Abstract
Flowering plants rely on double fertilization for their sexual reproduction. A pair of immotile sperm cells (SCs) are conveyed through a pollen tube into the female gametophyte, where they fertilize the egg cell and central cell to give rise to the embryo and endosperm in seeds, respectively1,2,3,4. The SCs and the pollen vegetative nucleus (VN) move as a male germ unit (MGU) in pollen tubes5,6. The tryptophan-proline-proline (WPP) domain-interacting tail-anchored proteins (WITs) and WPP domain-interacting proteins (WIPs) are involved in VN migration, and the cytoskeleton is required for MGU transportation in pollen tubes7,8,9,10,11. Here we report that two kinesins, referred to as HUG1 and HUG2, localize at the VN envelope in a WIT- and WIP-dependent manner and surround the SCs in pollen. Mutation of HUG1 and HUG2 leads to disconnected VN and SCs, impaired MGU transportation and reduced plant fertility, supporting key roles of HUG proteins in MGU formation and transportation in pollen.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The primer sequences used are provided in Extended Data Table 1. Biological materials can be obtained upon request. Source data are provided with this paper. All other data supporting the findings of this study are available within the paper and its Supplementary Information.
References
Sprunck, S. Twice the fun, double the trouble: gamete interactions in flowering plants. Curr. Opin. Plant Biol. 53, 106–116 (2020).
Johnson, M. A., Harper, J. F. & Palanivelu, R. A fruitful journey: pollen tube navigation from germination to fertilization. Annu. Rev. Plant Biol. 70, 809–837 (2019).
Zhong, S., Lan, Z. & Qu, L.-J. Ingenious male–female communication ensures successful double fertilization in angiosperms. Annu. Rev. Plant Biol. https://doi.org/10.1146/annurev-arplant-083123-071512 (2025).
Lord, E. M. & Russell, S. D. The mechanisms of pollination and fertilization in plants. Annu. Rev. Cell Dev. Biol. 18, 81–105 (2002).
McCue, A. D., Cresti, M., Feijó, J. A. & Slotkin, R. K. Cytoplasmic connection of sperm cells to the pollen vegetative cell nucleus: potential roles of the male germ unit revisited. J. Exp. Bot. 62, 1621–1631 (2011).
Dumas, C., Knox, R. B. & Gaude, T. The spatial association of the sperm cells and vegetative nucleus in the pollen grain of Brassica. Protoplasma 124, 168–174 (1985).
Motomura, K. et al. Persistent directional growth capability in Arabidopsis thaliana pollen tubes after nuclear elimination from the apex. Nat. Commun. 12, 2331 (2021).
Zhou, X. & Meier, I. Efficient plant male fertility depends on vegetative nuclear movement mediated by two families of plant outer nuclear membrane proteins. Proc. Natl Acad. Sci. USA 111, 11900–11905 (2014).
Åström, H., Sorri, O. & Raudaskoski, M. Role of microtubules in the movement of the vegetative nucleus and generative cell in tobacco pollen tubes. Sex. Plant Reprod. 8, 61–69 (1995).
Laitiainen, E., Nieminen, K. M., Vihinen, H. & Raudaskoski, M. Movement of generative cell and vegetative nucleus in tobacco pollen tubes is dependent on microtubule cytoskeleton but independent of the synthesis of callose plugs. Sex. Plant Reprod. 15, 195–204 (2002).
Wang, X. et al. Distinct functions of microtubules and actin filaments in the transportation of the male germ unit in pollen. Nat. Commun. 15, 5448 (2024).
Zhang, J. et al. Sperm cells are passive cargo of the pollen tube in plant fertilization. Nat. Plants 3, 17079 (2017).
Miller, D. D., Scordilis, S. P. & Hepler, P. K. Identification and localization of three classes of myosins in pollen tubes of Lilium longiflorum and Nicotiana alata. J. Cell Sci. 108, 2549–2563 (1995).
Heslop-Harrison, J. & Heslop-Harrison, Y. Myosin associated with the surfaces of organelles, vegetative nuclei and generative cells in angiosperm pollen grains and tubes. J. Cell Sci. 94, 319–325 (1989).
Heslop-Harrison, J. & Heslop-Harrison, Y. Conformation and movement of the vegetative nucleus of the angiosperm pollen tube: association with the actin cytoskeleton. J. Cell Sci. 93, 299–308 (1989).
Schattner, S., Schattner, J., Munder, F., Höppe, E. & Walter, W. J. A tug-of-war model explains the saltatory sperm cell movement in Arabidopsis thaliana pollen tubes by kinesins with calponin homology domain. Front. Plant Sci. 11, 601282 (2021).
Nebenführ, A. & Dixit, R. Kinesins and myosins: molecular motors that coordinate cellular functions in plants. Annu. Rev. Plant Biol. 69, 329–361 (2018).
Lee, Y. R. J., Qiu, W. & Liu, B. Kinesin motors in plants: from subcellular dynamics to motility regulation. Curr. Opin. Plant Biol. 28, 120–126 (2015).
Tian, J. et al. Orchestration of microtubules and the actin cytoskeleton in trichome cell shape determination by a plant-unique kinesin. eLife 4, e09351 (2015).
Yun, M. et al. Rotation of the stalk/neck and one head in a new crystal structure of the kinesin motor protein, Ncd. EMBO J. 22, 5382–5389 (2003).
Kocik, E., Skowronek, K. J. & Kasprzak, A. A. Interactions between subunits in heterodimeric Ncd molecules. J. Biol. Chem. 284, 35735–35745 (2009).
Liu, L. et al. Actomyosin and CSI1/POM2 cooperate to deliver cellulose synthase from Golgi to cortical microtubules in Arabidopsis. Nat. Commun. 14, 7442 (2023).
Dai, L. et al. The TOR complex controls ATP levels to regulate actin cytoskeleton dynamics in Arabidopsis. Proc. Natl Acad. Sci. USA 119, e2122969119 (2022).
Boavida, L. C., Qin, P., Broz, M., Becker, J. D. & Mccormick, S. Arabidopsis tetraspanins are confined to discrete expression domains and cell types in reproductive tissues and form homo and heterodimers when expressed in yeast. Plant Physiol. 163, 696–712 (2013).
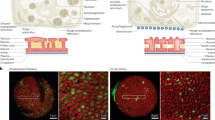
Sugi, N. et al. The peri-germ cell membrane: poorly characterized but key interface for plant reproduction. Nat. Plants 10, 1607–1609 (2024).
Tamura, K. et al. Myosin XI-i links the nuclear membrane to the cytoskeleton to control nuclear movement and shape in Arabidopsis. Curr. Biol. 23, 1776–1781 (2013).
Richardson, D. N., Simmons, M. P. & Reddy, A. S. N. Comprehensive comparative analysis of kinesins in photosynthetic eukaryotes. BMC Genomics 7, 18 (2006).
Xing, H. L. et al. A CRISPR/Cas9 toolkit for multiplex genome editing in plants. BMC Plant Biol. 14, 327 (2014).
Long, Y. et al. In vivo FRET–FLIM reveals cell-type-specific protein interactions in Arabidopsis roots. Nature 548, 97–102 (2017).
Acknowledgements
We thank Z. Wang and B. Zhu (Beijing Normal University) for helpful discussion. We acknowledge the experimental technology centre for life sciences at Beijing Normal University and thank X. Zhang for technical support. We thank Y.-L. Zhang (Imaging Core Facility of the Protein Research Center for Technology Development, Tsinghua University) for technical assistance for the FRET–FLIM assay. We thank L.-J. Qu (Peking University) for the drop1−/− drop2+/− pWIP1::WIP1-tdTomato seeds and D. Maruyama (Yokohama City University) for sharing the RHT plasmids. This work was supported by grants from the National Natural Science Foundation of China (nos 32270350 and 32070194 to Y. Zhang, 32100279 to T.W. and 32400563 to L.D.), the Fundamental Research Funds for the Central Universities (no. 2243200007 to Y. Zhang) and open funds of the State Key Laboratory of Plant Environmental Resilience (SKLPERKF2508). H.E.M. acknowledges funding from a Natural Sciences and Engineering Research Council of Canada Discovery Grant (no. 2020-05959), an Ontario Early Researcher Award (no. ER21-16-256) and the Canada Research Chairs programme (Canada Research Chair in Plant Cell Biology). Y. Zhu is supported by the China Scholarship Council (CSC no. 202106100034).
Author information
Authors and Affiliations
Contributions
Y. Zhang and C.X. initiated the project. Y. Zhang, Y.Y., L.D. and B.W. designed the experiments. Y.Y., L.D., B.W., Z.S., C.X., Y.W., J.Z., Y. Zhu, Y.H., H.E.M. and T.W. performed the experiments. X.L. and T.W. generated the wip1 wip2 wip3 mutants. X.Z., Y.G. and B.X. performed the phylogenetic analysis. Y. Zhang, Y.Y., L.D., B.W. and T.W. analysed the data. Y. Zhang, Y.Y., L.D., B.W. and H.R. wrote, reviewed and edited the manuscript with input from all the authors. Y. Zhang supervised the research and acquired funding.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Plants thanks Giampiero Cai, Iris Meier and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Subcellular localization analysis of 17 kinesin proteins with high expression in pollen.
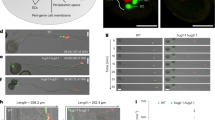
a, Representative images of 6 images for each kinesin proteins showing the subcellular localization of the 17 kinesins with high expression in pollen. The C-terminaly mNeonGreen-tagged constructs under the native promoters were transformed in the relevant mutant background for the subcellar localization analysis of each kinesin. Notably, two kinesins in the same subclade, referred to as HUG1 and HUG2, exhibited localization in large compartments within pollen tubes. Scale bars, 10 μm. b, c, Representative images of 4 images showing the localization of HUG1-mNeonGreen (b) and HUG2-mNeonGreen (c) surrounding the vegetative nucleus (VN, indicated by the arrows) and the two sperm cells (SCs). DNA in the VN and the SCs were stained with DAPI. Scale bars, 5 μm.
Extended Data Fig. 2 Phylogenetic analysis of kinesin genes in Arabidopsis and other plants.
a, Phylogenetic tree of 61 kinesins in Arabidopsis. Except for three kinesins that did not have family divisions, the other 58 kinesins are classified into 10 families, which are represented by different colors. The 17 bold marked kinesins are highly expressed in pollen based on data from eFP Browser (http://bar.utoronto.ca/). b, The phylogenetic relationships of the orthologs of HUG1 and HUG2 in green plants. Genes from distinct species are represented by differently colored branches, which correspond to the major taxonomic groups.
Extended Data Fig. 3 HUG1 and HUG2 are highly expressed in pollen.
a, Relative expression levels of HUG1 and HUG2 in different tissues and organs in Arabidopsis. The data were from publicly available RNA seq data (Arabidopsis eFP browser; http://bar.utoronto.ca/). The expression data was normalized with the Gene Chip operating software (GCOS) method with a Target intensity (TGT) and background (Bkg) threshold values of 100 and 20, respectively. The error bars represent means ± s.d. HUG1 and HUG2 are expressed in pollen. b, c, GUS activity assays in transgenic Arabidopsis plants that harbor HUG1 or HUG2 promoter-driven GUS construct. Representative images of 3 images showing GUS signals in inflorescence and pollen. Pollen grains were cultured in vitro for 2.5 h. Scale bars for inflorescence, 0.5 mm. Scale bars for pollen, 10 μm.
Extended Data Fig. 4 Molecular and phenotypical analysis of hug1, hug2 and their double mutants.
a, b, Gene structure and mutant alleles of HUG1 and HUG2. Boxes indicate coding region and UTRs; bold black lines indicate intron; black arrows indicate primers used for qRT-PCR. T-DNA positions in the mutant lines hug1-1 (SALK_023598) and hug2-1 (GABI_525E10) are indicated. CRISPR/Cas9 genome editing was applied to generate two independent double mutants of HUG1 and HUG2 (hugDKO-1 and hugDKO-2). hugDKO-1 and hugDKO-2 have a base G or T inserted into the fourth exon of HUG1, respectively, resulting in premature termination of HUG1 in both mutants. hugDKO-1 has an A base inserted into the fourth exon of HUG2 and hugDKO-2 has T base inserted into the fourth exon of HUG2, resulting in premature termination of HUG2 in both mutants. Red asterisks indicate the stop codons. c, qRT-PCR analysis of HUG1 and HUG2 transcript levels in Col-0, hug1-1, hug2-1 and hug1-1 hug2-1 pollen grains. Primer positions are indicated in (a). Two-tailed Students’ t-tests were used to determine significance. The error bars represent means ± s.e.m, n = 3. d, Representative images of Col-0, hug1-1, hug2-1 and hug1-1 hug2-1 pollen tubes. Pollen grains were cultured on germination media for 3.5 h. Scale bars, 100 μm. e, Quantitative analysis of pollen tube length. n = 40 pollen tubes from each genotype. The error bars represent means ± s.d. ns indicates not significant by two-tailed Student’s t-test, P value > 0.05.
Extended Data Fig. 5 HUGs are involved in establishing the VN-SCs connection but does not affect the length of cytoplasmic projection.
a, Representative images of 10 images showing the disconnection of the VN and the sperm cells (SCs) in pHUG2:HUG2-mNeonGreen-complementation lines in hug1-1 hug2-1 background. The blue arrow indicates the cytoplasmic projection tip of the SC, which was decoupled with the VN. Scale bars, 10 μm. b, Statistical analysis on the length of cytoplasmic projection in sperm cells. n = 27, 25, 23 and 25 for each genotype from left to right. In each box plot, the box indicates the 25th-75th percentiles, the center line indicates the median and the whiskers indicate the full data range. Two-tailed Students’ t-tests were used to determine significance.
Extended Data Fig. 6 The impaired functions of HUG1 and HUG2 causes major defects in male gametophytes.
a, Developing seeds in siliques 10 days after reciprocal crosses between Col-0 plants and different mutants or complementation lines. The images are representative of 5 samples for each genotype. Asterisks indicate unfertilized ovules, which were counted as aborted seeds. Scale bars, 0.5 mm. b, Satatistical analysis of seed abortion rates in (a). n = 5 siliques for each genotype. The error bars represent means ± s.d. Different letters [a-d] denote groups that show statistically significant differences amongst groups, P < 0.0001, One-way ANOVA. c, Representative images showing pollen tube length growing in Col-0 pistils. Pollen grains from Col-0, hug1-1 hug2-1 and hugDKO-1 were pollinated on WT stigma and aniline blue was applied to stain pollen tubes 4.5 h after pollination (HAP). The red lines indicated the longest pollen tubes in the pistils. Scale bars, 100 μm. d, Statistical analysis of the length of pollen tubes. n = 9, 12, 10 siliques for each genotype from left to right. The error bars represent means ± s.d. Two-tailed Student’s t-tests were used to determine significance. e, Images of semi in vitro pollen tube targeting to the ovule. The white arrow indicates the first pollen tube, and the white asterisk marks the second pollen tube. Scale bars, 0.1 mm. f, Quantification of pollen tube guidance efficiency to the ovules for each genotype. n = 200 ovules for each genotype. g, Quantification of the number of pollen tubes attracted by ovules for each genotype. The number of ovules with one or two pollen tubes is indicated.
Extended Data Fig. 7 Sequence alignments of the motor domain of HUG1, HUG2 and Drosophila melanogaster Ncd.
Multiple sequence alignment of the motor domain of Arabidopsis HUG1, HUG2 and a Drosophila melanogaster Ncd (non-claret disjunctional) protein. Black indicates identical amino acids, and gray indicates similar amino acids. The three conserved amino acids Arg552, Glu585 and Asn600, which are important for the motor activity of Ncd, are mutated to Ala (R838A), Ala (E871A/E749A) and Lys (N890K) in HUG1 or HUG2, respectively, in this study.
Extended Data Fig. 8 The restricted pollination assay for the point-mutated HUG1 complementation pollens and the subcellular localization of HUG1(N890K)-mNeonGreen, HUG1(R838A)-mNeonGreen and HUG1(E871A)-mNeonGreen in pollen tubes.
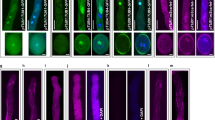
a, Domain composition and mutation sites of HUG1. Light green box, CC domain; pink box, motor domain; green box, mNeonGreen protein. b, Representative siliques from the WT, hug1-1 hug2-1, and the HUG1-mNeonGreen- or HUG1(N890K)-mNeonGreen-complementation lines. White asterisks indicated unfertilized ovules, which were counted as aborted seeds. Scale bars, 0.5 mm. c, Statistical analysis of seed abortion rates. n = 24, 24, 21, 20 and 20 siliques for each genotype from left to right. The error bars represent means ± s.d. Different letters denote groups that show statistically significant differences from other groups, P value < 0.0001, One-way ANOVA. d, Schematic drawing for separating mNeonGreen-positive pollen grains from those without mNeonGreen fluorescence. Pollen grains from the hemizygous complementation lines of HUG1(E871A)-mNeonGreen or HUG1(R838A)-mNeonGreen in the homozygous hug1-1 hug2-1 mutant background were used in these experiments. Fifteen mNeonGreen-positive or negative pollen grains were pollinated on WT stigma, respectively. The control was also tested with 15 isolated pollen grains. e, Developing seeds in siliques 10 days after pollination with pollen grains separated in (d). Scale bars, 0.5 mm. f, Statistical analysis of seed number. n = 6 siliques for each genotype. The error bars represent means ± s.d. Different letters denote groups that show statistically significant differences from other groups, P value < 0.0001, One-way ANOVA. g–j, Representative images of 8 images showing the subcellular localization of HUG1-mNeonGreen (g), HUG1(N890K)-mNeonGreen (h), HUG1(R838A)-mNeonGreen (i), HUG1(E871A)-mNeonGreen (j) and the nuclear marker H2B-tdTomato (the marker of VN and SN) in pollen tubes. Scale bars, 10 μm. k, Representative images of 3 time-lapse images showing that the elongated VN in the HUG1(R838A)-mNeonGreen complementation pollen tubes aligned with cytosolic microtubules. Scale bars, 5 μm. l, Relative fluorescent signal intensity along white lines in (k).
Extended Data Fig. 9 HUG proteins bind to microtubules in vitro and show additional localization along microtubules in pollen tubes.
a, Domain composition of HUG2. Light green box, CC domain; pink box, motor domain. b, Co-sedimentation assay for the binding of HUG2425-1025 with microtubules in vitro. Representative images from three independent experiments. c, Representative images of 8 images showing the dot-like localization of HUG1-mNeonGreen on microtubules (labelled by mCherry-MBD) in pollen tubes. The right panels show overexposed-images inside the white rectangle. d, Relative fluorescent signal intensity along white line in (c). e, Representative images of 3 time-lapse images showing the formation of HUG1-mNeonGreen-labelled dot-like compartments. f, Kymograph along the yellow line in (e). Note that HUG1-mNeonGreen-labelled dot-like compartments were derived from the VN and moved along the microtubules. Scale bar, 5 μm. g, Representative images of 3 images showing the colocalization of HUG2-mNeonGreen with the microtubule network in pollen tubes. h, Relative fluorescent signal intensity along white line in (g).
Supplementary information
Supplementary Video 1 (download AVI )
The movement of VN and SCs in Col-0, hug1-1-C (VN and SCs are connected), hug1-1-DC (VN and SCs are disconnected), hug1-1 hug2-1 and hugDKO-1 in pollen tubes.
Supplementary Video 2 (download AVI )
MGU movement in HUG1–mNeonGreen, HUG1(E871A)–mNeonGreen or HUG1(R838A)–mNeonGreen complementation lines in the hug1-1 hug2-1 mutant background in the absence or presence of oryzalin in pollen tubes.
Supplementary Video 3 (download AVI )
HUG1–mNeonGreen-labelled dot-like compartments are derived from the VN and move along microtubules in pollen tubes.
Source data
Source Data Figs. 1, 2 and 4 and Extended Data Figs. 3–6 and 8 (download XLSX )
Statistical source data for Figs. 1 and 2 and Extended Data Figs. 3–6 and 8, and statistical source data and unprocessed western blots for Fig. 4.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yan, Y., Dai, L., Wang, B. et al. Kinesin proteins HUG1 and HUG2 are essential for the formation and transportation of male germ units in Arabidopsis pollen tubes. Nat. Plants 11, 1489–1499 (2025). https://doi.org/10.1038/s41477-025-02064-z
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41477-025-02064-z