Abstract
Traumatic brain injury (TBI) is a complex condition in which multiple pathophysiological mechanisms influence the course of the disease. After the initial mechanical impact, neuroinflammatory reactions of glial cells along with infiltrating peripheral immune cells determine the overall clinical outcome. However, these secondary processes and their molecular determinants promoting either beneficial or detrimental consequences are not well-defined. Here, we show that TBI-mediated NF-κB activation in astrocytes impairs their homeostatic functions, amplifies the post-traumatic neuroimmune response and disturbs the multicellular CNS scar development in a male mouse model of TBI. Our results further demonstrate a specific deficit in the formation of the glial limitans border and establish that paracrine signaling pathways induced by NF-κB-activated astrocytes can prevent a beneficial restoration of the CNS integrity after TBI. These findings enhance our understanding on the NF-κB-mediated post-traumatic pathophysiology and provide information on future targeted therapies to improve TBI outcome.
Similar content being viewed by others
Background
Traumatic brain injury (TBI) is a leading cause of death and represents a major health burden in survivors. TBI can initiate the development of long-term neurocognitive and neuropsychiatric morbidities and has been associated with an increased risk of late-onset neurodegeneration1,2. After the initial mechanical insult directly causing the primary brain lesion, biochemical and neuroimmune reactions set up a secondary injury phase persisting for days or even months and years. Together with other confounding issues, such as age, sex, and comorbidities, these multi-dimensional secondary processes determine the overall clinical outcome with or without long-term consequences and represent a challenge for the development of effective therapies3,4. Therefore, it has to be clarified whether individual post-traumatic reactions can be guided in a way that allows efficient tissue regeneration while suppressing chronic neurodetrimental inflammation and degeneration.
Astrocytes, the most frequent glial cells in the mammalian brain, execute essential tasks in maintaining CNS homeostasis. Upon injury, astrocytes rapidly respond and adapt diverse functional states aiming to restrict tissue damage and promote neural tissue repair5,6. Reactive astrocytes take an integral part in the CNS scar formation process and—depending on the context—can initiate neurotoxic effects upon traumatic insults. Distinct functional polarization states of astrocytes have been observed in stroke, trauma, and various neurodegenerative diseases, such as Alzheimer’s disease. Specific, phenotype-defining signaling pathways and transcriptional regulators, however, are not well-defined7,8,9,10.
The IKK/NF-κB signaling pathway regulates immune and stress responses but also mediates neuroprotection in a context-dependent manner11,12,13. NF-κB-induced neuroinflammation is increasingly recognized as a central pathogenic mechanism in neurodegeneration14,15,16, and recent studies identified NF-κB signaling in astrocytes as a critical module triggering pathogenic reactive states17,18,19,20,21. In TBI, multiple danger signals (DAMPs) and stimuli activating innate immunity22,23 activate NF-κB in both neurons and glial cells. While NF-κB in neurons confers neuroprotection after TBI24, it remains to be tested whether and which NF-κB-regulated processes in glial cells promote a beneficial or detrimental post-traumatic outcome.
Here, we use a male mouse model of TBI to determine the transcriptomic and cellular changes in transgenic mice with NF-κB modulation in astrocytes25 that underwent closed-head, traumatic brain injury (CHI), the most common form of TBI found in human patients26. We identified prominent NF-κB-regulated gene expression programs in astrocytes and microglia cells after TBI, and especially a dynamic NF-κB activation pattern in astrocytes surrounding the insult area was found integral for adequate wound healing.
Results
TBI induces a prominent NF-κB-related gene expression signature
To gain insight into the regulation of gene expression after closed head injury, RNA sequencing was conducted with bulk tissue obtained from the insult area of CHI and sham-treated wildtype mice. We focused on the early secondary injury phase 3 days post-TBI, which is critical for the development of long-lasting consequences. Principal component analysis (PCA) revealed two distinct clusters representing sham-treated and TBI-induced mice, which show strong differential gene expression (Supplementary Fig. 1a, b). GO term analysis revealed multiple biological processes, molecular functions and cellular components (Supplementary Table 1) mainly associated with inflammatory processes, immune and stress responses, as well as extracellular matrix (ECM) organization, which were enriched 3 days post-TBI. The KEGG pathway analysis identified several top pathways enriched, which are associated with IKK/NF-κB signaling (Fig. 1a), including the “NF-kappa B signaling pathway” itself (Fig. 1a), that was further supported by the increased expression of NF-κB pathway components (Supplementary Table 2). Diverse gene expression signatures tightly connected to NF-κB signaling and the regulation of neuroimmune responses like cytokines and chemokines (Fig. 1b), as well as interferon-associated genes (Supplementary Fig. 1c), were significantly induced by TBI. Furthermore, the top-upregulated genes showed a prominent enrichment of genes (11 out of 30) regulated by or related to the IKK/NF-κB signaling pathway (Supplementary Table 3). Importantly, a temporal expression analysis of representative NF-κB-regulated target genes starting from 6 h until day 30 after TBI revealed that those genes (e.g., Tgm1, Lgals3, Spp1, among others) peak between day 3 and 7 and reach basal levels at day 30 post-TBI (Fig. 1c–j), indicating transient NF-κB activation.
a RNA sequencing was performed with RNA samples derived from traumatized bulk tissue (ipsilateral insult area) from TBI and sham-treated wildtype/control mice 3 days after CHI/TBI (3 dpi; n = 7 per group). Enrichment analysis using the pathway database of the Kyoto encyclopedia of genes and genomes (KEGG) shows an upregulation of multiple pathways linked to immune regulation at 3 dpi. Terms additionally associated with IKK/NF-κB signaling are underlined. Statistics: DESeq2 RNA-seq data analysis followed by KEGG enrichment analysis with PANTHER overrepresentation test (Fisher), FDR corrected and raw p < 0.05. KEGG database https://doi.org/10.1093/nar/28.1.27; https://doi.org/10.1002/pro.3715. b Heat map depicting the upregulation of multiple cytokines and chemokines in the insult area of TBI (green) and sham-treated animals (white). c–j Kinetic gene expression analyses of selected genes out of the top 30 DEGs, including NF-κB target genes (see Supplementary Table 3), using bulk tissue samples of the insult area (green) and the corresponding region of sham-treated mice (white) at 6 h, 3, 7, and 30 dpi as indicated. Quantification is shown for c Msr1, d Spp1, e Lyz2, f Tgm1, g Lgals3, h C3, i Cst7 and j Gpnmb. Bars indicate mean \(\pm \,\)SEM. Statistical comparison by ordinary two-way ANOVA with Tukey’s multiple comparisons test. Exact significances indicated in graphs. Six hours (Sham: n = 5; TBI: c, g n = 8, d, e, i, j n = 7, f n = 6), 3 d (Sham: n = 6; TBI: n = 5), 7 days (Sham: c, e, f n = 3, d, g, h, I, j n = 5; TBI: c, d, f–j n = 6, e n = 5) and 30 days (sham: n = 3; TBI: n = 7); dpi days post injury. Source data are provided as a Source data file.
Of note, TBI also decreased the expression of several genes (Supplementary Table 4) and processes, which mainly refer to synaptic signaling, neurotransmitter receptor and ion channel activity (Supplementary Table 5) (GO terms: e.g., “gated channel activity”; “synaptic signaling”; “chemical synaptic transmission”; and “neurotransmitter receptor activity”).
TBI-induced NF-κB activation is restricted to the insult area and follows a cell-type-specific pattern
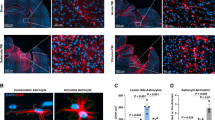
To characterize the spatio-temporal and cell-type-specific NF-κB activation pattern after CHI/TBI in situ, we used NF-κB reporter mice (kappa-eGFP)27,28. These analyses revealed a highly selective NF-κB activation pattern (EGFP+ cells) that is restricted to the immediate vicinity of the impact area. NF-κB activation follows a dynamic course (Fig. 2a, b) and is visible already 1 day post-injury, gets significantly increased at day 3 and stayed elevated until day 7, returning to the level of day 1 by day 15 post-TBI (Fig. 2a, b). Immunostainings with cell identity markers GFAP and ALDH1L1 for astrocytes, IBA1 for myeloid/microglia cells (Supplementary Fig. 1d), and NG2 for oligodendrocyte progenitor cells, allowed to determine the cell-type-specific NF-κB activation pattern. The majority of EGFP+ cells with activated NF-κB signaling represent IBA1+ macrophage/microglia cells, followed by astrocytes, with NG2-glia acting as minor contributors at day 3 and 7 post-TBI (Fig. 2c). The number of IBA1+ cells with active NF-κB did not change from day 3 to 7 (Fig. 2d), whereas almost 70% of the lesion-located astrocytes show activated NF-κB at day 3 after TBI. This portion declined to around 40% on day 7 (Fig. 2e), indicating a highly dynamic NF-κB activation kinetic in astrocytes. CHI/TBI mimics the nature and heterogeneity of human TBI and can lead to different degrees of injury severity, including fractures and bleedings24,26. Interestingly, NF-κB activation can only be detected when post-traumatic bleedings also take place. Thus, the expression strength of astrocyte marker genes like Gfap, Aqp4, and Aldh1l1 positively correlated with hematoma formation (Supplementary Fig. 1e–g).
a Representative images of EGFP expression in NF-κB reporter gene mice (kappa-EGFP) at different timepoints after TBI as indicated (scale bars: overview = 500 µm; zoom = 100 µm). b Quantification of EGFP positive cells per mm2 around the lesion side from (a) is shown (n = 3 per timepoint). c Distribution of EGFP/NF-κB positive cells (in %) in different cell populations at day 3 and 7 post TBI as indicated (n = 3 per timepoint). Quantification of d microglia/macrophages (% of IBA1+) and e astrocytes (% of ALDH1L1+ and GFAP+) expressing EGFP/NF-κB at day 3 (green) and 7 (white) post TBI. f Schematic representation of the workflow applied for flow cytometric analysis of EGFP/NF-κB expressing cells. Quantification of cell frequencies of g O4+ oligodendrocytes, h ACSA-2+ astrocytes, and i CD11b + CD45+ myeloid cells positive for EGFP after TBI (green) at day 1 (n = 8), 3 (n = 6, g n = 7), 7 (n = 3), 14 (n = 3) and 30 (n = 3, g n = 4) compared to sham treated (n = 8; white). j Quantification of CD45+-cells expressing EGFP and further subtyping in myeloid infiltrates (CD45highCD11b+; light green), resident microglia (CD45intermediate/lowCD11b+; dark green) and lymphoid cells (CD45+CD11-; red). Y-axis indicates the %percentage of positive events in the living CD45+ population. Bars indicate mean \(\pm \,\)SEM. Statistical comparison by ordinary one-way ANOVA with Tukey’s multiple comparisons test. Exact significances indicated in graphs; dpi days post injury, n.d. non-detectable; f was created in BioRender. Hein, T. (2026) https://BioRender.com/l4b4p8u. Source data are provided as a Source data file.
To further define the cell-type-specific NF-κB activation after TBI, we conducted flow cytometry using isolated cells from the wider insult area (Fig. 2f). Here, ACSA-2+ astrocytes showed pronounced dynamics in NF-κB activation, presenting with a major increase between day 1 and 3 to day 7 post-TBI, reaching residual activity at day 30 post-TBI (Fig. 2g and Supplementary Fig. 1h). O4+ oligodendrocyte lineage cells represent only a minor fraction with activated NF-κB (Fig. 2h and Supplementary Fig. 1h), while CD45+/CD11b+ myeloid/microglia cells already showed elevated basal NF-κB activity which significantly increases until day 7 post TBI and returning to starting levels at day 30 (Fig. 2i). To better differentiate between resident microglia and myeloid infiltrates besides IBA1 and CD11b expression, the status of the prototypical leukocyte marker CD45 was determined (Supplementary Fig. 1i, j). CD45low microglia were identified as the main contributors of post-traumatic NF-κB activation, while CD45high myeloid infiltrates only featured a comparably mild EGFP signal until day 30 post-TBI (Fig. 2j). Cells of the lymphoid lineage (CD11b−/CD45high) (Fig. 2j) do not show significant NF-κB activation. Among the myeloid subsets, post-traumatic NF-κB was prominently activated in CD45low/CD11c+ DAM cells at day 1 post TBI while CD45highCD11c+ cells show a transient downregulation of basal activity at day 3 (Supplementary Fig. 1k). Ly6G+ neutrophils exhibited only minimal post-traumatic NF-κB activation, while CD206+ CNS-associated macrophages (CAM) presented with marked NF-κB activity at sham and day 1 post TBI, a level continuously decreasing until day 30 (Supplementary Fig. 1k). CD3+ T-cells were almost free of NF-κB activity whereas B220+/MHC-II+ B lymphocytes show mild signs of NF-κB activation (Supplementary Fig. 1l).
Amplified NF-κB signaling in astrocytes impairs neurological recovery and compromises wound closure and reactive gliosis after TBI
To test the specific astrocytic NF-κB activation pattern on TBI outcome, we performed TBI with previously established loss-of-function (LoF; IKK2-DNGFAP) and gain-of-function (GoF; IKK2-CAGFAP) mouse models for NF-κB modulation in astrocytes25,29. These models use the tetracycline-regulated gene expression system (tet-off), where transgene expression was suppressed until the age of 3 and 5 weeks in the case of LoF and GoF animals via doxycycline application to avoid any developmental side effects. After doxycycline withdrawal, the IKK2-CAGFAP and IKK2-DNGFAP mouse models have been functionally validated and confirmed in detail prior to TBI induction (Supplementary Figs. 2 and 3).
After TBI, changes in body weight and motoric behavior were assessed as these parameters are frequently affected in human TBI patients30,31,32. Here, IKK2-CAGFAP animals experienced a persisting weight loss after TBI (Fig. 3a). Six hours after the mechanical insult, all groups, including sham-treated animals, showed a nonsignificant weight loss. While the sham cohort remained at this level for 1 day, both TBI-treated groups featured a further significant weight loss at this time point. At day 3, 7, and 30, TBI-treated controls regained weight to almost sham levels, whereas IKK2-CAGFAP animals exhibited a reduced body weight persisting until 30 days post-TBI, indicating an impaired recovery process in IKK2-CAGFAP animals.
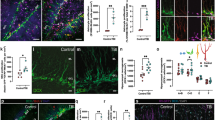
a Post-traumatic progression of body weight respective to the weight prior to TBI. Line plots show weight loss (% loss/gain) over time as indicated. b Assessment of the post-traumatic neurological performance of control and IKK2-CAGFAP animals as compared to their respective sham cohorts using the neurological severity score (NSS). Blue lines indicate comparison between TBI IKK2-CAGFAP and TBI control, red lines for TBI and sham IKK2-CAGFAP (c, d). Sham control: a 6 h n = 15, 24 h n = 15, day 3 n = 15, day 7 n = 3, day 30 n = 3; b 6 h n = 22, 24 h n = 22, day 3 n = 22, day 7 n = 3, day 30 n = 3; TBI control: a 6 h n = 22, 24 h n = 22, day 3 n = 22, day 7 n = 4, day 30 n = 4; b 6 h n = 34, 24 h n = 34, day 3 n = 34, day 7 n = 4, day 30 n = 4; Sham IKK2-CA: a 6 h n = 14, 24 h n = 14, day 3 n = 14, day 7 n = 4, day 30 n = 4; b 6 h n = 19, 24 h n = 19, day 3 n = 19, day 7 n = 4, day 30 n = 4; TBI IKK2-CA: a 6 h n = 23, 24 h n = 23, day 3 n = 23, day 7 n = 7, day 30 n = 7; b 6 h n = 29, 24 h n = 29, day 3 n = 29, day 7 n = 7 day 30 n = 7. Histological analysis (c) and quantification (d) of the post-traumatic lesion area over time as indicated. Sections were stained for GFAP (white) at day 1 (Ctrl: n = 6, IKK2-CAGFAP: n = 4, IKK2-DNGFAP: n = 4), 3 (Ctrl: n = 9, IKK2-CAGFAP: n = 3, IKK2-DNGFAP: n = 4), 7 (Ctrl: n = 6, IKK2-CAGFAP: n = 3, IKK2-DNGFAP: n = 3) and 15 (Ctrl: n = 7, IKK2-CAGFAP: n = 7, IKK2-DNGFAP: n = 4) post TBI. Scale bar = 500 µm. Statistical comparison by ordinary two-way ANOVA with Tukey’s multiple comparisons test. e Representative images and f quantification of longitudinal, lateral MRI in vivo analysis of control (n = 6), IKK2-CAGFAP (n = 4) and IKK2-DNGFAP (n = 3) animals at day 1, 15, and 30 post TBI. Yellow frames emphasize the lesion area used to measure the post-traumatic lesion size. Progression of the post-traumatic lesion healing process is visualized as percentages of the lesion volume remaining of the initial injury as of day 1 (calculate in mm3; set to 100%). Brightfield images show a representative overview of the perfused brains at day 30 post-TBI. Bars indicate mean ± SEM. Statistical comparison by ordinary two-way ANOVA with Tukey’s multiple comparisons test. Control: white–sham, blue–TBI; IKK2-CAGFAP: black–sham, red–TBI: IKK2-DNGFAP: yellow. Source data are provided as a Source data file.
A 10-point neurological severity score (NSS) was used to assess posttraumatic neurological impairments24,33. We also analyzed gait and locomotion utilizing an automated system (CatWalk™XT) to test changes in voluntary movement with high temporal and spatial resolution34,35. The NSS revealed a transient increase in neurological impairments—mainly motor dysfunction during beam walking and beam/round stick balancing—6 h after TBI in both control littermates and IKK2-CAGFAP mice compared to corresponding sham animals (Fig. 3b). After 24 h, the NSS already decreased in both controls and IKK2-CAGFAP, however, the NSS stayed significantly higher in IKK2-CAGFAP mice at this time point and persisted until day 3 post TBI. Later on, no significant differences could be detected between groups (Fig. 3b).
While several of the gait parameters investigated by CatWalk™XT showed only minor changes upon TBI compared to sham mice, the mean intensity of the maximal contact area of left and right front and hind paws was altered after TBI but without significant differences between genotypes (Supplementary Fig. 4a, b). Changes in the contact area can indicate an altered use of affected limbs. Indeed, the mechanical insult on the left hemisphere led to several impairments of the corresponding right paws in the IKK2-CAGFAP, such as length, width, area, and maximal contact area of the right front limb (Supplementary Fig. 4c). For the right hind paw, print positioning (Supplementary Fig. 4d) and print area (Supplementary Fig. 4e) were significantly altered in the IKK2-CAGFAP cohort. Overall, IKK2-CAGFAP animals showed significant differences, especially early after the initial trauma, but recovered until 30 days post-TBI.
To define the cellular and molecular changes influencing the TBI outcome of IKK2-CAGFAP mice, we first performed detailed histopathological analyses of the insult area at various time points after TBI. We found the formation of an initial astroglial border of GFAP-positive cells at day 3 post TBI in control mice, which evolves into a clearly defined glial scar encapsulating the core of the injury area in controls at day 7 and day 15 (Fig. 3c). In IKK2-CAGFAP mice, however, GFAP/IKK2 co-staining revealed a strongly increased and widespread expression pattern of GFAP already under sham conditions (Supplementary Fig. 5a). In response to TBI, the resulting wound borders in IKK2-CAGFAP mice were poorly defined and appeared with a vague and disorganized GFAP staining pattern, possibly indicating an impaired development of the glial limitans border (Fig. 3c, d). In control animals, the damaged trauma area gets slightly increased until day 3 post TBI, thereafter, remodeling of the tissue damage was steadily progressing, leading to a marked reduction in the overall lesion size at day 15 post TBI. In contrast, IKK2-CAGFAP animals showed a strong increase of the wound lesion size at day 3 with only limited reduction of the lesion size at day 7 and 15, thereby revealing a significant wound healing deficit over time until day 15. IKK2-DNGFAP animals do not develop obvious differences and behaved like controls with a clearly defined glial scar formation (Fig. 3c, d).
To further define TBI lesion progression, we also monitored the post-traumatic wound healing process via longitudinal, in-vivo MRI analysis at days 1, 15, and 30 post-TBI. In comparison to control littermates, GoF animals show a profound deficit in wound closure, exhibiting a significantly larger lesion size at day 15, which stayed enlarged until day 30 (Fig. 3e, f). Lesion size reduction in IKK2-DNGFAP animals again progresses similar to the control cohort (Fig. 3e, f).
To evaluate the changes in the glia scar formation of GoF mice in more detail, we performed NG2/CSPG4 and GFAP co-staining of the insult area (Fig. 4a and Supplementary Fig. 5) from day 1 to 15 after TBI. Here, the spread of the scar border zone, which is characterized by very strong NG2 and GFAP staining, was always on a lower level over time in the GoF model compared to controls (Fig. 4b, d). They develop a significantly thicker scar border, which then declines between day 7 and 15 post-TBI (Fig. 4b, d). The deposition of NG2/CSPG4 around the wound core was altered in the IKK2-CAGFAP model, showing a decreased presence over 15 days, while controls exhibit a dynamic course with a significant enrichment at day 7 (Fig. 4b). Reactivity of the NG2 glia was not affected as measured on the degree of hypertrophic NG2-glia (Fig. 4c). The GFAP expression in GoF mice always extends through the entire cortex due to high NF-κB activity, whereas controls again develop a distinctly dynamic course (Fig. 4e). Together, the ratio of injury size to the total area covered by an intense GFAP signal, was significantly lower for the IKK2-CAGFAP group at day 7 and 15 post TBI (Fig. 4f). Also, fibronectin staining revealed a locally restricted signal at the lesion side of controls, which disappeared until day 30 post TBI (Supplementary Fig. 6a, b). In the IKK2-CAGFAP cohort, however, a rather widespread fibronectin expression was observed, which peaked at day 3 post-TBI and declines afterwards until day 30 (Supplementary Fig. 6a, b).
a Representative images of the lesion area stained for GFAP and NG2 at different timepoints after TBI as indicated. Dotted lines illustrate the methodical approach for the quantification of the GFAP- and NG2-marked scar area. The yellow dotted line was used to measure the spread/thickness of the NG2-glia scar, while the distance until NG2-glia reactivity was present as hypertrophic NG2+ cells is indicated with the blue dotted line (days 3, 7, and 15). The spread of the GFAP scar is shown with the red dotted line (day 7, 15), while the GFAP expression thickness, defined as the distance until GFAP expression was detectable, is shown with the orange dotted arrow (day 1, 3, 7, and 15). The area, selected with yellow dotted lines, depicts the GFAP scar around the injury site. Purple squares mark the areas shown with higher magnification in Supplementary Fig. 5. b Quantification of the spread/thickness of the NG2 scar, and c the distance of the observed NG2 hypertrophy. d Quantification of the GFAP scar spread/thickness and e GFAP expression thickness (n = 3 per group). f Quantification of the GFAP scar area normalized to the injury area in control (blue) and IKK2-CAGFAP (red) mice. Scalebars: 500 µm; Bars indicate mean ± SEM. Statistical comparison by ordinary two-way ANOVA with Šidák’s ‘s multiple comparisons test. Exact significances indicated in graphs; control: n = 3, IKK2-CA: day 3 n = 3, day 7 + 15 n = 4. Source data are provided as a Source data file.
Furthermore, distinct differences in the dynamic expression pattern of ECM-associated genes were detected over the course of 30 days in the bulk insult area of control and GoF animals (Supplementary Fig. 6c). While several mediators (Fn1, Dcn, Chst11, Vcan, Tgfb1, and Cstb) showed a stronger expression in GoF animals specifically at day 3 post TBI, only Chst11 and Tgfb1 stay elevated until day 7 post TBI, whereas Mmp12 exhibited a lower level of expression in GoF mice compared to controls. Expression in Col1a2 showed no significant difference. Overall, this suggests a disturbance in the temporal progression of the post-traumatic wound tissue remodeling and ECM formation in mice with NF-κB hyperactivation in astrocytes.
NF-κB modulation in astrocytes disturbs their homeostatic maintenance and reactive function
To decipher the underlying molecular mechanisms for the wound closure deficit in the GoF model, transcriptomic profiling of primary astrocytes (ACSA-2+) and CD11b+ cells (myeloid infiltrates/microglia) isolated from the ipsilateral hemispheres of sham and TBI treated IKK2-CAGFAP, IKK2-DNGFAP and control animals was conducted (Supplementary Fig. 7a). PCA analysis showed a variable, but genotype-dependent clustering upon TBI (Supplementary Fig. 7b) indicating selective TBI responses when NF-κB is either repressed or pre-activated in astrocytes. Intriguingly, the transcriptome signature of TBI-treated IKK2-CAGFAP and sham-treated IKK2-CAGFAP astrocytes did not cluster differentially in PCA, suggesting that NF-κB activation in astrocytes is sufficient to trigger a TBI-like gene expression response (Supplementary Fig. 7b). Volcano plots of DEGs revealed specific genes differently regulated by TBI in GoF and LoF versus control astrocytes (Supplementary Fig. 7c, d). While a great number of DEGs overlapped (Supplementary Fig. 7e, f), TBI GoF and TBI LoF astrocytes showed several candidates (Top 30) significantly up- or downregulated as compared to the TBI control cohort. In the GoF model, the differential upregulation mostly included genes associated with immune functions, while in the LoF astrocytes, induction was mainly found for structural or nonclassified genes (Supplementary Tables S6 and 7). In case of GO term analysis, TBI induces an enrichment of processes associated with immune response/inflammation e.g., “positive regulation of cytokine production” and “regulation of leukocyte activation” (Supplementary Fig. 8a) in controls, a reaction which was significantly pronounced in TBI IKK2-CAGFAP astrocytes (Supplementary Fig. 8b). Importantly, a direct comparison of TBI IKK2-CAGFAP vs. TBI control group further confirmed hyperactivation of immune reactions (Fig. 5a) and increased expression of cytokines and chemokines (Supplementary Fig. 8c) but also revealed enrichment of biological processes related to cell cycle progression (Fig. 5a). When TBI IKK2-DNGFAP astrocytes were similarly compared to the TBI control group, the over-represented GO terms showed obvious differences and included “mitochondrial protein complex”, “cofactor metabolic process”, and “antioxidant activity” (Fig. 5b). However, cytokine expression was not affected in IKK2-DNGFAP astrocytes (Supplementary Fig. 8c).
Enrichment analysis using the Gene Ontology (GO) database showing the most significantly upregulated processes in a TBI IKK2-CAGFAP (GoF; red) and b IKK2-DNGFAP (LoF; yellow) astrocytes when compared to the TBI control (blue) group at day 3 post-TBI. Statistics: DESeq2 RNA sequencing analysis, |log2FoldChange| > 0, p < 0.05. https://doi.org/10.1038/75556, https://doi.org/10.1093/genetics/iyad031. c Schematic diagram of the insult area used for analysis of cell number and proliferation. Although the area of glial reactivity varied with trauma severity and post-traumatic injury timepoint, quantification of glial cells and their proliferation (BrdU+) was restricted to the orange area separated from the injury core (red area). Quantification of d GFAP+ astrocytes and e IBA1+ microglia/myeloid cells over time as indicated. f Representative images of proliferating astrocytes and microglia identified by either BrdU/GFAP or BrdU/IBA1 costaining at the indicated timepoints (sham; 1, 3, and 7 days). Scale bar = 50 µm for overview, scale bar = 20 µm for magnified inlet. Quantification of proliferating g GFAP+/BrdU+ astrocytes and h IBA1+/BrdU+ microglia/myeloid cells at different timepoints post TBI. Sham control d, g n = 6 e, h n = 7; sham IKK2-CAGFAP d n = 3, g, e n = 5, h n = 4; sham IKK2-DNGFAP d n = 2, e, g, h n = 3; day 1: control n = 6, IKK2-CAGFAP n = 3, IKK2-DNGFAP n = 3; day 3: control n = 6, IKK2-CAGFAP n = 3, IKK2-DNGFAP n = 3; day 7: control n = 7, IKK2-CAGFAP d, g n = 3 e, h n = 4, IKK2-DNGFAP n = 3; day 15: control: n = 5; IKK2-CAGFAP: n = 3; IKK2-DNGFAP: n = 3. Arrowheads mark GFAP+/BrdU+ astrocytes (yellow) and IBA1+/BrdU+ microglia/macrophages (red). Statistical comparison by ordinary two-way ANOVA with Tukey’s multiple comparisons test. Exact significances indicated in graphs. Source data are provided as a Source data file.
In line with the enriched proliferation processes in GoF astrocytes after TBI (Fig. 5a), we detected an overall increase in the cell number of GFAP+ astrocytes (Fig. 5c, d) and IBA1+ (Fig. 5c, e) cells around the lesion core of IKK2-CAGFAP mice compared to corresponding controls. Reactive astrocytes further increased until day 15 post-TBI, whereas IBA+ cells were elevated only until day 3. To further characterize the post-traumatic gliosis, proliferation of astrocytes and microglia was assessed by BrdU incorporation over time after trauma (Fig. 5f–h). Interestingly, astrocyte proliferation already peaked at day 3 in the IKK2-CAGFAP model, while controls reached maximal proliferation at day 7 (Fig. 5f, g). In IKK2-DNGFAP animals, astrocyte proliferation showed a mild attenuation without a clearly defined peak. A similar observation was made for IBA1+, where a significantly higher cell number and proliferation was detected at day 3 in IKK2-CAGFAP animals, while proliferation in the control group peaked again at day 7 post-TBI (Fig. 5f, h). Microgliosis and proliferation was significantly milder in the IKK2-DNGFAP model at day 7. These data indicate that astrocytes and microglia from IKK2-CAGFAP animals experience an accelerated post-traumatic proliferation response, leading to a specific reactive gliosis phenotype most likely promoting an irregular post-traumatic scar formation. In IKK2-DNGFAP animals, reactive gliosis seemed dampened but without significant effects on wound healing.
Genetic programs driving post-traumatic wound healing and regeneration are disrupted in IKK2-CAGFAP animals
To characterize the functional status of astrocytes and microglia cells in the IKK2-CAGFAP model, we assessed different transcriptomic signatures previously described in literature to be related to wound healing. Healthy astrocytes initiate specific genetic programs important for wound healing and regeneration as early as 2 days after spinal cord trauma36. When we tested this panel of cAEG genes (consensus healthy astrocyte-enriched genes)36 in our TBI model, we found them almost similarly downregulated in control and LoF astrocytes 3 days post-TBI. In contrast, astrocytes with activated NF-κB (sham GoF) already showed a prominent decrease in cAEG gene expression, which was further aggravated after TBI (Fig. 6a and Supplementary Fig. 8d). In addition, genes associated with homeostatic astrocyte functions, like ion channels and neurotransmitter receptors, were more strongly affected in the IKK2-CAGFAP model after TBI (Supplementary Fig. 9a, b). The epithelial-to-mesenchymal-transition (EMT)36, a signature associated with wound repair and regeneration36, gets significantly induced in controls, while IKK2-CAGFAP astrocytes did not mount this specific response (Fig. 6b), possibly resulting in an impaired border-forming capacity. GSEA of the GO-BP term “wound healing” (GO:0042060), on the other hand, was enriched in both TBI control and TBI IKK2-CAGFAP astrocytes without significant differences (Fig. 6c), indicating a general capability of the IKK2-CAGFAP model to initiate wound healing.
a Transcriptomic data of ACSA-2+ astrocytes purified from control, IKK2-DNGFAP (LoF) and IKK2-CAGFAP (GoF) at day 3 post-TBI, TBI and sham GoF were used for GSEA of a “healthy astrocyte-enriched genes“ (O’Shea et al.36) shown as representative heatmap and annotated with fold changes (log2 normalization) relative to sham control. Enrichment maps of b consensus epithelial-to-mesenchymal transition EMT genes (O’Shea et al.) and c the GO term “wound healing” reveal a lack in mounting significant responses in the GoF model. d Heatmap with corresponding fold changes of the gene signature of reactive, A1 and A2 astrocytes in TBI LoF, control and GoF, as well as sham GoF in comparison to sham control astrocytes. Respective significant (padj < 0.05) fold changes (FCs) are annotated. e Significantly downregulated GO terms associated with extracellular matrix (ECM) and post-traumatic wound healing in TBI GoF CD11b+ (green) and ACSA-2+ (yellow) cells compared to TBI controls, depicted as gene number and gene ratio. f KEGG enrichment chart depicting processes downregulated in ACSA-2 astrocytes derived of TBI IKK2-CAGFAP compared to the TBI control group. g Heatmap of genes associated with the GO term “glutathione peroxidase activity” (log2 normalization; padj = 0.0168) in ACSA-2+ astrocytes at day 3 post TBI. Bars indicate mean \(\pm \,\)SEM. Statistics: DESeq2 RNA sequencing analysis (two-sided Wald’s test), |log2FoldChange| > 0, p < 0.05. FDR correction of p-values (padj) by multiple comparisons test. Identification of over- or underrepresented GO and KEGG terms by hypergeometric distribution with subsequent multiple testing correction (padj). https://doi.org/10.1038/75556, https://doi.org/10.1093/genetics/iyad031. Over- or underrepresentation of gene sets (GSEA analysis) identified by phenotype-based permutation and subsequent multiple hypothesis testing. https://doi.org/10.1038/ng1180. Source data are provided as a Source data file.
Previous studies identified different subtypes of reactive astrocytes induced by either inflammatory insults producing a neurotoxic A1 phenotype or a rather neuroprotective and regenerative A2 phenotype initiated by ischemic injury37,38. As A1 neuroinflammatory reactive astrocytes are proposed to be induced by NF-κB, we found a significant enrichment of the A1 signature in astrocytes of IKK2-CAGFAP animals at sham and TBI conditions whereas astrocytes of control and IKK2-DNGFAP mice express lower levels of A1 genes (Fig. 6d). Interestingly, only IKK2-DNGFAP astrocytes develop a prominent upregulation in pan-reactive and neuroprotective A2 genes (Fig. 6d).
Importantly, we determined a negative, post-traumatic enrichment in various matrix-associated and regenerative GO terms in TBI IKK2-CAGFAP animals compared to the TBI controls, affecting both ACSA-2+ astrocytes and CD11b+ myeloid/microglia cells (Fig. 6e). Accordingly, the TBI-induced expression of multiple ECM component genes (GO: 0044420), such as Tgfb2, Col4a3, Lama or Ptn was significantly decreased in both cell populations derived from the IKK2-CAGFAP model (Supplementary Fig. 9c, d). Independent of the deficits in ECM remodeling, IKK2-CAGFAP astrocytes exhibited a negative enrichment in processes associated with protective and metabolic functions (Fig. 6f). This included glutathione peroxidase activity (Fig. 6g), important for the ROS-detoxifying processes. Furthermore, we could identify a SASP gene expression profile (senescence-associated secretory phenotype)39,40 after TBI (Supplementary Fig. 10a) previously linked to age-dependent wound repair deficits41. This signature was already present in non-lesioned IKK2-CAGFAP and further cemented by TBI (TBI IKK2-CAGFAP; Supplementary Fig. 10a). Notably, IKK2-DNGFAP animals develop a positive enrichment in those GO terms associated with ROS protection, such as “antioxidant activity” as well as an upregulation in components associated with their detoxification (Fig. 5b and Supplementary Table 8).
Astrocytic NF-κB activation exacerbates neuroinflammatory immune responses after TBI
Traumatic brain injury causes disruption and increased permeability of the blood-brain barrier (BBB), which supports easier entry of peripheral leukocytes into the brain. Immunoglobulin IgG staining in brain samples at different time points post TBI revealed a widespread parenchymal staining in IKK2-CAGFAP compared to control mice suggesting an increased BBB leakage in the GoF model (Supplementary Fig. 10b). Thus, we performed a detailed analysis of the post-traumatic neuroimmune response using flow cytometry to immunophenotype brain resident and infiltrating cells after TBI (gating strategies see Supplementary Fig. 11a, b). TBI induced a significant infiltration of myeloid cells (CD45highCD11b+) in all groups at day 3 of the post-traumatic immune response (Fig. 7a, b). Myeloid CD11b+ infiltrates further contained a population stained positive for CD11c, composed of CD45high/CD11c+ (Fig. 7c, e and Supplementary Fig. 11c) and CD45lowCD11c+ (Fig. 7d, e) subsets. A minor population of CD45highCD206+ macrophages (Fig. 7f) is present in the lesioned hemisphere as well. The frequency of the overall myeloid infiltration was significantly higher in the IKK2-CAGFAP cohort (Fig. 7b). This also applies to CD11b+/Ly6C+ infiltrates, which mainly include classical pro-inflammatory monocytes and Ly6G+ neutrophils (Fig. 7f, g). A detailed subtyping revealed a heterogeneous mixture of mainly CD45low/intermediate/CD11c+ microglia, a CD45high/CD11clow/MHC-II+/−/Ly6C+ population and CD45highCD11chighMHC-IIhigh dendritic cells (Supplementary Fig. 12a–c). In the microglia population (CD45low/intermediate CD11b+), expression of MHC class II and CD11c+ was significantly increased in the IKK2-CAGFAP cohort as well, thereby acting as an indicator of phenotypical changes (Supplementary Fig. 12d). All CD45high/CD11b+ cells co-expressed the migration marker CD44, which has previously been identified as an indicator of a brain-external origin42,43 (Supplementary Fig. 13a). Expression of this migration marker was also still present in residual infiltrates at day 14 post TBI, while expression density of the microglia marker P2RY12 further assists discrimination between resident and peripheral immune cells (Supplementary Fig. 13b). Remarkably, the peripheral CD45high/CD11b+ infiltrates in control and LoF mice were composed of CD11c+ cells and Ly6G+ neutrophils in equal parts, together with a smaller subset of mainly Ly6C+ cells and a minor population of CD206+ CAMs (Fig. 7h). GoF animals, however show an altered composition of peripheral infiltrates, featuring a marked preference of CD11c+ cells, while the proportion of Ly6G+ neutrophils was strongly decreased (Fig. 7h). Following up the qualitative and quantitative progression of the post-traumatic infiltration, we analyzed frequencies and total cell number of CD45high/CD11b- lymphoid (Fig. 7i, j), CD11b+/CD11c+ dendritic/DAM (Fig. 7k, l) and CD11b+/Ly6C+ (Fig. 7m, n) myeloid cells. Frequency and total cell number of lymphoid cells showed a mild increase in the course of 30 days post-TBI in both control and LoF mice. In contrast, GoF animals already presented with a significantly increased level of lymphoid cells (frequency and total number) at sham conditions, which gets prominently upregulated between day 3 and 14 post-TBI and stays elevated until day 30 post-TBI (Fig. 7i, j). CD11b+/CD11c+ cells exhibit a small peak in frequency and numbers at day 3 post-TBI in controls and LoF mice, which then declines to almost sham values at day 30. Again, GoF mice show an elevated basal level of CD11b+/CD11c+ cells and develop an obvious peak in total cell number from day 3 to 7, which then decreases again to basal levels at day 30 (Fig. 7k, l). Infiltration of CD11b+/Ly6C+ cells—primarily including monocytes and granulocytes—shows the highest frequencies at day 3 post-TBI in all three genotypes that then return to sham levels. This also applies to the absolute cell number, especially in GoF animals, which already start with a high basal level at sham conditions (Fig. 7m, n). The absolute cell counts clearly indicate a higher abundance of infiltrates in the GoF model. LoF animals do not exhibit significantly different population frequencies and cell counts but might exhibit a slight delay in the temporal organization of the post-traumatic leukocyte infiltration (Fig. 7i–n). Taking a closer look into the lymphoid compartment in the CNS, upregulation of astrocytic IKK/NF-κB signaling shows a profound effect on this population. The frequency of lymphocytes was sparse in controls at day 3 post trauma, while the GoF model already presents with changes in frequency and composition of the T cells (Supplementary Fig. 13c–f). Here, a significant increase in CD3+ T lymphocytes (Supplementary Fig. 13c) as well an altered CD4/CD8a cell ratio was determined, showing an increased frequency of CD3+/CD8a+ cytotoxic T lymphocytes (CTL) at day 3 post TBI (Supplementary Fig 13d–f). Interestingly, proportion of CD8a+ CTLs increases in the control cohort until day 14 post TBI, indicating that an adaption to the IKK2-CAGFAP group is taking place (Supplementary Fig. 13e). In contrast, there was no difference in the presence of B lymphocytes (B220+/MHC-II+) (Supplementary Fig. 13g). Overall, pre-existing neuroinflammation in IKK2-CAGFAP mice critically affects persistence and infiltration of different myeloid cell populations thereby interfering with regular post-traumatic wound healing.
Flow cytometry analysis of isolated brain leukocytes was performed at various time points post-TBI. a Representative scatter plots and b quantification of CD45highCD11b+ myeloid infiltrates derived of sham and TBI controls, IKK2-CAGFAP and IKK2-DNGFAP animals at day 3 post TBI. Representative scatter plots of c CD45highCD11b + CD11c+ and d CD45lowCD11b + CD11c+ as well as e quantification of CD45 + CD11b + CD11c+ myeloid cells measured in the ipsilateral hemisphere at day 3 post TBI. Y-axis shows percentage of living cells gated on CD45 + CD11b+. f Representative gating strategy for Ly6G+ neutrophils and CD206 + CNS-associated macrophages (CAMs) in the ipsilateral lesioned hemisphere TBI control at day 3 post TBI. g Frequency of CD11b+ cells also expressing Ly6C at day 3 post-TBI. Y-axis shows the percentage of cells gated on living CD11b+ cells. h Bar charts showing the relative composition of the CD45highCD11b + CD44+ compartment of infiltrates, comparing the control, IKK2-CAGFAP and IKK2-DNGFAP cohorts at day 3 post TBI. Frequencies and total cell counts of i, j CD45highCD11b- lymphoid cells (y-axis = % of gated living cells), k, l CD45 + CD11b + CD11c+ and m, n CD45 + CD11b + Ly6C+ myeloid cells progressing over time post trauma (x-axis). Bars indicate mean \(\pm \,\)SEM. Statistical comparison by ordinary one- or two-way ANOVA with Tukey’s multiple comparisons test. Exact significances indicated in graphs. i, k, m Blue: control (sham: n = 13, 3 dpi n = 13, 7 dpi n = 12, 14 dpi n = 10, 30 dpi n = 13); red: IKK2-CAGFAP (sham: n = 8, 3 dpi: n = 12, 7 dpi: n = 5, 14 dpi: n = 9, 30 dpi n = 6); yellow: IKK2-DNGFAP (sham: n = 6, 3 dpi n = 8, 7 dpi n = 7, 14 dpi n = 7, 30 dpi n = 3); j, l, n blue: control (sham: n = 6, 3 dpi n = 11–12, 7 dpi n = 12, 14 dpi n = 7, 30 dpi n = 9); red: IKK2-CAGFAP (sham: n = 3, 3 dpi: n = 10, 7 dpi: n = 3, 14 dpi: n = 8, 30 dpi n = 6); yellow: IKK2-DNGFAP (sham: n = 6, 3 dpi n = 8, 7 dpi n = 4, 14 dpi n = 5, 30 dpi n = 3). Source data are provided as a Source data file.
According to the TBI-induced changes in myeloid immune cell infiltration, we determined differential gene expression profiles in purified CD11b+ cells derived from TBI control, sham IKK2-CAGFAP, TBI IKK2-CAGFAP and TBI IKK2-DNGFAP at day 3 post-TBI TBI as shown by PCA (Supplementary Fig. 14a, b). NF-κB pre-activation in astrocytes exerts a profound effect on the CD11b+ myeloid compartment at day 3 post-TBI in IKK2-CAGFAP mice, whereas controls and IKK2-DNGFAP only showed slightly separated clusters at this time point. The transcriptomic changes in CD11b+ cells were reflected in a positive enrichment of immune cell activation terms that again were more pronounced in TBI IKK2-CAGFAP mice, such as responses to IFN-γ and pronounced cytokine signaling (Supplementary Fig. 14c). IFN-γ has a unique function in microglia activation, promoting their proliferation and driving them towards neurotoxic phenotypes44. Correspondingly, components of the Interferon-gamma mediated signaling pathway (GO:0060333), such as Jak2, Stat1, and Irf1, were significantly upregulated in the IKK2-CAGFAP myeloid population (Supplementary Fig. 14d).
Another important component of the innate immune response in the CNS is inflammasome signaling45,46, and especially the NLRP3 inflammasome reacts on PAMPs and DAMPs released upon TBI47. We detected an increased expression of NLRP3 inflammasome components in the ACSA-2+ population of TBI IKK2-CAGFAP (Fig. 8a) and determined a strong expression of NLRP3 and the associated ASC protein in the ipsilateral (ip) TBI insult area compared to the uninjured contralateral (co) site (Fig. 8b–d). AIM2 (absent in melanoma 2), another inflammasome factor responsive to double-stranded DNA, was downregulated after TBI (Fig. 8b, e). Interestingly, the TBI-induced NLRP3 and ASC levels were significantly increased in IKK2-CAGFAP mice, corresponding to their elevated neuroinflammatory conditions and wound healing deficits (Fig. 8c, d).
a Heat map (log2 normalization) of genes encoding inflammasome components expressed in GoF ACSA-2+ astrocytes at day 3 post-TBI compared to TBI controls. b Immunoblot analysis and corresponding quantification of c ASC, d NLRP3 and e AIM2 inflammasome components. Expression was normalized to eIF2 loading control. f Immunoblot analyses and the corresponding quantification of protein expression of g TGM1, h NRF2, i IBA1, j HO-1, k MAC2, l OPN and m LCN2 in the ipsilateral insult region (ip) and in the contralateral uninjured (co) cortical brain tissue from control (n = 3) and IKK2-CAGFAP (n = 3) animals. Tissue was isolated at day 3 post-TBI. n Ratio of normalized OPN and LCN2. For quantification, protein levels were normalized to β-actin loading control. Bars indicate mean ± SEM. Statistical comparison by ordinary one-way ANOVA with Tukey’s multiple comparisons test. Exact significances indicated in graphs. Protein expression presented in arbitrary units (arb. units). Protein data are presented as biological replicates. Blue: control ipsi, white: control contra, red: IKK2-CAGFAP ipsi, black: IKK2-CAGFAP contra. OPN: Osteopontin (Spp1), HO-1: Heme oxygenase 1 (Hmox1), MAC2: Galectin-3 (Lgals3). Source data are provided as a Source data file.
Factors determining TBI outcome get deregulated by NF-κB activation in astrocytes
Finally, we determined the protein expression status of top-regulated factors associated with disease progression and regeneration at day 3 post-TBI, including TGM1 (Tgm1), HO-1 (Hmox1), and OPN (Spp1), which are known to be involved in protective and remodeling processes after brain injury48,49,50. Furthermore, marker proteins indicative for inflammation and reactive gliosis, like MAC2, LCN2, and IBA1, were addressed, as well as NRF2 levels to get information on oxidative stress and glutamate homeostasis after TBI (Fig. 8f). Immunoblot analyses validated robust IKK2-CA transgene expression in IKK2-CAGFAP mice. TBI leads to a significant downregulation, especially in IKK2-CAGFAP animals, while NRF2 is only expressed upon TBI (ipsilateral) but without differences between controls and IKK2-CAGFAP mice. Importantly, all other factors tested were found with altered protein levels on the ipsilateral side, indicating an immediate response at the direct insult area (for quantification see Fig. 8g–m). TGM1 expression is stronger expressed in the IKK2-CAGFAP insult area, whereas OPN is significantly less upregulated in IKK2-CAGFAP mice after TBI. In the IKK2-DNGFAP model, however, OPN expression is significantly stronger (Supplementary Fig. 15a, b) while other mediators did not differ from the TBI control cohort (Supplementary Fig. 15a–e), indicating a specific regulation by IKK/NF-κB signaling after TBI. Notably, analysis of Spp1 mRNA expression in purified astrocytes (Supplementary Fig. 15g) and microglia (Supplementary Fig. 15h) revealed a cell-type-specific regulation pattern after TBI. While Spp1 expression was prominently induced in both cell types, the overall expression level was much more pronounced in microglia compared to astrocytes, suggesting that microglia are the main contributors of OPN protein after TBI. Remarkably, this upregulation was significantly lower in microglia of IKK2-CAGFAP animals. HO-1, MAC2, LCN2, and IBA1 protein expression are highly responsive to trauma, while their baseline expression (contralateral side) is low or at the detection limit. In the case of HO-1, IBA1, and LCN2, the TBI-induced protein level was more abundant in IKK2-CAGFAP animals. As osteopontin and lipocalin are proposed critical markers for CNS injury and astrocyte reactivity8,51,52 we also calculated the ratio between OPN/LCN2 protein levels at the insult area to get a scoring of TBI progression/outcome (Fig. 8n). Indeed, there was a clear correlation between beneficial wound healing progression in controls showing an OPN/LCN2 ratio of around one and the detrimental outcome in the IKK2-CAGFAP model with a significantly reduced ratio (mean value = 0.279). Interestingly, the OPN/LCN2 ratio correlated inversely with IKK2-CA transgene expression in astrocytes (Fig. 8f, n), meaning that the higher NF-κB activity after TBI, the lower the OPN/LCN2 ratio. Albeit the OPN/LCN2 ratio in the IKK2-DNGFAP model was found higher than 1, the increase was not significant according to the mild phenotype change (Supplementary Fig. 15f). However, at day 14 post-TBI, the IKK2-DNGFAP cohort still showed an enriched expression of OPN, HO-1 and LCN2 protein, suggesting an improved TBI outcome (Supplementary Fig. 15i).
Discussion
Traumatic brain injury represents a challenging, global health concern, which is characterized by high mortality and low recovery rates1,4. Importantly, secondary injury processes dominated by complex neuroinflammatory responses can lead to long-lasting complications. Therefore, a detailed understanding of the underlying molecular mechanisms is a prerequisite for the development of appropriate and timely treatment options53. Our study uncovers two IKK/NF-κB-dependent mechanisms in astrocytes which substantially contribute to an unfavorable TBI outcome: (i) a cell-intrinsic disturbance of astrocytic scar formation and (ii) alterations in post-traumatic leukocyte recruitment and immune activation. Proper organization and tuning of these two branches are essential for the adequate regulation of CNS wound healing and scar formation in mice (Supplementary Fig. 16)1,4,53.
In support of a key role of NF-κB in the regulation of astrocyte homeostasis and the pre-disposition to detrimental post-traumatic responses, recent studies showed that transcriptional changes underlying astrocyte reactivity can be highly heterogeneous in different neuropathological conditions and depend on context-specific transcription factor crosstalk9,17. The number of transcriptional regulators acting in this scenario is strongly limited but always includes the NF-κB transcription factor family, prominently placed18. Similar to our study, the canonical IKK2/NF-κB signaling pathway was identified as a crucial functional regulator unit promoting distinct inflammatory reactive states in human iPSC-derived astrocytes19. In addition, pro-inflammatory astrocyte epigenetic memory was shown to boost NF-κB-dependent transcriptional modules, thereby forcing pathogenic astrocyte reactivity20. So far, there is limited information on the exact consequences of TBI-induced NF-κB signaling in astrocytes on the resulting cell-autonomous and non-cell-autonomous reactions and their contribution to TBI outcome54,55. In cortical stab wound injury, a specific TBI paradigm, propagation of reactive astrogliosis was linked to distinct clusters of GFAP+ reactive astrocytes. These clusters could be enriched by LPS, a strong inducer of NF-κB, whereas pleiotropic NF-κB inhibition by sulfasalazine could suppress reactive astrogliosis after TBI56. In our IKK2-CAGFAP model, selective activation of IKK/NF-κB signaling in astrocytes was already sufficient to trigger a global reactive gliosis phenotype.
CNS wound healing and scar formation is a multicellular process depending on complex interdependent interactions amongst diverse neural and non-neural cells57,58. It also depends on the optimal timing and the exact sequence of action to guarantee successful restoration of the brain lesion and finally to achieve neuroprotection7,57,59. Scar formation encompasses a specific cytoarchitectural organization consisting of a central core of stromal cells together with fibrotic ECM and infiltrating peripheral immune cells that are surrounded by proliferating, reactive astrocytes and other glial cells. These cells form limitans borders to separate the central lesion core from the healthy CNS parenchyma26,58. In the past, glial cells were mainly regarded as producers of detrimental glial scars, overall interfering with neural regeneration, however, this view has been challenged, as reactive astrocytes can permit and even support CNS axon regeneration60. Our findings support the idea that NF-κB-activated astrocytes become primed for premature astrogliosis and paracrine microgliosis that disturbs the kinetics of the coordinated multicellular scar-forming process after TBI. The specific pathological response is defined by aberrant expression of distinct tissue repair- and remodeling-associated factors that repress appropriate glia limitans and fibrotic core scar formation while immune cell infiltration and activation are simultaneously aggravated. In particular, the proliferation kinetics of border-forming astrocytes and microglia at the wound lesion site are significantly accelerated in IKK2-CAGFAP mice.
Border-forming astrocytes are a damage-responsive cell population recently defined in a model of SCI36. They arise during post-traumatic gliosis and are characterized by suppressed homeostatic functions and interactions, while immune defense pathways are enriched, indicating a state of differentiation and functional immaturity36. Here, we identified a similar type of functional polarization in response to CHI/TBI. However, while downregulation of homeostatic functions and overrepresentation of immune-related pathways was initiated in control astrocytes after TBI, even without any injury, IKK2-CAGFAP astrocytes already showed an aggravation of these features due to NF-κB activation per se, an effect which was further boosted by TBI. IKK2-CAGFAP astrocytes acquire a specific reactivity status, including a gene expression pattern usually emerging only with advanced age61,62,63, including pre-existing alterations in neuroimmune function64, astrogliosis65, alterations in glutathione metabolism66,67 and SASP expression40. They also show an enrichment in neurotoxic A1 signature genes, which are frequently observed in various neurodegenerative diseases38 and physiological aging68. Interestingly, NF-κB-activated astrocytes can change the functional status of CD11b+ cells and initiate prominent downregulation of regenerative GO terms in these cells. In SCI, a specific healing-associated gene expression signature was highly enriched in astrocytes after injury, showing an early adaption of the transcriptome to regenerative processes36. This gene signature was also found responsive to TBI but not affected in the IKK2-CAGFAP model, suggesting NF-κB independent regulation.
To define the status of factors relevant for the regulation of post-traumatic wound healing and TBI outcome, we focused on osteopontin (OPN), lipocalin 2 (LCN2) and heme oxygenase (HO-1). Lack of OPN in spinal cord injury resulted in severe tissue damage and impaired motoric recovery, thus supported a neuroprotective function of OPN69. In line with this, OPN was determined as a beneficial factor for tissue repair and neuroprotection in different brain injury models48,49,70,71,72,73. Osteopontin is a multifunctional protein proposed to interact with CD44 and integrin receptors, and associated with cell vitality, proliferation, migration and immunity74. After brain injury, OPN expression is strongly induced and is linked to cell crosstalk and ECM homeostasis75,76. In addition, OPN is critically involved in the initiation of epithelial-to-mesenchymal transition (EMT), a process which plays a major role in wound healing77,78. Accordingly, we identified a clear TBI-dependent upregulation of OPN in the TBI insult area of control mice, coinciding with an enrichment of a specific consensus EMT gene signature36. In contrast, IKK2-CAGFAP mice exhibited a significantly reduced OPN expression after TBI and in parallel, astrocytes lacked the corresponding EMT gene enrichment, which together disturbs scar formation. In addition, we found an increased fibronectin deposition at the lesion area of IKK2-CAGFAP mice, a known pathology associated with wound healing deficits79,80,81.
Consistent with other studies82,83,84, we detected a massive increase of HO-1 in the insult region, a stress-induced enzyme mainly involved in the degradation of released hemoglobin, thereby forcing pronounced iron disposal in traumatized brains. Iron overload can increase oxidative stress and pro-inflammatory responses50, and may trigger ferroptosis85 associated with TBI86. Higher expression of HO-1 may also reflect a more detrimental TBI progression, similar to patients, where increased HO levels were related to higher injury severity87. Notably, elevated HO-1 levels coincided with higher LCN2 expression in IKK2-CAGFAP mice. LCN2 is an iron trafficking protein that acts as a major driver of post-traumatic neuroinflammation, and genetic inactivation of Lcn2 or LCN2 depletion reduced brain damage and improved neurological outcome after TBI88,89. Although LCN2 function in TBI and stroke is complex, upregulation has been associated with neurotoxicity and astrocyte reactivity. In this perspective, LCN2 has been proposed as a potential therapeutical target to counteract neurodegeneration90. Other factors known to influence TBI outcome are transglutaminases like Tgm1 (and Tgm2), which catalyze crosslink formation between proteins91. They get upregulated upon TBI and have been linked to neurodegeneration92. In stab wound injury, reduction of astrocyte-derived TGM1 protein was related to beneficial scar formation93,94. In our GoF model, TGM1 protein upregulation coincided with detrimental scar development, while TGM1 expression in LoF mice was reduced, albeit only by tendence.
The overall wound closure process of IKK2-DNGFAP mice with astrocyte-selective NF-κB inhibition did not undergo obvious beneficial alterations, although IKK2-DNGFAP-derived astrocytes showed several specific changes in their post-traumatic gene expression profile (e.g., development of a beneficial A2 signature, enrichment in mitochondrial and metabolic GO terms). Most likely, the paracrine effects usually mediated by reactive astrogliosis in TBI, like microgliosis, could not efficiently be suppressed by NF-κB inhibition in LoF astrocytes. In support of this, RNA-sequencing revealed that PCA of CD11b+ cells from TBI controls and TBI IKK2-DNGFAP do not clearly differentially cluster in PCA analysis. However, we also cannot exclude that dominant-negative acting IKK2-DN was not sufficient to achieve a complete blockade of TBI-induced NF-κB activation.
Our results revealed that pre-activation of IKK/NF-κB signaling in astrocytes initiated both quantitative and qualitative changes in the immune cell infiltration into the traumatized brain. Among these, we observed a massive preference of CD11c+ immune cells in the lesioned brain area of IKK2-CAGFAP animals. Subtyping of this cell population revealed a quite heterogeneous make-up of CD11c+/MHC class IIhigh/CD44+/Ly6Clo/hi cells likely including both conventional (cDC) and monocyte-derived (moDC) dendritic cells (DCs). The enrichment of IFN-γ-associated pathways found in the myeloid transcriptome of IKK2-CAGFAP mice could create conditions similar to EAE, promoting monocyte to moDC differentiation95,96 and may explain elevated DC levels already under sham conditions in the IKK2-CAGFAP model. We also detected alterations in the CD4/CD8 T cells ratio in IKK2-CAGFAP animals after TBI. Thus, the increased frequency of CD8a+ T cells (CTLs) may aggravate post-traumatic tissue damage, as these cells get potently activated by DCs97, and may trigger inflammatory neurodegeneration98,99,100 as well as neurological manifestations after TBI50,101.
The consequences of post-traumatic DC activity in the CNS are controversially discussed and may range from direct beneficial effects in the acute restoration of tissue homeostasis after injury to the initiation of long-term autoreactive T cell and autoantibody responses, which again have the capacity to generate positive or detrimental outcomes102,103. The presence of autoantibodies responses following TBI is well recognized, however, the exact correlation to trauma severity and its progression is less understood102,104. The exact contribution and underlying molecular mechanisms how DCs initiate and coordinate specific autoantibody responses in TBI is still ill-defined, thus, the IKK2-CAGFAP model is a valuable tool to characterize the underpinnings of DC-dependent autoantibody production.
In summary, our findings demonstrate that aggravated, post-traumatic IKK/NF-κB signaling in astrocytes, a scenario most likely arising with aging, dysregulates the balance between astrocyte-intrinsic processes and the overt neuroinflammatory response at the lesion area, thereby impairing effective wound healing and worsening the TBI outcome. Moreover, our findings encourage further prospective clinical testing of secreted OPN, LCN2, TGM1, and HO-1 levels in blood/serum to find out whether these factors are relevant prognostic parameters in TBI patients.
Methods
Transgenic mice
All animal experiments were performed in compliance with the institutional animal care guidelines at Ulm University (TFZ, Ulm) and the German animal protection law and approved by the Regierungspräsidium Tübingen (RP Tübingen, Germany; TVA1192).
Mice conditionally expressing either a constitutively active IKK2 allele (IKK2-CA) or a dominant-negative acting allele (IKK2-DN) in astrocytes (IKK2-CAGFAP, IKK2-DNGFAP) were previously generated and reported25,29,105. In brief, a transdominant (VSV-IKK2-DN; D145N) and a constitutively active version (CA-IKK2; IKK2-EE; S177E, S181E) of human IKK2 were cloned into the bidirectional tetracycline-dependent expression vector pBL5 to create the plasmids pBI5-VSV-IKK2-DN and pBL5-CA-IKK2. A bidirectional minigene was subsequently released by AatII-AseI, both fragments microinjected into the pronuclei of fertilized oocytes (NMRI outbred) and transplanted to pseudopregnant foster mothers106. To generate IKK2-CAGFAP and IKK2-DNGFAP double-transgenic mice, GFAP.tTA mice were crossed with (tetO)7.IKK2-CA) or (tetO)7.IKK2-DN)107. For transgene inactivation during complete development, doxycycline (0.5 g/L; MP Biomedicals) in 1% sucrose was given in the drinking water to the mothers during pregnancy from conception to weaning until the age of five weeks, then transgene expression was induced by Dox withdrawal. IKK2-CAGFAP and IKK2-DNGFAP mice were bred on the NMRI background, and single-transgenic and transgene-negative littermates were always used as controls. To monitor cell-type-specific NF-κB activation in response to TBI, we used an NF-κB-responsive reporter mouse line (kappa-EGFP) that uses EGFP as a read-out for in vivo NF-κB activity27,28. These mice were bred on pure 129/P2 Ola background.
Male mice were used in experiments if not otherwise stated. Mice were maintained under specific pathogen-free conditions in the animal facility of Ulm University (TFZ, Ulm). Animals were kept in groups of 3–5 mice in a 12-h light-dark cycle and were given ad libitum access to food and water.
CHI model
Adult male mice (12–20 weeks old) were subjected to experimental CHI with a standardized weight-drop device24,26. The animals were anesthetized with ketamine (Pfizer Pharma, Karlsruhe, Germany), with an intraperitoneal dose of 100 mg/kg body weight, and 2% xylazine (Bayer Health Care, Monheim, Germany), with an i.p. dose of 16 mg/kg body weight. Afterward, the skull was exposed by a longitudinal incision of the skin, and a focal blunt injury was induced in the left hemisphere by dropping a 330 g metal rod on the skull from a height of 2.7 cm. Only animals with a visible imprint of the falling rod on the skull were included for further analyses. After trauma, the mice received supporting oxygenation with 100% O2, the wound was sutured, and the animals were placed into a warmed recovery cage with ad libitum access to food and water. Buprenorphine analgesia (Temgesic; Essex Pharma, Munich, Germany) was administered subcutaneously (0.03 mg/kg body weight) immediately after trauma and again at 6 h until 24 h. Sham-procedure mice underwent anesthesia, scalp incision, suturing of the wound, and analgesia, but no experimental head trauma.
Gait analysis
Mouse gait analysis was performed using the CatWalk XT Automated Gait Analysis System (Noldus Technology, Wageningen, NL). Setup of the walkway area was done according to the manufacturer’s recommendations and adjusted for mice. Three compliant individual runs needed to be performed in 1–10 s for one successful trial. Prior to the experiments, mice were acclimated to the testing setup to ensure an uninterrupted performance. For the trial, home cages with food and nesting material were placed in the goal box, functioning as motivators for a successful run. Gait analysis was conducted by the provided software following acquisition of each trial, using a combination of automatic analysis and manual examination in slow motion. Runs displaying abnormalities, such as interruptions or sniffing by the analysis, were excluded. Trails were conducted using animals before and at day 2, 7, and 30 post-trauma or sham-treatment.
Primary cell isolation
Primary astrocytes and microglia from adult mice were isolated using the Adult Brain Dissociation Kit, gentleMACS Octo dissociator with heaters and MS columns in a magnetic field (Miltenyi Biotec, Bergisch-Gladbach, Germany) according to the manufacturer’s protocol. In brief, brains were dissected, enzymatically digested, and the debris was removed. Four to 6 hemispheres were pooled per sample. The remaining single cell suspension was first Fc-blocked and labeled with anti-ACSA-2 microbeads (130-097-679, Miltenyi Biotec), washed, and applied to an MS column. Purified ACSA-2-positive astrocytes were pelleted and snap-frozen in liquid nitrogen. The flowthrough was subsequently labeled with anti-CD11b microbeads (130-049,601, Miltenyi Biotec), washed, and purified using an MS column. The cell pellet containing CD11b positive cells was snap-frozen in liquid nitrogen. ACSA-2 and CD11b isolates were stored at −80 °C until further analysis. For primary astrocyte cell culture, purified ACSA-2+ astrocytes were plated on Poly-D-Lysin-coated plates using serum-free astrocyte medium. After 24 h, cells were left untreated or treated with a cocktail of LPS (10 µg/ml)/TNF-α (100 pg/ml)/PolyI:C (2 µg/ml) for 16 h to stimulate the IKK/NF-κB signaling pathway. Subsequently, cells were harvested and RNA isolated.
RNA extraction, cDNA synthesis, and qRT-PCR
RNA of cortical tissue and primary cells was isolated using a PeqGOLD® TriFast™ kit (peQlab, Darmstadt, Germany) or, for the day 15 and 30 cohort, Qiagen RNeasy Kits (Qiagen) according to the manufacturer’s instructions. For cDNA synthesis, the transcriptor high fidelity cDNA synthesis kit (Roche, Penzberg, Germany) was used according to the manufacturer’s instructions, but with only 0.5 µL instead of 1 µL reverse transcriptase mix and oligo-dT primers alone. To perform qRT-PCR, we used the LightCycler® 480 Instrument (Roche) according to the manufacturer’s instructions with intron-spanning primers and mono-color hydrolysis probes designed by Roche’s Universal Probe Library UPL assay design center. As a reference gene, the housekeeping gene hypoxanthine-guanine phosphoribosyl transferase (Hprt1) was used. Primer sequences and UPLs are available upon request.
RNA sequencing of primary cells
RNA sequencing was conducted by Novogene (London, UK). In summary, mRNA was purified from total RNA using poly-T oligo-attached magnetic beads. After the fragmentation step, random hexamer primers were used for first-strand cDNA synthesis, which was followed by the second-strand cDNA synthesis using dUTP for the directional library. The library was afterward checked by Qubit and real-time PCR for quantification, for size distribution detection, a bioanalyzer was used. Index-coded samples were clustered according to the manufacturer’s instructions. After cluster generation, the library preparations were subsequently sequenced on an Illumina platform. Paired-end reads were generated, and raw data (raw reads) of FASTQ format were processed through fastq. Reads containing adapter and poly-N sequences, as well as reads with low quality, were removed in this step to obtain clean data (clean reads). Q20, Q30, and GC content of the clean data were calculated, and for all downstream analyses, clean data of high quality were used. GRCm39 reference genome as well as gene model annotation files were directly downloaded from the genome website browser (NCBI/UCSC/Ensembl). Paired-end clean reads were aligned to the reference genome using the spliced transcript alignment to a reference (STAR) software. This software is based on a previously undescribed RNA-seq alignment algorithm that uses sequential maximum mappable seed search in uncompressed suffix arrays, which is followed by seed clustering and stitching procedure. Compared to other RNA sequencing aligners, STAR exhibits better alignment precision and sensitivity in both experimental and simulated data. FeatureCounts was used to count the read numbers mapped of each gene, reads per kilobase of exon model per million mapped reads (RPKM) were calculated based on the gene length and reads count mapped to this gene. With RPKM, the effects of sequencing depth and gene length for the read count are considered at the same time. Currently, it is the most commonly used method for estimation of gene expression levels108. DEGS were calculated by DESeq2, heatmaps of DEGS were generated using R (Version 3.0.3) ggplot2 package, pheatmap package. For gene set enrichment analysis, GSEA 4.3.3 by UC San Diego and Broad Institute (https://www.gsea-msigdb.org) was utilized109,110.
RNA sequencing for bulk tissue
RNA sequencing was conducted by BGI (Shenzen, China). RNA isolated from bulk tissue of ipsilateral and contralateral insult tissue (Wildtype sham n = 7, wildtype TBI ipsi n = 7 and contra n = 7) with a visible hematoma was subjected to 100-bp single-end sequencing with a BGISEQ-500 platform. For each sample, a sequencing depth of at least 48 million clean reads was applied. In brief, poly-A-containing mRNA molecules were purified using poly-T oligo-attached magnetic beads. Afterward, mRNA is fragmented into small pieces and copied into first-strand cDNA using reverse transcriptase and random primers, which is followed by second-strand cDNA synthesis via DNA Polymerase I and RNase H. Fragments were subsequently ligated to the adaptor, the products purified and enriched and pooled to generate a final library containing single-strand DNA circles (ssDNA circles). These ssDNA circles were used to generate DNA nanoballs, loaded into patterned nanoarrays, and read through the sequencing platform. Reads were filtered and cleaned using the SOAPnuke software, for genome mapping, HISAT was applied. Novel coding transcripts were merged with reference transcripts and mapped using Bowtie2, gene expression levels for each sample were calculated with RSEM. Data analysis was conducted using DESeq2 algorithms to detect DEGs. DEGs were used to perform Gene Ontology (GO) and KEGG pathway classification. GO analysis was conducted with PANTHER overrepresentation test, FDR correction, and raw p < 0.05 (release 24.02.2021).
Flow cytometry
Living cells were transferred into a fixed volume, and numbers were determined using Neubauer improved counting chamber. Staining for flow cytometric phenotyping was performed in PBS + 5% FCS + 0.05% Sodium Azide (FACS staining buffer). Dissociation of brains was performed using the Multi Tissue Dissociation Kit 1 (Miltenyi Biotec, Germany) according to the manufacturer’s protocol, Fc receptors were blocked for 15 min (FcR Blocking Reagent mouse, 130-092-575, Miltenyi Biotec, Bergisch Gladbach, Germany), and cells incubated with the respective conjugated antibodies for 20 min. Flow cytometry was performed with the BD FACSCanto II analysis was performed using FlowJo™ 10.7.1 (FlowJo LLC, BD). Living cells were gated by a combination of forward/side scatter and DAPI. Total counts of lymphoid, CD11c+ and Ly6C+ myeloid populations were calculated using the gated frequencies and the priorly determined number of living cells.
The following antibodies were used for flow cytometry analysis:
APC anti-mouse ACSA-2 (REAfinity™; Miltenyi Biotec #130-116-249), PE anti-mouse CD3e (clone: 145-2C11; BioLegend #100308), APC anti-mouse CD4 (clone: GK1.5; BioLegend #100412), FITC anti-mouse CD8a (clone: 53-6.7; BioLegend #100706), BV711 anti-mouse/human CD11b (clone M1/70; BioLegend #101242), PE anti-mouse/human CD11b (clone: M1/70.15.11.5; Miltenyi Biotec #130-113-797) APC anti-mouse CD11b (M1/70; BioLegend #101212), APC/Fire™750 anti-mouse/human CD11b (BioLegend #101261), FITC anti-mouse CD11c (N418; BioLegend #117306), PE anti-mouse CD11c (clone: N418; BioLegend #117308), APC/Cy7 anti-mouse CD14 (Sa14-2; BioLegend #123318), PE anti-mouse CD44 (IM-7; BioLegend #103007), APC/Cyanine7 anti-mouse CD44 (IM7; BioLegend #103028), APC anti-mouse CD45 (30-F11; BioLegend #103112), BV605 anti-mouse CD45 (clone: 30-F11; BioLegend #103140), APC anti-mouse CD45R/B220 (clone: RA3-6B2; BioLegend #103212), PE anti-mouse CD206 (MMR) (clone: C068C2; BioLegend #141706), APC/Cyanine7 anti-mouse I-A/I-E (M5/114.15.2; BioLegend #107627), PE anti-mouse Ly-6C (HK1.4; BioLegend #128007), FITC anti-mouse Ly-6G (clone: 1A8; BioLegend #127606), APC anti-mouse Ly-6G (clone: 1A8; BioLegend #127614), PE anti-human/mouse/rat O4 (REAfinity™, Miltenyi Biotec, 130-117-711), PE human IgG1 (REAfinity™, Miltenyi Biotec #130-113-450), PE Rat IgG2b (RTK4530; kappa, BioLegend #400636), PE Rat IgG2c (RTK4174; kappa, BioLegend #400707), FITC Armenian Hamster IgG (HTK888; BioLegend #400906), APC Rat IgG2b (RTK4530; kappa, BioLegend #400612), APC/Cyanin7 Rat IgG1 (RTK2071; kappa, BioLegend #107627)
Immunoblot analysis
Tissue samples (cortical impact area) were snap frozen in liquid nitrogen, homogenized with a mortar and pestle, and lysed in KA-lysis buffer [25 mM Tris-HCl, 150 mM NaCl, 25 mM sodium pyrophosphate, 50 mM β-glycerophosphate, 50 mM NaF, 2 mM EGTA, 2 mM EDTA, 1 mM DTT, 10% glycerol, 1% Triton X-100 (pH 8.0)] supplemented with protease inhibitors (1 mM PMSF and complete mini tablet; Roche Diagnostics, Mannheim, Germany). After centrifugation (30 min, 16,000 × g), the supernatant was used as the total protein extract. Equal amounts of protein (30 µg) were separated by SDS-PAGE and transferred to nitrocellulose membranes. After blocking with 5% nonfat dry milk in TBS buffer for 1 h at room temperature, primary antibodies were incubated in blocking solution overnight at 4 °C or for 2 h at room temperature. After 3 washing steps, membranes were incubated with horseradish peroxidase–coupled secondary antibody for 1 h at room temperature. Membranes were exposed to ECL detection reagent (Thermo Fisher Scientific) and developed by ECL.
The following antibodies were used for immunoblot stainings:
Anti-TGM1 rabbit pAb (1:500, Abcam #ab103814), anti-Lipocalin-2 goat pAb (1:2000, R&D Systems #AF1857), anti-Osteopontin goat pAb (1:1000, R&D Systems #AF808), anti-Heme Oxygenase 1 rabbit pAb (1:2000, Abcam #ab13243), anti-NRF2 mouse mAb (1:1000, SantaCruz Biotechnology #sc-365949), anti-Galectin-3 rabbit pAb (SantaCruz Biotechnology #sc-20157), anti-IBA1 rabbit pAb (1:2000, FUJIFILM Wako #019-19741), anti-IKKβ rabbit mAb (1:1000, Cell Signaling Technology #8943), β-Actin rabbit mAb (1:500, Cell Signaling Technology #8457), HRP-linked anti-rabbit IgG (1:2000, Cell Signaling Technology #7074), anti-EAAT1 rabbit mAb (1:1000, Cell Signaling Technology #56845).
In-vivo MRI analysis
Longitudinal in vivo MRI was performed at 1, 15, and 30 days post injury (dpi) using a Bruker BioSpec 11.7T small animal MRI system. Animals were anesthetized with 1.5–2% isoflurane in oxygen, administered via a nose cone, and positioned securely on an MRI-compatible animal bed during imaging. Axial and coronal T2-weighted images were acquired using rapid acquisition with relaxation enhancement (RARE) sequences. For axial imaging, the parameters were: repetition time (TR) of 250 ms, echo time (TE) of 2.2 ms, slice thickness of 0.5 mm, 33 slices, flip angle of 15°, and a field of view (FOV) of 17 × 17 mm² with a matrix size of 260 × 260. For coronal imaging, the TR was 193 ms, TE was 5.0 ms, slice thickness remained 0.5 mm, 15 slices were acquired, the flip angle was 17.5°, and the FOV and matrix size were the same as for axial imaging. Four signal averages were collected for each scan to improve the signal-to-noise ratio. The bandwidth for image acquisition was set at 96 KHz. Images from these sequences were used for longitudinal lesion volume quantification, analysis and assessment of wound healing in IKK2-CAGFAP (n = 4), IKK2-DNGFAP (n = 3) and control (n = 6) cohorts. For evaluation of the wound closure process, lesion volumes were first quantified using the RadiAnt DICOM Viewer 2025.2 (Medixant). For that, areas which showed an obvious loss of tissue, tissue perturbations or formation of white spots were measured and used for calculations of the total volume. The lesion volume of day 1 was set to 100%, volumes of day 15 and day 30 were calculated as % of the remaining lesion volumes compared to day 1.
Comparison between the cohorts within each time point is carried out with 2-way ANOVA and Tukey’s multiple comparisons test.
Post-mortem sample preparation
For the analysis of EGFP expression and glial reaction to trauma, mice were sacrificed 1, 3, 7, or 15 days after the CHI or sham procedure (at least 3 mice per group). Only mice showing a visible imprint (or fracture) and hematoma formation after CHI were included in the study. For post-mortem analyses, animals were anesthetized with an IP injection of a mixture of ketamine (100 mg/kg) and xylazine (10 mg/kg) diluted in a 0.9% NaCl solution. For tissue fixation, transcardial perfusion was performed, first with 25 ml of ice-cold PBS to allow blood removal, followed by 50 ml of ice-cold 4% PFA in PBS. Brains were extracted and further postfixed in 4% PFA for 2 h, transferred to a solution of 30 % saccharose in PBS at 4 °C overnight for cryoprotection, and stored at 4 °C until use.
Histology and immunofluorescence staining
For immunohistochemistry, 20 µm-thick cryosections were used, and only the slides that showed a visible trauma core were included in the study. Twenty-micrometer-thick cryosectioned brain slices were permeabilized and blocked in a solution of 5% BSA in PBS with 0.5% Triton and incubated with primary antibodies in blocking solution overnight at 4 °C. After washing with PBS, slices were incubated with secondary antibodies in blocking solution for 2 h at room temperature. For nuclear antibodies, heat-mediated antigen retrieval was needed and was performed with sodium citrate (10 mM, pH 6.0, 0.05% Tween 20) before the incubation with primary antibody. When both nuclear and nonnuclear antibodies were used at the same time, only nonnuclear antibodies were incubated at first and were then fixed with 4% PFA before antigen retrieval. After antigen retrieval, the slices were incubated with both nuclear and nonnuclear antibodies. Following primary antibodies were used: chicken α-GFP (1:250, Aves Lab #GFP-1020), rabbit α-NG2 (1:250, Merck #AB5320), rat α-BrdU (1:250, Abcam #ab6326), guinea pig α-Iba1 (1:500, Synaptic Systems #234004), mouse α-GFAP (1:500, Sigma-Aldrich #G3893), rabbit α-ALDH1L1 (1:250, Abcam #ab87117). The following secondary antibodies were used for immunodetection of the primary antibodies: α-chicken Alexa Fluor® 488 (1:500, Thermo Fisher #A11039), α-rat Alexa Fluor® 488 (1:500, Thermo Fisher #A21208), α-mouse IgG Alexa Fluor® 647 (1:500, Dianova #115-606-072), α-guinea pig IgG Alexa Fluor® 647 (1:500, Thermo Fisher #A21450), α-rabbit Cy® 3 (1:500, Dianova #711-165-152). Afterward, slices were washed with PBS and incubated with DAPI at a 1:1000 dilution for 10 min for nuclear detection and then mounted on a slide with aqua-poly/mount for imaging.
Image acquisition and cellular quantification analyses
For the study of EGFP-expressing cells, overview images of the trauma area were acquired with the Keyence BZ-9000 BioRevo microscope (Keyence, Neu-Isenburg, Germany) with filters for DAPI, FITC/ Alexa Fluor 488, Texas Red/ Alexa Fluor 568/594 and Cy5, in a ×20 magnification and 3 mice per time point were included in the study. For the quantification of EGFP-expressing cells, image acquisition was performed with a Leica SPE confocal microscope with ×40 magnification (area: 275,27 × 275,27 µm per image) and analyzed through the Fiji software (ImageJ). For each immunostaining quantified, at least 6 different images from 2 to 3 consecutive slides containing the trauma core were used for quantification. Cells were quantified if positive for EGFP and a cellular marker (NG2 for NG2-glia, GFAP or Aldh1l1 for astrocytes, Iba1 for microglia), and their nucleus was positive for DAPI counterstaining. The total number of individuated cells per mm2 was reported.
For the quantification of the trauma area, 3 images per mouse were acquired with the Keyence BZ-9000 BioRevo microscope at a ×10 or ×20 magnification, and the area of trauma was visible as an area with DAPI staining (although nuclei were smaller, indicating possible cell death), but no glial markers. Similarly, overviews of the ipsilateral hemisphere were imaged at a ×20 magnification with the Keyence BZ-9000 BioRevo microscope for the analysis of the fibronectin and immunoglobin stainings, as well as for the quantification of the GFAP/CSPG4 scar.
For the quantification of gliosis, mages were acquired in the area adjacent to the CHI, defined as an area with morphologically different nuclei and without clear cellular staining. The border of the CHI is where gliosis takes place, and the scar is forming. For this study, three different images per immunostaining were quantified, and 3–5 different mice per timepoint were analyzed. Image acquisition was performed with a Leica SPE confocal microscope with ×40 magnification (area: 275,27 × 275,27 µm per image) and analyzed through the Fiji software (ImageJ). Glial cells were identified by the presence of DAPI-positive nuclei and positivity to a cell-specific marker (NG2 for NG2-glia, GFAP for astrocytes, Iba1 for microglia). For the quantification of glial proliferation, a cell-specific marker was used for cellular identification and nuclear positivity for BrdU was double-checked with DAPI. To distinguish NG2-glia from NG2+ pericytes, the presence of cellular processes and distance from a blood vessel was assessed. All data were analyzed in a blinded manner, and quantifications are reported as mean ± SEM. No statistical method was used to determine the sample size, and they were purely based on previous studies in the field.
Statistical analysis
Analysis of RNAseq data was conducted using NovoMagic, a free online RNAseq tool provided by Novogene (Version of April 2024). DEGs were defined as p < 0.05, |log2FC| > 0, for GSEA and GO term analysis, significant hits were defined as FDR < 0.05. Immunoblot densitometry was analyzed with Image J. Statistical analyses were performed with Prism software (GraphPad, San Diego, CA, USA) and indicated in the specific figure legend. Normality and Gaussian distribution were assumed and not tested separately. Unless indicated otherwise, one- or 2-way ANOVA with subsequent Tukey’s multiple comparisons test was used to compare independent measurements at one or different time points, respectively. Data were considered significant at p < 0.05.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The authors declare that the data supporting this study are presented in the article and Supplementary information files. Source data are provided with this paper. RNA sequencing data generated in this study have been deposited in the GEO database under accession codes GSE276182 and GSE276422. Source data are provided with this paper.
References
Maas, A. I. R. et al. Traumatic brain injury: integrated approaches to improve prevention, clinical care, and research. Lancet Neurol. 16, 987–1048 (2017).
Brett, B. L., Gardner, R. C., Godbout, J., Dams-O’Connor, K. & Keene, C. D. Traumatic brain injury and risk of neurodegenerative disorder. Biol. Psychiatry 91, 498–507 (2022).
Jassam, Y. N., Izzy, S., Whalen, M., McGavern, D. B. & El Khoury, J. Neuroimmunology of traumatic brain injury: time for a paradigm shift. Neuron 95, 1246–1265 (2017).
Simon, D. W. et al. The far-reaching scope of neuroinflammation after traumatic brain injury. Nat. Rev. Neurol. 13, 171–191 (2017).
Dimou, L. & Götz, M. Glial cells as progenitors and stem cells: new roles in the healthy and diseased brain. Physiol. Rev. 94, 709–737 (2014).
Burda, J. E., Bernstein, A. M. & Sofroniew, M. V. Astrocyte roles in traumatic brain injury. Exp. Neurol. 275, 305–315 (2016).
Verkhratsky, A. et al. Astrocytes in human central nervous system diseases: a frontier for new therapies. Signal Transduct. Target. Ther. 8, 396 (2023).
Liddelow, S. A., Olsen, M. L. & Sofroniew, M. V. Reactive astrocytes and emerging roles in central nervous system (CNS) disorders. Cold Spring Harb. Perspect. Biol. 16, a041356 (2024)
Brandebura, A. N., Paumier, A., Onur, T. S. & Allen, N. J. Astrocyte contribution to dysfunction, risk and progression in neurodegenerative disorders. Nat. Rev. Neurosci. 24, 23–39 (2023).
Patani, R., Hardingham, G. E. & Liddelow, S. A. Functional roles of reactive astrocytes in neuroinflammation and neurodegeneration. Nat. Rev. Neurol. 19, 395–409 (2023).
Hinz, M. & Scheidereit, C. The IκB kinase complex in NF-κB regulation and beyond. EMBO Rep. 15, 46–61 (2014).
Kaltschmidt, B. & Kaltschmidt, C. NF-B in the nervous system. Cold Spring Harb. Perspect. Biol. 1, a001271–a001271 (2009).
Zhang, Q., Lenardo, M. J. & Baltimore, D. 30 years of NF-κB: a blossoming of relevance to human pathobiology. Cell 168, 37–57 (2017).
Glass, C. K., Saijo, K., Winner, B., Marchetto, M. C. & Gage, F. H. Mechanisms underlying inflammation in neurodegeneration. Cell 140, 918–934 (2010).
Baker, R. G., Hayden, M. S. & Ghosh, S. NF-κB, inflammation, and metabolic disease. Cell Metab. 13, 11–22 (2011).
Kaltschmidt, B., Helweg, L. P., Greiner, J. F. W. & Kaltschmidt, C. NF-κB in neurodegenerative diseases: recent evidence from human genetics. Front. Mol. Neurosci. 15, 954541 (2022).
Burda, J. E. et al. Divergent transcriptional regulation of astrocyte reactivity across disorders. Nature 606, 557–564 (2022).
Clayton, B. L. L. et al. A phenotypic screening platform for identifying chemical modulators of astrocyte reactivity. Nat. Neurosci. 27, 656–665 (2024).
Leng, K. et al. CRISPRi screens in human iPSC-derived astrocytes elucidate regulators of distinct inflammatory reactive states. Nat. Neurosci. 25, 1528–1542 (2022).
Lee, H.-G. et al. Disease-associated astrocyte epigenetic memory promotes CNS pathology. Nature 627, 865–872 (2024).
Linnerbauer, M., Wheeler, M. A. & Quintana, F. J. Astrocyte crosstalk in CNS inflammation. Neuron 108, 608–622 (2020).
Huber-Lang, M., Lambris, J. D. & Ward, P. A. Innate immune responses to trauma. Nat. Immunol. 19, 327–341 (2018).
Kusiak, A. & Brady, G. Bifurcation of signalling in human innate immune pathways to NF-kB and IRF family activation. Biochem. Pharmacol. 205, 115246 (2022).
Mettang, M. et al. IKK2/NF-κB signaling protects neurons after traumatic brain injury. FASEB J. 32, 1916–1932 (2018).
Oeckl, P., Lattke, M., Wirth, T., Baumann, B. & Ferger, B. Astrocyte-specific IKK2 activation in mice is sufficient to induce neuroinflammation but does not increase susceptibility to MPTP. Neurobiol. Dis. 48, 481–487 (2012).
Nespoli, E. et al. Glial cells react to closed head injury in a distinct and spatiotemporally orchestrated manner. Sci. Rep. 14, 2441 (2024).
Tomann, P., Paus, R., Millar, S. E., Scheidereit, C. & Schmidt-Ullrich, R. Lhx2 is a direct NF-κB target gene that promotes primary hair follicle placode down-growth. Development 143, 1512–1522 (2016).
Schlett, J. S. et al. NF-κB is a critical mediator of post-mitotic senescence in oligodendrocytes and subsequent white matter loss. Mol. Neurodegener. 18, 24 (2023).
Lattke, M. et al. Transient IKK2 activation in astrocytes initiates selective non-cell-autonomous neurodegeneration. Mol. Neurodegener. 12, 16 (2017).
Crenn, P. et al. Changes in weight after traumatic brain injury in adult patients: a longitudinal study. Clin. Nutr. 33, 348–353 (2014).
Sosnoff, J. J., Broglio, S. P. & Ferrara, M. S. Cognitive and motor function are associated following mild traumatic brain injury. Exp. Brain Res. 187, 563–571 (2008).
Corrigan, F., Wee, I. C. & Collins-Praino, L. E. Chronic motor performance following different traumatic brain injury severity—a systematic review. Front. Neurol. 14, 180353 (2023).
Flierl, M. A. et al. Mouse closed head injury model induced by a weight-drop device. Nat. Protoc. 4, 1328–1337 (2009).
Walter, J. et al. The CatWalk XT® is a valid tool for objective assessment of motor function in the acute phase after controlled cortical impact in mice. Behav. Brain Res. 392, 112680 (2020).
Timotius, I. K. et al. CatWalk XT gait parameters: a review of reported parameters in pre-clinical studies of multiple central nervous system and peripheral nervous system disease models. Front. Behav. Neurosci. 17, 1147784 (2023).
O’Shea, T. M. et al. Derivation and transcriptional reprogramming of border-forming wound repair astrocytes after spinal cord injury or stroke in mice. Nat. Neurosci. https://doi.org/10.1038/s41593-024-01684-6 (2024).
Zamanian, J. L. et al. Genomic analysis of reactive astrogliosis. J. Neurosci. 32, 6391–6410 (2012).
Liddelow, S. A. et al. Neurotoxic reactive astrocytes are induced by activated microglia. Nature 541, 481–487 (2017).
Saul, D. et al. A new gene set identifies senescent cells and predicts senescence-associated pathways across tissues. Nat. Commun. 13, 4827 (2022).
Coppé, J.-P., Desprez, P.-Y., Krtolica, A. & Campisi, J. The senescence-associated secretory phenotype: the dark side of tumor suppression. Annu. Rev. Pathol. Mech. Dis. 5, 99–118 (2010).
Pulido, T., Velarde, M. C. & Alimirah, F. The senescence-associated secretory phenotype: fueling a wound that never heals. Mech. Ageing Dev. 199, 111561 (2021).
Böttcher, C. et al. Human microglia regional heterogeneity and phenotypes determined by multiplexed single-cell mass cytometry. Nat. Neurosci. 22, 78–90 (2019).
Korin, B. et al. High-dimensional, single-cell characterization of the brain’s immune compartment. Nat. Neurosci. 20, 1300–1309 (2017).
Kann, O., Almouhanna, F. & Chausse, B. Interferon γ: a master cytokine in microglia-mediated neural network dysfunction and neurodegeneration. Trends Neurosci. 45, 913–927 (2022).
Guo, H., Callaway, J. B. & Ting, J. P.-Y. Inflammasomes: mechanism of action, role in disease, and therapeutics. Nat. Med. 21, 677–687 (2015).
Ravichandran, K. A. & Heneka, M. T. Inflammasomes in neurological disorders—mechanisms and therapeutic potential. Nat. Rev. Neurol. https://doi.org/10.1038/s41582-023-00915-x (2024).
Yao, J., Wang, Z., Song, W. & Zhang, Y. Targeting NLRP3 inflammasome for neurodegenerative disorders. Mol. Psychiatry 28, 4512–4527 (2023).
van Velthoven, C. T. J., Heijnen, C. J., van Bel, F. & Kavelaars, A. Osteopontin enhances endogenous repair after neonatal hypoxic–ischemic brain injury. Stroke 42, 2294–2301 (2011).
Shi, L. et al. Treg cell-derived osteopontin promotes microglia-mediated white matter repair after ischemic stroke. Immunity 54, 1527–1542 (2021).
Daglas, M. & Adlard, P. A. The involvement of iron in traumatic brain injury and neurodegenerative disease. Front. Neurosci. 12, 981 (2018).
Lan, Y. et al. Fate mapping of Spp1 expression reveals age-dependent plasticity of disease-associated microglia-like cells after brain injury. Immunity 57, 349–363 (2024).
Adler, O. et al. Reciprocal interactions between innate immune cells and astrocytes facilitate neuroinflammation and brain metastasis via lipocalin-2. Nat. Cancer 4, 401–418 (2023).
Khatri, N. et al. The complexity of secondary cascade consequent to traumatic brain injury: pathobiology and potential treatments. Curr. Neuropharmacol. 19, 1984–2011 (2021).
Wang, Z.-R. et al. Regulating effect of activated NF-κB on edema induced by traumatic brain injury of rats. Asian Pac. J. Trop. Med. 9, 274–277 (2016).
Khorooshi, R., Babcock, A. A. & Owens, T. NF-κB-driven STAT2 and CCL2 expression in astrocytes in response to brain injury. J. Immunol. 181, 7284–7291 (2008).
Cieri, M. B., Villarreal, A., Gomez-Cuautle, D. D., Mailing, I. & Ramos, A. J. Progression of reactive gliosis and astroglial phenotypic changes following stab wound-induced traumatic brain injury in mice. J. Neurochem. 167, 183–203 (2023).
Burda, J. E. & Sofroniew, M. V. Reactive gliosis and the multicellular response to CNS damage and disease. Neuron 81, 229–248 (2014).
Wahane, S. & Sofroniew, M. V. Loss-of-function manipulations to identify roles of diverse glia and stromal cells during CNS scar formation. Cell Tissue Res. 387, 337–350 (2022).
Guo, S. & DiPietro, L. A. Factors affecting wound healing. J. Dent. Res. 89, 219–229 (2010).
Anderson, M. A. et al. Astrocyte scar formation aids central nervous system axon regeneration. Nature 532, 195–200 (2016).
Wang, L. et al. Spatial transcriptomics of the aging mouse brain reveals origins of inflammation in the white matter. Nat. Commun. 16, 3231 (2025).
Wu, C. et al. Spatially resolved transcriptome of the aging mouse brain. Aging Cell 23, e14109 (2024).
Allen, W. E., Blosser, T. R., Sullivan, Z. A., Dulac, C. & Zhuang, X. Molecular and spatial signatures of mouse brain aging at single-cell resolution. Cell 186, 194–208 (2023).
Bauer, M. E., Pawelec, G. & Paganelli, R. Neuroimmunology and ageing–the state of the art. Immun. Ageing 21, 5 (2024).
Palmer, A. L. & Ousman, S. S. Astrocytes and aging. Front. Aging Neurosci. 10, 337 (2018).
Zhu, Y., Carvey, P. M. & Ling, Z. Age-related changes in glutathione and glutathione-related enzymes in rat brain. Brain Res. 1090, 35–44 (2006).
Detcheverry, F., Senthil, S., Narayanan, S. & Badhwar, A. Changes in levels of the antioxidant glutathione in brain and blood across the age span of healthy adults: a systematic review. Neuroimage Clin. 40, 103503 (2023).
Clarke, L. E. et al. Normal aging induces A1-like astrocyte reactivity. Proc. Natl. Acad. Sci. USA 115, E1896–E1905 (2018).
Hashimoto, M., Sun, D., Rittling, S. R., Denhardt, D. T. & Young, W. Osteopontin-deficient mice exhibit less inflammation, greater tissue damage, and impaired locomotor recovery from spinal cord injury compared with wild-type controls. J. Neurosci. 27, 3603–3611 (2007).
Ladwig, A. et al. Osteopontin attenuates secondary neurodegeneration in the thalamus after experimental stroke. J. Neuroimmune Pharmacol. 14, 295–311 (2019).
Rabenstein, M. et al. Osteopontin mediates survival, proliferation and migration of neural stem cells through the chemokine receptor CXCR4. Stem Cell Res. Ther. 6, 99 (2015).
Jin, Y.-C. et al. Intranasal delivery of RGD motif-containing osteopontin icosamer confers neuroprotection in the postischemic brain via αvβ3 integrin binding. Mol. Neurobiol. 53, 5652–5663 (2016).
Davaanyam, D., Kim, I.-D. & Lee, J.-K. Intranasal delivery of RGD-containing osteopontin heptamer peptide confers neuroprotection in the ischemic brain and augments microglia M2 polarization. Int. J. Mol. Sci. 22, 9999 (2021).
Icer, M. A. & Gezmen-Karadag, M. The multiple functions and mechanisms of osteopontin. Clin. Biochem. 59, 17–24 (2018).
Yim, A., Smith, C. & Brown, A. M. Osteopontin/secreted phosphoprotein-1 harnesses glial-, immune-, and neuronal cell ligand-receptor interactions to sense and regulate acute and chronic neuroinflammation. Immunol. Rev. 311, 224–233 (2022).
Lin, E. Y.-H., Xi, W., Aggarwal, N. & Shinohara, M. L. Osteopontin (OPN)/SPP1: from its biochemistry to biological functions in the innate immune system and the central nervous system (CNS). Int. Immunol. 35, 171–180 (2023).
Barriere, G., Fici, P., Gallerani, G., Fabbri, F. & Rigaud, M. Epithelial mesenchymal transition: a double-edged sword. Clin. Transl. Med. 4, 14 (2015).
Kothari, A. et al. Osteopontin—a master regulator of epithelial-mesenchymal transition. J. Clin. Med. 5, 39 (2016).
Patten, J. & Wang, K. Fibronectin in development and wound healing. Adv. Drug Deliv. Rev. 170, 353–368 (2021).
Budatha, M., Zhang, J. & Schwartz, M. A. Fibronectin-mediated inflammatory signaling through integrin α5 in vascular remodeling. J. Am. Heart Assoc. 10, e021160 (2021).
Stoffels, J. M. J., Zhao, C. & Baron, W. Fibronectin in tissue regeneration: timely disassembly of the scaffold is necessary to complete the build. Cell. Mol. Life Sci. 70, 4243–4253 (2013).
Morita, A. et al. Temporal evolution of heme oxygenase-1 expression in reactive astrocytes and microglia in response to traumatic brain injury. Brain Hemorrhages 1, 65–74 (2020).
Chang, E. F., Claus, C. P., Vreman, H. J., Wong, R. J. & Noble-Haeusslein, L. J. Heme regulation in traumatic brain injury: relevance to the adult and developing brain. J. Cereb. Blood Flow Metab. 25, 1401–1417 (2005).
Russell, N. H. et al. Time-dependent hemeoxygenase-1, lipocalin-2 and ferritin induction after non-contusion traumatic brain injury. Brain Res. 1725, 146466 (2019).
Roh, J. S. & Sohn, D. H. Damage-associated molecular patterns in inflammatory diseases. Immune Netw. 18, e27 (2018).
Wei, Z. et al. Broadening horizons: ferroptosis as a new target for traumatic brain injury. Burns Trauma 12, tkad051 (2024).
Cousar, J. L. et al. Heme oxygenase 1 in cerebrospinal fluid from infants and children after severe traumatic brain injury. Dev. Neurosci. 28, 342–347 (2006).
Zhao, R.-Y. et al. Role of lipocalin 2 in stroke. Neurobiol. Dis. 179, 106044 (2023).
Kim, J.-H. et al. Lipocalin-2 is a key regulator of neuroinflammation in secondary traumatic and ischemic brain injury. Neurotherapeutics 20, 803–821 (2023).
Jung, B.-K. & Ryu, K.-Y. Lipocalin-2: a therapeutic target to overcome neurodegenerative diseases by regulating reactive astrogliosis. Exp. Mol. Med. 55, 2138–2146 (2023).
Tatsukawa, H. & Hitomi, K. Role of transglutaminase 2 in cell death, survival, and fibrosis. Cells 10, 1842 (2021).
Tripathy, D. et al. Increased transcription of transglutaminase 1 mediates neuronal death in in vitro models of neuronal stress and Aβ1–42-mediated toxicity. Neurobiol. Dis. 140, 104849 (2020).
Frik, J. et al. Cross-talk between monocyte invasion and astrocyte proliferation regulates scarring in brain injury. EMBO Rep. 19, EMBR201745294 (2018).
Kjell, J. & Götz, M. Filling the gaps–a call for comprehensive analysis of extracellular matrix of the glial scar in region- and injury-specific contexts. Front. Cell. Neurosci. 14, 32 (2020).
Amann, L., Masuda, T. & Prinz, M. Mechanisms of myeloid cell entry to the healthy and diseased central nervous system. Nat. Immunol. https://doi.org/10.1038/s41590-022-01415-8 (2023).
Mundt, S., Greter, M. & Becher, B. The CNS mononuclear phagocyte system in health and disease. Neuron 110, 3497–3512 (2022).
Endo, S., Sakamoto, Y., Kobayashi, E., Nakamura, A. & Takai, T. Regulation of cytotoxic T lymphocyte triggering by PIR-B on dendritic cells. Proc. Natl. Acad. Sci. USA 105, 14515–14520 (2008).
Neumann, H. Cytotoxic T lymphocytes in autoimmune and degenerative CNS diseases. Trends Neurosci. 25, 313–319 (2002).
Hu, D., Xia, W. & Weiner, H. L. CD8+ T cells in neurodegeneration: friend or foe? Mol. Neurodegener. 17, 59 (2022).
Reagin, K. L. & Funk, K. E. The role of antiviral CD8+ T cells in cognitive impairment. Curr. Opin. Neurobiol. 76, 102603 (2022).
Zhang, Z., Duan, Z. & Cui, Y. CD8+ T cells in brain injury and neurodegeneration. Front. Cell. Neurosci. 17, 1281763 (2023).
Javidi, E. & Magnus, T. Autoimmunity after ischemic stroke and brain injury. Front. Immunol. 10, 686 (2019).
Worbs, T., Hammerschmidt, S. I. & Förster, R. Dendritic cell migration in health and disease. Nat. Rev. Immunol. 17, 30–48 (2017).
Needham, E. J. et al. Complex autoantibody responses occur following moderate to severe traumatic brain injury. J. Immunol. 207, 90–100 (2021).
Ouali Alami, N. et al. NF-κB activation in astrocytes drives a stage-specific beneficial neuroimmunological response in ALS. EMBO J. 37, EMBJ201798697 (2018).
Herrmann, O. et al. IKK mediates ischemia-induced neuronal death. Nat. Med. 11, 1322–1329 (2005).
Lattke, M., Magnutzki, A., Walther, P., Wirth, T. & Baumann, B. Nuclear factor κB activation impairs ependymal ciliogenesis and links neuroinflammation to hydrocephalus formation. J. Neurosci. 32, 11511–11523 (2012).
Mortazavi, A., Williams, B. A., McCue, K., Schaeffer, L. & Wold, B. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat. Methods 5, 621–628 (2008).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 102, 15545–15550 (2005).
Mootha, V. K. et al. PGC-1α-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat. Genet. 34, 267–273 (2003).
Acknowledgements
The authors thank Petra Weihrich, Mareike Wirth, Diana Giesler, Maysam Hassib Hussein and Sonja Braumüller for their excellent technical assistance. We also thank Dr. Gudrun Strauss for her help with optimization of gating strategies in flow cytometry analysis. Tabea M. Hein and Marsela Hakani are members of the International Graduate School in Molecular Medicine Ulm (IGradU). This research was funded by the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation) Collaborative Research Center CRC1149 (ID-251293561) to L.D., M.H.-L., and T.W. This work was further supported by the CRC1506 (ID-450627322) to L.D. and F.O./B.B.
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
B.B., T.W. conceptualized, coordinated, and supervised the project. T.W. administrated the project. B.B. planned and organized mouse breeding. T.M.H. performed Catwalk, FACS, biochemical experiments and data analysis. E.N., M.H. performed immunohistochemistry, brain imaging, cellular quantifications, and data analysis. T.M.H., E.N., H.W., M.L., F.O., L.D., T.W., and B.B. wrote and edited the original manuscript. T.M.H., H.W., E.N., M.M., S.N.M., M.H., and M.L. carried out CHI/TBI, M.H.-L. advised on TBI protocol and provided equipment. M.H. performed immunohistochemistry and data analysis. A.A., V.R. performed MRI experiments and analyzed data. D.Y., J.S.S., J.J., and V.P. performed biochemical experiments and analyzed data. T.M.H., M.T., and K.T. analyzed RNAseq data. L.D., F.O., and M.L. provided intellectual discussion and corrected the manuscript. All authors discussed the results, reviewed and edited the manuscript. All authors agreed to the submitted version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hein, T.M., Nespoli, E., Hakani, M. et al. NF-κB activation in astrocytes impairs wound healing after traumatic brain injury in male mice. Nat Commun 17, 2323 (2026). https://doi.org/10.1038/s41467-026-70304-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-70304-7