhe40 had high regulon activity in Trmp, Trm and Tint (Fig. 5a). Therefore, we used Tcf7−/− donor T cells to evaluate the changes of Tsm and their role in generating or maintaining the Tm pool in aGVHD and cGVHD overlapping target tissues liver and lung. We also evaluated histopathological changes in the salivary and lacrimal glands as cGVHD prototypic target tissues9. As compared with donor WT-T cells, Tcf7-deficient donor T cells (Tcf7−/−-T) showed significantly lower cGVHD severity, as judged by higher bodyweight and survival, especially during cGVHD period of 30–60 days after HCT (Fig. 5b), and by histopathology and fibrosis in the target tissues (Fig. 5c and Supplementary Fig. 7a, b). On day 60 after HCT, recipients of Tcf7−/−-T cells showed much lower yields of all four subsets of CD4+ Tm cells in the liver and lung (Fig. 5d) and markedly lower percentages of Tsm and Trmp compared with recipients of WT-T cells, although the percentages of Trm (Fig. 5d) and IFN-γ+GM-CSF+ cells were higher than in recipients of WT-T cells (Supplementary Fig. 7c). These results indicates that Tcf7 deficiency in donor T cells reduces Tsm quantity, leading to reduction of Trm quantity and attenuation of cGVHD.
a pySCENIC analysis of regulon activity among T cell subsets from the liver of BALB/c recipients with cGVHD at day 60 after HCT. b,e C57BL/6 CD45.2+ WT, Tcf7−/− T or Bcl6−/−-T cells were co-injected with CD45.1+ TCD-BM in irradiated BALB/c recipients. N = 10 per groups. %survival and %baseline body weight, respectively, with c,f representative Masion’s trichrome staining panel and pathology scores (N = 5 or 4), and d,g representative contour patterns and percentages/yields of four Tm subsets at day 60 after HCT, N = 4 or 5 per group. h BALB/c cGVHD recipients were established with spleen CD45.2+ WT or Bhlhe40−/− T cells co-injected with CD45.1+ TCD-BM. N = 10 or 20 per group. %survival and %baseline body weight, respectively, with i representative Masion’s trichrome staining panel (magnification, x100, N = 4), and j representative contour patterns and percentages/yields of four Tm subsets at day 60 after HCT, N = 4 or 5 per group. Data presented as mean ± SEM from at least 2 replicates. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001, as determined by unpaired two-tailed T-test and log-rank test for survival (b,e,h) and two-way ANOVA for multiple comparisons (c,d,f,g,i,j).
TCF1 is associated with induction of BCL6 expression, which supports the persistence of CD8+ Tm cells27, and BCL6 also plays an important role in maintenance of CD4+ T cell progenitors during chronic LCMV infection27. Thus, we also compared cGVHD severity and Tm subset changes in recipients given donor WT-T or Bcl6−/−-T cells, and the results show similar patterns observed in recipients given Tcf7−/−-T cells (Fig. 5e–g and Supplementary Fig. 7a, b, d). Taken collectively, these results indicate that TCF1 and BCL6 play important roles in generating or maintaining Tsm that maintain the pool of pathogenic Trm in the cGVHD target tissues.
Impairment of Tsm differentiation into Trm and amelioration of cGVHD by Bhlhe40-deficiency in donor T cells
Although previous studies have shown that formation and maintenance of CD4+ Trm in experimental colitis is regulated by HOBIT43, we observed scant expression of Hobit in Trm from cGVHD target tissues (Supplementary Fig. 1b). Instead, we observed the high regulon activity of Bhlhe40 in Trm (Fig. 5a). Thus, we assessed the role of BHLHE40 and HOBIT in Trm expansion in the cGVHD target tissues, using Bhlhe40−/− or Hobit−/− donor T cells. As compared to recipients given WT-T cells, recipients given Bhlhe40−/−-T cells showed marked reduction of cGVHD severity, as judged by the higher survival rate and bodyweight after HCT (Fig. 5h) and by histopathology and fibrosis on day 60 (Fig. 5i and Supplementary Fig. 7a, b). Compared to WT-T cells, Bhlhe40−/−-T cells showed much lower yields of Tm cells (Fig. 5j) and much lower percentages of Trm, Tint and IFN-γ+GM-CSF+ cells, although higher percentages of Tsm in the liver and lung (Fig. 5j and Supplementary Fig. 7e). In contrast, recipients given Hobit−/− donor T cells did not reduce cGVHD severity or reduce Trm or their production of IFN-γ or GM-CSF (Supplementary Fig. 7f–h). These results indicate that BHLHE40 but not HOBIT in donor T cells plays a critical role in generating or maintaining proinflammatory Trm cells in the cGVHD target tissues.
MHCII-TCR interactions maintain Tsm and augment their differentiation into Trm
Previous studies have shown that donor-type APCs are involved in the pathogenesis of cGVHD18,44. We observed that among Tm subsets in cGVHD target tissues clonal expansion was with Trm but not Tint subsets 60 days after HCT (Fig. 3), suggesting that differentiation and expansion of pathogenic Trm is MHCII-TCR interaction-dependent. Consistent with a previous report44, we observed that donor-type APCs in cGVHD upregulated expression of MHCII (Fig. 6a and Supplementary Fig. 8a). Thus, we evaluated the impact of donor and host MHCII on Tm subset expansion. As compared to results with WT-donor BM → WT-recipients, WT-donor BM → MHCII−/− recipients had modestly reduced severity of cGVHD, while MHCII−/−-donor BM → WT-recipients had markedly reduced severity of cGVHD, as indicated by body weight loss and histopathology (Supplementary Fig. 8b, c). We also explored the MHCII-TCR interaction in regulating the changes of the four Tm subsets in the GVHD target tissues of recipients with MHCII deficiency in donor- or host-type APCs during aGVHD on day 7 and during cGVHD on day 60 after HCT. Flow cytometry analysis of Tm subsets in target tissues liver and lung showed that, at day 7, Tm cells in the tissues were dominantly Tsm cells ( > 60%) with small percentage of Trmp, Trm, and Tint ( < 20%) (Fig. 6b, c and Supplementary Fig. 8d, e). As compared with WT-donor BM → WT-recipients, WT-donor BM → MHCII−/− recipients but not MHCII−/−-donor BM → WT-recipients had lower yields of Tsm although the percentage was relatively increased in the lung and liver (Fig. 6b, c and Supplementary Fig. 8d, e). In contrast, at day 60, Tsm among Tm were reduced to ~20%, and Trm and Tint increased to ~40%. As compared with WT-donor BM → WT-recipients, MHCII−/−-donor BM → WT-recipients but not WT-donor BM → MHCII−/− recipients had lower numbers of Trmp and Trm in both liver and lung tissues, although reduction of Tsm only occurred in the lung (Fig. 6b, c and Supplementary Fig. 8d, e). As compared with WT-donor BM → WT-recipients, the percentages of IFN-γ+GM-CSF+ Trm in the liver and lung of MHCII−/−-donor BM → WT-recipients did not differ, but they were lower in WT-donor BM → MHCII−/− recipients (Fig. 6d). These results indicate that recipient MHCII plays a key role in generating Tsm and their proinflammatory capacity during aGVHD. In contrast, donor MHCII plays a significant role in maintaining Tsm and their differentiation into Trm during cGVHD.
a Donor derived CD11b+ CD11c+ APCs gated from H2Kb+TCRβ− cells from the lung of No GVHD and cGVHD mice were examined at day 60 after HCT, expression of MHCII from APCs was also measured, N = 4. b,c cGVHD mice were established with CD45.2+ WT or MHCII−/− CD45.2+ BALB/c recipients given C57BL/6 CD45.2+ WT-BM or MHCII−/−- BM plus CD45.1+ WT-T cells. Four Tm subsets among injected donor-type (CD45.1+) CD4+ T cells from the lung (b) and liver (c) were examined on days 7 (N = 3 or 4 per group) and 60 after HCT (N = 6, 4 or 6 per group). d Proinflammatory IFN-γ and GM-CSF production among Trm cells was measured on day 60 after HCT (N = 6 per group). e BALB/c recipients were established with WT-T plus WT or IFNγR−/− TCD-BM. %survival and %baseline bodyweight. N = 10–15. f Representative Masion’ trichrome staining panel and pathology scores are shown (magnification, x200, N = 4). g Representative contour plots, percentages and yields of the four Tm subsets were analyzed on day 60 after HCT, N = 4 ~ 6. h Proinflammatory IFN-γ and GM-CSF production among CD4+ Tm cells, N = 4. i Percentages of CD11b+ CD11c+ APCs were also compared, and MHCII expression was analyzed, N = 4 ~ 6. Data presented as mean ± SEM from at least 2 replicates. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001, as determined by two-way ANOVA (a,b,f,g), one-way ANOVA (d), log-rank test comparison of survival (e) and unpaired two-tailed t-test (e,h,i).
In addition, we compared the Tm subsets derived from the injected and de novo generated donor-type CD4+ T cells in WT recipients given WT or MHCII−/− BM. As compared to the injected CD4+ Tm, the de novo developed CD4+ Tm showed much lower Trm percentages and lower percentages of IFN-γ+GM-CSF+ Trm, but higher percentage of Tsm and Trmp (Supplementary Fig. 8f, g). As compared to de novo-generated CD4+ Tm from WT-BM, the de novo-generated CD4+ Tm from MHCII−/− BM also showed lower percentages of Trm and IFN-γ+GM-CSF+ Trm (Supplementary Fig. 8f, g). These results indicate that Tsm derived from the injected CD4+ T cells but not de novo-generated CD4+ T cells play a critical role in supporting the pathogenic Trm pool.
IFN-γ induces MHCII expression by antigen-presenting cells and other cells45, but the impact of IFN-γ on regulation of tissue Tm subsets in cGVHD remains unclear. Recipients given IFN-γR−/− donor BM cells plus WT-T cells showed lower severity of cGVHD as judged with higher bodyweight and percentage of survival and lower pathology score and fibrosis on day 60 after HCT (Fig. 6e, f), which was associated with lower percentages of donor-type CD11c+CD11b+ DCs as well as associated with lower yields of Tsm, Trmp, and Trm in the lungs and liver but not reduction in the percentages of IFN-γ+GM-CSF+ Trm cells (Fig. 6g–i). These results indicate that IFN-γR signaling regulates expansion of MHCII-expressing donor DCs that maintains Tsm and augments their differentiation into pathogenic Trm cells to sustain cGVHD.
Tsm, Trmp, and Trm are present in the lymphoid-like structures in the liver tissues of cGVHD patients
To determine the clinical and translational implications of our experimental results, we investigated whether Tsm, Trmp, Trm could be identified in the blood and affected tissues from patients with cGVHD (Supplementary Table 1 and 2). We first analyzed CD4+ Tm subsets with TCF1 and T-bet in the blood of patients with or without cGVHD. CD45RA−FAS+ T cells are identified as Tm in human blood46. Among CD4+ Tm, the percentages and numbers of TCF1hiT-betlo stem-like memory T (Tsm) were lower and the percentages and numbers of TCF1loT-bethi Teff cells in the blood were higher in patients with cGVHD than in those without cGVHD (Fig. 7a). The TCF1hiT-betlo Tsm subset express stemness-related markers22 with high levels of CD27, TCF1 and CXCR5 (Fig. 7b, c), while the TCF1loT-bethi Teff subset express high levels of CD11b, CXCR3 and T-bet (Fig. 7b, c). These results show that cGVHD patients have high numbers of T-bethi Teff and lower numbers of TCF1hi Tsm cells in the blood when compared to controls without cGVHD.
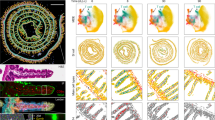
a Representative patterns, percentages and numbers of TCF1+ T-betlo Tsm and TCF1lo T-bethi Teff cells from the PBL of allo-HCT patients with No-GVHD and active cGVHD are shown and compared. N = 7. b Representative histograms show protein expression with (c) heatmap displaying relative expression among Tm subsets. d Tertiary lymphoid-like structures are shown with CD3, CD45RA, CD68, CD20, Collagen I and CD11c. e Merged images show staining of CD4, CD3, CD45RA, Foxp3 and DNA and f staining of IFN-γ, GM-CSF and CD4, and percentages of IFN-γ and GM-CSF producing CD4+ Tm cells and Treg cells were measured. g Merged image shows staining of CD69, Ly108 and CD4 in the liver of patients with active cGVHD. h Representative images of CCR7− CD69+ CD4+ CD3+ T cells of liver sections from cGVHD patients. i Representative images of TCF1 and CD69 expression among CD4+ CD45RA− cells of liver sections from cGVHD patients, N = 4. Data represented as means ± SEMs. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001, as determined by two-way ANOVA (a), one-way ANOVA (g,i) and unpaired two-tailed t-test (h).
We used imaging mass cytometry (IMC)47 to visualize Tm subsets in the liver biopsy specimens from patients with cGVHD (Supplementary Table 2). Liver tissues of cGVHD patients were stained with a 24-antibody panel (Supplementary Fig. 9a, Supplementary Table 3). With the UMAP algorithm and mapping, we found tertiary lymphoid-like structures (TLS) that consisted of Tm, dendritic cells, macrophages and B cells (Fig. 7d and Supplementary Fig. 9a-c). We analyzed 2 TLS regions from each of four patients (Fig. 7d and Supplementary Fig. 9c). CD4+ Tm cells in the liver of cGVHD patients had few Foxp3+ but many IFN-γ+GM-CSF+ cells (Fig. 7e, f). The Tm cells included Ly108+CD69− Tsm, Ly108+CD69+ Trmp, and Ly108−CD69+ Trm subsets, and the number of Tsm per field was significantly lower than the number of Trmp, or Trm (Fig. 7g and Supplementary Fig. 7d). We used immunofluorescent staining to validate the identification of TCF+CD69− Tsm, TCF1+CD69+ Trmp, and TCF1−CD69+ Trm cells among CCR7−CD45RA−CD4+CD3+ cells (Fig. 7h, i). These results indicate the presence of Tm subsets including Tsm, Trmp and proinflammatory Trm in lymphoid-like structures in the liver of patients contributing active cGVHD.
Discussion
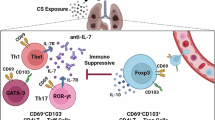
In the current studies, we showed that, as depicted in the diagram (Fig. 8), CD4+ Tm pool in cGVHD target tissues consists of Tsm, Trmp, and proinflammatory Trm subsets, and Tsm show greater capacity than Trmp in self-renewal/expansion and differentiation into Trm cells that mediate tissue damage. The generation and maintenance of Tsm and Tsm differentiation to Trm is regulated by TCR interaction with MHCII and by TCF1/BCL6 and BHLHE40. In addition, the Tsm, Trmp and Trm subsets contained in lymphoid-like structures were also observed in the tissues of patients with active cGVHD. These studies offer novel insights into how TCF1+CD4+ Tm progenitors and CD69+ Trm proinflammatory effectors in the tissues work together to sustain autoimmune-like cGVHD.
Based on distinct transcriptional and epigenomic features and flow cytometry of Ly108 versus CD69, donor graft CD4+ T-derived memory T cells (Tm) in the chronic GVHD target tissues can be divided into four subsets: Ly108+CD69− Tsm, Ly108+CD69+ Trmp, Ly108−CD69+ Trm, and Ly108−CD69− Tint. Trm are the critical effectors that mediate cGVHD, and Tsm are required for maintaining the Tm pool. Tsm differentiation into Trmp and then to Trm in a MHCII/TCR-dependent manner and produces biased clonal expansion of Trm cells with strong proinflammatory characteristics. TCF1/BCL6 and BHLHE40 differentially regulate the stemness and differentiation of Tsm cells. Solid lines with arrow mean strong indication and dashed lines with arrow means potential indication. Created in BioRender. Kong, W. (2026) https://BioRender.com/2extxe5.
A recent publication by Sacirbegovic et al showed that a tissue-resident TCF-1+ Tm subpopulation from acute GVHD tissue preferentially engrafted, expanded, and differentiated into effectors upon adoptive transfer36, but the subsets among TCF1+ Tm progenitors and the regulation of the progenitor differentiation remained unclear. Our current studies unraveled the characteristic features of four CD4+ Tm subsets during cGVHD: 1) Ly108+CD69− Tsm-like cells have strong repopulation and pathogenic capacity. 2) Ly108+CD69+ Trmp-like subset also has repopulation and pathogenic capacity but much lower than observed with Tsm. The Trmp express strong TCR signaling related markers, and some also express exhaustion-related markers (i.e., PD-1, TOX). 3) Ly108−CD69+ cells express elevated levels of terminally differentiated tissue-resident T markers and proinflammatory and cytokines (i.e., CD69, CD39, IFN-γ and GM-CSF), with low expression of anergy/exhaustion protein markers consistent with features of potent proinflammatory T effector cells39. Pathogenic effects depend on the numbers of Ly108−CD69+CD4+ Trm that require in situ replenishment by Tsm or Trmp. 4) Ly108−CD69− Tint cells show high expression of some NK features with low expression of TCR signaling markers. Tint cells have overlapping features of bystander T cells with little TCR signaling strength/exhaustion, poor clonal expansion and enhanced NKG2D expression48, although they have a role in anti-tumor immunity34, their role in cGVHD pathogenesis remains largely unknown.
Beltra JC et al31 used Ly108 vs CD69 to divide exhausted CD8+ T cells in the tumor and chronic infection tissues into four subsets including Ly108+CD69+, Ly108+CD69−, Ly108−CD69−, and Ly108−CD69+. They propose that the 108+CD69+ subset is the quiescent Texprog1 that give rise to 108+CD69− proliferative circulating Texprog2, Ly108−CD69− circulating Texint, and Ly108−CD69+ terminally exhausted Texterm subsets; and Texprog1 and Texprog2 can be interconverted. Adopting the Ly108 versus CD69 analysis, we also divided CD4+ Tm cells in cGVHD target tissues into four subsets, including Ly108+CD69−, Ly108+CD69+, Ly108−CD69+, and Ly108−CD69−. Using combined scRNA-Seq with ATAC-Seq, and flow cytometry analysis, we characterized Ly108+CD69− as Tsm, Ly108+CD69+ as Trmp, Ly108−CD69+ as Trm, and Ly108−CD69− as Tint. Based on the results of pseudotime single-cell trajectory analysis, kinetic analysis of the four subsets in the lymphoid and GVHD target tissues, and Tm subset transfer experiments, we propose that alloreactive Tm cells that have stem cell potential first give rise to Ly108+CD69− Tsm-like cells, and the Tsm then give rise to Trmp, Trm, and Tint. Tsm and Trmp can be partly interconverted, and Tsm and Trmp differentiation into Trm but not Tint depends on TCR/MHCII interaction. The Ly108 and CD69 differentiation pathways differ in some respects between CD4+ Tm cells in cGVHD target tissues and CD8+ Tm cells in tumor and chronic infection tissues. For example, the terminally differentiated Ly108−CD69+CD4+ Trm in GVHD target tissues are highly pathogenic and not anergic/exhausted, while the terminally differentiated Ly108−CD69+CD8+ Trm in tumor/chronic infection tissues are terminally exhausted31. Details in the stepwise differentiation among Tsm, Trmp, and Trm of CD4+ Tm subsets and the environmental cues that regulate the process remain to be elucidated.
In addition, the differentiation path of Tsm→Trmp→Trm was indicated by pseudotime single-cell trajectory analysis using combined scTCR-CDR3-Seq with scRNA-Seq, kinetic study of Tm subsets in the cGVHD target tissues, and transfer of sorted Tsm, Trmp, or Trm. Although the epigenetic analysis supported that Tsm and Trmp are more stem-like and Trm are more differentiated, the genes regulate the transitions from Tsm to Trmp to Trm remain unclear. Our data has not ruled out the possibility that Tsm can directly differentiation into Trm or that Trmp are not the direct progeny of Tsm; instead, they are established independently and maintained separately. Future studies are needed to address those questions.
BHLHE40 could serve as key differentiation checkpoint during CD8+ T cell differentiation as well as regulation of tissue-residency program49,50, while TCF-1 is required for CD4+ T cell persistence functions during alloimmunity51. We showed that Ly108+/TCF1+CD69− Tsm and Ly108+ /TCF1+CD69+ Trmp in the GVHD tissues are required to sustain cGVHD, because reduction of Tsm/Trmp generation by Tcf7/Bcl6-deficiency and reduction of Tsm differentiation into Trm cells by Bhlhe40-deficiency in donor T cells both markedly reduced the numbers of proinflammatory IFN-γ+GM-CSF+ Trm cells and prevented cGVHD. These observations are consistent with the reports that TCF1/BCL6 plays an important role in maintaining T cell stemness and BHLBH40 in regulating Tm differentiation49,50,51. We used Tcf7−/−, Bcl6−/−, or Bhlhe40−/− donor T cells to set up GVHD models, and amelioration of cGVHD by T cell deficiency in Tcf7, Bcl6, or Bhlhe40 could result from reduction in aGVHD. To address this question, we used depleting anti-CD4 mAb to treat the recipients after cGVHD was established and found that administration of depleting anti-CD4 that effectively depleted Tsm and their progeny ameliorated cGVHD. Relapse of cGVHD after stopping anti-CD4 treatment was associated with rapid rebound of Tsm and their progeny. In addition, we transferred sorted Tsm, Trmp, and Trm cells from the liver and lung of early cGVHD recipients into non-GVHD adoptive recipients and found that Tsm and Trmp but not Trm cells induced chronic inflammation with fibrosis in the liver and lung of the recipients. Therefore, while Trm cells are the pathogenic effectors, Tsm cells play a critical role in replenishing the Trm pool to sustain cGVHD pathogenesis.
Maintenance and differentiation of CD4+ Tsm cells to Trm requires TCR-MHCII interactions. Activation and differentiation of naive alloreactive donor CD4+ T cells require interaction with host-type MHCII early after HCT to trigger aGVHD16,19,52. We observed that maintenance and differentiation of Tsm into Trm cells in the target tissues during cGVHD required donor-type MHCII. This provides new mechanisms to previous observations that donor-type APCs and autoreactive-like donor-type CD4+ T cells mediated cGVHD pathogenesis44,53. We also observed that host MHCII was required to generate Tsm that differentiate into Trm cells with high level production of proinflammatory IFN-γ and GM-CSF cytokines, this may provide explanation to previous observations that MHC-mismatched HCT is associated with higher severity not only in aGVHD but also in cGVHD44. Although host parenchymal MHCII could mediate GVHD19, our studies with MHCII−/− or IFN-γR−/− donor BM showed that donor MHCII expressed by hematopoietic-derived APCs play an important role in maintaining Tsm and mediating their differentiation into Trm that sustain cGVHD.
Overall, our current studies have identified four distinct CD4+ Tm subsets including Tsm, Trmp, Trm, and Tint in the tissues of chronic inflammation such as cGVHD. We have also unraveled key differentiation factors that regulate maintaining and differentiation of Tsm cells in the tissues, including MHCII-TCR signaling, TCF1/BCL6, and BHLHE40. These studies indicate that Tsm may serve as the key therapeutic target for T cell-dependent chronic inflammation.
Methods
Study design
This study is designed to dissect the mechanisms about how the pathogenic CD4+ Trm pool is maintained to sustain the pathogenesis of cGVHD. First, with murine models of cGVHD, we applied flow cytometry sorting, combined single-cell RNA-Seq with TCR-CDR3-Seq, combined bulk RNA-Seq with ATAC-Seq, and in vivo transfer experiments to analyze, characterize, and validate the Tm subsets (i.e., Tsm) in cGVHD target tissues. Second, we used specific gene-deficient donor T cells to study the key that regulating the maintenance and differentiation of Tsm cells in the cGVHD target tissues. Finally, we attempt to link our observations with murine models to the pathogenesis of cGVHD in patients by using imaging mass cytometry (IMC) to visualize the Tsm cell subsets in tertiary lymphoid structures in the liver tissues of patients with active cGVHD.
Human samples
White blood cells were isolated and cryopreserved over in liquid nitrogen. Cells were thawed and T cells subsets were examined by flow cytometry. Patient tissue samples were also collected and analyzed with imaging mass cytometry or immunofluorescence staining to visualize regions of immune cell infiltration. City of Hope National Medical Center Institutional Review Board (IRB15172) and the Fred Hutchinson Cancer Center approved the study. All recruited volunteers provided written informed consent. The patients gave informed consent allowing sections from biopsies to be used for research if the material was no longer needed for clinical purposes or for any required retention.
Mice
BALB/c (NCI#555, H-2Kd), C57BL/6 (NCI#556, H-2Kb), BDF-1(NCI#99, H-2Kb/d) mice were purchased from the National Cancer Institute. Congenic CD45.1+(JAX# 002014), MHCII−/−(JAX# 003584), Nur77-GFP (JAX# 016617),CD4-cre (JAX# 017336), CD4−/−(JAX# 002663), Rag1−/−(JAX# 002216), IFNgR−/−(JAX# 003288), Tcf7f/f (JAX# 030909) or Bhlhe40−/−(JAX# 029732) C57BL/6 mice were purchased from the Jackson Laboratory (JAX). Hobit−/− C57BL/6 mice were kindly provided by Timothy O’Sullivan at UCLA, Bcl6f/f C57BL/6 mice were obtained from Dr Markus Muschen12,54. CD4-cre/Bcl6f/f or CD4-cre/Tcf7f/f C57BL/6, MHCII−/− BALB/c and C3H.SW (JAX#000438) mice were bred at City of Hope Animal Research Center (COH-ARC). CD4-cre/Bcl6f/f or CD4-cre/Tcf7f/f C57BL/6 were generated by crossing CD4-cre mice with Bcl6f/f or Tcf7f/f mice, respectively. MHCII−/− BALB/c mice were generated by backcrossing MHCII−/− C57BL/6 mice with BALB/c mice for more than 10 generations. All mice are maintained in a specific-pathogen-free room at COH-ARC. The experimental and control mice were housed in separate cages. The donor and host mice were generally male and 8-12 weeks old. The City of Hope Institutional Animal Care and Use Committee (IACUC) have approved all animal protocols.
Induction and evaluation of chronic GVHD
BALB/c cGVHD recipients (8 ~ 12 weeks old male, H2d) were exposed to 850 cGy total body irradiation (TBI) using a [137Cs] source 8–10 h prior to HCT and then received an intravenously (i.v.) injection of CD45.1+ or CD45.2+ C57BL/6 (8 ~ 12 weeks old male, H2b) donor spleen cells (1×106) or enriched Thy1.2+ spleen T cells (0.2 ×106) and CD45.2+ or CD45.1+ T cell-depleted BM cells (TCD-BM, 2.5 ×106). TCD-BM cells were depleted of T cells by incubation with biotin-conjugated anti-CD4 and anti-CD8 mAbs and anti-biotin Microbeads (Miltenyi Biotec# 130-097-046) and then pass through an autoMACS Pro cell sorter (Miltenyi Biotec, Germany). Enrichment of splenic Thy1.2+ T cells was achieved using mouse anti-CD90.2 microbeads (Miltenyi Biotec# 130-121-278). The purity of the Thy1.2+ T cells enrichment was > 98%, while the purity after T cell depletion was > 99%. The experimental and control animals were bred separately, evaluation and scoring of clinical chronic signs of GVHD and clinical cutaneous GVHD have been previously described12,55.
To establish MHC-haploidentical and minor mismatched cGVHD mouse models, B6D2F1 and C3H.sw recipient mice were irradiated with 1100 cGy and then engrafted with 5 M TCD-BM alone (No GVHD) or with 8 ×106 splenocytes added (cGVHD).
Isolation of mononuclear cells from lymph and non-lymph tissues
Mice were euthanized by CO₂ inhalation according to institutional guidelines and tissues were collected immediately after euthanasia. The spleens were meshed through a 70 μm nylon cell strainer (VWR, 10199-657) and cells were collected. Bone marrow cells were flushed from femurs with PBS and meshed through a 70 μm nylon cell strainer to collect the cells. Liver tissues were flushed with PBS and meshed through a 70 μm strainer and cells were collected and washed with PBS containing 2% BSA. Hepatic mononuclear cells were further enriched with a 40% and 70% Percoll gradient (Sigma-Aldrich# P1644). For isolation of MNC from lung and skin, tissues were cut into pieces and digested with collagenase VIII (Sigma-Aldrich# C2139) and DNase I (Fisher Scientific# NC0947151) in RPMI-1640(Fisher Scientific #21870092)/10%FBS (Fisher Scientific#A2720803) complete medium at 37 °C for 45 and 90 min, respectively. MNC were then isolated with Lympholyte-M (Cedarlane# CL5035). Mouse PBMCs were also enriched with mouse Lympholyte-M. Isolated cells were washed and counted with a Cellometer Auto 2000 Cell Viability Counter (Nextcelom).
Flow cytometry and antibody usage
Isolated mononuclear cells were used to measure surface antigens, transcription factors and intracellular cytokines. For surface antigen measurement, mononuclear cells were blocked with anti-CD16/CD32 mAbs for 15 min and cells were stained with a surface antibody mixture at 4˚C for 15 min. Cells were then washed and evaluated with the use of BD fortessa or Cytek Aurora instrument. For transcription factors analysis, mononuclear cells were stained with surface antigens and then fixed for 30 min with commercial Foxp3/Transcription Factor Staining Buffer Set (Fisher Scientific# 552300). Cells were then washed with permeabilization buffer (Fisher Scientific# 833356) and stained with an intracellular TF antibody mixture. For analysis of intracellular cytokines, isolated mononuclear cells were stimulated for 4 h with PMA (Fisher Scientific# 125491) plus ionomycin (Sigma Aldrich# I0634), and protein transport inhibitors (Fisher Scientific# BDB554724), and cells were then stained with a surface antibody mixture. Cells were later fixed with a commercial fixation and permeabilization kit (Fisher Scientific# 554722) and stained with a mixture of antibodies against intracellular cytokines. For BrDU staining, BrDU (Fisher Scientific#550891) was i.p. injected at 500 μg/mouse, and tissues were harvested 3 h later. BrdU measurement was applied with BD APC BrdU Flow Kit (Fisher Scientific# 552598). And Annexin V expressions were determined with Annexin V apoptosis detection kit (Biolegend# 88-8007-72).
The antibodies used to stain murine cells included anti-CD4 (RM4-5), anti-CD4 (GK1.5), anti-CD3 (17A2),anti-CD8 (53-6.7), anti-TCRβ (H57-597), anti-H-2Kb (AF6-88.5), anti-CD45.1 (A20), anti-CD45.2(#104), anti-CD62L (MEL-14), anti-CD69 (H1.2F3), anti-PD1 (29F.1A12), anti-CD44 (IM7), anti-Ly108 (13-G3), anti-CXCR6 (SA051D1), anti-CD39 (Duha59), anti-CD38 (#90), anti-TIGIT (Vstm3), anti-Fas (15A7), anti-TCF1 (C63D9), anti-T-bet (eBio4B10 (4B10)), anti-TOX (REA473), anti-ICOS (7E.17G9), anti-NKG2D (CX5),anti-B220 (RA3-6B2), anti-CD11b (M1/70), anti-CD11c (N418), anti-MHCII (M5/114.15.2), anti-CD45 (30-F11), anti-GM-CSF (MP1-22E9), anti-IFN-γ (XMG1.2), more details see Supplementary Table 4.
The antibodies used to stain human PBMCs included anti-human CD3/CD4/CD8a (Antibodies cocktail, Thermo Fisher Scientific), anti-human CD95 (DX2), anti-human CD45RA (HI100), anti-human CD29 (TS2/16), anti-human CXCR5 (MU5UBEE), anti-human CCR7 (4B12), anti-human CD27 (O323), anti-human CD56 (NCAM),anti-human CXCR3 (G025H7), anti-human CD11b (M1/70), anti-human CD197 (3D12), anti-human T-bet (4B10) and anti-human TCF1 (C63D9).
Intravascular labeling with anti-CD4 mAbs
Mice were intravenously injected with 2 µg aCD4 mAbs (Clone: RM-4-5) in 200 µl sterile PBS for 5 min and then euthanized to harvest tissues. Mononuclear cells were isolated from tissues for flow cytometry analysis. Lung circulating CD4+T cells are i.v. anti-CD4 (Clone: RM-4-5) positive, and lung tissue localized immune cells (Trm) are defined by i.v. anti-CD4 (Clone: RM-4-5) negative among H2Kb+TCRβ+CD4+ (Clone: GK1.5).
Immunofluorescent Staining
Mouse liver cryosections were fixed with 4% PFA, blocked by SuperBlock T20 (PBS) Blocking Buffer (Thermo Scientific) for 30 min and incubated overnight at 4 °C in primary antibody cocktails including rat anti-mouse CD4 (biolegend, clone RM4-5), rabbit anti-mouse Ly108 (Cell signaling, clone E2P7A,), goat anti-mouse CD69 (R&D, clone: Polyclonal). To measure CD4+ Tm cells infiltration in the liver of cGVHD patients, slides were stained with rabbit anti-hCD3 mAbs (Fluidigm :3170019D), Alexa Flour 488 conjugated anti-hCD4 mAbs (R&D, FAB8165G), mouse anti-hCCR7 (R&D, Clone # 150503), goat anti-hCD69 mAbs (R&D, AF2359) and DAPI or Alexa Flour 488 conjugated anti-hCD4 mAbs (R&D, FAB8165G), mouse anti-hCD45RA mAbs (Fluidigm,Clone:HI100), goat anti-hCD69 mAbs (R&D, AF2359), rabbit anti-TCF1 mAbs (Cell Signaling Technology, C63D9) and DAPI. Slides were washed with PBS-TB (PBS supplemented with 0.05% Tween 20 and 1%BSA) three times and stained with secondary antibody for 60 min at room temperature. Images were later acquired on a Confocal Zeiss LSM700 or LSM880 microscope at 200X or 400X magnification.
Histopathology
Liver, lung, salivary gland and lacrimal gland from No GVHD or cGVHD mice were harvested and fixed in 10% neutral formalin and processed in paraffin blocks. The tissue-embedded paraffin blocks were sectioned at 5 μm and sections were stained with haematoxylin and eosin (HE) and Masson’s trichrome by the Pathology Solid Tumor Core at the City of Hope. Slides were examined at 100x or 200x magnification and visualized with Zeiss Observer II at the City of Hope Light Microscopy Core.
Cell sorting
For adoptive transfer experiments, primary cGVHD recipients on day 30 after HCT were i.p. injected with 50ug Treg-protector (anti-ARTC2 nanobody, Biolegend) 30 min before euthanasia. Mononuclear cells from the liver and lung were mixed, and total CD4+ cells were enriched with anti-mouse CD4 beads (Miltenyi). Enriched cells were then stained with anti-H2kb mAb, anti-CD4 mAb, anti-TCRβ mAb, anti-CD45.1 mAb, anti-CD44 mAb, anti-CD62L mAb, anti-CD69 mAb and anti-Ly108 mAb cocktail. Four Tm cell subsets including CD69−Ly108+, CD69+Ly108+, CD69+Ly108− among injected H2kb+ CD45.1+ CD44+ CD62L−TCRβ+CD4+ T cells were sorted for adoptive transfer.
For scRNA-seq/scTCR-seq, cGVHD mice and No GVHD mice were euthanized on day 60 after HCT, mononuclear cells were isolated from the liver tissues and stained with a surface antibody mixture: anti-H2kb mAb, anti-CD4 mAb and anti-TCRβ mAb. Sorted H2Kb+ TCRβ+ CD4+ T cells were used for single cell sequencing.
For bulk RNA sequencing and ATAC sequencing, the liver tissues from cGVHD mice were harvested on day 60 after HCT. Isolated mononuclear cells were stained with anti-H2kb, anti-CD4, anti-TCRβ, anti-CD44, anti-CD62L, anti-CD69 and anti-Ly108. Four Tm cell subsets including CD69−Ly108+, CD69+Ly108+, CD69+Ly108−, CD69−Ly108− among injected CD45.1+ H2kb+ CD44+ CD62L−TCRβ+CD4+ T cells were sorted for Bulk RNA-Seq and ATAC-Seq.
Adoptive Transfer Experiments
Chronic GVHD was induced by transplanting CD45.2+ (or CD45.1+) TCD-BM cells (2.5×106) and splenocytes (1×106) from CD45.1+(or CD45.2+)-congenic C57BL/6 donors into lethally irradiated BALB/c recipients. 30 days after HCT, mononuclear cells from the liver and lung were isolated and stained with anti-H2Kb mAb, anti-CD45.1 mAb, anti-TCR mAb, anti-CD4 mAb, anti-CD69 mAb, anti-Ly108 mAb, anti-CD62L mAb, anti-CD44 mAb and then sorted by ARIA SORP. One million sorted Tsm, Trmp and Trm cells were transferred into mild GVHD adoptive recipients at 14 days after HCT with CD45.2+ C57BL/6 donor spleen cells (0.5 ×106) and T cell-depleted BM (TCD-BM) cells (2.5 ×106). 14 days after cell transfer, MNCs from the liver and lung were analyzed for percentage and yield of CD4+ T cells and four subsets of CD4+ Tm subsets.
To evaluate CD4+ Tm cells in regulation of cGVHD in secondary recipients, 0.25 ×106 Tsm, Trmp and Trm cells were adoptively transferred into BALB/c GVHD mice at day 15 after engraftment with 2.5 ×106 Rag1−/− BM and 0.1 ×106 CD8+ T cells from B6 donors.
Anti-CD4 mAbs treatment of cGVHD mice
cGVHD mice were treated with intraperitoneal (i.p.) anti-mouse CD4 mAbs (Bio X Cell, Clone GK1.5) weekly at 500 µg/mouse from day 29 to 50 after HCT or from day 29 to 99 after HCT. Control groups were given Rat IgG mAb purchased from Jackson ImmunoResearch Laboratories Inc.
10× Genomics single-cell RNA/TCR library preparation and sequencing
scRNA-seq library was generated using the Chromium Single Cell V(D)J Reagent Kits (10× Genomics) according to the manufacturer’s instructions and targeting 5,000 cells, and scTCR-seq library was generated using Chromium Single Cell V(D)J Enrichment Kit, mouse T and B Cell (10× Genomics), following all steps according to the manufacturer’s protocols. The cDNA library was sequenced on illumina HiSeq 2500 sequencing System platform at 26 + 8 + 101 (Read 1: 26 cycles, i7 Index: 8 cycles, Read 2: 101 cycles; Sequencing depth: 20,000/ read pairs per cell) for gene expression, and at 151 + 8 + 151 (Read 1: 151 cycles, i7 Index: 8 cycles, Read 2: 151 cycles; Sequencing depth: 5000/ read pairs per cell) for BCR and TCR sequencing. Sequencing reads were processed using the CellRanger pipeline with the default settings (10x Genomics).
scRNA/TCR-sequencing analysis
Analysis of mouse single cell RNA-seq data was performed using “Seurat”56. The datasets were subjected to quality control (elimination of cells with RNA features < 200 and mitochondrial RNA > 10%) and SCT normalization retaining the top 2000 variable genes, followed by integration using the Seurat workflow. The integrated data was scaled and subjected to principal component analysis (PCA), following by clustering using the top 12 PCs. UMAP was used to perform the dimensional reduction for visualization. Genes with differential expressions were identified by using the FindMarkers function in Seurat. Psuedo-time analysis was performed using the “Monocle3” package57. T cell stemness/progenitor signature were score via “AddModuleScore” with published genesets58. Single cell TCR-CDR3 data were processed by the 10x CellRanger pipeline, and cell barcodes were matched to the single cell RNA-seq data using R scripts. Visualization of single cell RNA-seq and TCR-CDR3-seq data was performed using “Seurat” and “scRepertoire”59. To explore key transcription factors in each cluster, 6000 cells were randomly selected, and active transcription factors were identified with the pySCENIC package60.
Bulk RNA-sequencing library preparation, sequencing and data analysis
RNA concentration was measured by NanoDrop 1000 (Thermo Fisher Scientific, Waltham Massachusetts, US), and RNA integrity was determined using Bioanalyzer (Agilent). Library construction of 280 ng total RNA for each sample was made using KAPA mRNA HyperPrep Kit (Illumina Platforms) (Kapa Biosystems, Wilmington, USA) using 10 cycles of PCR amplification. Libraries were purified using AxyPrep Mag PCR Clean-up kit (Thermo Fisher Scientific). Each library was quantified using a Qubit fluorometer (Life Technologies), and the size distribution was assessed by using the 2100 Bioanalyzer (Agilent Technologies, Santa Clara, USA). Sequencing was performed on NovaSeq 6000 Sequencing System (Illumina, San Diego, CA, USA) instrument using NovaSeq Reagent Kits to generate 101 bp Paired-end reads sequencing. Quality control of RNA-Seq reads was performed using FastQC.
For data analysis, raw RNA-seq sequences were subjected to adapter trimmed with Trimmomatic, and poly(A) tails were removed with FASTP. The trimmed reads were aligned to mouse genome mm10 using STAR with the default settings. The expression level of RefSeq genes were counted using “Rsubread”. The raw count data were normalized by trimmed mean of M-values (TMM) method using the Bioconductor package “edgeR”. Differential expression analysis was conducted using the quasi-likelihood (QL) F-test implemented in “edgeR” to identify differentially expressed genes, with the cutoff of at least two groups having average RPKM >= 1, FDR <= 0.05 and fold change >= 1.5. The gene set enrichment analysis was performed using ClusterProfiler to identify the affected Hallmark and KEGG pathways from MSigDB v7, using the pre-ranked gene list sorted by the -log10(p-value) with a sign determined by the fold change direction.
Bulk ATAC-sequencing library preparation, sequencing and data analysis
To prepare for the nuclei, we spun 50,000 cells at 300 × g for 5 min, followed by a wash using 50 μL of cold 1x PBS and centrifugation at 300 × g for 5 min. Cells were lysed using 50 μL cold lysis buffer (10 mM Tris-Cl, pH 7.4, 10 mM NaCl, 3 mM MgCl2, 0.1% NP-40, 0.1% Tween-20, 0.01% Digitonin and 0.5% BSA). Immediately after lysis, 500 μL wash buffer was added, and nuclei were spun at 500 × g for 10 min using a refrigerated centrifuge. To avoid losing cells during the nuclei prep, we used a fixed angle centrifuge and carefully pipetted away from the pellet after centrifugations. Nuclei tagmentation and adapter ligation by Tn5 was performed using the Nextera DNA Sample Preparation kit (Illumina), followed by purification with The Oligo Clean & Concentrator kit (ZYMO) according to the manufacturers’ instructions. Transposed DNA fragments were amplified using the NEBNext Q5 HotStart HiFi PCR Master Mix with regular forward and reverse barcoded primers. The numbers of additional amplification cycles were determined by quantitative-PCR using the NEBNext HiFi Master Mix, SYBR Green I (Applied Biosystems) and Custom Nextera Primers. The final product was prepared by using double-sided bead purification 0.5X and 1.8x volume AMPure XP beads (Beckman Colter Inc.) and quality checked on 2100 Bioanalyzer (Agilent). Sequencing was performed using a NovaSeq 6000 Sequencing System (Illumina, San Diego, CA, USA) instrument for 2 ×101 bp sequencing with ~50 million reads per sample.
Data analysis
ATAC-seq data were aligned to mouse genome mm10 using BWA-MEM with the default settings. Peak calling was performed by using Genrich. Peaks were annotated by the R package “ChIPseeker”. Kmeans clustering was generated using R package “pheatmap”. GO analysis of peaks in each cluster was conducted by “clusterProfiler”. Reads in peaks were scaled into RPKM values followed by log2 transformation with offset of 1 for fold change calculation. Fold changes >= 2 and p value <= 0.05 across conditions were considered significant. ATAC peak changes across sample groups were plotted using R package “ggalluvial”. Motifs enriched at the ATAC-seq peaks with 250 bp flanking region were identified using HOMER. The output from HOMER was parsed using R package “monaLisa” and volcano plots were generated using “EnhancedVolcano”.
Imaging Mass Cytometry
For performing imaging Mass cytometry (IMC), liver tissue sections from cGVHD patients, fixed in formalin and embedded in paraffin (FFPE), were dewaxed using fresh xylene, rehydrated with a series of alcohol dilutions, and washed with distilled water. Antigen retrieval was performed in a Biocare Medical decloaking chamber at 95 °C for 65 min in Tris-EDTA buffer (pH=9). The slides were cooled to room temperature, blocked with SuperBlock T20 (PBS) Blocking Buffer (Thermo Scientific) for 30 min, and incubated overnight at 4 °C with primary antibody mixtures as described below. After incubation, the slides were washed with PBS-TB (PBS supplemented with 0.05% Tween 20 and 1% BSA) three times and stained with a nuclear intercalator solution containing iridium 193 and 191 isotopes for 10 min at room temperature. Finally, the slides were washed with distilled water and dried at room temperature before data acquisition using a Hyperion imaging mass cytometer (Fluidigm). The data were stored as MCD files and txt files, and the MCD viewer, Fiji ImageJ61 software and Noise2Void62 were used for image processing.
IMC analysis
Raw IMC data were collected and stored in the MCD and txt file. Single cells were segmented with Steinbock guided by the ImcSegmentationPipeline shared by Bodenmiller’s group63. Output files were then imported into R using Rstudio interface for further analysis. Image and cell-level quality control and batch effect correction were conducted. Cell type labeling was applied by Shiny application of R package cytomapper. Briefly, the following markers were used to define cell types: B cells (CD20), Macrophage (CD68+ HLA-DR+), DCs (CD68− CD11c+ HLA-DR+), Endothelial cells (CD31+), epithelial cells (E-cadherin+), CD4+ Tsm (CD3+ CD4+ CD45RO+ Ly108+ CD69−), CD4+ Trmp cells (CD3+ CD4+ CD45RO+ Ly108+ CD69+), CD4+ Trm cells (CD3+ CD4+ CD45RO+ Ly108−CD69+), Tint cells (CD3+ CD4+ CD45RO+ Ly108−CD69−). Gated cells were then downloaded as SingleCellExperiment objects stored in the metadata. Label cells were then trained in a random forest classifier. Single cell visualization and spatial visualization were later performed with imcRtools package63.
Statistical analysis
The results presented in this study were obtained from at least two replicate experiments. Data are presented as mean ± SEM. Survival was compared using the log-rank test. When data were normally distributed, Student’s unpaired t-test was used to compare two groups. For multiple comparisons of one independent variable, ordinary one-way ANOVA with Holm-Šídák test was used, while for two or more independent variables, two-way ANOVA with Holm-Šídák test was used. Prism9 GraphPad Software was used for statistical analysis. Statistical significance was considered at P < 0.05 (*P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The bulk RNA-sequencing, ATAC-sequencing and single cell RNA-TCR-seq data have been deposited and publicly available under accession codes GSE263810, GSE263811, GSE263812, GSE263813. All data are included in the Supplementary Information or available from the authors, as are unique reagents used in this Article. The raw numbers for charts and graphs are available in the Source Data file whenever possible. Source data are provided with this paper.
References
Ferrara, J. L., Levine, J. E., Reddy, P. & Holler, E. Graft-versus-host disease. Lancet 373, 1550–1561 (2009).
Zeiser, R. & Blazar, B. R. Pathophysiology of chronic graft-versus-host disease and therapeutic targets. N. Engl. J. Med. 377, 2565–2579 (2017).
Shlomchik, W. D. Graft-versus-host disease. Nat. Rev. Immunol. 7, 340–352 (2007).
Buxbaum, N. P. et al. Chronic GvHD NIH consensus project biology task force: evolving path to personalized treatment of chronic GvHD. Blood Adv. 7, 4886–4902 (2023).
Flowers, M. E. & Martin, P. J. How we treat chronic graft-versus-host disease. Blood 125, 606–615 (2015).
Zhang, C. et al. Donor CD4+ T and B cells in transplants induce chronic graft-versus-host disease with autoimmune manifestations. Blood 107, 2993–3001 (2006).
Zhao, D. et al. In vivo-activated CD103+CD4+ regulatory T cells ameliorate ongoing chronic graft-versus-host disease. Blood 112, 2129–2138 (2008).
Zhao, D. et al. Alloimmune response results in expansion of autoreactive donor CD4+ T cells in transplants that can mediate chronic graft-versus-host disease. J. Immunol. 186, 856–868 (2011).
Wu, T. et al. Thymic damage, impaired negative selection, and development of chronic graft-versus-host disease caused by donor CD4+ and CD8+ T cells. J. Immunol. 191, 488–499 (2013).
Young, J. S. et al. Donor B cells in transplants augment clonal expansion and survival of pathogenic CD4(+) T cells that mediate autoimmune-like chronic graft-versus-host disease. J. Immunol. 189, 222–233 (2012).
Dertschnig, S., Hauri-Hohl, M. M., Vollmer, M., Hollander, G. A. & Krenger, W. Impaired thymic expression of tissue-restricted antigens licenses the de novo generation of autoreactive CD4+T cells in acute GVHD. Blood 125, 2720–2723 (2015).
Deng, R. et al. Extrafollicular CD4(+) T-B interactions are sufficient for inducing autoimmune-like chronic graft-versushost disease. Nat. Commun. 8, 978 (2017).
Song, Q., Kong, X., Martin, P. J. & Zeng, D. Murine models provide new insights into pathogenesis of chronic graft-versus-host disease in humans. Front. Immunol. 12, 700857 (2021).
Reddy, P., Negrin, R. & Hill, G. R. Mouse models of bone marrow transplantation. Biol. Blood Marrow Transpl. 14, 129–135 (2008).
Filipovich, A. H. et al. National Institutes of Health consensus development project on criteria for clinical trials in chronic graft-versus-host disease: I. Diagnosis and staging working group report. Biol. Blood Marrow Transpl. 11, 945–956 (2005).
Shlomchik, W. D. et al. Prevention of graft versus host disease by inactivation of host antigen-presenting cells. Science 285, 412–415 (1999).
Teshima, T. et al. Acute graft-versus-host disease does not require alloantigen expression on host epithelium. Nat. Med. 8, 575–581 (2002).
Chakraverty, R. & Sykes, M. The role of antigen-presenting cells in triggering graft-versus-host disease and graft-versus-leukemia. Blood 110, 9–17 (2007).
Koyama, M. et al. MHC class II antigen presentation by the intestinal epithelium initiates graft-versus-host disease and is influenced by the microbiota. Immunity 51, 885–898 (2019).
Kong, X. et al. Tissue-resident PSGL1loCD4+T cells promote B cell differentiation and chronic graft-versus-host disease-associated autoimmunity. J. Clin. Invest 131, e135468 (2021).
Zhang, Y., Joe, G., Hexner, E., Zhu, J. & Emerson, S. G. Host-reactive CD8+ memory stem cells in graft-versus-host disease. Nat. Med 11, 1299–1305 (2005).
Im, S. J. et al. Defining CD8+ T cells that provide the proliferative burst after PD-1 therapy. Nature 537, 417–421 (2016).
Zhou, X. et al. Differentiation and persistence of memory CD8(+) T cells depend on T cell factor 1. Immunity 33, 229–240 (2010).
Utzschneider, D. T. et al. T cell factor 1-expressing memory-like CD8(+) T cells sustain the immune response to chronic viral infections. Immunity 45, 415–427 (2016).
Zehn, D., Thimme, R., Lugli, E., de Almeida, G. P. & Oxenius, A. Stem-like’ precursors are the fount to sustain persistent CD8(+) T cell responses. Nat. Immunol. 23, 836–847 (2022).
Chen, Z. et al. TCF-1-centered transcriptional network drives an effector versus exhausted CD8 T cell-fate decision. Immunity 51, 840–855 (2019).
Wu, T. et al. The TCF1-Bcl6 axis counteracts type I interferon to repress exhaustion and maintain T cell stemness. Sci. Immunol. 1, eaai8593 (2016).
Masopust, D., Vezys, V., Marzo, A. L. & Lefrancois, L. Preferential localization of effector memory cells in nonlymphoid tissue. Science 291, 2413–2417 (2001).
Shiow, L. R. et al. CD69 acts downstream of interferon-alpha/beta to inhibit S1P1 and lymphocyte egress from lymphoid organs. Nature 440, 540–544 (2006).
Park, C. O. & Kupper, T. S. The emerging role of resident memory T cells in protective immunity and inflammatory disease. Nat. Med. 21, 688–697 (2015).
Beltra, J. C. et al. Developmental relationships of four exhausted CD8(+) T cell subsets reveals underlying transcriptional and epigenetic landscape control mechanisms. Immunity 52, 825–841 (2020).
Zou, D. et al. CD4(+) T cell immunity is dependent on an intrinsic stem-like program. Nat. Immunol. 25, 66–76 (2024).
Li, Y. et al. Stem-like T cells are associated with the pathogenesis of ulcerative colitis in humans. Nat. Immunol. 25, 1231–1244 (2024).
Beltra, J. C. et al. Stat5 opposes the transcription factor Tox and rewires exhausted CD8(+) T cells toward durable effector-like states during chronic antigen exposure. Immunity 56, 2699–2718 (2023).
Kong, X. et al. Trafficking between clonally related peripheral T-helper cells and tissue-resident T-helper cells in chronic GVHD. Blood 140, 2740–2753 (2022).
Sacirbegovic, F. et al. Graft-versus-host disease is locally maintained in target tissues by resident progenitor-like T cells. Immunity 56, 369–385 (2023).
Gearty, S. V. et al. An autoimmune stem-like CD8 T cell population drives type 1 diabetes. Nature 602, 156–161 (2022).
Khan, O. et al. TOX transcriptionally and epigenetically programs CD8(+) T cell exhaustion. Nature 571, 211–218 (2019).
Tugues, S. et al. Graft-versus-host disease, but not graft-versus-leukemia immunity, is mediated by GM-CSF-licensed myeloid cells. Sci. Transl. Med 10, eaat8410 (2018).
Anderson, K. G. et al. Intravascular staining for discrimination of vascular and tissue leukocytes. Nat. Protoc. 9, 209–222 (2014).
Yao, C. et al. Single-cell RNA-seq reveals TOX as a key regulator of CD8(+) T cell persistence in chronic infection. Nat. Immunol. 20, 890–901 (2019).
Borges da Silva, H., Wang, H., Qian, L. J., Hogquist, K. A. & Jameson, S. C. ARTC2.2/P2RX7 Signaling during Cell Isolation Distorts Function and Quantification of Tissue-Resident CD8(+) T Cell and Invariant NKT Subsets. J. Immunol. 202, 2153–2163 (2019).
Zundler, S. et al. Hobit- and Blimp-1-driven CD4(+) tissue-resident memory T cells control chronic intestinal inflammation. Nat. Immunol. 20, 288–300 (2019).
Anderson, B. E. et al. Distinct roles for donor- and host-derived antigen-presenting cells and costimulatory molecules in murine chronic graft-versus-host disease: requirements depend on target organ. Blood 105, 2227–2234 (2005).
Steimle, V., Siegrist, C. A., Mottet, A., Lisowska-Grospierre, B. & Mach, B. Regulation of MHC class II expression by interferon-gamma mediated by the transactivator gene CIITA. Science 265, 106–109 (1994).
Hamann, D. et al. Phenotypic and functional separation of memory and effector human CD8+ T cells. J. Exp. Med. 186, 1407–1418 (1997).
Hoch, T. et al. Multiplexed imaging mass cytometry of the chemokine milieus in melanoma characterizes features of the response to immunotherapy. Sci. Immunol. 7, eabk1692 (2022).
Lee, H., Jeong, S. & Shin, E. C. Significance of bystander T cell activation in microbial infection. Nat. Immunol. 23, 13–22 (2022).
Li, C. et al. The Transcription factor Bhlhe40 programs mitochondrial regulation of resident CD8(+) T cell fitness and functionality. Immunity 51, 491–507 (2019).
Wu, J. E. et al. In vitro modeling of CD8(+) T cell exhaustion enables CRISPR screening to reveal a role for BHLHE40. Sci. Immunol. 8, eade3369 (2023).
Mammadli, M., Suo, L., Sen, J. M. & Karimi, M. TCF-1 Is Required for CD4 T Cell Persistence Functions during AlloImmunity. Int J. Mol. Sci. 24, 4326 (2023).
Reddy, P. et al. A crucial role for antigen-presenting cells and alloantigen expression in graft-versus-leukemia responses. Nat. Med 11, 1244–1249 (2005).
Piper, C. et al. Pathogenic Bhlhe40+ GM-CSF+ CD4+ T cells promote indirect alloantigen presentation in the GI tract during GVHD. Blood 135, 568–581 (2020).
Geng, H. et al. Self-enforcing feedback activation between BCL6 and pre-B cell receptor signaling defines a distinct subtype of acute lymphoblastic leukemia. Cancer Cell 27, 409–425 (2015).
Song, Q. et al. IL-22-dependent dysbiosis and mononuclear phagocyte depletion contribute to steroid-resistant gut graft-versus-host disease in mice. Nat. Commun. 12, 805 (2021).
Hao, Y. et al. Dictionary learning for integrative, multimodal and scalable single-cell analysis. Nat. Biotechnol. 42, 293–304 (2024).
Qiu, X. et al. Reversed graph embedding resolves complex single-cell trajectories. Nat. Methods 14, 979–982 (2017).
Chen, Y. et al. BATF regulates progenitor to cytolytic effector CD8(+) T cell transition during chronic viral infection. Nat. Immunol. 22, 996–1007 (2021).
Borcherding, N., Bormann, N. L. & Kraus, G. scRepertoire: an R-based toolkit for single-cell immune receptor analysis. F1000Res 9, 47 (2020).
Aibar, S. et al. SCENIC: single-cell regulatory network inference and clustering. Nat. Methods 14, 1083–1086 (2017).
Rueden, C. T. et al. ImageJ2: imagej for the next generation of scientific image data. BMC Bioinforma. 18, 529 (2017).
A. Krull, T.-O. Buchholz., F. Jug. Noise2Void - Learning Denoising From Single Noisy Images. Proc. IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), pp. 2129-2137 (CVPR, 2019).
Windhager, J. et al. An end-to-end workflow for multiplexed image processing and analysis. Nat. Protoc. 18, 3565–3613 (2023).
Acknowledgements
This work was supported by NIH 1R01HL170099 and R01HL162847 and Excellence Award of The Beckman Research Institute (to D. Zeng), and institutional NCI P30CA033572. Graduate Academic Exchange Scholarship of Fujian Medical University and Riggs-Union International Exchange Scholarship of Fujian Medical University Union Hospital partially supported B. Wang’s living stipend. We thank Lucy Brown and her staff at the COH Flow Cytometry Facility; Raju Pillai, Aimin Li and staff at COH Pathology-Solid Tumor Core; Brian Armstrong and his staff at COH Light Microscopy Core; and Dr. Richard Ermel and his staff at COH Animal Research Center for providing excellent service. We also thank Dr. Cecelia Yeung and David Woolston at the Fred Hutchinson Cancer Center for assistance in providing sections of liver biopsies from patients with chronic GVHD.
Author information
Authors and Affiliations
Contributions
XK designed and performed experiments, acquired and analyzed data, prepared manuscript and wrote the draft manuscript. BW designed and performed experiments, acquired and analyzed data. XW and HQ performed scRNA/scTCR-sequencing, bulk-cell RNA-seq/ATAC-seq, and XW, HC, XK performed data analysis. WF, QL, RZ, MZ, UN and AW assisted in experiments. JL, TO provided Hobit−/− mice. RN, RP provided advice on human-related studies, organized human samples, and reviewed the manuscript. PJM provided advice on experimental design and critical review and editing of the manuscript. YC is BW’s PhD advisor. DZ designed and supervised the research and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Nu Zhang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Kong, X., Wang, B., Wu, X. et al. Stem-like memory-T maintenance and differentiation into tissue-resident T cells sustain chronic graft-versus-host disease in mice. Nat Commun 17, 3147 (2026). https://doi.org/10.1038/s41467-026-69975-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-69975-z