Abstract
Activation-induced cytidine deaminase (AID)-initiated immunoglobulin heavy chain (Igh) class switch recombination (CSR) replaces donor Cμ constant region exons (CHs) with a downstream acceptor CH to generate different functional antibody isotypes. However, mechanisms governing orientation-specific productive CSR remain incompletely understood. Through analysing the characteristics of evolved constant regions and constructing diversified constant regions to recapitulate productive CSR in jawed vertebrates to systematically dissect productive CSR determinants, we find that switch topological configuration (STC), including transcriptional orientation, chromatin distance, and chromatin domain of Igh, determines orientation-specific joining of AID-initiated breaks for productive CSR. Long-distance CHs under co-oriented transcription within Igh domain foster predominantly deletional joining-mediated productive CSR. In contrast, oppositely transcribed and short-distance CHs affect end-joining bias via promoting diffusion-mediated inversional joining for CSR with lower efficiency. Moreover, AID-initiated breaks in different domains, facilitate more diffusion-mediated orientation-unbiased end-joining for non-productive CSR. Our findings uncover chromatin-intrinsic mechanisms safeguarding orientation-specific productive CSR throughout evolution.
Similar content being viewed by others
Introduction
The functional efficacy of adaptive immunity is a defining feature of jawed vertebrates that equips them to counteract rapidly evolving pathogens (e.g., viruses, bacteria), which is crucially hinged on two synergistic processes: V(D)J recombination and class switch recombination (CSR)1. During B lymphocyte development, RAG endonuclease cleaves immunoglobulin heavy chain (Igh) V, D, and J gene segments to assemble a V(D)J exon adjacent to the Cμ constant region exons (CHs), encoding the default IgM antibody2,3. Upon B lymphocyte activation, Igh CSR replaces Cμ with one of several downstream acceptor CHs to change antibody isotypes and functions3,4. Each CH unit contains an I-promoter, a long repetitive switch (S) region intron, and several CH coding exons5. Chromatin loop extrusion6,7-mediated Igh 3′ regulatory region (3′RR) enhancer scanning activates transcription of acceptor CHs and promotes S-S synapsis for recombination5,8,9,10. Activation-induced cytidine deaminase (AID)11 targets the transcribed donor Sμ region and an acceptor S region to initiate deamination lesions12, which are converted to double-strand breaks (DSBs) and joined by DNA damage repair factors to accomplish CSR4,13,14,15. The Igh domain boundary, termed 3′CBEs downstream of 3′RR, consisted of ten consecutive CTCF-binding elements (CBEs)16, safeguards loop extrusion-mediated transcriptional and CSR activities within the Igh domain17,18.
Unlike the orientation-unbiased random joining of most chromosomal DSB ends genome-wide19,20,21, AID-initiated CSR within the Igh domain employs an orientation-specific joining mechanism to generate productive antibody isotypes22. The mouse Igh domain contains six acceptor CHs in the same transcriptional orientation as the upstream donor Cμ, indicating that only deletional joining between donor Sμ breaks and acceptor S breaks results in productive CSR10. DNA damage repair factors, such as ATM22,23,24,25, H2AX22,25,26, 53BP122,25,27,28, RIF122,25,29,30, Rev731,32, Shieldin33,34,35,36,37, and Ligase 425,38, have been found to promote deletional joining-mediated productive CSR by facilitating DSB tethering, suppressing DSB processing, and decreasing microhomology (MH) of CSR junctions in mice. Previous studies implicate the DNA damage repair factors in enforcing the deletional joining mechanism, but do not implicate them in providing a basis for the orientation-specific CSR22.
Chromatin loop extrusion has been proposed to be the underlying mechanism for CSR in Mus musculus and Homo sapiens, which promotes the formation of dynamic CSR centers where acceptor S regions get activated and synapsed with donor Sμ for CSR9,10,39. Moreover, chromatin loop extrusion has also been proposed to explain the predominantly deletional CSR in Mus musculus, supported by the increase of inversional joining from 3% to 19% during Sα CSR via inserting CBE-mediated chromatin loop extrusion blocks within the Igh constant region in murine CH12F3 cells9,10. However, in some birds and reptiles, CSR must proceed by inversional joining to generate productive antibody isotypes due to the opposite transcriptional orientation of donor Cμ and some acceptor CHs40,41. The chromatin loop extrusion-mediated predominantly deletional CSR in mice, and the structural diversity of constant regions across jawed vertebrates, suggest that regulation of orientation-specific joining for productive CSR is more than previously documented9,10,22,41. Thus, detailed mechanisms need to be investigated to comprehensively explain the joining features of productive CSR with different Igh architectures throughout evolution.
In this study, by comprehensively analysing the constant region across a series of species, we revealed that the distinct chromatin features play critical roles in regulating productive CSR. We constructed a series of murine CH12F3 cell mutants with distinct Igh configurations and found that switch topological configuration (STC), including transcription orientation, chromatin distance, and chromatin domain, determines orientation-specific joining of AID-initiated breaks for productive CSR.
Results
Intrinsic chromatin features affect end-joining of CSR
To investigate the potential influencing factors promoting productive CSR with different Igh architectures throughout evolution, we conducted a detailed comparative analysis of the constant region across multiple species. The sequence feature analysis of S regions indicated the GC content within S regions was increased in evolution (Supplementary Fig. 1a). In addition, RNA-seq analysis showed an adaptive fixation of unidirectional transcription polarity and a progressive elongation of Igh constant region along the evolutionary lineage from birds to mammals (Supplementary Fig. 1b). Moreover, Hi-C analysis indicated an increasing conservation of constant region within a domain across the Osteichthyes-to-Mammalia transition (Supplementary Fig. 1c). We further manipulated the Igh constant region of murine CH12F3 cells to investigate the roles of chromatin configurations in orientation-specific CSR.
CH12F3 cells undergo robust CSR between Sμ and Sα to generate predominantly deletional joining-mediated productive IgA antibodies upon αCD40/IL4/TGFβ activation22,42. To eliminate the potentially confounding effects of a non-productive Igh allele on the assessment of sequencing results, we employed Cas9/gRNA targeting to delete the entire non-productive Igh locus from upstream of the first VH to downstream of 3′CBEs in CH12F3NCΔ cells9, thereby generating CH12F3PO cells (Supplementary Fig. 2a, b). We placed the Cα unit next to the donor Cμ with opposite transcriptional orientation by inverting the cluster of six acceptor CHs from upstream of Cγ3 to downstream of Cα in CH12F3PO cells to generate γ3-αInv cells (Fig. 1a, Supplementary Fig. 2c, d). Then we employed high-sensitive CSR-HTGTS-seq to assay the joining features of CSR junctions (Fig. 1b) and found that γ3-αInv cells exhibited predominantly deletional Sγ3 CSR and a small but nonnegligible number of Sα CSR (Fig. 1c). Remarkably, in contrast to the predominantly deletional Sα CSR in CH12F3PO cells, Sα CSR underwent orientation-unbiased joining with 46% inversional joining of Sμ–Sα junctions in γ3-αInv cells without perturbing DNA damage repair factors (Fig. 1d, e).
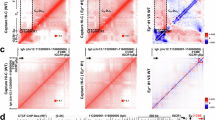
a Schematic of the Igh locus from V(D)J exon to 3′CBEs in CH12F3PO cells and an illustration of the generation of γ3-αInv cells. b Illustration of deletional and inversional joining between the donor Sμ and an acceptor Sx in CSR. The CSR joining features were measured by CSR-HTGTS-seq with 5′Sμ bait. c CSR-HTGTS-seq analysis of break joining between 5′Sμ and acceptor S regions in CH12F3PO and γ3-αInv cells stimulated with αCD40/IL4/TGFβ for 72 h. Junctions within the Igh constant region are plotted at a 2.25 kb bin size. Data are presented as mean ± s.e.m. from three independent experiments. d Joining features of Sμ–Sα junctions in CH12F3PO and γ3-αInv cells. Data are presented as mean ± s.e.m. from three independent experiments. e Bar graph shows the inversional joining of Sμ–Sα junctions in CH12F3PO and γ3-αInv cells. Data are presented as mean ± s.e.m. from six independent &experiments. P-values were calculated via an unpaired two-tailed Student’s t-test. f Pro-seq and 3C-HTGTS with iEμ/Iμ or 3′RR bait (blue asterisks) show the transcription status and chromatin interaction of the Igh locus from CH12F3PO cells. g Pro-seq and 3C-HTGTS with iEμ/Iμ or 3′RR bait (blue asterisks) show the transcription status and chromatin interaction of the Igh locus from γ3-αInv cells. h Bar graph shows the relative transcription activity of the Cγ3 unit and the Cα unit in CH12F3PO and γ3-αInv cells. Data are presented as mean ± s.e.m. from three independent experiments. P-values were calculated via a paired two-tailed Student’s t-test. i Bar graph shows the relative interaction frequency between iEμ/Iμ bait with Cγ3 unit and Cα unit in CH12F3PO and γ3-αInv cells. Data are presented as mean ± s.e.m. from three independent experiments. P-values were calculated via a paired two-tailed Student’s t-test. Source data are provided as a Source Data file.
We further used Pro-seq and 3C-HTGTS analyses to check the transcription status and chromatin interaction of the Igh locus to reveal the underlying mechanisms. In contrast to the predominantly transcription of the downstream Cα unit in CH12F3PO cells, γ3-α cluster inversion significantly activated transcription of the downstream Cγ3 unit and repressed upstream Cα transcription in γ3-αInv cells (Fig. 1f–h, Supplementary Fig. 2e, f), suggesting that chromatin loop extrusion-mediated 3′RR scanning predominantly activates the Cγ3 unit instead of the upstream Cα unit in γ3-αInv cells. And the low transcription of the Cα unit corresponds to the low level of AID-initiated Sα CSR in γ3-αInv cells (Fig. 1c, g). In addition, unlike the dominant interaction between the Cα unit with iEμ and 3′RR in CH12F3PO cells, the Cγ3 unit interacted with iEμ and 3′RR within the CSR center in γ3-αInv cells, facilitating Sμ−Sγ3 synapsis-mediated predominant deletional joining of Sμ−Sγ3 junctions (Fig. 1c, f, g, i, Supplementary Fig. 2e, f). Although the rarity of interactions between donor Cμ and acceptor Cα unit limited Sμ–Sα synapsis formation in γ3-αInv cells, low-frequent short-range diffusion could promote orientation-unbiased random joining of Sμ–Sα junctions, accounting for 46% inversional Sα CSR for generating productive IgA antibodies (Fig. 1d–i, Supplementary Fig. 2e, f). These data suggest that chromatin configuration, including transcriptional orientation and chromatin distance of CHs, affects orientation-specific joining of CSR within the Igh domain.
Transcription orientation of CHs affects end-joining of CSR
Given that both transcriptional orientation and chromatin distance of Cμ and Cα units were changed in γ3-αInv cells by inverting the entire acceptor CH region (Fig. 1a), we further inverted different acceptor CH units individually to change the relative transcription orientation between donor Cμ and acceptor CHs. We first inverted the Cα unit, the farthest downstream acceptor CH from donor Cμ in CH12F3PO cells, to generate αInv cells (Fig. 2a, Supplementary Fig. 3a, b). The results showed that αInv cells slightly increased inversional joining of Sμ–Sα junctions from 4% to 8% (Fig. 2b–d). And this increase of inversional joining in αInv cells had little effect on the MH of Sμ–Sα junctions (Supplementary Fig. 3c), suggesting a distinct regulation from DNA damage repair factor-deficiency-mediated increase of inversional joining during CSR. Then we took advantage of the activated CSR from upstream acceptor CHs upon Iα deletion in IαΔ CH12F3 cells9. We inverted Cγ2a unit in IαΔ cells to generate γ2aInv cells (Fig. 2e, Supplementary Fig. 3d, e) and found that γ2aInv cells significantly increased inversional joining of Sμ−Sγ2a junctions from 12% to 25% without affecting MHs (Fig. 2f–h, Supplementary Fig. 3f). We further inverted Cγ1 unit in IαΔ cells to generate γ1Inv cells (Fig. 2i, Supplementary Fig. 3g, h) and found that γ1Inv cells significantly increased inversional joining of Sμ−Sγ1 junctions from 5% to 43% without affecting MHs (Figs. 2j–l, Supplementary Fig. 3i). We further inverted Cα unit in γ3-αInv cells to generate γ3-αInv-αInv cells (Fig. 2m, Supplementary Fig. 3j, k). Notably, γ3-αInv-αInv cells significantly decreased inversional joining of Sμ–Sα junctions from 43% to 18% without affecting MHs (Fig. 2n–p, Supplementary Fig. 3l). All these data indicate that transcription orientation plays distinct roles from DNA damage repair factors in regulating orientation-specific joining of CSR, with more inversional joining from two opposite transcribed CHs.
a Schematic of the Igh locus from V(D)J exon to 3′CBEs in CH12F3PO cells and an illustration of the generation of αInv cells. Joining features of Sμ–Sα junctions in CH12F3PO (b) and αInv (c) cells. Data are presented as mean ± s.e.m. from three independent experiments. d Bar graph shows the inversional joining of Sμ–Sα junctions in CH12F3PO and αInv cells. Data are presented as mean ± s.e.m. from five independent experiments. e Schematic of the Igh locus from the V(D)J exon to 3′CBEs in IαΔ cells and an illustration of the generation of γ2aInv cells. Joining features of Sμ−Sγ2a junctions in IαΔ (f) and γ2aInv (g) cells. h Bar graph shows the inversional joining of Sμ−Sγ2a junctions in IαΔ and γ2aInv cells. i Schematic of the Igh locus from V(D)J exon to 3′CBEs in IαΔ cells and illustration of the generation of γ1Inv cells. Joining features of Sμ−Sγ1 junctions in IαΔ (j) and γ1Inv (k) cells. l Bar graph shows the inversional joining of Sμ−Sγ1 junctions in IαΔ and γ1Inv cells. m Schematic of Igh locus from V(D)J exon to 3′CBEs in γ3-αInv cells and illustration of the generation of γ3-αInv-αInv cells. Joining features of Sμ–Sα junctions in γ3-αInv (n) and γ3-αInv-αInv (o) cells. p Bar graph shows the inversional joining of Sμ–Sα junctions in γ3-αInv and γ3-αInv-αInv cells. For (f–h), (j–l), and (n–p), data are presented as mean ± s.e.m. from three independent experiments. P-values were calculated via an unpaired two-tailed Student’s t-test. Source data are provided as a Source Data file.
Chromatin distance of CHs affects end-joining of CSR
To further assess the effects of chromatin distance between donor Cμ and acceptor CH on orientation-specific joining of CSR, we changed the chromatin distance between Cμ and Cγ3 unit. We first inverted the Cγ3 unit in IαΔ cells to generate γ3Inv cells, which have a shorter distance between Cμ and Cγ3 (Fig. 3a, Supplementary Fig. 4a, b), and γ3Inv cells significantly increased inversional joining of Sμ−Sγ3 junctions from 8% to 20% (Fig. 3b–d). Given that γ3Inv cells changed both chromatin distance and transcription orientation of Cμ and Cγ3 compared to IαΔ cells, we further inverted the Cγ3 unit in γ3Inv cells to generate γ3Inv-γ3Inv cells (Fig. 3a, Supplementary Fig. 4a, c). We found that γ3Inv-γ3Inv cells decreased inversional Sμ−Sγ3 junctions from 20% to 14% compared to γ3Inv cells (Fig. 3c–e), consistent with the conclusion that opposite transcriptional orientation of CHs increases inversional joining of CSR (Fig. 2). Notably, compared to IαΔ cells, γ3Inv-γ3Inv cells increased inversional Sμ−Sγ3 junctions from 8% to 14% (Fig. 3b, d, e), indicating that chromatin distance contributes to orientation-specific joining of CSR. We also deleted the intergenic region between Cμ and Cγ3 unit directly to generate γ3IRD cells (Fig. 3a, Supplementary Fig. 4d, e), and γ3IRD cells significantly increased inversional joining of Sμ−Sγ3 junctions from 8% to 15% (Fig. 3b, d, f), exhibiting a similar trend as γ3Inv-γ3Inv cells. And this short-distance-mediated increase of inversional joining was not dependent on changing MH-related DNA damage repair pathways (Supplementary Fig. 4f). Altogether, these data indicate that chromatin distance of CHs plays roles in regulating orientation-specific joining of CSR, with more diffusion-mediated inversional joining from two short-distance CHs.
a A Schematic of the Igh locus from V(D)J exon to 3′CBEs in IαΔ cells and an illustration of the generation of γ3Inv, γ3Inv-γ3Inv, and γ3IRD cells. Joining features of Sμ−Sγ3 junctions in IαΔ (b), γ3Inv (c), γ3Inv-γ3Inv (e), and γ3IRD (f) cells. d Bar graph shows the inversional joining of Sμ−Sγ3 junctions in IαΔ, γ3Inv, γ3Inv-γ3Inv, and γ3IRD cells. Data are presented as mean ± s.e.m. from three independent experiments. P-values were calculated via an unpaired two-tailed Student’s t-test. Source data are provided as a Source Data file.
Inverted I-promoter-mediated block activates inversional CSR
To reveal the regulatory mechanism of transcriptional orientation in orientation-specific joining of CSR, we further fused the full Sγ3 and Sα in CH12F3PO cells and αInv cells to generate Iγ3-FS-Cα and Iγ3-FS-Iα cells, respectively (Supplementary Fig. 5a–c). In Iγ3-FS-Cα cells, both Sγ3 and Sα were transcribed under Iγ3 promoter with the same transcription orientation as donor Cμ, synapsed with donor Cμ, and underwent robust deletional Sγ3 CSR and Sα CSR at similar levels (Fig. 4a, Supplementary Fig. 5d). However, in Iγ3-FS-Iα cells, both Sγ3 and Sα were transcribed under inverted Iα promoter instead of the upstream Iγ3 promoter, suggesting that 3′RR enhancer scans to activate the closer promoter for transcriptional activation (Fig. 4b, Supplementary Fig. 5e). Compared to the robust synapsis between donor Sμ and both Sγ3 and Sα in Iγ3-FS-Cα cells (Fig. 4a, Supplementary Fig. 5d), Iγ3-FS-Iα cells exhibited major interaction between donor Sμ with inverted Iα promoter and adjacent Sα, which blocked transcriptional activation of Iγ3 promoter, leading to predominantly Sα CSR and less frequent Sγ3 CSR (Fig. 4b, Supplementary Fig. 5e). Meanwhile, cohesin significantly accumulated at the inverted Iα promoter and this inverted Iα promoter with cohesin accumulation-mediated block effects increased inversional joining of Sμ–Sα junctions from 8% to 19% and of less frequent Sμ−Sγ3 junctions from 10% to 36% (Fig. 4a, b, Supplementary Fig. 5f).
a CSR-HTGTS-seq, Pro-seq, 3C-HTGTS with iEμ/Iμ (blue asterisks), and Rad21 ChIP-seq to show the CSR junctions, transcription status, chromatin interaction, and cohesin binding of the Igh locus from Iγ3-FS-Cα cells. b CSR-HTGTS-seq, Pro-seq, 3C-HTGTS with iEμ/Iμ bait (blue asterisks), and Rad21 ChIP-seq to show the CSR junctions, transcription status, chromatin interaction, and cohesin binding of the Igh locus from Iγ3-FS-Iα cells. c CSR-HTGTS-seq, Pro-seq, 3C-HTGTS with iEμ/Iμ (blue asterisks), and Rad21 ChIP-seq to show the CSR junctions, transcription status, chromatin interaction, and cohesin binding of the Igh locus from Iγ3-PS-Cα cells. d CSR-HTGTS-seq, Pro-seq, 3C-HTGTS with iEμ/Iμ bait (blue asterisks), and Rad21 ChIP-seq to show the CSR junctions, transcription status, chromatin interaction, and cohesin binding of the Igh locus from Iγ3-PS-Iα cells. For (a–d), data are presented as mean ± s.e.m. from three independent experiments. Source data are provided as a Source Data file.
We also fused the partial Sγ3 and Sα in CH12F3PO cells and αInv cells to generate Iγ3-PS-Cα and Iγ3-PS-Iα cells, respectively (Supplementary Fig. 5a, g, and h). In Iγ3-PS-Cα cells, both Sγ3 and Sα were transcribed under the Iγ3 promoter with the same transcription orientation as donor Cμ, synapsed with donor Cμ, and underwent robust Sγ3 CSR and less frequent Sα CSR (Fig. 4c, Supplementary Fig. 5i). Compared to the robust Sα CSR in Iγ3-FS-Cα and Sγ3 CSR in Iγ3-PS-Cα, the less frequent Sα CSR in Iγ3-PS-Cα cells also suggests that AID targets more 5′S region for CSR (Fig. 4a, c). Iγ3-PS-Iα cells exhibited a similar trend of transcription status, chromatin interaction, cohesin accumulation, and CSR as Iγ3-FS-Iα cells (Fig. 4b, d, Supplementary Fig. 5e, j). The inverted Iα promoter in Iγ3-PS-Iα cells dominated the transcription orientation, chromatin interaction, cohesin accumulation, and CSR with more diffusion-mediated inversional Sμ–Sα and Sμ−Sγ3 junctions (Fig. 4d, Supplementary Fig. 5j). These results indicate that unidirectional transcription polarity within the Igh constant region orchestrates 3′RR-mediated transcriptional activation, cohesin accumulation, and chromatin interaction to regulate loop extrusion- and diffusion-mediated end-joining of CSR.
Chromatin domains of Igh affect end-joining of CSR
To investigate the joining features of AID-initiated breaks located in different chromatin domains, we placed 3′RR and its associated Cα unit outside the Igh domain by inverting the Cα-3′CBEs region in αInv cells to generate α-3CBEInv cells, in which 3′RR and Cα are located in a different domain from the donor Cμ unit (Fig. 5a, Supplementary Fig. 6a, b). Remarkably, α-3CBEInv cells significantly increased inversional Sμ–Sα junctions from 8% to 53% without affecting MHs (Fig. 5b–d, Supplementary Fig. 6c). In contrast to the predominantly Sμ–Sα synapsis for robust deletional Sα CSR in αInv cells (Fig. 5e, Supplementary Fig. 6d), Cα unit in α-3CBEInv cells was isolated by the new topologically-associated domain (TAD) formed by the inverted 3′CBEs and downstream CBE, which prevented Sμ–Sα synapsis even Cα unit was highly transcribed (Fig. 5f–j, Supplementary Fig. 6e). These data indicate that the donor Cμ and acceptor CH located in different TADs undergo diffusion-mediated orientation-unbiased end-joining of CSR.
a Schematic of the Igh locus from the V(D)J exon to downstream CBE outside the Igh domain in αInv cells and an illustration of the generation of α−3CBEInv cells. b CSR-HTGTS-seq analysis of break joining between 5′Sμ and acceptor Sα in αInv and α−3CBEInv cells stimulated with αCD40/IL4/TGFβ for 72 h. Junctions within the Igh constant region are plotted at a 2.25 kb bin size. Data are presented as mean ± s.e.m. from three independent experiments. c Joining features of Sμ–Sα junctions in αInv and α−3CBEInv cells. Data are presented as mean ± s.e.m. from three independent experiments. d Bar graph shows the inversional joining of Sμ–Sα junctions in αInv and α−3CBEInv cells. Data are presented as mean ± s.e.m. from three independent experiments. P-values were calculated via an unpaired two-tailed Student’s t-test. e 3C-HTGTS with 3′CBEs, iEμ/Iμ or 3′RR bait (blue asterisks), and Pro-seq to show the chromatin interaction and transcription status of the indicated region from αInv cells. f 3C-HTGTS with 3′CBEs, iEμ/Iμ or 3′RR bait (blue asterisks), and Pro-seq to show the chromatin interaction and transcription status of the indicated region from α−3CBEInv cells. Relative interaction frequency between 3′CBEs (g), iEμ/Iμ (h), or 3′RR (i) bait with the indicated regions in αInv and α−3CBEInv cells. Data are presented as mean ± s.e.m. from three independent experiments. P-values were calculated via paired two-tailed Student’s t-test. j Relative transcription activity of Cμ and Cα unit in αInv and α-3CBEInv cells. Data are presented as mean ± s.e.m. from three independent experiments. P-values were calculated via paired two-tailed Student’s t-test. Source data are provided as a Source Data file.
Given that the above CSR occurs between classical donor Sμ and acceptor S regions, we also wanted to investigate the general joining mechanism between AID-targeted classical S region and off-targets. Within the Igh domain, chromatin loop extrusion-mediated 3′RR scanning activates Cα for CSR in CH12F3PO cells and scans upstream CHs to activate their CSR activity upon the elimination of Cα competition in IαΔ cells9. In α-3CBEInv cells, 3′RR activates Cα for CSR in a new TAD, and we further deleted the Cα unit in α-3CBEInv cells to generate α-3CBEInv-αΔ cells (Fig. 6a, Supplementary Fig. 7a, b). The data indicated that 3′RR activated an ectopic S (eS) region immediately downstream of the deleted Cα unit and donor Sμ recombined, without orientation bias, to this eS region in α-3CBEInv-αΔ cells (Fig. 6b–g). This eS region indicated a significantly lower GC content than classical Sμ and Sα regions (Supplementary Fig. 7c), corresponding to the lower AID targeting frequency (Fig. 6b, c). These data indicate that AID-initiated on- and off-targets located in different domains promote diffusion-mediated orientation-unbiased recombination.
a Schematic of the Igh locus from the V(D)J exon to downstream CBE outside the Igh domain in α-3CBEInv cells and an illustration of the generation of α−3CBEInv-αΔ cells. b CSR-HTGTS-seq and Pro-seq analyses to show the CSR junctions and transcription status of the indicated region from α−3CBEInv cells. CSR junctions are plotted at a 3.55 kb bin size. Data are presented as mean ± s.e.m. from three independent experiments. c CSR-HTGTS-seq and Pro-seq analyses to show the CSR junctions and transcription status of the indicated region from α−3CBEInv-αΔ cells. CSR junctions are plotted at a 3.55 kb bin size. Data are presented as mean ± s.e.m. from three independent experiments. CSR-HTGTS-seq and Pro-seq to show the CSR junctions and transcription status around the eS region in α−3CBEInv (d) and α−3CBEInv-αΔ (e) cells. f Bar graph shows the deletional and inversional joining of eS region junctions in α−3CBEInv-αΔ cells. Data are presented as mean ± s.e.m. from six independent experiments. P-values were calculated via an unpaired two-tailed Student’s t-test. g Bar graph shows the transcription activity of the eS region in α−3CBEInv and α−3CBEInv-αΔ cells. Data are presented as mean ± s.e.m. from three independent experiments. P-values were calculated via an unpaired two-tailed Student’s t-test. Source data are provided as a Source Data file.
Discussion
In this study, through analysing chromatin features of constant region across different species (Fig. 7a) and constructing a series of Igh constant regions to recapitulating productive CSR in jawed vertebrates, we have revealed some evolutionary adaptations of the constant region facilitating CSR in evolutionary species, including that transcription polarity, chromatin distance, and TAD of Igh, termed as switch topological configuration (STC), plays critical roles in regulating orientation-specific joining of AID-initiated CSR for generating productive antibodies (Fig. 7b, c). Some species, such as Mus musculus and Homo sapiens, have the same transcription orientation between acceptor CHs with donor Cμ within the Igh locus and undergo orientation-biased deletional joining of CSR, leading to efficient generation of productive antibodies22,39. Chromatin loop extrusion can promote long-range S-S synapsis for predominantly orientation-biased deletional joining-mediated productive CSR in some species, such as mice (Fig. 7b). In contrast, some species have opposite transcribed CHs, short-distance CHs, as well as CHs located in different TADs, which strongly activate short-range diffusion-mediated random joining of AID-initiated breaks, leading to more inversional joining-mediated productive CSR (Fig. 7c).
a Chromatin features of the Igh constant region in different species (such as Ginglymostoma cirratum, Danio rerio, Xenopus laevis, Alligator sinensis, Gallus gallus, Mus musculus, and Homo sapiens) throughout evolution. The columns with different colors indicate different CH units. The green arrows indicate the transcription orientation. b long-distance CHs under the same transcriptional orientation within the Igh domain promote orientation-biased deletional joining of CSR for generating productive antibodies in some species, such as Mus musculus. Chromatin loop extrusion might promote this orientation-biased end joining for highly efficient CSR. c Opposite transcribed CHs, short-distance CHs, or different chromatin domain-isolated CHs promote inversional joining-mediated productive CSR in some species, such as Gallus gallus. Short-range diffusion might promote this relatively less-efficient CSR. The red bricks indicate the anchors for mediating TAD formation. The blue circles indicate the cohesin rings. The light red circle indicates the CSR center. The pink circle indicates the short-range diffusion area. The black arrows indicate the transcription orientation. The blue arrows indicate chromatin loop extrusion-mediated pulling force. The dotted arrows indicate the potential joining events between donor Sμ and acceptor Sx breaks. The key elements, such as donor Sμ (red oval), activated acceptor Sx (green oval), iEμ (atrovirens oval), and 3′RR (blue oval), are also indicated.
Cohesin-mediated chromatin loop extrusion has been proposed to explain the packaging of large amounts of chromatin DNA into TADs within the nucleus6,7. Cohesin can be loaded onto chromatin and promote the formation of chromatin loop domains when cohesin reaches a pair of convergently oriented CBEs bound by CTCF6,7,43. In addition to CTCF, some factors (such as transcription factors, MCM, and dCas9)44,45,46 and chromatin structures (such as R-loops and replication forks)47,48 have also been suggested as potential dynamic impediments during chromatin loop extrusion-mediated physiological processes. Our work indicates that the promoter polarity not only determines the transcriptional orientation but also affects the chromatin loop extrusion and related DNA recombination process. Interestingly, elimination of chromatin loop extrusion only modestly affects gene expression, and chromatin loop extrusion has been found to be required for long-range enhancer action49,50,51. And previous studies also suggest that chromatin loop extrusion plays a more dominant role in long-distance D to JH recombination, and short-range diffusion dominates the DQ52 to JH recombination within V(D)J recombination center46. Moreover, a recent study suggests that chromatin loop extrusion brings Vκ and Jκ into close proximity around Cer and Sis elements, and local diffusion promotes the final Vκ to Jκ recombination within the Igκ recombination center52. Our work suggests that chromatin loop extrusion plays a more dominant role in promoting orientation-specific end-joining of long-range DNA recombination. Although inversion of the individual CH unit, which has a long distance from the donor Cμ unit, increases the inversion end-joining for various degrees, deletional end-joining is still the dominant outcome due to the long distance. Thus, the intrinsic chromatin features such as transcriptional orientation, chromatin distance, and chromatin domains orchestrate chromatin loop extrusion- and diffusion-mediated DNA damage repair to balance repair efficiency and outcomes.
Early studies exhibited that numerous DNA repair factors participate in promoting deletional joining-mediated productive CSR by regulating DSB processing and repair pathways22,23,24,25,26,27,28,29,30,31,32,33,34,35,36,37,38, but the roles of these factors still fail to fully account for the regulation of orientation-specific joining mechanisms. Chromatin loop extrusion-mediated CSR model greatly improves the understanding of CSR, including the formation of dynamic CSR centers, transcription activation of acceptor CH units, synapsis of acceptor CH and donor Cμ units, and orientation-biased deletional joining of CSR in mice9,10. The chromatin loop extrusion-mediated predominantly deletional joining of CSR junction in mice extends the end-joining regulation from DNA damage repair-related trans-factors to dynamic chromatin regulation9,10. By further revealing the chromatin features of the Igh constant region across different species, we reveal that STC-related intrinsic chromatin features of the Igh locus fulfil another layer of regulation in orientation-specific joining of AID-initiated CSR for generating productive antibodies. These detailed STC-mediated mechanisms of orientation-specific joining of DSBs not only deepen the understanding of productive CSR in different species throughout evolution and repair of pathogenic DSB genome-wide, but also provide new insights for antibody-directed evolution.
Methods
Experimental procedures
No statistical methods were used to predetermine sample size, and experiments were not randomized in this study. Investigators were not blinded to allocation during experiments and outcome assessment.
Cell culture
CH12F3 cells were cultured in medium R15 and stimulated with αCD40 (1 μg/ml, eBioscience), IL4 (20 ng/ml, PeproTech), and TGFβ (0.5 ng/ml, R&D Systems) for 72 h. CSR-HTGTS-seq was performed in AID-proficient CH12F3 cells. Pro-seq, 3C-HTGTS, and ChIP-seq were performed in AID-deficient CH12F3 cells to obviate confounding effects of CSR-related genomic rearrangements.
Generation of mutant CH12F3 cell lines
gRNA design and vector construction: a Cas9/gRNA approach was employed to generate all the various mutant strains used in this study, as described previously9. gRNAs were designed using the E-CRISP web tool (v5.4) with default parameters for SpCas9, targeting the Mus musculus genome. Candidate gRNAs were selected based on high predicted on-target efficiency scores (>80) and the absence of predicted off-target sites (with ≤3 mismatches) within protein-coding exons. gRNAs were individually cloned into the BbsI and BsaⅠ sites of the p332 vector following the standard protocol. The specific sequences for all gRNAs used are listed in Supplementary data.
Nucleofection and clonal selection: to generate mutant clones, 2 × 10⁶ CH12F3 cells were nucleofected with 2 µg of p332-based Cas9/gRNA plasmids containing two gRNA sequences using the Lonza 4D-Nucleofector™ system (Solution SF, program CA-137). Twenty-four hours post-nucleofection, single-cell subcloning was performed by fluorescence-activated cell sorting (FACS) on a BD FACS Aria™ SORP cell sorter to isolate one cell into one well of 96-well plates. Genomic DNA from individual clones was subsequently extracted for PCR-based genotyping and sequencing to confirm the desired genetic modification. Two pairs of genotyping primers were designed on either side of the two break sites for each mutant. Different combinations of these four primers can be used for PCR genotyping to detect WT bands, deletion bands, and inversion bands. The detailed primer pairs for genotyping were described in the corresponding figure legends. Based on the PCR genotyping and sequencing results, we can only detect WT bands for the control cells and positive bands (deletion or inversion) for the mutant clones. Detailed primers for selecting various mutant CH12F3 cell lines are listed in Supplementary data.
CH12F3PO cells were generated by deleting the entire non-productive Igh locus from upstream of the first VH to downstream of 3′CBEs in CH12F3NCΔ cells9 and confirmed by PCR genotyping and sequencing. γ3-αInv cells were generated by inverting the six acceptor CHs region from upstream of Cγ3 to downstream of Cα in CH12F3PO cells and confirmed by PCR genotyping and sequencing. αInv cells were generated by inverting the Cα unit in CH12F3PO cells and confirmed by PCR genotyping and sequencing. We generated the IαΔ cells by deleting the Iα promoter in CH12F3 cells in the previous study9. γ2aInv cells were generated by inverting the Cγ2a unit in IαΔ cells and confirmed by PCR genotyping and sequencing. γ1Inv cells were generated by inverting the Cγ1 unit in IαΔ cells and confirmed by PCR genotyping and sequencing. γ3-αInv-αInv cells were generated by inverting the Cα unit in γ3-αInv cells and confirmed by PCR genotyping and sequencing. γ3Inv cells were generated by inverting the Cγ3 unit in IαΔ cells and confirmed by PCR genotyping and sequencing. γ3-αInv-γ3Inv cells were generated by inverting the Cγ3 unit in γ3-αInv cells and confirmed by PCR genotyping and sequencing. γ3IRD cells were generated by deleting the intergenic region between Cγ3 and Cμ to position Cγ3 ~ 17 kb away from Cμ in IαΔ cells and confirmed by PCR genotyping and sequencing. Iγ3-FS-Cα cells were generated by fusing the entire Sγ3 with Sα in CH12F3PO cells and confirmed by PCR genotyping and sequencing. Iγ3-FS-Iα cells were generated by fusing the entire Sγ3 with Sα in αInv cells and confirmed by PCR genotyping and sequencing. Iγ3-PS-Cα cells were generated by fusing the partial Sγ3 with Sα in CH12F3PO cells and confirmed by PCR genotyping and sequencing. Iγ3-PS-Iα cells were generated by fusing the partial Sγ3 with Sα in αInv cells and confirmed by PCR genotyping and sequencing. α-3CBEInv cells were generated by inverting the α-3′CBEs region in αInv cells and confirmed by PCR genotyping and sequencing. α-3CBEInv-αΔ cells were generated by deleting the Cα unit in α-3CBEInv cells and confirmed by PCR genotyping and sequencing. The corresponding AID-deficient cells were generated by deleting the Aicda gene from the above cells and confirmed by PCR genotyping and Western blotting. At least two independent clones were obtained for each derivative mutant genotype.
Chromatin feature analyses of the Igh locus across species
Reference genome assemblies used in this paper included Danio rerio (GRCz10), Xenopus tropicalis (UCB_Xtro_10.0), Gallus gallus (GRCg7w), and Mus musculus (mm9). Alligator sinensis was derived from Pan et al.53. Splicing regions in Mus musculus have been well-annotated. For other species, we aligned the constant region sequences obtained from IMGT54. The splicing regions were defined as 4 kb upstream of the first exon of the constant regions. Splicing region sequences were extracted by bedtools (v2.31.1), and the GC contents were calculated with seqkit (v2.8.2). Background GC content was derived genome-wide by sliding 4-kb windows.
Reads of RNA-seq from all species53,55,56,57,58 were aligned to their respective reference genomes using STAR (v2.7.11a). Strand-specific bigwig files were generated with deeptools (v3.5.3), and strand orientation was defined from the alignment described above.
All Hi-C data53,59,60,61 were processed with Hi-C-Pro (v3.1.0) under default settings, except the LIGATION_SITE was species-specific, to generate valid interaction pair files. The output pair files were then converted to .hic files using the hicpro2juicebox utility. Hi-C files were also converted to .cool files using Hicexplorer (v3.7.2). Except for Gallus gallus, all species were analyzed at 25 kb resolution. Gallus gallus was analyzed at 5 kb resolution because the Igh locus is located on the microchromosome. Insulation scores and TADs were calculated with cooltools (v0.7.1) with a window size of 7 bins. All visualizations were generated by pyGenomeTracks (v3.8).
CSR-HTGTS-seq library preparation and data analysis
CSR-HTGTS-seq libraries with 5′Sμ bait were prepared from different CH12F3 mutants stimulated with αCD40/IL4/TGFβ for 72 h as described previously22. Briefly, 25 μg gDNA from αCD40/IL4/TGFβ-stimulated CH12F3 cells was sonicated on the Covaris M220 sonicator. The sonicated DNA segments were amplified by LAM-PCR with biotinylated 5′Sμ primer. The amplified biotin-labeled LAM-PCR products were enriched with streptavidin C1 beads (Thermo Fisher Scientific, #65001) for 4 h at room temperature, followed by adaptor ligation with the following PCR program: 25 °C 1 h, 22 °C 3 h, 16 °C overnight. The adaptor-ligated products were subjected to nested-PCR with barcode primers and followed by tag-PCR with Illumina platform-matched P5-I5 and P7-I7 primers. 500–1000 bp tag-PCR products were selected by separation on 1% TAE gel. CSR-HTGTS-seq libraries were sequenced by paired-end 150 bp on a NovaSeq 6000 (Illumina) sequencer.
Libraries were processed via the published pipeline and mapped against the mm9 genome or the corresponding modified genome22. Data were analyzed and plotted after removing the duplicates as described22. Each experiment was repeated at least three times from at least two independent clones. To plot the MH pattern, direct joins (MH = 0) and junctions with MH were pooled and sorted by length, and the number of junctions with indicated length of MH were counted and calculated as a percentage of the total number of junctions mapped to the region of interest. Primers used for CSR-HTGTS-seq are listed in Supplementary Table 1.
3C-HTGTS library preparation and data analysis
3C-HTGTS analyses were performed as previously described on AID-/- CH12F3 cells9 stimulated with αCD40/IL4/TGFβ for 72 h. Briefly, 10 million cells were crosslinked with 2% formaldehyde for 10 min at room temperature and quenched with glycine at a final concentration of 125 mM. Then, the crosslinked cells were lysed in the 3C lysis buffer, and nuclei were digested with NlaIII enzyme (NEB, R0125) at 37 °C overnight. The digested samples were ligated by T4 DNA ligase (NEB, M1801) at room temperature for at least 6 h. The ligated products were de-crosslinked with Proteinase K (Roche, #03115852001) at 56 °C overnight, and the 3C templates were purified by phenol/chloroform. The 3 C templates were used for making 3C-HTGTS with different baits, including the iEμ-Iμ, 3′RR(HS4), or 3′CBEs locale. Besides the DNA templates and baits, the 3C-HTGTS library preparation procedures were similar to CSR-HTGTS-seq described above.
The 3C-HTGTS libraries were then sequenced by paired-end 150 bp sequencing on a NovaSeq 6000 (Illumina) sequencer, and data were processed as previously described9. Each experiment was repeated from at least two independent clones. Before plotting the data for comparison, libraries were size-normalized to the total junctions of the smallest library in the set of libraries for comparison. For statistical analyses, we counted the number of junctions within the indicated bait-interacting locals for each sample. For bar graph presentations in Figs. 1 and 5, the junction number from control samples was normalized to represent 100%, and relative experimental values are listed as a percentage of the control values. Primers used for 3C-HTGTS are listed in Supplementary Table 1.
Pro-seq library preparation and data analysis
Pro-seq libraries were prepared as described previously62 from AID-/- CH12F3 cells stimulated with αCD40/IL4/TGFβ for 72 h. Briefly, 10 million cells were collected and permeabilized with permeabilization buffer. The permeabilized cells were resuspended in 100 μl of storage buffer for nuclear run-on with 2× run-on mix at 37 °C for 5 min. RNA was extracted using Trizol and followed by hydrolysis. The RNA was incubated with streptavidin C1 beads (Thermo Fisher Scientific, #65001), and the enriched run-on samples were incubated with hydroxyl repair with T4 PNK (NEB, M0201S) and RppH (NEB, M0356S), followed by ligating the 5′ and 3′ RNA adaptor. RT-PCR was performed from the adaptor-ligated RNA to obtain cDNA. The cDNA was subjected to making Pro-seq libraries by two rounds of PCR with barcode primers. 200–500 bp products from the first round of PCR were subjected to the second round of PCR, with the number of PCR cycles determined by test PCR amplification. The second round of PCR products was size-selected by SPRIselect beads (Beckman Coulter, B23318). Pro-seq libraries were sequenced via single-end 50 bp sequencing on a HiSeq2500 (Illumina) sequencer or paired-end 150 bp sequencing on a NovaSeq 6000 (Illumina) sequencer. The processed reads were mapped to the mm9 genome or the corresponding modified genome using the Bowtie2 software. The read coverage for each sample was normalized to a coverage of 10 million 100nt reads for display. Each experiment was repeated three times from at least two independent clones.
ChIP-seq library preparation and data analysis
ChIP-seq was carried out in AID-deficient cells stimulated with αCD40/IL4/TGFβ for 72 h based on a prior protocol63 with some modifications. Briefly, 20 million cells were collected and fixed with 1% formaldehyde for 10 min at room temperature, and quenched with glycine at a final concentration of 125 mM. Cells were lysed and sonicated with a Covaris M220 sonicator for 5–8 min until the size of most fragments was in the range of 200–700 bp. The fragmented chromatin was incubated with Rad21 antibody (ab992, Abcam) overnight, followed by incubating with magnetic protein A (Invitrogen, 10002D) for 4 h at 4 °C. The enriched chromatin was used for tagmentation, followed by decrosslinking and elution. Purified DNA was used to make ChIP-seq libraries. The amplified libraries were purified by SPRIselect beads (Beckman Coulter, B23318) and were sequenced via paired-end 150 bp sequencing on a NovaSeq 6000 (Illumina) sequencer. The processed reads were mapped to the modified genome using the Bowtie2 software. The uniquely aligned reads were retained, while duplicate reads and those located in blacklist regions were excluded prior to further analyses. The read coverage for each sample was normalized by RPKM.
Quantification and statistical analysis
Statistical analyses for CSR-HTGTS-seq, 3C-HTGTS, and Pro-seq between two samples were performed via two-tailed, unpaired, or paired Student’s t-test. At least three biological repeats were done for each statistical analysis. p < 0.05 is considered significant. P-value shown in the bar graphs in the main and Supplementary Figs. The statistical analyses were also described in the corresponding Figure legends and Supplementary Fig. Legends. Unless indicated otherwise, all agarose gel electrophoresis images are representative of at least two independent experiments, with uncropped gels shown in the Source Data file.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
CSR-HTGTS-seq, 3C-HTGTS, Pro-seq, and ChIP-seq sequencing data analyzed here have been deposited in the GEO database. The GEO accession number for CSR-HTGTS-seq is GSE299392. The GEO accession number for 3C-HTGTS is GSE299499. The GEO accession number for Pro-seq is GSE299975. The GEO accession number for ChIP-seq is GSE299391. All other data can be found in the Supplementary Data of this paper or in the Source Data. This includes all uncropped gel electrophoresis images and data shown in graphs throughout the manuscript, including the Supplementary Figs. All data are available from the authors upon reasonable request. Source data are provided with this paper.
Code availability
V(D)J-HTGTS-seq, CSR-HTGTS-seq, and 3C-HTGTS data was processed through published pipelines (http://robinmeyers.github.io/transloc_pipeline/). Pro-Seq was aligned to either the mm9 genome or modified genomes with bowtie2 (v.2.4.4) (http://bowtie-bio.sourceforge.net/bowtie2/index.shtml), processed by samtools (v.1.15.1) (https://sourceforge.net/projects/samtools/fles/samtools/1.15.1/), and generated graph files via the RseqQ (v.5.0.1) (https://rseqc.sourceforge.net/) package. ChIP-Seq was aligned to the mm9-derived modified genomes with bowtie2 (v.2.4.4), processed by samtools (v.1.15.1), generated bigwig files via Deeptools (v.3.5.1) (https://github.com/deeptools/deepTools), and generated peak files via MACS2 (v.2.2.7.1) (https://github.com/taoliu/MACS/).
References
Bonilla, F. A. & Oettgen, H. C. Adaptive immunity. J. Allergy Clin. Immunol. 125, S33–S40 (2010).
Teng, G. & Schatz, D. G. Regulation and evolution of the RAG recombinase. Adv. Immunol. 128, 1–39 (2015).
Alt, F. W., Zhang, Y., Meng, F. L., Guo, C. & Schwer, B. Mechanisms of programmed DNA lesions and genomic instability in the immune system. Cell 152, 417–429 (2013).
Methot, S. P. & Di Noia, J. M. Molecular mechanisms of somatic hypermutation and class switch recombination. Adv. Immunol. 133, 37–87 (2017).
Oudinet, C., Braikia, F. Z., Dauba, A. & Khamlichi, A. A. Mechanism and regulation of class switch recombination by IgH transcriptional control elements. Adv. Immunol. 147, 89–137 (2020).
Fudenberg, G. et al. Formation of chromosomal domains by loop extrusion. Cell Rep. 15, 2038–2049 (2016).
Sanborn, A. L. et al. Chromatin extrusion explains key features of loop and domain formation in wild-type and engineered genomes. Proc. Natl. Acad. Sci. USA 112, E6456–E6465 (2015).
Bruzeau, C., Moreau, J., Le Noir, S. & Pinaud, E. Panorama of stepwise involvement of the IgH 3’ regulatory region in murine B cells. Adv. Immunol. 149, 95–114 (2021).
Zhang, X. et al. Fundamental roles of chromatin loop extrusion in antibody class switching. Nature 575, 385–389 (2019).
Zhang, Y., Zhang, X., Dai, H. Q., Hu, H. & Alt, F. W. The role of chromatin loop extrusion in antibody diversification. Nat. Rev. Immunol. 22, 550–566 (2022).
Muramatsu, M. et al. Class switch recombination and hypermutation require activation-induced cytidine deaminase (AID), a potential RNA editing enzyme. Cell 102, 553–563 (2000).
Feng, Y., Seija, N., Di Noia, J. M. & Martin, A. AID in antibody diversification: there and back again. Trends Immunol. 42, 89 (2021).
Luo, S., Qiao, R. & Zhang, X. DNA damage response and repair in adaptive immunity. Front. Cell Dev. Biol. 10, 884873 (2022).
Saha, T., Sundaravinayagam, D. & Di Virgilio, M. Charting a DNA repair roadmap for immunoglobulin class switch recombination. Trends Biochem. Sci. 46, 184–199 (2021).
Yu, K. AID function in somatic hypermutation and class switch recombination. Acta Biochim. Biophys. Sin. 54, 759–766 (2022).
Garrett, F. E. et al. Chromatin architecture near a potential 3’ end of the igh locus involves modular regulation of histone modifications during B-Cell development and in vivo occupancy at CTCF sites. Mol. Cell Biol. 25, 1511–1525 (2005).
Zhang, X., Yoon, H. S., Chapdelaine-Williams, A. M., Kyritsis, N. & Alt, F. W. Physiological role of the 3’IgH CBEs super-anchor in antibody class switching. Proc. Natl. Acad. Sci. USA https://doi.org/10.1073/pnas.2024392118 (2021).
Yu, K. An insulator that regulates chromatin extrusion and class switch recombination. Proc. Natl. Acad. Sci. USA https://doi.org/10.1073/pnas.2026399118 (2021).
Chiarle, R. et al. Genome-wide translocation sequencing reveals mechanisms of chromosome breaks and rearrangements in B cells. Cell 147, 107–119 (2011).
Wei, P. C. et al. Long neural genes harbor recurrent DNA break clusters in neural stem/progenitor cells. Cell 164, 644–655 (2016).
Frock, R. L. et al. Genome-wide detection of DNA double-stranded breaks induced by engineered nucleases. Nat. Biotechnol. 33, 179–186 (2015).
Dong, J. et al. Orientation-specific joining of AID-initiated DNA breaks promotes antibody class switching. Nature 525, 134–139 (2015).
Reina-San-Martin, B., Chen, H. T., Nussenzweig, A. & Nussenzweig, M. C. ATM is required for efficient recombination between immunoglobulin switch regions. J. Exp. Med. 200, 1103–1110 (2004).
Lumsden, J. M. et al. Immunoglobulin class switch recombination is impaired in Atm-deficient mice. J. Exp. Med. 200, 1111–1121 (2004).
Panchakshari, R. A. et al. DNA double-strand break response factors influence end-joining features of IgH class switch and general translocation junctions. Proc. Natl. Acad. Sci. USA 115, 762–767 (2018).
Reina-San-Martin, B. et al. H2AX is required for recombination between immunoglobulin switch regions but not for intra-switch region recombination or somatic hypermutation. J. Exp. Med. 197, 1767–1778 (2003).
Bothmer, A. et al. 53BP1 regulates DNA resection and the choice between classical and alternative end joining during class switch recombination. J. Exp. Med. 207, 855–865 (2010).
Bothmer, A. et al. Regulation of DNA end joining, resection, and immunoglobulin class switch recombination by 53BP1. Mol. Cell 42, 319–329 (2011).
Chapman, J. R. et al. RIF1 is essential for 53BP1-dependent nonhomologous end joining and suppression of DNA double-strand break resection. Mol. Cell 49, 858–871 (2013).
Di Virgilio, M. et al. Rif1 prevents resection of DNA breaks and promotes immunoglobulin class switching. Science 339, 711–715 (2013).
Xu, G. et al. REV7 counteracts DNA double-strand break resection and affects PARP inhibition. Nature 521, 541–544 (2015).
Boersma, V. et al. MAD2L2 controls DNA repair at telomeres and DNA breaks by inhibiting 5’ end resection. Nature 521, 537–540 (2015).
Ghezraoui, H. et al. 53BP1 cooperation with the REV7-shieldin complex underpins DNA structure-specific NHEJ. Nature 560, 122–127 (2018).
Mirman, Z. et al. 53BP1-RIF1-shieldin counteracts DSB resection through CST- and Polalpha-dependent fill-in. Nature 560, 112–116 (2018).
Noordermeer, S. M. et al. The shieldin complex mediates 53BP1-dependent DNA repair. Nature 560, 117–121 (2018).
Dev, H. et al. Shieldin complex promotes DNA end-joining and counters homologous recombination in BRCA1-null cells. Nat. Cell Biol. 20, 954–965 (2018).
Vincendeau, E. et al. SHLD1 is dispensable for 53BP1-dependent V(D)J recombination but critical for productive class switch recombination. Nat. Commun. 13, 3707 (2022).
Han, L. & Yu, K. Altered kinetics of nonhomologous end joining and class switch recombination in ligase IV-deficient B cells. J. Exp. Med. 205, 2745–2753 (2008).
Du, L. et al. Orientation regulation of class-switch recombination in human B cells. J. Immunol. 213, 1093–1104 (2024).
Kitao, H., Arakawa, H., Yamagishi, H. & Shimizu, A. Chicken immunoglobulin mu-chain gene: germline organization and tandem repeats characteristic of class switch recombination. Immunol. Lett. 52, 99–104 (1996).
Sun, Y., Huang, T., Hammarstrom, L. & Zhao, Y. The immunoglobulins: new insights, implications, and applications. Annu. Rev. Anim. Biosci. 8, 145–169 (2020).
Nakamura, M. et al. High frequency class switching of an IgM+ B lymphoma clone CH12F3 to IgA+ cells. Int. Immunol. 8, 193–201 (1996).
Fudenberg, G., Abdennur, N., Imakaev, M., Goloborodko, A. & Mirny, L. A. Emerging evidence of chromosome folding by loop extrusion. Cold Spring Harb. Symp. Quant. Biol. 82, 45–55 (2017).
Stadhouders, R. et al. Transcription factors orchestrate dynamic interplay between genome topology and gene regulation during cell reprogramming. Nat. Genet. 50, 238–249 (2018).
Dequeker, B. J. H. et al. MCM complexes are barriers that restrict cohesin-mediated loop extrusion. Nature 606, 197–203 (2022).
Zhang, Y. et al. The fundamental role of chromatin loop extrusion in physiological V(D)J recombination. Nature 573, 600–604 (2019).
Zhang, H. et al. CTCF and R-loops are boundaries of cohesin-mediated DNA looping. Mol. Cell 83, 2856–2871.e2858 (2023).
Jeppsson, K. et al. Cohesin-dependent chromosome loop extrusion is limited by transcription and stalled replication forks. Sci. Adv. 8, eabn7063 (2022).
Rao, S. S. P. et al. Cohesin loss eliminates all loop domains. Cell 171, 305–320.e324 (2017).
Kane, L. et al. Cohesin is required for long-range enhancer action at the locus. Nat. Struct. Mol. Biol. 29, 891–897 (2022).
Rinzema, N. J. et al. Building regulatory landscapes reveals that an enhancer can recruit cohesin to create contact domains, engage CTCF sites and activate distant genes. Nat. Struct. Mol. Biol. 29, 563–574 (2022).
Zhang, Y. W. et al. Molecular basis for differential Igk versus Igh V(D)J joining mechanisms. Nature https://doi.org/10.1038/s41586-024-07477-y (2024).
Pan, T. et al. Genomic insights and the conservation potential of captive breeding: the case of Chinese alligator. Sci. Adv. 11, eadm7980 (2025).
Manso, T. et al. IMGT® databases, related tools and web resources through three main axes of research and development. Nucleic Acids Res. 50, D1262–D1272 (2022).
Yang, H. et al. A map of cis-regulatory elements and 3D genome structures in zebrafish. Nature 588, 337–343 (2020).
Buisine, N. et al. Xenopus tropicalis genome re-scaffolding and re-annotation reach the resolution required for in vivo ChIA-PET analysis. PLoS ONE 10, e0137526 (2015).
Nuthalapati, N. K. et al. Transcriptomic analysis of early B-cell development in the chicken embryo. Poult. Sci. 98, 5342–5354 (2019).
He, P. et al. The changing mouse embryo transcriptome at whole tissue and single-cell resolution. Nature 583, 760–76 (2020).
Hellsten, U. et al. The genome of the Western clawed frog Xenopus tropicalis. Science 328, 633–636 (2010).
Rhie, A. et al. Towards complete and error-free genome assemblies of all vertebrate species. Nature 592, 737–746 (2021).
Rivas, M. A. et al. Smc3 dosage regulates B cell transit through germinal centers and restricts their malignant transformation. Nat. Immunol. 22, 240–U278 (2021).
Mahat, D. B. et al. Base-pair-resolution genome-wide mapping of active RNA polymerases using precision nuclear run-on (PRO-seq). Nat. Protoc. 11, 1455–1476 (2016).
Schmidl, C., Rendeiro, A. F., Sheffield, N. C. & Bock, C. ChIPmentation: fast, robust, low-input ChIP-seq for histones and transcription factors. Nat. Methods 12, 963–965 (2015).
Acknowledgements
We thank Drs. Frederick W. Alt and Tasuku Honjo for providing reagents. We thank Drs. Xiaoliang Sunney Xie, Zemin Zhang, Fuchou Tang, Guoqiang Li, Jiazhi Hu, and Hsiang-Ying Lee for some technical support. We thank all Zhang lab members for helpful discussions. This work was supported by the National Key R&D Program of China (2022YFA1305000 to X.Z.), the National Natural Science Foundation of China (32270930 to X.Z.), the Beijing Natural Science Foundation (5232007 to X.Z.), and the Beijing Advanced Innovation Center for Genomics at Peking University.
Author information
Authors and Affiliations
Contributions
S.L. (Sha Luo), R.Q., and X.Z. designed the study; S.L. (Sha Luo), R.Q., X.S., S.L. (Shuchan Li), Z.Y., and A.M performed the experiments; H.Z., and L.Y. designed some of the bioinformatics pipelines; S.L. (Sha Luo), R.Q., H.Z., and X.Z. analysed the data; S.L. (Sha Luo), R.Q., H.Z., L.Y., and X.Z. prepared the figures with some input from Y.C. and Y-F.Z.; S.L. (Sha Luo), R.Q., H.Z., L.Y., and X.Z. wrote the manuscript. X.Z. managed and supervised the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Luo, S., Qiao, R., Zha, H. et al. Chromatin-intrinsic mechanisms determine orientation-specific class switch recombination. Nat Commun 17, 3319 (2026). https://doi.org/10.1038/s41467-026-70031-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-70031-z