Abstract
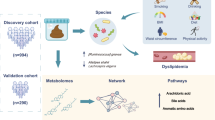
In this 12-week trial, 136 participants with moderately dyslipidemia were randomly assigned to receive Lactiplantibacillus plantarum (LP) or placebo. In the intention-to-treat analysis, the group-by-time interaction did not reach statistical significance. However, in per-protocol set (adherence ≥85% and no antibiotic use), LP supplementation reduced low-density lipoprotein cholesterol (LDL-C) (−0.118 mmol/L) and total cholesterol (TC) (−0.163 mmol/L), compared with the placebo (both Pgroup×time < 0.05). Post-intervention group differences were identified in gut microbial genera and species, correlated with changes in bile acids, which in turn were jointly related to lipid reduction. Microbiota-based machine learning models well-predicted the lipid reductions. Subjects with lower genetic risk scores experienced large decreases in LDL-C (Mean ± SD: −0.749 ± 0.632 mmol/L) and TC (−1.306 ± 0.436 mmol/L) (both Ptrend and Pinteraction < 0.001). Our data supported the beneficial effects of LP in patients with moderate dyslipidemia involving gut microbiota and host genetics.
Similar content being viewed by others
Introduction
Hypercholesterolemia, particularly elevated low-density lipoprotein cholesterol (LDL-C), was causally associated with cardiovascular events1. Globally, elevated plasma LDL-C was attributable to more than a third of deaths from ischemic heart disease or ischemic stroke2. While healthy dietary patterns effectively reduce blood lipids3,4, reversing unhealthy dietary habits could be challenging given dyslipidemia per se is imperceptible if no other related outcomes happen.
Consequently, there is a growing interest in alternative interventions. Supplementation of specific probiotics has emerged as a potential strategy, as suggested by clinical trials5,6,7. The underlying mechanisms vary by taxa, including deconjugation of bile acids (BA) by bile salt hydrolases8, binding cholesterol, embedding cholesterol in the probiotic cell membrane9, and converting cholesterol to coprostanol10. Among the various probiotic strains investigated, Lactobacillus (sensu lato) has garnered significant attention for its lipid-lowering potential. While, its beneficial effects on LDL-C have been well-documented11,12,13, current evidence implies significant knowledge gaps regarding specific target populations. First, previous studies primarily focused on individuals with high lipid levels. It remains to be elucidated whether probiotic Lactobacillus would be effective among people with moderately elevated lipids before the onset of lipid-related complications. Second, ethnic differences play a crucial role. Previous trials were largely conducted in Western populations. However, the Chinese population has relatively lower LDL-C levels, and different genetic susceptibility to dyslipidemia1 compared to Western populations. Given that an estimated 400 million people in China suffer from dyslipidemia14, little is known about the potential benefit of Lactobacillus in this specific demographic.
Within this genus, Lactiplantibacillus plantarum (L. plantarum, LP)—formerly known as Lactobacillus plantarum—has emerged as a promising candidate. Previous studies suggested that supplementation with Lactiplantibacillus plantarum was able to alter the gut microbiota profiles15. However, little has been documented regarding the detailed correlations of microbial change and their metabolites with host lipid change, which, we believe, are very informative in elucidating the mechanisms in host-microbial interactions for lipid optimization. Moreover, individual differences in gut microbiota may interfere with the effect of probiotic intervention on the improvement of serum lipids16. Determining the initial gut microbial panels predictive of lipid responsiveness may facilitate the development of precision strategies of probiotic supplementation and clarify the underlying mechanisms. In addition to the microbiome, host genetics represent another layer of complexity influencing intervention outcomes. Recent studies also revealed that lipid changes in response to dietary and lifestyle interventions were partially determined by host genetic background17,18. However, the genetic effects on blood lipid decline driven by L. plantarum are unclear.
Thus, this study aimed to evaluate the effects of a L. plantarum supplement for 12 weeks on lipid profile and gut microbiota in individuals with mild lipid abnormalities but no medication use. Moreover, to elucidate the responsiveness differences determined by host genetic and gut microbiota background.
Results
Participants and characteristics
Overall, a total of 120 subjects completed the entire intervention, with 16 subjects dropped out (Fig. S1). Among the 120 completers, 19 participants were excluded from the per-protocol set due to antibiotic use (n = 9) or adherence rate < 85% (n = 10), resulting in a final per-protocol set of 101 participants. In terms of general adherence, the mean adherence score was 0.93 ± 0.08, with no between-group difference. In total, 39.3% of the participants were female, with a mean age of 45.2 years (Table 1). All the baseline characteristics were comparable.
Effects on plasma lipid profiles
Intention-to-treat analysis showed no statistically significant time-by-group effect for LDL-C and total cholesterol (TC) (P group×time = 0.075 and 0.053, respectively) (Table 2), while in the per-protocol set (adherence ≥ 85% and not taking antibiotics during the trial), LDL-C and TC levels in the LP group showed a significant reduction over the 12-week intervention compared to the placebo (PLA) group (Pgroup×time = 0.041 and 0.023, respectively) (Table 3). No significant time-by-group interaction was observed for other outcome variables.
Safety and adverse events
There was no variation within or between groups for liver and kidney biomarkers (Table S1). During the study, no severe adverse event was reported (Table S2); mild adverse events, mainly gastrointestinal discomfort, were recorded, with no between-group difference.
Effects on gut microbiota
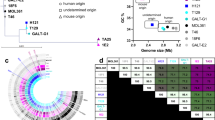
There was no group-difference in baseline species community richness (Chao1 estimate) and diversity (Shannon index, ACE estimate), while the Chao1 estimate and Shannon index were lower in the LC group than in the PLA group at 12 weeks (Fig S2). No between-group difference can be detected for the genus and species levels at baseline and 12 weeks (all false discovery rate [FDR] P > 0.05), when the taxa abundance was compared individually. However, when different taxa levels were assessed jointly by the linear discriminant analysis (LDA) combined effect size (LEfSe) analysis, 21 bacterial taxa showed between-group differences after interventions (Fig. 1A, Fig S3). In particular, Lactiplantibacillus plantarum was significantly enriched in the LP group. In total, 5 differential genera and 2 differential species were confirmed when compared individually(Fig. 1B, C, FDR P < 0.05). At the genus level, the relative abundance of Alistipes and Pseudoflavonifractor were lower, that of Solibaculum, Lactiplantibacillus, and Howardella were higher in the LP group than the PLA group. At the species level, the relative abundance of Solibaculum mannosilyticum was higher, whereas that of Bacteroides caccae was lower in the LP group.
A Taxa with linear discriminant analysis (LDA) scoreså 2 and P < 0.05 in bacterial communities by group. Circles indicate phylogenetic levels from phylum to species. B, C Differential genera (B), and species (C) confirmed with Wilcoxon rank-sum test. The color of the origin is that of the group with higher relative abundance. * FDR P < 0.05.
Correlations of changes in microbial taxa, bile acids, and blood lipid profiles in the LP group
Among all the differential microbial taxa (genera and species), we observed a positive correlation between the change in Enterococcus faecalis and the change in TC, and negative correlations for the changes in Solibaculum and Solibaculum mannosilyticum with the change in triglycerides (TG) in the LP group (Fig. 2A). We further characterized the alterations in fecal bile acid profiles (Table S3). Moreover, the changes in differential microbial taxa were primarily correlated with changes in fecal bile acids (Fig. 2A, Table S4). In particular, we identified positive correlations between changes in Lactiplantibacillus and L.plantarum and changes in glycocholic acid and isolithocholic acid. Moreover, the change in taurodehydrocholic acid was inversely correlated with the change in Enterococcus faecalis, related to TC improvement. In addition, we also observed that BA scores were associated with LDL-C and TG (P = 0.047 and 0.010, respectively), whereas the association with TC did not reach statistical significance (P = 0.052) (Fig. 2B–D).
A Significant partial spearman’s correlations among changes in differential genera(Δ)or species(▯), bile acids, and changes in lipid profiles, adjusted for age and sex. The line colors indicate the direction of the association, with red for positive correlations and blue for negative correlations. The width of the line represents the strength of the association. B–D Fitted line plot of the linear regression between the measured lipid value (i.e., dependent variable) and bile acids scores of lipid profiles (i.e., independent variable). TC total cholesterol, TG triglyceride.
Predisposition to the intervention responsiveness by gut microbiota and genetic backgrounds
The predictive performances of baseline gut microbiota are presented in Fig. 3A, B. The highest area under the receiver operator characteristics curves (AUC) (95% CI) values of machine learning (ML) models were 0.905 and 0.810 for predicting responders ( >5% decrease) in LDL-C and TC, with unique top ten contributing features, respectively. Genetic risk scores (GRS) for LDL-C and TC were developed with 12 and 11 single-nucleotide polymorphisms (SNPs) that interacted with intervention on the change of LDL-C, and TC, respectively (Tables S5, S6). Lower GRSs were associated with significant decreases in LDL-C (−0.749 ± 0.632 mmol/L) and TC (−1.306 ± 0.436 mmol/L) in subjects receiving L.plantarum (both P for trend and interaction <0.001, Fig. 3C, D). Correlation networks were identified among the top ten predicting species, genetic background (GRS, individual SNPs), and changes in bile acids for LDL-C and TC, respectively (Fig. 3E, F, Table S7, S8). In particular, the GRS of LDL-C was correlated with variations in Bifidobacterium longum and the changes in glycocholic acid. Interestingly, Roseburia intestinalis, contributing to the predicting of both LDL-C and TC reduction, was connected to both genetic variance in HLA-C (rs3873379) and changes in bile acids, including glycohyocholic acid, isolithochomic acid, and glycocholic acid. In addition, GRS for TC was linked with an unclassified species in the genus Acetivibrio, which in turn was correlated with changes in taurocholic acid and 12-ketochenodeoxycholic acid. The explained variances of LDL-C change upon intervention were 26.4% by microbiome risk score (MRS) and 11.7% by GRS; while for TC, it was 18.5% and 21.9%, respectively (Fig. 3G, H).
A, B Baseline gut microbiota predicting the improvement of lipid profiles (A, LDL-C; B, TC) upon interventions using machine learning models. C, D The interactions between GRS and intervention on changes in LDL-C (C), TC (D). Data points represent the mean change from baseline, and error bars indicate the standard error. The median across the tertile of GRSLDL-C was 12.23 (range: 8.07–14.01), 15.48 (range: 14.02–16.63), and 17.95 (range: 16.64–21.21). The median across the tertile of GRSTC was 11.16 (range: 6.82–13.58), 14.89 (range: 13.59–16.17), and 17.37 (range: 16.18–19.69). P-values for trend and interaction were calculated using generalized linear models adjusting for age and sex. E, F The network work of genetic background, gut microbiota, and bile acids (E, LDL-C; F, TC). Partial spearman’s correlations among changes in relative genera or species abundance, genetic background, and changes in bile acids with adjustment for age and sex. The line color indicated the direction of the association, with red for positive correlation and blue for negative correlation. The width of the line represented the strength of the association; as the line widens, the association grows stronger. G, H Degree of interpretability of genetic background and gut microbiota for interventions to improve blood lipids (G, LDL-C; H TC).
Discussion
In this randomized trial, the intention-to-treat analysis showed a borderline but non-significant trend in lipid reduction. However, among participants strictly adhering to the protocol (per-protocol analysis), L. plantarum supplementation for 12 weeks was effective in lowering LDL-C and TC compared to the placebo. Both initial gut microbiota and genetic background predisposed individual differences in lipid reduction after the intervention.
Previous meta-analyses suggested that L.plantarum was able to reduce lipid levels in patients with hyperlipidemia11,12,13, primarily in participants with relatively high total cholesterol levels. A trial by Fuentes et al.19. suggested that in subjects with high baseline TC (6.49 mmol/L to 7.76 mmol/L), L.plantarum CECT 7527, CECT 7528, and CECT 7529 significantly reduced TC and LDL-C; however, in their subjects with low baseline TC (5.17 mmol/L to 6.47 mmol/L), LDL-C reduction was not significant, and the TC reduction effect was smaller. In clinical practice, the American Heart Association, American College of Cardiology20, and Chinese Medical Association21 recommend those with a high risk of 10-year Atherosclerotic Cardiovascular Disease (ASCVD) to actively consider drug therapy for lipid control, yet recommend those with moderate or low risks to consider lifestyle modification firstly. This clinical practice, in fact, calls for randomized controlled trial-based evidence for lifestyle strategies among people with moderate and low ASCVD risk. Regarding L.plantarum, evidence was rare for this sub-clinical population. In our study, we found significant time-by-group effects in the per-protocol set, and the mean difference of the change in LDL-C (−0.118 mmol/L) and TC (−0.163 mmol/L) account for 3.0% and 2.8% of their mean initial values. We consider the per-protocol results to be the most relevant because of instances of poor treatment compliance and of antibiotic use in the full intent-to-treat population, which are likely to reduce the effects of the probiotic. Previous studies have suggested that each 1% reduction in TC may reduce the risk of coronary heart disease by 2% to 3%22, and that each 0.26 mmol/L reduction in LDL-C may contribute to a 7.1% reduction in the relative risk of coronary heart disease23. Therefore, our results supported the lipid-lowering effects of adhering to L.plantarum in subjects with moderately elevated lipids.
At 12 weeks, the LP group was remarkably enriched in Firmicutes, including genus Lactiplantibacillus at the genus level and L.plantarum at the species level, indicating a traceable adherence of the L.plantarum supplementation. Interestingly, the LP group also showed an enrichment in Enterococcus faecalis (Fig. 1). In a rat model, supplementation of Enterococcus faecalis significantly reduced serum TC, TG, and LDL-C levels24, and in a mice model, oral administration of Enterococcus faecalis was able to reduce serum TC by means of increasing the liver cholesterol transporter expression and modulating the composition of gut microbiota25. Moreover, Alistipes was lowered after the LP intervention. A previous study showed that transplant of the microbiota from humans with elevated cholesterol to mice induced a high cholesterol phenotype associated with elevated abundance of Alistipes26. In addition, our baseline data showed an inverse correlation between abundance of Alistipes and plasma insulin level (r = −0.31, P = 0.001). Notably, the enrichment of Lactiplantibacillus/L.plantarum in the LP group provides biological corroboration of exposure and supports a causal interpretation of downstream microbial shifts. At the same time, the co-enrichment of other taxa (e.g., Enterococcus faecalis) should be interpreted as ecosystem-level responses rather than direct probiotic engraftment.
One of the major mechanisms underlining the cholesterol-lowering effects of L.plantarum is the reduction of bile salt reabsorption through the enterohepatic cycle (acting through bile salt hydrolases, BSHs)27. In the small intestine, conjugated BA could be hydrolyzed by microbial BSHs to produce unconjugated BA, which are less soluble and more hydrophobic, making it difficult to passively enter the enterocytes. This leads to increased excretion of BA via the feces. Consequently, liver is required to mobilize more systemic cholesterol to resynthesize bile salts for the next digestive cycle, thereby lowering serum cholesterol levels27. In our study, the post-intervention differential genera or species were largely correlated with changes in fecal bile acids, mostly with conjugated structure, which were jointly related to the changes in plasma lipid (Fig. 2). Our results endorsed the involvement of microbial BA metabolism in the L.plantarum-derived lipid optimization in human subjects, with measured changes in gut microbiome and fecal bile acids.
Despite a general reduction in plasma lipids after the L.plantarum supplementation, the degree of responsiveness varied considerably between individuals. Using ML models, we found that specific baseline microbial features performed well in predicting intervention responsiveness of LDL-C and TC with excellent AUCs. We found that baseline Bacteroides stercoris was more abundant among those responders for lowering LDL-C. Similarly, from a case-control study of metabolic dysfunction-associated fatty liver disease, gut Bacteroides stercoris showed good diagnostic performance28. Interestingly, we noticed a lower abundance of Bifidobacterium longum in responders of TC reduction. Previous clinical trials demonstrated that Bifidobacterium longum significantly alleviated hyperlipidemia by modulating the composition of the gut microbiota29,30. Our results suggested that suboptimal gut microbiota composition, such as low abundance in Bifidobacterium longum, could somewhat be compensated by supplementation with exogenous L.plantarum. To the best of our knowledge, the current study is the first one to demonstrate a strong interaction between GRS and L.plantarum intervention on lipid changes. Differences up to 1.0 ~ 1.5 mmol/L in LDL-C or TC reductions were found between participants with the highest and lowest lipid-GRS. Those genetic traits may provide a reference for the development of precision intervention strategies in L.plantarum supplementation for lipid health. The complex correlation networks between the predictive microbial species, genetic traits, and bile acid changes may have well-informed the host-microbiome interactions in decoding individualized effects upon L.plantarum supplementation. We found a higher explained variance of LDL-C reduction by MRS than that by GRS, highlighting the plasticity of gut microbiome in achieving optimal LDL-C, in contrast to the inborn genetic background. Conceptually, these findings align with the emerging precision-nutrition framework in which host genetics and baseline microbiota act as effect modifiers of dietary or probiotic interventions.
Clinically, these findings suggest that a well-characterized L. plantarum formulation may be considered as an adjunct, non-pharmacologic option for adults with moderately elevated lipids who are not yet receiving lipid-lowering medication, within a lifestyle-first management framework. The per-protocol benefit underscores the importance of adherence when evaluating probiotic effects in practice.
The limitations of the study may include the short intervention duration that restricted assessment of sustained effects. However, the 12-week period was sufficient to observe significant metabolic changes, and strict compliance monitoring was implemented to ensure the validity of these short-term observations. Possible fluctuations in plasma lipid could be affected by seasonal temperature fluctuation; to overcome this, we employed a randomized, double-blind, placebo-controlled design, which ensures that environmental factors such as seasonality affect both groups equally, thereby isolating the treatment effect of the probiotic. The sample size may not be powered for the per-protocol analysis given smaller between-group difference than assumed. Moreover, the limited sample size may have reduced the power in maximizing the SNPs in GRS; nevertheless, the sample size was strictly calculated based on the primary outcome (LDL-C reduction), and the GRS analysis is presented as an exploratory finding to generate hypotheses for future large-scale studies. Finally, regarding the inherent limits of 16S rRNA sequencing in distinguishing some genera, we utilized the PacBio full-length 16S rRNA sequencing platform to maximize taxonomic resolution compared to traditional V3-V4 region sequencing, and integrated untargeted metabolomics to provide functional validation of the microbial shifts. Additionally, given that probiotic effects are strain-specific, our findings are pertinent to the specific L. plantarum strain used in this study and should not be generalized to the entire species.
In summary, our study suggested that adherence to L.plantarum supplementation may effectively reduce LDL-C and TC levels, and optimize gut microbiota profiles and metabolism in subjects with moderately elevated lipids but no medication use. The initial gut microbiota features and genetic background of the subject may predispose the maximum effects of the intervention, suggesting a necessity to establish tailored probiotic interventions by considering both host genetics and individual gut microbiota potentials.
Methods
Study design and participants
This study is a 12-week double-blind, randomized, placebo-controlled trial conducted in Xi’an, China, from April 2022 to December 2022. The protocol was finalized prior to participant recruitment, and all procedures were conducted under ethics approval with written informed consent obtained before any study-related assessments. The inclusion criteria were: (1) age between 18 and 70 years, (2) blood lipid levels: 3.4 ≤ LDL-C < 4.9 mmol/L, 5.18 ≤ TC < 7.2 mmol/L, and 1.7 ≤ TG < 5.6 mmol/L; (3) have not received lipid-lowering drug treatment or having ceased such treatment for more than three months. The exclusion criteria included: (1) pregnancy, planning for conception, or lactation; (2) difficult in verbal expression; (3) unable to take the intervention products during the run-in period; (4) allergies to the intervention products; (5) ASCVD risk assessed as high based on “Guidelines for the Prevention and Treatment of Dyslipidemia in Chinese Adults 2016; 6) have undergone gastrointestinal surgery within 30 days, or have received antibiotic treatment (excluding medications for skin or genital tract issues) within 30 days; 7) hospitalized or acute gastrointestinal diseases in recent 2 weeks; 8) presence of severe conditions affecting the heart, liver, kidneys, hematopoietic system, or mental health; 9) regular consumption of probiotic products, or recent medication use affecting blood lipid within the last 2 weeks; 10) abnormal liver and kidney function, or a history of disorders affecting lipid metabolism, severe gastric hyperacidity, or severe constipation; 11) age <30 years and evidence suggesting a genetic predisposition to high blood lipids, defined as meeting ≥2 of the following 3 criteria: (i) untreated LDL-C ≥ 4.7 mmol/L; (ii) tendon/skin xanthomas or corneal arcus before age 45; (iii) a first-degree relative with familial hypercholesterolemia or premature ASCVD; 12) other health issues unsuitable to participate. This study was approved by the Medical Ethics Committee of the Second Affiliated Hospital and Xi’an Jiaotong University (ref no: 2021-009) according to the Declaration of Helsinki. All participants provided written informed consent before the intervention. The study was registered at https://www.chictr.org.cn (ChiCTR2200058631). Any protocol deviations and concomitant medication use during follow-up were documented at each study visit.
The participants were recruited by the research center (Health Management Department of the Second Affiliated Hospital, Xi’an Jiaotong University) through WeChat advertising and phone calls. A REDCap online system (https://www.wcrcnet.cn/redcap/) was used to collect the eligibility-related questionnaire information, followed by a face-to-face confirmation of their health examination report at the research center. Eligibility was assessed in a two-step process (online prescreening followed by in-person confirmation) to ensure that biochemical eligibility and exclusion criteria were verified before randomization. A total of 589 individuals who met the age requirement and blood lipid criteria were contacted by phone calls for the appointment of face-to-face confirmation. 453 were excluded due to exclusion criteria, personal schedule conflicts, or other reasons. Finally, 136 participants were successfully enrolled in the study (Fig S1).
Randomization and blinding
Before baseline examination, a simple block randomization (block size 4, ratio 1:1) was conducted by an independent statistician who was not involved in other parts of the study. All allocation details were contained in opaque sealed envelopes, which were opened and used sequentially at baseline visits. The intervention products for both groups were prepared by By-Health Co., Ltd. (Zhuhai, China) with identical attributes, including shape, color, smell, and outer packaging, with only one difference in group number label (1 or 2). Participants and researchers were blinded to treatment assignments until the entire study was completed.
Sample size
The sample size was determined by setting a between-group difference in LDL-C change of 0.38 mmol/L after the intervention31, assuming a standard deviation (SD) of LDL-C as 0.65 mmol/L for the target population. With 90% statistical power and a two-sided significance level (α) of 5% and allowing for 10% participant attrition, a total of 136 participants were needed (68 participants in each group).
Intervention
Before the baseline visit, there was a 7-day run-in period during which participants were required to take 1 sachet (placebo without informing the participants) per day to get used to the intervention. The run-in period was designed to familiarize participants with daily intake procedures and to identify potential non-adherence before randomization, under ethics approval. A total of 136 eligible subjects were randomly assigned to the LP group or the PLA group at the baseline (Week 0) and continued the intervention over the next 12 weeks, a period that allows sufficient time for stable probiotic colonization and to encompass the necessary physiological turnover period for host lipid pools. The LP group received a mixture of L. plantarum strains CECT 7527, CECT 7528, and CECT 7529 (1:1:1 ratio; 1.2 × 109 CFU per sachet/day, Life-space product), while the PLA group received maltodextrin powder (1 g per sachet/day). During the 12-week intervention phase, participants were required to take 1 sachet with water ( ≤37 °C) every day. All the participants received reading materials (both hard copy and electronic version) of lifestyle counseling consisting of 6 elements: balanced diet, physical activity, weight control, smoking, and alcohol restriction, and healthy sleep (Table S9). Participants were required to visit the research center at weeks 0, 6, and 12 to receive the intervention products and complete all necessary physical measurements, biological sample collection, and questionnaire surveys. Participants were also instructed to return any remaining intervention products at each visit and to report any adverse events, medication usage, or hospitalization events. The compliance score was defined as the number of consumed intervention product packets divided by the total number of intervention packets provided.
Outcomes and data collection
The primary outcome is the between-group difference in the change in LDL-C from baseline to week 12. The secondary outcomes are exploratory outcomes including changes in plasma TC, high-density lipoprotein cholesterol (HDL-C), TG, blood glucose, insulin levels, body mass index (BMI), and gut microbiota over the 12 weeks of intervention.
At baseline (week 0), information on demographics, lifestyle (smoking and alcohol drinking habits), physical activity, medical history, and food frequency questionnaires were collected from participants using the Redcap online questionnaire. At baseline, 6, and 12 weeks, Height and weight were measured to the nearest 0.1 cm and 0.1 kg, respectively, using an electronic scale (SK-CK/TCS-160D-W/H; Shuangjia, Shenzhen, China) while wearing lightweight clothing without shoes. After 5 min of rest, blood pressure was measured three times, with a 30 s interval between each measurement, using a digital monitor (Omron HEM-7124), and the mean value was used in the analysis. Blood samples were obtained by trained health workers after a 10 h fasting at each visit. Then, the blood samples were centrifuged and stored at −80 °C until measurement. Plasma levels of LDL-C, HDL-C, TC, TG, glucose, insulin, as well as liver function markers (alanine aminotransferase, aspartate aminotransferase, gamma-glutamyl transferase) and kidney function markers (creatinine, uric acid, urea nitrogen) were measured by an automatic biochemical analyzer (Beckman Coulter AU680). All measurements were performed following the manufacturer’s instructions and routine laboratory quality-control procedures.
Fecal sample collection, assessment of gut microbiota and BAs
Fecal samples were self-collected by participants at baseline (within 3 days before the intervention) and week 12 (the day of the visit, or within 3 days before or after the visit). A pair of fecal sample collection tubes equipped with ice packs and insulation bags, as well as printed instructions for sample collection, were provided to the participants. Fresh fecal samples were required to hand over to the research center within 4 hours after collection to ensure an immediate freezing to a −80 °C fridge until experiment. A video version of sample collection instruction was developed and provided to ensure the sample quality (online video: https://doi.org/10.6084/m9.figshare.25864888.v1). DNA was extracted from stool samples (200 mg) using the PF Mag-Bind Stool DNA Kit (Omega Bio-tek, Georgia, U.S.) following the manufacturer’s instructions. The bacterial 16S rRNA genes were amplified using the universal bacterial primers 27 F (5’-AGRGTTYGATYMTGGCTCAG-3’) and 1492 R (5’-RGYTACCTTGTTACGACTT-3’). Purified products were pooled in equimolar and DNA library was constructed using the SMRTbell prep kit 3.0 (Pacifc Biosciences, CA, USA) according to PacBio’s instructions. Purified SMRTbell libraries were sequenced on the Pacbio Sequel IIe System (Pacifc Biosciences, CA, USA) at the Majorbio Bio-Pharm Technology Co. Ltd. (Shanghai, China). After quality control, the average number of full-length 16S reads per sample was 25807. Rarefaction curves of observed features approached saturation across samples, supporting sufficient sequencing depth for downstream diversity analyses (Fig S4). The fecal metabolites were measured by LC-MS/MS32. Firstly, an aliquot of approximately 50 mg of each fecal sample was transferred into a 2 mL centrifuge tube, into which a single 6 mm grinding bead was placed. Metabolites were then extracted using 400 μL of a methanol-water mixture (4:1, v/v) that contained an internal standard (L-2-chlorophenylalanine, 0.02 mg/mL). The samples were homogenized at low-temperature conditions (using a Wonbio-96c frozen tissue grinder, Shanghai Wanbo Biotechnology Co., Ltd.; parameters: −10 °C, 50 Hz, 6 min). Following homogenization, the samples underwent ultrasonic extraction at 5 °C for 30 min (40 kHz). After extraction, the samples were incubated at −20 °C for 30 minutes to promote protein precipitation and then centrifuged at 13,000 g for 15 min at 4 °C. The resulting supernatants were obtained and transferred into autosampler vials forthe LC-MS/MS analysis. Chromatographic separation was performed using a Thermo UHPLC-Q Exactive HF-X system (Thermo Fisher Scientific, USA) coupled with an ACQUITY HSS T3 column (100 mm × 2.1 mm i.d., 1.8 μm; Waters, USA) at Majorbio Bio-Pharm Technology Co. Ltd. (Shanghai, China). Mass spectrometric detection was carried out on the Thermo UHPLC-Q Exactive HF-X Mass Spectrometer equipped with an electrospray ionization (ESI) source, operated in both positive and negative ion modes. Instrument parameters were adjusted for optimal performance: the ion source temperature was set at 425 °C; sheath gas flow rate at 50 arbitrary units; auxiliary gas flow rate at 13 arbitrary units; and the ion-spray voltage floating (ISVF) set to 3500 V (positive mode) and −3500 V (negative mode). Normalized collision energy was ramped (20-40-60 V) for MS/MS fragmentation. Data was acquisited in Data Dependent Acquisition (DDA) mode with a mass scan range of 70–1050 m/z. The resolution for Full MS was 60,000, while MS/MS scans were performed at a resolution of 7500. Quality control samples, generated by pooling equal volumes of all fecal samples, were run at regular intervals to check system stability and reproducibility of the analyses. The raw LC-MS/MS data were processed using Progenesis QI software (Waters Corporation, Milford, USA) for baseline filtering, peak identification, alignment, integration, and retention time correction. Bile acids were identified by matching the accurate molecular mass (mass error < 10 ppm) and MS/MS fragmentation patterns against the Human Metabolome Database (HMDB), METLIN, and a self-built standard library from Majorbio Bio-Pharm Technology. Quantification was performed based on the relative peak area of each identified bile acid, normalized to the internal standard and sample weight.
Clinical characteristics statistical analyses
As prespecified, the results were analyzed with both intention-to-treat and per-protocol principles. Descriptive statistics are presented as mean ± SD for continuous variables or as numbers and percentages for qualitative variables. Baseline characteristics were compared using t-test, Wilcoxon rank-sum test, or Fisher’s exact test for normal, skewed continuous variables and categorical variables when appropriate. Generalized estimating equation (GEE) was used to assess the effect of time, group, and time×group on outcomes, adjusted for age, sex, and BMI. Within-arm differences were also analyzed using GEE adjusted for age, sex, and baseline measurement. Analysis of covariance was used to evaluate between-arm differences at each visit with adjustment for age, sex, and BMI.
16S rRNA gene and BA statistical analyses
The Wilcoxon rank-sum test was used to compare the α-diversity indices at the species level of gut microbial community richness (Ace and Chao1 estimates) and community diversity (Shannon’s index) between groups. Differences in relative genus or species abundance between the two groups were analyzed using the Wilcoxon rank-sum test at baseline and 12 weeks. The LEfSe was performed to analyze variance at the taxonomic level of phylum, class, order, family, genus, and species to identify between-group difference in taxa (LDA > 2 and P < 0.05).
The optimal BA panels for predicting lipid improvement were selected impartially with a random forest algorithm by integrating it into a repetitive double cross-validation framework33. Subsequently, BA scores predicting the improvement in LDL-C (BAS LDL-C) and TC (BAS TC) were generated using ridge regression followed by tenfold cross-validation. The association between blood lipid alterations and BAS LDL-C or BAS TC was assessed using a general linear model, with adjustment for age and sex.
Three ML algorithms (lightGBM, logistic regression, xgboost) were tested to select optimal models predicting responders (reduction >5% of their baseline values in LDL-C or TC) upon the LP intervention based on the baseline gut microbiota. After excluding variables with variance 0, 85% of the samples were randomly selected as the training set. The training set was used to construct the model through fivefold cross-validation. The AUCs was obtained based on the cross-validation error of the 10,000-times self-help method (bootstrap), and was used to determine the optimal ML model. Partial Spearman’s correlation analysis was used to assess the correlations between BA, microbiota, genetic backgrounds and lipid profiles with adjustment of age and sex, and was visualized using Cytoscape version 3.10.2.
Genotyping and GRS calculation
The participants’ DNA was extracted from the buffy coat fraction of centrifuged blood using the QIAamp DNA Mini Kit (Qiagen, Valencia, CA). Genotyping was conducted using a customized Illumina Infinium® Asian Screening Array (ASA-CHIA) (Illumina, San Diego, CA, USA; Guoke Biotechnology, Beijing, China). The chip is designed with more than 738,000 variants for medical research and related disease studies in East Asian populations. We excluded the SNPs with a call rate <90%, minor allele frequency (MAF) < 1%, or not in Hardy-Weinberg equilibrium (P < 10−6) using PLINK(v1.07)34. The SNPs were first selected from those significantly associated with lipid profiles in a genome-wide association analysis conducted in an East Asian population (Table S10). To identify SNPs significantly interacting with lipid improvement, we examined genotype, intervention, and genotype–intervention interactions as independent predictors of changes in lipid traits, adjusted for age, and sex in the generalized linear models. Those showing a significant P for interaction were used to calculate the GRS with the following equation:
Where β is the effect size of each SNP on plasma cholesterol, n is the number of SNPs for each GRS, and sum of the β is the sum of effect sizes of SNPs included in GRSs35. A higher GRS indicates a higher genetic predisposition to poorer lipid response after the trial. A generalized linear model was fitted to examine the interaction of the GRS with the intervention on lipid changes adjusting for age and sex.
Estimation of the variance in blood lipid reduction explained by GRS and gut microbiota
The MRS was derived from the Shapley additive explanation (SHAP) values of the top ten microbial species from the optimal ML algorithms36. The SHAP assigned each variable an importance value predicted by the ML algorithm for quantifying the prediction results. The SHAP value indicates the degree to which the variable contributes to the model predictions. SHAP values were between −1 and 1, with a value greater than 0 indicating an increasing effect on lipid levels. The detailed calculation formula is as follows:
If SHAPij \(\le 0\), then sij is assigned to 0, otherwise 1. In the formula, i indicates the ith individual and MRSi specifies the MRS of the ith individual, j indicates the jth gut microbiota, and sij refers to the MRS of the jth gut microbiota in the ith individual. To assess the proportion of lipid improvement explained by genetic background (Rg, based on GRS) and initial gut microbiota (Rm, based on MRS)37, the following three linear regression models were developed:
Blood lipid change explained by Rg (Vg) was calculated by using the below formula:
Blood lipid change explained by Rm (Vm) was calculated by using the below formula:
Statistical analyses were performed using the R (version 4.3.1) or Python (version 3.12.3). Two-sided P-value or Q-value < 0.05 were considered significant. FDR-adjusted P-value (Q-value) was checked in post-hoc analyses when appropriate.
Data availability
Data will be made available on request.
References
Sun, L. et al. Causal Associations of Blood Lipids with Risk of Ischemic Stroke and Intracerebral Hemorrhage in Chinese Adults. Nat. Med 25, 569–574 (2019).
Vaduganathan, M., Mensah, G. A., Turco, J. V., Fuster, V. & Roth, G. A. The Global Burden of Cardiovascular Diseases and Risk: A Compass for Future Health. J. Am. Coll. Cardiol. 80, 2361–2371 (2022).
Papadaki, A., Nolen-Doerr, E. & Mantzoros, C. S. The Effect of the Mediterranean Diet on Metabolic Health: A Systematic Review and Meta-Analysis of Controlled Trials in Adults. Nutrients 12, 3342 (2020).
Chiavaroli, L. et al. DASH Dietary Pattern and Cardiometabolic Outcomes: An Umbrella Review of Systematic Reviews and Meta-Analyses. Nutrients 11, 338 (2019).
Ettinger, G., MacDonald, K., Reid, G. & Burton, J. P. The Influence of the Human Microbiome and Probiotics on Cardiovascular Health. Gut Microbes 5, 719–728 (2014).
DiRienzo, D. B. Effect of Probiotics on Biomarkers of Cardiovascular Disease: Implications for Heart-Healthy Diets. Nutr. Rev. 72, 18–29 (2014).
Guo, Z. et al. Influence of Consumption of Probiotics on the Plasma Lipid Profile: A Meta-Analysis of Randomised Controlled Trials. Nutr. Metab. Cardiovasc Dis. 21, 844–850 (2011).
Jia, B., Park, D., Hahn, Y. & Jeon, C. O. Metagenomic Analysis of the Human Microbiome Reveals the Association between the Abundance of Gut Bile Salt Hydrolases and Host Health. Gut Microbes 11, 1300–1313 (2020).
Gérard, P. Metabolism of Cholesterol and Bile Acids by the Gut Microbiota. Pathogens 3, 14–24 (2013).
Lye, H.-S., Rusul, G. & Liong, M.-T. Removal of Cholesterol by Lactobacilli via Incorporation and Conversion to Coprostanol. J. Dairy Sci. 93, 1383–1392 (2010).
Wu, Y., Zhang, Q., Ren, Y. & Ruan, Z. Effect of Probiotic Lactobacillus on Lipid Profile: A Systematic Review and Meta-Analysis of Randomized, Controlled Trials. PLoS One 12, e0178868 (2017).
Ya, C., J, K. Effect of Probiotics on Blood Lipid Concentrations: A Meta-Analysis of Randomized Controlled Trials. Medicine 94, https://doi.org/10.1097/MD.0000000000001714 (2015).
Wang, L. et al. The Effects of Probiotics on Total Cholesterol: A Meta-Analysis of Randomized Controlled Trials. Medicine 97, e9679 (2018).
Lu, Y. et al. Prevalence of Dyslipidemia and Availability of Lipid-Lowering Medications Among Primary Health Care Settings in China. JAMA Netw. Open 4, e2127573 (2021).
M, S. et al Efficacy and Safety of Lactobacillus Plantarum K50 on Lipids in Koreans With Obesity: A Randomized, Double-Blind Controlled Clinical Trial. Front. Endocrinol. 12, https://doi.org/10.3389/fendo.2021.790046 (2022).
Kerlikowsky, F. et al. Distinct Microbial Taxa Are Associated with LDL-Cholesterol Reduction after 12 Weeks of Lactobacillus Plantarum Intake in Mild Hypercholesterolemia: Results of a Randomized Controlled Study. Probiot. Antimicro. Prot. https://doi.org/10.1007/s12602-023-10191-2 (2023).
Pollin, T. I. et al. Genetic Modulation of Lipid Profiles Following Lifestyle Modification or Metformin Treatment: The Diabetes Prevention Program. PLoS Genet 8, e1002895 (2012).
San-Cristobal, R. et al. Mediterranean Diet Adherence and Genetic Background Roles within a Web-Based Nutritional Intervention: The Food4Me Study. Nutrients 9, 1107 (2017).
Fuentes, M. C., Lajo, T., Carrión, J. M. & Cuñé, J. Cholesterol-Lowering Efficacy of Lactobacillus Plantarum CECT 7527, 7528 and 7529 in Hypercholesterolaemic Adults. Br. J. Nutr. 109, 1866–1872 (2013).
Grundy, S. M. et al. 2018 AHA/ACC/AACVPR/AAPA/ABC/ACPM/ADA/AGS/APhA/ASPC/NLA/PCNA Guideline on the Management of Blood Cholesterol: A Report of the American College of Cardiology/American Heart Association Task Force on Clinical Practice Guidelines. Circulation 139, e1082–e1143 (2019).
Joint committee for guideline revision 2016 Chinese Guideline for the Management of Dyslipidemia in Adults. J. Geriatr. Cardiol. 15, 1–29 (2018).
Manson, J. E. et al. The Primary Prevention of Myocardial Infarction. N. Engl. J. Med 326, 1406–1416 (1992).
Briel, M. et al. Association between Change in High Density Lipoprotein Cholesterol and Cardiovascular Disease Morbidity and Mortality: Systematic Review and Meta-Regression Analysis. BMJ 338, b92 (2009).
Gao, Y. & Li, D. Screening of Lactic Acid Bacteria with Cholesterol-Lowering and Triglyceride-Lowering Activity in Vitro and Evaluation of Probiotic Function. Ann. Microbiol 68, 537–545 (2018).
Zhu, Y. et al. Beneficial Effects of Enterococcus Faecalis in Hypercholesterolemic Mice on Cholesterol Transportation and Gut Microbiota. Appl Microbiol Biotechnol. 103, 3181–3191 (2019).
Le Roy, T. et al. The Intestinal Microbiota Regulates Host Cholesterol Homeostasis. BMC Biol. 17, 94 (2019).
Padro, T. et al. Lactiplantibacillus Plantarum Strains KABP011, KABP012, and KABP013 Modulate Bile Acids and Cholesterol Metabolism in Humans. Cardiovasc. Res. 120, 708–722 (2024).
Zhang, Y. et al. Comparison of Gut Microbiota in Male MAFLD Patients with Varying Liver Stiffness. Front Cell Infect. Microbiol 12, 873048 (2022).
Chu, C. et al. Bifidobacterium Longum CCFM1077 Attenuates Hyperlipidemia by Modulating the Gut Microbiota Composition and Fecal Metabolites: A Randomized, Double-Blind, Placebo-Controlled Clinical Trial. Engineering 28, 193–205 (2023).
Al-Sheraji, S. H. et al. Hypocholesterolaemic Effect of Yoghurt Containing Bifidobacterium Pseudocatenulatum G4 or Bifidobacterium Longum BB536. Food Chem. 135, 356–361 (2012).
Fuentes, M. C., Lajo, T., Carrión, J. M. & Cuñé, J. A Randomized Clinical Trial Evaluating a Proprietary Mixture of Lactobacillus Plantarum Strains for Lowering Cholesterol. Mediterr. J. Nutr. Metab. 9, 125–135 (2016).
Xie, M. et al. Metabolic Responses and Their Correlations with Phytochelatins in Amaranthus Hypochondriacus under Cadmium Stress. Environ. Pollut. 252, 1791–1800 (2019).
Shi, L., Westerhuis, J. A., Rosén, J., Landberg, R. & Brunius, C. Variable Selection and Validation in Multivariate Modelling. Bioinformatics 35, 972–980 (2019).
Purcell, S. et al. PLINK: A Tool Set for Whole-Genome Association and Population-Based Linkage Analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Huo, S. et al. Genetic Susceptibility, Dietary Cholesterol Intake, and Plasma Cholesterol Levels in a Chinese Population. J. Lipid Res. 61, 1504–1511 (2020).
Zeng, F. et al. Gut Microbiome Features and Metabolites in Non-Alcoholic Fatty Liver Disease among Community-Dwelling Middle-Aged and Older Adults. BMC Med 22, 104 (2024).
Wu, C. et al. Obesity-Enriched Gut Microbe Degrades Myo-Inositol and Promotes Lipid Absorption. Cell Host Microbe 32, 1301–1314.e9 (2024).
Acknowledgements
We thank all the participants for their involvement in this study, and all the members of Xin Liu Laboratory for assisting with the intervention procedure and discussing the manuscript. This study was supported by Xi’an Jiaotong University (DW080038K0000004), BYHEALTH Nutrition and Health Research Foundation (TY202104001), National Natural Science Foundation of China (82173504, 82011530197), and the Science and Technology Resources Open Sharing Platform of the Shaanxi Province (2023-CX-PT-47).
Author information
Authors and Affiliations
Contributions
Conceptualization, X.L., X.Z. and Q.W.; methodology, X.L. and X.Z.; software, X.L., B.M.; validation, C.H.; formal analysis, G.M.; investigation, X.L., Q.W., G.M., Y.L., C.H., J.L., J.X., K.X., X.F., J.H., K.J., Y.H., W.L., N.Y.; resources, X.L., X.Z. and Q.W.; data curation, G.M., Y.L., C.H., J.L., J.X., K.X., X.F., J.H., K.J., Y.H., W.L., N.Y., B.Z.; writing—original draft preparation, G.M.; writing—review and editing, Y.L. M.A.; visualization, G.M.; supervision, X.L.; project administration, X.L.; funding acquisition, X.L. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ma, G., Li, Y., He, C. et al. Effects of Lactiplantibacillus plantarum on moderate dyslipidemia before medication involving gut microbiota and host genetics. npj Sci Food 10, 95 (2026). https://doi.org/10.1038/s41538-026-00749-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41538-026-00749-z