Abstract
Human neural organoids are essential platforms for fundamental and applied research due partly to their complex, three-dimensional neuronal circuit geometries. Standard and recently developed neural interface technologies have shortcomings in their ability to electrically characterize and control neural activity in these systems, owing to their limited accessibility to neuron populations and microelectrode densities. Here we report a shape-matched, soft, three-dimensional mesoscale framework with nearly full surface coverage to neural organoids that supports high channel count interfaces for precision electrophysiology and programmed electrical stimulation. The neural interface is designed via inverse modelling techniques and self-assembles three-dimensionally around the organoids. Three-dimensional reconstruction of neural activities allows high-resolution spatial electrophysiology to reveal network-level characteristics in neural organoids. The porous framework offers options for simultaneous fluorescence imaging, localized optogenetic neuromodulation, longitudinal monitoring, pharmacological evaluations and modelling of neural disease phenotypes, demonstrating broad applicability for studies of human-derived cortical and spinal organoids.
Similar content being viewed by others
Main
Neural organoids feature elaborate neural interconnections and three-dimensional (3D) cytoarchitectures that recapitulate certain essential characteristics of functional brains and nervous systems not present in traditional two-dimensional (2D) cell cultures1,2,3. When produced from human pluripotent stem cells, these constructs also avoid effects of phylogenetic differences between human and the most widely used animal models. As a result, human-derived neural organoids are of increasing interest for both fundamental research and translational studies. For instance, the collective neural activity in cortical and spinal organoids provides versatile platforms for investigating neurodevelopment4, circuit formation5,6 and neural disease modelling7,8,9, with additional potential for use in precision medicine based on patient-specific genetics10,11. Furthermore, the neuronal networks and human brain-like cellular composition in these systems offer opportunities in the development of living platforms for computation and artificial intelligence12.
These and other use cases demand neural interfaces that can support high accessibility to neural populations, high-resolution electrophysiological measurements and spatiotemporal stimulation protocols. An essential goal is to characterize and control network-level activity across most or all of the overall neuron population, at or near the cellular scale. Recent progress in organoid interface technology overcomes fundamental geometrical constraints associated with the 2D designs of traditional multielectrode array (MEA)13 methods and complementary metal–oxide–semiconductor-based circuits12,14. Specifically, the latest systems support 3D coverage by use of stretchable mesh constructs and/or deformable electronics15,16,17, multilayer MEAs18, self-folding structures19,20 and buckled frameworks21, each of which integrates arrays of electrodes that conform to the 3D features of organoids. Shortcomings of these methods are in their ability to access only a relatively small fraction of the overall neuronal population owing to limitations in the extent of areal/volumetric coverage, the numbers of electrodes, the electrode densities and/or addressability.
This work presents a method for high-accessibility and high-resolution spatial neural electrophysiology of organoids (Fig. 1a). A combined collection of strategies in 3D assembly techniques, computational design strategies22 and organoid growth strategies qualitatively advances those published previously21. These resulting interfaces feature complex and continuously curved supports that conform to targeted 3D geometries of organoids with nearly complete surface coverage, therefore allowing one to distribute large numbers of microelectrodes in dense arrays. Key capabilities are in high-resolution electrophysiology and versatile options in patterned electrical stimulation, as well as 3D reconstruction of neural activities for advanced spatial and network-level analysis. User-definable 3D geometries allow modulation of organoids into artificial shapes. These functionalities establish this method as a unique vehicle for measurements and manipulations of neural organoids in fundamental neuroscience investigations, biomedical studies and neural intelligence research.
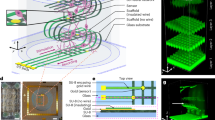
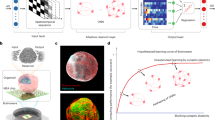
a, A schematic illustration of a representative system. An engineered mechanical buckling method applied to a 2D precursor in a microlattice geometry leads to the formation of a desired 3D shape, guided by inverse computational modelling. The microlattice offers large areas to support dense microelectrode arrays but with sufficient porosity to allow diffusive transport for natural metabolic processes. b, A schematic diagram of the design strategy to simultaneously realize targeted surface geometries and 3D curvatures matching those of an organoid. c, A micro-CT image of an interface that offers 91% surface coverage across the spherical surface of an organoid with 240 independently addressable electrodes for measurement and/or simulation. d, A fluorescence image of the microelectrode array interfaced with cells in an organoid inserted into and grown within the 3D microlattice framework. e, Schematic illustration and image (inset) of an organoid cultured within this framework with external connecting circuits for neural recording and stimulation. f, A collection of neural spike waveforms from hSOs and hCOs. Data are presented as mean values ± s.d. of all spikes within each cluster. Scale bars, 1 ms (horizontal) and 20 μV (vertical). g, Comparison of 3D neural interface in this work with other organoid interface technology, plotted as a function of surface coverage and number of electrodes. Data points are estimated from refs. 12,13,14,15,16,17,18,19,20,21,46.
Realizing complex 3D biological surfaces from 2D-fabricated functional devices is a long-standing engineering challenge due to the coupling between geometry design and the 2D-to-3D transformation processes. Mechanically guided 3D assembly and microlattice design strategies establish the fundamental basis for forming the advanced interfaces introduced in this study. The former transforms 2D precursors (Supplementary Fig. 1) into 3D frameworks through processes in controlled mechanical buckling21,23,24 (Fig. 1a and Supplementary Video 1). The latter enables high coverage across the surfaces of targeted organoids by providing means for continuous, independent control of the geometries and stiffness distributions across the 2D precursors. The result deterministically yields versatile user-defined 3D geometries through inverse computational design, shown in Fig. 1a as an example of nearly complete coverage across the surface of a spherical organoid (Fig. 1a). Strategic choices of microlattice geometries also provide surface areas sufficiently large to accommodate large, dense collections of microelectrodes but with adequate porosity to enable minimally restricted diffusive transport in support of natural metabolic processes. An additional strategy in organoid growth within these frameworks ensures intimate, low-impedance contacts in both spherical and complex 3D shapes defined by the geometry of the interface. Detailed biological evaluations with both cortical and spinal organoids derived from human pluripotent stem cells, referred to as hCOs and hSOs respectively, validate these features and demonstrate feasibility for high-density neural recording, surface activity mapping, 3D correlation analysis and location-specific neural stimulation. These options appear in longitudinal measurements of organoid activity and real-time evaluation of neural disease phenotype models, along with responses to pharmacological perturbations, localized optogenetic stimulation and manipulation of neural circuits.
Results
Design strategy for full surface coverage and shape-conformal interface
A key advance is in the ability to achieve diverse 3D geometries in these organoid interfaces, each with capabilities for nearly full area coverage through an inverse design strategy that uses component elements with customized microlattice layouts22 (Fig. 1a and Extended Data Fig. 1). Figure 1b illustrates this method for the case of a spherical shape. Discretization of the organoid surface into n (for example, n = 8 in Fig. 1b) pieces of interconnecting subsurfaces with 2D shapes that enable continuous surface coverage forms a projection in the geometry space, thereby determining a beam width distribution wg (x). An adaptive genetic algorithm (AGA) based on Euler–Bernoulli beam theory (Extended Data Fig. 1) allows determination of an optimal bending stiffness distribution K (x) of the beam that yields a buckled configuration z (x) with the highest degree of match with the 3D curvature of the surface of a targeted organoid. For a beam made of solid material with uniform thickness distribution, the stiffness distribution K (x) is proportional to the width distribution ws (x), that is, K (x) ∝ ws (x). Generally, wg (x) ≠ ws (x), such that the shape matching at full surface coverage is nearly impossible. A solution to this problem involves patterning these materials into well-defined microlattices in the form of triangle-shaped micropores with spatially varying sizes, to define an engineered distribution of porosity φ (x), and thus the effective modulus E (x) ∝ (1 − φ (x)). A microlattice beam with porosity distribution of [φ (x) = 1 − ws (x)/wg (x)] allows both full surface coverage and targeted 3D curvatures. Finite element analysis (FEA) validates the mechanical feasibility of this approach, demonstrating a maximum strain <2.2% throughout the mechanically guided assembly, below the critical strain of typical engineering polymers (Extended Data Fig. 2a–e). Optimizing the device thickness and adding chamfers to relieve stress concentrations ensures low strain levels, even for organoids with dimensions down to a few hundred micrometres (Extended Data Fig. 2f,g).
The microlattice design enables free distribution of microelectrodes at lattice nodes with interconnects along triangle edges, therefore transforming the polymer scaffold into a 3D neural interface. Figure 1c and Supplementary Video 2 show micro-computed tomography (micro-CT) imaging of assembled interfaces. This structure covers 91% of the surface of a millimetre-scale spherical organoid with 240 individually addressable microelectrodes (diameters ≤30 µm) and reference electrodes (Supplementary Fig. 2 and Supplementary Note 1), distributed uniformly to offer an interface with spatial resolution beyond 100 µm for a millimetre-sized neural organoid (Fig. 1c,d). Coatings of platinum (Pt) black lead to microelectrodes with low electrochemical impedances (~10 kΩ) at 1 kHz (Extended Data Fig. 3a–c), high charge injection capacities (~200 µA) (Extended Data Fig. 3d–f) and excellent stability for long-term operation (Extended Data Fig. 3g,h) in electrical recording and/or stimulation for neural organoids (Fig. 1e–g).
High-density neural recording
Experiments with hCOs derived from human-induced pluripotent stem (hiPS) cells demonstrate the function of these neural interfaces25. Immunostaining of MAP2+ neurons, SOX+ neural progenitors and S100B+ glial cells from organoid sections on days 30, 60 and 120 upon differentiation reveals uniform cell distributions in these hCOs, with notable neural interconnections after day 60 (Fig. 2a,b and Extended Data Fig. 4a–c). The studies involve simultaneous electrophysiological recordings and calcium fluorescence imaging of hCOs after day 60 of differentiation. The latter is enabled by the use of transparent substrates of polydimethylsiloxane (PDMS) and polymers (polyimide (PI) or parylene-C) as scaffolds (Supplementary Fig. 3), with GCaMP6 to induce production of the green fluorescence protein (GFP). Figure 2c and Supplementary Video 3 present typical results of hCO (day 88). The accumulated firing rate across all channels from electrophysiological recording aligns with the brightness from calcium fluorescence imaging, with identical oscillatory patterns of neural activity.
a, Immunostaining of an hCO at day 60 upon differentiation reveals uniform cell distributions. b, Magnified image of the tissue, consisting of neurons, astrocytes and neural progenitors. c, Neural activity of an hCO detected by simultaneous recording of electrophysiology and imaging with calcium fluorescence. a.u., arbitrary units. d, A schematic illustration of a grow-in-place strategy that ensures intimate, gentle contacts between the neural interface and the organoid. e, Monitoring the growth curve of the hCO defines the targeted size of the 3D neural interface. Data are presented as mean values ± s.d. of six different hCOs. f, Electrophysiological recordings through various stages of the grow-in-place process reveal an increasing number of active channels. g–i, High-density neural recordings of an hCO using an interface with 240 individual channels. g, Schematic illustrations of eight constituent beams of the interface, each labelled with a specific colour. h, Neural potential measured at 240 electrode sites. i, Raster plots of the recorded neural activity reveal complex oscillatory rhythms.
Effective physical coupling, and associated low-impedance contacts, between the interface and the organoid are crucial for high-quality electrophysiological recordings. Large separations between microelectrodes and the organoid surface may lead to redirecting of neural currents through low-resistance pathways and hinder spike recording. The strategy to ensure contact involves two aspects: (1) an inversely designed 3D structure that matches the shape of the targeted organoid, and (2) a process of coculturing the organoid after insertion as a grow-in-place process to fill any remaining gaps (Fig. 2d). Tracking the evolution of a representative hCO reveals a fast growth period from day 0 to day 60, followed by slow growth. An interface with a slightly larger inner diameter closely matches this size, while minimizing the risk for mechanical compression during insertion (Fig. 2e). Subsequent monitoring of the grow-in-place process reveals a gradual filling of the remaining space within several days (Extended Data Fig. 4d,e and Supplementary Fig. 4). During this process, the porous nature of the interface allows diffusive transport needed to support natural cell proliferation, but with a boundary that prevents growth of cells through the pores. Data from immunostaining indicate a similar distribution of cell types between organoids integrated into the interface in this manner versus controls (Extended Data Fig. 4f). Simultaneous measurements indicate a gradually increasing number of channels in the interface that show spiking activity, consistent with a corresponding gradual increase in the contact (see Fig. 2f and Supplementary Fig. 5 for cases of hCO and hSO).
Figure 2g–i shows examples of recording from an hCO (day 87), collected across 240 individual channels, where the colours mark channels from different beams. Signal processing through filtering, peak detecting, unsupervised clustering and artefact removal (Extended Data Fig. 5) yields data that clearly capture the rhythmic dynamics of synchronous activities across most surface neurons in the hCO26. Variation in the activity levels across individual channels reflects heterogeneities in the surface activity across the organoid (Supplementary Fig. 6 and Supplementary Note 2). Approaches for evaluating spike waveforms and spatial variations in tissue–electrode coupling (Methods; Extended Data Fig. 5 and Supplementary Note 2) can contribute to bands of silent channels in the raster plots. The raster plot demonstrates repeating patterns of one strong wave followed by three weaker waves in this case (Fig. 2i). These highly synchronized, periodic yet unequally spaced neural events suggest organoid-level neuronal interconnections that resemble coded activity patterns of developing human brains27.
3D spatial electrophysiology
Complete and high-resolution spatial electrophysiology requires dense electrodes distributed across a 3D interface with maximal coverage. Assuming that each microelectrode has an effective detection distance of 100 µm (refs. 28,29), a circle-covering model determines the fraction of the interface surface that is detectable as a function of the number of microelectrodes with optimized locations (Supplementary Note 3). Calculation results indicate a minimum of 240 microelectrodes to realize a detectable fraction above 90% (Fig. 3a). Probing the activity of an hCO (day 90) with a selected portion of the microelectrodes experimentally validates the crucial role of channel number (Fig. 3b). A uniformly distributed array of 8 microelectrodes (3.4% detection range) fails to capture any oscillatory burst events, whereas the case of 32 microelectrodes (13.6% detection range) detects only local synchronous events in 5 out of 8 beams (Fig. 3b). By sharp contrast, the full array of 240 microelectrodes (92% detection range) reveals complete rhythmic oscillatory waves across the organoid (Figs. 2j and 3b).
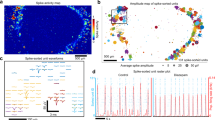
a, Modelling of detectable portions of the interface as a function of electrode numbers, by assuming a 100 µm detection distance, across an area coverage of 91%. b, Neural activity measured from an hCO based on data from a subset of 8, 32 and 240 electrodes in the array, to illustrate the numbers necessary to capture meaningful network-level neural processes. c, A schematic illustration of the spatial locations of 240 microelectrodes for 3D reconstruction of neural electrophysiology. d, A surface map of electrophysiological potential across a hCO at a particular moment. e, A surface map of the mean amplitude of neural spikes. f, A surface map of the mean firing rate. g, Recordings and corresponding characteristic waveforms of neural spikes of four different locations within one synchronous wave associated with an hCO. Waveform data are presented as mean values ± s.d. of all spikes within each cluster. Scale bars, 50 ms (horizontal) and 50 μV (vertical) for the potential plots, and 1 ms (horizontal) and 20 μV (vertical) for the waveforms. h, Correlation analysis among 389 recorded neural units of an hCO. i, A 3D connectivity map of correlated units of one example neural unit (left), and strong correlations among all channels (right) of the same hCO as in h. j, A schematic illustration of electrical stimulation and simultaneous calcium fluorescence imaging. k, Brightness analysis of the fluorescence images reveals neural activity of a hCO with no stimulation, with 60-s interval stimulation, with 30-s interval stimulation, and after stimulation. l, Fluorescence imaging of baseline and brightness difference maps with stimulation delivered from a single microelectrode, all microelectrodes of one beam, and all microelectrodes of two beams. Experiments were repeated in three independent hCOs with similar results. a.u., arbitrary units.
The system captures recordings with direct spatial labelling of neural signals without traditional channel-barcode indexing and section imaging. The result provides a practical and straightforward means for high-resolution spatial recording and electrical stimulation15,30 (Fig. 3c). Recording results associated with the spatial electrode positions form high-density 3D point cloud data. Neural activity reconstruction across the surface of the organoid uses a radial basis function for interpolation between these point cloud data. Figure 3d–f illustrates 3D reconstructed surface mapping of neural potential at a specific time, the mean spike amplitude and the mean firing rate for a typical hCO (day 87). The correlated patterns between amplitude and firing rate suggest persistent rather than transient strong and weak activity regions. The 3D interfaces also allow shape-matched recording from organoids spanning a broad range of ages, to reveal these characteristics of neural activity (Supplementary Figs. 7–10). Other examples show activities that are asynchronous, strongly synchronized oscillatory and localized (Extended Data Fig. 6).
Further investigations involve examination of specific locations (Fig. 3g) and 3D analysis of cross-correlations (Fig. 3h,i). Figure 3g shows recordings from four locations across the organoid surface, each with three channels, and plots the corresponding potentials within one synchronous wave alongside typical single unit spike waveforms. The data indicate varying initiation times for the same synchronous wave at different regions. Specifically, the two locations in the lower half initiate early, while the two in the upper half follow with an approximate 100-ms delay. This phenomenon suggests signal propagation through the interconnected neural network of the hCO with mappable temporal and spatial distribution of neural events (Supplementary Fig. 11 and Supplementary Note 4). The recorded waveforms reflect details of neural activities in specific locations in organoids and comparable to the reports in vivo (Supplementary Figs. 12 and 13 and Supplementary Note 5). Cross-correlation analysis examines these 3D interconnections. Figure 3h shows the correlation matrix among 389 recorded neural units from the same hCO as in Fig. 3g. The results illustrate notable correlations throughout the organoid and fine structures of stronger and weaker correlated unit pairs. Reconstruction visualizes these correlations in a 3D connectivity map (Fig. 3i). Notably, correlated neural units of a given neuron are distributed across the entire hCO rather than localized to neighbouring regions. Strong correlations connect individual channels throughout the hCO. Volumetric fluorescence imaging through two-photon microscopy (Supplementary Fig. 14 and Supplementary Note 6) and synaptic staining (Supplementary Fig. 15) validate the highly interconnected neuronal structures in this hCO. These highly interconnected, complex neural circuits that span 3D space resemble the organization of the developing brain, distinct from previous observations in 2D organoid slices26. The capability of the 3D interface for neural recording and analysis from large neuronal population with high spatiotemporal resolution offers important advantages over existing and conventional methods (Supplementary Fig. 16).
In addition to recording, the same interface allows spatially selective electrical stimulation, with simultaneous live calcium imaging (Fig. 3j, Supplementary Figs. 17 and 18 and Supplementary Note 7). Biphasic charge-balanced exocellular stimulation in a current-control mode delivers single stimuli with 200-µs-width periodic pulses (Extended Data Fig. 7a,b). Brightness analysis of calcium fluorescence indicates a threshold current of 3 µA to initiate neural activity (Extended Data Fig. 7c–h). Figure 3k depicts hCO (day 92) activity by calcium fluorescence before, during and after stimulation with 20-µA pulses across all channels, revealing phase-locked responses to stimulations at both 60-s and 30-s periods. Location-specific stimulation, using selected microelectrodes with simultaneous fluorescence monitoring (Fig. 3l, hCO at day 80), confirms aligned neural activities. This type of spatiotemporal electrostimulation is crucial for efforts in neural computing and organoid intelligence12.
Versatility in 3D structure and shape modulation
Figure 4 illustrates aspects of versatility in design, for various electrode numbers, electrode densities, interface porosities and surface coverages. Reducing the sizes of the micropores increases the number of microlattice nodes and corresponding microelectrodes, while preserving the desired 2D geometry and stiffness distribution (Fig. 4a). The design options include four factors: the local mesh density (ρ (x)), the width of a microlattice edge (b), the local porosity (φ (x)) and the bending stiffness (K (x)) of the beam. Fixing the cross-section area leads to the following dependence among these variables: ρ (x)b ∝ wg (x) (1 − φ (x)) ∝ K (x). The accessible range of bending stiffness is limited by the resolution of microfabrication and geometric constraint of overlapping between adjacent beam components. Introducing a minimum microlattice edge width bmin = 10 µm, set by practical microfabrication constraints, defines the overall design space (Supplementary Note 8). Figure 4b–d shows the results of FEA that follow this principle. Refinements of microlattice layouts towards these limits allow up to 1,800 microelectrodes with a spatial resolution of 40 µm, comparable to state-of-the-art complementary metal–oxide–semiconductor-based high-density 2D MEAs31,32 (Fig. 4b). This strategy enables a broadly adjustable range of average porosity up to 70% (Fig. 4c) and a surface coverage between 53% and 91% to concentrate the array of microelectrodes to regions of interest (Fig. 4d). The versatile design space for these 3D structures and electrode configurations also allows advanced application to assembloids and internal neural interconnections (Supplementary Figs. 19 and 20 and Supplementary Notes 9 and 10). In addition, as mentioned previously, the porosity facilitates efficient diffusion of nutrients and waste products essential for the health of the interfaced organoid. Quantitative analysis of a typical 240-channel spherical design demonstrates minimal restrictions on diffusive mass transport (Supplementary Note 11). Specifically, simulations that compare an interfaced organoid with the control case reveal similar steady-state concentration distributions of oxygen, carbon dioxide and glucose (Extended Data Fig. 8).
a, Schematic illustrations of the process of computationally guided refinement of microlattice layouts to maximize the density of microelectrodes while conserving the desired 2D geometry and stiffness distribution. b–d, Practically accessible levels of spatial resolution as a function of number of electrodes (b), porosity as a function of number of electrodes (c) and porosity as a function of surface coverage (d). e, Diverse geometries and corresponding micro-CT images of 3D interfaces. f, A schematic illustration of the grow-in-place process to realize organoids with shapes defined by these 3D interfaces. g, Fluorescence images of neural organoids with spherical, hexagonal prismatic and cubic shapes.
Other options include non-spherical shapes such as hexagonal prisms, cubes, ellipsoids, pyramids and even asymmetrical geometries, each with optimized distributions of micropores and microelectrode arrays. Reconstructed micro-CT imaging of some examples are shown in Fig. 4e. Growth within these constructs leads to releasable organoids with corresponding shapes (Fig. 4f). Figure 4g shows bottom-view fluorescence images of hCOs with spherical, hexagonal prismatic and cubic geometries, demonstrating the ability to fill sharp corners for precise geometric transformation from the interface to organoids, including those with high aspect ratios33,34. These capabilities are relevant to neural biomechanics research35 and engineering of assembloids with precisely tailored building blocks36,37.
Manipulation of activity and disease modelling
Other demonstrations of this technology include longitudinal monitoring, pharmacological studies, optogenetic neural modulations, manipulation of neural circuits and phenotype of neural disease models. Longitudinal recordings of spontaneous neural activity in organoids for 20 days reveal characteristic waveforms of each channel by single-unit activity analysis (Fig. 5a). Most single-unit spike waveforms remain stable throughout the recording period. Some channels exhibit emergent units over time, consistent with growth-induced gap filling at the neural interface. Figure 5b demonstrates the ability to monitor organoid responses to different pharmacological manipulations. Applying the potassium channel blocker 4-aminopyridine (4-AP) increases the average firing rate fourfold from 0.28 ± 0.06 Hz to 1.2 ± 0.2 Hz, while sodium channel blockers lidocaine and tetrodotoxin (TTX) lead to nearly complete elimination of the spontaneous activity.
a, Longitudinal monitoring of neural organoids shows stable tracking of single neural units with days after organoid insertion. Scale bars, 1 ms (horizontal) and 20 μV (vertical). b, Response of neural organoids, as recorded from n = 96 individual channels of integrated 3D interface, from exposure to 4-AP, lidocaine and TTX, measured as heatmaps of firing rates to highlight changes in activity before and after the application of these drugs. Two-sided t-tests on average firing rates indicate that the changes are statistically significant with ***P = 1.57 × 10−7 for 4-AP, ***P = 1.18 × 10−7 for lidocaine and **P = 2.01 × 10−3 for TTX. c, A schematic illustration of localized optogenetic stimulation with a position-tunable fibre laser. d, Monte Carlo simulations of the intensity distribution for local illumination of four microelectrodes. e, A raster plot of the responses of the exposed microelectrodes and several unexposed neighbouring microelectrodes, showing phase-locked responses only in the former. f, A schematic illustration of the effect of the elimination of neural transmitters by exposure to BTX. g, A raster plot of the responses at representative channels before and after exposure to BTX, consistent with the abolishment of oscillatory synchronous neural activities. h, A schematic illustration of the enhancement of neural transmission due to exposure to glutamate. i, Heatmaps of firing rates of an integrated organoid before and after dosing of glutamate show extreme neuronal excitatory effects. j, Longitudinal monitoring of n = 96 individual channels from an integrated 3D interface reveals an increase followed by a decrease of average firing rate within 5 days after exposure to glutamate. The data are presented as mean ± s.e.m. for b and j. Panels created in BioRender: f and h, Wu, M. (2025) https://BioRender.com/8wd9f21.
Additional studies illustrate the value of spatial mapping of electrophysiology in response to localized optogenetic modulation, as demonstrated with neural organoids transfected with lentivirus expressing channelrhodopsin-2. Illumination with a 470-nm laser delivered through an optical fibre allows precise control of the location, distance and angle of light delivery (Fig. 5c). Figure 5d represents Monte Carlo modelling results for a given position of the incident laser beam on the organoid surface, where the bright region covers four microelectrodes while leaving others unexposed (Supplementary Note 12). Optogenetic stimulation involves trains of pulses with durations of 200 ms separated by 10 s. The raster plot in Fig. 5e reveals simultaneous recordings from channels within this illuminated region and adjacent to it. The latter shows unperturbed, random spontaneous neural spikes throughout the test, whereas the former exhibit spikes phase locked to the optical stimulation.
The ability to probe oscillatory waves forms a promising means to study neural circuits. These oscillatory activities originate from network-level interconnections and neurotransmitter signalling7,27. Figure 5f summarizes investigations of neural circuit manipulation with botulinum toxin (BTX), a species that can inhibit release of excitatory neurotransmitters in the central nervous system, including glutamate and neuropeptides38. Application of BTX eliminates pronounced oscillatory rhythms, with complete abolishment of synchronous events (Fig. 5g). These results suggest a substantial elimination of chemical connections within synapses and corresponding reduction in network-level activities. These findings suggest a paradigm for organoid neural circuit research based on spatial electrophysiology, distinct from that for in vivo models39,40.
A final example involves studies of neural diseases41, where electrophysiology at the network level is crucial. Figure 5h presents results in monitoring a developing disease phenotype caused by an imbalance of the neurotransmitter, glutamate. Excessive glutamate transmission induces abnormal neuronal firing and extreme calcium flux into postsynaptic neurons, eventually resulting in excitotoxicity and nerve damage. This system has potential relevance to disease conditions such as multiple sclerosis, amyotrophic lateral sclerosis and Parkinson’s disease42. To mimic this process, longitudinal electrophysiological monitoring of hCOs tracks disease progression following glutamate application. Heatmaps of neural firing rates before and after application in Fig. 5i indicate increased neuronal excitation. Continuous tracking of the average firing rate over 6 days shows an initial increment of firing rate from 0.19 ± 0.06 Hz to 0.6 ± 0.2 Hz within 24 h, followed by a dramatic decline to 0.05 ± 0.03 Hz over subsequent days, consistent with glutamate excitotoxicity (Fig. 5j), which is a process associated with neurodegenerative disease and neurotrauma.
Conclusions
This work introduces a 3D mesoscale electronics technology that enables near-complete surface coverage, high-resolution electrophysiological measurements and electrical stimulation protocols for neural organoids. Unusual material architectures and design approaches, taken together with confined growth techniques, yield spatially continuous, low-impedance interfaces to complex organoid surfaces, with sufficient permeability to allow diffusive transport needed for natural metabolic processes. A computationally guided design space accommodates not only broad options in microelectrode number, distribution, resolution, porosity and coverage, but also engineering shape control in matured organoids. Hundreds of individually addressable microelectrodes enable single-unit neuronal recording of network-level activity in organoids, as well as direct 3D reconstruction for spatial electrophysiology. Such systems also offer capabilities in simultaneous fluorescence imaging, longitudinal coculturing and monitoring, and detection of neural responses in pharmacological evaluations and optogenetic neuromodulation, as additional features that are important for many studies. Examples in localized optogenetic activation, neural circuit manipulation and development of neural disease phenotypes suggest wide-ranging utility in investigations of neurodevelopment, neurodegeneration and neural disease progression. Uses in systems for neural organoid intelligence and in other classes and combinations of organoids represent additional areas of opportunity.
Methods
Inverse design strategies
Inverse design used an AGA to determine 2D microlattice patterns, defined by spatially varying triangular voids, for targeted 3D geometries (Extended Data Fig. 1). The optimization process used the Python framework DEAP, a common package for the deployment of AGAs. Discretization of the surface of the target geometry into a finite number of segments formed 2D shapes with width distributions (wg) for continuous surface coverage. These geometries served as the initial population for optimization of the stiffness distribution (ws) through AGA. For each generation, Python codes determined coordinate errors between simulated 3D shapes (ABAQUS) created by the buckling-guided assembly process and the targeted geometry. A set of 50 individuals per generation ensured an accurate determination of ws, defined by an error below a threshold value. The resulting geometries and stiffness distributions defined 2D precursor structures in microlattice layouts of triangular voids, for mechanically guided assembly into the desired 3D configurations.
3D assembly
Fabrication of the 2D precursors involved the following process steps (Supplementary Fig. 1). (1) Spin coating (3,000 rpm for 40 s) and baking (180 °C for 150 s) of poly(methyl methacrylate) on a substrate of glass or a silicon wafer formed a sacrificial release layer. (2) Spin coating (750 rpm for 60 s), baking (120 °C for 600 s, 150 °C for 300 s, 180 °C for 300 s) and vacuum baking (260 °C for 1 h) of a precursor to PI (PI-2545, HD Microsystems) defined the first layer of the polymer scaffold. (3) Spin coating (5,000 rpm for 40 s) and baking (110 °C for 60 s) formed a layer of negative photoresist (AZ nLOF 2035). (4) Photolithography with a maskless aligner (MLA150, Heidelberg Instruments) defined patterns for microelectrodes and interconnects, by 375-nm ultraviolet exposure with a dose of 140 mJ mm−2, followed by post baking (110 °C for 90 s) and developing (immersion in AZ 300MIF for 55 s). (5) Electron beam evaporation yielded a uniform 10-nm Cr adhesion layer and a 200-nm Au film. (6) Lift-off by immersion in a photoresist stripper (AZ 400T) overnight resulted in corresponding patterns of Au/Cr. This step defines the microelectrodes, traces and reference electrodes. (7) Repeating step 2 yielded the second layer of the polymer scaffold. (8) Repeating steps 3 and 4 formed a pattern of photoresist to define the 2D outlines of the polymer scaffold. (9) Thermal evaporation (Denton Vacuum Explorer 14) formed a 50-nm Cu film. (10) The lift-off process resulted in six defined Cu patterns that served as hard masks for plasma etching. (11) Plasma dry etching transferred this Cu pattern into the underlying polymer to form the scaffolds and to expose the microelectrodes. (12) Wet etching removed the remaining Cu, thereby completing the fabrication.
The mechanically guided process for a 3D framework involved the following sequence. (1) Immersion in acetone dissolved the poly(methyl methacrylate) sacrificial layer and released the 2D precursor from the supporting substrate. (2) A piece of polyvinyl alcohol tape allowed retrieval of the precursor and exposure of its backside for sputter deposition of 30-nm SiO2 through a shadow mask, to define the selective bonding sites. (3) Exposure to ultraviolet ozone for 5 min followed by baking at 75 °C for 15 min enabled strong chemical bonds to form between the bonding sites and the surface of a prestretched substrate of PDMS (SYLGARD 184 Silicone Elastomer, with 20:1 mixing ratio) upon physical contact. (4) Release of the prestretch caused the 2D precursor to buckle into a corresponding 3D geometry.
Mechanical analysis
Three-dimensional FEA quantified all aspects of the post-buckling responses and the maximum principal stress/strain distributions in the 3D structures. Four-node 2D shell elements (S4R) modelled the polymer scaffolds, and eight-node 3D solid elements (C3D8R) captured the elastomeric substrates. Refined meshes ensured computational accuracy and efficiency, conducted with commercial software (ABAQUS). Linear elastic constitutive models described the properties of the metal (Au) and polymer (PI), where the elastic modulus and Poisson’s ratio are EAu = 70 GPa and νAu = 0.44; EPI = 2.5 GPa and νPI = 0.35. Modelling of the prestretched silicone substrate relied on an isotropic hyperelastic structure using the Mooney–Rivlin law (EPDMS = 2 MPa and νPDMS = 0.49). The organoids were modelled as an incompressible solid material with an elastic modulus of ~1 kPa. Applying linear buckling analyses for the 2D pattern determined the critical buckling strain and the corresponding lowest buckling mode, which were then considered as initial geometric imperfections in the post-buckling simulations to achieve the final deformed configurations and stress/strain distributions. Notably, the geometric nonlinearity was accounted for in the post-buckling analyses of the 3D configuration.
Electrical characterization
A three-electrode configuration with a Pt wire as the counter electrode and Ag/AgCl as the reference electrode enabled electroplating of Pt black (Autolab PGSTAT128N, Metrohm AG, -0.1 V for 30 s) with a solution of 3 wt% of chloroplatinic acid and 0.1 wt% of lead acetate. A similar three-electrode configuration in phosphate-buffered saline (PBS) enabled electrochemical impedance spectroscopy and charge injection limit measurements on each Pt-coated microelectrode. Measurements of charge injection limits employed biphasic pulses (a cathodic pulse followed by anodic pulse) with a pulse duration of 200 µs. A 2-ms interval between the two pulses defined the access voltage (Va) as the voltage drop at the end of each pulse that originates from ionic conductivity of the electrolyte. The highest (lowest) voltage minus (plus) the access voltage yielded the most positive (negative) voltage, Emax (Emin), as plotted with an example case in Extended Data Fig. 3d. Gradually increasing the pulse current until either Emax = 0.9 V or Emin = −0.6 V defined the maximum charge injection without exceeding the water window. Long-term monitoring of the 3D structures immersed in 37 °C PBS used a commercial system (Intan RHS system, Intan Technologies) to track changes in electrochemical impedance and noise levels for each channel.
Generation of human cortical organoids
This study used human cortical organoids derived from WTC-11 (Coriell, GM25256), BJFF.6 (RRID: CVCL_VU02, Genome Engineering & Stem Cell Center (GESC), Washington University School of Medicine) and AN1.1 (Tech ID: T-019532, GESC, Washington University School of Medicine) hiPS cell lines. The genome integrity of all lines are confirmed by karyotyping (Supplementary Figs. 21, 23 and 24). The process began with culturing cells in mTeSR Plus medium (StemCell Technologies, 08620) plus 1% penicillin and streptomycin (Thermofisher, 15140122) in six-well plates coated with Matrigel (Sigma-Aldrich, CLS354277) for 5 days, followed by dissociation into single-cell suspensions using gentle cell dissociation reagent (StemCell Technologies, 100-0485) and centrifugation at 300g for 5 min. Dorsal Forebrain Organoid Kits (StemCell Technologies, 08620) formed the organoids following a previously published protocol25. In brief, hiPS cells formed uniform aggregates after transfer to Aggrewell 800 microwell culture plates (StemCell Technologies, 34815) at a density of 10,000 cells per microwell. Treatment of the aggregates involved exposure to dual SMAD inhibition for 6 days and with partial daily media changes. On day 7, transfer occurred to ultralow-attachment six-well plates (30–40 organoids per well), supplied with serum-free media containing fibroblast growth factor-2 (FGF-2) and epidermal growth factor (EGF) until day 25. They were then transferred to media containing brain-derived neurotrophic factor and neurotrophic factor 3 until day 43, after which the media was supplied without growth factors. A mixture of four small molecules—GSK2879552 (Selleck Chemicals, S7796), EPZ5676 (Selleck Chemicals, S7062), Bay K 8644 (Selleck Chemicals, S7924) and NMDA (Selleck Chemicals, S7072)—each at a concentration of 1 mM, was added to the media according to a previously published protocol43 to enhance neuron maturation into cortical organoids from day 15 until the day of experiments.
Generation of human spinal organoids
Generation of human spinal organoids (hSCOs) followed an established protocol that was similar to that of the cortical case37, but with differences in the supplements. On day 1, cells (HUES 3 line) (genome integrity confirmed by karyotyping; Supplementary Fig. 22) were cultured with mTeSR Plus Medium (StemCell Technologies). Dissociation of cells using Accutase (Fisher Scientific, 50-112-9055), followed by centrifugation and seeding into the AggreWell 800 (StemCell Technologies, 34815) plates at 10,000 cells per microwell, initiated the aggregation. After a centrifugation at 100g for 3 min, incubation of the plates occurred for 24 h at 37 °C with 5% CO2. From day 0 to day 6, organoid culture was in MTeSR plus medium supplemented with dorsomorphin (2.5 μM, Sigma-Aldrich, P5499) and SB-431542 (10 μM, DNSK, KI-12) with daily media changes, then replaced on day 7 with Neurobasal-A medium containing N-2 (Life Technologies, 17502048), B-27 (50×, minus vitamin A, Gibco, 12-587-001), CHIR (3 μM, MTocris, 4423409), retinoic acid (0.1 μM; Sigma-Aldrich, R2625) and EGF (20 ng ml−1; R&D Systems, 236-EG-200) as growth factors for neural induction. From day 19, the medium was enriched with BDNF (20 ng ml−1, R&D Systems, 248-BDB-250/CF), IGF-1 (10 ng ml−1, Peprotech, 100-11), DAPT (2.5 μM; Tocris, 2634/10), N-2 (Life Technologies, 17502048), L-ascorbic acid (AA, 200 nM; Sigma-Aldrich, A4403) and cAMP (50 nM; Sigma-Aldrich, D0627) to promote further neural maturation. The feeding schedule was every other day until day 41 and subsequently every 4–5 days after day 43 without growth factors.
Immunofluorescence and imaging
Fixation of the organoids occurred through overnight immersion in 4% paraformaldehyde (Thermofisher, J61899.AK) at 4 °C, followed by rinsing three times with Dulbecco’s PBS (DPBS; Thermofisher, 14190136) and storage in PBS (Thermofisher, 10-010-031) at 4 °C. Organoids were transferred to 30% sucrose/PBS solution for 24–48 h before sectioning and placing in cryomolds filled with OCT (Leica Biosystems, 3801480). Cryostat sectioning of samples occurred sequentially, at 14 µm thickness, using a Microm HM525 Cryostat. The sections were then placed on microscope glass slides and stored at −20 °C for subsequent immunostaining.
Immunofluorescence began with DPBS washing of the slides before incubation for 1 h at room temperature in blocking solution (1% BSA (Thermofisher, 15260-037) and 0.1% Triton-X (Sigma-Aldrich, 9036-19-5 diluted in PBS), and then followed by incubation with primary antibodies diluted in the blocking solution overnight at 4 °C. Primary antibodies used in this study included mouse anti-S100B (Sigma-Aldrich, S2532, 1:100), guinea pig anti-MAP2 (SYSY, 188 004, 1:100), rabbit anti-SOX2 (Sigma-Aldrich, AB5603, 1:100), PSD95 (Abcam, ab18258, 5 μg ml−1) and Synaptotagmin-1 (Developmental Studies Hybridoma Bank (DSHB), AB_2199314, 1:100). Subsequently, after washing of the primary antibodies with three 5-min cycles of DPBS, incubation with secondary antibodies occurred at room temperature and protected from light. The secondary antibodies used in this study were Alexa Fluor 488 goat anti-guinea pig (Jackson, 106-545-003; 1:500), Alexa Fluor 647 goat anti-mouse (Jackson, 115-605-003; 1:500), Alexa Fluor 568 goat anti-rabbit (Thermo Fisher Scientific, A-11011), Alexa Fluor 647 goat anti-rabbit (Jackson, 111-605-144; 1:200) and Alexa Fluor 488 goat anti-mouse (Jackson, 115-545-205; 1:200). Adding 4′,6-diamidino-2-phenylindole (Sigma-Aldrich, 10236276001, 1:10,000) for 5 min yielded labelled nuclei. Air drying and placing the slices on a cover slip with mounting medium (Vector Labs, H-1700-10) prepared them for fluorescent imaging with a FV3000 Olympus confocal microscope. Laser settings and exposure times were consistent for each channel throughout all samples.
Integration of organoids with 3D interfaces
Insertion typically happened after day 60 of differentiation for hCOs and day 40 for hSOs. Sterilization of the 3D structure occurred through autoclaving, followed by rinsing with PBS immediately before integration. Placing the interface on a plastic ridge and applying gentle pressure opened the 3D interfaces, allowing insertion of organoids using wide-bore pipettes. Integrated organoids were kept at 37 °C with 5% CO2, with media changes every 2–3 days to support the grow-in-place process before electrophysiological recordings.
Simultaneous electrophysiological and fluorescence recordings and data analysis
Simultaneous recordings of electrophysiology and calcium fluorescence occurred on a confocal microscope. Application of a semi-permanent fluorescent calcium reporter, GCaMP6f (Addgene, 100837-AAV1), induced GFP fluorescence to allow measurements of calcium dynamics by live imaging. Organoids were transduced with adeno-associated virus particles at a 5 × 105 multiplicity of infection and incubated for 72 h. After removing the viral particles, GCaMP6f expressed in 3–5 days. A commercial system (Intan RHD, Intan Technologies) with multiple headstages connected in parallel enabled high-channel number recordings. The Intan RHX software recorded and stored real-time electrophysiological potentials at a sampling rate of 20 kHz, with a bandwidth of 1–9,000 Hz and a notch filter at 60 Hz. Each recording session concluded in less than 5 min to minimize the effects of cooling on organoid activities during the measurements.
The software package ImageJ with a customized Java code served as the basis for evaluating calcium fluorescence images and videos. Each frame was discretized into 100 × 100 units to quantify brightness at each location. Normalizing these brightness data with the average brightness of a baseline image of the same session revealed the spatiotemporal characteristics of organoid activity. Locally weighted scatterplot smoothing with frac = 0.5 in range [0:1] to reduce effects of environmental noise and random fluctuations preceded filtering with a Savitzky–Golay filter with window length = 9, polyorder = 8. Computations of the differences of the normalized brightness between the unit matrix of two imaging frames formed a 2D heatmap map to indicate the 2D distribution of activities.
Data analysis of neural activities used a customized Python code (Extended Data Fig. 5) that followed a multistep procedure. Conversion of the raw data into .npy format preceded processing with a third-order Butterworth bandpass filter between 250 Hz and 3,000 Hz and application of a peak detection algorithm. These latter processes involved calculating the r.m.s. noise of the recording channels and detecting peaks with amplitudes larger than five times of the r.m.s. Defining a 5-ms minimum interval between any two labelled peaks prevented duplication in peak counting. Segmentation of each detected peak by a 3-ms window (60 data points of neural potential for 20 kHz sampling rate) formed a high-dimensional vector. Use of an unsupervised clustering algorithm two-component uniform manifold approximation and projection reduced the dimension of the 60-dimensional vector to 2. Subsequently, density-based spatial clustering of applications with noise (DBSCAN) categorized these 2D vectors into clusters of neural spikes.
Calculations of the average potential and standard deviations (s.d.) of all the neural spikes within each cluster defined the overall characteristic waveforms. Such waveforms collected throughout all recording sessions formed a comprehensive dataset, including both neural activities and artefacts. Systematic recordings with control groups of empty interfaces and interfaces integrated with fixed organoids marked the waveforms associated with mechanical and optical noise in the environment. Excluding spike clusters with these artefact waveforms from the recording data ensured artefact-free raster plots of neural activity.
Additional calculations of the basic characteristics included firing rates and spike amplitudes of each individual channel. Associating the data of each channel with the designed 3D coordinates of each corresponding microelectrode resulted in a 3D spatial cloud data distributed across the surface of a sphere that mimics the geometry of the organoid. A radial basis function (scipy.interpolate.Rbf) then interpolated this cloud data onto a mesh on the spherical surface with points uniformly distributed in spherical coordinates (600 grids in θ and 300 grids in φ). Three-dimensional cross-correlation analysis used pairwise cross-correlations among all recorded neural spike clusters. For the spike times of each pair of recorded clusters, cross-correlograms (CCGs) with a specified range of −200 to 200 ms and a bin width of 0.5 ms followed by boxcar-filtering of 3 ms defined histograms of the amplitudes of delay correlations as a function of time delay. CCG of shuffled spike trains of each cluster in the pair that filtered with the same condition defined the baseline of the correlation. Significant correlations corresponded to cases where two consecutive binned data points of the CCG of one cluster pair exceed the means plus five times of the s.d. of the baseline. In such scenarios, the correlation strength of the cluster pair represents the peak amplitudes of CCG normalized by the baseline mean value, and the lag represents the bin location of the peak.
In vivo recording in mice
Mice were anesthetized using vaporized isoflurane (3% for induction and 1.5–2% for maintenance) and positioned in a stereotaxic apparatus designed for small animals (David Kopf Instruments). A craniotomy (~1.0 mm diameter) was made over the somatosensory cortex at anterior–posterior coordinates 0.0 to –1.5 mm and medial–lateral coordinates ±2 mm. For reference, a stainless-steel screw (18-8 button head Torx screw, #90910A600, McMaster-Carr) with a silver wire lead (0.375 mm diameter, #64-1320, Warner Instruments) was implanted in the contralateral occipital bone.
A 64-channel microelectrode array (4 shanks spaced 250 µm apart, 16 electrodes per shank at 50-µm vertical intervals; model A4x16_5mm_250-177, NeuroNexus Technologies) was then inserted through the craniotomy, targeting a depth of 750 µm for the deepest electrode on each shank. The array was interfaced with an analogue-to-digital amplifier (#C3325, Intan Technologies) mounted to the stereotaxic frame.
Pharmacological tests, optogenetic stimulations, neural circuit manipulations and disease phenotypes
Evaluations of pharmacological responses included exposure to 4-AP (Sigma-Aldrich, 275875-1G) at a concentration of 100 μM, which evoked immediate synchronous activity in spinal organoids. Lidocaine, introduced at a concentration of 2 mM (Sigma-Aldrich, L7757-25G) and then washed out after 20 min, inhibited calcium dynamics and disrupted connectivity. TTX at a concentration of 100 μM produced notable inhibitory effects, reducing overall activity and decreasing the number of active channels. Optogenetic transfection used Hues 3 ESC-derived spinal organoids at least 40 days old and approximately 1.5 mm diameter in size. Transfection was performed at a multiplicity of infection of 20, assuming 30,000 live cells, using a lentivirus from VectorBuilder (Vector ID: VB900122-3326aya) as the basis for studies of optogenetic activation. The EF1A promoter of the lentivirus delivered channelrhodopsin-2 to the cellular genome, resulting in EGFP expression in transfected cells. After 24 h of incubation, fully changing the cell culture media removed the lentivirus. Feeding of the virally transfected organoids occurred every Monday, Wednesday and Friday. Integration of transfected organoids with interfaces occurred 4 days after transfection, with optogenetic activity induced and recorded 10 days after transfection. For neural circuit manipulation, application of two clinical doses (2 U) of BTX (OnabotulinumtoxinA, AbbVie) diluted in 2 ml biomedia to neural organoids formed a concentration of 1 U ml−1. Washing removed BTX after incubation at 37 °C for 48 h. Recordings were made before and on each day after exposure to BTX. Disease modelling used glutamate (Thermofisher, J63424.14) at a concentration of 25 μM, followed by 2 h incubation and washing. Monitoring the neural activity captured the responses in all the tests above before the dosing, and at specific time after dosing (1 h for 4-AP, lidocaine and TTX, and every day for BTX and glutamate).
Statistical analysis
Statistical comparisons between experimental datasets were performed using a two-sided t-test. Statistical significance corresponded to P values of less than 0.05. Results shown in the figures use symbols to indicate statistical significance: NS (not significant), **P < 0.01 or *P < 0.001. These analyses used OriginLab.
Ethics statement
hiPS cell culture activities at the University of Illinois Chicago were conducted in accordance with Protocol 22-020, which was approved by the university’s Institutional Biosafety Committee. The WTC-11 line was derived from a donor who gave voluntary informed consent to a qualified professional. The cells used in this study were obtained from The Coriell Institute.
The human embryonic stem cell lines used in this study are National Institutes of Health-approved, and the human embryonic stem and hiPS cell research was conducted in compliance with the Shirley Ryan AbilityLab as well as Northwestern University’s Committee on Human Stem Cell Research registration requirements, which oversee the provenance, use and safekeeping of human pluripotent stem cells.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data are available within the Article and its Supplementary Information. Source data are provided with this paper.
Code availability
The custom code used in this study is available via Zenodo at https://doi.org/10.5281/zenodo.17794914 (ref. 44) and https://doi.org/10.5281/zenodo.17687597 (ref. 45).
References
Quadrato, G. et al. Cell diversity and network dynamics in photosensitive human brain organoids. Nature 545, 48–53 (2017).
Velasco, S. et al. Individual brain organoids reproducibly form cell diversity of the human cerebral cortex. Nature 570, 523–527 (2019).
Pașca, S. P. Constructing human neural circuits in living systems by transplantation. Cell 187, 8–13 (2024).
Kelley, K. W. & Pașca, S. P. Human brain organogenesis: toward a cellular understanding of development and disease. Cell 185, 42–61 (2022).
Revah, O. et al. Maturation and circuit integration of transplanted human cortical organoids. Nature 610, 319–326 (2022).
Giandomenico, S. L. et al. Cerebral organoids at the air–liquid interface generate diverse nerve tracts with functional output. Nat. Neurosci. 22, 669–679 (2019).
Samarasinghe, R. A. et al. Identification of neural oscillations and epileptiform changes in human brain organoids. Nat. Neurosci. 24, 1488–1500 (2021).
Bowles, K. R. et al. ELAVL4, splicing, and glutamatergic dysfunction precede neuron loss in MAPT mutation cerebral organoids. Cell 184, 4547–4563 (2021).
Martins, J.-M. F. et al. Self-organizing 3D human trunk neuromuscular organoids. Cell Stem Cell 26, 172–186 (2020).
Vlachogiannis, G. et al. Patient-derived organoids model treatment response of metastatic gastrointestinal cancers. Science 359, 920–926 (2018).
Paulsen, B. et al. Autism genes converge on asynchronous development of shared neuron classes. Nature 602, 268–273 (2022).
Cai, H. et al. Brain organoid reservoir computing for artificial intelligence. Nat. Electron. 6, 1032–1039 (2023).
Osaki, T. et al. Complex activity and short-term plasticity of human cerebral organoids reciprocally connected with axons. Nat. Commun. 15, 2945 (2024).
Beaubois, R. et al. BiœmuS: a new tool for neurological disorders studies through real-time emulation and hybridization using biomimetic Spiking Neural Network. Nat. Commun. 15, 5142 (2024).
Yang, X. et al. Kirigami electronics for long-term electrophysiological recording of human neural organoids and assembloids. Nat. Biotechnol. 42, 1836–1843 (2024).
Le Floch, P. et al. Stretchable mesh nanoelectronics for 3D single-cell chronic electrophysiology from developing brain organoids. Adv. Mater. 34, 2106829 (2022).
Li, T. L. et al. Stretchable mesh microelectronics for the biointegration and stimulation of human neural organoids. Biomaterials 290, 121825 (2022).
Wu, Y. et al. Three-dimensional liquid metal-based neuro-interfaces for human hippocampal organoids. Nat. Commun. 15, 4047 (2024).
Huang, Q. et al. Shell microelectrode arrays (MEAs) for brain organoids. Sci. Adv. 8, eabq5031 (2022).
Martinelli, E. et al. The e-Flower: a hydrogel-actuated 3D MEA for brain spheroid electrophysiology. Sci. Adv. 10, eadp8054 (2024).
Park, Y. et al. Three-dimensional, multifunctional neural interfaces for cortical spheroids and engineered assembloids. Sci. Adv. 7, eabf9153 (2021).
Cheng, X. et al. Programming 3D curved mesosurfaces using microlattice designs. Science 379, 1225–1232 (2023).
Yan, Z. et al. Deterministic assembly of 3D mesostructures in advanced materials via compressive buckling: a short review of recent progress. Extreme Mech. Lett. 11, 96–104 (2017).
Zhang, Y. et al. Printing, folding and assembly methods for forming 3D mesostructures in advanced materials. Nat. Rev. Mater. 2, 1–17 (2017).
Paşca, A. M. et al. Functional cortical neurons and astrocytes from human pluripotent stem cells in 3D culture. Nat. Methods 12, 671–678 (2015).
Sharf, T. et al. Functional neuronal circuitry and oscillatory dynamics in human brain organoids. Nat. Commun. 13, 4403 (2022).
Trujillo, C. A. et al. Complex oscillatory waves emerging from cortical organoids model early human brain network development. Cell Stem Cell 25, 558–569 (2019).
Segev, R., Goodhouse, J., Puchalla, J. & Berry, M. J. Recording spikes from a large fraction of the ganglion cells in a retinal patch. Nat. Neurosci. 7, 1155–1162 (2004).
Henze, D. A. et al. Intracellular features predicted by extracellular recordings in the hippocampus in vivo. J. Neurophysiol. 84, 390–400 (2000).
Yang, X. et al. Bioinspired neuron-like electronics. Nat. Mater. 18, 510–517 (2019).
Antón-Bolaños, N. et al. Brain Chimeroids reveal individual susceptibility to neurotoxic triggers. Nature 631, 142–149 (2024).
Cartiglia, M. et al. A 4096 channel event-based multielectrode array with asynchronous outputs compatible with neuromorphic processors. Nat. Commun. 15, 7163 (2024).
Kheiri, S. et al. Microfluidic platform for generating and releasing patient-derived cancer organoids with diverse shapes: insight into shape-dependent tumor growth. Adv. Mater. 36, 2410547 (2024).
Nikolaev, M. et al. Homeostatic mini-intestines through scaffold-guided organoid morphogenesis. Nature 585, 574–578 (2020).
Lancaster, M. A. et al. Guided self-organization and cortical plate formation in human brain organoids. Nat. Biotechnol. 35, 659–666 (2017).
Bagley, J. A., Reumann, D., Bian, S., Lévi-Strauss, J. & Knoblich, J. A. Fused cerebral organoids model interactions between brain regions. Nat. Methods 14, 743–751 (2017).
Andersen, J. et al. Generation of functional human 3D cortico-motor assembloids. Cell 183, 1913–1929 (2020).
Luvisetto, S. Botulinum neurotoxins in central nervous system: an overview from animal models to human therapy. Toxins 13, 751 (2021).
Steinmetz, N. A. et al. Neuropixels 2.0: a miniaturized high-density probe for stable, long-term brain recordings. Science 372, eabf4588 (2021).
Liu, Y. et al. A high-density 1,024-channel probe for brain-wide recordings in non-human primates. Nat. Neurosci. 27, 1620–1631 (2024).
Sun, A. X. et al. Potassium channel dysfunction in human neuronal models of Angelman syndrome. Science 366, 1486–1492 (2019).
Teleanu, R. I. et al. Neurotransmitters—key factors in neurological and neurodegenerative disorders of the central nervous system. Int. J. Mol. Sci. 23, 5954 (2022).
Hergenreder, E. et al. Combined small-molecule treatment accelerates maturation of human pluripotent stem cell-derived neurons. Nat. Biotechnol. 42, 1515–1525 (2024).
liu, N. Shape-conformal porous frameworks for full coverage of neural organoids and high-resolution electrophysiology. Zenodo https://doi.org/10.5281/zenodo.17794914 (2025).
Jin, T. Shape-conformal porous frameworks for full coverage of neural organoids and high-resolution electrophysiology. Zenodo https://doi.org/10.5281/zenodo.17687597 (2025).
Soscia, D. A. et al. A flexible 3-dimensional microelectrode array for in vitro brain models. Lab Chip 20, 901–911 (2020).
Acknowledgements
We thank members of the Rogers Lab, Finan Lab, Franz Lab and Huang Lab for helpful discussions and technical support. We thank T. Osaki for helpful discussions on data analysis. Microfabrication in this work was performed at the NUFAB facility of Northwestern University’s NUANCE Center, with support from the SHyNE Resource (NSF ECCS-2025633), the IIN, and Northwestern’s MRSEC programme (NSF DMR-2308691). Micro-CT characterizations were performed at the Imaging Technology Group at Beckman Institute of Advanced Science and Technology, University of Illinois Urbana-Champaign and the Northwestern University Center for Advanced Molecular Imaging (RRID: SCR_021192) generously supported by NCI CCSG P30 CA060553 awarded to the Robert H Lurie Comprehensive Cancer Center. This work was supported by the Querrey-Simpson Institute for Bioelectronics (N.L., J.Liu, Z.Z., J.Li, Y.W., Q.L., Z.X., L.Z., M.W., Ya.Z., H.L. and J.A.R.), the National Institutes of Health under award number R011NS113935 (N.L., S.S., Y.S., S.K., C.K.F. and J.D.F.), the National Natural Science Foundation of China under grant number 12225206 (T.J., X.C. and Yi.Z.), the New Cornerstone Science Foundation through the XPLORER PRIZE (T.J., X.C. and Yi.Z.), the Belle Carnell Regenerative Neurorehabilitation Fund (I.C., M.J.Q. and C.K.F.), the National Science Foundation Graduate Research Fellowship Program (A.V.), the NIH/NIBIB Center for Label-free Imaging and Multiscale Biophotonics (CLIMB), P41EB031772 (S.A.B. and J.A.R.), and the Haythornthwaite Foundation Research Initiation Grant (H.L.).
Author information
Authors and Affiliations
Contributions
These authors contributed equally: Z.Z. and J.Li. J.A.R., J.D.F, C.K.F., Yi.Z. and N.L. conceived and designed the project. S.S. and I.C. maintained and differentiated hiPS cells into hCOs. I.C., A.V., M.J.Q., Y.S. and S.K. maintained and differentiated hiPS and hES cells into hSOs. T.J., C.X. and Yi.Z. performed the inverse design. T.J. performed mechanical analysis. T.J., N.L. and H.L. designed the devices. N.L., Z.Z., J.Li, Q.L., Z.X, L.Z. and W.-Y.M. performed microfabrication and 3D assembly of devices. N.L., J.Liu, Z.Z., J.Li and Ya.Z. performed the electrical characterization of devices. Y.W. designed circuit boards. N.L., S.S., Z.Z., J.Li, Q.L. and M.W. performed electrophysiology and imaging characterization of neural activities. N.L., Z.Z. and J.Li analysed the electrophysiology data. H.Z., L.Z. and Y.H. performed diffusion simulations. A.V. and S.B. performed genomic analysis. S.L. and S.A.B. performed two-photon imaging. N.L., J.Liu, Yi.Z., C.K.F., J.D.F. and J.A.R. wrote the manuscript with inputs from S.S., T.J. and H.Z. Yi.Z., C.K.F., J.D.F. and J.A.R. supervised the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Biomedical Engineering thanks Stefano Vassanelli and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Inverse design strategy for 3D neural interface.
Discretization of the targeted organoid geometries defines the 2D outline of each beam, as well as the initial population that goes through an optimization process with AGA algorithm to obtain the distribution of normalized stiffness. Microlattice design transfers this stiff distribution into the predefined 2D outlines, forming the scaffold design of 3D neural interface.
Extended Data Fig. 2 Mechanical analysis of 3D neural interface.
a, FEA simulation of the displacement through the 3D assembly processes. b, FEA simulation of the stress distribution in the polymer layer of assembled interface. c, FEA simulation of the strain distribution in the polymer layer of assembled interface. d, FEA simulation of the stress distribution in the Au traces of assembled interface. e, FEA simulation of the strain distribution in the Au traces of the assembled interface. f, Introducing chamfer design eliminates the stress concentration around the micropores. g, FEA modeling of the highest strain within the polymer scaffold with different targeted sizes of organoids and with different thicknesses. A thin device thickness ensures the compatibility to interface with small organoids below 500 µm in diameter.
Extended Data Fig. 3 Electrical characterization of microelectrodes.
a, Electro impedance spectrum (EIS) of 96 individual microelectrodes of one interface. b, Histogram of the impedance at 1 kHz frequency of the 96 microelectrodes in (a), showing an average value around 10 kΩ. c, The impedance of each microelectrodes (at 10 kHz frequency) showing in its original location. d, Charge injection limit test of one example microelectrode showing increasing voltage with increasing current injection. The current when associated voltage approaches the water window defines the charge injection limit. e, Histogram of the charge injection limit of 96 microelectrodes of one interface, showing an average value around 200 µA. f, The change injection limit of each microelectrodes showing in its original location. g and h, Stability test of all microelectrodes of one interface in 37 °C PBS solution for 90 days shows stable impedance (g) and rms noise level (h) of the microelectrodes. Data are presented as mean values ± SD of 96 individual microelectrodes.
Extended Data Fig. 4 Interface integration with hCO.
a, Bright field images of generated hCOs. Images are representative of 6 organoids. b, Immunostaining images of day 30 hCO. c, Immunostaining images of day 120 hCO. Immunostaining images are representative images of 3 organoids. d, Bright field images monitoring the grow-in-place process of one integrated hCO. This experiment was repeated with 6 devices. e, Electrophysiological recording of the same hCO in (d) showing increasing number of active channels following the filling of gaps along the interface from day 64 to day 79. f, Immunostaining images of control group hCO and hCO grown-in-place for two weeks show negligible interface effect on cell distributions. Images are representative of 3 sets of organoids.
Extended Data Fig. 5 Data analysis of the electrophysiological signals.
a, Overview of 30 example channels of one recording after filtering. Scale bars: 100 ms (horizontal) and 50 μV (vertical). b, 5 times of the rms noise defines the thresholds of the single processing, exceeding which defines a peak. Scale bars: 5 ms (horizontal) and 20 μV (vertical). c, Overview of all the detected peaks of one example channel. Scale bars: 0.5 ms (horizontal) and 20 μV (vertical). d, UMAP visualization of clustered peaks. e, The characteristic waveforms of each cluster in (d), showing different scenarios of neural signals, optical artifacts (from microscope Laser source), and mechanical artifacts. Scale bars: 0.5 ms (horizontal) and 20 μV (vertical). f, Labeling the detected peaks with their cluster information allows to remove artifacts in processed recording data. Scale bars: 100 ms (horizontal) and 20 μV (vertical) for the overview plot, and 10 ms (horizontal) and 20 μV (vertical) for the superimposed zoom-in plot. g, Raster plots of the neural spikes from 240 channels of one recording (hCO, day 88) before and after artifact removing.
Extended Data Fig. 6 hCO with different scenarios of detected network-level activity.
Raster plots of neural spikes, spike amplitude of example channels (with error bars define the standard deviation all spikes detected by the microelectrode), and 3D reconstructed surface mapping of average spike amplitude and firing rates of a, hCO (day 90) with asynchronous activities. b, hCO (day 89) with synchronous activities. And c, hCO (day 120) with localized and partial activities. The data are presented as mean ± s.e.m. for spike amplitude.
Extended Data Fig. 7 Electro stimulation with 3D neural interface.
a, Schematic illustration of simultaneous calcium fluorescence imaging and electro stimulation. b, Details of each stimulation train with biphasic pulses. c, Calcium fluorescence image of integrated hCO for stimulation. d, Average fluorescence brightness as a function of stimulating current. e–h, heatmap of brightness change under stimulation and brightness plot with time during 5 stimulations with 60 s interval, with current of 1 µA (e), 3 µA (f), 5 µA (g), and 20 µA (h), revealing a current threshold for neural response around 3 µA.
Extended Data Fig. 8 Effect of 3D neural interface on diffusion.
a, Calculated contour of the steady-state concentration distribution of O2 of one integrated neural organoid. b, Comparation of the O2 concentration along the depth between control organoid and integrated organoid. c, Varying the reaction rate of O2 across a broad range of magnitudes reveals a limited difference between the depth integrated organoid to achieve the same concentration of control organoid at the depth of 200 µm. d, Calculated contour of the steady-state concentration distribution of CO2 of one integrated neural organoid. e, Comparation of the CO2 concentration along the depth between control organoid and integrated organoid. f, Varying the reaction rate of CO2 across a broad range of magnitudes reveals a limited difference between the depth for integrated organoid to achieve the same concentration of control organoid at the depth of 200 µm. g, Calculated contour of the steady-state concentration distribution of glucose of one integrated neural organoid. h, Comparation of the glucose concentration along the depth between control organoid and integrated organoid. i, Varying the reaction rate of glucose across a broad range of magnitudes reveals a limited difference between the depth for integrated organoid to achieve the same concentration of control organoid at the depth of 200 µm.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–28, Notes 1–12 and References 1–14.
Supplementary Video 1 (download MOV )
3D assembly of organoid interfaces.
Supplementary Video 2 (download MOV )
Micro-CT imaging of an assembled organoid interface.
Supplementary Video 3 (download MOV )
Simultaneous recording of organoid activity with electrophysiology and fluorescence.
Source data
Source Data Figs. 2–5 (download XLSX )
Statistical source data of Figs. 2–5.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Liu, N., Shiravi, S., Jin, T. et al. Shape-conformal porous frameworks for full coverage of neural organoids and high-resolution electrophysiology. Nat. Biomed. Eng (2026). https://doi.org/10.1038/s41551-026-01620-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41551-026-01620-y