Abstract
The symbiogenetic origin of organelles, such as chloroplasts, is established, and organelle genomes provide evidence of prokaryotic ancestry. Nevertheless, most organelle proteins are nuclear-encoded and function in concert with those expressed from the organelle genomes, representing constitutive molecular chimerism. The evolutionary forces generating chimerism have been widely discussed, but without much evidence. Here we provide biochemical evidence of transient molecular chimerism in nature, along with a possible mechanistic explanation for chimerism. In the flagellate, Rapaza viridis, nuclear-encoded proteins support photosynthesis in transient, xenogeneic chloroplasts (kleptoplasts) acquired from the green alga Tetraselmis sp. We focused on two putative kleptoplast-targeted proteins: a RuBisCO small subunit-like protein (RvRbcS-like) and a RuBisCO activase homologue. Immunofluorescence microscopy confirmed the kleptoplast localization of the proteins, and the knockdown and knockout experiments demonstrated impaired photosynthesis, particularly for RvRbcS-like. The unique carboxyl-terminal extension of the RvRbcS-like protein suggests that it has an additional role in pyrenoid reorganization, a key step in kleptoplast remodelling. Protein translocation into kleptoplasts requires rapid, de novo assembly of transport systems after each acquisition, unlike the constitutive chimerism of established organelles. This previously unreported phenomenon in eukaryotes positions R. viridis as a unique, genetically tractable model for investigating the molecular and evolutionary origins of organelles.
Similar content being viewed by others
Introduction
Eukaryotes are inherently chimeric. Although the core system originated in archaea, many eukaryote proteins were derived from different bacteria of another domain1, and eukaryote organelles, such as mitochondria and chloroplasts, resulted from the cellular-level integration of other bacteria. Each integration arose in a specific physiological context, which was probably endosymbiotic, where selection operated at that moment, and the chimeric states found in modern eukaryotes are constitutive. Therefore, it is only natural that theories on organellogeneses2,3,4,5,6,7,8,9 hinge on two extremes: modern endosymbiotic examples with little molecular-level integration10 and genetically fully integrated organelles, although the proposed evolutionary processes are not well evidenced. Nevertheless, there are some exceptional and unique opportunities for evolutionary studies into the transient integrations of xenogeneic bodies in some eukaryotic cells, such as the phenomenon called kleptoplasty.
Kleptoplasty represents a distinct form of algal exploitation, in which the host cells temporarily acquire chloroplasts from other algal cells, presumably to exploit them for phototrophy, and the stolen plastids are called kleptoplasts11. Kleptoplasty has been reported in a wide range of eukaryotes and, importantly, demonstrates a wide variety of symbiogenetic modes. Some well-known examples of multicellular host organisms include the sacoglossan sea slugs12,13. However, unicellular hosts are more common and have been reported in taxa with diverse dinoflagellates14,15,16,17,18, ciliates19,20,21, and foraminiferans22,23, as well as a kathablepharid24, a centrohelid25, and a euglenozoan9. Unlike mutualistic algal endosymbiosis, the kleptoplasts that confer phototrophy to the host are extracted from the original algal cells and can be considered as symbiogenetic in nature, functioning as transient organelles. However, the molecular basis of kleptoplast maintenance differs among cases26,27. For example, in the ciliate Mesodinium rubrum21,28 and the dinoflagellates Nusuttodinium aeruginosum29 and Durinskia capensis18, the nuclei of the kleptoplast donor algae are retained in the host cytoplasm along with the kleptoplasts. These kleptoplasts seem to be temporarily maintained by the algal nuclei-encoded proteins, whose expression may or may not be modulated by the host29,30.
In other intriguing cases, even without algal nuclei, partial genetic integrations have been proposed based on transcriptomic data to be involved in prolonged maintenance of active kleptoplasts by asserting the expression of host nuclear-encoded proteins in the xenogeneic kleptoplast interior9,26,31,32,33. However, these hypotheses lacked biochemical confirmation. Approximately 60 candidate kleptoplast-targeted proteins were proposed in Dinophysis acuminate, which included several components of the thylakoid photosynthetic apparatus, sugar transporter, and some enzymes involved in pigment biosynthesis31,32. In the Ross Sea dinoflagellate, some photosynthetic proteins encoded by the host nuclear genome may target and function in the kleptoplasts, especially the photosystem I subunits, allowing temporary establishment of cyclic electron transport33.
Recent findings have revealed an outstanding case of kleptoplasty in the euglenozoan flagellate Rapaza viridis9,34,35. This organism obtains its kleptoplasts from the green alga Tetraselmis sp. through phagocytosis. Although it retains the chloroplasts, the algal nucleus is expelled together with the cytoplasm shortly after its ingestion. The resulting kleptoplast is then split into smaller pieces and distributed to the daughter cells, without being enlarged, maintaining high photosynthetic performance for about 2 weeks before declining (Fig. 1)9. Most importantly, the R. viridis transcriptome contained more chloroplast-related genes than ever reported, with 274 identified according to the most conservative estimate9, each encoding distinctive amino-terminal (N-terminal) sequences similar to the bipartite targeting signals found in Euglenophyceae algae, the sister clade of R. viridis possessing secondary chloroplasts derived from Pyramimonas9,36. Therefore, the phenomena observed in the kleptoplasty of R. viridis, such as pyrenoid reorganization and multiplication, persistence of photosynthetic activity, and accumulation of photosynthetic products in the host cytoplasm, were tentatively attributed to genes that are presumably acquired by horizontal gene transfer from various algae9. However, in silico predictions alone are insufficient to prove protein import and elucidate its mechanistic underpinnings, highlighting the need for cellular-level biochemical investigations34.
Chloroplasts/kleptoplasts are shown in green, and pyrenoids are shown in yellow. a R. viridis ingests whole cells of a specific Tetraselmis strain by phagocytosis (ingestion event), typically consuming them within several hours at a 1:3 cell ratio. b In the kleptoplast transformation stage (15–20 h after ingestion), R. viridis expels the algal cytoplasm and nucleus while retaining the chloroplasts as three-membrane-bound kleptoplasts. The large, starch-sheathed algal pyrenoid then transforms into multiple Rapaza pyrenoids associated with thylakoid membranes35. The kleptoplasts are further subdivided, possibly by constricting the outermost phagosomal membrane. c During the phototrophic stage, kleptoplasts containing one or more pyrenoids are inherited by daughter cells during exponential growth. R. viridis remains highly photosynthetic for approximately 2 weeks, remaining virtually autotrophic through a 1-week growth phase and a 1-week stationary phase. This autotrophy is supported by suspended growth without external inorganic nitrogen34, culture decline in the absence of light9, and cytosolic polysaccharide accumulation in stationary phase cells9,34. Kleptoplasts do not grow or replicate during this period. d In the absence of new kleptoplast acquisition, vacuoles begin to form after 3 weeks and gradually expand to occupy most of the cell. The cells ultimately die after 4–5 weeks (declining stage). See Karnkowska et al.9 for further details. e A Venn diagram showing the distribution of genes encoding proteins involved in key photosynthetic processes in the chloroplast genome, as well as in the nuclear genomes of R. viridis and Tetraselmis sp.
Here, we investigated kleptoplasty in R. viridis at the protein level, focusing on the intracellular localization and function of host nuclear-encoded genes. Using genetic engineering and biochemical approaches, we show that selected highly expressed proteins with predicted chloroplastic functions are translocated into kleptoplasts, where they are crucial for the effective operation of this xenogeneic organelle.
Results
Host chloroplastic gene expression
From the time-series transcriptome data taken from four distinct kleptoplastic stages (Fig. 1 and Supplementary data), we identified 37 R. viridis transcripts encoding proteins with conserved domains indicative of chloroplastic functions37. The expression levels of these transcripts were comparable to those of key mitochondrial metabolic genes (Supplementary Table 1 and Fig. 2a). The presence of a distinctive euglenid 5′-spliced leader sequence38 in these transcripts (Supplementary Table 2) and the mapping of their open reading frames (ORFs) to contigs in the draft genome, with or without introns conforming to the canonical GT(GC)-AG splice rule39 (Supplementary Table 3), verifies their genuine expression from the R. viridis nuclear genome. Phylogenetic analysis of the translated peptide sequences showed that about 40% clustered with Euglenophyceae proteins, consistent with previous reports9, and about 20% with Tetraselmis spp. (Supplementary Table 4). However, none of the 37 sequences were identical or highly homologous to the current kleptoplast donor strain Tetraselmis sp. NIES-4478. This suggests any horizontal transfers of these important genes occurred in the distant past, and the genes evolved as unique components of R. viridis.
a Thirty-seven R. viridis transcripts encoding well-predicted full-length domains associated with putative chloroplast functions37 (solid bars), along with six mitochondria-targeted transcripts corresponding to components of complex III and the pyruvate dehydrogenase complex (open bars), all exhibited similarly high maximum expression levels. b Normalized transcript counts across four timepoints in a time-series transcriptome dataset. The vertical axis shows normalized counts (mean; n = 3 biological replicates per time point) scaled to the maximum value of each gene; the horizontal axis marks the sampling points corresponding to the early and late kleptoplast transformation phases and two later timepoints in the growth and stationary phases of the phototrophic stage. The genes are grouped according to the timepoint of peak expression. c RNA-seq counts and qPCR measurements for RvRbcS-like (black) and RvRca-like (red). Solid squares with error bars represent normalized RNA-seq counts (mean ± SEM; n = 3 biological replicates per time point; left axis), and the solid circles represent normalized qPCR values (single measurement per time point; n = 1; right axis). qPCR confirmed that RvRbcS-like peaked during the late kleptoplast transformation phase, whereas RvRca-like peaked slightly later, between the late kleptoplast transformation phase and the growth phase. Source data are provided as a Source data file.
These putative proteins include core components of the thylakoid membrane photosynthetic machinery, including light-harvesting proteins and subunits of photosystem II, cytochrome b6f, and ATP synthase. Transcripts were also identified for proteins involved in the Calvin–Benson cycle, including ribulose 1,5-bisphosphate carboxylase/oxygenase (RuBisCO) and its related factors, thylakoid membrane proteases (FtsH homologs), a triose-phosphate/phosphate translocator (TPT), pentose-phosphate/phosphate translocators, and other enzymes. The peak transcript levels varied by gene (Supplementary Table 1). Although many transcripts, including those for the electron transport chain, peaked during the growth or stationary phases of the phototrophic stage, others peaked in the late kleptoplast transformation stage (Fig. 2b). From the late kleptoplast transformation stage, we selected two genes for further analysis: a homolog of the RuBisCO small subunit (RvRbcS-like) and of RuBisCO activase (RvRca-like). Quantitative polymerase chain reaction (qPCR) confirmed that RvRbcS-like expression peaked during the transformation stage, whereas RvRca-like expression peaked in the early to mid-growth phase (Fig. 2c). If transported into kleptoplasts and functionally active, these proteins are likely to be essential for carbon fixation in kleptoplastic photosynthesis.
Host protein kleptoplast localization
We confirmed the expression of RvRbcS-like and RvRca-like in R. viridis and their absence in their respective knockout strains by immunoblotting, with Tetraselmis sp. as a control (Fig. 3c). Custom anti-peptide antibodies were synthesized to recognize the corresponding amino acid sequences from R. viridis NIES-4477 specifically (Fig. 3a), and both proteins were detected in wild-type R. viridis but not in the knockout strains (ΔRvRbcS-like and ΔRvRca-like, respectively).
a, b Schematic representation of the translated products of RvRbcS-like and RvRca-like ORFs, respectively. Both peptides include an extended NtLCD. RvRbcS-like comprises four tandemly concatenated RbcS domains, each with slightly different sequences, connected by unique linker sequences, and an extended CtLCD enriched in repeated tetrapeptide motifs, such as SY[G/A/E/D/-]Q and SY[R/Q]P (highlighted in blue). The anti-peptide antibody target sequences are highlighted in red. c, d Immunoblotting of RvRbcS-like and RvRca-like using respective anti-peptide antibodies. The signals were exclusively detected in wild-type R. viridis and were absent in Tetraselmis sp. and the corresponding gene knockout strains, confirming the specificity of the antibodies. e Immunoblotting of wild-type and RvRbcS-uCt-insHA3 strains using anti-RvRbcS-like peptide antibody (1, 2) and anti-HA tag antibody (3, 4). No anti-HA signal was detected in the wild-type cells. By contrast, the HA-tagged strains showed a major band matching the RvRbcS-like signal, along with additional fragments, which were more clearly detected by anti-HA antibody. Equal protein loading was verified using Coomassie brilliant blue staining (Supplementary Fig. 1). f Fluorescence and bright-field images of RvRbcS-uCt-insHA3 and wild-type cells, respectively. Immunofluorescence signals indicating HA tag expression (yellow) were only detected in RvRbcS-uCt-insHA3 cells and were predominantly localized in the kleptoplast, as shown by the colocalization with chlorophyll autofluorescence (magenta). The HA-tagged RvRbcS-like signals (yellow) were centered and largely overlapped with RbcL (blue), suggesting their colocalization. Scale bars: 10 μm. g Merged fluorescence and bright-field images of wild-type and ΔRvRca-like cells, respectively. Immunofluorescence signals from the anti-RvRca-like peptide antibody (green) were exclusively detected in wild-type cells and localized within kleptoplasts, as indicated by chlorophyll autofluorescence (red). Scale bars: 10 μm. Source data are provided as a Source data file.
The observed peptide sizes were much smaller than predicted. RvRbcS-like encodes a 942-residue protein composed of four tandem RbcS domains (cd0352737) flanked by a 177-residue N-terminal low-complexity domain (NtLCD) and a 228-residue C-terminal low-complexity domain (CtLCD), with an expected molecular weight of ~105 kDa (Fig. 3a). However, immunoblotting revealed a predominant band a ~90 kDa, consistent with the removal of the ~18 kDa NtLCD. Full-length peptides were rarely observed; only one instance of a ~105 kDa band was found in an early experiment (Supplementary Fig. 2). Similarly, RvRca-like encodes a 542-residue protein containing a conserved Rca domain (PLN00020) and a 171-residue NtLCD, with a predicted size of ~59 kDa (Fig. 3b). However, the detected band was ~42 kDa (Fig. 3d), consistent with cleavage of the ~17 kDa NtLCD.
These findings strongly suggest that both proteins undergo NtLCD cleavage during or after translocation into the kleptoplast. Because chloroplast-targeting sequences are typically removed during import40,41, our data imply that these nuclear-encoded peptides are translocated into kleptoplasts via their NtLCDs and subsequently processed to their mature forms.
Next, we demonstrated the localization of these peptides within the kleptoplasts using immunofluorescence microscopy. We generated a genome-edited strain in which an epitope tag was fused to RvRbcS-like because the custom anti-peptide antibody targeting RvRbcS-like used above is unlikely to recognize the protein in its natural conformation, forming the RuBisCO complex. A 3×HA tag was inserted immediately downstream of the fourth putative RbcS domain, ensuring that the tag was exposed outside the putative complex (strain RvRbcS-uCt-insHA3) (Supplementary Fig. 3). The resulting peptide, ~90 kDa after NtLCD removal, was detected by immunoblotting of the total cellular proteins using an anti-HA antibody (Fig. 3e), confirming successful tag insertion. After staining with the anti-HA antibody, immunofluorescence microscopy of the HA-tagged RvRbcS strain revealed fluorescent signals localized just inside the kleptoplasts, demonstrating RvRbcS-like translocation into the kleptoplast (Fig. 3f). Multiplex observations using a generic anti-RuBisCO large subunit (RbcL) antibody showed significant overlap between the HA-tagged protein and RbcL signals (Fig. 3f). This observation strongly suggests that RvRbcS-like may function together with RuBisCO in the kleptoplast pyrenoid9,35. No HA signal was detected in wild-type cells (Fig. 3e, g), validating the specificity of the reporter immunofluorescence.
Similarly, immunofluorescence microscopy using the anti-RvRca-like antibody revealed signals localized in the kleptoplasts in wild-type cells (Fig. 3g), but not in ΔRvRca-like cells (Fig. 3g), confirming that the detected signal represented naturally folded RvRca-like. These findings demonstrate that RvRca-like is also translocated into the kleptoplasts.
Function of host-derived proteins
Next, we examined the phenotypic consequences of knocking out RvRbcS-like and RvRca-like by genome editing to assess their functional importance. In batch cultures of the ΔRvRbcS-like strain with precisely equal initial cell concentrations at the ingestion event (Fig. 1), ΔRvRbcS-like exhibited markedly reduced growth, reaching less than half of the wild-type level (Fig. 4a). Most ΔRvRbcS-like cells died by day 22, much earlier than the wild-type survival of approximately 5 weeks9. Additionally, the accumulation of cytoplasmic polysaccharide grains9,34,35 was markedly diminished in ΔRvRbcS-like compared with that in wild-type cells (Fig. 4b, d). Photosynthetic activity per cell in ΔRvRbcS-like was suppressed to about half of that of the wild-type activity (Fig. 4c). Dark oxygen consumption, a measure of respiratory activity, was also significantly reduced, suggesting suppressed cellular activity due to the reduced supply of photosynthetic products. Supporting these findings, RNA interference targeting RvRbcS-like also impaired photosynthesis (Supplementary Fig. 4). Although gene silencing was incomplete, the light–response curves showed that the net oxygen evolution per cell in the RNA interference-treated culture was significantly reduced to about half of the control level (Supplementary Fig. 4).
a Cell growth curves over 2 weeks following the ingestion event, normalized to the initial cell density of R. viridis (8.0 × 104 cells/mL), in batch cultures with three equal numbers of Tetraselmis sp. cells for phagocytosis (mean ± SEM; n = 3 biological replicates per time point; some error bars are smaller than the markers). b Quantification of insoluble polysaccharide grains per 105 R. viridis cells (mean ± SEM; n = 3 biological replicates per time point). c Light–response curves based on the oxygen evolution rates per 106 R. viridis cells. Negative values indicate oxygen consumption exceeding production, and the values at zero light intensity represent mitochondrial oxygen consumption (dark respiration rate) (mean ± SEM; n = 3 biological replicates per time point; some error bars are smaller than the markers). d Time-course observations of cell morphology and intracellular polysaccharide accumulation using differential interference contrast microscopy. Scale bars: 10 μm. The arrowheads on day 10 indicate host cytosolic polysaccharide grains. Wild-type cells accumulated polysaccharides after day 10, whereas ΔRvRbcS-like lacked grains and ΔRvRca-like showed partial reduction. e Immunoblotting of protein expression in wild-type and ΔRvRbcS-like strains. The anti-RvRbcS-like peptide antibody detected a ~91 kDa band in the wild-type strain only (top panel). The anti-RbcL antibody detected signals in all cells, but the levels decreased significantly after day 4 in ΔRvRbcS-like strains (second panel). The anti-TsRbcS peptide antibody detected signals in all cells but decreased markedly in both wild-type and ΔRvRbcS-like strains after day 4 (third panel). Equal protein loading was assessed by detecting α-tubulin on the same membrane after stripping and reprobing (bottom panel). f Time-series measurements of relative TsRbcS and RbcL levels per unit volume of sampled batch cultures (mean ± SEM; n = 3 biological replicates per time point). Source data are provided as a Source data file.
These results highlight the critical role of RvRbcS-like in maintaining photosynthetic efficiency in kleptoplasts. Because orthodox RbcS is an essential subunit of RuBisCO in algal and plant chloroplasts, RvRbcS-like is likely to be involved in carboxylation activity in kleptoplasts. Although RbcL is encoded by the chloroplast genome and may continue to be expressed in kleptoplasts, the original Tetraselmis sp. nuclear-encoded RbcS protein (TsRbcS) is unlikely to be produced after the kleptoplast transformation stage (Fig. 1), during which the algal nucleus is eliminated from the R. viridis cells9.
A time-series immunoblot analysis of total cellular proteins using a general plant RbcL antibody showed no significant changes in the RbcL levels in the first week after the ingestion event (Fig. 1) in wild-type cells (Fig. 4e), indicating maintenance of intact RuBisCO. Because RbcL and RbcS are balanced in a stoichiometric ratio to form the RbcL8RbcS8 heterohexadecameric RuBisCO complex, the RbcL levels are expected to be regulated by the availability of RbcS42. Thus, the maintenance of RbcL levels after the ingestion event is expected to depend on RbcS or its functional equivalent. Indeed, in wild-type cells, TsRbcS levels gradually declined, whereas RvRbcS-like levels increased over 7 days, as detected using anti-peptide antibodies specific to each protein (Fig. 4e). This interpretation was further supported when the immunoblot signals were evaluated in terms of protein abundance per culture volume (i.e., the total amount of each protein within a unit volume of culture; Fig. 4f), a measure independent of cell proliferation and associated variation in total protein content. By contrast, in the ΔRvRbcS-like cells, which lacks the supply of RvRbcS-like, RbcL levels also decreased (Fig. 4e, f), strongly indicating the effective loss of RuBisCO.
The phenotype of another knockout strain, ΔRvRca-like, was considerably milder, despite the well-recognized essential role of RuBisCO activase in RuBisCO function43. The growth rate of ΔRvRca-like cells during the first week after the ingestion event was nearly comparable to that of wild-type cells, with only a slightly lower final cell concentration (Fig. 4a). However, phenotypic differences became apparent during the stationary phase, because the ΔRvRca-like cells did not show the accelerated accumulation of cytoplasmic polysaccharide grains that was observed in wild-type cells (Fig. 4b). Notably, the photosynthetic rate of ΔRvRca-like cells on day 8 was significantly lower than that of wild-type cells (Fig. 4c). These observations suggest that the ΔRvRca-like phenotype manifests more gradually than the ΔRvRbcS-like phenotype, reflecting a delayed but substantial reduction in RuBisCO efficiency.
NtLCDs in kleptoplast-targeted proteins
We demonstrated that the NtLCD of RvRbcS-like alone is sufficient to mediate the translocation of a downstream peptide into the kleptoplast. Specifically, based on the results indicating that the NtLCDs of RvRbcS-like function as translocation signals and are cleaved during maturation (Fig. 3c and Supplementary Fig. 2), we tested this hypothesis using a NanoLuc luciferase reporter system44. We generated a transformant line expressing luciferase without an NtLCD (Fig. 5a) and another line expressing luciferase fused to the NtLCD of RvRbcS-like at the N-terminus (Fig. 5b). In the absence of the NtLCD, luciferase bioluminescence was detected throughout the cytosol but not in kleptoplasts (Fig. 5c). In contrast, when fused to the NtLCD, luciferase was mainly localized in kleptoplasts (Fig. 5d). These results provide direct evidence that the NtLCD is sufficient for transporting nuclear-encoded proteins into kleptoplasts.
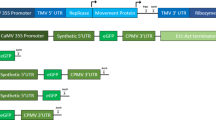
a Gene construct used to induce the expression of codon-optimized luciferase (optNluc) and the G418 resistance marker gene (neor), flanked by the E. gracilis pyruvate: NADP(+)-oxidoreductase gene (EgPNO) promoter and the Arabidopsis thaliana heat shock protein gene (AtHSP) terminator. b Gene construct incorporating an NtLCD sequence from RvRbcS-like (519 bp) upstream of the optNluc-neor cassette. c Bioluminescence microscopy of R. viridis transformant expressing luciferase without an NtLCD sequence, showing cytosolic bioluminescence spatially separated from chlorophyll fluorescence. d Bioluminescence microscopy of the transformant expressing luciferase fused to the NtLCD of RvRbcS-like, showing overlapping bioluminescence and chlorophyll fluorescence, indicating localization to the kleptoplast.
Importantly, NtLCDs ranging from 134 to 320 amino acids (mean 194 ± 37) were identified in 35 of the 37 putative kleptoplast-targeted proteins listed above (Fig. 2a and Supplementary Table 3). These NtLCDs structurally resemble chloroplast-targeting presequences in Euglena gracilis36,45, which typically include one or two transmembrane helices (TMHs). Based on this similarity, the R. viridis NtLCDs could be classified into four types (Fig. 6a): the most common RvClass IA, which features two TMHs flanking a 90–100 residue hydrophilic segment enriched in hydroxylated residues at both ends, basic residues in the center, and acidic residues toward the C-terminus (Fig. 6b); RvClass IB, which includes an additional TMH in the C-terminal extension; RvClass IC, which has a shorter spacer (40–80 residues) lacking the acidic C-terminal region (Fig. 6b); and RvClass II, which lacks a second TMH and has a simplified hydrophilic structure similar to RvClass IC (Fig. 6b). The NtLCD of RvRbcS-like falls into RvClass IA.
a Of the 37 sequences examined, 35 exhibited extended NtLCDs. These were classified according to the number of predicted TMHs and the features of the hydrophilic peptide regions located between the TMHs. b Characteristics of RvClass IA1, IA2, and II NtLCD sequences. The figure shows the averaged frequencies of amino acid categories calculated using a 15-residue sliding window for nonpolar (A, F, G, I, L, M, P, V, W), polar (C, N, Q), hydroxylated (S, T, Y), acidic (D, E), and basic (H, K, R) amino acids. White capital letters indicate the peaks of enrichment: H (hydroxylated), A (acidic), and B (basic). The sequences in each class were aligned and averaged by position. c Features of class IA targeting sequences of E. gracilis36,45 from selected chloroplast-targeted proteins corresponding to the R. viridis kleptoplast proteins discussed here. Peptide sequence alignments underlying the analyses shown in Fig. 6b, c are provided in the Supplementary data. Source data are provided as a Source data file.
These findings suggest that at least 20 candidate proteins with RvClass IA NtLCDs, including RvRbcS-like and RvRca-like, and possibly three with RvClass IB NtLCDs (Fig. 6a), are likely transported into kleptoplasts via an evolutionarily conserved but unidentified translocation mechanism. All NtLCD classes identified in R. viridis are structurally comparable to the N-terminal targeting signals of E. gracilis chloroplast proteins (Fig. 6c)36,45. Based on this similarity, the remaining 12 proteins with RvClass IC or II NtLCDs (Fig. 6a) are also likely to be transported into kleptoplasts, because Class II signals similar to RvClass II were sufficient for chloroplast import in E. gracilis46. By contrast, the two proteins lacking extended NtLCDs showed constitutive expression patterns across all kleptoplastic stages (Supplementary Table 1). RvTPT1 encodes a predicted TPT, which may function on the outermost kleptoplast membrane of phagosomal origin9,35, rather than within the kleptoplast itself. Among four additional TPT genes with lower expression (Supplementary Table 5), three (RvTPT2–4) carry RvClass IA NtLCDs, suggesting that they are directed to inner membranes derived from algal chloroplasts.
Discussion
R. viridis is the first organism in which nuclear-encoded proteins were biochemically shown to be expressed in a transient xenogeneic organelle. This finding indicates that the mature kleptoplasts in R. viridis exhibit molecular chimerism, not merely structural incorporation. The progressive formation of this chimerism appears to be physiologically necessary to sustain photosynthesis because the kleptoplast is no longer replenished with authentic Tetraselmis-derived proteins once the algal nucleus is lost. Moreover, chimerism develops heterogeneously, whereby different host factors are incorporated at distinct stages. For example, although RvRca-like is transcriptionally upregulated early (Fig. 2), the knockout experiments reveal that the disruption causes functional deficiency only after the stationary culture phase, about 1 week after the ingestion event (Fig. 4). By contrast, RvRbcS-like is critical throughout the phototrophic stage, accumulating as TsRbcS levels rapidly decline (Fig. 4e). This contrast likely reflects the different stabilities of these proteins, with TsRca persisting longer and delaying phenotypic effects in the absence of host-derived RvRca-like.
Among these host-derived proteins, RvRbcS-like stands out and implies that the molecular chimerism in R. viridis involves host-driven remodeling of kleptoplasts, not just to compensate for the lost Tetraselmis proteins. A striking example is the rapid pyrenoid reorganization during the kleptoplast transformation stage (Fig. 1)9,35, a phenomenon absent in the original algal chloroplast, that is likely to be driven by host-derived proteins. Notably, RvRbcS-like comprises four tandem RbcS domains flanked by extended NtLCDs and CtLCDs (Fig. 3a). This structure superficially resembles the eight-domain RbcS precursor in E. gracilis (EgRbcS), which is cleaved in chloroplasts to yield single-domain RbcS peptides40. However, RvRbcS-like appears to remain intact, aside from the removal of NtLCD, and its linker sequences differ markedly from the conserved decapeptide cleavage motifs in EgRbcS, which likely precludes similar processing.
We propose that the unique 228-residue CtLCD of RvRbcS-like contributes to liquid–liquid phase separation (LLPS)-driven pyrenoid reorganization. This domain contains repeated tetrapeptide motifs, including serine-tyrosine-[glycine/alanine/glutamate/aspartate/-]-glutamine (SY[G/A/E/D/-]Q) and serine-tyrosine-[arginine/glutamine]-proline (SY[R/Q]P), and SYGQ-resembling motifs are found in LLPS-prone proteins, such as the human RNA-binding protein FUS47. Pyrenoids are LLPS-mediated assemblies essential for carbon fixation, and a low-complexity repeat protein is crucial for their formation in the green alga Chlamydomonas reinhardtii48. The CtLCD may similarly facilitate pyrenoid remodeling in R. viridis. Thus, RvRbcS-like may have dual roles of supporting RuBisCO function and of enabling the formation and inheritance of multiple pyrenoids in daughter kleptoplasts.
This remodeling requires effective delivery of host-derived proteins into kleptoplasts after each ingestion event, necessitating rapid assembly of a translocation mechanism. To achieve effective delivery, the system that recognizes the NtLCD as a signal sequence must be established de novo either in or even prior to the kleptoplast transformation stage. The de novo establishment of the system must be a fundamental difference from the constitutive chimerism observed among established organelles. However, because the only clue to this mechanism lies in the structural properties of the NtLCDs of the putative kleptoplast-targeted proteins, the protein translocation system in the chloroplasts of Euglenophyceae should be considered. Despite representing constitutive chimerism, Euglenophyceae share targeting sequences that are structurally similar to the NtLCDs of R. viridis.
Insights into the underlying mechanisms can be drawn from studies of E. gracilis, where two elements are thought to be involved in protein translocation into its chloroplast. The first is an endoplasmic reticulum-targeting signal peptide containing the first TMH, which enables vesicular delivery to the outermost membrane of the triple-membrane envelope. The second is a downstream transit peptide-like region (Fig. 6c). This second segment is thought to mediate import via a presumed translocon analogous to the outer/inner chloroplast membrane (TOC/TIC) complexes of primary chloroplasts, although nearly all of these components, except one, remain unidentified in E. gracilis46.
In R. viridis, a key distinction is that the original TOC/TIC complexes from Tetraselmis appear to be retained in the inner two membranes, at least initially. Accordingly, the first TMH of the NtLCD may act as the endoplasmic reticulum-targeting signal, whereas the acidic residue-depleted N-terminal half of the hydrophilic region in RvClass IA resembles a transit peptide (Fig. 6a). However, relying solely on Tetraselmis machinery cannot account for the additional NtLCD features: an acidic residue-rich C-terminal segment, a second TMH, and a downstream low-complexity domain extension (Fig. 6b). These features are absent from green algal chloroplast-targeting signals, including those of Tetraselmis, suggesting the presence of a distinct, previously unrecognized translocation mechanism.
Understanding the molecular protein transport mechanism may reveal a more broadly conserved eukaryotic system underlying kleptoplasty in R. viridis. It is impossible to reconstruct the ecophysiological state of the last common ancestor of Rapaza and Euglenophyceae definitively, or determine whether the last common ancestor possessed transient kleptoplasts or permanent chloroplasts. However, it is likely that this ancestor had already developed translocation machinery, which was later inherited and adapted for Euglenophyceae chloroplasts and R. viridis kleptoplasts, given their apparently shared signals. The ancestral machinery probably evolved through modification or recombination of pre-existing systems, possibly related to intraphagosomal processing. For example, differential degradation of ingested components is common among algivorous heterotrophs, where the chloroplast structure often degrades after phototoxic chlorophylls49, suggesting that specific protein delivery into engulfed cells may have pre-dated kleptoplasty. Interestingly, the putative chloroplast-targeting sequences in dinoflagellates, organisms from a distinct eukaryotic supergroup, share structural features with those in Euglenophyceae chloroplast-targeting presequences36, and hence with R. viridis NtLCDs. Although this may reflect convergent evolution driven by the need to transport proteins across triple-membrane-bound plastids, an underlying molecular homology based on a more fundamental eukaryotic mechanism cannot be excluded.
All organisms exist in evolutionarily transient states, shaped by continuous evolutionary refinements as they adapt to ever-changing conditions to ensure their survival through successive generations, and R. viridis is no exception to this. This species cannot be definitively placed on a trajectory toward acquiring chloroplasts. Nevertheless, R. viridis represents a new example of the unexplored ecophysiological potential of eukaryotic cells. Thus, the introduction of R. viridis as an experimental model phototroph that may be amenable to genetic engineering, alongside other kleptoplastic phototrophs that have been and will be studied, will expand our understanding of the molecular mechanisms by which eukaryotes interact with the bacterium-derived photosynthetic machinery. Beyond the direct structural manipulation by host-supplied proteins, the interaction may involve precise anterograde and retrograde signaling between the host nucleus and the organelle50,51,52, which would be essential for regulating photosynthesis to prevent the xenogeneic object from merely becoming a source of reactive oxygen species. We believe that a deeper, multidimensional understanding of the transient molecular chimerism demonstrated in R. viridis will provide valuable insights into the evolutionary potential of eukaryotic cells, which have given rise to a diverse array of organelles throughout their evolutionary history.
Methods
Cell culture
R. viridis (NIES-4477) and Tetraselmis sp. (NIES-4478) were cultured under axenic conditions in Daigo IMK medium (Nihon Pharmaceutical, Tokyo, Japan; maintenance medium) at 20 °C under illumination of 100–150 μmol photons m−2 s−1 with a 14-h light/10-h dark cycle (14 L:10D). R. viridis cultures were generally established by mixing Tetraselmis sp. cells into fresh medium at a ratio of 1:1–1:3 (depending on the experiment) as a source of new kleptoplasts. After inoculation, the Tetraselmis sp. cells were typically completely consumed by R. viridis within 0.5–3 h, depending on the initial ratio, resulting in monospecific cultures containing only R. viridis. The maintenance culture conditions were described in previous reports9,34. Experimental R. viridis cultures used for the time-series transcriptome and qPCR analysis were initiated by mixing Tetraselmis sp. cells at a 1:1 ratio, resulting in rapid consumption of the kleptoplast donor within 0.5 h. The remaining experiments were initiated by mixing Tetraselmis sp. cells at a 1:3 ratio. The cell numbers were counted using a particle counter/analyzer (CDA-1000, Sysmex, Kobe, Japan).
Microscopy
Differential interference contrast images were obtained using an inverted microscope (IX71, Olympus, Tokyo, Japan) equipped with a color CCD camera (FX630, Olympus) and a Flovel Image Filing System (Flovel, Chofu, Japan). Live cells were placed between two glass coverslips (0.13–0.17 mm) and observed.
Time-series transcriptomic analysis
To obtain the transcriptomes of R. viridis NIES-4477, we also analyzed the transcriptome of the kleptoplast donor alga Tetraselmis sp. NIES-4478 from the unialgal culture. This was done to subtract any potential contaminating sequences, although all Tetraselmis cells are typically consumed shortly after inoculation, and the cytoplasm and nucleus are eliminated early during kleptoplast formation. R. viridis cells were sampled from quintuplicate batch cultures at 8, 18, 120, and 360 h after initial inoculation, and Tetraselmis was sampled from quintuplicate unialgal cultures. The collected cells were pelleted by centrifugation and immediately frozen in liquid nitrogen. Total RNA was extracted from all samples using the NucleoSpin RNA XS kit (TaKaRa, Kusatsu, Japan), and the three replicates with the highest RNA yields were selected for downstream experiments. Then, 2.0‒22.4 μg of the total RNA samples were sent to Macrogen Co. (Tokyo, Japan) for library construction and sequencing using a sequencing system (HiSeq X, Illumina, San Diego, CA, USA).
The RNA-seq raw reads were cleaned using cutadapt ver. 2.853 by trimming low-quality-value ends (<QV20) and adapter sequences and discarding any reads shorter than 50 bp. The trimmed reads from the R. viridis samples were assembled de novo using Trinity ver. 2.11.054 in the paired-end mode with the option “--min_contig_ length 300”. When splicing variants of a gene were found, the longest transcript was selected as the representative mRNA sequence. The ORFs were predicted using TransDecoder ver. 5.5.0 (http://transdecoder.github.io). The de novo transcriptome sequences were then filtered against the genome data from Tetraselmis sp. (methods described below; Supplementary data deposited in Figshare55), although reads attributable to the Tetraselmis sp. genome were, in fact, miscellaneous in the R. viridis RNA-seq reads. Therefore, the contigs with 97% identity to Tetraselmis nucleotide sequences evaluated by BLASTN search were omitted, and the remaining contigs were used as a reference for the R. viridis transcripts. We confirmed that none of the omitted sequences contained the euglenid 5′-spliced leader sequence38. To estimate the function of the predicted orf, BLASTP searches were conducted using the Uniprot/Swiss-Prot database (Uniprot release 2020_6, downloaded from https://www.uniprot.org/uniprotkb). To obtain gene expression scores, one side of the trimmed reads was mapped to the reference using Bowtie2 ver. 2.4.256. SAMtools ver. 1.1157, BEDtools ver. 2.92.258, and R ver. 4.0.359 were used to calculate the number of reads mapped to contigs (raw count) and the reads per kilobase of transcript per million mapped reads. To analyze the time-series transcriptomic changes of R. viridis, genes with low mapped read counts were omitted using the filterLowCountGenes command (low.count = 10) in the TCC package ver. 1.30.060 in R. The data were then normalized using the calcNormFactors command, and differentially expressed genes (DEGs) were identified using the estimateDE command of TCC in R. R. viridis contigs were defined as DEGs if the false discovery rate was <0.05 (Supplementary data deposited in Figshare55). The normalized count was obtained using the getNormalizedData command of TCC.
For genome assembly of Tetraselmis sp. NIES-4478 in association with the above R. viridis RNA-seq analysis, genomic DNA was extracted from a stationary-phase monoxenic culture using the DNeasy Plant Mini Kit (QIAGEN, Venlo, Netherlands). The purified DNA was sent to Macrogen Co. (Tokyo, Japan) for library preparation and sequencing on an Illumina HiSeq X system. Raw reads were processed with cutadapt ver. 2.153, trimming adapter sequences and low-quality bases (<QV20), and discarding reads shorter than 50 bp. A total of 19 million paired-end reads were assembled using SPAdes version 3.13.061, and the resulting scaffold FASTA file was used to eliminate Tetraselmis contigs from the R. viridis transcriptome reference, as described above.
Nuclear genomic DNA sequencing and assembling
Genomic DNA was purified from a stationary phase monoxenic culture of R. viridis containing Tetraselmis sp. using the standard phenol/chloroform/isoamyl alcohol method. The purified DNA sample was sent to Macrogen Co. for library construction and sequencing using a sequencing system (NovaSeq6000, Illumina). After trimming and quality control procedures using the FASTX-Toolkit (http://hannonlab.cshl.edu/fastx_toolkit/), 152 million paired-end reads were retained. Genome assembling was carried out using MaSuRCA version 4.0.762 as follows. To determine the reliable insert size and standard deviation (SD) of the insert size for paired-end reads, a preliminary scaffold was constructed using 10% of paired-end reads with SPAdes version 3.15.461 and default setting. Mapping analysis against the preliminary scaffold was performed using Bowtie256, and the insert size and SD were estimated to be 120 bp and 110 bp, respectively. The MaSuRCA assembly was subsequently performed using all 152 million quality-controlled paired-end reads with the CABOG assembly option. Finally, a total primary scaffold length of 508 Mbp with an average coverage depth of 91× was obtained.
Amplification and sequencing of specific cDNA and gDNA loci
The full-length cDNA and genomic DNA (gDNA) sequences of RvRbcS-like and RvRca-like, including the flanking regions, were amplified and sequenced (DDBJ accession numbers: LC877979‒LC877982). Total RNA (>200 bases) was extracted from R. viridis using ISOGEN II (Nippon Gene, Toyama, Japan) and reverse-transcribed with SuperScript IV (Thermo Fisher Scientific, Waltham, MA, USA) and the GeneRacer oligo-dT24 primer (Thermo Fisher Scientific). Rapid amplification of cDNA ends (RACE) was performed to determine the 5′ and 3′ termini using primers for the spliced leader sequence and the GeneRacer adapter, as described by Nakazawa et al.63. The target gDNA regions were identified by BLASTN searches against the DNA-seq contigs, and amplicons spanning from the first to the last exon were cloned and sequenced by next-generation sequencing (Plasmid-EZ, Azenta, South Plainfield, NJ, USA), which also identified missing gaps in the DNA-seq data, typically enriched short repeats. The RvPsbR gene was also partially re-sequenced due to discrepancies between its cDNA and contig sequences. All primers used in this study are listed in Supplementary Table 6. All resulting sequence data have been deposited in Figshare55.
Phylogenetic analysis
BLASTP searches were conducted against the NCBI nr database using the translated peptide sequences of each highly expressed gene listed in Supplementary Table 1 as queries. Homologs from the chloroplast donor Tetraselmis sp. and the Euglenophyceae algae Eutreptiella spp. and E. gracilis were retrieved from in-house RNA-seq data and manually incorporated into the dataset. Sequence alignment was performed using the MAFFT algorithm with default parameters (MAFFT package v7.520)64, followed by trimming with trimAl 1.2rev5765. Short and ambiguously aligned positions were manually removed. Maximum likelihood trees were inferred using IQ-TREE software v1.6.1266, and the number of nonparametric bootstrap replicates and substitution models are specified in each figure. The resulting trees were manually inspected to assess the relationships between R. viridis sequences and those of other organisms in the database.
qPCR analysis
Wild-type R. viridis cells were sampled from a batch cultures at 1, 6, 12, 18, 24, 48, 72, 96, 144, 192, 240, 288, and 336 h after initial inoculation. The cells were pelleted by centrifugation and immediately frozen in liquid nitrogen. Total RNA was extracted using ISOGEN II (Nippon Gene), and cDNA was synthesized by reverse transcription using the PrimeScript RT Reagent Kit (TaKaRa). The RvRbcS-like and RvRca-like transcript levels were quantified using a real-time PCR system (LightCycler 96 system, NIPPON Genetics, Tokyo, Japan) with KAPA SYBR Fast Universal (NIPPON Genetics) with the primers listed in Supplementary Table 6.
RNAi
Silencing experiments targeting RvRbcS-like were performed following the method described by Maruyama et al.34. Double-stranded DNA templates were amplified from R. viridis cDNA using specific primers, each containing a 5′ extension with a T7 promoter sequence (Supplementary Table 6). Double-stranded RNA (dsRNA) corresponding to a 434 bp fragment of RvRbcS-like was synthesized and purified using the MEGAscript RNAi Kit (Thermo Fisher Scientific). The dsRNA was introduced into R. viridis cells through two consecutive electroporation treatments. For the first electroporation, R. viridis cells 5 days after inoculation with the prey Tetraselmis sp. were used. A total of 4 × 106 cells were resuspended in 200 µL of the RNAi electroporation buffer (1/20-strength artificial seawater (ASW; 5% (v/v)) with 500 mM trehalose) containing 15 µg of dsRNA and electroporated using a GENE PULSER II system (Bio-Rad, Hercules, CA, USA) at 0.45 kV and 50 µF in a 0.2 cm gap cuvette. Immediately afterward, the cells were transferred to fresh IMK medium and incubated for 2 days (half a day in the dark followed by a light–dark cycle at 20 °C). Negative control experiments were conducted in parallel without dsRNA. The electroporation was repeated once, 2 days later, using the same procedure. The treated cells were again kept in the dark for half a day and then fed with Tetraselmis sp. to acquire new kleptoplasts, thereby establishing RNAi-knockdown batch cultures.
CRISPR/Cas9 genome editing
CRISPR/Cas9 genome editing experiments were performed following the general procedure described by Maruyama et al.34, with the following design-specific modifications. Target CRISPR RNAs (crRNAs) were designed using the CHOPCHOP (version 3) webtool67 and are listed in Supplementary Table 7. Custom crRNAs were synthesized by Integrated DNA Technologies (Coralville, IA, USA). Guide RNAs were prepared by hybridizing each crRNA with tracrRNA (Alt-R CRISPR-Cas9 tracrRNA, Integrated DNA Technologies) and then were incubated with Alt-R S.p. HiFi Cas9 Nuclease V3 (Integrated DNA Technologies) to assemble the ribonucleoprotein (RNP) complexes. The expected cleavage sites in the RvRbcS-like and RvRca-like coding regions of the gDNA are shown in Supplementary Fig. 5.
To generate knockout strains (ΔRvRbcS-like and ΔRvRca-like), pairs of RNPs targeting two distinct loci in the first exon of each gene (Supplementary Fig. 5), together with a single-stranded oligonucleotide (ssODN) donor (Supplementary Table 7), were introduced into R. viridis cells through an electroporation treatment described in the next section. This procedure enabled precise deletions of 200 and 120 bp at the target sites in RvRbcS-like and RvRca-like, respectively, while simultaneously introducing stop codons at the resulting junctions (Supplementary Fig. 5). To generate the HA-tagged RvRbcS-like strain, an RNP targeting a single site on the 3′ side of the sequence encoding the fourth RbcS domain in the second exon was used (Supplementary Table 7). In this case, four partially overlapping ssODN fragments were introduced together with the RNPs, allowing for seamless in-frame integration of the HA tag sequence at the cleavage site through complementary base pairing (Supplementary Fig. 5).
For electroporation, R. viridis cells 1 week after inoculation with the prey Tetraselmis sp. were used. A total of 2 × 105 cells were resuspended in 50 µL of the CRISPR/Cas9 electroporation buffer (1/10-strength ASW (10% (v/v)) supplemented with 450 mM trehalose) containing 100 µM of each RNP and 200 µM of each ssODN, and electroporated using a GENE PULSER II system at 0.45 kV and 50 µF in a 0.2 cm gap cuvette. Immediately afterward, the cells were transferred to fresh IMK medium and incubated at 26 °C in the dark for 24 h. The RNP-transfected R. viridis cells were then fed with an equal number of Tetraselmis sp. cells and further cultured for 72 h in a 14 L:10D cycle at 26 °C before being transferred to the standard culture condition at 20 °C (primary culture). Clonal strains were generated from the primary culture by aseptically separating individual R. viridis cells into culture plates using a microcapillary method using glass tubes. The targeted genomic region of each clone was PCR-amplified and sequenced to identify successfully edited clones.
Immunoblotting
R. viridis cultures were harvested by centrifugation at designated timepoints following the kleptoplast acquisition and unialgal Tetraselmis sp. cultures, and the cell pellets were immediately snap frozen in liquid nitrogen. For protein extraction, the pellets were resuspended in lysis buffer containing 1 mM phenylmethanesulfonyl fluoride and 5 mM 6-aminocaproic acid, followed by sonication. The lysates were centrifuged at 20,000 × g for 1 min, and the resulting supernatants were mixed with 2× sodium dodecyl sulfate (SDS) sample buffer (500 mM Tris-HCl, pH 6.8, 10% [w/v] SDS, 0.01% [w/v] bromophenol blue, 10% [v/v] β-mercaptoethanol, 20% [v/v] glycerol). The samples were heat-denatured for 10 min at 90 °C or 3 min at 100 °C. Protein concentrations were determined by the Bradford assay (XL-Bradford KY-1040; Apro Science Inc., Tokushima, Japan). Equal amounts (3 μg) of total protein were loaded onto SDS-polyacrylamide gels (12.5%) and electrophoresed. The proteins were transferred to polyvinylidene difluoride membranes (Immobilon-P, Merck Millipore, Burlington, MA, USA) by electroblotting. The membranes were probed with the following antibodies: rabbit polyclonal anti-RvRbcS-like (epitope: HSLAKAHRREGGET, amino acids [aa] 491–505; Eurofins Genomics KK, Tokyo, Japan), rabbit polyclonal anti-RvRca-like (epitope: AIASDDNVERFGKK, aa 181–189; Eurofins Genomics KK), rabbit polyclonal anti-TsRbcS (epitope: RISTALPIEKRSVA, aa 131–144; Eurofins Genomics KK), rabbit polyclonal anti-RbcL (#AS03-037; Agrisera, Vännäs, Sweden), and mouse monoclonal anti-HA (clone 16B12, #901513; BioLegend, San Diego, CA, USA). For reprobing, antibodies were stripped from the membranes using Restore Western Blot Stripping Buffer (Thermo Fisher Scientific), followed by incubation with monoclonal anti-α-tubulin antibody produced in mouse (clone B-5-1-2, #T5168; Sigma-Aldrich). Immunoreactive bands were detected using horseradish peroxidase (HRP)-conjugated secondary antibodies (anti-mouse immunoglobulin [Ig]G[H + L] or anti-rabbit IgG[H + L]; Cell Signaling Technology, Danvers, MA, USA) and Immobilon western chemiluminescent HRP substrate (Merck KGaA, Darmstadt, Germany). Signals were visualized with an image analyzer (LAS-4000, GE Healthcare, Chicago, IL, USA) and a multi imager (MultiImager II ChemiBOX, BioTools, Gunma, Japan). The signal intensities were quantified using ImageJ software68.
Immunofluorescence microscopy
Immunostaining of fixed cells was performed using mouse monoclonal anti-HA (clone 16B12, #901513; BioLegend) and rabbit polyclonal anti-RbcL (#AS03-037; Agrisera) antibodies to examine the colocalization of HA-tagged RvRbcS-like and RbcL, and with rabbit polyclonal anti-RvRca-like (epitope: AIASDDNVERFGKK, Eurofins Genomics KK) antibody to examine the localization of RvRca-like. Secondary antibodies included Alexa Fluor 555-conjugated anti-mouse IgG and Alexa Fluor 488-conjugated anti-rabbit IgG (Thermo Fisher Scientific). Images for colocalization analysis of HA-tagged RvRbcS-like and RbcL, as well as for RvRca-like localization, were acquired using a confocal laser scanning microscope (LSM700, Zeiss, Oberkochen, Germany) equipped with a 100× oil-immersion objective lens (EC Plan-Neofluar 100×/1.3 Oil Ph3 M27, Zeiss). Alexa Fluor 488-labeled RbcL or RvRca-like was excited with a 488 nm laser, and fluorescence was recorded between 505 and 550 nm. Alexa Fluor 555-labeled HA-tagged RvRbcS-like was excited at 555 nm, and fluorescence was recorded between 560 and 615 nm. Chlorophyll autofluorescence was excited at 488 nm, and fluorescence above 650 nm was recorded. Z-stack images were captured at 0.4-μm intervals and processed by maximum intensity projection using ImageJ software.
Photosynthetic activity measurement
Wild-type and knockout strains of R. viridis (2 × 106 cells), as well as RNAi-knockdown cells, were harvested by gentle centrifugation from batch cultures at 8 days after initial inoculation and immediately resuspended in fresh IMK medium (2 mL). To avoid carbon limitation during measurement, 1 M NaHCO3 (10 µL) was added to the suspension. Oxygen evolution was then measured using a temperature-controlled liquid-phase oxygen electrode system (OXYT-1, Hansatech Instruments, Kings Lynn, UK).
Quantification of polysaccharide grains
Polysaccharide grains were extracted from R. viridis cells and purified using a previously described method9. The resulting insoluble polysaccharide precipitate was dissolved in 1 M NaOH, and its concentration was determined using the phenol–sulfuric acid assay69, with glucose solution as the standard reference.
NanoLuc reporter assay
Transformants expressing NanoLuc luciferase were generated using the procedure described by Nakazawa et al.44. For the construct lacking an NtLCD sequence, a DNA cassette was assembled consisting of the E. gracilis PNO promoter region (−118 to −1), the NanoLuc gene codon-optimized for E. gracilis (optNluc), and a G418 resistance gene (neor) fused downstream via a linker encoding the peptide GSSGAIA, followed by the Arabidopsis thaliana heat shock protein (AtHSP) terminator. For the construct containing the NtLCD of RvRbcS-like, the same expression cassette was used, with the NtLCD sequence inserted upstream of the optNluc gene. Both constructs were introduced into R. viridis cells using the electroporation protocol described by Nakazawa et al.44. Following electroporation, the cells were inoculated into Daigo IMK medium (15 mL), cultured for 10 days, and then transferred twice at weekly intervals. After complete consumption of Tetraselmis cells in the third culture, G418 was added at a final concentration of 10 μg/mL. Eight days later, a fourth transfer was performed, and the G418 concentration was increased to 20 μg/mL. Weekly transfers continued for 1 month, after which the clonal strains were isolated using glass microcapillaries and maintained in G418-free medium. To screen for successful transformants, 25 μL of the supernatant from the R. viridis lysates were assessed using a luciferase assay system (Nano-Glo, Promega, Madison, WI, USA) with a 20/20n Luminometer (Promega). Positive clones were further validated by cDNA PCR amplification of the inserted gene constructs. For luminescence imaging, the cells were briefly suspended in 100% methanol to immobilize them and increase substrate permeability, and then they were resuspended in IMK medium before imaging with a luminescence imaging system (LV200, Olympus).
Statistics and reproducibility
Statistics and reproducibility. No statistical method was used to predetermine sample size. No outliers were excluded from the analyses. For the time-series RNA-seq experiment, three biological replicates with the highest RNA yields were selected from five independent cultures for library preparation and sequencing. The experiments were not randomized. Investigators were not blinded to allocation during experiments and outcome assessment.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All next-generation sequencing data generated in this study are available at Figshare (https://doi.org/10.6084/m9.figshare.29356226)55. This includes the time-series transcriptome data (FASTA files containing 69,444 cDNA and predicted peptide sequences, and differential expression analysis in Excel format), the draft genome assembly of Rapaza viridis (FASTA file of 269,570 contigs), and sequences of amplified cDNA and gDNA loci. All raw RNA-seq data for R. viridis NIES-4477, DNA-seq data for Tetraselmis sp. NIES-4478, and DNA-seq data for R. viridis generated in this study, along with their assembled contig sequences, have been deposited in the DDBJ Sequence Read Archive under BioProject accession codes PRJDB35573, PRJDB35574, and PRJDB35577, respectively. Molecular phylogenetic trees corresponding to Supplementary Tables 1–5 are also available in Figshare (https://doi.org/10.6084/m9.figshare.29356226)55. Source data are provided with this paper55.
References
Pittis, A. A. & Gabaldón, T. Late acquisition of mitochondria by a host with chimaeric prokaryotic ancestry. Nature 531, 101–104 (2016).
Gray, M. W., Burger, G. & Lang, B. F. Mitochondrial evolution. Science 283, 1476–1481 (1999).
Sibbald, S. J. & Archibald, J. M. Genomic insights into plastid evolution. Genome Biol. Evol. 12, 978–990 (2020).
Martin, W. et al. Gene transfer to the nucleus and the evolution of chloroplasts. Nature 393, 162–165 (1998).
Timmis, J. N., Ayliffe, M. A., Huang, C. Y. & Martin, W. Endosymbiotic gene transfer: organelle genomes forge eukaryotic chromosomes. Nat. Rev. Genet. 5, 123–135 (2004).
Larkum, A. W., Lockhart, P. J. & Howe, C. J. Shopping for plastids. Trends Plant Sci. 12, 189–195 (2007).
Ku, C. et al. Endosymbiotic origin and differential loss of eukaryotic genes. Nature 524, 427–432 (2015).
Ponce-Toledo, R. I., López-García, P. & Moreira, D. Horizontal and endosymbiotic gene transfer in early plastid evolution. New Phytol. 224, 618–624 (2019).
Karnkowska, A. et al. Euglenozoan kleptoplasty illuminates the early evolution of photoendosymbiosis. Proc. Natl. Acad. Sci. USA 120, e2220100120 (2023).
Shinzato, C. et al. Using the Acropora digitifera genome to understand coral responses to environmental change. Nature 476, 320–323 (2011).
Waugh, G. R. & Clark, K. B. Seasonal and geographic variation in chlorophyll level of Elysia tuca (Ascoglossa: Opisthobranchia). Mar. Biol. 92, 483–487 (1986).
Serôdio, J., Cruz, S., Cartaxana, P. & Calado, R. Photophysiology of kleptoplasts: photosynthetic use of light by chloroplasts living in animal cells. Philos. Trans. R. Soc. Lond. B Biol. Sci. 369, 20130242 (2014).
Maeda, T. et al. Chloroplast acquisition without the gene transfer in kleptoplastic sea slugs, Plakobranchus ocellatus. eLife 10, e60176 (2021).
Park, M. G. et al. First successful culture of the marine dinoflagellate Dinophysis acuminata. Aquat. Microb. Ecol. 45, 101–106 (2006).
Kim, M., Nam, S. W., Shin, W., Coats, D. W. & Park, M. G. Dinophysis caudata (Dinophyceae) sequesters and retains plastids from the mixotrophic ciliate prey Mesodinium rubrum. J. Phycol. 48, 569–579 (2012).
Gast, R. J., Moran, D. M., Dennett, M. R. & Caron, D. A. Kleptoplasty in an Antarctic dinoflagellate: caught in evolutionary transition? Environ. Microbiol. 9, 39–45 (2007).
Onuma, R. & Horiguchi, T. Kleptochloroplast enlargement, karyoklepty and the distribution of the cryptomonad nucleus in Nusuttodinium (= Gymnodinium) aeruginosum (Dinophyceae). Protist 166, 177–195 (2015).
Yamada, N. et al. Discovery of a kleptoplastic ‘dinotom’ dinoflagellate and the unique nuclear dynamics of converting kleptoplastids to permanent plastids. Sci. Rep. 9, 10474 (2019).
Yih, W., Kim, H. S., Jeong, H. J., Myung, G. & Kim, Y. G. Ingestion of cryptophyte cells by the marine photosynthetic ciliate Mesodinium rubrum. Aquat. Microb. Ecol. 36, 165–170 (2004).
Johnson, M. D., Oldach, D., Delwiche, C. F. & Stoecker, D. K. Retention of transcriptionally active cryptophyte nuclei by the ciliate Myrionecta rubra. Nature 445, 426–428 (2007).
Hughes, E. A., Maselli, M., Sorensen, H. & Hansen, P. J. Metabolic reliance on photosynthesis depends on both irradiance and prey availability in the mixotrophic ciliate, Strombidium cf. basimorphum. Front. Microbiol. 12, 642600 (2021).
Jauffrais, T. et al. Effect of light on photosynthetic efficiency of sequestered chloroplasts in intertidal benthic foraminifera (Haynesina germanica and Ammonia tepida). Biogeosci 13, 2715–2726 (2016).
Tsuchiya, M. et al. Acquisition, maintenance, and ecological roles of kleptoplasts in Planoglabratella opercularis (Foraminifera, Rhizaria). Front. Mar. Sci. 7, 585 (2020).
Okamoto, N. & Inouye, I. Hatena arenicola gen. et sp. nov., a katablepharid undergoing probable plastid acquisition. Protist 157, 401–419 (2006).
Sørensen, M. E. S. et al. A novel kleptoplastidic symbiosis revealed in the marine centrohelid Meringosphaera with evidence of genetic integration. Curr. Biol. 33, 3571–3584 (2023).
Bodył, A. Did some red alga-derived plastids evolve via kleptoplastidy? A hypothesis. Biol. Rev. Camb. Philos. Soc. 93, 201–222 (2018).
Miyagishima, S. Taming the perils of photosynthesis by eukaryotes: constraints on endosymbiotic evolution in aquatic ecosystems. Comm. Biol. 6, 1150 (2023).
Kim, G. H. et al. Cryptophyte gene regulation in the kleptoplastidic, karyokleptic ciliate Mesodinium rubrum. Harmful Algae 52, 23–33 (2016).
Onuma, R. et al. Changes in the transcriptome, ploidy, and optimal light intensity of a cryptomonad upon integration into a kleptoplastic dinoflagellate. ISME J. 14, 2407–2423 (2020).
Johnson, M. D. et al. Functional control and metabolic integration of stolen organelles in a photosynthetic ciliate. Curr. Biol. 33, 973–980 (2023).
Wisecaver, J. H. & Hackett, J. D. Transcriptome analysis reveals nuclear-encoded proteins for the maintenance of temporary plastids in the dinoflagellate Dinophysis acuminata. BMC Genom. 11, 366 (2010).
Hongo, Y., Yabuki, A., Fujikura, K. & Nagai, S. Genes functioned in kleptoplastids of Dinophysis are derived from haptophytes rather than from cryptophytes. Sci. Rep. 9, 9009 (2019).
Hehenberger, E., Gast, R. J. & Keeling, P. J. A kleptoplastidic dinoflagellate and the tipping point between transient and fully integrated plastid endosymbiosis. Proc. Natl. Acad. Sci. USA 116, 17934–17942 (2019).
Maruyama, M. et al. Horizontally acquired nitrate reductase realized kleptoplastic photoautotrophy of Rapaza viridis. Plant Cell Physiol. 64, 1082–1090 (2023).
Yamaguchi, A., Yubuki, N. & Leander, B. S. Morphostasis in a novel eukaryote illuminates the evolutionary transition from phagotrophy to phototrophy: description of R. viridis n. gen. et sp. (Euglenozoa, Euglenida). BMC Evol. Biol. 12, 29 (2012).
Durnford, D. G. & Gray, M. W. Analysis of Euglena gracilis plastid-targeted proteins reveals different classes of transit sequences. Eukaryot. Cell 5, 2079–2091 (2006).
Wang, J. et al. The conserved domain database in 2023. Nucleic Acids Res. 51, D384–D388 (2023).
Tessier, L. H. et al. Short leader sequences may be transferred from small RNAs to premature mRNAs by transsplicing in Euglena. EMBO J. 10, 2621–2625 (1991).
Sheth, N. et al. Comprehensive splice-site analysis using comparative genomics. Nucleic Acids Res. 34, 3955–3967 (2006).
Chan, R. L. et al. Eight small subunits of Euglena ribulose 1-5 bisphosphate carboxylase/oxygenase are translated from a large mRNA as a polyprotein. EMBO J. 9, 333–338 (1990).
Chen, K., Chen, X. & Schnell, D. J. Initial binding of preproteins involving the Toc159 receptor can be bypassed during protein import into chloroplasts. Plant Physiol. 122, 813–822 (2000).
Rodermel, S., Haley, J., Jiang, C. Z., Tsai, C. H. & Bogorad, L. A mechanism for intergenomic integration: abundance of ribulose bisphosphate carboxylase small-subunit protein influences the translation of the large-subunit mRNA. Proc. Natl. Acad. Sci. USA 93, 3881–3885 (1996).
Bracher, A., Whitney, S. M., Hartl, F. U. & Hayer-Hartl, M. Biogenesis and metabolic maintenance of Rubisco. Annu. Rev. Plant Biol. 68, 29–60 (2017).
Nakazawa, M. et al. Stable nuclear transformation methods for Euglena gracilis and its application to a related Euglenida. Algal Res. 75, 103292 (2023).
Schwartzbach, S. D. & Shigeoka, S. Euglena: Biochemistry, Cell and Molecular Biology (Springer, 2017).
Novák Vanclová, A. M. et al. Metabolic quirks and the colourful history of the Euglena gracilis secondary plastid. New Phytol. 225, 1578–1592 (2020).
Murthy, A. C. et al. Molecular interactions underlying liquid−liquid phase separation of the FUS low-complexity domain. Nat. Struct. Mol. Biol. 26, 637–648 (2019).
Mackinder, L. C. et al. A repeat protein links Rubisco to form the eukaryotic carbon-concentrating organelle. Proc. Natl. Acad. Sci. USA 113, 5958–5963 (2016).
Kashiyama, Y. et al. Taming chlorophylls by early eukaryotes underpinned algal interactions and the diversification of the eukaryotes on the oxygenated Earth. ISME J. 13, 1899–1910 (2019).
Hwang, Y. et al. Anterograde signaling controls plastid transcription via sigma factors separately from nuclear photosynthesis genes. Nat. Commun. 13, 7440 (2022).
Goldbecker, E. S. & de Vries, J. Systems biology of streptophyte cell evolution. Annu. Rev. Plant Biol. 76, 493–522 (2025).
Duanmu, D. et al. Retrograde bilin signaling enables Chlamydomonas greening and phototrophic survival. Proc. Natl. Acad. Sci. USA 110, 3621–3626 (2013).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 17, 10–12 (2011).
Grabherr, M. G. et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 29, 644–652 (2011).
Kashiyama, Y. et al. Data for: transient molecular chimerism for exploiting xenogeneic organelles. Figshare https://doi.org/10.6084/m9.figshare.29356226 (2025).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Ihaka, R. & Gentleman, R. R. A language for data analysis and graphics. J. Comput. Graph. Stat. 5, 299–314 (1996).
Sun, J., Nishiyama, T., Shimizu, K. & Kadota, K. T. C. C. An R package for comparing tag count data with robust normalization strategies. BMC Bioinform. 14, 219 (2013).
Bankevich, A. et al. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 19, 455–477 (2012).
Zimin, A. V. et al. The MaSuRCA genome assembler. Bioinformatics 29, 2669–2677 (2013).
Nakazawa, M. et al. Alteration of wax ester content and composition in Euglena gracilis with gene silencing of 3-ketoacyl-CoA thiolase isozymes. Lipids 50, 483–492 (2015).
Katoh, K., Rozewicki, J. & Yamada, K. D. MAFFT online service: multiple sequence alignment, interactive sequence choice and visualization. Brief. Bioinform. 20, 1160–1166 (2019).
Capella-Gutiérrez, S., Silla-Martínez, J. M. & Gabaldón, T. trimAl: a tool for automated alignment trimming. Bioinformatics 25, 1972–1973 (2009).
Trifinopoulos, J., Nguyen, L. T., von Haeseler, A. & Minh, B. Q. W-IQTREE: a fast online phylogenetic tool for maximum likelihood analysis. Nucleic Acids Res. 44, W232–W235 (2016).
Labun, K. et al. CHOPCHOP v3: expanding the CRISPR web toolbox beyond genome editing. Nucleic Acids Res. 47, W171–W174 (2019).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Hodge, J. E. & Hofreiter, B. T. Methods in Carbohydrate Chemistry Vol. 1 (eds. Whistler, R. L. & Wolfrom, M. L.) 380–394 (Academic Press, 1962).
Acknowledgements
We thank Ms. Tomomi Munekyo for her technical assistance. This study was supported, in part, by the Japan Society for the Promotion of Science (JSPS) KAKENHI to Y.K. (18H03743, 21H02273, 24K09042), Y.K., M.N. (21K19240; 23K17996), Y.K., H.A., and N.I. (24K02048), S.M. (24H00579).
Author information
Authors and Affiliations
Contributions
M.M. performed all light microscopy and qPCR analyses. M.M., Y.K., and R.O. contributed to the generation and assembly of time-series transcriptomic data. Y.K., R.O. analyzed the transcriptomic data. Y.K. performed nuclear genome sequencing, and G.T. conducted genome assembly. Y.K., M.M. amplified and sequenced specific cDNA and gDNA loci. Y.K. performed phylogenetic analyses of protein sequences. S.Y., M.M. carried out RNAi experiments. Y.K., M.N., M.M., T.K., and H.I. performed CRISPR/Cas9 genome editing. M.N., M.I., and Y.K. conducted immunoblotting. M.M., Y.K., and N.I. performed immunofluorescence microscopy. M.M. measured photosynthetic activity. Y.K. quantified intracellular polysaccharide grains. Y.K., M.N. conducted the NanoLuc reporter assays. Y.K. and M.M. were responsible for culturing associated with all experiments. Y.K., M.N., H.A., K.A., and S.M. contributed to study design. Y.K. wrote the manuscript. All authors discussed the results and contributed to the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kashiyama, Y., Maruyama, M., Nakazawa, M. et al. Transient molecular chimerism for exploiting xenogeneic organelles. Nat Commun 17, 2371 (2026). https://doi.org/10.1038/s41467-026-70516-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-70516-x