Abstract
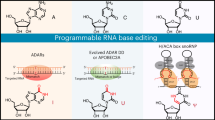
Base and prime editing technologies precisely install defined nucleotide edits in both dividing and non-dividing cells, offering potential for correcting pathogenic mutations directly in organisms. However, to fully leverage their therapeutic potential, accurately measuring editing rates with high spatial resolution is crucial. Here we use imaging-based in situ sequencing (ISS) to map base and prime editing events within native tissues. We establish and validate this technology in mouse brains treated with intein-split adenine base editors or prime editors delivered via adeno-associated viral vectors. We next apply ISS in the liver of mice and macaques treated with adenine base editors encoded on lipid nanoparticle-encapsulated mRNA and guide RNA (RNA-LNP). Effective editing was observed across all metabolic zones of liver lobules. Moreover, in experiments where repeated doses of RNA-LNP are administered, the initial dose does not affect the editing efficiency and distribution of the subsequent dose. Our results demonstrate how ISS can visualize gene editing events in vivo and suggest that base editor delivery using RNA-LNP could be used to address a wide spectrum of metabolic liver diseases.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All oligonucleotides and padlock probes used in the study are provided in Supplementary Table. HTS data have been deposited at the Sequence Read Archive (PRJNA1102160)49. Source data are provided with this paper.
Code availability
The CellProfiler pipeline and R script is available via GitHub at https://github.com/sharan-j/genome_editing_ISS (ref. 50).
References
Komor, A. C., Kim, Y. B., Packer, M. S., Zuris, J. A. & Liu, D. R. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature 533, 420–424 (2016).
Gaudelli, N. M. et al. Programmable base editing of A·T to G·C in genomic DNA without DNA cleavage. Nature 551, 464–471 (2017).
Anzalone, A. V. et al. Search-and-replace genome editing without double-strand breaks or donor DNA. Nature 576, 149–157 (2019).
Villiger, L. et al. Treatment of a metabolic liver disease by in vivo genome base editing in adult mice. Nat. Med. 24, 1519–1525 (2018).
Villiger, L. et al. In vivo cytidine base editing of hepatocytes without detectable off-target mutations in RNA and DNA. Nat. Biomed. Eng. 5, 179–189 (2021).
Rothgangl, T. et al. In vivo adenine base editing of PCSK9 in macaques reduces LDL cholesterol levels. Nat. Biotechnol. 39, 949–957 (2021).
Musunuru, K. et al. In vivo CRISPR base editing of PCSK9 durably lowers cholesterol in primates. Nature 593, 429–434 (2021).
Koblan, L. W. et al. In vivo base editing rescues Hutchinson–Gilford progeria syndrome in mice. Nature 589, 608–614 (2021).
Arbab, M. et al. Base editing rescue of spinal muscular atrophy in cells and in mice. Science 380, eadg6518 (2023).
Chemello, F. et al. Precise correction of Duchenne muscular dystrophy exon deletion mutations by base and prime editing. Sci. Adv. 7, eabg4910 (2021).
Yeh, W. H. et al. In vivo base editing restores sensory transduction and transiently improves auditory function in a mouse model of recessive deafness. Sci. Transl. Med. 12, eaay9101 (2020).
Böck, D. et al. In vivo prime editing of a metabolic liver disease in mice. Sci. Transl. Med. 14, eabl9238 (2022).
Davis, J. R. et al. Efficient prime editing in mouse brain, liver and heart with dual AAVs. Nat. Biotechnol. 42, 253–264 (2024).
Fu, Y. et al. In vivo prime editing rescues photoreceptor degeneration in nonsense mutant retinitis pigmentosa. Nat. Commun. 16, 2394 (2025).
Kulkarni, J. A. et al. The current landscape of nucleic acid therapeutics. Nat. Nanotechnol. 16, 630–643 (2021).
Espina, V. et al. Laser-capture microdissection. Nat. Protoc. 1, 586–603 (2006).
Levy, J. M. et al. Cytosine and adenine base editing of the brain, liver, retina, heart and skeletal muscle of mice via adeno-associated viruses. Nat. Biomed. Eng. https://doi.org/10.1038/s41551-019-0501-5 (2020).
Ke, R. et al. In situ sequencing for RNA analysis in preserved tissue and cells. Nat. Methods 10, 857–860 (2013).
Mignardi, M. et al. Oligonucleotide gap-fill ligation for mutation detection and sequencing in situ. Nucleic Acids Res. 43, e151(2015).
Lomakin, A. et al. Spatial genomics maps the structure, nature and evolution of cancer clones. Nature 611, 594–602 (2022).
Chen, X., Sun, Y. C., Church, G. M., Lee, J. H. & Zador, A. M. Efficient in situ barcode sequencing using padlock probe-based BaristaSeq. Nucleic Acids Res. 46, 22 (2018).
Feldman, D. et al. Optical pooled screens in human cells. Cell 179, 787–799 (2019).
Grundberg, I. et al. In situ mutation detection and visualization of intratumor heterogeneity for cancer research and diagnostics. Oncotarget 4, 2407–2418 (2013).
Larsson, C., Grundberg, I., Söderberg, O. & Nilsson, M. In situ detection and genotyping of individual mRNA molecules. Nat. Methods 7, 395–397 (2010).
Richter, M. F. et al. Phage-assisted evolution of an adenine base editor with improved Cas domain compatibility and activity. Nat. Biotechnol. 38, 883–891 (2020).
Maïno, N. et al. A microfluidic platform towards automated multiplexed in situ sequencing. Sci. Rep. 9, 3542 (2019).
Stirling, D. R. et al. CellProfiler 4: improvements in speed, utility and usability. BMC Bioinform. 22, 433 (2021).
Hanna, R. E. et al. Massively parallel assessment of human variants with base editor screens. Cell 184, 1064–1080 (2021).
Böck, D. et al. Prime editing of the β1 adrenoceptor in the brain reprograms mouse behavior. Preprint at bioRxiv https://doi.org/10.1101/2023.05.19.541410 (2023).
Ma, R., Martínez-Ramírez, A. S., Borders, T. L., Gao, F. & Sosa-Pineda, B. Metabolic and non-metabolic liver zonation is established non-synchronously and requires sinusoidal Wnts. Elife 9, e46206 (2020).
Ben-Moshe, S. et al. Spatial sorting enables comprehensive characterization of liver zonation. Nat. Metab. 1, 899–911 (2019).
Zincarelli, C., Soltys, S., Rengo, G. & Rabinowitz, J. E. Analysis of AAV serotypes 1-9 mediated gene expression and tropism in mice after systemic injection. Mol. Ther. 16, 1073–1080 (2008).
Karikó, K., Buckstein, M., Ni, H. & Weissman, D. Suppression of RNA recognition by Toll-like receptors: the impact of nucleoside modification and the evolutionary origin of RNA. Immunity 23, 165–175 (2005).
Baiersdörfer, M. et al. A facile method for the removal of dsRNA contaminant from in vitro-transcribed mRNA. Mol. Ther. Nucleic Acids 15, 26–35 (2019).
Ryan, D. E. et al. Phosphonoacetate modifications enhance the stability and editing yields of guide RNAs for Cas9 editors. Biochemistry 62, 3512–3520 (2023).
Kudo, T. et al. Multiplexed, image-based pooled screens in primary cells and tissues with PerturbView. Nat. Biotechnol. https://doi.org/10.1038/s41587-024-02391-0 (2024).
Gu, J. et al. Mapping multimodal phenotypes to perturbations in cells and tissue with CRISPRmap. Nat. Biotechnol. https://doi.org/10.1038/s41587-024-02386-x (2024).
Duff, C. & Baruteau, J. Modelling urea cycle disorders using iPSCs. npj Regen. Med. 7, 56 (2022).
Zabulica, M. et al. Correction of a urea cycle defect after ex vivo gene editing of human hepatocytes. Mol. Ther. 29, 1903–1917 (2021).
Wang, B., Zhao, L., Fish, M., Logan, C. Y. & Nusse, R. Self-renewing diploid Axin2+ cells fuel homeostatic renewal of the liver. Nature 524, 180–185 (2015).
He, L. et al. Proliferation tracing reveals regional hepatocyte generation in liver homeostasis and repair. Science 371, eabc4346 (2021).
Wei, Y. et al. Liver homeostasis is maintained by midlobular zone 2 hepatocytes. Science 371, eabb1625 (2021).
Marquart, K. F. et al. Predicting base editing outcomes with an attention-based deep learning algorithm trained on high-throughput target library screens. Nat. Commun. 12, 5114 (2021).
Mathis, N. et al. Predicting prime editing efficiency and product purity by deep learning. Nat. Biotechnol. https://doi.org/10.1038/s41587-022-01613-7 (2023).
Grieger, J. C. & Samulski, R. J. Packaging capacity of adeno-associated virus serotypes: impact of larger genomes on infectivity and postentry steps. J. Virol. 79, 9933–9944 (2005).
Vadovics, M., Muramatsu, H., Sárközy, A. & Pardi, N. Production and evaluation of nucleoside-modified mRNA vaccines for infectious diseases. Methods Mol. Biol. 2786, 167–181 (2024).
Conway, A. et al. Non-viral delivery of zinc finger nuclease mRNA enables highly efficient in vivo genome editing of multiple therapeutic gene targets. Mol. Ther. 27, 866–877 (2019).
Gyllborg, D. et al. Hybridization-based in situ sequencing (HybISS) for spatially resolved transcriptomics in human and mouse brain tissue. Nucleic Acids Res. 48, E112 (2020).
Janjuha, S., Haenggi, T. & Schwank, G. Spatial profiling of base editing by in situ sequencing in mice and macaques. NCBI SRA https://www.ncbi.nlm.nih.gov/bioproject/PRJNA1102160 (2025).
Janjuha, S., Haenggi, T. & Schwank, G. Genome editing ISS. GitHub https://github.com/sharan-j/genome_editing_ISS (2025).
Acknowledgements
We thank the Functional Genomics Center Zurich for technical support and access to instruments at the University of Zurich; and members of the laboratory of G.S. and Acuitas for discussions and comments on the paper. This work was supported by the European Molecular Biology Organization long-term fellowship EMBO ALTF 873-2019 (to S.J.), a Novartis Young Investigator Grant (to S.J.), Swiss National Science Foundation grant number 310030_185293 (to G.S.), a European Research Council Consolidator Grant (to G.S.) and the Helmut Horten Foundation (to G.S.). The Pardi Laboratory was supported by the National Institute of Allergy and Infectious Disease of the US National Institutes of Health under award numbers R01AI153064, P01AI158571 and P01AI172531.
Author information
Authors and Affiliations
Contributions
G.S., S.J., S.C.S. and Y.K.T. conceived the study. S.J. and T.H. set up and performed all ISS experiments and analysis. T.R., L.K., M.W. and D.B. performed in vivo injections into mice. N.M. and K.M. performed experiments on GFP reporter cells. W.J.M. performed RNA-LNP formulations; H.M., M.V. and N.P. performed the mRNA production. T.C.C., S.C.S., Y.K.T., S.J., T.H. and G.S. analysed data from the macaque experiments. G.S. and S.J. wrote and edited the paper with input from all co-authors. All authors read and approved the final version of the paper.
Corresponding authors
Ethics declarations
Competing interests
T.C.C., W.J.M., S.C.S. and Y.K.T. are employees of Acuitas Therapeutics. G.S. and K.M. are co-founders of Nerai Bio. G.S. is a scientific adviser to Prime Medicine. The other authors declare no competing interests.
Peer review
Peer review information
Nature Biomedical Engineering thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Detection of prime editing by ISS in cultured cells.
(a) Schematic of the experimental setup to detect prime editing in GFP reporter cells treated with PE2. (b) Quantification of the percentage of edited reads using NGS. Each datapoint represents an independent well of separately transfected cells. (c) Image shows edited (green) and unedited GFP reporter cells (left panel), and RCPs (right panel). Scale bar: 10 um. (d) Quantification of the fraction of GFP positive cells and edited reads detected using ISS. Each point represents an independent well (replicate 1, n = 805 cells; replicate 2 = 704 cells). (e) Quantification of the concordance between GFP expression and ISS in prime edited GFP reporter cells.
Extended Data Fig. 2 Detection of base editing by ISS in the mouse brain after intracranial AAV injection.
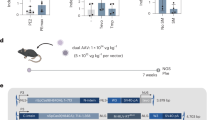
(a) (Left) Schematic of the experimental set-up. 2 μl AAV PHP.eB with a concentration of 1×1013 vg/mL encoding for the ActbV30 targeting ABEmax were injected into the hypothalamus of adult mice (Right). Image from the Allen Brain Atlas. Depicting areas illustrate the areas analyzed by NGS (yellow) or ISS (red). (b) Representative brain tissue region cut by laser microdissection from mice injected in hypothalamus. (c) Quantification of editing efficiencies in the hypothalamus of treated mice detected by NGS and ISS. Datapoints represent different mice. n = 3922 and 1703 RCPs. (d) Image shows RCPs in the hypothalamus of mice treated with ABE targeting ActbV30. Insets show the RCPs as single channel images. Scale bar: 10 um. (e) Quantification of editing rates in distinct sub-regions of the hypothalamus. Datapoints represent separate areas in the hypothalamus from at least two mice. Left: n = 521 and 659 RCPs; mid: n = 1623 and 807 RCPs; right: n = 1778 and 237 RCPs. Bars represent mean ± s.d. a, Created with BioRender.com.
Extended Data Fig. 3 Biodistribution of AAV9 and RNA-LNP in mice liver.
(a) (Top) Schematic of the experimental set-up. Adult mice were injected via the tail-vein with either AAV9 TagRFP-expressing vector or LNP-mCherry. (Bottom) Representative images of the liver lobule zonation from treated mice. Pericentral cells are counterstained with Glutamine Synthetase (GS) (magenta) antibody and periportal cells are counterstained with ECAD (green) antibody. Scale bar: 100 um. (b) Quantification of average intensities in arbitrary units of either TagRFP or mCherry in the three liver zones. Datapoints represent independently treated animals. a, Created with BioRender.com.
Extended Data Fig. 4 Analysis of editing in extrahepatic tissues liver and toxicity markers after systemic RNA-LNP delivery into macaques.
Quantification of (a) liver toxicity markers and (b) inflammatory biomarkers or cytokines before and during treatment. Bars represent mean ± s.d. Empty datapoints represent animals from strategy 1 and filled datapoints represent animals from strategy 2. Circle and square datapoints represent values of independently treated animals (2 animals per strategy). Vertical dotted lines represent the time point of I.V. infusion. (c-d) Quantification of editing in extrahepatic organs in animals from strategy 1 and 2. Each datapoint represents the values of independently treated animals. A control group was not included due to the unavailability of liver tissue from untreated macaques.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–10.
Supplementary Table 1 (download XLSX )
Oligonucleotide and padlock probe sequences used in the paper, and percentage editing rates from NGS and ISS.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Janjuha, S., Haenggi, T., Chamberlain, T.C. et al. Spatial profiling of gene editing by in situ sequencing in mice and macaques. Nat. Biomed. Eng (2025). https://doi.org/10.1038/s41551-025-01512-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41551-025-01512-7